Abstract
This study investigates the sensitivity and specificity of predicting epileptic seizures from intracranial electroencephalography (iEEG). A monitoring system is studied to generate an alarm upon detecting a precursor of an epileptic seizure. The iEEG traces of ten patients suffering from medically intractable epilepsy were used to build a prediction model. From the iEEG recording of each patient, power spectral densities were calculated and classified using support vector machines. The prediction results varied across patients. For seven patients, seizures were predicted with 100% sensitivity without any false alarms. One patient showed good sensitivity but lower specificity, and the other two patients showed lower sensitivity and specificity. Predictive analytics based on the spectral feature of iEEG performs well for some patients but not all. This result highlights the need for patient-specific prediction models and algorithms.
Keywords: Predictive analytics, Seizure prediction, EEG, Cross-validation, Personalized healthcare
Introduction
The focus of healthcare service is shifting from cure to prevention. Key enabling factors include sensor technology and data analytics. Various sensors are embedded into mobile devices, providing rich physiological measurements. Data analytics plays a key role in turning the abundant data into actual improvements in the quality of life. For example, a personalized healthcare system can be set up to monitor patients with a chronic disease [1] or acute symptoms such as predicting heart diseases [2].
For this purpose, predictive analytics has gained significant attention. Predictive analytics is a statistical method for learning patterns in the obtained data and predicting outcomes for new inputs. This tool supports data-driven decision-making in numerous fields such as manufacturing, marketing, and governments. This paper further explores a new application domain: predictive analytics for brain monitoring systems. The focus is on the feasibility of predicting abnormal brain conditions such as epileptic seizures.
The remainder of this paper is organized as follows. Section 2 outlines the brain monitoring systems and their application to seizure prediction. The experimental method and its results are described in Sects. 3 and 4, respectively. In Sect. 5, some concluding remarks and discuss future works are provided.
Predictive analytics for epileptic seizures
Figure 1 shows a schematic diagram of a brain monitoring system. Brain signals are measured by a sensor. Using the measured brain signals and a prediction model, the processor makes a prediction regarding the future state of the brain and produces an alarm when an abnormality is expected to occur.
Fig. 1.
A schematic diagram of a brain monitoring system
The electroencephalogram (EEG) has been the most widely used method for brain monitoring. The main advantages of an EEG over other modalities such as a CT or MRI are a low cost and high temporal resolution. However, the downside of an EEG is a low signal-to-noise ratio and poor spatial resolution. Therefore, the standard EEG has limitations in monitoring deeper areas of the brain (e.g., the temporal lobe). To overcome this limitation, EEG sensors are implanted into the brain to monitor deeper areas, which is called an intracranial EEG (iEEG).
Epileptic seizures are a commonly observed brain disorder, and epilepsy is the name of such a disorder characterized by an enduring occurrence of epileptic seizures [3]. About one-percent of the world’s population suffers from epilepsy [3]. The quality of life of an epileptic would be greatly improved if precursors to their seizures can be proactively detected and appropriate interventions applied.
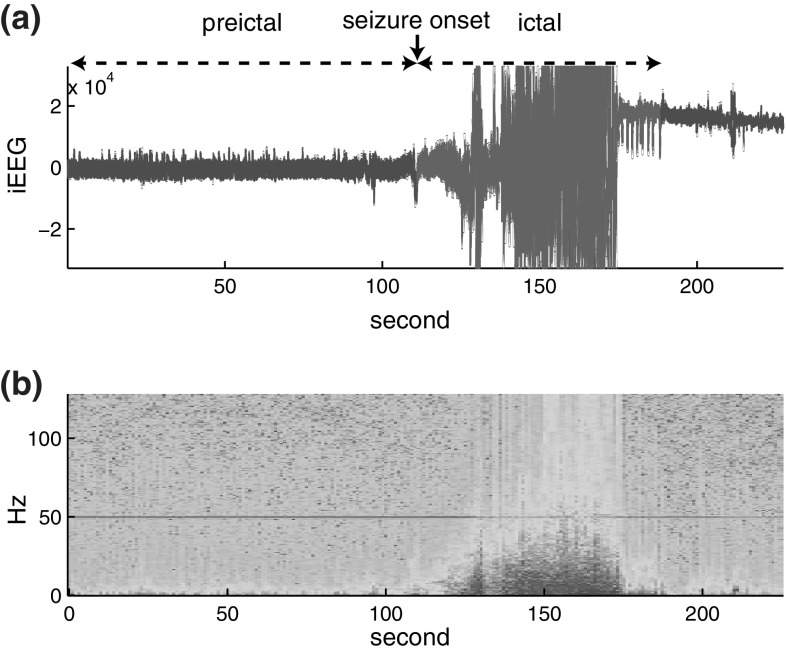
Figure 2 shows a typical intracranial EEG recording and its spectrogram of an epileptic seizure. In Fig. 2(a), the seizure response is shown in red, the duration of which is called the ictal period. During this ictal period, the spectrogram in Fig. 2(b) shows a significant increase in high frequency components. The duration of several minutes preceding the seizure onset is called the preictal period, as shown in Fig. 2(a).
Fig. 2.
Examples of intracranial EEG (a) and its spectrogram (b) that contains an epileptic seizure
A seizure predictor works as follows. Temporal or spectral patterns of the preictal period are learned and stored in the prediction model, which classifies EEG inputs into two classes: normal or preictal. When the level of the predicted preictal state is above a certain threshold, an alarm is raised.
Even after decades of hard work, it remains elusive whether a seizure prediction from an EEG is even possible [4–6]. Early approaches made use of autoregressive models to identify unstable states of the brain [4]. Later, notions of stability were adopted from theories on dynamic systems to quantify the brain states. Such measures include the largest Lyapunov exponent [7], the dynamical similarity index [8], and the phase coherence [9]. Unfortunately, all of these studies are limited to retrospectively detecting the presence or absence of a seizure. Iasemidis et al. [10] first attempted to predict epileptic activities in a prospective manner. Specifically, they use a Lyapunov exponent to predict a seizure from EEG recordings, and achieved a sensitivity of 84% and false alarm rate of 0.12 per hour. Unfortunately, this work and the following studies failed to meet requirements for clinical applications [4]. Some authors have even argued that the detection or prediction techniques developed to date are no better than a random guess [5, 6].
This controversy over the predictability of seizures is due to the poor signal-to-noise ratio of EEG recordings and the inherent uncertainty of brain signals. Whereas the former has been addressed by iEEGs, the need remains for the latter. Thus, this study focuses on the accuracy of predicting an epileptic seizure from an iEEG.
Another contribution of this paper is to rigorously analyze the sensitivity and specificity of seizure prediction. In this study, prediction accuracy was measured by cross-validations where training and data samples were separated in such a way that all the samples related to one seizure event are used for test and never used for training. This forces the monitoring system to predict truly unseen seizures and allows us to accurately quantify predictability of seizure events.
Experimental method
Brain signal dataset
For the experiments, iEEG recordings of the patients diagnosed with temporal lobe epilepsy from Freiburg EEG Database [11] were used. For each patient, six iEEG channels were available, and three channels among them were infocal. The sampling frequency was 256 samples per second. Patient information, the electrode type, and the number of seizures are summarized in Table 1.
Table 1.
Summary of EEG dataset for experiments
| Patient ID | Age | Sex | Origin | Electrodes | Number of seizures |
|---|---|---|---|---|---|
| 2 | 38 | M | Temporal | Depth | 3 |
| 4 | 26 | F | Temporal | Depth, grid, strip | 5 |
| 6 | 31 | F | Temporo/ocipital | Depth, grid, strip | 3 |
| 7 | 42 | F | Temporal | Depth | 3 |
| 10 | 47 | M | Temporal | Depth | 5 |
| 12 | 42 | F | Temporal | Depth, grid, strip | 4 |
| 13 | 22 | F | Temporo/occipital | Depth, strip | 2 |
| 14 | 41 | F | Fronto/temporal | Depth, strip | 4 |
| 15 | 31 | M | Temporal | Depth, strip | 4 |
| 16 | 50 | F | Temporal | Depth, strip | 5 |
The dataset comprises blocks of about one-hour iEEG recordings. Ictal and interictal blocks contain recordings with and without a seizure, respectively. In each ictal block, five minutes of preictal durations were used for the positive samples (). All interictal blocks were used for negative samples ().
From the samples chosen above, the spectral features of an EEG were extracted as follows. The power spectral density (PSD) for each channel was calculated using the short-time Fourier transform with 10-s Hamming windows (50% overlap). The resulting PSD had a frequency range of 0 to 128 Hz, which was averaged within widely used frequency bands [12, 13] as described in Table 2. For slow and fast gamma bands, frequency ranges of 47 to 53 Hz and 97 to 103 Hz were excluded to remove power line noises (harmonics of 50 Hz). The average PSD in eight bands was normalized by the sum of all bands in each channel. Therefore, a 48-dimensional spectral feature was produced for 6-channel iEEG signals every 5 s.
Table 2.
Frequency bands used for experiments
| Band | Frequency (Hz) |
|---|---|
| Delta | 0.5–4 |
| Theta | 4–8 |
| Alpha | 8–13 |
| Beta | 13–30 |
| Slow gamma | 30–47 and 53–75 |
| Fast gamma | 75–97 and 103–128 |
Leave-one-seizure-out cross-validation
Using the obtained spectral features, a “leave-one-seizure-out” cross-validation was performed for each patient using the linear support vector machines (SVMs) [14].
One cross-validation involves partitioning a given iEEG dataset into training and test datasets, each of which contains positive (preictal) and negative (interictal) samples. In previous studies, training and test datasets were manually selected without clearly defined criterion [13] or randomly partitioned in a manner by which some samples related to the same seizure event may be included for both training and test datasets [12], which may result in over- or under-estimation of seizure predictability. In contrast, we insured that samples related to a seizure were included into either of training or test datasets, but not into both. Specifically, we first partitioned the iEEG dataset into blocks according to seizure events so that each block either contains a seizure event (positive block) or was segregated any seizure by at least two hours (negative block). Thus, the number of positive blocks was the same as the number of seizures, but we had far more negative blocks. A positive block (corresponding to a single seizure event) and randomly selected negative blocks comprised a test dataset, and the remaining blocks were used for training an SVM. Such cross-validations were repeated ten times for one seizure left out for each test and different random partitions for training datasets.
The parameter settings for training a linear SVM were as follows. During training, positive samples were weighted higher to counteract the lack of samples, as proposed in [12]. Specifically, the weight for positive samples was set to be inversely proportional to the ratio of positive to negative samples in the training dataset. Another SVM parameter set by the user is the cost of a miss-classification, denoted as C. Increasing C places a higher penalty for a miss-classification, and tends to improve the accuracy but at the potential risk of an over fitting. Experiments were conducted for a wide range of C, from to on a log scale.
For each test sample, the trained SVM produces an estimate or , corresponding to a preictal (seizure predicted) or interictal (free of seizure) state, respectively.
Quantification of prediction accuracy
The predictability for each patient is quantified in terms of sensitivity and specificity. Estimate is correct when . This can be further divided into true positive () and true negative (). An incorrect estimate can occur in two ways: a miss-classification () and a false alarm (). The probabilities of a miss () and false alarm () are defined as follows:
| 1 |
| 2 |
where is the number of corresponding events. A lower and indicate a higher sensitivity and specificity, respectively.
Experimental results
Figure 3 shows the sensitivities and specificities of seizure prediction for a wide range of SVM parameters C. The error bars in Figs. 3(a) and 3(b) show the means and standard deviations of and as functions of C. As C increases, both and decrease and then saturate. Once , there is no significant difference in sensitivity or specificity. Thus, SVM parameter C was set to 1.0 in what follows. Figure 3(c) shows as a function of when , which is also shown in Table 3.
Fig. 3.

Experimental results: probabilities of a a miss-classification () and b false alarm () are shown for a range of SVM parameters C. In c, is shown as a function of when
Table 3.
Sensitivity and specificity of seizure prediction
| Patient ID | ) | () |
|---|---|---|
| 2 | ||
| 4 | ||
| 6 | ||
| 7 | ||
| 10 | ||
| 12 | ||
| 13 | ||
| 14 | ||
| 15 | ||
| 16 |
The sensitivity and specificity varied across patients. Seven patients with IDs of 2, 4, 6, 7, 10, 12, and 13 showed a perfect sensitivity () and high specificity (). Patient 15 showed a good sensitivity () but worse specificity (). Patients 14 and 16 showed a lower sensitivity and specificity.
Rather surprisingly, the predictability is not related to the number of seizures in the dataset. One might expect that more seizures in the training and testing datasets would lead to a higher sensitivity and specificity. However, this does not necessarily hold in seizure prediction. The dataset of Patients 4, 10, and 16 have five seizures (Table 1). Whereas Patients 4 and 10 showed a perfect sensitivity and specificity, patient 16 showed a much worse predictability ( and , as shown in Table 3 and Fig. 3(c)). Similarly, Patient 12 with four seizures showed a perfect predictability, but Patients 14 and 15 with the same number of seizures showed a worse predictability.
What is even more interesting is that Patient 13 with only two seizures showed a perfect predictability. Using the leave-one-seizure-out cross-validation, iEEGs of one seizure were trained to build a prediction model that predicts another seizure. This implies that the spectral feature combined with SVM is able to identify a predictive pattern that is common to the distinct seizure events of Patient 13.
Conclusions
This paper investigated predictability of an epileptic seizure from an iEEG. A monitoring system was designed to predict impending abnormalities such as seizures from brain signals. A prediction model based on iEEG spectral features and linear SVMs was obtained for each patient. The predictability was measured by testing this model with samples not used for training.
To accurately evaluate the predictive analytics for brain signal monitoring, we explored some cross-validation methods. Abnormalities that one wishes to predict are rare events but may lead to serious clinical impact to the patient. To evaluate the predictability from such imbalanced data, we used a cross-validation framework, called leave-one-seizure-out cross-validation.
The results demonstrate that the prediction model using spectral features and SVMs shows a high sensitivity and specificity for a majority of patients. Among the ten patients studied, seven showed almost perfect sensitivity and specificity. One patient showed a good sensitivity but a lower specificity. Finally, the sensitivity and specificity were lower for two of the patients.
The present study was limited by the small number of patients. More data are clearly needed to fully understand the predictability of epileptic seizures. Nonetheless, even for the small dataset of ten patients, the predictability varied significantly across patients, which demonstrates the variety of abnormal brain activities and potential advantages of patient-specific methods for monitoring brain signals.
Further study will be necessary to determine the cause of low predictability for the two patients. One possibility is the inherent limitation of an EEG. While an EEG has an excellent temporal resolution, its spatial resolution is poor. For this reason, even trained experts often find it difficult to diagnose brain diseases from an EEG. Thus, combining other sensing modalities to monitor the brain state is a promising research direction.
Despite such limitations, a high variability of sensitivity and specificity across patients supports the benefits of personalized health care. Brain signals, similar to other biometric signals, show a high level of uncertainty and variability. Thus, patient-specific features and algorithms are required to extract any useful information from brain signals. This paper focused on utilizing the spectral features of an EEG and SVMs, which showed a good sensitivity and specificity for eight out of the ten patients studied. We are currently exploring other types of features and algorithms that can potentially lead to a higher sensitivity and specificity for the three patients who showed a lower predictability.
Acknowledgements
This work was supported by 2016 Hongik University Research Fund.
References
- 1.Jeong S, Kim S, Kim D, Youn CH, Kim YW. A personalized healthcare system for chronic disease care in home-hospital cloud environments. In: International conference on ICT convergence. 2013. p. 371–76.
- 2.Gandhi M, Singh S. Predictions in heart disease using techniques of data mining. In: International conference on futuristic trends on computational analysis and knowledge management. 2015. p. 520–25.
- 3.Fisher RS, Boas WvE, Blume W, Elger C, Genton P, Lee P, Engel J. Epileptic seizures and epilepsy: definitions proposed by the international league against epilepsy and the international bureau for epilepsy. Epilepsia. 2005;46(4):470–472. doi: 10.1111/j.0013-9580.2005.66104.x. [DOI] [PubMed] [Google Scholar]
- 4.Mormann F, Andrzejak RG, Elger CE, Lehnertz K. Seizure prediction: the long and winding road. Brain. 2007;130(2):314–333. doi: 10.1093/brain/awl241. [DOI] [PubMed] [Google Scholar]
- 5.Scheffer M, Bascompte J, Brock WA, Brovkin V, Carpenter SR, Dakos V, Held H, van Nes EH, Rietkerk M, Sugihara G. Early-warning signals for critical transitions. Nature. 2009;461(7260):53–59. doi: 10.1038/nature08227. [DOI] [PubMed] [Google Scholar]
- 6.Andrzejak RG, Chicharro D, Elger CE, Mormann F. Seizure prediction: any better than chance? Clin Neurophysiol. 2009;120(8):1465–1478. doi: 10.1016/j.clinph.2009.05.019. [DOI] [PubMed] [Google Scholar]
- 7.Iasemidis LD, Sackellares JC, Zaveri HP, Wlliams WJ. Phase space topography and the Lyapunov exponent of electrocorticograms in partial seizures. Brain Topogr. 1990;2:187–201. doi: 10.1007/BF01140588. [DOI] [PubMed] [Google Scholar]
- 8.Le Van Quyen ML, Martinerie J, Baulac M, Varela F. Anticipating epileptic seizures in real time by a non-linear analysis of similarity between EEG recordings. NeuroReport. 1999;10:2149–2155. doi: 10.1097/00001756-199907130-00028. [DOI] [PubMed] [Google Scholar]
- 9.Mormann F, Lehnertz K, David P, Elger C E. Mean phase coherence as a measure for phase synchronization and its application to the EEG of epilepsy patients. Phys D: Nonlinear Phenom. 2000;144(3–4):358–369. doi: 10.1016/S0167-2789(00)00087-7. [DOI] [Google Scholar]
- 10.Iasemidis L, Shiau DS, Chaovalitwongse W, Sackellares J, Pardalos P, Principe J, Carney P, Prasad A, Veeramani B, Tsakalis K. Adaptive epileptic seizure prediction system. IEEE Trans Biomed Eng. 2003;50(5):616–627. doi: 10.1109/TBME.2003.810689. [DOI] [PubMed] [Google Scholar]
- 11.Freiburg seizure prediction project. http://epilepsy.uni-freiburg.de/freiburg-seizure-prediction-project/eeg-database.
- 12.Park Y, Luo L, Parhi KK, Netoff T. Seizure prediction with spectral power of EEG using cost-sensitive support vector machines. Epilepsia. 2011;52(10):1761–1770. doi: 10.1111/j.1528-1167.2011.03138.x. [DOI] [PubMed] [Google Scholar]
- 13.Bandarabadi M, Teixeira CA, Rasekhi J, Dourado A. Epileptic seizure prediction using relative spectral power features. Clin Neurophysiol. 2015;126(2):237–248. doi: 10.1016/j.clinph.2014.05.022. [DOI] [PubMed] [Google Scholar]
- 14.Cortes C, Vapnik V. Support-vector networks. Mach Learn. 1995;20:273–297. [Google Scholar]