Abstract
Apical–basal polarity is essential for the formation and function of epithelial tissues, whereas loss of polarity is a hallmark of tumours. Studies in Drosophila have identified conserved polarity factors that define the apical (Crumbs, Stardust, Par-6, atypical protein kinase C [aPKC]), junctional (Bazooka [Baz]/Par-3), and basolateral (Scribbled [Scrib], Discs large [Dlg], Lethal [2] giant larvae [Lgl]) domains of epithelial cells. Because these conserved factors mark equivalent domains in diverse types of vertebrate and invertebrate epithelia, it is generally assumed that this system underlies polarity in all epithelia. Here, we show that this is not the case, as none of these canonical factors are required for the polarisation of the endodermal epithelium of the Drosophila adult midgut. Furthermore, like vertebrate epithelia but not other Drosophila epithelia, the midgut epithelium forms occluding junctions above adherens junctions (AJs) and requires the integrin adhesion complex for polarity. Thus, Drosophila contains two types of epithelia that polarise by fundamentally different mechanisms. This diversity of epithelial types may reflect their different developmental origins, junctional arrangement, or whether they polarise in an apical–basal direction or vice versa. Since knock-outs of canonical polarity factors in vertebrates often have little or no effect on epithelial polarity and the Drosophila midgut shares several common features with vertebrate epithelia, this diversity of polarity mechanisms is likely to be conserved in other animals.
A developmental study in the fruit fly Drosophila melanogaster shows that apical-basal polarity in the midgut epithelium is established by a different mechanism from other Drosophila epithelia, but more akin to that of vertebrate epithelia.
Author summary
The Drosophila midgut is lined by a single-layered epithelium that acts as a barrier to the environment while allowing for nutrient uptake and related physiological processes. To fulfil these roles, midgut epithelial cells are highly polarised, with a pronounced asymmetric distribution of cellular components. Previous work in Drosophila revealed a conserved set of factors governing cell polarity, and it is thought that this network of proteins underlies all examples of polarity in Drosophila (and other organisms). Here, we demonstrate that the epithelial cells in the Drosophila midgut are not polarised by these canonical polarity factors but instead rely on the integrin adhesion complex. Thus, there are two types of epithelia in Drosophila that polarise using fundamentally different mechanisms. This diversity may reflect a difference in developmental origin (endodermal versus ectodermal), a difference in junctional arrangement, or the direction in which the respective cells polarise. Since knock-outs of canonical polarity factors often have little or no effect on epithelial polarity in vertebrate model systems, this diversity of polarity mechanisms might be conserved in other organisms.
Introduction
Most animal organs and tissues are composed of epithelial cells that adhere laterally to each other to form sheets that act as barriers between compartments. The formation of epithelial monolayers depends on the coordinated polarisation of each cell along its apical–basal axis, and this polarity underlies all aspects of epithelial biology [1,2]. For example, the function of epithelia as barriers to fluids and pathogens depends on the correct positioning of the occluding cell–cell junctions (septate junctions [SJs] in invertebrates and tight junctions in vertebrates), whereas the adhesion between cells depends on the lateral localisation of cadherin-dependent adherens junctions (AJs).
Much of our understanding of how epithelial cells polarise comes from studies of Drosophila melanogaster that have identified a conserved set of epithelial polarity factors that define different cortical domains along the apical–basal axis of the cell. The apical domain is specified by the transmembrane protein Crumbs, the adaptor protein Stardust, and the Par-6/atypical protein kinase C (aPKC) complex; the boundary between the apical and lateral domains is defined by Bazooka (Baz, Par-3 in other organisms), which positions the apical-most lateral junction; and the rest of the lateral domain is marked by Scribbled (Scrib), Discs large (Dlg), and Lethal (2) giant larvae (Lgl) [3]. Null mutations in any of these factors disrupt epithelial polarity in the primary epithelium that forms from the cellular blastoderm of the Drosophila embryo and gives rise to most of the structures of the larva and adult [4–11]. Similarly, loss of any of these genes disrupts the secondary epithelium formed by the follicle cells that surround the developing oocyte [12–14]. In each tissue, Baz seems to play a pivotal role in positioning the apical AJs and in localising the apical factors, which then exclude Baz from the apical domain [15–19]. The identity of the apical and lateral domains is then maintained by mutual antagonism between apical and lateral factors [20,21]. The requirement for some of these factors becomes less stringent in polarised epithelia as they mature. For example, Crumbs is particularly important in epithelial tissues that are remodelling their cell junctions as they undergo morphogenetic rearrangements, and Scrib, Dlg, and Lgl are not required to maintain polarity in mid-embryogenesis, as the Yurt group of lateral proteins takes over the antagonism of the apical factors, although Scrib and Dlg are still required for the formation of the SJs [20–24].
Epithelial cells are thought to have evolved at the origin of multicellularity, as cells first started to adhere to each other to form sheets, suggesting that apical–basal polarity is an ancient invention that is controlled by a conserved mechanism [25]. In support of this view, all of the canonical epithelial factors in Drosophila are conserved in vertebrates and localise to the equivalent cortical domains [26–33]. Knock-downs or knock-outs of some of these factors impair polarity in certain contexts, such as in Xenopus embryonic blastomeres and in some cultured cell lines [34–37]. In most cases, however, knock-outs of canonical polarity factors have little or no effect on polarity or cause unrelated phenotypes. For example, mice homozygous for a null mutation in PAR-3 die in mid-gestation from a heart defect caused by a failure of the epicardial cell migration, but other embryonic epithelia appear to form normally [38]. Similarly, knock-out of PAR-3 in mouse mammary stem cells disrupts the morphogenesis of the mammary gland but not the ability of the cells to polarise [39,40]. Finally, Scrib functions as a planar cell polarity gene in mice but has no obvious effect on apical–basal polarity [41]. Although this lack of polarity phenotypes in mammals could be a result of redundancy between paralogues, it raises the possibility that at least some vertebrate epithelia polarise by different mechanisms from the model Drosophila epithelia. In support of this view, many mammalian epithelia require integrin adhesion to the basement membrane to orient their polarity, whereas the well-characterised Drosophila epithelia do not [42–44]. Furthermore, vertebrate epithelial cells have an inverted arrangement of junctions compared to insect epithelia: the apical junction in vertebrates is the occluding, tight junction, which forms at the apical/lateral boundary above a lateral zonula adherens (ZA), whereas insect cells form lateral SJs, below an apical ZA [45–47].
The possibility that the polarisation of some epithelia is independent of the canonical polarity system prompted us to ask if all Drosophila epithelia polarise in the same way. We therefore examined polarity in the Drosophila adult midgut epithelium, which is mainly absorptive rather than secretory and is endodermal in origin, unlike the well-characterised epithelia, which are secretory and arise from the ectoderm or mesoderm [48]. The adult midgut is a homeostatic tissue in which basal intestinal stem cells (ISCs in Fig 1F) divide to produce new cells that integrate into the epithelium to replace dying enterocytes (ECs), which are shed into the gut lumen. This has the advantage that one can generate homozygous mutant stem cell clones in heterozygotes at the late pupal or adult stage to produce clonal patches of mutant ECs in the adult midgut without disrupting the development of the tissue. Our results reveal that the polarisation of the midgut epithelium does not require any of the canonical polarity factors and has several features in common with vertebrate epithelia, making it a useful model for investigating alternative polarity pathways.
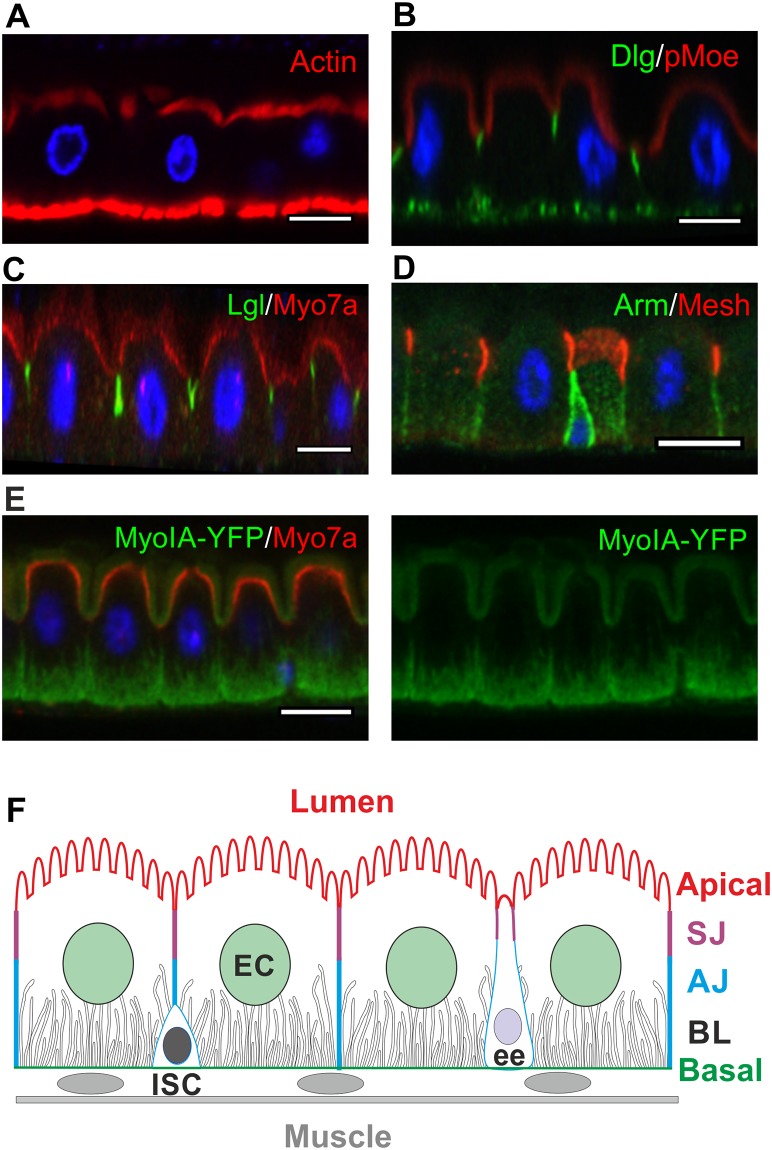
Fig 1. Drosophila midgut epithelial cells have a reversed arrangement of intercellular junctions compared to other Drosophila epithelia.
All micrographs in this and succeeding figures show apical–basal confocal sections of the adult Drosophila midgut epithelium, with apical on top. (A–E) The adult Drosophila midgut epithelium stained for the following: (A) F-actin (phalloidin; red) and DNA (DAPI; blue), revealing the apical brush border. The strong basal signal corresponds to the visceral musculature. (B) pMoe (red), Dlg (green), and DNA (DAPI; blue). (C) Lgl (green), Myo7a (red), and DNA (DAPI, blue). The signal detected by the anti-Myo7a antibody in EC nuclei is unspecific, as it is also present in ck13 (Myo7a) mutant clones (S1G Fig). (D) The AJ component Armadillo (green) and the SJ component Mesh (red), revealing the organization of intercellular junctions in the midgut epithelium. (E) Myo31DF/MyoIA (green) and Myo7a (red). Myo31DF/MyoIA is mainly localised to the apical brush border and the BL, whereas Myo7a is localised to the apical domain, slightly below the brush border. (F) Diagram of the midgut epithelium, which is mainly composed of ECs with a lower frequency of ee cells, both of which turn over in the adult and are replaced by the progeny of basal ISCs. The diagram shows the localisation of the SJs above the AJs and the BL. Scale bars, 10 μm. AJ, adherens junction; Arm, Armadillo; BL, basal labyrinth; Dlg, Discs large; EC, enterocyte; ee, enteroendocrine cell; ISC, intestinal stem cell; Lgl, Lethal (2) giant larvae; MyoIA, Myosin IA; Myo7a, Myosin 7a; Myo31DF, Myosin 31DF; pMoe, phospho-Moesin; SJ, septate junction; YFP, yellow fluorescent protein.
Results
The midgut is a typical epithelium with an apical brush border marked by F-actin (Fig 1A) and phospho-Moesin (pMoe) (Fig 1B) as well as Myosin IA (Fig 1E) and an apical domain marked by Myosin 7a(Fig 1C). However, our analysis led us to rediscover an interesting property of this epithelium: the smooth SJs, which form the occluding barrier to paracellular diffusion, form at the apical side of the lateral domain, above AJs, which are diffusely distributed over the lateral domain [49–52] (Fig 1D and 1F). This is the opposite way around compared to other Drosophila epithelia, in which the AJs condense to form a ZA around the apical margin of the cell, with the SJs, if present, positioned more basally in the lateral membrane. The organisation of junctions in the midgut therefore resembles the junctional arrangement in mammals, in which the occluding tight junctions form above the AJ.
In secretory epithelia, the Crumbs/Stardust complex defines the apical and marginal region and anchors the Par-6/aPKC complex in this domain [5,9,53]. Apical Crb and aPKC then exclude Baz/Par-3 to define the apical/lateral boundary by positioning the apical AJs [17–19]. This raises the question of whether these factors mark the same positions or the same structures when the junctions are reversed in the midgut.
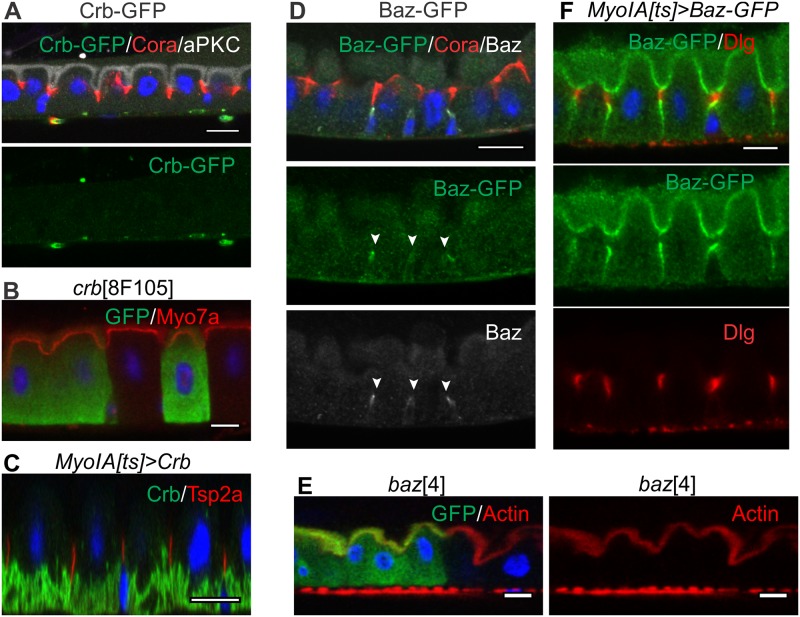
Crumbs is not detectable in the adult midgut epithelium, as has previously been observed in the embryo [4] (Fig 2A). We used the mosaic analysis with a repressible cell marker (MARCM) technique [54] to generate positively marked clones in the adult midgut epithelium for null mutations in crumbs or stardust (crb8F–105, crb11A22; sdtk85). The adults were then dissected 8 to 10 days after clone induction to allow the stem cells, which divide about once a day, to go through multiple divisions because this rules out any perdurance of wild-type proteins expressed before the clones were induced. However, clones of crb and sdt null mutations give no obvious phenotypes (Fig 2B, S1A, S1D and S1E Fig). Overexpression of Crumbs expands the apical domain in other Drosophila epithelia [55]. By contrast, ectopic expression of Crumbs in the adult midgut epithelium does not affect EC polarity and the Crumbs protein does not localise apically, concentrating instead in the basal labyrinth (BL), an extensive set of tubular membrane invaginations from the EC basal surface (Fig 2C). Baz/Par-3 is also not detectable in ECs, although it is expressed in the ISCs, in which it localises apically as previously reported [56] (Fig 2D). MARCM clones homozygous for baz null alleles (baz815–8; baz4) occur at a similar frequency to wild-type clones and contain a normal number of cells, arguing against a role for Baz in asymmetric stem cell division, consistent with the view that these divisions are random and largely symmetric [57]. More importantly, baz null mutant ECs integrate into the epithelium and develop normal apical–basal polarity, in contrast to other epithelia in Drosophila (Fig 2E, S1B, S1C and S1F Fig). Baz-green fluorescent protein (GFP) localises apically and to the AJs when ectopically expressed in the midgut epithelium, consistent with binding to the Par-6/aPKC complex and to E-cadherin complexes (Fig 2F). However, ectopic Baz expression has no effect on the polarity of the ECs or on the formation of an apical SJ.
Fig 2. Crb and Baz are neither detectably expressed nor required for apical–basal polarity in adult midgut ECs.
(A) Adult midgut epithelium from a female carrying a GFP protein trap insertion in the endogenous Crb locus stained with anti-GFP (green). Crb-GFP is not detectable in the adult midgut epithelium. Cora (red) and aPKC (white) are also shown. (B) ECs homozygous for crb8F105 (marked by GFP, green). The apical domain forms normally as revealed by the apical localization of Myo7a (red). (C) Conditional mis-expression of UAS-Crb in adult midgut ECs using MyoIA-GAL4; tubP-GAL80ts. Crb (green) localises to the BL but does not perturb SJ formation as revealed by Tsp2a (red). (D) Adult midgut epithelium expressing Baz-GFP under endogenous control (protein trap line), stained with anti-GFP (green) and anti-Baz (white). Baz/Baz-GFP is not detectable in ECs but is expressed in ISCs, in which it localises apically (white arrowhead). Cora (red) labels the SJ. (E) ECs homozygous mutant for baz4 (marked by GFP, green). The apical brush border forms normally as revealed by staining for F-actin with phalloidin (red). (F) Ectopic expression of UAS-Baz-GFP in adult midgut ECs using MyoIA-GAL4; tubP-GAL80ts has no effect on the formation of the apical SJ marked by Dlg (red). Baz-GFP localises to the apical domain and AJs (green). Scale bars, 10 μm. AJ, adherens junction; aPKC, atypical protein kinase C; Baz, Bazooka; BL, basal labyrinth; Cora, Coracle; Crb, Crumbs; Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; MyoIA, Myosin IA; Myo7a, Myosin 7a; SJ, septate junction; Tsp2a, Tetraspanin 2a; tubP, tubulin promoter; UAS, Upstream Activation Sequence.
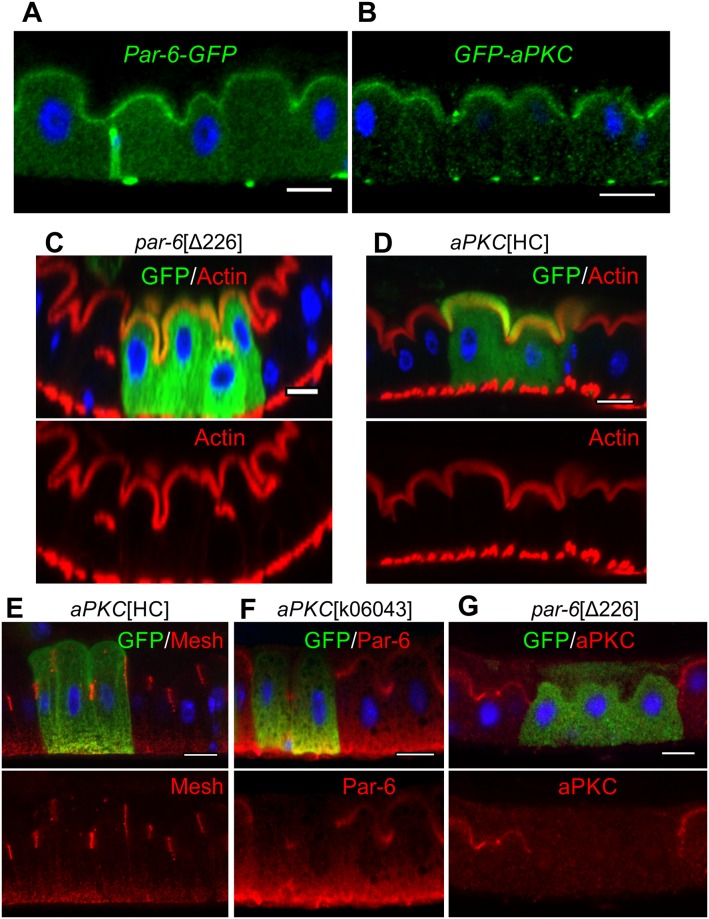
Both aPKC and Par-6 are expressed in the midgut and localise apically, as they do in all other epithelia (Fig 3A and 3B). In most polarised cells, the apical localisation of the Par-6/aPKC complex depends on Baz/Par-3, and in epithelia, this also requires Crumbs and Stardust [2]. Consistent with our observation that these proteins are absent from ECs, neither Baz nor Crb are required for the localisation of Par-6, indicating that the latter must be targeted apically by a distinct mechanism (S1A and S1B Fig). Surprisingly, the apical domain forms normally in par-6Δ226 and aPKCk06043 mutant clones, and the morphology of the cells is unaffected (Fig 3C, S1H and S1J Fig). Although aPKCk06043 is considered a null allele, the corresponding P element insertion does not disrupt the shorter isoforms of aPKC. Thus, it is conceivable that some aPKC activity remains in aPKCk06043 homozygous mutant cells. We therefore used CRISPR to generate a complete null, aPKCHC, a frameshift mutation resulting in a premature stop codon and a truncation of aPKC at amino acid 409, which is located in the middle of the kinase domain (S1L Fig). Homozygous aPKCHC clones also show no phenotype, forming normal actin-rich brush borders and SJs, confirming that aPKC is dispensable for EC polarity (Fig 3D and 3E). Nevertheless, Par-6 is lost from the apical domain in aPKCk06043 clones and aPKC is not apical in par-6Δ226 clones, showing that their localisations are interdependent (Fig 3F and 3G). Thus, the Par-6/aPKC complex still marks the apical domain of the midgut epithelium but is not required for its formation or maintenance.
Fig 3. aPKC and Par-6 are not required for EC polarity.
Subcellular localisation of endogenously tagged Par-6–GFP (A) and GFP-aPKC (B), as revealed by anti-GFP staining (green). MARCM clones (marked by GFP, green) homozygous mutant for par-6Δ226 (C) and aPKCHC (D, E) show normal apical actin brush borders (F-actin in red) and SJ localisation (E; Mesh, red). The apical localisation of Par-6 (red) is lost in aPKCK06043 MARCM clones (marked by GFP, green) (F) as is the apical localisation of aPKC (red) in par6Δ226 MARCM clones (marked by GFP, green) (G). Scale bars, 10 μm. aPKC, atypical protein kinase C; EC, enterocyte; GFP, green fluorescent protein; MARCM, mosaic analysis with a repressible cell marker; SJ, septate junction.
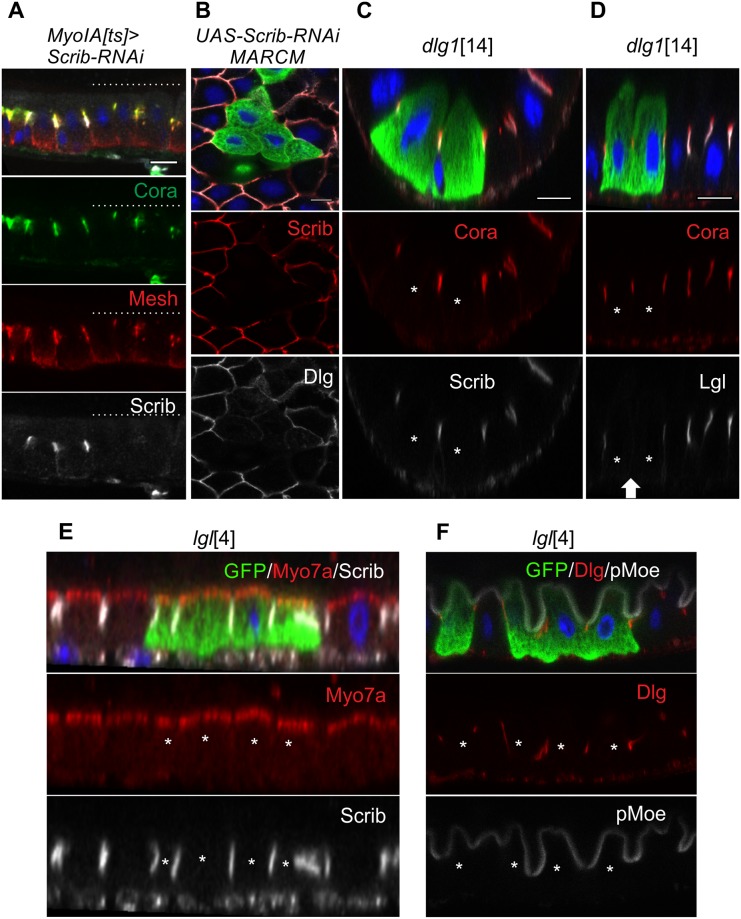
The lateral polarity factors Scrib, Dlg, and Lgl are all expressed in the midgut and colocalise with each other to the SJs, marked by the conserved SJ component, Coracle [58] (Fig 1B and 1C, S2A Fig). Since the SJs form at the apical side of the lateral membrane in the midgut, in the position occupied by the AJs in other Drosophila epithelia, these proteins mark a conserved structure rather than a conserved position. The lateral epithelial polarity factors are required for the formation of pleated SJs in the embryo [20,21,59]. However, the smooth SJs form normally in scrib, dlg, and lgl mutant clones or when these factors are depleted by RNA interference (RNAi) (Fig 4A and 4C, S2B Fig). The apical domain is also unaffected in scrib, dlg, and lgl mutant or knock-down cells, in contrast to other epithelia in which apical factors are mislocalised to the basolateral domain (Fig 4E and 4F, S2C and S2D Fig). In stage 13 embryos, the Yurt complex excludes apical factors from the lateral membrane in place of Scrib, Dlg, and Lgl [23,24]. We therefore also examined the role of Yurt in the midgut. Yurt localises to the SJs, as it does in the embryo, but yurt mutant clones still polarise normally and form SJs that recruit Lgl (Fig 5). Finally, we examined Par-1, which localises laterally in other epithelia in which it plays a role in localising AJs through phosphorylation of Baz and in organising the microtubule cytoskeleton [60,61]. Par-1 is expressed in the ISCs, in which it localises laterally but is not detectable in the ECs. Consistent with its lack of expression, par-19 mutant ECs show normal apical–basal polarity (S3A and S3B Fig). Thus, all the canonical epithelial polarity factors are dispensable for the polarisation of the midgut epithelium, even though Par-6, aPKC, Scrib, Dlg, Lgl, and Yurt are expressed and localise to equivalent positions to secretory epithelia.
Fig 4. Scrib, Dlg, and Lgl localise to SJs but are not required for SJ formation or apical–basal polarity.
(A) Mosaic knock-down of Scrib by RNAi in adult ECs. The SJ markers Cora (green) and Mesh (red) localise normally in cells depleted of Scrib (white). The ECs lacking Scrib are indicated by dashed lines above. (B) Dlg (red) does not localise to the SJs in Scrib–RNAi MARCM clones (marked by GFP, green). (C) Cora (red) and Scrib (white) localise to the SJs in dlg114 MARCM clones (marked by GFP, green). (D) Lgl does not localise to the SJ between dlg114 mutant cells (marked by GFP, green). The white arrow indicates the SJ between the dlg114 mutant cells. (E) Myo7a (red) and Scrib (white) localise normally to the apical cortex and SJs respectively in lgl4 MARCM clones (marked by GFP, green). (F) Dlg (red) and pMoe (white) localise normally in lgl4 mutant cells (marked by GFP, green). White asterisks mark the mutant cells. Scale bars, 10 μm. Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; MARCM, mosaic analysis with a repressible cell marker; Myo7a, Myosin 7a; pMoe, phospho-Moesin; RNAi, RNA intereference; Scrib, Scribbled; SJ, septate junction; UAS, Upstream Activation Sequence.
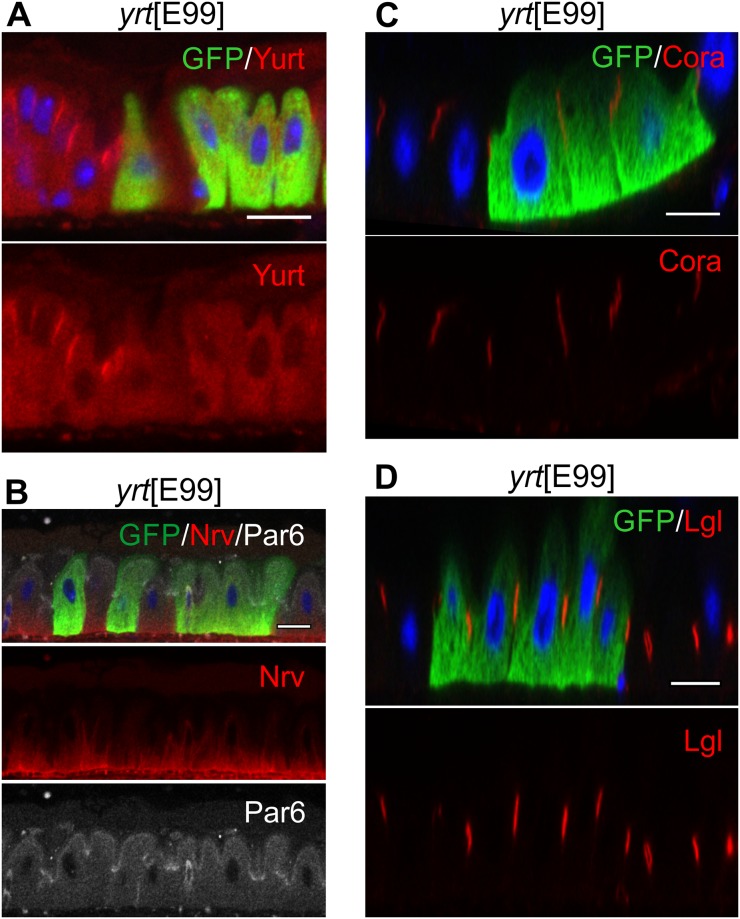
Fig 5. Yurt is not required for EC polarisation.
(A) Yurt (red) localises to SJs and is not detectable in ECs mutant for the loss-of-function allele yrt[E99] (marked by GFP, green). (B) In yurt mutant clones (marked by GFP, green), Par-6 (white) still localises to apical domain and Na+ K+ ATPase (stained with the pan-Na–pump β-subunit (Nrv) [62] antibody Nrv5F7 (red)) still localises to BL. (C) and (D) Yurt is not required for SJ formation, as Cora (C) (red) and Lgl (D) (red) still localise to SJ in yurt mutant clones (marked by GFP, green). Scale bars, 10 μm. BL, basal labyrinth; EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; Nrv, Nervana; SJ, septate junction.
The relationships between the lateral factors has been difficult to assess in other epithelia because mutants in scrib, dlg, and lgl give rise to round, unpolarised cells without an identifiable lateral domain [11]. We took advantage of the normal EC polarisation in these mutants to investigate the interdependence of their recruitment to the SJs. Neither Dlg nor Lgl are recruited to SJs in cells depleted of Scrib by RNAi (Fig 4B and S2D Fig). Scrib localises normally in dlg mutant clones, whereas both Scrib and Dlg localise normally to the SJs in lgl mutant clones (Fig 4C–4F). Thus, there is a simple hierarchical relationship between these factors in the midgut epithelium, in which Scrib is required to recruit Dlg, which is needed for Lgl localisation.
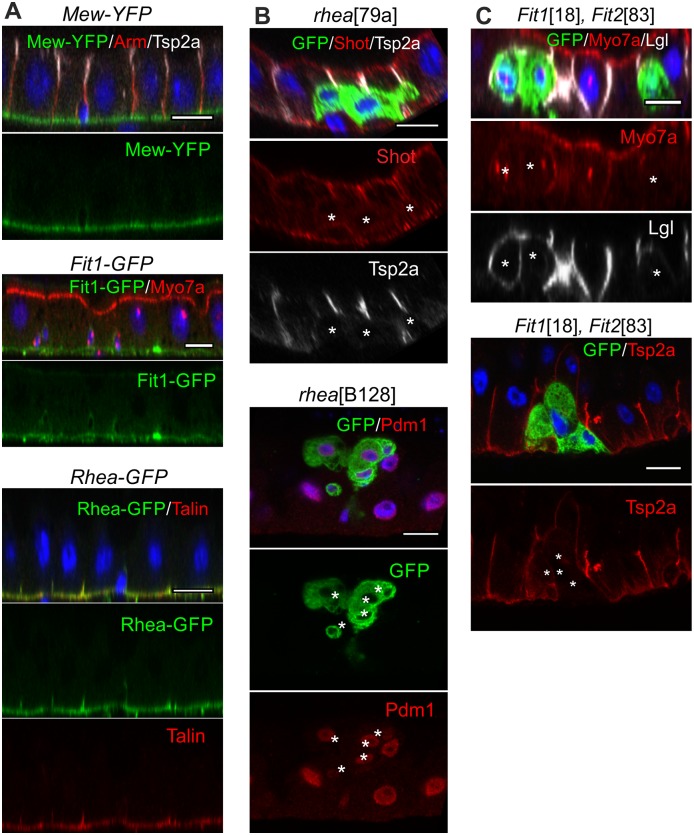
The surprising observation that none of the classical epithelial polarity factors are required for the apical–basal polarisation of the midgut epithelium raises the question of how polarity is generated and maintained. Given the similar junctional arrangement to mammalian epithelia, we addressed whether polarity in midgut ECs depends on integrin-dependent adhesion to the extracellular matrix, as it does in several mammalian epithelia [42–44]. Components of the integrin adhesion complex, such as the α-integrin Mew and the essential cytoplasmic adaptor proteins Talin (Drosophila Rhea) [63] and Kindlin (Drosophila Fit 1 [64]; Fit 2 is not detectable expressed in the midgut) are highly localised to the basal side of the midgut epithelium (Fig 6A). The expression of two α-integrins and two β-integrins in the midgut complicates the genetic analysis of their function, so we focused on the cytoplasmic components of the integrin adhesion complex. Clones of cells homozygous for null alleles of rhea (rhea79a and rheaB128) detach from the basement membrane and fail to polarise, forming irregularly shaped cells that do not form SJs or an apical domain (Fig 6B). Most rhea mutant cells remain within the epithelial layer, below the SJs of the wild-type cells, probably because they do not form SJs themselves (Fig 6B and S4A Fig). Despite their inability to polarise and integrate into the epithelium, the mutant cells still appear to differentiate: they become polyploid, the size of their nuclei is not significantly different from that of wild-type cells (rhea nuclear long axis: 6.0 μm ± 1.0 μm versus heterozygous cells: 6.9 μm ± 0.9 μm), and they express the marker for differentiating ECs, Pdm1 [65] (Fig 6B). Cells mutant for both Fit1 and Fit2 show a similar phenotype to Talin mutants. Mutant cells have normal nuclear dimensions (Fit1 Fit2 nuclear long axis: 6.6 μm ± 1.2 μm versus heterozygous cells: 6.9 μm ± 1.0 μm) and express Pdm1 but fail to localise apical markers or form SJs (Fig 6C and S4B Fig).
Fig 6. The integrin adhesion complex is required for EC polarisation and integration.
(A) The α-integrin Mew and the cytoplasmic adaptor proteins of the integrin adhesion complex, Talin (Rhea) and Fermitin (Fit, also known as Kindlin), localise to the basal surface of the midgut epithelium. Mew-YFP, Rhea-GFP (protein trap insertions), and Fit1-GFP (a genomic fosmid construct) [66] were detected with an anti-GFP antibody (green), whereas the subcellular localization of Talin was also revealed with an anti-Talin antibody (red). (B) rhea79a mutant cells (marked by GFP, green) detach from the basement membrane and fail to polarise. Shot (red) is apically enriched in neighbouring wild-type ECs but is not localised in rhea mutant cells, which fail to form SJs marked by Tsp2a (white). Most rheaB128 mutant cells (marked by GFP, green) express Pdm1 (red), a marker for differentiating ECs. (C) Fit118 Fit283 double mutant clones show a similar phenotype: Myo7a is not enriched apically (red), Lgl (white) spreads around the whole plasma membrane, and SJs fail to form as shown by the loss of Tsp2a localization (red). White asterisks * mark the mutant clones. Scale bars, 10 μm. EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; Myo7a, Myosin 7a; SJ, septate junction; Tsp2a, Tetraspanin 2a; YFP, yellow fluorescent protein.
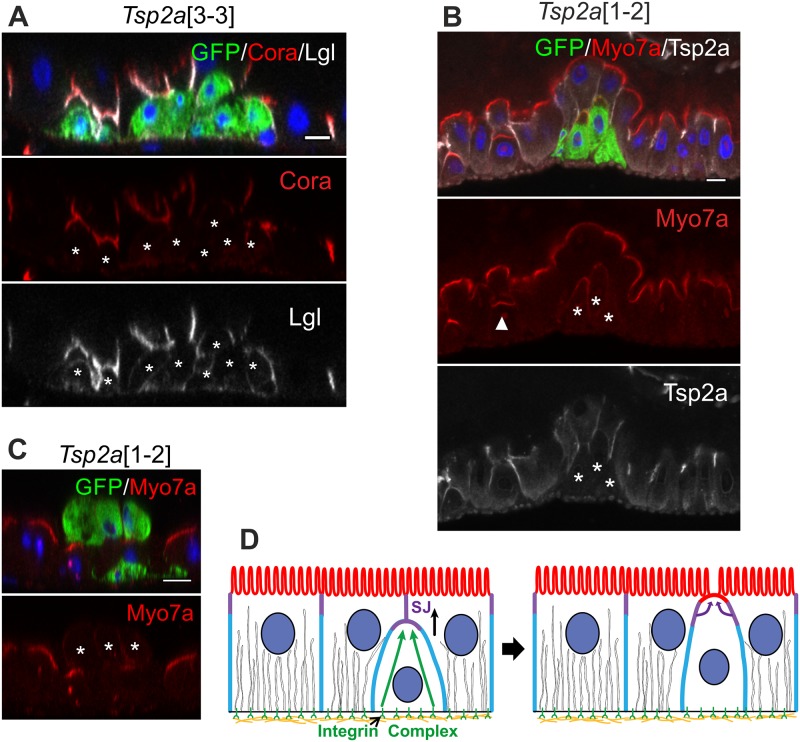
The ISCs lie beneath the epithelium and differentiating ECs must therefore integrate into the epithelium from the basal side, inserting between the SJs of the flanking ECs while maintaining an intact barrier. We therefore examined the effects of mutations in the core SJ components Tsp2a and Mesh [51,52]. More than 90% of mutant cells fail to integrate through the SJs of the neighbouring wild-type cells, and the clones form clusters on the basal side of the epithelium (Fig 7A, S3A and S4C Figs). Nevertheless, the mutant cells still appear to differentiate normally, as shown by their nuclear size (Tsp2a nuclear long axis: 6.0 μm ± 0.9 μm versus heterozygous cells: 6.6 μm ± 0.8 μm) and Pdm1 expression (S4D Fig). Wild-type cells start to form an apical domain as they integrate, before they have a free apical surface, as shown by the enrichment of apical components such as Myo7a (arrowhead in Fig 7B). By contrast, apical markers only weakly localise in cells mutant for SJ components and never form a clear apical domain, even if the cells are extruded from the apical side of the epithelium (Fig 7B and 7C). Thus, the midgut epithelium appears to polarise in a basal to apical manner, in which adhesion to the ECM is required for the formation of the SJs and the SJs are needed for the formation of the apical domain (Fig 7D).
Fig 7. SJ proteins are required for EC polarisation and integration.
(A) Tsp2a mutant clones (marked by GFP, green) fail to integrate into the epithelium, forming basal clusters that lack SJs, as indicated by the loss of Cora (red) localization and diffuse Lgl staining (white). (B) and (C) Neither Tsp2a mutant cells (marked by GFP, green) that cluster on the basal side (B) or that face the lumen of the gut (C) form an apical domain, as shown by the lack of apical Myo7a (red) enrichment. (D) Model of the steps in EC polarisation: adhesion to the basement membrane is required for SJ formation, which in turn is necessary for the formation of an apical domain. White asterisks * mark the mutant clones. White arrowhead in (B) marks the integrating EC. Scale bars, 10 μm. Cora, Coracle; EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; Myo7a; Myosin 7a; SJ, septate junction; Tsp2a, Tetraspanin 2a.
Discussion
Our results reveal that the intestinal epithelium polarises by a fundamentally different mechanism from other Drosophila epithelia, as none of the classical epithelial polarity genes are required for its apical–basal polarisation. This cannot be attributed to redundancy between paralogues, as might be the case in vertebrates, because all of the polarity factors are single-copy genes in Drosophila. Thus, our observations provide strong evidence against the idea that there is a universal system for polarising epithelial cells. Nevertheless, a core set of the polarity factors (Par-6, aPKC, Scrib, Dlg, and Lgl) is expressed in the midgut epithelium, and these proteins localise to the equivalent positions to other Drosophila epithelia. Thus, they may still serve important functions in the midgut epithelium that are not essential for the overall apical–basal polarisation of the cells.
Given that all other epithelia in Drosophila use the canonical polarity pathway, our observations raise the question of why the midgut epithelium is different. This is unlikely to reflect the fact that the midgut is absorptive rather than secretory, as secretory cells in the midgut, such as the enteroendocrine (ee) cells and the acid-secreting copper cells, polarise in the same way as the ECs (S1J and S1K Fig). One key difference between the midgut epithelium and other epithelia is that it is the only epithelial tissue of endodermal origin in the fly, whereas all other epithelia are ectodermal or mesodermal. Thus, it is possible that endodermal epithelia are intrinsically different in the way that they polarise. In support of this view, it has recently been found that PAR-3, PAR-6, and aPKC are degraded in the invaginating endomesoderm of the Cnidarian Nematostella vectensis and are not required for this tissue to form an epithelium [67]. Thus, the difference between endodermal and ectodermal polarity systems may have evolved before the origin of Bilateria. The Drosophila midgut arises from the cellular blastoderm of the embryo and initially polarises in the same way as other embryonic epithelia before it undergoes an epithelial-to-mesenchymal transition (EMT) under the control of the endodermal GATA family transcription factor, Serpent [68]. Serpent drives EMT at least in part by inhibiting the transcription of crumbs and stardust and might therefore contribute to the switch in polarity mechanisms. However, Serpent is turned off as the midgut primordium migrates and is not expressed when the cells reform an epithelium to generate the midgut tube. Indeed, continued expression of Serpent blocks the cells from re-epithelising after migration. Thus, a pulse of Serpent expression may trigger the switch in polarity mechanisms, perhaps through downstream transcription factors, but Serpent itself prevents epithelial polarisation.
A second important difference between the adult midgut epithelium and other epithelia in Drosophila is its reversed arrangement of occluding junctions and AJs, with apical SJs forming above lateral AJs. In other Drosophila epithelia, Baz plays a key role in concentrating the AJs at the apical margin of the lateral membrane to form the ZA [69]. Thus, the absence of Baz in midgut ECs may contribute to the absence of an apical ZA. This cannot be the only factor making the midgut different, however, as ectopic expression of Baz in midgut ECs does not alter the position of the SJs. One reason why the SJs may have evolved to form above the AJs in the midgut is that this places the barrier to paracellular diffusion apically, thereby preventing the contents of the gut lumen from accessing the lateral sides of the cell, which is presumably important because the gut is full of digestive enzymes and potential pathogens. Other Drosophila epithelia that face the external environment—such as the epidermis, trachea, foregut, and hindgut—secrete an impermeable cuticle, which provides a protective covering to prevent pathogens from accessing the cell surface [70]. The development of the typical arthropod cuticular exoskeleton may therefore have freed these epithelia from the need to place their occluding junctions apically, allowing the apical positioning of the ZA.
The third important difference between the midgut epithelium and other epithelia is that the ECs polarise in a basal-to-apical direction as they integrate into the epithelium, whereas all other Drosophila epithelia polarise in an apical-to-basal direction. For example, the primary embryonic epithelium forms during cellularisation, as the furrow canals grow in from the apical surface of the embryo between the nuclei, while the follicle cells polarise in response to an apical cue from the germ line [14,71]. Thus, it is possible that the mechanism by which cells polarise depends on the order in which they generate the basal, lateral, and apical domains. Since the midgut is the only endodermal epithelium in Drosophila, the only epithelium with apical SJs, and the only epithelium that polarises from basal to apical, it is not possible to determine which of these characteristics underlies its alternative mechanism of cell polarisation, and this will require analysis in other organisms with a greater diversity of epithelial cell types.
Whatever the reason for the alternative polarity mechanism in the Drosophila midgut epithelium, its polarity is much more similar to that of well-characterised vertebrate epithelia than other Drosophila epithelia. Firstly, the midgut and vertebrate epithelia have apical occluding junctions above lateral AJs, whereas other Drosophila epithelia have a reversed arrangement of junctions. Secondly, the midgut epithelium does not require the canonical epithelial polarity factors that are essential in other Drosophila epithelia, and this also seems to be the case in some vertebrate epithelia, although this may be due to redundancy between paralogues. Thirdly, the midgut epithelium and a number of vertebrate epithelia depend on signals from the integrin adhesion complex to polarise correctly [42–44]. These similarities suggest that the Drosophila midgut epithelium may prove a better in vivo model for at least some types of vertebrate epithelia. It will therefore be important to determine whether vertebrates also contain distinct types of epithelia whose polarity is controlled by different factors.
Materials and methods
D. melanogaster genetics
w1118 or y2 or OreR flies were used as wild type unless otherwise specified. Other stocks used in this study were as follows:
Fluorescently tagged protein lines: EGFP-aPKC (this paper), Par6-EGFP [72], Mew-YFP [73] (Kyoto DGRC #115524), Baz-EGFP [74] (Bloomington #51572), Myo31DF-YFP [73] (Kyoto DGRC #115611), Crb-EGFP [75] (gift from Y. Hong, University of Pittsburgh, United States of America), Fit1-EGFP [66] (gift from B. Klapholz and N. Brown, Department of Physiology, Development and Neuroscience, University of Cambridge, United Kingdom), Rhea-GFP (Bloomington #39650), Par-1-GFP [61,76].
Mutant stocks: aPKCk06043 [6], aPKCHC (this paper), par6Δ226 [77], dlg1A (Bloomington #57086), dlg114 [59], lgl4 [78], baz815-8 [79], baz4 [8], crb8F105 [4], crb11A22 [4], yurtE99 [80], ck13 (Bloomington #5259), Tsp2A1-2, Tsp2A3-3, Tsp2A2-9 [52] (gift from M. Furuse, Kobe University, Japan), meshf04955 [51] (Kyoto DGRC #114660), Fit118, Fit283, Fit118Fit283 [64], rhea79a [63], rheaB28, rheaB128 [64], ilk54 [81] (gifts from B. Klapholz and N. Brown, Department of Physiology, Development and Neuroscience, University of Cambridge, UK), par-19 (a missense mutation that changes Valine 279 to Aspartate adjacent to the ATP binding pocket of the Par-1 kinase domain. This allele was characterised by Teresa Niccoli and behaves like a null mutation genetically).
UAS responder lines: UAS-Scrib–RNAi (Bloomington #35748), UAS-Crb (Bloomington #5544), UAS-Baz-GFP [17].
The following stocks were used to generate (positively labelled) MARCM [54] clones:
MARCM FRTG13: y w, UAS-mCD8::GFP, Act5C-GAL4, hsFLP[1]; FRTG13 tubP-GAL80.
MARCM FRT82B: y w, UAS-mCD8::GFP, Act5C-GAL4, hsFLP[1];; FRT82B tubP-GAL80.
MARCM FRT19A: w, hsFLP, tubP-GAL80, FRT19A;; tubP-GAL4, UAS-mCD8::GFP/TM3, Sb.
MARCM FRT2A: hsFLP[1]; tubP-GAL4, UAS-mCD8::GFP/CyO, GFP; FRT2A tubP-GAL80 (gift from B. Klapholz and N. Brown).
MARCM FRT40A: y w, UAS-mCD8::GFP, Act5C-GAL4, hsFLP[1]; FRT40A tubP-GAL80.
Negatively marked clones on the X chromosome were generated using the following stock:
y w His2Av::GFP hsFLP[12] FRT19A/FM7a (Bloomington #32045).
Clones mutant for mesh were generated using the following stock: esg-GAL4, UAS-FLP, tubP-GAL80[ts]/CyO; FRT82B nlsGFP (referred to as esg > FLP[ts] in S4C Fig; gift from G. Kolahgar, Department of Physiology, Development and Neuroscience, University of Cambridge, UK).
UAS-Crb and Baz-GFP were expressed in the adult midgut epithelium using the following driver line: y w; MyoIA-GAL4, tubP-GAL80[ts] (referred to as MyoIA[ts] in Figs 2C, 2F and 4A and S2D Fig; gift from G. Kolahgar).
Stock maintenance
Standard procedures were used for Drosophila husbandry and experiments. Flies were reared on standard fly food supplemented with live yeast at 25 °C. For the conditional expression of UAS responder constructs (e.g., RNAi) in adult flies, parental flies were crossed at 18 °C and the resulting offspring reared at the same temperature until eclosion. Adult offspring were collected for 3 days and then transferred to 29 °C to inactivate the temperature-sensitive GAL80 protein. To generate MARCM or GFP-negative clones, flies were crossed at 25 °C and the resulting offspring subjected to heat shocks either as larvae (from L2 until eclosion) or as adults (5–9 days after eclosion). Heat shocks were performed at 37 °C for 1 h (twice daily). Flies were transferred to fresh food vials every 2–3 days and kept at 25 °C for at least 9 days after the last heat shock to ensure that all wild-type gene products from the heterozygous progenitor cells had turned over. For this study, all (midgut) samples were obtained from adult female flies.
Formaldehyde fixation
Samples were dissected in PBS and fixed with 8% formaldehyde (in PBS containing 0.1% Triton X-100) for 10 min at room temperature. Following several washes with PBS supplemented with 0.1% Triton X-100 (washing buffer), samples were incubated in PBS containing 3% normal goat serum (NGS, Stratech Scientific Ltd, Cat. #005-000-121; concentration of stock solution: 10 mg/ml) and 0.1% Triton X-100 (blocking buffer) for 30 min. This fixation method was only used for samples in which F-actin was stained with fluorescently labelled phalloidin, as phalloidin staining is incompatible with heat fixation.
Heat fixation
The heat fixation protocol is based on a heat–methanol fixation method used for Drosophila embryos [82]. Samples were dissected in PBS, transferred to a wire mesh basket, and fixed in hot 1X TSS buffer (0.03% Triton X-100, 4 g/L NaCl; 95 °C) for 3 s before being transferred to ice-cold 1X TSS buffer and chilled for at least 1 min. Subsequently, samples were transferred to washing buffer and processed for immunofluorescence stainings.
Immunofluorescence staining
After blocking, samples were incubated with the appropriate primary antibody/antibodies diluted in blocking buffer at 4 °C overnight. Following several washes, samples were incubated with the appropriate secondary antibody/antibodies either at room temperature for 2 h or at 4 °C overnight. Samples were then washed several times in washing buffer and mounted in Vectashield containing DAPI (Vector Laboratories) on borosilicate glass slides (No. 1.5, VWR International). All antibodies used in this study were tested for specificity using clonal analysis (MARCM) or RNAi.
Primary antibodies:
Mouse monoclonal antibodies: anti-Dlg (4F3), anti-Cora (c615.16), anti-αSpec (3A9), anti-Arm (N2 7A1), anti-Talin (A22A, E16B), anti-Pros (MR1A), anti-Crb (Cq4), anti-Nrv (Nrv5F7), anti-Mys (CF.6G11). All monoclonal antibodies were obtained from the Developmental Studies Hybridoma Bank and used at 1:100 dilution.
Rabbit polyclonal antibodies: anti-pEzrin (NEB Cat. #3726S, 1:200 dilution); anti-Lgl (Santa Cruz Biotechnoloy Inc., d-300, Cat. #SC98260, 1:200 dilution); anti-aPKC (Santa Cruz Biotechnoloy Inc., Cat #SC216, 1:100 dilution); anti-βHSpec [83] (gift from C. Thomas, The Pennsylvania State University, USA, 1:1,000 dilution); anti-Baz [6] (gift from A. Wodarz, University of Cologne, Germany, 1:200 dilution); anti-Par6 [84] (gift from D. J. Montell, UCSB, USA, 1:500 dilution); anti-Mesh [51] and anti-Tsp2A [52] (gift from M. Furuse, 1:1,000 dilution); anti-Scrib [85] (gift from C. Q. Doe, University of Oregon, USA, 1:1,000 dilution); anti-Pdm1 [86] (gift from F. J. Diaz-Benjumea, Centre for Molecular Biology "Severo Ochoa" (CBMSO), Spain, 1:1,000 dilution); anti-Cno [87] (gift from M. Peifer, UNC, USA, 1:1,000 dilution).
Other antibodies used: chicken anti-GFP (Abcam, Cat. #ab13970, 1:1,000 dilution); guinea pig anti-Yurt [88] (gift from U. Tepass, University of Toronto, Canada, 1:1,000 dilution); guinea pig anti-Myo7a [89] (gift from D. Godt, University of Toronto, Canada, 1:1,000 dilution); guinea pig anti-Shot [90] (1:1,000 dilution); rat anti-Mesh [51] (gift from M. Furuse, 1:1,000 dilution).
Secondary antibodies:
Alexa Fluor secondary antibodies (Invitrogen) were used at a dilution of 1:1,000.
Alexa Fluor 488 goat anti-mouse (#A11029), Alexa Fluor 488 goat anti-rabbit (#A11034), Alexa Fluor 488 goat anti-guinea pig (#A11073), Alexa Fluor 488 goat anti-chicken IgY (#A11039), Alexa Fluor 555 goat anti-rat (#A21434), Alexa Fluor 555 goat anti-mouse (#A21422), Alexa Fluor 555 goat anti-rabbit (#A21428), Alexa Fluor 568 goat anti-guinea pig (#A11075), Alexa Fluor 647 goat anti-mouse (#A21236), Alexa Fluor 647 goat anti-rabbit (#A21245), Alexa Fluor 647 goat anti-rat (#A21247). Only cross-adsorbed secondary antibodies were used in this study to eliminate the risk of cross-reactivity.
F-Actin was stained with phalloidin conjugated to Rhodamine (Invitrogen, Cat. #R415, 1:500 dilution).
Imaging
Images were collected on an Olympus IX81 (40× 1.35 NA Oil UPlanSApo, 60× 1.35 NA Oil UPlanSApo) using the Olympus FluoView software Version 3.1 and processed with Fiji (ImageJ).
Generation of endogenous EGFP-aPKC
Endogenously tagged aPKC with EGFP fused to the N-terminus was generated by CRISPR-mediated homologous recombination. In vitro synthesised gRNA [91] to a CRISPR target approximately 60 nucleotides downstream from the aPKC start codon (target sequence GAATAGCGCCAGTATGAACATGG) and a plasmid donor containing the ORF of EGFP as well as appropriate homology arms (1.5 kb upstream and downstream) were coinjected into nos-Cas9–expressing embryos (Bloomington #54591; also known as CFD2) [92]. Single F0 flies were mated to y w flies and allowed to produce larvae before the parent was retrieved for PCR analysis. Progeny from F0 flies in which a recombination event occurred (as indicated by PCR) were further crossed and analysed to confirm integration. Several independent EGFP-aPKC lines were isolated. Recombinants carry the EGFP coding sequence inserted immediately downstream of the endogenous start codon and a short linker (amino acid sequence: Gly-Ser-Gly-Ser) between the coding sequence for EGFP and the coding sequence for aPKC. Homozygous flies are viable and healthy.
Generation of aPKC[HC]
We used the CRISPR/Cas9 method [91] to generate a null allele of aPKC. sgRNA was in vitro transcribed from a DNA template created by PCR from two partially complementary primers: forward primer: 5′-GAAATTAATACGACTCACTATAggattacggcatgtgtaaggGTTTTAGAGCTAGAAATAGC-3′; reverse primer: 5′- AAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAACGGACTAGCCTTATTTTAACTTGCTATTTCTAGCTCTAAAAC-3′. The sgRNA was injected into Act5c-Cas9 embryos [92]. Putative aPKC mutants in the progeny of the injected embryos were recovered, balanced, and sequenced. The aPKCHC allele contains a small deletion around the CRISPR site, resulting in one missense mutation and a frameshift that leads to stop codon at amino acid 409 in the middle of the kinase domain, which is shared by all isoforms (S1L Fig). The aPKCHC allele was subsequently recombined onto FRTG13 to generate MARCM clones. No aPKC protein was detectable by antibody staining in both midgut and follicle cell clones, and follicle cells homozygous for aPKCHC display a phenotype that is indistinguishable from that observed in follicle cells homozygous for the aPKCK06403 allele.
Quantification and statistical analysis
The proportions of rhea, Fit, and Tsp2a mutant cells inside the epithelial layer were calculated as follows: images were taken of different regions of several midguts containing MARCM clones stained with an apical marker. Cells that were above the neighbouring cells or had a clear apical domain were counted as ‘cells NOT inside the layer’, whereas cells without a detectable free apical surface were counted as ‘cells inside the layer’. Data were analysed with Graphpad Prism software. The graph in S4A Fig shows the average percent of cells inside the layer ± SD%.
The nuclear long axis of rhea, Fit, and Tsp2a mutant clones was manually measured in 20 random mutant cells (stained with DAPI, excluding obvious ISCs) and 20 surrounding heterozygous cells using Fiji.
Reproducibility of experiments
All experiments were repeated multiple times with independent crosses: Baz-EGFP (4 independent experiments), EGFP-aPKC (9 independent experiments), Mew-YFP (5 independent experiments), Par6-EGFP (4 independent experiments), Crb-EGFP (5 independent experiments), Fit1-EGFP (4 independent experiments), Myo31DF-YFP (4 independent experiments), MyoIA[ts] > UAS-Scrib-RNAi (11 independent experiments), MyoIA[ts] > UAS-Crb (6 independent experiments), and MyoIA[ts] > UAS-Baz-GFP (3 independent experiments).
The phenotypes of homozygous mutant clones were analysed in multiple guts from independent experiments as follows: baz4 (13 independent experiments, 3,078 mutant cells analysed); baz815-8 (4 independent experiments, 734 cells analysed); aPKCk06043 (8 independent experiments, 3,681 mutant cells analysed); aPKCHC (9 independent experiments, 15,984 mutant cells analysed); par6Δ226 (4 independent experiments, 2,558 mutant cells analysed); crb11A22 (4 independent experiments, 1,478 mutant cells analysed); crb8F105 (5 independent experiments, 3,288 mutant cells analysed); lgl4 (6 independent experiments, 6,790 mutant cells analysed); dlg114 (5 independent experiments, 3,092 mutant cells analysed); rhea79a, rheaB28, rheaB128 (4 independent experiments for each genotype, 184 mutant cells analysed in total); Fit118Fit283 (7 independent experiments, 608 double mutant cells analysed); Fit118 (4 independent experiments, 854 mutant cells analysed); ilk54 (5 independent experiments, 65 mutant cells analysed); Tsp2a1-2, Tsp2a2-9, and Tsp2a3-3 (7, 4, and 6 independent experiments, respectively; a total of 1,205 Tsp2a mutant cells were analysed); and meshf04955 (5 independent experiments, 643 mutant cells analysed).
Supporting information
(A) and (B) Par-6 (red) localises normally to the apical surface of crb8F105 (A) and baz4 (B) mutant cells (marked by GFP, green). (C) baz815–8 mutant cells (marked by GFP, green) form a normal apical brush border as revealed by phalloidin staining of F-actin (red). (D) crb11A22 MARCM clones (marked by GFP, green) show normal Mesh (red) and Cno (white) localisation. (E) sdtK85 MARCM clones (marked by GFP, green) show normal Arm (red) and Lgl (white) localisation. (F) baz4 MARCM clones (marked by GFP, green) show normal Mesh (red) and Cno (white) localization. (G) Unspecific signals of Myo7a staining (red) in nuclei are present in ck13 MARCM clones (marked by GFP, green). (H) aPKCk06043 MARCM clones show normal Cora (red) at SJs and βHSpec (white) at the apical domain. (I) aPKCk06043 MARCM clones show normal αSpec (red) localization at the cortex. (J) The copper cell region has similar polarity to other regions of the midgut, with pMoe (red) at the apical domain, although folded inside, and Dlg (green) at apical SJs. (K) ee cells, which are labelled with Pros (green in nucleus), have Tsp2a (red) at the apical SJ and Arm (green) at the lateral AJ. (L) Schematic genomic map showing the aPKCHC allele and corresponding protein sequence. Scale bars, 10 μm. AJ, adherens junction; aPKC, atypical protein kinase C; Arm, Armadillo; αSpec, α-Spectrin; Baz, Bazooka; βHSpec, βH-Spectrin; Cno, Canoe; Crb, Crumbs; Dlg, Discs large; EC, enterocyte; ee, enteroendocrine; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; MARCM, mosaic analysis with a repressible cell marker; Myo7a, Myosin 7a; pMoe, phospho-Moesin; Pros, Prospero; SJ, septate junction.
(TIF)
(A) Scrib (red) localises to the EC SJs, marked by Cora (green). (B) Mesh (red) localises normally to the SJs of lgl4 mutant cells (marked by GFP, green). (C) dlg114 MARCM clones (marked by GFP, green) show normal apical localization of α-Spectrin (red) and βH-Spectrin (white). (D) RNAi knock-down of Scrib in adult midgut ECs has no effect on the subcellular localization of Myo7a (red) but disrupts Dlg (green) and Lgl (white) localisation to the SJs. Scale bar, 10 μm. Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; MARCM, Mosaic analysis with a repressible cell marker; Myo7a, Myosin 7a; RNAi, RNA interference; Scrib, Scribbled; SJ, septate junction.
(TIF)
(A) Par-1 is not detectable in ECs, although it is expressed in ISCs, as revealed by staining for GFP (green) in a Par-1 GFP protein trap line. Dlg is in red and aPKC in white. (B) A MARCM clone of par-19 (marked by GFP, green). The mutant cells form normal SJs marked by Lgl (red). Scale bar, 10 μm. aPKC, atypical protein kinase C; Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; ISC, intestinal stem cell; Lgl, Lethal (2) giant larvae; MARCM, mosaic analysis with a repressible cell marker; SJ, septate junction.
(TIF)
(A) Most rhea, Fit1, Fit1 Fit2, and Tsp2a mutant cells remain inside the epithelia layer. The graph is based on the analysis of 299 cells in wild-type MARCM clones (13 images), 1,205 Tsp2a mutant cells (25 images from Tsp2a1–2, Tsp2a 3–3, and Tsp2a 2–9 clones), 608 Fit118 Fit283 (FitD) double mutant cells (23 images), 854 Fit118 mutant cells (24 images), and 184 rhea mutant cells (18 images from rheaB28, rhea79a, and rhea B128). The underlying data can be found in S1 Data. (B) Fit118 Fit283 double mutant cells (marked by GFP, green) differentiate as ECs, as revealed by the expression of Pdm1 (white). Talin is in red. (C) A meshf04955 mutant clone (marked by the loss of GFP) stained for α–Spectrin (red) and Mesh (white). (D) Tsp2a1–2 mutant cells (marked by GFP, green) differentiate as ECs, as revealed by the expression of Pdm1 (red). White asterisks * and lines mark the mutant clones. Scale bars, 10 μm. EC, enterocyte; GFP, green fluorescent protein; MARCM, mosaic analysis with a repressible cell marker; SJ, septate junction.
(TIF)
Images were taken of different regions of several midguts containing MARCM clones stained with an apical marker. Cells that were above the neighbouring cells or had a clear apical domain were counted as ‘cells NOT inside the layer’, whereas cells without a detectable free apical surface were counted as ‘cells inside the layer’. For each image, the percentage of clones inside the layer among all clones was calculated. Data were analysed with Graphpad Prism software. The graph in S4A Fig shows the average percent of cells inside the layer ± SD%.
(XLSX)
Acknowledgments
We are grateful to C. Mendes for discussions and help with experiments; G. Kolahgar, Y. Izumi, M. Furuse, B. Klapholz and N. Brown for fly lines and reagents; U. Tepass, C. Thomas, A. Wodarz, D. J. Montell, C. Q. Doe, F. J. Diaz-Benjumea, M. Peifer and D. Godt for antibodies; Teresa Niccoli for characterising the par-19 and other present and past members of the St Johnston laboratory for discussions and comments on the manuscript.
Abbreviations
- AJ
adherens junction
- aPKC
atypical protein kinase C
- Arm
Armadillo
- Baz
Bazooka
- BL
basal labyrinth
- Cno
Canoe
- Cora
Coracle
- Crb
Crumbs
- CRISPR
Clustered Regularly Interspaced Short Palindromic Repeats
- Dlg
Discs large
- EC
enterocyte
- ee
enteroendocrine
- EMT
epithelial to mesenchymal transition
- GFP
green fluorescent protein
- ISC
intestinal stem cell
- Lgl
Lethal (2) giant larvae
- MARCM
Mosaic analysis with a repressible cell marker
- Myo31DF
Myosin31DF
- Myo7a
Myosin 7a
- MyoIA
Myosin IA
- NGS
normal goat serum
- Nrv
Nervana
- pMoe
phospho-Moesin
- Pros
Prospero
- RNAi
RNA inteference
- Scrib
Scribbled
- SJ
septate junction
- Tsp2a
Tetraspanin 2A
- tubP
tubulin promoter
- UAS
Upstream Activation Sequence
- YFP
yellow fluorescent protein
- ZA
zonula adherens
- αSpec
α-Spectrin
- βHSpec
βH-Spectrin
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
This work was supported by a Wellcome Trust Principal Fellowship to DStJ (080007, 207496) and by core support from the Wellcome Trust (092096, 203144) and Cancer Research UK (A14492, A24823). JC was supported by a Royal Society K.C. Wong Postdoctoral Fellowship. ACS was supported by a Wellcome Trust 4 year PhD studentship (083348). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Rodriguez-Boulan E, Macara IG (2014) Organization and execution of the epithelial polarity programme. Nat Rev Mol Cell Biol 15: 225–242. 10.1038/nrm3775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.St Johnston D, Ahringer J (2010) Cell polarity in eggs and epithelia: parallels and diversity. Cell 141: 757–774. 10.1016/j.cell.2010.05.011 [DOI] [PubMed] [Google Scholar]
- 3.Assemat E, Bazellieres E, Pallesi-Pocachard E, Le Bivic A, Massey-Harroche D (2008) Polarity complex proteins. Biochim Biophys Acta 1778: 614–630. 10.1016/j.bbamem.2007.08.029 [DOI] [PubMed] [Google Scholar]
- 4.Tepass U, Theres C, Knust E (1990) crumbs encodes an EGF-like protein expressed on apical membranes of Drosophila epithelial cells and required for organization of epithelia. Cell 61: 787–799. [DOI] [PubMed] [Google Scholar]
- 5.Tepass U, Knust E (1993) Crumbs and stardust act in a genetic pathway that controls the organization of epithelia in Drosophila melanogaster. Dev Biol 159: 311–326. 10.1006/dbio.1993.1243 [DOI] [PubMed] [Google Scholar]
- 6.Wodarz A, Ramrath A, Grimm A, Knust E (2000) Drosophila atypical protein kinase C associates with Bazooka and controls polarity of epithelia and neuroblasts. J Cell Biol 150: 1361–1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Petronczki M, Knoblich JA (2001) DmPAR-6 directs epithelial polarity and asymmetric cell division of neuroblasts in Drosophila. Nat Cell Biol 3: 43–49. 10.1038/35050550 [DOI] [PubMed] [Google Scholar]
- 8.Muller HA, Wieschaus E (1996) armadillo, bazooka, and stardust are critical for early stages in formation of the zonula adherens and maintenance of the polarized blastoderm epithelium in Drosophila. J Cell Biol 134: 149–163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bachmann A, Schneider M, Theilenberg E, Grawe F, Knust E (2001) Drosophila Stardust is a partner of Crumbs in the control of epithelial cell polarity. Nature 414: 638–643. 10.1038/414638a [DOI] [PubMed] [Google Scholar]
- 10.Bilder D, Perrimon N (2000) Localization of apical epithelial determinants by the basolateral PDZ protein Scribble. Nature 403: 676–680. 10.1038/35001108 [DOI] [PubMed] [Google Scholar]
- 11.Bilder D, Li M, Perrimon N (2000) Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science 289: 113–116. [DOI] [PubMed] [Google Scholar]
- 12.Goode S, Perrimon N (1997) Inhibition of patterned cell shape change and cell invasion by Discs large during Drosophila oogenesis. Genes Dev 11: 2532–2544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Benton R, St Johnston D (2003) A conserved oligomerization domain in Drosophila Bazooka/PAR-3 is important for apical localization and epithelial polarity. Curr Biol 13: 1330–1334. [DOI] [PubMed] [Google Scholar]
- 14.Tanentzapf G, Smith C, McGlade J, Tepass U (2000) Apical, lateral, and basal polarization cues contribute to the development of the follicular epithelium during Drosophila oogenesis. J Cell Biol 151: 891–904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Harris TJ, Peifer M (2005) The positioning and segregation of apical cues during epithelial polarity establishment in Drosophila. J Cell Biol 170: 813–823. 10.1083/jcb.200505127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Franz A, Riechmann V (2010) Stepwise polarisation of the Drosophila follicular epithelium. Dev Biol 338: 136–147. 10.1016/j.ydbio.2009.11.027 [DOI] [PubMed] [Google Scholar]
- 17.Morais-de-Sa E, Mirouse V, St Johnston D (2010) aPKC phosphorylation of Bazooka defines the apical/lateral border in Drosophila epithelial cells. Cell 141: 509–523. 10.1016/j.cell.2010.02.040 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Walther RF, Pichaud F (2010) Crumbs/DaPKC-dependent apical exclusion of Bazooka promotes photoreceptor polarity remodeling. Curr Biol 20: 1065–1074. 10.1016/j.cub.2010.04.049 [DOI] [PubMed] [Google Scholar]
- 19.Krahn MP, Buckers J, Kastrup L, Wodarz A (2010) Formation of a Bazooka-Stardust complex is essential for plasma membrane polarity in epithelia. J Cell Biol 190: 751–760. 10.1083/jcb.201006029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tanentzapf G, Tepass U (2003) Interactions between the crumbs, lethal giant larvae and bazooka pathways in epithelial polarization. Nat Cell Biol 5: 46–52. 10.1038/ncb896 [DOI] [PubMed] [Google Scholar]
- 21.Bilder D, Schober M, Perrimon N (2003) Integrated activity of PDZ protein complexes regulates epithelial polarity. Nat Cell Biol 5: 53–58. 10.1038/ncb897 [DOI] [PubMed] [Google Scholar]
- 22.Campbell K, Knust E, Skaer H (2009) Crumbs stabilises epithelial polarity during tissue remodelling. J Cell Sci 122: 2604–2612. 10.1242/jcs.047183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Laprise P, Lau KM, Harris KP, Silva-Gagliardi NF, Paul SM, et al. (2009) Yurt, Coracle, Neurexin IV and the Na(+),K(+)-ATPase form a novel group of epithelial polarity proteins. Nature 459: 1141–1145. 10.1038/nature08067 [DOI] [PubMed] [Google Scholar]
- 24.Gamblin CL, Hardy EJ, Chartier FJ, Bisson N, Laprise P (2014) A bidirectional antagonism between aPKC and Yurt regulates epithelial cell polarity. J Cell Biol 204: 487–495. 10.1083/jcb.201308032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Alberts B, Johnson A, Lewis J, Raff M, Roberts K, et al. (2002) Molecular Biology of the Cell. New York: Garland Science. [Google Scholar]
- 26.Makarova O, Roh MH, Liu CJ, Laurinec S, Margolis B (2003) Mammalian Crumbs3 is a small transmembrane protein linked to protein associated with Lin-7 (Pals1). Gene 302: 21–29. [DOI] [PubMed] [Google Scholar]
- 27.Yamanaka T, Horikoshi Y, Sugiyama Y, Ishiyama C, Suzuki A, et al. (2003) Mammalian Lgl forms a protein complex with PAR-6 and aPKC independently of PAR-3 to regulate epithelial cell polarity. Curr Biol 13: 734–743. [DOI] [PubMed] [Google Scholar]
- 28.Ebnet K, Suzuki A, Horikoshi Y, Hirose T, Meyer Zu Brickwedde MK, et al. (2001) The cell polarity protein ASIP/PAR-3 directly associates with junctional adhesion molecule (JAM). EMBO J 20: 3738–3748. 10.1093/emboj/20.14.3738 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Laprise P, Viel A, Rivard N (2004) Human homolog of disc-large is required for adherens junction assembly and differentiation of human intestinal epithelial cells. J Biol Chem 279: 10157–10166. 10.1074/jbc.M309843200 [DOI] [PubMed] [Google Scholar]
- 30.Navarro C, Nola S, Audebert S, Santoni MJ, Arsanto JP, et al. (2005) Junctional recruitment of mammalian Scribble relies on E-cadherin engagement. Oncogene 24: 4330–4339. 10.1038/sj.onc.1208632 [DOI] [PubMed] [Google Scholar]
- 31.Yamanaka T, Horikoshi Y, Izumi N, Suzuki A, Mizuno K, et al. (2006) Lgl mediates apical domain disassembly by suppressing the PAR-3-aPKC-PAR-6 complex to orient apical membrane polarity. J Cell Sci 119: 2107–2118. 10.1242/jcs.02938 [DOI] [PubMed] [Google Scholar]
- 32.Stucke VM, Timmerman E, Vandekerckhove J, Gevaert K, Hall A (2007) The MAGUK protein MPP7 binds to the polarity protein hDlg1 and facilitates epithelial tight junction formation. Mol Biol Cell 18: 1744–1755. 10.1091/mbc.E06-11-0980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kohjima M, Noda Y, Takeya R, Saito N, Takeuchi K, et al. (2002) PAR3beta, a novel homologue of the cell polarity protein PAR3, localizes to tight junctions. Biochem Biophys Res Commun 299: 641–646. [DOI] [PubMed] [Google Scholar]
- 34.Chalmers AD, Pambos M, Mason J, Lang S, Wylie C, et al. (2005) aPKC, Crumbs3 and Lgl2 control apicobasal polarity in early vertebrate development. Development 132: 977–986. 10.1242/dev.01645 [DOI] [PubMed] [Google Scholar]
- 35.Roh MH, Fan S, Liu CJ, Margolis B (2003) The Crumbs3-Pals1 complex participates in the establishment of polarity in mammalian epithelial cells. J Cell Sci 116: 2895–2906. 10.1242/jcs.00500 [DOI] [PubMed] [Google Scholar]
- 36.Chen X, Macara IG (2005) Par-3 controls tight junction assembly through the Rac exchange factor Tiam1. Nat Cell Biol 7: 262–269. 10.1038/ncb1226 [DOI] [PubMed] [Google Scholar]
- 37.Mertens AE, Rygiel TP, Olivo C, van der Kammen R, Collard JG (2005) The Rac activator Tiam1 controls tight junction biogenesis in keratinocytes through binding to and activation of the Par polarity complex. J Cell Biol 170: 1029–1037. 10.1083/jcb.200502129 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hirose T, Karasawa M, Sugitani Y, Fujisawa M, Akimoto K, et al. (2006) PAR3 is essential for cyst-mediated epicardial development by establishing apical cortical domains. Development 133: 1389–1398. 10.1242/dev.02294 [DOI] [PubMed] [Google Scholar]
- 39.McCaffrey LM, Macara IG (2009) The Par3/aPKC interaction is essential for end bud remodeling and progenitor differentiation during mammary gland morphogenesis. Genes Dev 23: 1450–1460. 10.1101/gad.1795909 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.McCaffrey LM, Montalbano J, Mihai C, Macara IG (2012) Loss of the Par3 polarity protein promotes breast tumorigenesis and metastasis. Cancer Cell 22: 601–614. 10.1016/j.ccr.2012.10.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Montcouquiol M, Rachel RA, Lanford PJ, Copeland NG, Jenkins NA, et al. (2003) Identification of Vangl2 and Scrb1 as planar polarity genes in mammals. Nature 423: 173–177. 10.1038/nature01618 [DOI] [PubMed] [Google Scholar]
- 42.Yu W, Datta A, Leroy P, O'Brien LE, Mak G, et al. (2005) Beta1-integrin orients epithelial polarity via Rac1 and laminin. Mol Biol Cell 16: 433–445. 10.1091/mbc.E04-05-0435 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Myllymaki SM, Teravainen TP, Manninen A (2011) Two distinct integrin-mediated mechanisms contribute to apical lumen formation in epithelial cells. PLoS ONE 6: e19453 10.1371/journal.pone.0019453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Akhtar N, Streuli CH (2013) An integrin-ILK-microtubule network orients cell polarity and lumen formation in glandular epithelium. Nat Cell Biol 15: 17–27. 10.1038/ncb2646 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Farquhar MG, Palade GE (1963) Junctional complexes in various epithelia. J Cell Biol 17: 375–412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tepass U, Tanentzapf G, Ward R, Fehon R (2001) Epithelial cell polarity and cell junctions in Drosophila. Annu Rev Genet 35: 747–784. 10.1146/annurev.genet.35.102401.091415 [DOI] [PubMed] [Google Scholar]
- 47.Woods DF, Wu JW, Bryant PJ (1997) Localization of proteins to the apico-lateral junctions of Drosophila epithelia. Dev Genet 20: 111–118. 10.1002/(SICI)1520-6408(1997)20:2<111::AID-DVG4>3.0.CO;2-A [DOI] [PubMed] [Google Scholar]
- 48.Naszai M, Carroll LR, Cordero JB (2015) Intestinal stem cell proliferation and epithelial homeostasis in the adult Drosophila midgut. Insect Biochem Mol Biol 67: 9–14. 10.1016/j.ibmb.2015.05.016 [DOI] [PubMed] [Google Scholar]
- 49.Baumann O (2001) Posterior midgut epithelial cells differ in their organization of the membrane skeleton from other drosophila epithelia. Exp Cell Res 270: 176–187. 10.1006/excr.2001.5343 [DOI] [PubMed] [Google Scholar]
- 50.Furuse M, Izumi Y (2017) Molecular dissection of smooth septate junctions: understanding their roles in arthropod physiology. Ann N Y Acad Sci 1397: 17–24. 10.1111/nyas.13366 [DOI] [PubMed] [Google Scholar]
- 51.Izumi Y, Yanagihashi Y, Furuse M (2012) A novel protein complex, Mesh-Ssk, is required for septate junction formation in the Drosophila midgut. J Cell Sci 125: 4923–4933. 10.1242/jcs.112243 [DOI] [PubMed] [Google Scholar]
- 52.Izumi Y, Motoishi M, Furuse K, Furuse M (2016) A tetraspanin regulates septate junction formation in Drosophila midgut. J Cell Sci 129: 1155–1164. 10.1242/jcs.180448 [DOI] [PubMed] [Google Scholar]
- 53.Hurd TW, Gao L, Roh MH, Macara IG, Margolis B (2003) Direct interaction of two polarity complexes implicated in epithelial tight junction assembly. Nat Cell Biol 5: 137–142. 10.1038/ncb923 [DOI] [PubMed] [Google Scholar]
- 54.Lee T, Luo L (2001) Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci 24: 251–254. [DOI] [PubMed] [Google Scholar]
- 55.Wodarz A, Hinz U, Engelbert M, Knust E (1995) Expression of crumbs confers apical character on plasma membrane domains of ectodermal epithelia of Drosophila. Cell 82: 67–76. [DOI] [PubMed] [Google Scholar]
- 56.Goulas S, Conder R, Knoblich JA (2012) The Par complex and integrins direct asymmetric cell division in adult intestinal stem cells. Cell Stem Cell 11: 529–540. 10.1016/j.stem.2012.06.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.de Navascues J, Perdigoto CN, Bian Y, Schneider MH, Bardin AJ, et al. (2012) Drosophila midgut homeostasis involves neutral competition between symmetrically dividing intestinal stem cells. EMBO J 31: 2473–2485. 10.1038/emboj.2012.106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Fehon RG, Dawson IA, Artavanis-Tsakonas S (1994) A Drosophila homologue of membrane-skeleton protein 4.1 is associated with septate junctions and is encoded by the coracle gene. Development 120: 545–557. [DOI] [PubMed] [Google Scholar]
- 59.Woods DF, Hough C, Peel D, Callaini G, Bryant PJ (1996) Dlg protein is required for junction structure, cell polarity, and proliferation control in Drosophila epithelia. J Cell Biol 134: 1469–1482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Benton R, St Johnston D (2003) Drosophila PAR-1 and 14-3-3 inhibit Bazooka/PAR-3 to establish complementary cortical domains in polarized cells. Cell 115: 691–704. [DOI] [PubMed] [Google Scholar]
- 61.Doerflinger H, Benton R, Shulman JM, St Johnston D (2003) The role of PAR-1 in regulating the polarised microtubule cytoskeleton in the Drosophila follicular epithelium. Development 130: 3965–3975. [DOI] [PubMed] [Google Scholar]
- 62.Sun B, Salvaterra PM (1995) Characterization of nervana, a Drosophila melanogaster neuron-specific glycoprotein antigen recognized by anti-horseradish peroxidase antibodies. J Neurochem 65: 434–443. [DOI] [PubMed] [Google Scholar]
- 63.Brown NH, Gregory SL, Rickoll WL, Fessler LI, Prout M, et al. (2002) Talin is essential for integrin function in Drosophila. Dev Cell 3: 569–579. [DOI] [PubMed] [Google Scholar]
- 64.Klapholz B, Herbert SL, Wellmann J, Johnson R, Parsons M, et al. (2015) Alternative mechanisms for talin to mediate integrin function. Curr Biol 25: 847–857. 10.1016/j.cub.2015.01.043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Beebe K, Lee WC, Micchelli CA (2010) JAK/STAT signaling coordinates stem cell proliferation and multilineage differentiation in the Drosophila intestinal stem cell lineage. Dev Biol 338: 28–37. 10.1016/j.ydbio.2009.10.045 [DOI] [PubMed] [Google Scholar]
- 66.Maartens AP, Wellmann J, Wictome E, Klapholz B, Green H, et al. (2016) Drosophila vinculin is more harmful when hyperactive than absent, and can circumvent integrin to form adhesion complexes. J Cell Sci 129: 4354–4365. 10.1242/jcs.189878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Salinas-Saavedra M, Rock AQ, Martindale MQ (2018) Germ layer-specific regulation of cell polarity and adhesion gives insight into the evolution of mesoderm. Elife 7: pii: e36740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Campbell K, Whissell G, Franch-Marro X, Batlle E, Casanova J (2011) Specific GATA factors act as conserved inducers of an endodermal-EMT. Dev Cell 21: 1051–1061. 10.1016/j.devcel.2011.10.005 [DOI] [PubMed] [Google Scholar]
- 69.Harris TJ, Peifer M (2004) Adherens junction-dependent and -independent steps in the establishment of epithelial cell polarity in Drosophila. J Cell Biol 167: 135–147. 10.1083/jcb.200406024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Hillman R, Lesnik LH (1970) Cuticle formation in the embryo of Drosophila melanogaster. Journal of Morphology 131: 383–395. [Google Scholar]
- 71.Figard L, Xu H, Garcia HG, Golding I, Sokac AM (2013) The plasma membrane flattens out to fuel cell-surface growth during Drosophila cellularization. Dev Cell 27: 648–655. 10.1016/j.devcel.2013.11.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Wirtz-Peitz F, Nishimura T, Knoblich JA (2008) Linking cell cycle to asymmetric division: Aurora-A phosphorylates the Par complex to regulate Numb localization. Cell 135: 161–173. 10.1016/j.cell.2008.07.049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Lowe N, Rees JS, Roote J, Ryder E, Armean IM, et al. (2014) Analysis of the expression patterns, subcellular localisations and interaction partners of Drosophila proteins using a pigP protein trap library. Development 141: 3994–4005. 10.1242/dev.111054 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Buszczak M, Paterno S, Lighthouse D, Bachman J, Planck J, et al. (2007) The Carnegie protein trap library: a versatile tool for Drosophila developmental studies. Genetics 175: 1505–1531. 10.1534/genetics.106.065961 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Huang J, Zhou W, Dong W, Watson AM, Hong Y (2009) From the Cover: Directed, efficient, and versatile modifications of the Drosophila genome by genomic engineering. Proc Natl Acad Sci U S A 106: 8284–8289. 10.1073/pnas.0900641106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lighthouse DV, Buszczak M, Spradling AC (2008) New components of the Drosophila fusome suggest it plays novel roles in signaling and transport. Dev Biol 317: 59–71. 10.1016/j.ydbio.2008.02.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Huynh JR, Petronczki M, Knoblich JA, St Johnston D (2001) Bazooka and PAR-6 are required with PAR-1 for the maintenance of oocyte fate in Drosophila. Curr Biol 11: 901–906. [DOI] [PubMed] [Google Scholar]
- 78.Ohshiro T, Yagami T, Zhang C, Matsuzaki F (2000) Role of cortical tumour-suppressor proteins in asymmetric division of Drosophila neuroblast. Nature 408: 593–596. 10.1038/35046087 [DOI] [PubMed] [Google Scholar]
- 79.Kuchinke U, Grawe F, Knust E (1998) Control of spindle orientation in Drosophila by the Par-3-related PDZ-domain protein Bazooka. Curr Biol 8: 1357–1365. [DOI] [PubMed] [Google Scholar]
- 80.Hoover KB, Bryant PJ (2002) Drosophila Yurt is a new protein-4.1-like protein required for epithelial morphogenesis. Dev Genes Evol 212: 230–238. 10.1007/s00427-002-0231-6 [DOI] [PubMed] [Google Scholar]
- 81.Zervas CG, Psarra E, Williams V, Solomon E, Vakaloglou KM, et al. (2011) A central multifunctional role of integrin-linked kinase at muscle attachment sites. J Cell Sci 124: 1316–1327. 10.1242/jcs.081422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Muller HA (2008) Immunolabeling of embryos. Methods Mol Biol 420: 207–218. 10.1007/978-1-59745-583-1_12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Thomas GH, Kiehart DP (1994) Beta heavy-spectrin has a restricted tissue and subcellular distribution during Drosophila embryogenesis. Development 120: 2039–2050. [DOI] [PubMed] [Google Scholar]
- 84.Pinheiro EM, Montell DJ (2004) Requirement for Par-6 and Bazooka in Drosophila border cell migration. Development 131: 5243–5251. 10.1242/dev.01412 [DOI] [PubMed] [Google Scholar]
- 85.Albertson R, Doe CQ (2003) Dlg, Scrib and Lgl regulate neuroblast cell size and mitotic spindle asymmetry. Nat Cell Biol 5: 166–170. 10.1038/ncb922 [DOI] [PubMed] [Google Scholar]
- 86.Terriente J, Perea D, Suzanne M, Diaz-Benjumea FJ (2008) The Drosophila gene zfh2 is required to establish proximal-distal domains in the wing disc. Dev Biol 320: 102–112. 10.1016/j.ydbio.2008.04.028 [DOI] [PubMed] [Google Scholar]
- 87.Sawyer JK, Harris NJ, Slep KC, Gaul U, Peifer M (2009) The Drosophila afadin homologue Canoe regulates linkage of the actin cytoskeleton to adherens junctions during apical constriction. J Cell Biol 186: 57–73. 10.1083/jcb.200904001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Laprise P, Beronja S, Silva-Gagliardi NF, Pellikka M, Jensen AM, et al. (2006) The FERM protein Yurt is a negative regulatory component of the Crumbs complex that controls epithelial polarity and apical membrane size. Dev Cell 11: 363–374. 10.1016/j.devcel.2006.06.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Glowinski C, Liu RH, Chen X, Darabie A, Godt D (2014) Myosin VIIA regulates microvillus morphogenesis and interacts with cadherin Cad99C in Drosophila oogenesis. J Cell Sci 127: 4821–4832. 10.1242/jcs.099242 [DOI] [PubMed] [Google Scholar]
- 90.Nashchekin D, Fernandes AR, St Johnston D (2016) Patronin/Shot Cortical Foci Assemble the Noncentrosomal Microtubule Array that Specifies the Drosophila Anterior-Posterior Axis. Dev Cell 38: 61–72. 10.1016/j.devcel.2016.06.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Bassett A, Liu JL (2014) CRISPR/Cas9 mediated genome engineering in Drosophila. Methods 69: 128–136. 10.1016/j.ymeth.2014.02.019 [DOI] [PubMed] [Google Scholar]
- 92.Port F, Chen HM, Lee T, Bullock SL (2014) Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc Natl Acad Sci U S A 111: E2967–2976. 10.1073/pnas.1405500111 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(A) and (B) Par-6 (red) localises normally to the apical surface of crb8F105 (A) and baz4 (B) mutant cells (marked by GFP, green). (C) baz815–8 mutant cells (marked by GFP, green) form a normal apical brush border as revealed by phalloidin staining of F-actin (red). (D) crb11A22 MARCM clones (marked by GFP, green) show normal Mesh (red) and Cno (white) localisation. (E) sdtK85 MARCM clones (marked by GFP, green) show normal Arm (red) and Lgl (white) localisation. (F) baz4 MARCM clones (marked by GFP, green) show normal Mesh (red) and Cno (white) localization. (G) Unspecific signals of Myo7a staining (red) in nuclei are present in ck13 MARCM clones (marked by GFP, green). (H) aPKCk06043 MARCM clones show normal Cora (red) at SJs and βHSpec (white) at the apical domain. (I) aPKCk06043 MARCM clones show normal αSpec (red) localization at the cortex. (J) The copper cell region has similar polarity to other regions of the midgut, with pMoe (red) at the apical domain, although folded inside, and Dlg (green) at apical SJs. (K) ee cells, which are labelled with Pros (green in nucleus), have Tsp2a (red) at the apical SJ and Arm (green) at the lateral AJ. (L) Schematic genomic map showing the aPKCHC allele and corresponding protein sequence. Scale bars, 10 μm. AJ, adherens junction; aPKC, atypical protein kinase C; Arm, Armadillo; αSpec, α-Spectrin; Baz, Bazooka; βHSpec, βH-Spectrin; Cno, Canoe; Crb, Crumbs; Dlg, Discs large; EC, enterocyte; ee, enteroendocrine; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; MARCM, mosaic analysis with a repressible cell marker; Myo7a, Myosin 7a; pMoe, phospho-Moesin; Pros, Prospero; SJ, septate junction.
(TIF)
(A) Scrib (red) localises to the EC SJs, marked by Cora (green). (B) Mesh (red) localises normally to the SJs of lgl4 mutant cells (marked by GFP, green). (C) dlg114 MARCM clones (marked by GFP, green) show normal apical localization of α-Spectrin (red) and βH-Spectrin (white). (D) RNAi knock-down of Scrib in adult midgut ECs has no effect on the subcellular localization of Myo7a (red) but disrupts Dlg (green) and Lgl (white) localisation to the SJs. Scale bar, 10 μm. Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; Lgl, Lethal (2) giant larvae; MARCM, Mosaic analysis with a repressible cell marker; Myo7a, Myosin 7a; RNAi, RNA interference; Scrib, Scribbled; SJ, septate junction.
(TIF)
(A) Par-1 is not detectable in ECs, although it is expressed in ISCs, as revealed by staining for GFP (green) in a Par-1 GFP protein trap line. Dlg is in red and aPKC in white. (B) A MARCM clone of par-19 (marked by GFP, green). The mutant cells form normal SJs marked by Lgl (red). Scale bar, 10 μm. aPKC, atypical protein kinase C; Dlg, Discs large; EC, enterocyte; GFP, green fluorescent protein; ISC, intestinal stem cell; Lgl, Lethal (2) giant larvae; MARCM, mosaic analysis with a repressible cell marker; SJ, septate junction.
(TIF)
(A) Most rhea, Fit1, Fit1 Fit2, and Tsp2a mutant cells remain inside the epithelia layer. The graph is based on the analysis of 299 cells in wild-type MARCM clones (13 images), 1,205 Tsp2a mutant cells (25 images from Tsp2a1–2, Tsp2a 3–3, and Tsp2a 2–9 clones), 608 Fit118 Fit283 (FitD) double mutant cells (23 images), 854 Fit118 mutant cells (24 images), and 184 rhea mutant cells (18 images from rheaB28, rhea79a, and rhea B128). The underlying data can be found in S1 Data. (B) Fit118 Fit283 double mutant cells (marked by GFP, green) differentiate as ECs, as revealed by the expression of Pdm1 (white). Talin is in red. (C) A meshf04955 mutant clone (marked by the loss of GFP) stained for α–Spectrin (red) and Mesh (white). (D) Tsp2a1–2 mutant cells (marked by GFP, green) differentiate as ECs, as revealed by the expression of Pdm1 (red). White asterisks * and lines mark the mutant clones. Scale bars, 10 μm. EC, enterocyte; GFP, green fluorescent protein; MARCM, mosaic analysis with a repressible cell marker; SJ, septate junction.
(TIF)
Images were taken of different regions of several midguts containing MARCM clones stained with an apical marker. Cells that were above the neighbouring cells or had a clear apical domain were counted as ‘cells NOT inside the layer’, whereas cells without a detectable free apical surface were counted as ‘cells inside the layer’. For each image, the percentage of clones inside the layer among all clones was calculated. Data were analysed with Graphpad Prism software. The graph in S4A Fig shows the average percent of cells inside the layer ± SD%.
(XLSX)
Data Availability Statement
All relevant data are within the paper and its Supporting Information files.