Abstract
The large-scale chemical spill on January 9, 2014 from coal processing and cleaning storage tanks of Freedom Industries in Charleston affected the drinking water supply to 300,000 people in Charleston, West Virginia metropolitan, while the short-term and long-term health impacts remain largely unknown and need to be assessed and monitored. There is a lack of publically available toxicological information for the main contaminant 4-methyl-1-cyclohexanemethanol (4-MCHM). Particularly, little is known about 4-MCHM metabolites and their toxicity. This study reports timely and original results of the mechanistic toxicity assessment of 4-MCHM and its metabolites via a newly developed quantitative toxicogenomics approach, employing proteomics analysis in yeast cells and transcriptional analysis in human cells. These results suggested that, although 4-MCHM is considered only moderately toxic based on the previous limited acute toxicity evaluation, 4-MCHM metabolites were likely more toxic than 4-MCHM in both yeast and human cells, with different toxicity profiles and potential mechanisms. In the yeast library, 4-MCHM mainly induced chemical stress related to transmembrane transport and transporter activity, while 4-MCHM metabolites of S9 mainly induced oxidative stress related to antioxidant activity and oxidoreductase activity. With human A549 cells, 4-MCHM mainly induced DNA damage-related biomarkers, which indicates that 4-MCHM is related to genotoxicity due to its DNA damage effect on human cells and therefore warrants further chronic carcinogenesis evaluation.
Graphical Abstract

■ INTRODUCTION
On January 9, 2014, approximately 10,000 US gal (38,000 L) of chemicals used in coal processing and cleaning leaked from the storage tanks of Freedom Industries, about 1.5 miles upstream from West Virginia American Water’s drinking water intake, affecting 300,000 people in Charleston, West Virginia metropolitan area.1 By the evening of January 10, nearly 700 residents had contacted West Virginia’s poison control center, reporting a range of symptoms including nausea and rashes.2 Detectable 4-methyl-1-cyclohexanemethanol (4-MCHM) was reported in house tap water samples and WV American Water effluent.3–5 West Virginia-American Water Company (WVAW) and the State issued water ban and restriction of water use for a period following the spill. In March 2014, 4-MCHM was found to be absorbed on the activated carbon filters and onto plastic water pipes; therefore, it likely continued to be released into water over a relatively long period of time after the spill.4
According to information provided by Freedom Industries, the leaked tank contained 88.5% crude MCHM, 7.3% PPH Stripped basic, and 4.2% water. The crude MCHM is a mixture containing the primary component 4-MCHM at 68–89% (w/w) which exists as two diastereomers, cis-4-MCHM [3937-48-2] and trans-4-MCHM [3937-49-3] according to the Material Safety Data Sheet (MSDS).6,7 There are a number of other chemicals in the crude MCHM, including 4-(methoxymethyl) cyclohexanemethanol, methyl 4-methylcydohexanecarboxylate (MMCHC; 5%), dimethyl-1,4-cyclohexanedicarboxylate (1%), and 1, 4-cyclohexanedimethanol (1–2%) that all exist as cis- and trans-isomers.5,8 The material PPH Stripped is reported to be primarily composed of dipropylene glycol phenyl ether (DiPPH) and propylene glycol phenyl ether (PPH).8 Given the small percentage of PPH, DiPPH, and other chemicals in the tank, as well as their relatively lower reported toxicity (e.g., LD50 > 2000 mg/kg for PPH versus 825 mg/kg for crude MCHM, 707–1768 mg/kg for pure 4-MCHM in different reports),6,9 we focused on the toxicity of the main pollutant 4-MCHM in this study.
4-Methyl-1-cyclohexanemethanol (4-methylcyclohexanemethanol, 4-MCHM) is an organic compound classified as a saturated higher alicyclic primary alcohol. It has been patented for use in air fresheners10 and a frothing agent for coal processing and cleaning.11,12 There is a lack of publically available toxicological information for 4-MCHM. A limited number of reports, including the report released by Eastman Chemical Co. and several recent studies after the spill, indicated that the crude MCHM is moderately toxic to aquatic organisms based on acute study with fathead minnow and Daphnia magna,9,13 has no severe genotoxicity with the Ames test,9 has relatively low systematic toxicity to mammalian species based on receptor studies,9 and can cause moderate skin and eye irritation in rabbits.9 A recent study suggested that the crude MCHM may be more toxic to Daphnia magna than previous reported.13 A toxicity study of pure (distilled) 4-MCHM focused on acute single dose oral and dermal toxicity in rats, dermal irritation and skin sensitizing in guinea pig, eye irritation in rabbits, and a 28-day oral feeding study in rats.9 According to these studies, pure MCHM is moderately toxic, and the effect focused on skin and eye irritation. Several recent investigations of The National Toxicology Program (NTP) are ongoing for more toxicological data for pure 4-MCHM. A structure—activity relationship (SAR) analysis suggested that 4-MCHM might result in effects on the development of offspring and might be an eye and skin irritant.14 The initial update of cellbased HTS (high throughput screening) assays suggested that 4-MCHM “exhibited no biological activity toward the evaluated subset of cellular and molecular targets of toxicological concern”.15 Maternal toxicity and decreased fetal weight of offspring were observed in a rat prenatal developmental toxicity study.16 Centers for Disease Control and Prevention (CDC) determined no adverse effect-concentration (NOAEC) of 1 ppm in drinking water based on a 28-day rats study.6,17 Another independent toxicologist determined a 0.120 ppm of NOAEC based on the same data CDC reviewed with inhalation and dermal exposures.18 Recently, the EPA announced a healthbased 30-day air screening level of 0.010 ppm for 4-MCHM.5,19
However, little is yet known about 4-MCHM metabolites and their toxicity. A WHO study of alicyclic primary alcohols suggested that 4-MCHM would primarily be metabolized to its corresponding carboxylic acid: 4-methylcyclohexanecarboxylic acid (CAS 13064-83-0, a naphthenic acid).20 Naphthenic acids have been reported as major contaminants in water used for extraction of oil from tar sands. Both acute and chronic toxicity, including hepatotoxicity to fish and other organisms, has been reported for naphthenic acids mixtures. 21,22 However, the potential toxicity effects of 4-methylcyclohexanecarboxylic acid, especially on human health, are still unknown. In summary, previous studies provide a limited understanding of 4-MCHM and its metabolites’ toxicology profiles. Timely and mechanistic toxicity screening and assessment of 4-MCHM and its metabolites, as pointed out by the recent report of Whelton et al., is in urgent need to provide critical information for evaluating the short-term and long-term health impacts of this chemical spill on the affected people, which largely remain unknown and need to be assessed and monitored.
This study reports timely results of mechanistic toxicity assessment of 4-MCHM and its metabolites via a newly developed quantitative toxicogenomics approach, employing proteomics analysis in yeast cells and transcriptional analysis in human cells. Yeast has been commonly used for toxicity assessment because it shares a lot of functional homologues with mammalian cells.23 The yeast genome is fully characterized, and the primary biological functions are conserved, which makes it a suitable model for toxicology.24–27 The stress response ensemble-based toxicogenomics assay with GFP-fused yeast cell library has been recently proposed and demonstrated by our group.27,28 The selected stress response genes are present and highly conserved in most cell types of metazoans and are activated at significantly lower toxicant concentrations than those causing overt cellular injury,29,30 which make them suitable to reveal potential toxicity mechanisms comprehensively. Specific toxicity and stress response-based studies have been used by both our group28,31 and others, with specific focus on oxidative damage,32,33 DNA damage,34 and inflammation and apoptosis,35,36 to provide important insights into the toxicological potential elicited by chemicals. These in vitro high throughput molecular assays provide fast yet informative results for initial toxicity screening and risk identification to guide further extensive toxicity evaluation, as needed for the case of the 4-MCHM chemical spill. Proteomics analysis of 148 proteins involved in known cellular response pathways in yeast cells was performed using GFP-tagged yeast reporters’ library.27,28 In addition, toxicity in human lung epithelial cells was also conducted, using gene expression analysis of 12 key biomarkers via RT q-PCR Although 4-MCHM has been considered moderate for acute toxicity based on limited data, our molecular assays provide more subcytotoxic mechanistic toxicity information on 4-MCHM and its metabolites mixture, to help better understand its potential chronic effects. The results provide useful information for guiding further 4-MCHM toxicity study, public health protection strategy, and regulation formation.
METHODS
Cytotoxicity of 4-MCHM in Yeast and Human Lung Epithelial Cells.
24 h cytotoxicity of 4-MCHM (product M1412, 98.0% (GC), CAS Number 34885-03-5, cis and trans mixture) was obtained from TCI America (Portland, OR), and its metabolites (4-MCHM mixed with 1% rat liver extraction (S9 fraction, Sprague–Dawley Rat, Invitrogen, NY, US)) were tested in yeast S.cerevisiae (ATCC 201388) and human lung epithelial cells (A549), respectively. For yeast cells, cytotoxicity was evaluated by growth inhibition in yeast for 24 h in triplicates. For human lung epithelial A549 cells, 2 × 104/well of cells was seeded in 96-well plates (Costar) for 24 h to reach continuous monolayer growth in F12 medium with 10% fetal bovine serum (FBS). 4-MCHM prepared in F12 medium with 1% FBS with or without S9 was added in PBS-washed cells for 24 h in triplicates. The exposed cells were fixed and stained in a 20% methanol/0.2% crystal violet solution, washed by distilled water, and then extracted by lysis buffer (0.9 g of sodium citrate and 1.25 mL of 1 N HCl in 98 mL of 47.5% ethanol). The cell amount was quantified via Micro plate Reader (Synergy H1Multi-Mode, Biotech, Winooski, VT) as OD630.37,38
Proteomics Analysis in Yeast Stress Response Ensemble Library.
A library of 148 in-frame GFP fusion proteins involved in known cellular stress response pathways of S.cerevisiae (Invitrogen, no. 95702, ATCC 201388) (Table S1, Supporting Information), constructed by oligonucleotidedirected homologous recombination to tag each open reading frame (ORF) with Aequorea victoria GFP (S65T) in its chromosomal location at the 3’ end,39 was employed according to our previous publications.27,28 The biomarkers covered key proteins involved in general stress response, oxidative stress, chemical stress, protein stress, and DNA stress. Yeast strains selected were grown in clear bottom black 384-well plates (Costar) with SD medium (60 μL per well) for 4–6 h at 30 °C to reach early exponential growth (OD600 about 0.2 to 0.4). Six-log concentrations of 4-MCHM stock solution (70 μg/L to 7 g/L, 7× stock) dissolved in PBS with or without S9 or control (PBS with or without 1% S9) based on the maximum noncytotoxic concentration (over 95% cell survival tested by growth inhibition in yeast for 24 h as shown in Table 2) were added at 10 μL per well to obtain the final concentrations (10 μg/L to 1 g/L). For treatments with S9, 4-MCHM or PBS was preincubated with S9 for 6 h at 37 °C to be consistent with human cell exposure. The plates were then placed in a Micro plate Reader (Synergy H1Multi-Mode, Biotech, Winooski, VT) for absorbance (OD600 for cell growth) and GFP signal (filters with 485 nm excitation and 535 nm emission for protein expression) measurements every 5 min for 2 h after a fast shake for 1 min. All experiments were performed in triplicate.
Table 2.
Cytotoxic Endpoints (IC50 and IC5) of 4-MCHM and 4-MCHM Metabolites (4-MCHM+S9) in Yeast (S.cerevisiae (ATCC 201388)) and Human Lung Epithelial Cells (A549)a
| IC50 (mg/L) | IC5 (mg/L) | R2 (p-value) | hill slope | |
|---|---|---|---|---|
| Yeast (24 h) | ||||
| 4-MCHM | 4702 ± 1.3 | 453.3 ± 2.2 | 0.9472 (0.0052) | −1.31 ± 0.35 |
| 4-MCHM with 1%S9 | 1722 ± 1.2 | 390.9 ± 1.3 | 0.9898 (<0.0001) | −1.96 ± 0.56 |
| Human Cells (24 h) | ||||
| 4-MCHM | 148.7 ± 1.2 | 11.2 ± 1.8 | 0.9827 (<0.0001) | −1.14 ± 0.24 |
| 4-MCHM with 1%S9 | 55.7 ± 1.4 | 0.2 ± 3.0 | 0.9502 (0.0009) | −0.50 ± 0.10 |
IC50 and IC5 values are determined via Four Parameter Logistic nonlinear regression model fitting of cell survival assay in Figure 1.
Data Processing and Analysis for Proteomics Assay in Yeast Cell Library.
Data Processing.
A detailed description of data processing for the yeast reporters library assay was provided in our previous report.28 Briefly, OD and GFP raw data from the measurements were first corrected by subtracting the background OD and GFP signal of the medium control (with or without chemicals) and referred to as ODcorrected and GFPcorrected. The protein expression for each measurement was then normalized by cell number (ODcorrected) as P = GFPcorrected/ODcorrected. Then, the P level was further normalized among replicate plates based on mean PPGK1 of medium control on each plate. The alteration in expression for a given protein at each time point due to chemical exposure, also referred as induction factor I, was represented as I = Pe/Pc, where Pe was defined as corrected and normalized P in the experiment condition with chemical exposure and Pc was defined as corrected and normalized P in the vehicle control condition without any chemical exposure.
Molecular Endpoint Derivation.
To quantifiy the chemical-induced protein expression level effect, the real-time protein expression profile was then integrated into the Protein Effect Level Index (PELI) in a similar manner as the Transcriptional Effect Level Index (TELI) proposed for transcriptomics analysis in our previous studies.27,28,40 On the basis of the function and nature of the proteins selected in this study that are essential in yeast cellular stress responses (Table S1, Supporting Information), the extent of up-regulation reflecting yeast cellular stress responses is quantified as follows.27,28,40
For protein (ORF) i, the protein effect level was quantified as the accumulative protein expression changes averaged over a 2 h exposure time as follows:
| (1) |
where t was exposure time in hours. Ii = I when I ≥ 1.0 (up-regulation); Ii = 1.0 when I < 1.0 (down-regulation).
An assay noise cutoff treshold was determined as 1.5 for PELIORF, on the basis of 2SD (the average SD for PELIORFi in our yeast assay) and in reference to literature.23,27,28 All tests were conducted in triplicates.
PCA Analysis.
Principal component analysis (PCA) was performed for the toxicogenomics data for both 4-MCHM and 4-MCHM metabolites (+S9) to simplify the complex data sets of categories by analyzing the components with the greatest amount of variance based on their protein expression profiles in yeast. The analysis was conducted on the basis of protein expression levels of each chemical (ln I, average of triplicates) during a 2 h exposure by software suite MeV (MutiExperiment Viewer) v4.8,41 with centering mode as mean and number of neighbors for KNN imputation as 10.
Gene Set Enrichment Analysis.
Stress category enrichment analysis was computed using the original Gene Set Enrichment Analysis (GSEA) algorithm by Subramanian et al.,42 based on mean PELIORF values for all the treatments of 4-MCHM and 4-MCHM+S9 with multiple doses. The proteins in each stress category from Table S1, Supporting Information, were used to define the stress response gene sets.
Gene Ontology.
Gene ontology (GO) analysis was performed with novel Network Ontology Analysis method (NOA) for enrichment analysis to determine the significantly represented GO biological categories and to analyze functions of gene network, as it allows enrichment analysis with a user defined reference set.43 In this study, the stress library was used as the reference set, and activated ORFs were used as test sets, which were defined as mean PELIORF > 1.5 based on signal-to-noise ratio as described in the Data Processing section. A p-value threshold of 0.01 was used for significance. GO analysis was performed for the concentration that had the most proteins (ORFs) activated at 10 mg/L for 4-MCHM and 100 mg/L for 4-MCHM+S9, respectively.
Transcriptional Analysis of Key Stress Biomarkers in Human Cells.
Gene expression level changes of key genes involved in cellular stress and toxicity responses including inflammation, apoptosis, DNA damage, and oxidative damage (Table 1)31 were analyzed in A549 cells at subcytotoxic doses (>95% of cells were alive based on 24 h cytotoxic tests, details in Figure 1). 2 × 105/well of human lung epithelial A549 cells were seeded in 6-well plates (Costar) for 24 h to reach a continuous monolayer in F12 medium with 10% FBS. 4-MCHM (final concentrations at 1, 10, and 100 mg/L) prepared in F12 medium with 1% FBS or 4-MCHM with 1%S9 (final concentration at 0.1, 1, and 10 mg/L) were added in PBS-washed cells, and F12 medium with 1% FBS with or without S9 was added to reach a final volume of 2 mL in each well in triplicate for 6 h. RNA was then extracted from exposed cells using a one step RNA reagent (BS410A, Bio Basic Inc., Canada) and reverse transcribed to cDNA with a Verso cDNA Synthesis Kit (Thermo Scientific, US). Q-PCR was performed using SYBR Green Supermix (Bio-Rad, US) on the iQ5 Multicolor Real-Time PCR detection system (Bio-Rad, US). PCR primers targeting the selected genes indicating inflammation, apoptosis, DNA damage, and oxidative damage were obtained on the basis of literature review and the NCBI database as reported previously (Table S2, Supporting Information),31 with housekeeping genes GAPDH and 18s RNA used as internal control. The changes in gene expression were normalized by the geometric average of GAPDH and 18s RNA based on multiple internal control normalization.44 The fold change that reflects relative gene expression change due to treatment compared to vehicle control, also referred to as induction factor I, was determined by the comparative CT method (2−ΔΔCT).45 All experiments were performed in triplicate.
Table 1.
List of Genes Involved in Cellular Stress and Toxicity Responses for Human Cells in This Study31
| stress type | gene |
|---|---|
| inflammation | TNF-α, IL-1β, NFKB1, NFKB2 |
| apoptosis | P53, CASP3, CASP8 |
| DNA damage (double strand break) | KU70, RAD51 |
| oxidative damage | HO1, SOD1, GPX1 |
| internal control | GAPDH, 18S rRNA |
Figure 1.
Dose responses based on a 24 h cytotoxicity (survival) assay of 4-MCHM and its metabolites (4-MCHM+S9) in yeast (A) and human A549 cells (B). X-axis: chemical concentration (mg/L); Y-axis: percentage of surviving cells compared to vehicle control. Mean ± SD, n = 3.
■ RESULTS
Cytotoxicity of 4-MCHM and Its Metabolites.
The potential acute toxicity of 4-MCHM and its metabolites with rat liver extract (S9 fraction) was evaluated by a 24 h cytotoxicity assay quantifying the survival percentage of yeast S.cerevisiae (ATCC 201388) and human lung epithelial cells (A549) (Figure 1). 4-MCHM metabolites (4-MCHM+S9) were more toxic than 4-MCHM only, for both yeast and A549 cells, although they were more pronounced in human cells, as indicated by the lower IC50 and IC5 values of 4-MCHM metabolites than that of the original 4-MCHM (Table 2). This difference indicated potential generation of more toxic metabolites by S9 than parent 4-MCHM. Human cells were more sensitive to the toxicity of 4-MCHM and its metabolites than yeast cells with much lower IC50 and IC5 values than those for yeast cells.
Distinct Toxicity Profiles of 4-MCHM and Its Metabolites in Yeast Assay.
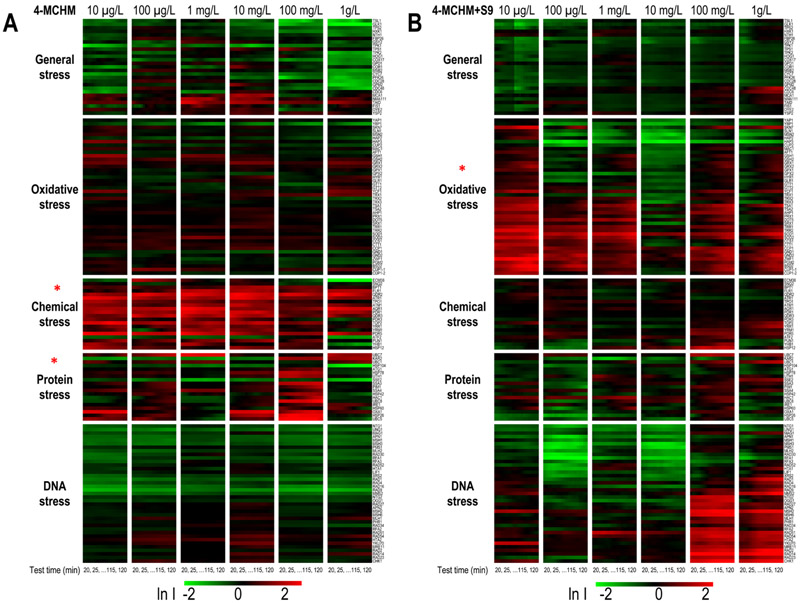
The real-time differential protein expression profiles of 148 proteins in yeast indicative of cellular stress pathway activities were distinctive for 4-MCHM and its metabolites (Figure 2), suggesting compound(s)-specific cellular responses as results of their different toxicity mechanisms. The temporal protein expression profiles were dose dependent, and protein activation shift in response to dose concentrations has been reported previously.27 For 4-MCHM exposure, strong chemical stress was induced for all the six concentrations and protein stress responses were observed at concentration higher than 100 mg/L. In comparison, the toxicity profile for 4-MCHM metabolites (+S9) showed strong oxidative stress induction indicating oxidants generation from the S9-mediated metabolism of 4-MCHM. The metabolites mixture seemed to contain DNA-damaging chemicals since activation of DNA damage and repair-relevant pathways were observed at higher dose concentrations of 100 mg/L and 1 g/L.
Figure 2.
Temporal altered protein expression profiles of 148 biomarkers indicative of different stress responses upon exposure to 4-MCHM (A) and 4-MCHM metabolites (4-MCHM+S9) (B) across six-log concentrations. The mean natural log of induction factor (ln I) indicates the magnitude of altered protein expression (represented by a green–black–red color scale at the bottom. Red spectrum colors indicate up-regulation; green spectrum colors indicate down-regulation. Values beyond ±2 are shown as ±2). X-axis top: concentrations for each chemical; X-axis bottom: testing time in minutes; the first data point shown is at 20 min after exposure due to data smoothing with a moving average of every five data points. Y-axis left: clusters of proteins by stress response pathways; Y-axis right: list of proteins (ORFs) tested, with details in Table S1, Supporting Information. n = 3. The stress category is labeled with “*” for those with FDR < 25% for gene set enrichment analysis (the statistical significance of the enrichment score for a single set).
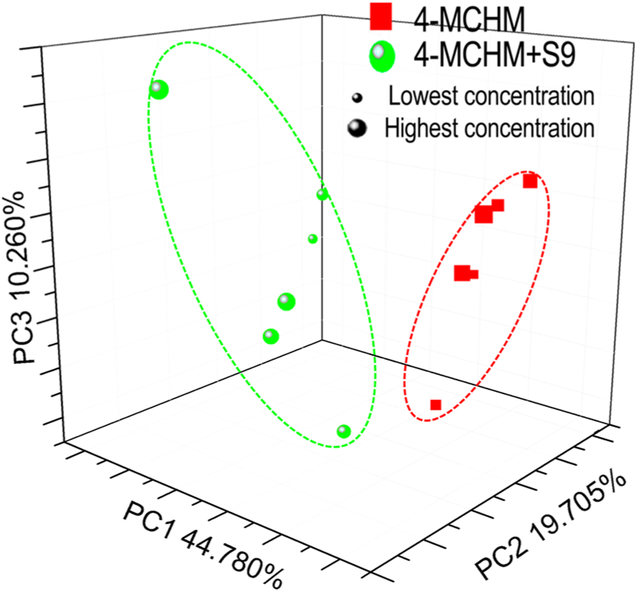
The chemical-specific and concentration-dependent real time protein expression profiles of 4-MCHM and its metabolites may serve as fingerprints to allow chemicals clustering based on their specific toxicity mechanisms. The principle component analysis (PCA) (Figure 3) also showed the clear separation of 4-MCHM and its metabolites based on their altered stress response protein expression profiles, suggesting that 4-MCHM was likely transformed by S9 into other metabolites with different toxicity mechanisms from those exhibited by 4-MCHM, as discussed in more depth in the later section.
Figure 3.
PCA graphical representations of the three major components of protein expression change profiles for 4-MCHM (red square) and 4-MCHM with S9 (green sphere), based on induction factor (ln I) during a 2 h exposure across six-log concentrations. Samples are color coded according to chemicals. Each square or sphere represents one treatment (a chemical at a given concentration) with symbol size indicating the relative level of concentration.
Insights into the Toxicity Mechanism of 4-MCHM and Its Metabolites.
To further analyze and compare the toxicity responses and potential mechanisms of 4-MCHM and its metabolites, stress category responses were analyzed using a gene set enrichment analysis (GSEA) algorithm, and the proteins were ranked on the basis of their PELIORF values (Table 3). For 4-MCHM, the protein set related to “chemical stress”, including transporters and proteins involved in the membrane, cell wall, and cell structure, was significantly enriched compared to the untreated control with FDR = 0 (false discovery rate, FDR < 25% is defined as significant enrichment). Protein stress related to unfolded protein responses and autophagy to turn over misfolded or damaged proteins was also significantly enriched with FDR = 4.22%. This suggested that the toxicity mechanism of 4-MCHM is likely related to chemical transport, membrane damage, and protein stress. In comparison, for 4-MCHM+S9, the oxidative stress was significantly enriched with changes in biomarkers indicative of oxidant regulation and defense systems of oxidative stress responses (FDR = 0). The 4-MCHM metabolites resulting from S9 transformation likely contained oxidizing chemicals that induce oxidative stress.
Table 3.
Summary of GSEA Results for 4-MCHM and 4-MCHM+S9 Based on Toxicogenomics Assay in Yeast Cells
| stress category |
gene set size |
enrichment score (ES) |
normalized enrichment score (NES) |
nominal p-value |
FDR-q valuea |
|---|---|---|---|---|---|
| 4-MCHM | |||||
| chemical | 23 | −0.8132 | −1.7474 | 0 | 0 |
| protein | 19 | −0.6216 | −1.3134 | 0.05305 | 0.0422 |
| oxidative | 44 | −0.3586 | −0.8076 | 0.906 | 0.8715 |
| general | 30 | −0.2780 | −0.6088 | 0.995 | 0.7950 |
| 4-MCHM+S9 | |||||
| oxidative | 44 | −0.6979 | −1.9776 | 0 | 0 |
| protein | 19 | −0.3975 | −1.0379 | 0.4326 | 1 |
| DNA | 38 | −0.3392 | −0.9559 | 0.5872 | 0.9494 |
| general | 30 | −0.3012 | −0.8229 | 0.7887 | 0.9866 |
| chemical | 23 | −0.2347 | −0.6268 | 0.9589 | 0.9649 |
FDR: false discovery rate, indicative of probability that a gene set with a given NES represents a false positive finding. FDR < 25% is defined as significant enrichment according to GSEA software, and FDR of 25% indicates that the result is likely to be valid 3 out of 4 times.42
In addition to the stress response categories-based GSEA, representative GO categories-based enrichment analysis was also performed for the concentration that had the most ORFs activated at 10 mg/L for 4-MCHM and 100 mg/L for 4-MCHM+S9, respectively, in order to identify the significantly overrepresented biological functional categories among these activated ORFs, with the whole stress library selected as the reference (Table 4). The results indicated that the main biological process induced by a 10 mg/L 4-MCHM exposure was related to transmembrane transport, while 100 mg/L 4-MCHM+S9 mainly induced cellular responses to stimulus and stress, including DNA stress. Many genes (see Table 4 for details) related to transporter activity were induced by 10 mg/L 4-MCHM exposure, which was consistent with the GSEA results for gene sets of stress response categories, which indicated chemical stress as the significantly enriched stress category. For 100 mg/L 4-MCHM+S9, the main induced molecular function was mostly related to antioxidant activity and oxidoreductase activity, which was also consistent with previous stress response GSEA results that oxidative stress was significantly enriched for 4-MCHM+S9. In summary, the two independent gene enrichment analysis of yeast cell assays suggested that 4-MCHM mainly induced chemical stress related to transmembrane transport and transporter activity, while 4-MCHM metabolites of S9 mainly induced oxidative stress and DNA stress.
Table 4.
Overrepresented (p-value <0.01) Biological Categories Based on GO Databases for a 2 h Exposure with 4-MCHM (10 mg/L) and 4-MCHM+S9 (100 mg/L) in Yeast, Using the Stress Library Serving as a Reference Set
| category | p-value | names of observed genes in categorya | number of genes observed in category |
number of genes in category |
|---|---|---|---|---|
| 4-MCHM 10 mg/L | ||||
| GO Biological Process | ||||
| drug transport [GO:0015893] | 9.20 × 10−5 | YRR1; YRM1; TPO1; AQR1; PDR5; FLR1; QDR3; ATR1; QDR2 | 9 | 10 |
| drug transmem-brane transport [GO:0006855] | 1.60 × 10−4 | YRR1; YRM1; AQR1; FLR1; QDR3; ATR1; QDR2 | 7 | 7 |
| transmembrane transport [GO:0055085] | 0.002 | HSP78; YRR1; YRM1; TPO1; AQR1; FLR1; ATM1; BPT1; QDR3; ATR1; QDR2 | 11 | 17 |
| GO Cellular Component | ||||
| plasma membrane [GO:0005886] | 1.20 × 10−4 | CDC6; TPO1; PUN1; HSP12; AQR1; PDR5; FLR1; QDR3; ATR1; QDR2 | 10 | 12 |
| integral to membrane [GO:0016021] | 0.0012 | ECM38; YSP2; TPO1; PUN1; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; ATR1; OXA1; QDR2 | 13 | 21 |
| intrinsic to membrane [GO:0031224] | 0.0012 | ECM38; YSP2; TPO1; PUN1; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; ATR1; OXA1; QDR2 | 13 | 21 |
| membrane [GO:0016020] | 0.002 | CDC6; TRX1; ECM38; YSP2; COR1; TPO1; PUN1; HSP12; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; | 17 | 32 |
| membrane part [GO:0044425] | 0.0053 | ATR1; OXA1; QDR2 ECM38; YSP2; COR1; TPO1; PUN1; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; ATR1; OXA1; QDR2 |
14 | 26 |
| GO Molecular Function | ||||
| transporter activity [GO:0005215] | 3.60 × 10−5 | COR1; TPO1; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; ATR1; OXA1; QDR2 | 11 | 13 |
| transmembrane transporter activity [GO:0022857] | 1.20 × 10−4 | COR1; TPO1; AQR1; PDR5; FLR1; ATM1; BPT1; QDR3; ATR1; QDR2 | 10 | 12 |
| drug transmembrane transporter activity [GO:0015238] | 0.001 | TPO1; AQR1; PDR5; FLR1; QDR3; ATR1; QDR2 | 7 | 8 |
| substrate-specific transporter activity[GO:0022892] | 0.001 | COR1; TPO1; AQR1; BPT1; ATR1; OXA1; QDR2 | 7 | 8 |
| substrate-specific transmembrane transporter activity [GO:0022891] | 0.0033 | COR1; TPO1; AQR1; BPT1; ATR1; QDR2 | 6 | 7 |
| endopeptidase activity [GO:0004175] | 0.0077 | PIM1; COR1; NMA111; MCA1 | 4 | 4 |
| peptidase activity, acting on Lamino acid peptides [GO:0070011] | 0.0077 | PIM1; COR1; NMA111; MCA1 | 4 | 4 |
| peptidase activity [GO:0008233] | 0.0077 | PIM1; COR1; NMA111; MCA1 | 4 | 4 |
| 4-MCHM+S9 100 mg/L | ||||
| GO Biological Process | ||||
| response to stimulus [GO:0050896] | 5.60 × 10−5 | RAD4; UTH1; RAD34; GPX2MAG1; HSP60; GSH1; TRR2; GRX2; GRX1; CUP1-2; TRX2MMS2; PRX1;DOT5; RAD9; PGM2; ATF2; HAC1; YCF1; UBC1; SKN7MLH1; YHB1; PDR5; BSD2; TRR1; TSA2; SOD2; PHR1; YKU70; CUP1-1; AHP1; ZWF1; GND1; SOD1; TSA1; CCP1; APN2; UBC7; RFA2MRE11; HTA2; RAD27; RAD23; OGG1; HSP12; RAD51; NTG2; RAD2; RAD54; MSH6MSH2; CHK1; RAD14 | 55 | 106 |
| cellular response to stimulus [GO:0051716] | 5.90 × 10−4 | RAD4; RAD34; GPX2MAG1; TRR2; GRX2; GRX1; CUP1-2; TRX2MMS2; PRX1; DOT5; RAD9; HAC1;SKN7MLH1; TRR1; TSA2; PHR1; YKU70; CUP1-1; AHP1; ZWF1; GND1; TSA1; CCP1; APN2; RFA2MRE11; HTA2; RAD27; RAD23; OGG1; HSP12; RAD51; NTG2; RAD2; RAD54; MSH6MSH2;CHK1; RAD14 | 42 | 76 |
| response to stress [GO:0006950] | 7.80 × 10−4 | RAD4; UTH1; RAD34; GPX2MAG1; HSP60; GSH1; TRR2; GRX2; GRX1; CUP1-2; TRX2MMS2; PRX1;DOT5; RAD9; HAC1; UBC1; SKN7MLH1; YHB1; BSD2; TRR1; TSA2; SOD2; PHR1; YKU70; CUP1-1; AHP1; ZWF1; GND1; SOD1; TSA1; CCP1; APN2; RFA2MRE11; HTA2; RAD27; RAD23; OGG1; HSP12; RAD51; NTG2; RAD2; RAD54; MSH6MSH2; CHK1; RAD14 | 50 | 97 |
| GO Biological Process | ||||
| cellular response to stress [GO:0033554] | 0.0011 | RAD4; RAD34; GPX2MAG1; TRR2; GRX2; GRX1; CUP1-2; TRX2MMS2; PRX1; DOT5; RAD9; HAC1; SKN7MLH1; TRR1; TSA2; PHR1; YKU70; CUP1-1; AHP1; GND1; TSA1; CCP1; APN2; RFA2MRE11; HTA2; RAD27; RAD23; OGG1; HSP12; RAD51; NTG2; RAD2; RAD54; MSH6MSH2; CHK1; RAD14 | 41 | 75 |
| response to DNA damage stimulus [GO:0006974] | 0.0058 | RAD4; RAD34; MAG1MMS2; RAD9MLH1; PHR1; YKU70; TSA1; APN2; RFA2MRE11; HTA2; RAD27; RAD23; OGG1; RAD51; NTG2; RAD2; RAD54; MSH6MSH2; CHK1; RAD14 | 24 | 40 |
| cellular homeostasis [GO:0019725] | 0.0058 | MAC1; TRR2; GRX2; GRX1; TRX2; PRX1; DOT5; PGM2; TRR1; TSA2; AHP1; SOD1; TSA1 | 13 | 18 |
| homeostatic process [GO:0042592] | 0.0072 | MAC1; TRR2; GRX2; GRX1; TRX2; PRX1; DOT5; PGM2; YCF1; TRR1; TSA2; YKU70; AHP1; SOD1; TSA1; RFA2; RAD51; RAD54 | 18 | 28 |
| GO Molecular Function | ||||
| antioxidant activity [GO:0016209] | 0.0033 | GPX2; TRR2; GRX2; GRX1; CUP1-2; PRX1; DOT5; TRR1; TSA2; SOD2; CUP1-1; AHP1; SOD1; TSA1; CCP1 | 15 | 21 |
| oxidoreductase activity [GO:0016491] | 0.0039 | GPX2; TRR2; GRX2; GRX1; CUP1-2; TRX2; PRX1; DOT5; YHB1; TRR1; TSA2; SOD2; CUP1-1; GND2; AHP1; ZWF1; GND1; SOD1; TSA1; CCP1 | 20 | 31 |
Observed genes refer to genes up regulated with PELIORF over 1.5
Insights into Toxicity of 4-MCHM and Its Metabolites in Human Lung Cells.
Gene expression analysis of 12 selected toxicity and stress-response related genes in human lung epithelial cells (A549) was performed to investigate the potential adverse effect of 4-MCHM and its metabolites in human cells. Altered gene expression changes after a 6 h exposure to 4-MCHM and its metabolites (+S9) in human lung epithelial cells (A549) are shown in Figures 4 and 5. The distinctive gene expression change profiles of 4-MCHM and 4-MCHM+S9 showed a different toxicity nature between the two.
Figure 4.
Altered gene expression profiles of biomarkers indicative of toxicity and stress responses in human cells, upon a 6 h exposure to 4-MCHM only and 4-MCHM +1% S9, respectively, across three concentrations in human A549 cells. The mean natural log of induction factor (ln I) indicates the magnitude of altered gene expression (represented by a green–black–red color scale at the bottom. Red spectrum colors indicate up-regulation; green spectrum colors indicate down-regulation; values beyond ±2 are shown as ±2, values in Figure 5). X-axis top: concentrations tested; Y-axis left: clusters of genes by stress categories; Y-axis right: list of genes tested, with details of gene descriptions in Table 1. n = 3.
Figure 5.
Gene expression change in A549 cells after a 6 h exposure of 4-MCHM and 4-MCHM metabolites (4-MCHM+S9). RT-qPCR indicated gene alteration by fold change (Y-axis), compared to untreated control; *: I >1 with p < 0.05, n = 3.
In human cells, 4-MCHM exposure at 10 mg/L led to DNA damage, as indicated by the significant up-regulation of Rad51 (I = 7.1115, p = 0.0005) and Ku70 (I = 3.1457, p = 0.0299), two key genes involved in DNA double strand break repair. In contrast, 4-MCHM+S9 exposure at the same concentration (10 mg/L) led to significant up-regulation of genes mostly involved in apoptosis indicated by P53 (controlling cell cycle and apoptosis, I = 7.1630, p = 0.0498)46 compared to the untreated control, indicating higher toxicity of the metabolites mixture than the 4-MCHM. In summary, the stress gene expression in human cells suggested that 4-MCHM mainly induced DNA damage, while 4-MCHM metabolites of S9 seemed more toxic. The 4-MCHM metabolites of S9 affected biomarkers indicative of oxidative damage and inflammation at lower concentrations and affected apoptosis biomarkers at higher concentration.
■ DISCUSSION
This study reports the first investigation of cytotoxicity, molecular toxicity potential, and mechanisms of 4-MCHM and its metabolites by rat liver extraction. Previous toxicity reports of 4-MCHM focused on conventional in vivo endpoints, with limited mechanistic information reported. There has been no toxicity report about its metabolites. This study investigated cytotoxicity and molecular mechanistic toxicity profiles of the 4-MCHM and its metabolites using a toxicogenomics approach in two cell lines with yeast and a human epithelial cell line (A549) (relevant to respiratory exposures in humans), respectively.
This study shows that 4-MCHM metabolites were more toxic than 4-MCHM in both yeast and human cells, with different toxicity profiles and potential mechanisms. In the yeast library, 4-MCHM mainly induced chemical stress related to transmembrane transport and transporter activity, while 4-MCHM metabolites of S9 mainly induced oxidative stress related to antioxidant activity and oxidoreductase activity. With human A549 cells, 4-MCHM mainly induced DNA damage-related biomarkers, suggesting potential chronic effects that warrant further genotoxicity evaluation. This study also pointed out the need to consider metabolism for further 4-MCHM toxicity assessment.
On the basis of previous toxicology studies of alicyclic primary alcohols, to which 4-MCHM belongs, 4-MCHM was likely transformed by S9 into the corresponding aldehyde and carboxylic acid: 4-methylcyclohexanecarboxylic acid.20 Indeed, 4-methylcyclohexanecarboxylate was detected at drinking water taps by Foreman et al.8 Currently, there is no toxicity report for 4-methylcyclohexanecarboxylic acid. This study suggested that 4-methylcyclohexanecarboxylic acid or further metabolite(s) may have the potential to be more toxic and need further investigation.
Besides 4-MCHM metabolites, further assessment of possible transformation products from 4-MCHM through drinking water treatment is also necessary. The oxidative transformation pathways of organic contaminants are rather complex and dynamic, and the degradation intermediate mixtures can be even more toxic than the original contaminant(s).47,48 An oxidation transformation study of the spilled product (crude MCHM and stripped PPH) using free chlorine and KMnO4 oxidation did not detect any transformation products with the liquid–liquid extraction and GCMS method.3 However, the oxidation transformation process is rather complex and further investigation is needed to identify the transformation products.
It should be noted that this study focused on toxicity of pure 4-MCHM purchased from TCI America. Great differences existed among the spill, the crude MCHM, and pure 4-MCHM. The chemical spill at Elk River contains crude MCHM, PPH, and DiPPH. The crude MCHM contains other chemicals, which may lead to various toxicity responses. The crude MCHM also has a different ratio of cis- and trans-MCHM isomers compared to the standard 4-MCHM used in this study,8 with different odor thresholds.49 Isomers and their metabolites may have different activities, leading to different toxicity, which require further investigation. According to the previous study of the Eastman Company, crude MCHM seems to be less toxic than 4-MCHM with a weaker effect on dermal irritation and skin sensitization. The different toxicity of crude MCHM and pure 4-MCHM may be a result of multiple reasons, including other chemicals in crude MCHM, their interactions, and the difference of isomers.
In conclusion, this study revealed different toxicity and potential mechanisms of 4-MCHM and its metabolites by S9 in yeast and human cells (A549). These results suggested that, although 4-MCHM is considered only moderately toxic based on previous limited acute toxicity evaluation, its metabolites may be more toxic than 4-MCHM and are more relevant to human exposure. Our study at the molecular level revealed some subcytotoxic molecular mechanisms such as DNA damage potential, which indicates that 4-MCHM is related to carcinogenesis and reproductive toxicity due to its DNA damage effect on human cells. Our results suggested that longterm medical monitoring should be considered for the population. It may also provide insights into potential longterm aquatic toxicity issues. The toxicogenomics-based molecular toxicity screening assay employed in this study provides timely information regarding the underlying mechanisms of toxic action of 4-MCHM and its metabolites, especially related to low-dose and chronic exposures, which makes it a useful tool for public health protection and health monitoring needs.
Supplementary Material
ACKNOWLEDGMENTS
This study was supported by the National Science Foundation (NSF) (EEC-0926284, CAREER CBET-0953633, CBET-1440764) and the National Institute of Environmental Health Sciences (NIEHS) (PROTECT P42ES017198).
Footnotes
Supporting Information
List of proteins in stress response pathways ensemble yeast cell library; primers used for measuring gene expression by RT q-PCR in A549 cells. The Supporting Information is available free of charge on the ACS Publications website at DOI: 10.1021/acs.est.5b00371.
Notes
The authors declare no competing financial interest.
■ REFERENCES
- (1).Cooper WJ Responding to crisis: The West Virginia chemical spill. Environ. Sci. Technol 2014, 48 (6), 3095. [DOI] [PubMed] [Google Scholar]
- (2).Elk River Chemical Spill Health Effects Findings of Emergency Department Record Review, 2014; http://www.wvdhhr.org/Elk%20River%20Chemical%20Spill%20Health%20Effects%20-%20Findings%20of%20Emergency%20Department%20Record%20Review.pdf.
- (3).Rosen JS; Whelton AJ; McGuire MJ; Clancy JL; Bartrand T; Eaton A; Patterson J; Dourson M; Nance P; Adams C WV TAP Final Report; WV TAP, 2014.
- (4).Whelton AJ; Rosen JS; Clancy J; Clancy T; Ergul A The Crude MCHM Chemical Spill 10-Home Study: Tap Water Chemical Analysis, 2014; http://www.wvgazette.com/assets/PDF/CH619157.pdf.
- (5).Whelton AJ; McMillan L; Connell M; Kelley KM; Gill JP; White KD; Gupta R; Dey R; Novy C Residential tap water contamination following the Freedom Industries chemical spill: Perceptions, water quality, and health impacts. Environ. Sci. Technol. 2014, 49 (2), 813–823. [DOI] [PubMed] [Google Scholar]
- (6).Eastman Chemical Company. MSDS for Crude MCHM. http://www.eastman.com/Products/Pages/ProductHome.aspx?Product= 71014291&list=Chemicals.
- (7).Adams C; Whelton A; Rosen J Health Effects for Chemicals in 2014 West Virginia Chemical Release: Crude MCHM Compounds, PPH and DiPPH, 2014; https://s3.amazonaws.com/s3.documentcloud.org/documents/1088254/health-effects-lit-review-wvtap-15mar14-v1-4.pdf.
- (8).Foreman WT; Rose DL; Chambers DB; Crain AS; Murtagh LK; Thakellapalli H; Wang KK Determination of (4-methylcyclohexyl) methanol isomers by heated purge-and-trap GC/MS in water samples from the 2014 Elk River, West Virginia, chemical spill. Chemosphere 2014, DOI: 10.1016/j.chemosphere.2014.11.006. [DOI] [PubMed] [Google Scholar]
- (9).Eastman Chemical Company. Eastman Crude MCHM Studies; http://www.eastman.com/Products/Pages/ProductHome.aspx?Product=71014291&list=Chemicals.
- (10).Nagamura Y; Satoh Y; Tatsumi J; Yamamura K Producing method of alcohols such as cyclohexanedimethanol. U.S. Patent US6600080 B1, 2003.
- (11).Christie RD; Fortin RJ; Gross AE Process for coal flotation using 4-methyl cyclohexane methanol frothers. U.S. Patent US4915825 A, 1990.
- (12).Perkin WH; Pope WJ CIX. Experiments on the synthesis of 1-methylcyclo-hexylidene-4-acetic acid,[graphic omitted]. Part I. J. Chem. Soc., Trans 1908, 93, 1075–1085. [Google Scholar]
- (13).Whelton AJ You Can Make A Difference: University Students, Faculty, & Practitioners from the US & Abroad Provide Assistance Following the Elk River Spill; National Association of City and County Health Officials Conference, Atlanta, Georgia, 2014. [Google Scholar]
- (14).West Virginia Chemical Spill: Structure-Activity Relationship Analysis, December 2014. NTP Update; http://ntp.niehs.nih.gov/ntp/research/areas/wvspill/sar_ntp-wv_dec14_508.pdf.
- (15).West Virginia Chemical Spill: High Throughput Screening Assays, December 2014. NTP Update; http://ntp.niehs.nih.gov/ntp/research/areas/wvspill/ntp_wv_hts_dec2014_508.pdf.
- (16).West Virginia Chemical Spill: Prenatal Developmental Toxicity Evaluation, December 2014. NTP Update; http://ntp.niehs.nih.gov/ntp/research/areas/wvspill/ntp_wv_prenatal_dec2014_508.pdf.
- (17).CDC 2014. West Virginia Chemical Release; http://www.bt.cdc.gov/chemical/MCHM/westvirginia2014/.
- (18).Toxicological Excellence in Risk Assessment, Report of Expert Panel Review of Screening Levels for Exposure to Chemicals from the January 2014 Elk River Spill; 2014; http://www.tera.org/Peer/WV/WV%20Expert%20Report%2012%20May%202014.pdf.
- (19).Derivation of an Extrapolated Short-term Inhalation Screening Level for 4-Methylcyclohexanemethanol (MCHM – CAS# 34885-03-5). US EPA, Office of Research and Development, National Center for Environmental Assessment (NCEA), Superfund Health Risk Technical Support Center (STSC), Eds., Washington, D.C. USA, July 3, 2014; http://media.wvgazette.com/static/watchdog/EPA%20MCHM%20Air%20Screening%20Level.pdf. [Google Scholar]
- (20).Speijers G; Renwick A Alicyclic Primary Alcohols, Aldehydes, Acids, and Related Esters; Europe 8, (1.1), 0.019; http://www.inchem.org/documents/jecfa/jecmono/v50je10.htm.
- (21).Rogers VV; Wickstrom M; Liber K; MacKinnon MD Acute and subchronic mammalian toxicity of naphthenic acids from oil sands tailings. Toxicol. Sci 2002, 66 (2), 347–355. [DOI] [PubMed] [Google Scholar]
- (22).Allen EW Process water treatment in Canada’s oil sands industry: I. Target pollutants and treatment objectives. J. Environ. Eng. Sci 2008, 7 (2), 123–138. [Google Scholar]
- (23).Välimaa A-L; Kivistö A; Virta M; Karp M Real-time monitoring of non-specific toxicity using a Saccharomyces cerevisiae reporter system. Sensors 2008, 8 (10), 6433–6447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (24).Hughes T R Yeast and drug discovery. Funct. Integr. Genomics 2002, 2 (4–5), 199–211. [DOI] [PubMed] [Google Scholar]
- (25).Mager WH; Ferreira PM Stress response of yeast. Biochem. J. 1993, 290, 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (26).McGary KL; Park TJ; Woods JO; Cha HJ; Wallingford JB; Marcotte EM Systematic discovery of nonobvious human disease models through orthologous phenotypes. Proc. Natl. Acad. Sci. U.SA 2010, 107 (14), 6544–6549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).O’Connor ST; Lan J; North M; Loguinov A; Zhang L; Smith MT; Gu AZ; Vulpe C Genome-wide functional and stress response profiling reveals toxic mechanism and genes required for tolerance to benzo[a]pyrene in S. cerevisiae. Front. Genet 2012, 3, 316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Lan J; Gou N; Gao C; He M; Gu AZ Comparative and mechanistic genotoxicity assessment of nanomaterials via a quantitative toxicogenomics approach across multiple species. Environ. Sci. Technol. 2014, 48 (21), 12937–12945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (29).Simmons SO; Fan CY; Ramabhadran R Cellular stress response pathway system as a sentinel ensemble in toxicological screening. Toxicol. Sci 2009, 111 (2), 202–225. [DOI] [PubMed] [Google Scholar]
- (30).Kultz D Molecular and evolutionary basis of the cellular stress response. Annu. Rev. Physiol 2005, 67, 225–257. [DOI] [PubMed] [Google Scholar]
- (31).Khatri M; Bello D; Pal AK; Cohen JM; Woskie S; Gassert T; Lan J; Gu AZ; Demokritou P; Gaines P Evaluation of cytotoxic, genotoxic and inflammatory responses of nanoparticles from photocopiers in three human cell lines. Part. Fibre Toxicol. 2013, 10 (1), 42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (32).Quincozes-Santos A; Bobermin LD; Latini A; Wajner M; Souza DO; Goncalves CA; Gottfried C Resveratrol protects C6 astrocyte cell line against hydrogen peroxide-induced oxidative stress through heme oxygenase 1. PLoS One 2013, 8 (5), No. e64372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (33).Utomo A; Jiang X; Furuta S; Yun J; Levin DS; Wang YC; Desai KV; Green JE; Chen PL; Lee WH Identification of a novel putative non-selenocysteine containing phospholipid hydroperoxide glutathione peroxidase (NPGPx) essential for alleviating oxidative stress generated from polyunsaturated fatty acids in breast cancer cells. J. Biol. Chem 2004, 279 (42), 43522–43529. [DOI] [PubMed] [Google Scholar]
- (34).Lieber MR The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem 2010, 79, 181–211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (35).Fridman JS; Lowe SW Control of apoptosis by p53. Oncogene 2003, 22 (56), 9030–9040. [DOI] [PubMed] [Google Scholar]
- (36).Komiyama J; Nishimura R; Lee H-Y; Sakumoto R; Tetsuka M ; Acosta TJ; Skarzynski DJ; Okuda K Cortisol is a suppressor of apoptosis in bovine corpus luteum. Biol. Reprod 2008, 78 (5), 888–895. [DOI] [PubMed] [Google Scholar]
- (37).Saotome K; Morita H; Umeda M Cytotoxicity test with simplified crystal violet staining method using microtitre plates and its application to injection drugs. Toxicol. In Vitro 1989, 3 (4), 317–321. [DOI] [PubMed] [Google Scholar]
- (38).Liu W; Li H; Chen YH N-terminus of M2 protein could induce antibodies with inhibitory activity against influenza virus replication. FEMS Immunol. Med. Microbiol 2003, 35 (2), 141–146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (39).Huh WK; Falvo JV; Gerke LC; Carroll AS; Howson RW; Weissman JS; O’Shea EK Global analysis of protein localization in budding yeast. Nature 2003, 425 (6959), 686–691. [DOI] [PubMed] [Google Scholar]
- (40).Gou N; Gu AZ A new transcriptional effect level index (TELI) for toxicogenomics-based toxicity assessment. Environ. Sci. Technol 2011, 45 (12), 5410–5417. [DOI] [PubMed] [Google Scholar]
- (41).Saeed AI; Bhagabati NK; Braisted JC; Liang W; Sharov V; Howe EA; Li J; Thiagarajan M; White JA; Quackenbush J TM4 Microarray Software Suite. Methods Enzymol 2006, 411, 134–193. [DOI] [PubMed] [Google Scholar]
- (42).Subramanian A; Tamayo P; Mootha VK; Mukherjee S; Ebert BL; Gillette MA; Paulovich A; Pomeroy SL; Golub TR; Lander ES; Mesirov JP Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U.SA 2005, 102 (43), 15545–15550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (43).Wang JG; Huang Q; Liu ZP; Wang Y; Wu LY; Chen LN; Zhang XS NOA: A novel network ontology analysis method. Nucleic Acids Res. 2011, 39 (13), No. e87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (44).Vandesompele J; De Preter K; Pattyn F; Poppe B; Van Roy N; De Paepe A; Speleman F Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3 (7), RESEARCH0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (45).Schmittgen TD; Livak KJ Analyzing real-time PCR data by the comparative C-T method. Nat. Protoc 2008, 3 (6), 1101–1108. [DOI] [PubMed] [Google Scholar]
- (46).Polyak K; Xia Y; Zweier JL; Kinzler KW; Vogelstein B A model for p53-induced apoptosis. Nature 1997, 389 (6648), 300–305. [DOI] [PubMed] [Google Scholar]
- (47).Boxall ABA Transformation products of synthetic chemicals in the environment; Springer: Dordrecht; New York, 2009; p xiv, 249 pages. [Google Scholar]
- (48).Wols BA; Hofman-Caris CH Review of photochemical reaction constants of organic micropollutants required for UV advanced oxidation processes in water. Water Res. 2012, 46 (9), 2815–2827. [DOI] [PubMed] [Google Scholar]
- (49).Gallagher DL; Phetxumphou K; Smiley E; Dietrich AM Tale of two isomers: Complexities of human odor perception for cis-and trans-4-methylcyclohexane methanol from the chemical spill in West Virginia. Environ. Sci. Technol 2015, 49 (3), 1319–1327. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.