Abstract
The mortality rate of non-small cell lung cancer (NSCLC) remains high worldwide. miR-21-5p plays an important part in many cancer types, including NSCLC. However, the effect of miR-21-5p in NSCLC tumorigenesis remains poorly understood. The present study investigated whether miR-21-5p promoted NSCLC cell proliferation in vitro. In order to study the molecular mechanism by which miR-21-5p contributes to NSCLC progression, three bioinformatics algorithms were used to predict the genes which miR-21-5p targeted. TGFBI was identfieid as a putative direct target in NSCLC cells via the luciferase reporter assay. Furthermore, miR-21-5p downregulated TGFBI protein expression by a post-transcriptional mechanism via western blotting and a reverse transcription-quantitative polymerase chain reaction analysis. Finally, TGFBI exhibited opposing effects to those of miR-21-5p on NSCLC cells, suggesting that miR-21-5p may promote cell proliferation by negative regulation of TGFBI. These results suggest miR-21-5p promote the proliferation of NSCLC cells via inhibiting TGFBI expression.
Keywords: miR-21-5p, TGFBI, NSCLC
Introduction
More than 85% of lung cancer cases are non-small cell lung cancer (NSCLC), and the mortality rate of NSCLC remains high all over the world (1). In USA, NSCLC was the second most prevalent cancer among all new cancer cases and cancer deaths in 2016 (2). Traditional chemotherapy regimens for NSCLC have many disadvantages such as limited efficacy, high recurrence rate, and high toxicity (3). These disadvantages limit the efficacy of drug therapy for NSCLC, so an improved understanding of the exact mechanisms of this disease and developing new, targeted therapy drugs for NSCLC is urgent.
Transforming growth factor b-induced protein (TGFBI), known as βIg-h3 or keratoepithelin, contains a carboxyl-terminal Arg-Gly-Asp (RGD) integrin-binding sequence and four conserved fasciclin-1 (FAS1) domains (4). TGFBI plays an important role in many cellular disease processes, for example, tumorigenesis, progression, and metastasis (5). TGFBI is down-regulated in many cancer types such as lung cancer (6), breast cancer (7) and ovary carcinoma (8). TGFBI functions as a suppressive role via inhibition of cell proliferation, delay of the G1-S phase transition, and induction of senescence in the prevention of breast cancer and mesothelioma cells (9). TGFBI-derived peptides might be used as possible therapeutic adjuvants to enhance the response to chemotherapy in NSCLC (10). Recent studies suggested that loss of TGFBI expression has been described in lung cancer (11). It has been reported that downregulation of TGFBI protein is a frequent event and related to the tumor progression in human lung cancer through comparing 130 primary lung carcinomas to normal lung tissues (6,12). However, the upstream regulatory mechanism of TGFBI is not fully understood.
During the past decades, microRNAs (miRNAs, small non-coding RNA molecules), which is about 22 nucleotides in length, have been found to act in post-transcriptional regulation and RNA silencing of gene expression via binding the 3′-untranslated region (3′-UTR) of target mRNAs (13). Many reports have indicated miRNAs function as tumor suppressors or oncogenes in many cancer types (14). Many researches have indicated that miRNAs regulate many cellular processes including differentiation, proliferation, migration, and apoptosis (15). Recent researches have indicated miRNAs also function in the initiation and progression of NSCLC. For example, miR-455-3p was shown to regulate NSCLC cell proliferation and migration by downregulation of HOXB5 (16). Cell proliferation and invasion of NSCLC were inhibited by miR-504 by targeting LOXL2 (17). miR-142-5p inhibited tumorigenesis of NSCLC by targeting PIK3CA expression (18). Cell survival and metastasis of NSCLC were promoted by CXCL6 via down-regulation of miR-515-5p (19). Recent studies have reported miR-21-5p is upregulated in NSCLC patients (20). However, how miR-21-5p regulates cell proliferation in NSCLC and the involved molecular mechanisms remain poorly understood.
In our research, we show that miR-21-5p directly regulated TGFBI in two NSCLC cell lines. Additionally, we found that miR-21-5p promoted the proliferation of NSCLC cells via inhibiting TGFBI expression.
Materials and methods
Cell culture
We obtained two human NSCLC cell lines A549 and H1299 and human lung cell line HLF from the Cell Bank, Chinese Academy of Sciences (Shanghai, China). We cultured the cells using 10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA) in DMEM (HyClone; GE Healthcare Life Sciences, Logan, UT, USA) at 37°C with a humidified atmosphere of 5% CO2.
miRNA overexpression and knockdown
Either a miR-21-5p mimic or a miR-21-5p inhibitor were transfected into NSCLC cells to overexpress or knockdown miR-21-5p, respectively. We purchased the synthetic miR-21-5p mimic, miR-21-5p inhibitor, and negative control mimic and control inhibitor RNAs from Guangzhou RiboBio Co., Ltd., (Guangzhou, China). We seeded A549 and H1299 cells in 6-well plates using 10% FBS in DMEM. When the cells reached 60–70% confluence, the media was switched to Opti-MEM Reduced Serum Medium (Gibco; Thermo Fisher Scientific, Inc.) before transfection. Then, we transfected equal amounts of the miR-21-5p mimic, the control mimic, the miR-21-5p inhibitor, or the control inhibitor into the cells using Lipofectamine 3000 (Invitrogen; Thermo Fisher Scientific, Inc.) according to the manufacturer's protocols. After 48 h, the cells were harvested. The TGFBI expression levels were determined by western blotting, and the miR-21-5p expression levels were determined by RT-qPCR.
miR-21-5p expression in NSCLC
We used The Cancer Genome Atlas data (https://cancergenome.nih.gov/) to analyze and compare the miR-21-5p expression in NSCLC tissues and normal solid tissues. This database compared the miR-21-5p expression between 91 normal solid tissues and 792 primary solid tumors of NSCLC.
RNA isolation and reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
We extracted total RNA from transfected cells using TRIzol Reagent (Invitrogen; Thermo Fisher Scientific, Inc.) under the guidance of the manufacturer's protocols. We employed TaqMan miRNA probes (Applied Biosystems; Thermo Fisher Scientific, Inc.) to detect and quantify the levels of miRNAs under the guidance of the manufacturer's protocols. We performed RT-qPCR on a 7500 Real-time PCR System (Applied Biosystems; Thermo Fisher Scientific, Inc.) using a TaqMan PCR kit. Each reaction was run in triplicate. After that, we determined the cycle threshold (Cq) data using fixed threshold settings, then calculated the average Cq of triplicate PCR reactions. The relative amount of miRNAs were confirmed by the comparative Cq method (21). We used U6 snRNA as an internal control. The relative amount of miR-21-5p, which was normalized to U6, was calculated with the equation 2−ΔΔCq, in which ΔΔCq=(CqmiR-21-5p-CqU6) tumor-(CqmiR-21-5p-CqU6) control.
To quantify the amount of TGFBI mRNA, we used oligo dT primers (Takara Biotechnology Co., Ltd., Dalian, China) to reverse transcribe total RNA to cDNA. Next, RT-qPCR was run on the samples using SYBR Green Dye (Invitrogen; Thermo Fisher Scientific, Inc.) and specific primers targeting TGFBI and GAPDH (sequences listed in Table I). When the reactions were finished, we determined the Cq values with fixed threshold setting. The relative amount of TGFBI mRNA, which was normalized to GAPDH, was calculated through a similar method as described above.
Table I.
Sequences of primers.
| Primer | Sense | Antisense |
|---|---|---|
| TGFBI | 5′-GTGCGGCTAAAGTCTCTCCA-3′ | 5′-AAGCCCTGGAAAACGCTGAT-3′ |
| GAPDH | 5′-CGAGCCACATCGCTCAGACA-3′ | 5′-GTGGTGAAGACGCCAGTGGA-3′ |
TGFBI, transforming growth factor β induced.
Luciferase reporter assay
Direct targeting by miR-21-5p of the TGFBI gene was tested using luciferase reporter assays (22). Human genomic DNA was used as the template to amplify the entire 3′-UTR of human TGFBI by PCR. Next, the amplified PCR product was directly cloned into the p-MIR-reporter plasmid (Ambion; Thermo Fisher Scientific, Inc.), then we confirmed the successful insertion via sequencing. We mutated the sequences (interacted with the miR-21-5p seed region) from ATAAGCTA to TATTCGAT to test binding specificity, then inserted the TGFBI 3′-UTR mutant fragment into the p-MIR-reporter plasmid. Then, we seeded A549 cells into 24-well plates. The next day, we used Lipofectamine 3000 to transfect equal amounts (0.3 µg) of β-galactosidase (β-gal) expression plasmid (Ambion; Thermo Fisher Scientific, Inc.) and the firefly luciferase reporter plasmid together with 20 pmol of the miR-21-5p mimic, mimic control, miR-21-5p inhibitor, or inhibitor control. We used the β-gal plasmid as a control of transfection efficiency. The cells were assayed after 24 h post-transfection via a luciferase assay kit (Promega Corporation, Madison, WI, USA).
Plasmid construction and siRNA interference assays
A plasmid containing the human TGFBI open reading frame was purchased from Shanghai Genechem Co., Ltd., (Shanghai, China) and was used to overexpress TGFBI. We used an empty plasmid as a negative control. To silence TGFBI expression, TGFBI siRNA was obtained from Guangzhou RiboBio Co., Ltd. We used a scrambled siRNA sequence as a negative control. We then transfected the TGFBI overexpressing plasmid and TGFBI siRNA into A549 and H1299 cells using Lipofectamine 3000. After 48 h of transfection, total protein and RNA were extracted and evaluated by western blotting and RT-qPCR, respectively.
Protein extraction and western blotting
RIPA lysis buffer (Sigma-Aldrich; Merck KGaA, Darmstadt, Germany), which contained a protease inhibitor, was used to lyse protein from the cells. We separated proteins by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (Bio-Rad Laboratories, Inc., Hercules, CA, USA). The TGFBI expression levels were determined by western blotting. We normalized the protein levels by probing the same blots with a GAPDH antibody and analyzed the protein bands using Bandscan software Image J. The TGFBI antibody (A2561) was purchased from ABclonal Biotechnology (Woburn, MA, USA), and the GAPDH antibody (FL-335, sc-25778) was purchased from Santa Cruz Biotechnology, Inc., (Dallas, TX, USA).
Cell proliferation assay
For the CCK-8 assay, A549 cells were seeded into 96-well plates at 1×104 cells per well and cultured for 12 h using 10% FBS in DMEM. Cell proliferation was assessed under the guidance of the manufacturer's protocols of the Cell Counting Kit-8 (Nanjing KeyGen Biotech Co., Ltd., Nanjing, China). After transfection, we added 10 µl aliquot of CCK-8 solution to the test wells at 12, 24, 36, 48, and 60 h. Absorbance at 450 nm wavelength was measured after a 2 h incubation.
For the EdU assay, we seeded A549 cells into 48-well plates (Corning Incorporated, Corning, NY, USA). Until the transfected A549 cells reached 80% confluency, we used an EdU assay kit (Guangzhou RiboBio Co., Ltd.) to measure the cell proliferation rate. We followed the manufacturer's protocols except the nucleus staining dye was changed from Hoechst 33342 (supplied with the kit) to DAPI (Beyotime Institute of Biotechnology, Haimen, China) (23). When the stain was finished, the cells were imaged by fluorescence microscopy (BX51; Olympus Corporation, Tokyo, Japan).
Statistical analysis
Date shown are the mean ± standard deviation (SD) of at least three independent experiments, with P<0.05 considered to indicate a statistically significant difference. SPSS software, v.17.0 (SPSS, Inc., Chicago, IL, USA) was used to perform statistical analysis. The results were compared using one-way analysis of variance followed by a post hoc Tukey test for multiple comparisons.
Results
miR-21-5p expression is upregulated in NSCLC cell lines and tissues
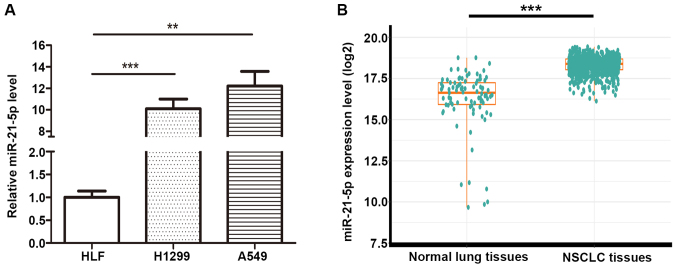
We have measured miR-21-5p expression in two NSCLC cell lines (A549 and H1299) and one normal lung cell lines (HLF). And we found miR-21-5p expression in A549 is upregulated compared with that in HLF and H1299, and we also found miR-21-5p expression in H1299 is upregulated compared with that in HLF (Fig. 1A). To investigate miR-21-5p levels compare between NSCLC vs. normal lung in clinical tissue samples, the TCGA database was employed to analyze miR-21-5p expression of 91 normal solid tissues and 792 primary solid tumors of NSCLC (Fig. 1B). The results showed that miR-21-5p was significantly upregulated in the solid tumors compared to the normal tissues (fold change=4.24 and P-value <0.001).
Figure 1.
miR-21-5p expression level is up-regulated in NSCLC tissues and cell lines. (A) miR-21-5p expression levels in two NSCLC cell lines (A549, H1299) and one normal lung cell line (HLF). (n=3; **P<0.01, ***P<0.001). (B) miR-21-5p expression levels in 792 NSCLC tissues were compared with 91 normal lung tissues obtained from the TCGA database and miR-21-5p was upregulated in NSCLC patients. The results were expressed as mean ± SD of three independent experiments (***P<0.001).
miR-21-5p promotes NSCLC cell proliferation
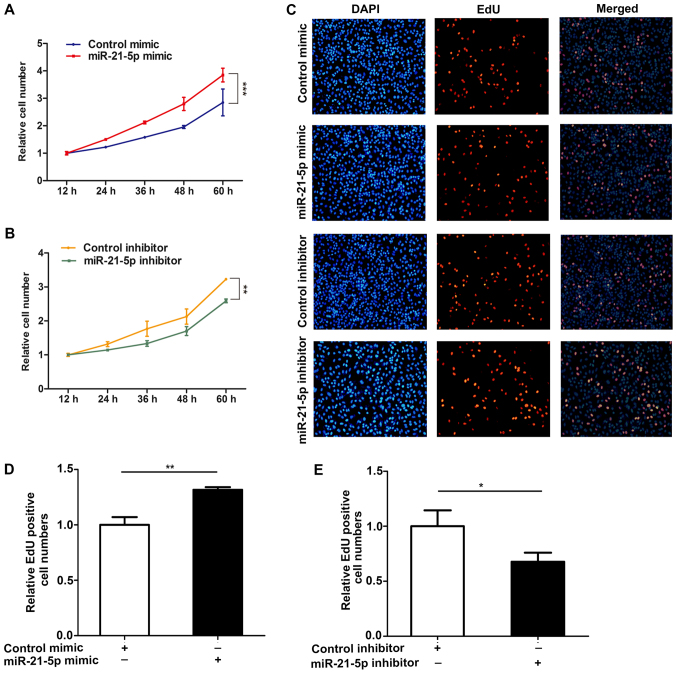
To better study the effect of miR-21-5p on the proliferation of NSCLC cells, we analyzed the role of miR-21-5p on the proliferation of NSCLC cells (A549) via CCK-8 and EdU assay methods. The results showed that overexpression of miR-21-5p resulted in an increase in proliferation of A549 cells, but that the proliferation of A549 cells was inhibited when the expression of miR-21-5p was inhibited (Fig. 2). These results indicate that miR-21-5p induces the proliferation of NSCLC cells.
Figure 2.
miR-21-5p promotes cell proliferation. (A) Cell proliferation curve of A549 cells transfected with the control mimic or miR-21-5p mimic. (B) Cell proliferation curve of A549 cells after transfection with equal doses of the control inhibitor or miR-21-5p inhibitor. (C-E) EdU assays performed in A549 cells transfected with equal doses of control mimic, miR-21-5p mimic, control inhibitor, or miR-21-5p inhibitor, and the EdU positive cells were observed at 565 nm using a fluorescence inverted microscope (magnification, ×200). (C) Representative images; (D) mimic and (E) inhibitor quantitative analysis. The results were expressed as mean ± SD of three independent experiments (n=3; *P<0.05; **P<0.01; ***P<0.001).
Prediction of TGFBI as a miR-21-5p target gene
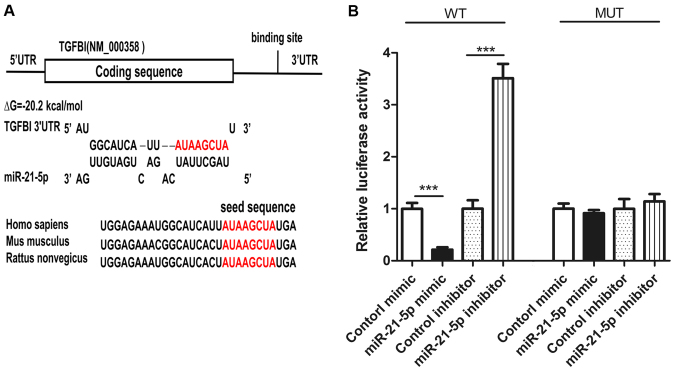
In order to elucidate the specific mechanism by which miR-21-5p induces the proliferation of NSCLC cells, we used three different computer software programs, TargetScan (24), miRanda (25), and PicTar (26) to predict genes that might bind to miR-21-5p. Among these candidate genes (zinc finger protein 367, G protein-coupled receptor 64, SMAD family member 7 and so on), TGFBI was predicted as miR-21-5p target-gene by all three programs. The potential for binding of TGFBI and miR-21-5p is shown in Fig. 2A. The binding energy of the hairpin structure is −20.2 kj/mol, and this binding energy is within the binding energy range for miRNA and target genes. In addition, the 3 ′UTR region of TGFBI binding with miR-21-5p is highly conserved across species (Fig. 3A).
Figure 3.
TGFBI is predicted to be a miR-21-5p target. (A) Graphical representation of the predicted duplex of the TGFBI 3′-UTR and miR-21-5p. The red sequence represents the miR-21-5p seed-recognition sequence in the 3′-UTR of TGFBI. The seed-recognition sequence is conserved. The predicted Gibbs free energy value is also listed. (B) Luciferase reporter assay. Firefly luciferase reporters which contain either WT or MUT miR-21-5p binding sites in the 3′-UTR of TGFBI were co-transfected into A549 cells with equal doses of the miR-21-5p mimic, control mimic, miR-21-5p inhibitor, or control inhibitor. The results were expressed as mean ± SD of three independent experiments (n=3; ***P<0.001). WT, wild-type; MUT, mutant.
Identification of the TGFBI as a miR-21-5p direct target gene
To verify whether miR-21-5p binds to the 3 ′UTR region of TGFBI, two luciferase reporter gene plasmids, TGFBI wild-type and TGFBI mutant-type, were constructed. We transfected these plasmids with miR-21-5p mimic, control mimic, miR-21-5p inhibitor, or control inhibitor into A549 cells. In wild type, the activity of luciferase was decreased after the overexpression of miR-21-5p (Fig. 3B) and the activity of luciferase was increased after inhibiting the expression of miR-21-5p (Fig. 3B). In the mutant, the activity of luciferase did not change after overexpression or inhibition of miR-21-5p (Fig. 3B). These findings indicate that the binding sequence is required for interaction of miR-21-5p and TGFBI mRNA.
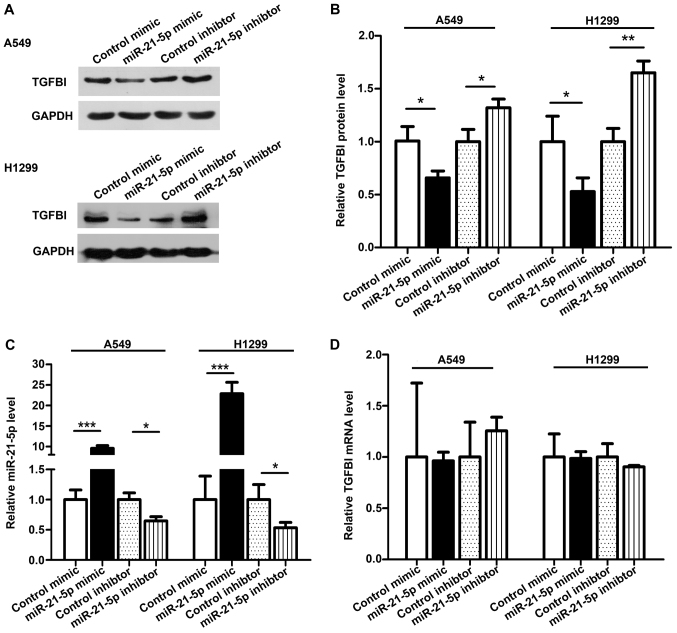
To further investigate whether TGFBI is a miR-21-5p target gene, we measured the expression of TGFBI in NSCLC cell lines A549 and H1299. To alter the miR-21-5p levels, we transfected a miR-21-5p mimic or a miR-21-5p inhibitor into A549 and H1299. As expected, the miR-21-5p expression increased and the TGFBI expression decreased significantly after miR-21-5p was overexpressed in A549 and H1299 cell lines (Fig. 4A-C). When we transfected the miR-21-5p inhibitor in A549 and H1299 cells, the miR-21-5p expression was decreased, and the TGFBI expression was increased (Fig. 4A-C). To better explain the effect of miR-21-5p on TGFBI expression, we measured the expression of TGFBI mRNA from transfected cells. Neither overexpression nor inhibition of miR-21-5p affected the mRNA level of TGFBI (Fig. 4D), indicating that miR-21-5p regulates the expression of the TGFBI gene at the post-transcriptional level.
Figure 4.
TGFBI is a direct miR-21-5p target gene. (A and B) Western blot analysis of TGFBI protein levels in A549 cells and H1299 cells after transfection with the control mimic, miR-21-5p mimic, control inhibitor, or miR-21-5p inhibitor. (A) Representative image. (B) Statistical analysis of the image. (C) RT-qPCR analysis of miR-21-5p level in A549 cells and H1299 cells transfected with the control mimic, miR-21-5p mimic, control inhibitor, or miR-21-5p inhibitor. (D) RT-qPCR analysis of TGFBI mRNA level in A549 cells and H1299 cells transfected with the control mimic, miR-21-5p mimic, control inhibitor, or miR-21-5p inhibitor. The results were expressed as mean ± SD of three independent experiments (n=3;*P<0.05; **P<0.01; ***P<0.001).
miR-21-5p promotes NSCLC cell proliferation by targeting TGFBI
Because miRNA can act on multiple genes, we studied whether the regulation of TGFBI expression by miR-21-5p affects the proliferation of NSCLC. To alter the expression of TGFBI, we transfected TGFBI siRNA or an overexpression plasmid into A549 cells. The efficiency of over expression and knockdown of TGFBI was measured and was shown in Fig. 5. The results showed that the proliferation of A549 cells is promoted after knockdown of TGFBI (Fig. 6A-C), but overexpression of TGFBI inhibited the proliferation of A549 cells (Fig. 6D-F). These results demonstrate that miR-21-5p and TGFBI exhibit opposite effect on the proliferation of NSCLC cells. Importantly, the proliferation rates of cells co-transfected with the miR-21-5p mimic and the TGFBI overexpressing plasmid were significantly lower than that of cells transfected with the miR-21-5p mimic alone (Fig. 7). These results indicate that the additional expression of TGFBI gene can restore the effect of miR-21-5p on cell proliferation. This result suggests that the regulation of TGFBI gene expression by miR-21-5p may explain how miR-21-5p exerts its function as an oncomiR, a microRNA (miRNA) that is associated with cancer.
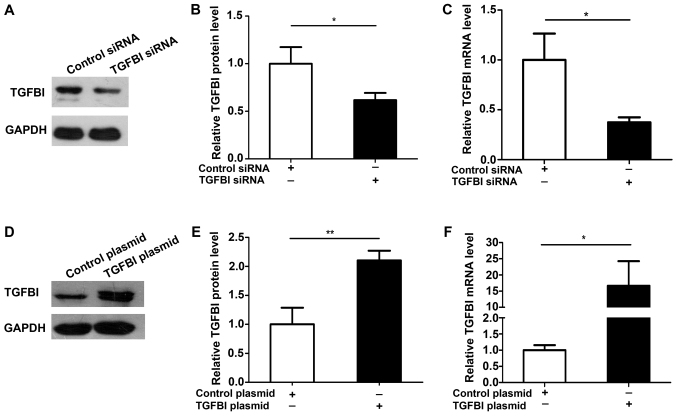
Figure 5.
Overexpression and knockdown efficiency of TGFBI. (A and B) Western blotting analysis of the TGFBI protein expression levels in A549 cells transfected with control siRNA or TGFBI siRNA. (A) Representative image; (B) quantitative analysis. (C) RT-qPCR analysis of the TGFBI mRNA expression levels in A549 cells transfected with control siRNA or TGFBI siRNA. (D and E) Western blotting analysis of the TGFBI protein expression levels in A549 cells transfected with control plasmid or TGFBI plasmid. (D) Representative image; (E) quantitative analysis. (F) RT-qPCR analysis of the TGFBI mRNA expression levels in A549 cells transfected with control plasmid or TGFBI plasmid. The results were expressed as mean ± SD of three independent experiments (n=3; *P<0.05; **P<0.01).
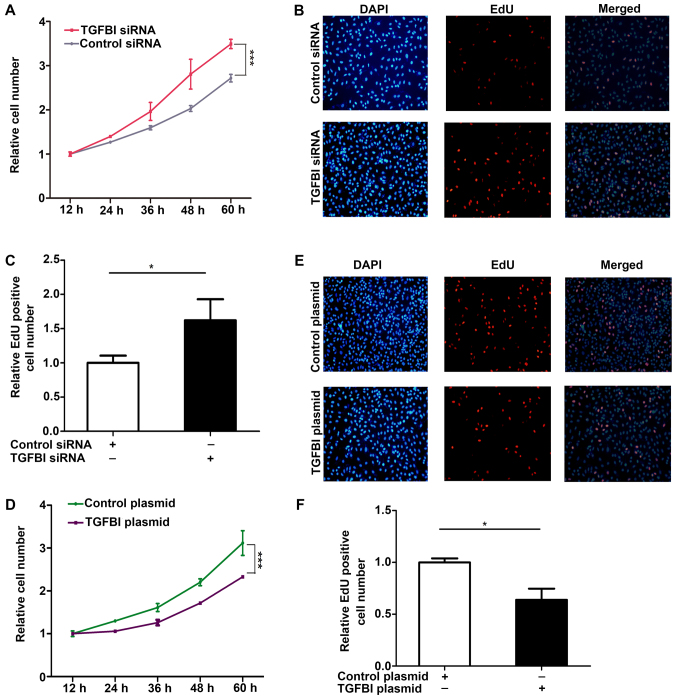
Figure 6.
TGFBI inhibits NSCLC cell proliferation. (A) Cell proliferation curve of A549 cells after transfection with equal doses of control siRNA or TGFBI siRNA. (B and C) EdU assays performed in A549 cells transfected with equal doses of control siRNA, TGFBI siRNA. (B) Representative image; (C) quantitative analysis. (D) Cell proliferation curve of A549 cells transfected with equal doses of the control plasmid or the TGFBI overexpressing plasmid. (E and F) EdU assays performed in A549 cells transfected with equal doses of the control plasmid or the TGFBI overexpressing plasmid, and the EdU positive cells were observed at 565 nm using a fluorescence inverted microscope (magnification, ×200). (E) Representative image; (F) quantitative analysis. The results were expressed as mean ± SD of three independent experiments (n=3; *P<0.05; ***P<0.001). NSCLC, non-small cell lung cancer.
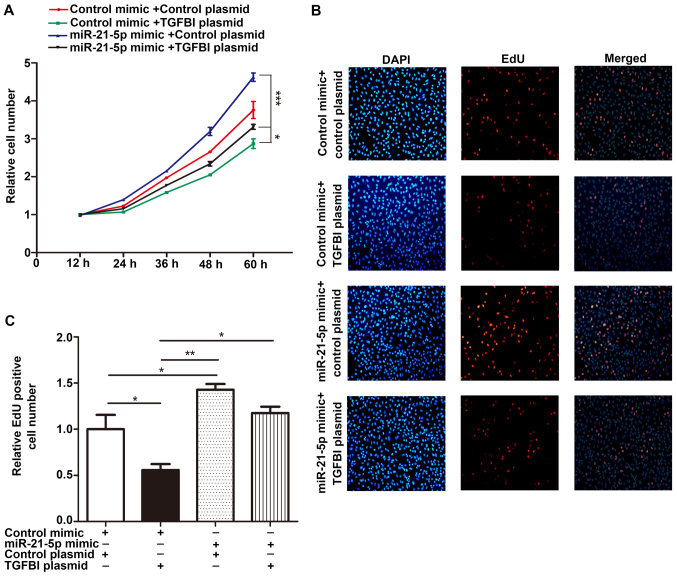
Figure 7.
miR-21-5p promotes NSCLC cell proliferation by targeting TGFBI. (A) Cell proliferation curve of A549 cells transfected with the control mimic plus the control plasmid, the control mimic plus the TGFBI overexpressing plasmid, the miR-21-5p mimic plus the control plasmid, or the miR-21-5p mimic plus the TGFBI overexpressing plasmid. (B and C) EdU assays performed in cells described in (A), and the EdU positive cells were observed at 565 nm using a fluorescence inverted microscope (magnification, ×200). (B) Quantitative analysis; (C) representative images. The results were expressed as mean ± SD of three independent experiments (n=3; *P<0.05; **P<0.01; ***P<0.001). NSCLC, non-small cell lung cancer.
Discussion
TGFBI is expressed at low levels in many types of cancers, including lung cancer (27). There have been many studies of TGFBI in cancer, but most studies have mainly focused on TGFBI expression levels, and the specific role of TGFBI in NSCLC is not clear. Recent studies have shown that TGFBI may serve as a predictive factor for chemotherapy response and suggest that TGFBI-derived peptides can be used as adjuvant for the treatment of NSCLC (10). In this study, silencing TGFBI by siRNA promoted the proliferation of NSCLC cells. After overexpression of TGFBI, the opposite results were obtained, suggesting that TGFBI may be a tumor suppressor gene that acts in the prevention of NSCLC. The lncRNA H19/miR-675 axis inhibits metastasis of prostate cancer cells by targeting TGFBI (28). Recent study has shown that in corneal fibroblasts, TGF-β regulated the expression of the TGFBI protein through miR-21- and miR-181a-coordinated activity and Smad signaling (29). Several miRNA can be highly expressed in cancer, and can act on tumor suppressor genes, decreasing gene expression. Therefore, we used three analysis programs to predict miRNAs that could target TGFBI. Among these candidate miRNAs (such as miR-489-3p, miR-21-5p, miR-590-5p and miR-9-5p), miR-21-5p has been reported as an oncogene in a variety of cancers including lung cancer (30). We demonstrated the interaction of miR-21-5p and TGFBI by luciferase experiment. To verify miR-21-5p regulation of TGFBI gene expression, we overexpressed miR-21-5p in NSCLC cell lines A549 and H1299. The expression of the TGFBI protein decreased significantly and the expression of TGFBI increased after miR-21-5p was inhibited, but the expression level of TGFBI mRNA did not change. These results show that miR-21-5p can inhibit the TGFBI expression by post-transcriptional regulation. Finally, we indicate that miR-21-5p promotes the proliferation of NSCLC cells by inhibiting the expression of TGFBI. These findings reveal the important role of miR-21-5p targeting TGFBI as a novel regulatory pathway in the development of NSCLC.
Previous researches have shown that miRNAs may be an important regulator of cancer (31). The miRNA miR-21-5p is highly expressed in a variety of cancers, such as laryngeal squamous cell carcinoma (32), gastric cancer (33), colorectal cancer (34), breast cancer (35), and NSCLC (36). Multiple genes can be targeted by one single miRNA, and the same gene can be regulated by multiple miRNA. Therefore, miR-21-5p might have different mRNA binding partners other than TGFBI, and these different target genes may also play an important role in carcinogenesis. For example, miR-21-5p has been reported to promote ox-LDL-induced endothelial cell senescence by down-regulation of the mitochondrial fission protein Drp1 (37). PDCD4 may be a functional target of miR-21-5p in osteosarcoma cells (38). Therefore, it is important to study the mechanistic details of this new pathway in the process of NSCLC cells. In our research, we showed that overexpression of miR-21-5p promotes the proliferation of NSCLC cells, and the reduction of TGFBI can mimic the induction of miR-21-5p. Interestingly, although miR-21-5p has many other targeted genes, overexpression of TGFBI can successfully counter the role of miR-21-5p to induce the proliferation of NSCLC cells. Our results suggest targeting TGFBI may be the important mechanism by which miR-21-5p exhibit its oncomiR roles. Thus, the reason why the up-regulation of miR-21-5p in NSCLC can induce cell proliferation might be that TGFBI is regulated by miR-21-5p.
In summary, this study describes a new regulatory network in which miR-21-5p and TGFBI can precisely tune the NSCLC cells proliferation. This study suggest new directions for NSCLC treatment in the future. Future studies should pay more attention on the effect of miR-21-5p on other cellular functions of NSCLC such as apoptosis, migration ability or invasiveness.
Acknowledgements
Not applicable.
Funding
The present study was supported by grants from the fund for outstanding talented young persons in colleges and universities of Anhui province (grant no. 2012SQRL119), the natural science research project in colleges and universities of Anhui province (grant nos. KJ2016A734, KJ2016SD59 and KJ2017A256), the National Natural Science Foundation of China (grant nos. 81601380, 81802651 and 31372198), and the Outstanding Young Talent Support Program Key Projects in Anhui Colleges and Universities (grant no. gxyqZD2016173).
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Authors' contributions
LN and YaZ were involved in the study concept and design, and analysis and interpretation of data; LY, JM, JZ, and YW were involved in acquisition of data, analysis and interpretation of data, statistical analysis, and drafting of the manuscript; YuZ performed bioinformatics analysis; ZW and YiZ performed the luciferase reporter assay; LC and LX performed RNA isolation and RT-qPCR; LL and SL performed the cell proliferation assay and western blotting; ZQ and LX revised the manuscript, and performed analysis and interpretation of data.
Ethics approval and consent to participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
References
- 1.Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108. doi: 10.3322/caac.21262. [DOI] [PubMed] [Google Scholar]
- 2.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30. doi: 10.3322/caac.21332. [DOI] [PubMed] [Google Scholar]
- 3.Hung MS, Chen IC, You L, Jablons DM, Li YC, Mao JH, Xu Z, Hsieh MJ, Lin YC, Yang CT, et al. Knockdown of Cul4A increases chemosensitivity to gemcitabine through upregulation of TGFBI in lung cancer cells. Oncol Rep. 2015;34:3187–3195. doi: 10.3892/or.2015.4324. [DOI] [PubMed] [Google Scholar]
- 4.Kim JE, Kim SJ, Lee BH, Park RW, Kim KS, Kim IS. Identification of motifs for cell adhesion within the repeated domains of transforming growth factor-beta-induced gene, betaig-h3. J Biol Chem. 2000;275:30907–30915. doi: 10.1074/jbc.M002752200. [DOI] [PubMed] [Google Scholar]
- 5.Zhang Y, Wen G, Shao G, Wang C, Lin C, Fang H, Balajee AS, Bhagat G, Hei TK, Zhao Y. TGFBI deficiency predisposes mice to spontaneous tumor development. Cancer Res. 2009;69:37–44. doi: 10.1158/0008-5472.CAN-08-1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhao Y, El-Gabry M, Hei TK. Loss of Betaig-h3 protein is frequent in primary lung carcinoma and related to tumorigenic phenotype in lung cancer cells. Mol Carcinog. 2006;45:84–92. doi: 10.1002/mc.20167. [DOI] [PubMed] [Google Scholar]
- 7.Wen G, Partridge MA, Li B, Hong M, Liao W, Cheng SK, Zhao Y, Calaf GM, Liu T, Zhou J, et al. TGFBI expression reduces in vitro and in vivo metastatic potential of lung and breast tumor cells. Cancer Lett. 2011;308:23–32. doi: 10.1016/j.canlet.2011.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ahmed AA, Mills AD, Ibrahim AE, Temple J, Blenkiron C, Vias M, Massie CE, Iyer NG, McGeoch A, Crawford R, et al. The extracellular matrix protein TGFBI induces microtubule stabilization and sensitizes ovarian cancers to paclitaxel. Cancer Cell. 2007;12:514–527. doi: 10.1016/j.ccr.2007.11.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li B, Wen G, Zhao Y, Tong J, Hei TK. The role of TGFBI in mesothelioma and breast cancer: Association with tumor suppression. BMC Cancer. 2012;12:239. doi: 10.1186/1471-2407-12-239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Irigoyen M, Pajares MJ, Agorreta J, Ponz-Sarvisé M, Salvo E, Lozano MD, Pio R, Gil-Bazo I, Rouzaut A. TGFBI expression is associated with a better response to chemotherapy in NSCLC. Mol Cancer. 2010;9:130. doi: 10.1186/1476-4598-9-130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhao Y, Shao G, Piao CQ, Berenguer J, Hei TK. Down-regulation of Betaig-h3 gene is involved in the tumorigenesis in human bronchial epithelial cells induced by heavy-ion radiation. Radiat Res. 2004;162:655–659. doi: 10.1667/RR3270. [DOI] [PubMed] [Google Scholar]
- 12.Shao G, Berenguer J, Borczuk AC, Powell CA, Hei TK, Zhao Y. Epigenetic inactivation of Betaig-h3 gene in human cancer cells. Cancer Res. 2006;66:4566–4573. doi: 10.1158/0008-5472.CAN-05-2130. [DOI] [PubMed] [Google Scholar]
- 13.Shimono Y, Zabala M, Cho RW, Lobo N, Dalerba P, Qian D, Diehn M, Liu H, Panula SP, Chiao E, et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell. 2009;138:592–603. doi: 10.1016/j.cell.2009.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hu X, Zhang M, Miao J, Wang X, Huang C. miRNA-4317 suppresses human gastric cancer cell proliferation by targeting ZNF322. Cell Biol Int. 2018;42:923–930. doi: 10.1002/cbin.10870. [DOI] [PubMed] [Google Scholar]
- 15.Mei Q, Li X, Guo M, Fu X, Han W. The miRNA network: Micro-regulator of cell signaling in cancer. Expert Rev Anticancer Ther. 2014;14:1515–1527. doi: 10.1586/14737140.2014.953935. [DOI] [PubMed] [Google Scholar]
- 16.Gao X, Zhao H, Diao C, Wang X, Xie Y, Liu Y, Han J, Zhang M. miR-455-3p serves as prognostic factor and regulates the proliferation and migration of non-small cell lung cancer through targeting HOXB5. Biochem Biophys Res Commun. 2018;495:1074–1080. doi: 10.1016/j.bbrc.2017.11.123. [DOI] [PubMed] [Google Scholar]
- 17.Ye MF, Zhang JG, Guo TX, Pan XJ. MiR-504 inhibits cell proliferation and invasion by targeting LOXL2 in non small cell lung cancer. Biomed Pharmacother. 2018;97:1289–1295. doi: 10.1016/j.biopha.2017.11.005. [DOI] [PubMed] [Google Scholar]
- 18.Wang Z, Liu Z, Fang X, Yang H. MiR-142-5p suppresses tumorigenesis by targeting PIK3CA in non-small cell lung cancer. Cell Physiol Biochem. 2017;43:2505–2515. doi: 10.1159/000484459. [DOI] [PubMed] [Google Scholar]
- 19.Li J, Tang Z, Wang H, Wu W, Zhou F, Ke H, Lu W, Zhang S, Zhang Y, Yang S, et al. CXCL6 promotes non-small cell lung cancer cell survival and metastasis via down-regulation of miR-515-5p. Biomed Pharmacother. 2018;97:1182–1188. doi: 10.1016/j.biopha.2017.11.004. [DOI] [PubMed] [Google Scholar]
- 20.Zhou Y, Sheng B, Xia Q, Guan X, Zhang Y. Association of long non-coding RNA H19 and microRNA-21 expression with the biological features and prognosis of non-small cell lung cancer. Cancer Gene The. 2017;24:317–324. doi: 10.1038/cgt.2017.20. [DOI] [PubMed] [Google Scholar]
- 21.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 22.Chen X, Wang K, Chen J, Guo J, Yin Y, Cai X, Guo X, Wang G, Yang R, Zhu L, et al. In vitro evidence suggests that miR-133a-mediated regulation of uncoupling protein 2 (UCP2) is an indispensable step in myogenic differentiation. J Biol Chem. 2009;284:5362–5369. doi: 10.1074/jbc.M807523200. [DOI] [PubMed] [Google Scholar]
- 23.Liu Y, Liu R, Yang F, Cheng R, Chen X, Cui S, Gu Y, Sun W, You C, Liu Z, et al. miR-19a promotes colorectal cancer proliferation and migration by targeting TIA1. Mol Cancer. 2017;16:53. doi: 10.1186/s12943-017-0625-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mazière P, Enright AJ. Prediction of microRNA targets. Drug Discov Today. 2007;12:452–458. doi: 10.1016/j.drudis.2007.04.002. [DOI] [PubMed] [Google Scholar]
- 25.John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS. Human MicroRNA targets. PLoS Biol. 2004;2:e363. doi: 10.1371/journal.pbio.0020363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M, Rajewsky N. Combinatorial microRNA target predictions. Nat Genet. 2005;37:495–500. doi: 10.1038/ng1536. [DOI] [PubMed] [Google Scholar]
- 27.Ivanov SV, Ivanova AV, Salnikow K, Timofeeva O, Subramaniam M, Lerman MI. Two novel VHL targets, TGFBI (BIGH3) and its transactivator KLF10, are up-regulated in renal clear cell carcinoma and other tumors. Biochem Biophys Res Commun. 2008;370:536–540. doi: 10.1016/j.bbrc.2008.03.066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhu M, Chen Q, Liu X, Sun Q, Zhao X, Deng R, Wang Y, Huang J, Xu M, Yan J, Yu J. lncRNA H19/miR-675 axis represses prostate cancer metastasis by targeting TGFBI. FEBS J. 2014;281:3766–3775. doi: 10.1111/febs.12902. [DOI] [PubMed] [Google Scholar]
- 29.Choi SI, Jin JY, Maeng YS, Kim TI, Kim EK. TGF-beta regulates TGFBIp expression in corneal fibroblasts via miR-21, miR-181a, and Smad signaling. Biochem Biophys Res Commun. 2016;472:150–155. doi: 10.1016/j.bbrc.2016.02.086. [DOI] [PubMed] [Google Scholar]
- 30.Liu XG, Zhu WY, Huang YY, Ma LN, Zhou SQ, Wang YK, Zeng F, Zhou JH, Zhang YK. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med Oncol. 2012;29:618–626. doi: 10.1007/s12032-011-9923-y. [DOI] [PubMed] [Google Scholar]
- 31.Ruvkun G. Clarifications on miRNA and cancer. Science. 2006;311:36–37. doi: 10.1126/science.311.5757.36d. [DOI] [PubMed] [Google Scholar]
- 32.Re M, Magliulo G, Gioacchini FM, Bajraktari A, Bertini A, Ceka A, Rubini C, Ferrante L, Procopio AD, Olivieri F. Expression levels and clinical significance of miR-21-5p, miR-let-7a and miR-34c-5p in laryngeal squamous cell carcinoma. Biomed Res Int. 2017;2017:3921258. doi: 10.1155/2017/3921258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chan SH, Wu CW, Li AF, Chi CW, Lin WC. miR-21 microRNA expression in human gastric carcinomas and its clinical association. Anticancer Res. 2008;28:907–911. [PubMed] [Google Scholar]
- 34.Zhu M, Huang Z, Zhu D, Zhou X, Shan X, Qi LW, Wu L, Cheng W, Zhu J, Zhang L, et al. A panel of microRNA signature in serum for colorectal cancer diagnosis. Oncotarget. 2017;8:17081–17091. doi: 10.18632/oncotarget.15059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Huo D, Clayton WM, Yoshimatsu TF, Chen J, Olopade OI. Identification of a circulating microRNA signature to distinguish recurrence in breast cancer patients. Oncotarget. 2016;7:55231–55248. doi: 10.18632/oncotarget.10485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang K, Chen M, Wu W. Analysis of microRNA (miRNA) expression profiles reveals 11 key biomarkers associated with non-small cell lung cancer. World J Surg Oncol. 2017;15:175. doi: 10.1186/s12957-017-1244-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhang JJ, Liu WQ, Peng JJ, Ma QL, Peng J, Luo XJ. miR-21-5p/203a-3p promote ox-LDL-induced endothelial cell senescence through down-regulation of mitochondrial fission protein Drp1. Mech Ageing Dev. 2017;164:8–19. doi: 10.1016/j.mad.2017.03.009. [DOI] [PubMed] [Google Scholar]
- 38.Zhang R, Xia T. Long non-coding RNA XIST regulates PDCD4 expression by interacting with miR-21-5p and inhibits osteosarcoma cell growth and metastasis. Int J Oncol. 2017;51:1460–1470. doi: 10.3892/ijo.2017.4127. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data generated or analyzed during this study are included in this published article.