Abstract
Background
The use of inedible lignocellulosic biomasses for biomanufacturing provides important environmental and economic benefits for society. Efficient co-utilization of lignocellulosic biomass-derived sugars, primarily glucose and xylose, is critical for the viability of lignocellulosic biorefineries. However, the phenomenon of glucose repression prevents co-utilization of both glucose and xylose in cellulosic hydrolysates.
Results
To circumvent glucose repression, co-utilization of cellobiose and xylose by Bacillus coagulans NL01 was investigated. During co-fermentation of cellobiose and xylose, B. coagulans NL01 simultaneously consumed the sugar mixtures and exhibited an improved lactic acid yield compared with co-fermentation of glucose and xylose. Moreover, the cellobiose metabolism of B. coagulans NL01 was investigated for the first time. Based on comparative genomic analysis, two gene clusters that encode two different operons of the cellobiose-specific phosphoenolpyruvate-dependent phosphotransferase system (assigned as CELO1 and CELO2) were identified. For CELO1, five genes were arranged as celA (encoding EIIAcel), celB (encoding EIIBcel), celC (encoding EIICcel), pbgl (encoding 6-phospho-β-glucosidase), and celR (encoding a transcriptional regulator), and these genes were found to be ubiquitous in different B. coagulans strains. Based on gene knockout results, CELO1 was confirmed to be responsible for the transport and assimilation of cellobiose. For CELO2, the five genes were arranged as celR, celB, celA, celX (encoding DUF871 domain-containing protein), and celC, and these genes were only found in some B. coagulans strains. However, through a comparison of cellobiose fermentation by NL01 and DSM1 that only possess CELO1, it was observed that CELO2 might also play an important role in the utilization of cellobiose in vivo despite the fact that no pbgl gene was found. When CELO1 or CELO2 was expressed in Escherichia coli, the recombinant strain exhibited distinct cellobiose uptake and consumption.
Conclusions
This study demonstrated the cellobiose-assimilating pathway of B. coagulans and provided a new co-utilization strategy of cellobiose and xylose to overcome the obstacles that result from glucose repression in a biorefinery system.
Electronic supplementary material
The online version of this article (10.1186/s13068-018-1323-5) contains supplementary material, which is available to authorized users.
Keywords: Bacillus coagulans, Glucose repression, Cellobiose, Xylose, Cellobiose operon, Phosphoenolpyruvate-dependent phosphotransferase system, Escherichia coli
Introduction
Lignocellulosic biomasses, in particular agricultural and forest residues, represent an attractive resource for replacing fossil fuels. This is because lignocellulosic biomasses are renewable, abundant, and enriched with convertible polysaccharides. Model processes for converting biomass into utilizable fuels, chemicals, and materials are known as lignocellulosic biorefineries. Within a conventional biorefinery process, structural cellulose and hemicellulose are degraded to various monosaccharides after pretreatment and hydrolysis. These resultant monosaccharides can then be further transformed by various microorganisms to produce liquid fuels and platform chemicals [1–4]. The rendered hexose monosaccharides, mainly glucose, mostly hail from cellulose hydrolysis; whereas, the pentoses (mainly xylose if hardwoods and non-woods are processed) are formed from hemicelluloses degraded during pretreatment [5]. However, most microorganisms are unable to ferment xylose, presenting a processing challenge for maximizing the monosaccharides generated by a biorefinery process [6, 7]. Even if a given microorganism can metabolize pentoses, glucose repression is a significant barrier toward effective co-utilization of glucose and xylose mixtures. Specifically, glucose repression causes a sequential or diauxic fermentation of the mixed sugars, resulting in low productivity and, therefore, high processing expenses [7–9]. In general, the phenomenon of glucose repression presents to be a problematic bottleneck for lignocellulosic biorefineries and must be overcome to achieve economic viability.
The common strategy to overcome glucose repression is metabolic engineering of microorganism strains, including deletion of the ptsG gene [7], mutation of the crp or ccpA gene [10], or mutation of the catabolite response element sequence [11]. This partially relieves glucose repression, but co-fermentation of glucose and xylose has been shown to remain inferior relative to single substrate fermentation. However, up to now, investigations into solving the issue of glucose repression by genetic modification have not yielded great breakthroughs. To bypass the problems caused by glucose repression, an alternative strategy is to co-ferment cellobiose, a dimer of glucose, simultaneously with xylose [8]. However, most wild-type microorganisms, such as Escherichia coli and yeasts, are unable to ferment cellobiose. As a result, numerous metabolic engineering approaches have been carried out to construct recombinant microorganisms with cellobiose consumption ability. The first strategy was through display of β-glucosidase on the cell surface, which hydrolysed cellobiose into two molecules of glucose outside the cells. However, the co-fermentation process required accurate control of both cellobiose hydrolysis and glucose consumption rates to avoid glucose accumulation [12]. Thereafter, the extracellular hydrolysis of cellobiose was substituted by intercellular hydrolysis [13]. By integration of a cellodextrin transporter, an intracellular β-glucosidase, and the xylose metabolic pathway into Saccharomyces cerevisiae, the engineered S. cerevisiae was capable of co-fermenting cellobiose and xylose simultaneously [14]. In addition, the role of β-glucosidase can be replaced by cellobiose phosphorylase, which cleaves cellobiose into glucose and glucose-1-phosphate [9, 15].
Bacillus coagulans has been demonstrated to be a suitable microorganism for the efficient conversion of biomass-derived monosaccharides into lactic acid, which is an important platform chemical [16, 17]. Many studies reported that B. coagulans strains can ferment xylose to lactic acid via the pentose phosphate pathway. In addition, the xylose operon of B. coagulans has been analyzed, heterologously expressed, and characterized [18]. Moreover, our previous report showed that B. coagulans might also possess an effective cellobiose transportation and metabolism pathway [19]. Therefore, unlike recombinant strains, B. coagulans may be an ideal wild-type microorganism with the capability to ferment both cellobiose and xylose. However, co-fermentation of cellobiose and xylose by B. coagulans and the pathway of cellobiose metabolism in B. coagulans have not yet been studied.
With that, we aimed to evaluate co-utilization of cellobiose and xylose by B. coagulans for lactic acid production and to identify the genes responsible for cellobiose catabolism in B. coagulans. Firstly, the ability of B. coagulans to convert cellobiose into lactic acid was investigated. Next, the fermentation performance of B. coagulans was examined using mixture of cellobiose and xylose. Based on these results, we intended to then probe for the cellobiose operons present in B. coagulans. The goal of this study was to advance the utilization of cellobiose and xylose mixtures in a biorefinery setting with hopes that B. coagulans will be recognized as a strong candidate for improving the efficiency and overall economics of biorefinery processes.
Materials and methods
Bacterial strains and media
All strains, plasmids, and primers used in this study are listed in Table 1. B. coagulans strains were maintained on a GYC agar slant containing (g/L): glucose 20, yeast extract 1, corn steep liquor powder 2.5, NH4Cl 1, MgSO4·7H2O 0.2, and CaCO3 10. The pH was adjusted to 7.2 using NaOH before adding CaCO3. The slant was incubated at 50 °C for 14 h and stored at 4 °C. For genetic engineering, B. coagulans ATCC 7050 was grown in BC medium at 45 °C and 120 rpm [20].
Table 1.
Strains, plasmids and primers used in this study
| Designation | Relevant characteristics | Source or reference |
|---|---|---|
| Strains | ||
| B. coagulans NL01 | Undomesticated wild strain | Laboratory preservation |
| B. coagulans ATCC 7050 | Undomesticated wild strain | American Type Culture Collection |
| B. coagulans XZL4 | Undomesticated wild strain | Professor Ping Xua |
| B. coagulans ATCC 7050 (ΔcelR) | celR deletion strain of B. coagulans ATCC 7050 | This study |
| L. lactis MG1363 | Host for gene cloning | Laboratory preservation |
| E. coli BL21(DE3) | Host for protein expression | Invitrogen |
| E. coli BL21 (pET-celo1) | E. coli BL21(DE3) harboring the plasmid pET-celo1 | This study |
| E. coli BL21 (pET-celo2) | E. coli BL21(DE3) harboring the plasmid pET-celo2 | This study |
| E. coli BL21 (pETDuet-1) | E. coli BL21(DE3) harboring the plasmid pETDuet-1 | This study |
| Plasmids | ||
| pMH77 | pSH71 replication containing temperature-sensitive vector, Cmr | Professor Oscar P. Kuipersb |
| pMD-19T | Gene cloning vector, Ampr | TaKaRa |
| 19T-celR1 | pMD-19T containing the flanking regions of celR | This study |
| pMH77-celR1 | pMH77 containing the flanking regions of celR | This study |
| pETDuet-1 | Protein expression vector, Ampr | Novagen |
| pET-celo1 | pETDuet-1 containing the CELO1 from B. coagulans NL01 | This study |
| pET-celo2 | pETDuet-1 containing the CELO2 from B. coagulans NL01 | This study |
| Primers | Sequences (5′ → 3′) and properties | |
| celR1up.f | GATCCGGAATTCCCACACAAAGGGAAATTACGTAAGCCAC (EcoRI) | This study |
| celR1up.r | GTAAATGGCGGCGGCACTGAGCTCATGCCAACAACTCCCTTTTATCATTG | This study |
| celR1down.f | CAATGATAAAAGGGAGTTGTTGGCATGAGCTCAGTGCCGCCGCCATTTAC | This study |
| celR1down.r | CATCCGCTCGAGCAATATCATCCAGTTTCACCATATTCGG (XhoI) | This study |
| celo1.f | AGATCTCTTGAACATAGAAGAAATCAGTTTTC (BglII) | This study |
| celo1.r | GGTACCTTATTTTATACATGCCATTAACTCAT (KpnI) | This study |
| celo2.f | GAGCTCGTTGATTTCGAATCGGCAAAAAC (SacI) | This study |
| celo2.r | CTCGAGTTAAGCAATGCTATCCTTCGTCGT (XhoI) | This study |
aFrom State Key Laboratory of Microbial Metabolism, Shanghai Jiao Tong University, People’s Republic of China
bFrom Groningen Biomolecular Sciences and Biotechnology Institute, Department of Genetics, University of Groningen, Netherlands
Escherichia coli strains were cultivated in Luria Broth (LB) medium, which contained 10 g/L tryptone, 5 g/L yeast extract, and 10 g/L NaCl. When needed, 100 mg/L ampicillin was added for plasmid maintenance.
Fermentation experiments of B. coagulans strains
A loop of cells collected from a fully grown slant was inoculated into a 150-mL flask containing 50 mL of GYC liquid medium as a seed culture. The culture was incubated for 14 h at 50 °C with 150 rpm and then inoculated with a 10% (v/v) inocula into Erlenmeyer flasks with for lactic acid production. Fermentations were conducted in 250-mL Erlenmeyer flasks containing 100 mL of medium with shaking at 50 °C and 150 rpm on a rotary shaker. The medium for cellobiose fermentation contained the following (g/L): various amount of cellobiose (20–120), yeast extract 2.5, corn syrup powder 1.2, MgSO4·7H2O 0.4, (NH4)2SO4 3, KH2PO4 0.22, MnSO4·H2O 0.03, FeSO4·H2O 0.03, and CaCO3 at 1/2 of the added sugar by weight. For the glucose/xylose and cellobiose/xylose co-utilization experiments, the sugars were replaced by 5 g/L glucose and 5 g/L xylose, 5 g/L cellobiose and 5 g/L xylose, 40 g/L glucose and 40 g/L xylose, and 40 g/L cellobiose and 40 g/L xylose, respectively. Samples were taken periodically to determine the concentrations of residual sugars and the produced lactic acid. All experiments were performed in triplicate.
RNA isolation, cDNA generation, and RT-PCR
Total RNA was isolated from the B. coagulans NL01 cells grown to the exponential phase under inducing conditions (with cellobiose as the sole carbon source), using a Qiagen RNeasy total RNA kit (Qiagen, Germany). The integrity of the RNA was guaranteed by electrophoresis of total RNA in a 1.5% agarose gel. The gDNA was erased in advance, and cDNA was generated by PrimeScript™ RT reagent kit with a gDNA eraser (TaKaRa, China). Reverse transcription PCR (RT-PCR) was performed in accordance with the standard procedures using 1 μM each specific primer (see Additional file 1: Table S1). The genomic DNA of B. coagulans NL01 was used as a positive control.
Gene knockout procedure of B. coagulans ATCC 7050
The celR gene knockout of B. coagulans ATCC 7050 was performed using plasmid pMH77. All procedures were conducted according to a previous Ref. [21]. Genomic DNA of B. coagulans ATCC 7050 was extracted using the Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA). The flanking regions of the celR gene were amplified from B. coagulans ATCC 7050 genomic DNA using the primers celR1up.f and celR1up.r (upstream, 1000 bp) and celR1down.f and celR1down.r (downstream, 1000 bp), respectively. After gel purification, overlap extension PCR was performed, wherein the upstream and downstream regions were fused using the primers celR1up.f and celR1down.r. The resulting PCR product was gel purified and ligated into pMD-19T to form a new plasmid, 19T-celR1. The plasmid 19T-celR1 was digested by EcoRI and XhoI and cloned into the corresponding sites of pMH77 to obtain another plasmid, pMH77-celR1. The plasmid pMH77-celR1 was first transformed into Lactococcus lactis MG1363. The extracted plasmid from L. lactis MG1363 was then as transformed into B. coagulans ATCC 7050 by electroporation.
A colony of B. coagulans ATCC 7050 harboring pMH77-celR1 was cultured at 45 °C for 12 h and then shifted to 60 °C for further incubation for 12 h. The diluted cells were plated on a BC plate supplemented with 7 mg/L chloramphenicol and cultured at 60 °C to select the first crossover cells. The right single crossover colony was incubated in BC liquid broth without chloramphenicol overnight at 45 °C and then spread onto BC plates without chloramphenicol. Plenty of colonies were sequentially spread onto BC plates with and without chloramphenicol and incubated overnight at 45 °C. The colony that grew on BC plates without chloramphenicol, but did not grow on BC plates with chloramphenicol, was the double crossover cell. The resulting deletion mutant was designated as B. coagulans ATCC 7050 (ΔcelR). All the constructed strains were validated by PCR and DNA sequencing.
Utilization of cellobiose by recombinant E. coli strains
The cellobiose operons of B. coagulans NL01 were amplified from the genomic DNA with primers celo1.f/celo1.r and celo2.f/celo2.r. The PCR product was cloned into the expression plasmid pETDuet-1 between unique BglII/KpnI and SacI/XhoI sites to yield the recombinant plasmid pET-celo1 and pET-celo2, respectively. The plasmids pET-celo1 and pET-celo2 were transformed into E. coli BL21(DE3) to construct the recombinant E. coli BL21 (pET-celo1) and E. coli BL21 (pET-celo2).
The resulting E. coli BL21 (pET-celo1), E. coli BL21 (pET-celo2) and a control with the same host bearing an empty plasmid were cultured in LB medium overnight. This culture was inoculated with 5% into 30-mL LB medium in the presence of 10 g/L cellobiose, 100 mg/L ampicillin, and 0.2 mM IPTG. The cells were further cultured at 37 °C and 200 rpm. The samples were collected periodically for determination of the cell density and the residual cellobiose. All experiments were performed in triplicate.
Analytic methods
Glucose, xylose, cellobiose, and lactic acid were analyzed using an Agilent HPLC 1260 system with a Bio-Rad Aminex HPX-87H column (300 × 7.8 mm) and a refractive index detector. The mobile phase was 5 mM H2SO4 at a flow rate of 0.6 mL/min, and the column temperature was maintained at 55 °C [6, 19]. The cell density of E. coli was estimated by determining the optical density at 600 nm.
Productivity (g/L/h) was calculated as the concentration of lactic acid produced per liter divided by the fermentation time (h). Lactic acid yield was calculated as the percentage of measured lactic acid relative to the theoretical amount of lactic acid producible. The theoretical amount was calculated as follows: glucose (1 g lactic acid/g glucose), xylose (1 g lactic acid/g xylose) and cellobiose (1.05 g lactic acid/g cellobiose) [1].
The activity of 6-phospho-β-glucosidase was determined using phosphorylated p-nitrophenyl-β-d-glucopyranoside-6-phosphate (pNPβG6P) as a substrate. The assay was carried out with a total volume of 1 mL, containing 50 mM PBS (pH 7), 1 mM pNPβG6P and 50 μL of suitably diluted crude enzyme. The reaction was incubated at 50 °C for 2 min and terminated by the addition of 500 μL of 1 M Na2CO3. The released p-nitrophenol (pNP) was determined by the absorbance at 410 nm. One unit of 6-phospho-β-glucosidase was defined as the amount of enzyme catalyzing the appearance of 1 μmol of pNP per min under experimental conditions.
Results and discussion
Efficient cellobiose utilization by B. coagulans NL01
As the basis for the current work, our previous studies found that B. coagulans strain NL01 had the ability to use cellobiose as carbon source to produce lactic acid [16, 19]. In this study, we first carried out the pure sugar fermentation at a concentration of 40 g/L glucose, cellobiose or xylose. The results showed that all three substrates were consumed quickly, and the lactic acid yields from glucose, cellobiose, and xylose were 94.1%, 91.1%, and 89.5%, respectively. These demonstrated that B. coagulans NL01 can efficiently utilize the lignocellulosic biomass-derived sugars.
To evaluate the maximum potential of B. coagulans NL01 for cellobiose utilization, fermentation was conducted with different initial concentrations of cellobiose, ranging from 40 g/L to 120 g/L. As shown in Fig. 1a, B. coagulans NL01 was capable of fermenting cellobiose efficiently regardless of its initial concentration. With the increased initial cellobiose concentration, the maximum concentration of lactic acid was raised from 40.8 to 114.5 g/L, and the yield varied from 91.1 to 94.8% (Fig. 1b). It should be noted that no fermentation byproducts, such as acetic acid or ethanol, were found present in the fermentation broth. Several other B. coagulans strains have been reported to have the capacity to ferment cellobiose for lactic acid production. For example, B. coagulans 36D1 was applied to production of lactic acid from cellulose through simultaneous saccharification and fermentation (SSF) in a fed-batch mode, which obtained a yield rate of approximately 88% [22]. B. coagulans LA204 produced lactic acid with a yield of 85.5% from 50 g/L cellobiose [23]. B. coagulans IPE22 can also use cellobiose; however, the fermentation performance of this strain under high cellobiose concentrations was not reported [24]. The best fermentation was performed by B. coagulans WCP10-4, which produced 196.3 g/L lactic acid from 200 g/L cellobiose in a 2-L fermentor [25]. Although previous papers have reported cellobiose fermentation, they did not focus on simultaneous consumption of cellobiose and xylose. Moreover, these studies did not investigate the genes responsible for cellobiose metabolism.
Fig. 1.
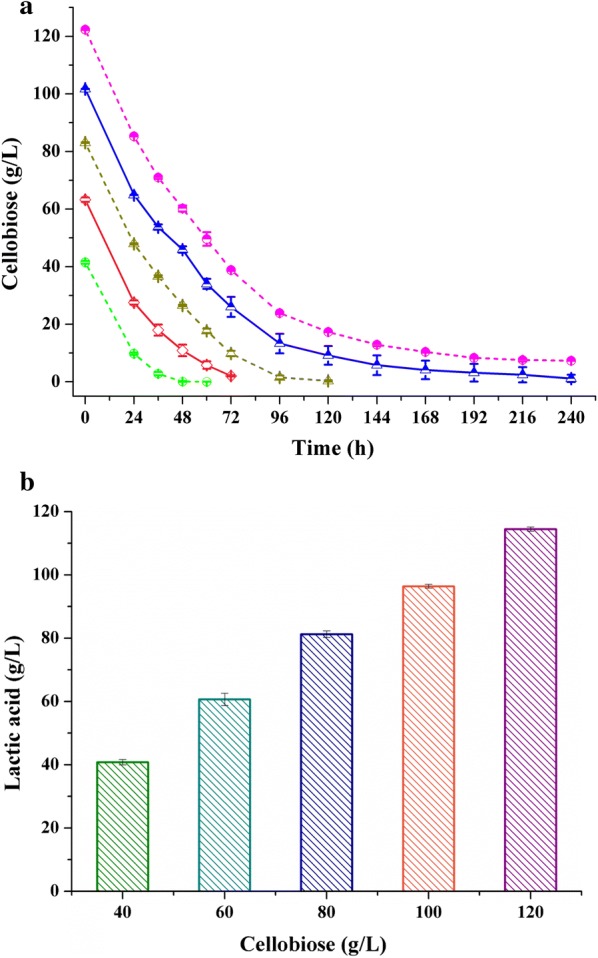
Fermentation of cellobiose by B. coagulans NL01. a Time course of cellobiose utilization under different initial concentrations. Green, 40 g/L; Pink, 60 g/L; Dark yellow, 80 g/L; Blue, 100 g/L; Magenta, 120 g/L. b Effect of cellobiose concentration on lactic acid production. The maximum concentration of lactic acid was obtained after 60, 72, 120, 240, and 240 h, respectively. Fermentations were carried out in 250-mL Erlenmeyer flasks containing 100-mL medium at 50 °C and 150 rpm on a rotary shaker. The medium contained the following (g/L): various amounts of cellobiose (40–120), yeast extract 2.5, corn syrup powder 1.2, MgSO4·7H2O 0.4, (NH4)2SO4 3, KH2PO4 0.22, MnSO4·H2O 0.03, FeSO4·H2O 0.03, and CaCO3 at 1/2 of the added sugar by weight. Values are the average of three independent experiments ± SD
Comparison of glucose/xylose and cellobiose/xylose co-utilization
As we introduced, xylose utilization by microorganisms is inhibited when glucose accumulates massively during SSF of lignocellulosic biomass. Interestingly, many recombinant strains demonstrated that cellobiose and xylose can assimilate simultaneously. However, few similar experiments have been conducted in wild strains. To demonstrate the superiority of cellobiose/xylose co-utilization in B. coagulans NL01, co-fermentation of glucose/xylose and cellobiose/xylose mixtures was carried out and compared.
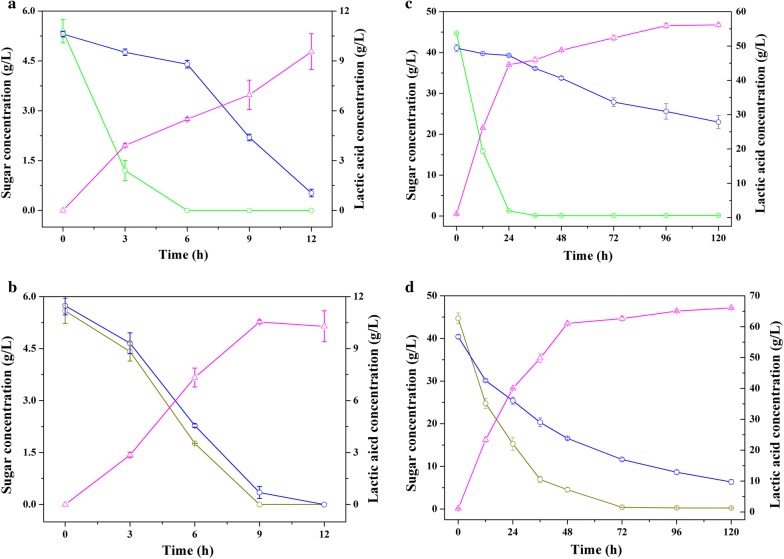
Figure 2 shows both sugar consumption and lactic acid production by B. coagulans NL01 at a total sugar concentration of approximately 10 g/L (Fig. 2a, b) and 80 g/L (Fig. 2c, d). During co-fermentation of glucose and xylose, as expected, xylose utilization was retarded during the early phases of fermentation. When glucose was depleted, the xylose uptake rate was markedly hastened. This observation indicated that glucose repression did occur. Figure 2a shows that the final lactic acid concentration was 9.6 g/L at a 12-h fermentation, representing 94.9% lactic acid yield. However, there remained some unutilized xylose in the fermentation medium at the conclusion of fermentation time. With the concentrations of sugars increasing, xylose consumption would be more inhibited. Moving on to the 80 g/L fermentation experiment, after a 120-h fermentation, approximately 23.0 g/L xylose remained (Fig. 2c). For the fermentation containing cellobiose and xylose, B. coagulans NL01 consumed both sugars simultaneously during fermentation (Fig. 2b, d). It was also be observed that both sugars were almost completely consumed, translating to a high lactic acid concentration [10.5 g/L at 9 h fermentation (Fig. 2b)]. In contrast, only 7 g/L lactic acid was produced within 9 h when co-fermentation involved 5 g/L glucose and 5 g/L xylose (Fig. 2a). As for co-fermentation of 40 g/L cellobiose and 40 g/L xylose, approximately 66.1 g/L lactic acid was produced at the fermentation end-point. The residual xylose was 6.4 g/L, which was much lower than that in Fig. 2c, d). The observation of incomplete xylose consumption could be explained by a lessened ability of B. coagulans NL01 regarding fermentation of xylose relative to fermentation of cellobiose. In fact, when using 80 g/L cellobiose or xylose as the substrate, cellobiose was consumed completely (Fig. 1a), whereas xylose remained in surplus (above 5 g/L) [26]. In all, the results of Fig. 2 indicate that B. coagulans NL01 was able to co-ferment cellobiose and xylose rapidly and efficiently (despite residual xylose).
Fig. 2.
Co-fermentation of glucose/xylose or cellobiose/xylose mixtures for lactic acid production by B. coagulans NL01. Green, glucose; Dark yellow, cellobiose; Blue, xylose; Magenta, lactic acid. Fermentations were carried out in 250-mL Erlenmeyer flasks containing 100-mL medium at 50 °C and 150 rpm on a rotary shaker. The medium contained the following (g/L): glucose 5 and xylose 5 (a)/cellobiose 5 and xylose 5 (b)/glucose 40 and xylose 40 (c)/cellobiose 40 and xylose 40 (d), yeast extract 2.5, corn syrup powder 1.2, MgSO4·7H2O 0.4, (NH4)2SO4 3, KH2PO4 0.22, MnSO4·H2O 0.03, FeSO4·H2O 0.03, and CaCO3 at 1/2 of the added sugar by weight. Values are the average of three independent experiments ± SD
Two different cellobiose metabolic operons found in B. coagulans NL01
So far, cellobiose metabolism in B. coagulans has not been reported. The excellent fermentation performance using cellobiose led us to further investigate the specific catabolic pathways in B. coagulans NL01 that provide the demonstrated superiority. At present, two different pathways of cellobiose degradation are known in microorganisms: hydrolysis (depending on β-glucosidase) and phosphorolysis (depending on cellobiose phosphorylase). These pathways have been elucidated as well as exploited in various instances of metabolic engineering [9, 15, 27]. Therefore, we have attempted to determine the activities of both β-glucosidase and cellobiose phosphorylase enzymes in B. coagulans NL01. However, no β-glucosidase or cellobiose phosphorylase activity was detected in any subcellular locations of B. coagulans NL01 in our studies.
Currently, numerous B. coagulans strains have undergone genomic sequencing and annotation, including B. coagulans NL01. In agreement with our previous discussion, no protein predicted as β-glucosidase or cellobiose phosphorylase could be found in the B. coagulans NL01 genome. Instead, a protein annotated as a putative 6-phospho-β-glucosidase (Genbank accession No. WP_046721809) was searched, which belongs to glycosyl hydrolase family 1. Previous studies reported that the β-1,4-glycosidic bond can be broken down by phosphoglycosyl hydrolases, such as 6-phospho-β-glucosidase, by the aid of the phosphoenolpyruvate-dependent phosphotransferase system (PEP-PTS) [28]. Therefore, we next attempted to determine 6-phospho-β-glucosidase enzymatic activity in B. coagulans NL01. When B. coagulans NL01 was cultivated using glucose, xylose or cellobiose as the sole carbon source, 6-phospho-β-glucosidase activity was detected in the presence of cellobiose (0.29 U/mg), but not in the presence of glucose or xylose. This observation indicated that 6-phospho-β-glucosidase may play an essential role in cellobiose metabolism and that expression of this enzyme required cellobiose for induction.
PEP-PTS is usually composed of EI, HPr and EII(s). EI and HPr are the general cytoplasmic PTS components. In contrast, EII(s) are substrate-specific permeases that consist of three or four domains IIA, IIB, IIC and IID (present only in a few families). They can be encoded by a single or multiple genes organized in an operon [28–30]. For catabolism of cellobiose, the cellobiose operon has been reported in some lactic acid bacteria, such as Streptococcus mutans [31] and Lactobacillus plantarum [32]. The genetic organization of cellobiose operons from different strains was quite different, but all included genes at least for 6-phospho-β-glucosidase (pbgl), a transcriptional regulator (celR), IIA (celA), IIB (celB), and IIC (celC).
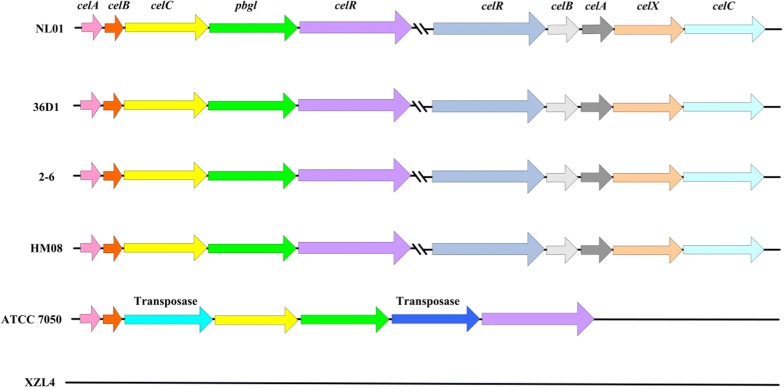
Up to now, no data related to a cellobiose operon in B. coagulans have been disclosed. Therefore, we sought to reveal all relevant components of the cellobiose operon in the B. coagulans NL01 genome. Figure 3 shows that this strain possesses two different potential cellobiose operons (assigned as CELO1 and CELO2) and that they were found to be not adjacent to one another. Each operon was composed of five genes and arranged in the same direction. For CELO1, the five genes were arranged as celA, celB, celC, pbgl, and celR. For CELO2, unexpectedly, no pbgl gene was found. Instead, a gene annotated as a DUF871 domain-containing protein (hereafter denoted as celX) was identified (DUF represents domains of unknown function). They were organized as celR, celB, celA, celX, and celC. Moreover, there was no sequence identity between the corresponding components of CELO1 and CELO2.
Fig. 3.
Organization of cellobiose operons in different B. coagulans strains. Arrows indicate the direction of gene translation
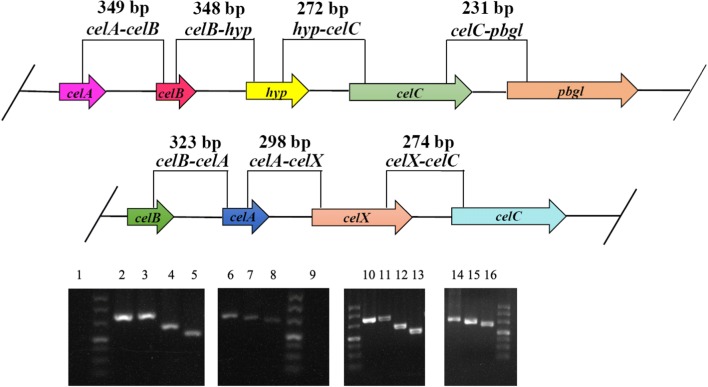
The co-transcription of structural genes in the CELO1 and CELO2 was certified by an RT-PCR approach. The cDNA product generated from total RNA extracted from cells induced by cellobiose was used as template in RT-PCR to examine the amplification of overlapping regions spanning each two adjacent genes. For CELO1, because the length from cleB to celC was longer than other intervals and a hypothetic gene (hyp) was predicted between them, the amplified overlapping region spanning cleB–celC was substituted by regions spanning cleB–hyp and hyp–celC (Fig. 4). Figure 4 shows that only a single PCR product was obtained from each amplification reaction, and the size of each fragment from cDNA was consistent with those from genomic DNA. These results proved that both of the gene clusters are co-transcribed as an operon.
Fig. 4.
Identification of the co-transcription of structural genes in the CELO1 and CELO2 operon using RT-PCR. cDNA was isolated and RT-PCR was performed as described in the text (for lane 2–8). The RNA and genomic DNA of B. coagulans NL01 was used as negative (for lane 1 and 9) and positive control (for lane 10–16), respectively. The amplified overlapping regions spanned celA–celB (lane 1, 2, and 10), celB–hyp (lane 3 and 11), hyp–celC (lane 4 and 12), celC–pbgl (lane 5 and 13), celB–celA (lane 6, 9, and 14), celA–celX (lane 7 and 15), celX–celC (lane 8 and 16)
The role of CELO1 and CELO2 in cellobiose utilization
To investigate whether other B. coagulans strains possess similar cellobiose operons, another five typical strains that had been sequenced and reported were selected. The chosen strains were B. coagulans 36D1, 2-6, HM-08, ATCC 7050 and XZL4. Figure 3 indicates that the cellobiose operons were ubiquitous in most B. coagulans strains. For strains NL01, 36D1, 2-6, and HM-08, all of them had two cellobiose operons, and comparisons between them revealed a high degree of synteny. For strain ATCC 7050, only the cellobiose operon CELO1 was found, and it was interspersed with two transposase genes. Strain XZL4 was the sole exception which had no cellobiose operon.
Next, strains NL01, ATCC 7050 and XZL4 were selected to determine whether the difference in the cellobiose operons affected their cellobiose fermentation ability. Figure 5 shows that strains NL01 and ATCC 7050 could utilize cellobiose, whereas XZL4 could not. Specifically, the utilization rate of cellobiose by NL01 was found to be much more rapid than that of ATCC 7050. Through comparison among NL01, ATCC 7050 and XZL4, it was clear that the above cellobiose operons were crucial to the cellobiose fermentation ability of B. coagulans strains. In addition, the CELO2 also seems to play a vital role in enhancing cellobiose metabolism, in spite of this operon lacking the component annotated as the 6-phospho-β-glucosidase encoding gene. Therefore, CELO2 might be an incomplete operon, which hastens the transportation and phosphorylation of cellobiose into cells, but lacks the ability to hydrolyse cellobiose-6-phosphate. Another possibility is that the DUF871 domain-containing protein is a novel 6-phospho-β-glucosidase, and CELO2 plays a stronger role in cellobiose uptake when in vivo.
Fig. 5.
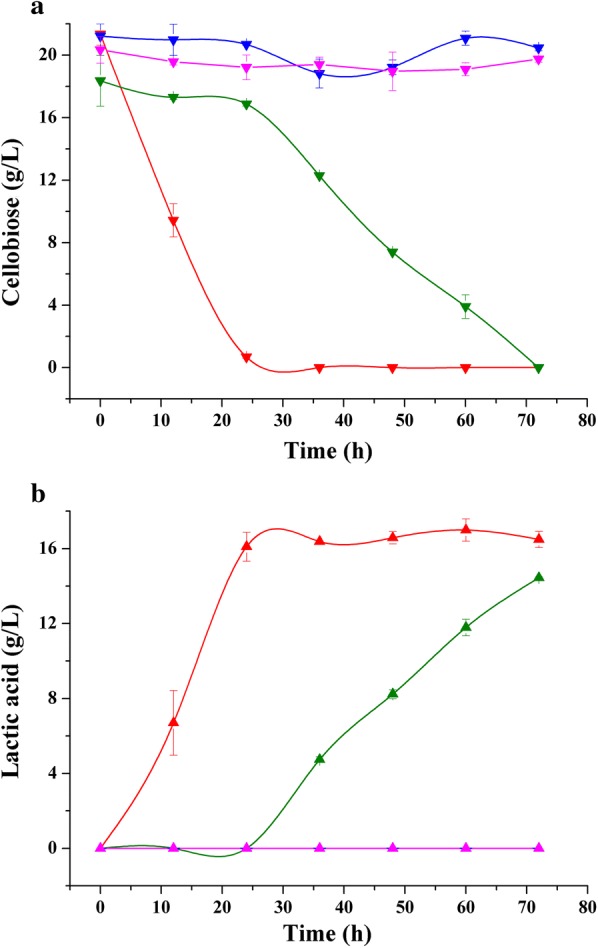
Cellobiose fermentation by B. coagulans NL01, ATCC 7050, XZL4, and ATCC 7050(ΔcelR). a Time courses of cellobiose consumption. b Time courses of lactic acid production. Red, B. coagulans NL01; Olive, B. coagulans ATCC 7050; Blue, B. coagulans XZL4; Magenta, B. coagulans ATCC 7050(ΔcelR). Fermentations were carried out in 250-mL Erlenmeyer flasks containing 100 mL medium at 50 °C and 150 rpm on a rotary shaker. The medium contained the following (g/L): cellobiose 20, yeast extract 2.5, corn syrup powder 1.2, MgSO4·7H2O 0.4, (NH4)2SO4 3, KH2PO4 0.22, MnSO4·H2O 0.03, FeSO4·H2O 0.03, and CaCO3 10
To clarify the specific role of CELO1 and CELO2 in vivo, different mutants of B. coagulans NL01 lacking CELO1, CELO2, and both CELO1 and CELO2 should be constructed as a prerequisite. However, B. coagulans was reported to be recalcitrant to genetic engineering [20, 21]. Up to now, genetic manipulation of most B. coagulans strains was currently unable to be performed, except B. coagulans ATCC 7050 and B. coagulans P4-102B [20, 21, 33]. Compared with B. coagulans P4-102B, B. coagulans ATCC 7050 is a standard and genome-sequenced strain and has been knocked out by several different research groups. Therefore, only the CELO1 mutant derived from B. coagulans ATCC 7050 can presently be constructed in our lab. It is significant, however, that the PEP-PTS components of CELO1 from B. coagulans NL01 and B. coagulans ATCC 7050 share a high level of identity, although two extra transposase genes exist in ATCC 7050. The protein sequences of IIA, IIB, IIC, 6-phospho-β-glucosidase, and the transcriptional regulator from B. coagulans NL01 were found to be 96%, 96%, 99%, 96%, and 94% homologous with those from B. coagulans ATCC 7050, respectively. Given these facts, the CELO1 mutant of B. coagulans ATCC 7050 was used as an alternative for further studying the function of B. coagulans’ cellobiose operon in vivo.
Because CELO1 in B. coagulans ATCC 7050 is composed of five different and long components (up to 10,397 bp), full knockout of CELO1 was very difficult. Instead, only celR was selected as a knockout target to obtain a mutant strain of B. coagulans ATCC 7050 (ΔcelR). Analogously, fermentation of cellobiose by this mutant was carried out and the results are shown in Fig. 5. The fermentation profile of B. coagulans ATCC 7050 (ΔcelR) was similar to that of B. coagulans XZL4, which indicates that the incomplete CELO1 lost the function of cellobiose utilization by partial gene knockout. In NCBI, celR of B. coagulans is annotated as a PRD-containing protein, which means that this protein contains a PTS regulation domain (PRD). So far, the PRD-containing transcription regulators are divided into two groups: antiterminators and transcription activators, and the latter can be further divided into two classes, NifA/NtrC- and DeoR-like PRD-containing transcription activators [34]. In some sequenced bacterial genomes, many transcription activators are incorrectly referred to as antiterminators [34]. This is also the case in B. coagulans genomes. The PRDs of B. coagulans should be DeoR-like because the molecular weight of different types of PRD-containing transcription regulators is notably different: 30 kDa for antiterminators, 65 kDa for DeoR-like, and 90–100 kDa for NifA/NtrC-like transcription activators [34]. The result of B. coagulans ATCC 7050 (ΔcelR) in Fig. 5 also certifies that celR is an indispensable component for CELO1 to perform its function.
Although the CELO1 and CELO2 of B. coagulans NL01 could not be mutated in vivo, they could be cloned and expressed in heterologous hosts. E. coli was reported to be without cellobiose consumption ability except for when it was cultivated using cellobiose as the sole carbon source for a long time. This observation was believed to be due to the cryptic nature of the bgl and cel operons [35, 36]. Hence, two recombinant strains, E. coli BL21(pET-celo1) and E. coli BL21(pET-celo2) were constructed to investigate whether CELO1 and CELO2 could enable a genetically modified E. coli to ferment cellobiose. An E. coli strain bearing an empty plasmid (pETDuet-1) was designated as E. coli BL21(pETDuet-1) and used as control. The cellobiose utilizing capacities of the three engineered strains were tested in shaker flasks. Figure 6 showed that both E. coli BL21(pET-celo1) and E. coli BL21(pET-celo2) had better cellobiose metabolism than E. coli BL21(pETDuet-1). Both E. coli BL21(pET-celo1) and E. coli BL21(pET-celo2) consumed almost all added cellobiose, which indicated that CELO2 is a complete operon and may have a great influence on cellobiose catabolism for B. coagulans NL01. In contrast, E. coli BL21(pETDuet-1) did not show significant cellobiose consumption. Accordingly, the cell growth of E. coli BL21(pETDuet-1) was much weaker compared with E. coli BL21(pET-celo1) and E. coli BL21(pET-celo2).
Fig. 6.
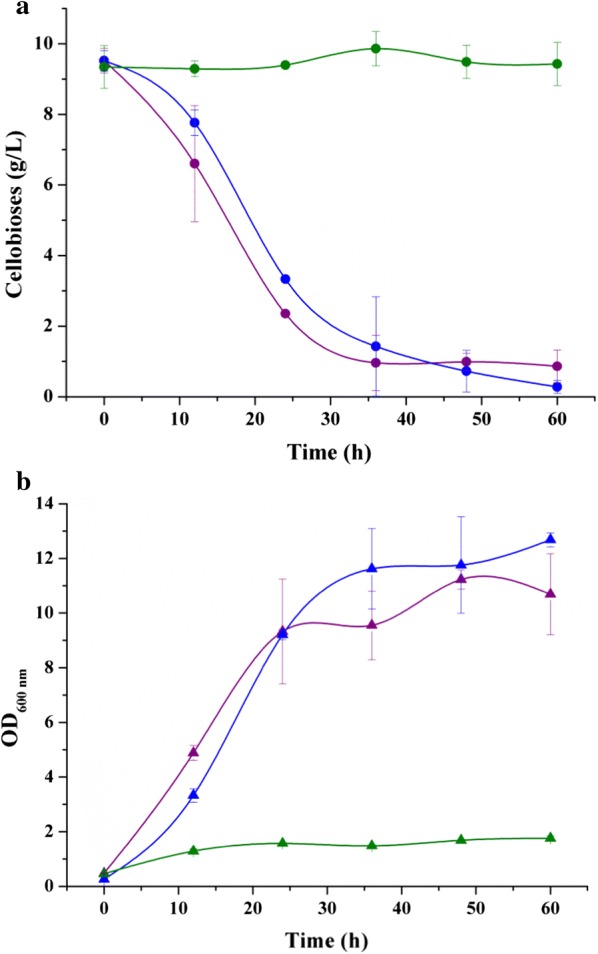
Cellobiose fermentation of E. coli BL21(pET-celo1), E. coli BL21(pET-celo2) and E. coli BL21(pETDuet-1). a Time courses of cellobiose consumption. b Time course of optical density. Purple, E. coli BL21(pET-celo1); Blue, E. coli BL21(pET-celo2); Olive, E. coli BL21(pETDuet-1). Fermentations were carried out in 100-mL Erlenmeyer flasks containing 30-mL LB medium in the presence of 10 g/L cellobiose, 100 mg/L ampicillin, and 0.2 mM IPTG at 37 °C and 200 rpm on a rotary shaker
Due to the lack of a mature tool for genetic engineering of B. coagulans NL01, knockout of CELO1 and CELO2 in NL01 is different to execute now. However, this study has provided a new research field in terms of increasing our understanding of cellobiose metabolism in B. coagulans. Continuing the study of CELO2 and DUF871 domain-containing protein in B. coagulans NL01 might provide new insights into the diversity of microbial cellobiose utilization mechanisms and the diversity of 6-phospho-β-glucosidase.
Conclusions
In this study, a wild type of B. coagulans (NL01) was found to employ cellobiose-specific PEP-PTS for translocation and phosphorylation of cellobiose, and intracellular hydrolysis of cellobiose-6-phosphate. With the aid of this unique strategy for intracellular assimilation of cellobiose, co-fermentation of cellobiose and xylose was successfully performed to circumvent glucose repression. Moreover, two different cellobiose operons were identified by comparative genomic analysis and the role of CELO1 and CELO2 was certified, respectively. Overall, B. coagulans NL01 would be recognized as a robust microorganism for a biorefinery process and not be limited to lactic acid production. The cellobiose operon described herein can be a promising prototype for development of alternative approach aimed at circumventing the sophisticated glucose repression.
Additional file
Additional file 1: Table S1. The primers used for RT-PCR.
Authors’ contributions
All authors contributed to the conception and planning of the study. ZZ, TJ, LZ, SOY, JZ, XL, QH executed the experimental work. ZZ, TJ, HX, and JOY analyzed the data. ZZ, LM, BY, HX, and JOY contributed regents and materials. ZZ, TJ and JOY wrote and revised the manuscript. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Consent for publication
All authors consented on the publication of this work.
Ethics approval and consent to participate
Not applicable.
Funding
We thank Dr. Ping Xu from Shanghai Jiao Tong University for offering B. coagulans XZL4, and Dr. Oscar P. Kuipers from University of Groningen for offering pMH77. This study was supported by the National Natural Science Foundation of China (51561145015, 31300487), the Major Program of the Natural Science Foundation of Jiangsu Higher Education of China (16KJA220004). We also kindly acknowledge partial support from the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- SSF
simultaneous saccharification and fermentation
- LB
Luria Broth
- RT-PCR
reverse transcription PCR
- PEP-PTS
phosphoenolpyruvate-dependent phosphotransferase system
- CELO1
cellobiose operon 1
- CELO2
cellobiose operon 2
- pbgl
6-phospho-β-glucosidase gene
- PRD
PTS regulation domain
References
- 1.Jiang T, Qiao H, Zheng Z, Chu Q, Li X, Qiang Y, Jia O. Lactic acid production from pretreated hydrolysates of corn stover by a newly developed Bacillus coagulans strain. PLoS ONE. 2016;11:e0149101. doi: 10.1371/journal.pone.0149101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Huang C, Ragauskas AJ, Wu X, Huang Y, Zhou X, He J, Huang C, Lai C, Li X, Yong Q. Co-production of bio-ethanol, xylonic acid and slow-release nitrogen fertilizer from low-cost straw pulping solid residue. Bioresour Technol. 2017;250:365–373. doi: 10.1016/j.biortech.2017.11.060. [DOI] [PubMed] [Google Scholar]
- 3.Huang C, Jeuck B, Du J, Yong Q, Chang HM, Jameel H, Phillips R. Novel process for the coproduction of xylo-oligosaccharides, fermentable sugars, and lignosulfonates from hardwood. Bioresour Technol. 2016;219:600–607. doi: 10.1016/j.biortech.2016.08.051. [DOI] [PubMed] [Google Scholar]
- 4.Li X, Yang L, Gu X, Lai C, Huang C, Yong Q. A combined process for production of fumaric acid and xylooligosaccharides from corncob. BioResources. 2017;13:399–411. [Google Scholar]
- 5.Limayem A, Ricke SC. Lignocellulosic biomass for bioethanol production: current perspectives, potential issues and future prospects. Prog Energy Combust Sci. 2012;38:449–467. doi: 10.1016/j.pecs.2012.03.002. [DOI] [Google Scholar]
- 6.Zheng Z, Cai C, Jiang T, Zhao M, Ouyang J. Enhanced l-lactic acid production from biomass-derived xylose by a mutant Bacillus coagulans. Appl Biochem Biotechnol. 2014;173:1896–1906. doi: 10.1007/s12010-014-0975-2. [DOI] [PubMed] [Google Scholar]
- 7.Wu Y, Shen X, Yuan Q, Yan Y. Metabolic engineering strategies for co-utilization of carbon sources in microbes. Bioengineering. 2016;3:10. doi: 10.3390/bioengineering3010010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kim SR, Ha SJ, Wei N, Oh EJ, Jin YS. Simultaneous co-fermentation of mixed sugars: a promising strategy for producing cellulosic ethanol. Trends Biotechnol. 2012;30:274–282. doi: 10.1016/j.tibtech.2012.01.005. [DOI] [PubMed] [Google Scholar]
- 9.Chomvong K, Kordić V, Li X, Bauer S, Gillespie AE, Ha SJ, Oh EJ, Galazka JM, Jin YS, Cate JH. Overcoming inefficient cellobiose fermentation by cellobiose phosphorylase in the presence of xylose. Biotechnol Biofuels. 2014;7:1–11. doi: 10.1186/1754-6834-7-85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wu Y, Yang Y, Ren C, Chen Y, Yang S, Yang G, Jiang W. Molecular modulation of pleiotropic regulator CcpA for glucose and xylose coutilization by solvent-producing Clostridium acetobutylicum. Metab Eng. 2015;28:169. doi: 10.1016/j.ymben.2015.01.006. [DOI] [PubMed] [Google Scholar]
- 11.Bruder M, Moo-Young M, Chung DA, Chou CP. Elimination of carbon catabolite repression in Clostridium acetobutylicum-a journey toward simultaneous use of xylose and glucose. Appl Microbiol Biotechnol. 2015;99:7579–7588. doi: 10.1007/s00253-015-6611-4. [DOI] [PubMed] [Google Scholar]
- 12.Nakamura N, Yamada R, Katahira S, Tanaka T, Fukuda H, Kondo A. Effective xylose/cellobiose co-fermentation and ethanol production by xylose-assimilating S. cerevisiae via expression of β-glucosidase on its cell surface. Enzyme Microb Technol. 2008;43:233–236. doi: 10.1016/j.enzmictec.2008.04.003. [DOI] [Google Scholar]
- 13.Parisutham V, Sathesh-Prabu C, Mukhopadhyay A, Lee SK, Keasling JD. Intracellular cellobiose metabolism and its applications in lignocellulose-based biorefineries. Bioresour Technol. 2017;239:496–506. doi: 10.1016/j.biortech.2017.05.001. [DOI] [PubMed] [Google Scholar]
- 14.Ha SJ, Galazka JM, Kim SR, Choi JH, Yang X, Seo JH, Glass NL, Cate JH, Jin YS. Engineered Saccharomyces cerevisiae capable of simultaneous cellobiose and xylose fermentation. Proc Natl Acad Sci USA. 2011;108:504–509. doi: 10.1073/pnas.1010456108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Shin HD, Wu J, Chen R. Comparative engineering of Escherichia coli for cellobiose utilization: hydrolysis versus phosphorolysis. Metab Eng. 2014;24:9–17. doi: 10.1016/j.ymben.2014.04.002. [DOI] [PubMed] [Google Scholar]
- 16.Ouyang J, Ma R, Zheng ZJ, Cai C, Zhang M, Jiang T. Open fermentative production of L-lactic acid by Bacillus sp. strain NL01 using lignocellulosic hydrolyzates as low-cost raw material. Bioresour Technol. 2013;135:475–480. doi: 10.1016/j.biortech.2012.09.096. [DOI] [PubMed] [Google Scholar]
- 17.Zhang C, Zhou C, Assavasirijinda N, Yu B, Wang L, Ma Y. Non-sterilized fermentation of high optically pure d-lactic acid by a genetically modified thermophilic Bacillus coagulans strain. Microb Cell Fact. 2017;16:213. doi: 10.1186/s12934-017-0827-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zheng Z, Xi L, Jiang T, Ye W, Ouyang J. Genomic analysis of a xylose operon and characterization of novel xylose isomerase and xylulokinase from Bacillus coagulans NL01. Biotechnol Lett. 2016;38:1331–1339. doi: 10.1007/s10529-016-2109-0. [DOI] [PubMed] [Google Scholar]
- 19.Zhou J, Ouyang J, Xu Q, Zheng Z. Cost-effective simultaneous saccharification and fermentation of l-lactic acid from bagasse sulfite pulp by Bacillus coagulans CC17. Bioresour Technol. 2016;222:431–438. doi: 10.1016/j.biortech.2016.09.119. [DOI] [PubMed] [Google Scholar]
- 20.Kovács AT, Van HM, Kuipers OP, Van KR. Genetic tool development for a new host for biotechnology, the thermotolerant bacterium Bacillus coagulans. Appl Environ Microbiol. 2010;76:4085–4088. doi: 10.1128/AEM.03060-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sun L, Zhang C, Lyu P, Wang Y, Wang L, Yu B. Contributory roles of two L-lactate dehydrogenases for l-lactic acid production in thermotolerant Bacillus coagulans. Sci Rep. 2016;6:37916. doi: 10.1038/srep37916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ou MS, Mohammed N, Ingram LO, Shanmugam KT. Thermophilic Bacillus coagulans requires less cellulases for simultaneous saccharification and fermentation of cellulose to products than mesophilic microbial biocatalysts. Appl Biochem Biotechnol. 2009;155:76–82. doi: 10.1007/s12010-008-8509-4. [DOI] [PubMed] [Google Scholar]
- 23.Hu JL, Zhang ZT, Lin YX, Zhao SM, Mei YX, Liang YX, Peng N. High-titer lactic acid production from NaOH-pretreated corn stover by Bacillus coagulans LA204 using fed-batch simultaneous saccharification and fermentation under non-sterile condition. Bioresour Technol. 2015;182:251–257. doi: 10.1016/j.biortech.2015.02.008. [DOI] [PubMed] [Google Scholar]
- 24.Zhang YM. An efficient process for lactic acid production from wheat straw by a newly isolated Bacillus coagulans strain IPE22. Bioresour Technol. 2014;158:396. doi: 10.1016/j.biortech.2014.02.128. [DOI] [PubMed] [Google Scholar]
- 25.Ong SA, Ng ZJ, Wu JC. Production of high concentration of l-lactic acid from cellobiose by thermophilic Bacillus coagulans WCP10-4. Appl Microbiol Biotechnol. 2016;100:6501–6508. doi: 10.1007/s00253-016-7600-y. [DOI] [PubMed] [Google Scholar]
- 26.Ouyang J, Cai C, Chen H, Jiang T, Zheng Z. Efficient non-sterilized fermentation of biomass-derived xylose to lactic acid by a thermotolerant Bacillus coagulans NL01. Appl Biochem Biotechnol. 2012;168:2387–2397. doi: 10.1007/s12010-012-9944-9. [DOI] [PubMed] [Google Scholar]
- 27.Rutter C, Chen R. Improved cellobiose utilization in E. coli by including both hydrolysis and phosphorolysis mechanisms. Biotechnol Lett. 2014;36:301–307. doi: 10.1007/s10529-013-1355-7. [DOI] [PubMed] [Google Scholar]
- 28.Jamal Z, Miotsertier C, Thibau F, Dutilh L, Lonvaudfunel A, Ballestra P, Marrec CL, Dolslafargue M. Distribution and functions of phosphotransferase system genes in the genome of the lactic acid bacterium Oenococcus oeni. Appl Environ Microbiol. 2013;79:3371. doi: 10.1128/AEM.00380-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Viana R, Monedero V, Dossonnet V, Vadeboncoeur C, Pérezmartínez G, Deutscher J. Enzyme I and HPr from Lactobacillus casei: their role in sugar transport, carbon catabolite repression and inducer exclusion. Mol Microbiol. 2000;36:570–584. doi: 10.1046/j.1365-2958.2000.01862.x. [DOI] [PubMed] [Google Scholar]
- 30.Deutscher J, Aké FM, Derkaoui M, Zébré AC, Cao TN, Bouraoui H, Kentache T, Mokhtari A, Milohanic E, Joyet P. The bacterial phosphoenolpyruvate: carbohydrate phosphotransferase system: regulation by protein phosphorylation and phosphorylation-dependent protein-protein interactions. Microbiol Mol Biol Rev. 2014;78:231–256. doi: 10.1128/MMBR.00001-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zeng L, Burne RA. Transcriptional regulation of the cellobiose operon of Streptococcus mutans. J Bacteriol. 2009;191:2153–2162. doi: 10.1128/JB.01641-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Michlmayr H, Kneifel W. β-Glucosidase activities of lactic acid bacteria: mechanisms, impact on fermented food and human health. FEMS Microbiol Lett. 2014;352:1–10. doi: 10.1111/1574-6968.12348. [DOI] [PubMed] [Google Scholar]
- 33.Rhee MS, Kim JW, Qian Y, Ingram LO, Shanmugam KT. Development of plasmid vector and electroporation condition for gene transfer in sporogenic lactic acid bacterium, Bacillus coagulans. Plasmid. 2007;58:13–22. doi: 10.1016/j.plasmid.2006.11.006. [DOI] [PubMed] [Google Scholar]
- 34.Joyet P, Bouraoui H, Aké FMD, Derkaoui M, Zébré AC, Cao TN, Ventroux M, Nessler S, Noirot-Gros MF, Deutscher J. Transcription regulators controlled by interaction with enzyme IIB components of the phosphoenolpyruvate: sugar phosphotransferase system. Biochim Biophys Acta. 2013;1834:1415–1424. doi: 10.1016/j.bbapap.2013.01.004. [DOI] [PubMed] [Google Scholar]
- 35.Parker LL, Hall BG. Mechanisms of activation of the cryptic cel operon of Escherichia coli K12. Genetics. 1990;124:473. doi: 10.1093/genetics/124.3.473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sekar R, Shin HD, Chen R. Engineering Escherichia coli cells for cellobiose assimilation through a phosphorolytic mechanism. Appl Environ Microbiol. 2012;78:1611–1614. doi: 10.1128/AEM.06693-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Table S1. The primers used for RT-PCR.
Data Availability Statement
All data generated or analyzed during this study are included in this published article.