Shade avoidance involves complex regulation of multiple hormone network modules, and salicylic acid pathway genes are required for petiole shade avoidance.
Abstract
Plants have sophisticated mechanisms for sensing neighbor shade. To maximize their ability to compete for light, plants respond to shade through enhanced elongation and physiological changes. The shade avoidance response affects many different organs and growth stages, yet the signaling pathways underlying this response have mostly been studied in seedlings. We assayed transcriptome changes in response to shade across a 2-d time course in the wild type and 12 Arabidopsis (Arabidopsis thaliana) mutants. The resulting temporal map of transcriptional responses to shade defines early and late responses in adult plants, enabling us to determine connections between key signaling genes and downstream responses. We found a pervasive and unexpectedly strong connection between shade avoidance and genes related to salicylic acid, suggesting salicylic acid signaling to be an important shade avoidance growth regulator. We tested this connection and found that several mutants disrupting salicylic acid levels or signaling were defective in shade avoidance. The effect of these mutations on shade avoidance was specific to petiole elongation; neither hypocotyl nor flowering time responses were altered, thereby defining important stage-specific differences in the downstream shade avoidance signaling pathway. Shade treatment did not change salicylic acid levels, indicating that the mediation of shade avoidance by salicylic acid is not dependent on the modulation of salicylic acid levels. These results demonstrate that salicylic acid pathway genes also are key components of petiole shade avoidance.
Plants perceiving neighbor shade exhibit a suite of developmental and physiological responses known as the shade avoidance syndrome (SAS). SAS includes promotion of organ growth, acceleration of flowering time, suppression of shoot branching, and differential resource allocation. In agricultural settings, dense planting can cause shade avoidance, reducing yields due to changes in carbon allocation that favor stem elongation over seed, fruit, or tuber production (Boccalandro et al., 2003; Chincinska et al., 2008). Therefore, it is important to understand shade avoidance signaling pathways throughout the lifespan of the plant to develop strategies for maximizing yield in dense plantings.
Canopy shade is perceived by phytochrome and cryptochrome photoreceptors that initiate signaling through an extensive signaling network (Casal, 2013). Light transmitted through or reflected from leaves has reduced red to far-red light (R/FR). This low R/FR inactivates phytochrome, allowing the accumulation of phytochrome-interacting factor (PIF) transcription factors. PIFs, in turn, activate the transcription of YUCCA (YUC) auxin biosynthesis genes, thereby increasing auxin production and growth. Indeed, in the case of the hypocotyl, rapid auxin production is seen within a few hours of shade treatment, and growth promotion is seen with similar timing, although there is additional growth promotion at later time points (Cole et al., 2011). Both phases of growth are dependent on an increase in YUC transcripts by PIF7 (Li et al., 2012). In addition to increasing auxin production, shade also increases auxin sensitivity (Hersch et al., 2014; de Wit et al., 2015). In shade, cyptochromes detect the decrease in blue light and interact physically with PIFs to alter PIF activity (Pedmale et al., 2016).
As part of the shade avoidance response, plants appear to prioritize growth over defense to compete with neighbors for light (Ballaré, 2014). Extreme examples that demonstrate prioritized growth and reduced defense are phytochrome-deficient mutants in Arabidopsis (Arabidopsis thaliana; Genoud et al., 2002; de Wit et al., 2013) and tomato (Solanum lycopersicum; Izaguirre et al., 2006) that show both constitutive shade avoidance and greater susceptibility to pathogen attack than the wild type. Low R/FR reduces plant immunity by modulating the jasmonic acid and salicylic acid hormone defense pathways (Ballaré, 2014). Resistance against a hemibiotrophic pathogen (Pseudomonas syringae pv tomato) and a necrotrophic pathogen (Botrytis cinerea) was found to be suppressed by shade treatment (Cerrudo et al., 2012; de Wit et al., 2013). The reduced resistance to B. cinerea under shade was dependent on jasmonic acid signaling but independent of salicylic acid signaling (Cerrudo et al., 2012). Low R/FR also has been shown to reduce downstream transcriptional responsiveness to jasmonic acid and salicylic acid (Cerrudo et al., 2012; de Wit et al., 2013; Leone et al., 2014). While most previous work has focused on how shade reduces defense, we recently found that jasmonic acid pathway genes are required for full growth responses to low R/FR (Nozue et al., 2015). Although low R/FR also reduces salicylic acid signaling, it is unknown whether salicylic acid signaling components are important for growth responses to low R/FR.
Most studies on shade avoidance signaling pathways have focused on hypocotyl elongation in young seedlings. Given that shade avoidance responses are observed in many different organs throughout the plant life cycle, it is important to ask if there are different downstream signaling pathways operating in different organs or life stages. Our genetic analysis revealed multiple adult-specific shade avoidance genes (Nozue et al., 2015); here, we use time-course transcriptome profiling of the wild type and 12 shade avoidance mutants to dissect the genetic and temporal control of shade-induced transcriptome changes in mature plants. These data also allow us to address an additional unsolved challenge: understanding the temporal activation sequence of hormone pathways during shade avoidance. For example, even though the up-regulation of auxin pathway genes is a hallmark of SAS, here we found that many auxin pathway-related and other early-response genes were expressed normally in the constitutively shade-avoiding phytochrome B (phyB) mutant. The time gap between the rapid regulation of auxin pathways and constitutive shade avoidance responses in phyB illustrates the importance of understanding how temporal changes of SAS genes upon shade treatment relate to long-term growth responses. We found that auxin, jasmonic acid, and salicylic acid hormone pathways are each regulated by low R/FR but with distinct temporal patterns. Surprisingly, we found that transcriptional network modules related to salicylic acid were more sensitive to mutant perturbation than auxin network modules, suggesting a connection between salicylic acid signaling and growth control. Testing this idea revealed that several mutants with altered salicylic acid levels or signaling are defective in petiole shade avoidance, demonstrating a role for salicylic acid pathway components in shade avoidance growth control.
RESULTS AND DISCUSSION
Auxin Production Is Increased Transiently But Not Sustained
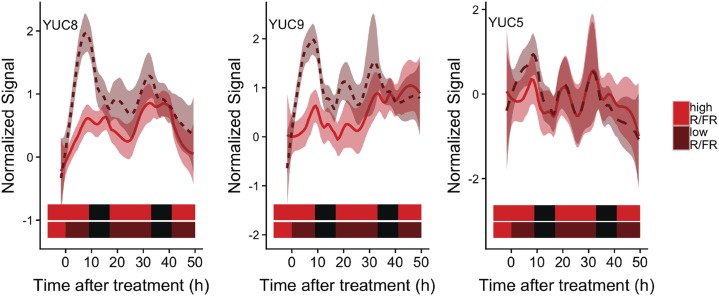
Low R/FR light treatment inactivates phytochrome photoreceptors, up-regulating auxin-regulated pathways in seedlings (Sessa et al., 2005; Tao et al., 2008) and the apical region of young adult plants (Nozue et al., 2015). Furthermore, auxin is indispensable for full SAS (Nozue et al., 2015; Müller-Moulé et al., 2016). However, examining published data (Hu et al., 2013), we found no enrichment of auxin pathway genes among phyA/B/D/E differentially expressed genes (Supplemental Fig. S1A), and most shade-induced auxin pathway genes (Sessa et al., 2005) are not differentially expressed in phyA/B/D/E mutants. The lack of auxin gene up-regulation in the phyA/B/D/E mutant is surprising, since these plants exhibit a strong, constitutive SAS phenotype (Franklin et al., 2003). Similarly, auxin response genes were not enriched among differentially expressed genes after 1 d of low R/FR treatment (Ciolfi et al., 2013). These discrepancies raised a question: is auxin synthesis through YUC biosynthesis genes only transiently activated during SAS? To answer this question, we used YUC promoter::LUCIFERASE2 (LUC2) reporter constructs to monitor YUC activation during SAS. YUC8 and YUC9 promoter activity increased upon low R/FR treatment, peaking 4 to 7 h after the start of the shade treatment and then dissipating such that there were no differences between low R/FR and high R/FR by day 2. In contrast, YUC5 promoter activity was not influenced by low R/FR treatment (Fig. 1), consistent with previous transcriptome data from low R/FR-treated leaves (Nozue et al., 2015). Since the induction of YUC5 was observed in 8-d-old seedlings (Müller-Moulé et al., 2016), YUC5 might be specific to seedling shade avoidance responses. Our data indicate that auxin production transiently increases upon low R/FR treatment, but up-regulation is sustained only on day 1 of the treatment, thereby explaining why the constitutive shade avoidance phy quadruple mutant did not show the activation of auxin pathway genes.
Figure 1.
Auxin biosynthetic genes are transiently induced by shade. Luciferase was driven by promoters of auxin biosynthesis genes that are involved in the SAS (YUC8, YUC9, and YUC5) and monitored upon low R/FR treatment for 2 d. Luminescence from individual plants was normalized by dividing by luminescence at Zeitgeber time 7 (ZT7; just before low R/FR treatment). Red, brown, and black horizontal bars indicate high R/FR, low R/FR, and dark, respectively. The solid red lines (high R/FR) and dashed brown lines (low R/FR) indicate localized regression (loess) smoothing of data from 13 to 25 plants; shading indicates 95% confidence interval. y axis units are arbitrary.
Each Hormone Pathway Has a Distinct Temporal Response to Shade
Having found that auxin pathway activation is transient, we wanted to determine subsequent, longer term transcriptional responses to shade. To address this question, we used a time-course transcriptome analysis to define shade-responsive gene expression changes. We assayed the response in wild-type (Columbia [Col]) plants and then used a series of 12 mutant lines (Table 1) to define important determinants of the wild-type response.
Table 1. Genotypes used for RNA sequencing.
| Genotype | Gene Name (Arabidopsis Genome Initiative) | Purpose | Reference |
|---|---|---|---|
| aos | AOS (AT5G42650) | Known jasmonic acid-deficient mutant | Park et al. (2002) |
| co-9 | CO (AT5G15840) | Impaired flowering time pathway | Balasubramanian et al. (2006) |
| Col | – | Wild type | – |
| hy5-215 | HY5 (AT5G11260) | Shade avoidance mutant | Holm et al. (2002) |
| jar1 | JAR1 (AT2G46370) | Shade avoidance and jasmonic acid-deficient mutant | Guranowski et al. (2007) |
| kat1-1 | KAT1 (AT5G46240) | Shade avoidance and potassium channel mutant | Nozue et al. (2015) |
| mida9-4 | MIDA9 (AT5G02760) | Shade avoidance mutant | Nozue et al. (2015) |
| PAR1 RNAi | PAR1 (AT2G42870) | Shade avoidance mutant | Roig-Villanova et al. (2007) |
| phyB-9 | PHYB (AT2G18790) | Shade avoidance mutant | Reed et al. (1993) |
| pif3-1 | PIF3 (AT1G09530) | Shade avoidance mutant | Kim et al. (2003) |
| pif4/5 | PIF4 (AT2G43010) | Shade avoidance mutant | Lorrain et al. (2008) |
| PIF5 (AT3G59060) | |||
| spt-11 | SPT (AT4G36930) | Shade avoidance mutant | Ichihashi et al. (2010) |
| yuc2/5/8/9 | YUC2 (AT4G13260) | Shade avoidance mutant | Nozue et al. (2015); Müller-Moulé et al. (2016) |
| YUC5 (AT5G43890) | |||
| YUC8 (AT4G28720) | |||
| YUC9 (AT1G04180) |
Our first analysis focused on low-R/FR-responsive genes because low-R/FR-responsive genes contain a high proportion of functionally important SAS genes (Nozue et al., 2015). Specifically, we used the 276 genes that were low R/FR responsive in at least one genotype (Col or the 12 mutants; Table 1). We used t-distributed stochastic noise embedding (t-SNE) to define clusters with a distinct time-course pattern of responsiveness to low R/FR (Supplemental Fig. S2; Supplemental Data Set S1). Characteristic features of each cluster in Col were determined by enrichment analysis with Gene Ontology (GO) terms and a curated list of hormone-responsive genes and known transcription factor targets (see “Materials and Methods”; Supplemental Data Sets S2–S5; Supplemental Fig. S3). To better illustrate the temporal shade response, we focused on genes present in curated hormone/transcription factor categories that had significant overlap with the t-SNE clusters (Fig. 2A; Supplemental Data Set S6). To account for possible stochastic variation and the effect of starting parameters on clustering runs, we repeated these procedures 10 times to identify common results (Supplemental Data Sets S1–S6; for details, see “Materials and Methods”).
Figure 2.
Time-course visualization of shade-responsive genes in custom categories. For each custom category, all genes enriched in t-SNE clusters (P < 0.05 in Supplemental Fig. S3) were selected, and their mean shade responsiveness for each genotype was visualized in heat maps for the wild type (A) and for each mutant (B). Log2 fold change is shown: magenta and green represent up- and down-regulated, respectively. Asterisks indicate significantly different responses for the category of genes compared with Col (false discovery rate [FDR] < 0.0001). Minpts of 13 and epsilon of 3.25 were used for this figure. ABAup or ABAdown, abscisic acid up- or down-regulated; ACCup or ACCdown, 1-aminocyclopropane-1-carboxylic acid up- or down-regulated; BLup or BLdown, blassinolide up- or down-regulated; IAAup or IAAdown, indole-3-acetic acid (IAA) up- or down-regulated; MJup or MJdown, methyl jasmonate (MJ) up- or down-regulated; MYC234up or MYC234down, up- or down-regulated by MYC234 genes; PIFtarget, PIF target genes; SAup or SAdown, salicylic acid up- or down-regulated. Details are given in Supplemental Data Set S3.
Consistent with the behavior of YUCp::LUC2 activity (Fig. 1), auxin-up-regulated genes were up-regulated by R/FR at the 1-h time point and returned to baseline by the 49-h time point (Fig. 2A; Supplemental Data Set S7). Interestingly, no YUC genes were found in these clusters. The lack of YUC genes is likely due to the leaf blade- or cotyledon-specific (Nito et al., 2015) induction of these genes, while our transcriptome data are derived from the apical region excluding the first and second leaves (Supplemental Fig. S4). In agreement with this observation, it has been shown that auxin generated in cotyledon under low R/FR is required for low-R/FR-induced hypocotyl growth or gene expression (Kozuka et al., 2010; Procko et al., 2014; Nito et al., 2015).
MJ-up-regulated genes showed a unique pattern: repression specifically at 4 h. Interestingly, this matches the pattern of MYC2/3/4-up-regulated genes (Fig. 2A; Supplemental Data Set S7). Because MYC2/3/4 positively regulates jasmonic acid response genes (Kazan and Manners, 2013), this co-occurrence suggests that low R/FR represses MJ-up-regulated genes through the inhibition of MYC2/3/4 transcription factors. The pattern of MJ-down-regulated genes does not match that of MYC2/3/4-regulated genes, suggesting that these genes are not regulated through MYC2/3/4 in our condition.
In contrast to auxin- and jasmonic acid-responsive genes, genes responsive to salicylic acid were repressed at a later time point (49 h) in adult plants. It should be noted that the repression of salicylic acid pathway genes was not seen in phyA/B/D/E mutants assayed as seedlings (Hu et al., 2013), which reflects the well-known developmental regulation of the salicylic acid pathway (Gaffney et al., 1993; Xie et al., 2011) and highlights the importance of examining later plant stages, as we did here.
Distinct Patterns of Shade-Responsive Genes in Shade Avoidance Mutants
Having defined the wild-type temporal response to shade, next we wanted to determine how genes in the SAS pathway influence downstream gene expression. To do this, we determined which genes were misexpressed in response to low R/FR in each of 10 SAS mutants as compared with the wild type. Specifically, we focused on how the temporal pattern of shade-responsive gene expression of each custom gene category (Fig. 2A; Supplemental Data Set S7) was altered in each mutant as compared with Col (Fig. 2B; Supplemental Data Set S7).
Most genotypes had significantly reduced responses as compared with the wild type in multiple categories. Shade-responsive IAA-up-regulated genes were induced rapidly in Col (Fig. 2A; Supplemental Data Set S7). This induction was reduced severely in yuc2/5/8/9, hy5, and phyB and reduced significantly in all mutants, indicating that IAA-up-regulated genes are a common target. Shade-responsive PIF target genes were affected in pif4/5, confirming that the shade responsiveness of those genes is mediated via PIF4/5. The responsiveness to shade of those genes also was reduced in yuc2/5/8/9, confirming a model in which auxin and PIF4/5 work in a common pathway in this response. It should be noted that, as a group, the expression of shade-responsive PIF targets in phyB was not statistically different from that of the wild type, even though the shade induction of IAA-up-regulated genes was reduced in phyB. However, a subset of shade-responsive PIF targets did have reduced shade responses in phyB (Supplemental Fig. S5). This result suggests that there is a subset of PIF targets whose shade response is regulated by phytochrome photoreceptors other than phyB and that this subset of PIF targets is not important for the shade induction of IAA-up-regulated genes.
Examining the defense-related gene categories, in wild-type plants we found that shade-responsive MJ-up-regulated genes were repressed by early low-R/FR treatment (4 h; Fig. 2A; Supplemental Data Set S7). AOS encodes an enzyme that synthesizes a precursor of jasmonic acid (Laudert and Weiler, 1998), while JAR1 encodes an enzyme that conjugates Ile to jasmonic acid, and JA-Ile is the major active jasmonic acid (Staswick and Tiryaki, 2004; Westfall et al., 2012). Therefore, one would expect that mutations in these genes would have similar effects. Surprisingly, while jar1 showed significantly altered responsiveness to shade compared with Col, the alteration could not be explained simply by a deficiency of active jasmonic acid because aos did not show a significant defect. The degree of alteration in aos and jar1 is consistent with their SAS phenotype: jar1 showed reduced SAS while aos did not (Nozue et al., 2015). In the auxin-deficient mutant yuc2/5/8/9, the pattern of MJ up-regulation was not altered, indicating that the jasmonic acid pathway is upstream or independent of the auxin pathways in SAS. However, since jasmonic biosynthesis mutants (aos and jar1) showed altered auxin-responsive gene expression patterns, there is an interaction between jasmonic acid and auxin pathways in SAS. Surprisingly, shade-responsive MJ-up-regulated genes were constitutively up-regulated in low-R/FR-grown phyB, a pattern not observed in other SAS mutants.
The responsiveness of salicylic acid-up-regulated genes to low R/FR was altered in only a subset of mutants (jar1, kat1, mada9, and spt11). In some mutants, the shade responsiveness of abscisic acid- or ethylene-responsive genes was altered. At present, the link between abscisic acid or ethylene and shade avoidance is limited (Reddy et al., 2013).
In conclusion, this analysis has identified specific gene categories whose shade response is defective in the various mutants and revealed that the gene expression pattern of custom categories (Supplemental Data Set S6) could be useful as biomarkers of shade avoidance.
Distinct Coexpression Modules Are Affected in Morphological Shade Avoidance Mutants
The above analyses focused on low-R/FR-responsive genes. However, it is possible that SAS mutants also alter the expression of non-low-R/FR-responsive genes, and such genes also could be important for SAS. To find these genes, we first examined coexpressed gene modules in all of the wild-type Col samples (four time points and both light conditions). Because we were using actual expression values instead of shade responsiveness as input to clustering, we were able to meet the minimum sample requirement (20 samples) for performing a weighted gene coexpression network analysis (WGCNA; Langfelder and Horvath, 2008). WGCNA defined 28 gene expression modules, and we characterized each of them by examining the enrichment of GO terms (Supplemental Data Set S8), hormone-responsive genes, and selected gene targets (Fig. 3; Supplemental Data Set S9). To define the modules involved in SAS and understand their regulation, we determined which modules were significantly misexpressed in each of the SAS mutants (Fig. 4; Supplemental Data Set S10). We found that module salmon was altered significantly in nine of the 10 SAS mutants examined (eight of them were enriched for genes misexpressed in these mutants even under high R/FR), suggesting that this is a core downstream module regulated by SAS genes in adult plants.
Figure 3.
Overrepresentation analysis of WGCNA gene modules. P values for overrepresentation were calculated by GOseq (Young et al., 2010). The P values were transformed into −log10 (P value) and color coded from magenta (highly enriched) to white (no enrichment). Numbers in cells indicate the number of genes in the intersection between each custom category and WGCNA module.
Figure 4.
Shade avoidance mutants misexpress specific WGCNA gene modules. The map and associated numbers in cells display the intersection between genes in each WGCNA coexpression module and genes misexpressed in a given genotype at least at one time point. P values for overlap were calculated by GOseq (Young et al., 2010). The P values were transformed into −log10 (P value) and color coded from magenta (highly enriched) to white (no enrichment). Genotypes highlighted in red and brown indicate high R/FR and low R/FR, respectively.
The Salmon Module Is Related to Salicylic Acid Signaling
Overrepresentation analysis showed that the salmon module is related to salicylic acid (Fig. 3; Supplemental Data Set S5). Among the 155 salmon module genes, PATHOGEN AND CIRCADIAN CONTROLLED1 (PCC1) and one of its closest homologs, AT3G22235 (Sauerbrunn and Schlaich, 2004; Segarra et al., 2010), are particularly interesting, because PCC1 also was down-regulated by shade (belonging to the salicylic acid-up-regulated group in the t-SNE; Fig. 2; Supplemental Data Set S7) and the expression of PCC1 and its homolog AT3G22235 was altered in almost all SAS mutants (Supplemental Data Set S10). Furthermore, PCC1 is involved in defense (Segarra et al., 2010) and light-regulated growth (Mir and León, 2014); thus, PCC1 may be an important regulator of salmon module genes during SAS. Another interesting set of genes in the salmon module are six WRKY transcription factor genes (Phukan et al., 2016) known to be important for defense signaling, including WRKY33 (Andreasson et al., 2005; Birkenbihl et al., 2012), WRKY51 (Gao et al., 2011), and WRKY70 (Ulker et al., 2007), abiotic stress, including WRKY25, WRKY33, and WRKY46 (Li et al., 2011; Chen et al., 2017), and leaf senescence, including WRKY54 and WRKY70 (Ulker et al., 2007). Among these, WRKY46, WRKY54, and WRKY70 are particularly interesting because they are involved in brassinosteroid-regulated plant growth (Chen et al., 2017). Notably, genes regulated by WRKY33 also are enriched in the salmon module (Fig. 3), implying that WRKY33 is a hub gene controlling the expression of other salmon module genes (Birkenbihl et al., 2012). PHYTOALEXIN-DEFICIENT4, which encodes a lipase-like protein that is important for salicylic acid signaling, also is in the salmon module. The salmon module contains SYSTEMIC ACQUIRED RESISTANCE DEFICIENT1 (SARD1) and CALMODULIN BINDING PROTEIN60g (CBP60g), both of which have been shown to be important activators of the salicylic acid biosynthesis gene ISOCHORISMATE SYNTHASE1 (ICS1; Zhang et al., 2010; Wang et al., 2011, 2015). Another salicylic acid-related gene in the salmon module is ACCELERATED CELL DEATH6 (ACD6), involved in resistance to P. syringae (Lu et al., 2003).
Salicylic Acid Plays an Unexpected Role in Adult SAS
The extensive misregulation of salicylic acid pathway genes in SAS mutants suggests that the salicylic acid signaling pathway might be important for growth during SAS. To determine if salicylic acid pathway genes are important for growth during SAS, we measured the shade avoidance response of seven salicylic acid pathway mutant strains: NahG-OX (Gaffney et al., 1993), enhanced disease susceptibility5 (eds5)/salicylic acid induction-deficient1 (sid1; Nawrath and Métraux, 1999; Nawrath et al., 2002), sid2/ics1 (Nawrath and Métraux, 1999; Wildermuth et al., 2001), npr1, npr3, npr4, and npr3/4 (Zhang et al., 2006).
Induced defense responses are known to be impaired in these salicylic acid-related mutants; by contrast, responses to shade in leaf were altered in a complex manner (Fig. 5). Three mutant lines (NahG-OX, npr1, and sid2) showed a constitutive sun phenotype with reduced shade avoidance elongation in two indices (petiole length and petiole length-blade length ratio). Among these lines, the salicylic acid-deficient mutant sid2 showed the most severe phenotype. Although NPR3 and NPR4 can act as salicylic acid receptors (Zhang et al., 2010; Fu et al., 2012; Ding et al., 2018), neither of them is required for responses to shade (Fig. 5, A and B). These findings suggest that a basal level of salicylic acid biosynthesis is required for proper shade avoidance and that shade influences downstream salicylic acid signaling via an NPR3/4-independent mechanism. There is precedent for an NPR3/4-independent model: NPR1 can bind to salicylic acid directly (Wu et al., 2012; Manohar et al., 2015; Ding et al., 2018), salicylic acid promotes NPR1 accumulation even in npr3/4 double mutants (Fu et al., 2012), and salicylic acid regulates pollen tube growth independently of NPR3/4 (Rong et al., 2016). Previous data have shown that while effector-triggered immunity (ETI) is compromised in npr3/4 double mutants (Fu et al., 2012), basal defenses are maintained or enhanced (Zhang et al., 2006), in contrast to npr1 mutants, where both ETI and basal defenses are disrupted (Cao et al., 1994). Our data are consistent with a model wherein NPR3 and NPR4 play relatively specific roles in ETI but NPR1 is required for a broader spectrum of salicylic acid signaling responses. Another noticeable effect of mutations in salicylic acid-related genes is short petiole elongation, which has been observed under both light conditions, consistent with a proposed role of salicylic acid as a general growth regulator (Rivas-San Vicente and Plasencia, 2011). An alternative possibility is that SID2 and NPR1 are moonlighting in the SAS pathway, in other words, that their role in SAS is independent of their role in salicylic acid signaling. This hypothesis seems unlikely, however, given that plants overexpressing NahG exhibit similar SAS phenotypes. Given that NahG is a bacterial enzyme, the most parsimonious explanation for the similar phenotypes of sid2 and NahG-OX plants is that, in both genotypes, the phenotypes are due to reduced levels of salicylic acid.
Figure 5.
Salicylic acid-related mutants have reduced SAS as well as general reduced growth. A, Petiole length. B, Ratio of petiole length. C, Hypocotyl length. D, Flowering time (days at bolting). Significant differences were evaluated using a linear mixed-effects model with genotype, treatment, and the genotype-by-treatment interaction as fixed effects. Asterisks over sun columns indicate significant differences from Col in sun (a linear mixed-effects model; P < 0.05). Asterisks for shade on Col indicate significant responses to shade, and asterisks for mutants indicate significantly different shade responses compared with Col (a linear mixed-effects model; P < 0.05). The third to sixth leaves were measured from 15 to 33 plants per genotype/condition from 13 experimental trials.
To test if salicylic acid signaling components are important for other aspects of SAS, we phenotyped salicylic acid-related mutants with responses in hypocotyl and flowering time. Neither salicylic acid-up-regulated genes nor genes in the salicylic acid-related salmon module were misexpressed in phyA/B/D/E mutant seedlings (Supplemental Figs. S1A and S6), whereas they are misexpressed in phyB mutant adults (Fig. 4; Supplemental Fig. S1B), suggesting that the interaction between shade avoidance and salicylic acid signaling may be stage or organ specific. Indeed, we found that salicylic acid pathway mutants showed hypocotyl shade avoidance responses similar to those of the wild type (Fig. 5C). Another SAS response is an acceleration of flowering time (Franklin et al., 2003; Kim et al., 2008; Wollenberg et al., 2008). None of the salicylic acid-related mutants showed altered flowering responses (Fig. 5D), indicating that salicylic acid does not mediate this response. Thus, the requirement of salicylic acid signaling components is specific to petiole shade avoidance and demonstrates how downstream signaling modules vary between organs and developmental stages.
Since two salmon module genes, SARD1 and CBP60g, activate salicylic acid biosynthesis genes, we asked whether the modulation of salicylic acid-related pathways by low R/FR was simply due to an alteration in the amount of salicylic acid by low R/FR treatment. Salicylic acid levels were measured in 18-d-old Col, phyB, and sid2 plants maintained in high R/FR or exposed to 4 d of low R/FR. A significant reduction in salicylic acid was observed in phyB (P = 0.0006), but the R/FR ratio did not alter salicylic acid levels in any genotype (P = 0.849; Supplemental Fig. S7). Thus, the modulation of salicylic acid-related pathways by low R/FR was not due to changes of salicylic acid levels. Since the transcriptome signatures in Col treated with low R/FR for 2 d and in phyB grown under high R/FR are similar, the significant reduction in salicylic acid in phyB is surprising. This reduction may be the result of longer term effects of low R/FR specific to the adult stage, since phytochrome mutants have been reported to have normal levels of salicylic acid in seedlings (Genoud et al., 2002).
It is interesting that eds5 mutants showed normal shade avoidance elongation even though their leaves are small and their petiole-leaf blade length ratio was smaller than that in Col. This means that the influence of salicylic acid on growth can be separated from its effect on shade avoidance responses. SID2 encodes a salicylic acid biosynthesis enzyme that is imported to and functions in plastids (Strawn et al., 2007; Garcion et al., 2008). eds5 plants have a mutation in a multidrug and toxin extrusion-like transporter that may be required for the export of salicylic acid from the chloroplast to the cytoplasm (Nawrath et al., 2002; Serrano et al., 2013; Yamasaki et al., 2013). If the proposed function of EDS5 is true, then export of salicylic acid from the chloroplast to the cytoplasm is important for basal leaf and petiole growth, but EDS5-mediated export is not required for a normal shade avoidance response. This observation could be explained by a plastid salicylic acid-related retrograde signal required for shade avoidance responses, whereas exported cytoplasmic salicylic acid from chloroplast would be required only for basal growth both under low and high R/FR. It should be noted that there is a known link between light signaling and the defense system through the chloroplast in FHY3- and FAR1-mediated phyA signaling (Wang et al., 2016).
CONCLUSION
Analysis of our time-course transcriptome data revealed that three hormone pathways are differentially regulated and involved in adult shade avoidance responses. First, there is transient activation of the auxin pathway, followed by transient repression of the jasmonic acid pathway, and finally late repression of the salicylic acid pathway. While the effects of shade-mediated repression on jasmonic and salicylic acid signaling have been studied with respect to decreased defense, there is little information on whether this down-regulation is important for the proper growth responses during shade avoidance. The pervasive misregulation of the salmon salicylic acid-related module genes in shade avoidance mutants shows that there is a much stronger connection between these pathways than was suspected previously. Our subsequent analysis of salicylic acid pathway mutants demonstrates that genes in the salicylic acid pathway are necessary for shade-regulated growth in adult plants. Interestingly, salicylic acid pathway mutants do not show defects in seedling SAS, and salicylic acid-responsive genes are only affected by phytochrome mutants in adults, demonstrating important differences between seedling and adult shade avoidance. Our findings provide insight into the stage- and organ-specific downstream signaling components and a novel framework of growth-defense prioritization in adult plants (Fig. 6). We hypothesize that low R/FR does not just inhibit NPR1 signaling but instead alters its function, perhaps by protein modification. It has been shown previously that low R/FR causes NPR1 translocation into the nucleus but inhibits its phosphorylation in the presence of salicylic acid (de Wit et al., 2013). Under this model, shade-modified, nucleus-localized NPR1 is no longer able to promote defense responses (i.e. is unable to up-regulate canonical salicylic acid-up-regulated genes) but instead is diverted to promoting elongation. This model can explain both our transcriptome data and salicylic acid-related mutant phenotypes. Although this model is not able to explain why other SAS components (e.g. YUCs and PIFs) are required for proper salmon module gene expression, within this framework, salicylic acid-related component(s) in the salmon module are strong candidates for linking growth-defense prioritization in the SAS.
Figure 6.
Schematic representation of the plant growth-defense signaling network and the effects of low R/FR. Proposed new connections based on our systems analysis are shown in thick dotted lines. NPR1* is a proposed modified NPR1, such as by phosphorylation.
MATERIALS AND METHODS
Plant Growth Condition
The plant growth condition for the analysis of shade avoidance was described previously in detail (Nozue et al., 2015). Briefly, Arabidopsis (Arabidopsis thaliana) plants were grown under a 16-h-light/8-h-dark photoperiod under simulated sun condition (R/FR = 1.5), and 2-week-old plants were used for the following experiments. Most plants used in this study were described by Nozue et al. (2015), except NahG-OX (Gaffney et al., 1993), npr1-1 (Cao et al., 1997), eds5-3/sid1 (Nawrath and Métraux, 1999; Nawrath et al., 2002), and sid2-1 (Nawrath and Métraux, 1999; Wildermuth et al., 2001), all provided by Clare Casteel (University of California, Davis), and salicylic acid receptor mutants npr3-1, npr4-3, npr3-1, and npr4-3 (Zhang et al., 2006), all provided by Xinnian Dong (Duke University).
For YUC::LUC2 assays, 13-d-old plants containing the LUC2 gene (pGL4.10; Promega) driven by the promoter of YUC5, YUC8, or YUC9 (see below) were sprayed with luciferin and transferred to simulated sun condition in a light-emitting diode chamber (R/FR = 1.3; Nozue et al., 2015) and then subjected to a luciferase assay for 2 d (see below).
For RNA sequencing (RNAseq) library construction, 2-week-old Col and 12 mutants (aos, co, hy5, jar1, kat1-2, mida9, PAR1_RNAi09, phyB, pif3, pif4pif5, spt, and yuc2yuc5yuc8yuc9) grown under simulated sun condition (R/FR = 1.5) were split into simulated sun condition and simulated shade condition (R/FR = 0.5; Nozue et al., 2015), and young aboveground tissues (aboveground tissues excluding hypocotyl, cotyledons, and first and second leaves) were collected at 1, 4, 25, or 49 h after the onset of low R/FR treatment at 7 h after dawn. Two plants were pooled for each sample, and three biological replicate samples for each genotype/treatment combination were collected. Samples were frozen quickly in liquid nitrogen and kept at −80°C.
For shade avoidance phenotyping, 2-week-old plants of Col, NahG-OX, npr1-1, eds5-3, sid2, npr3, npr4, and npr3npr4 were grown for a further 12 d in each light condition for leaf phenotyping as described (Nozue et al., 2015).
For salicylic acid measurement, 2-week-old plants of Col, phyB, and sid2 were grown for a further 4 d in each light condition, and young aboveground tissues described for RNA library construction were collected.
YUC Promoter::LUC2 Assay
YUC5::LUC2, YUC8::LUC2, and YUC9::LUC2 constructs were made by amplifying the promoters (about 2 kb before the start site) of these genes (YUC5 [AT5G43890], YUC8 [AT4G28720], and YUC9 [AT1G04180]) by PCR (see below for primers) and cloning into the pENTR-D/TOPO entry vector (Invitrogen). The resulting entry vectors were combined with the pEarlyGate301::Luc2 destination vector (Rawat et al., 2009), which confers kanamycin resistance in bacteria and Basta resistance in plants, using LR Clonase mix (Invitrogen). Transgenic plants were identified by their Basta resistance or by luciferase screening if they were already Basta resistant.
Homozygous YUC::LUC2 transgenic plants that were sprayed with luciferin were split into high and low R/FR at ZT7. From ZT6, luciferase activity was monitored for 2 d using an iKon M-934 CCD camera (Andor) controlled by LabView software (National Instruments) with Snap-Lite LED illumination (Quantum Devices). Luminescence at each time point was measured by ImageJ with a multi-measure plugin (OptiNav; see detailed protocols in http://malooflab.phytonetworks.org/wiki/Analyzing_luminiscence/), normalized by dividing luminescence one time point before low/high R/FR treatment (as a reference time point), and then increasing relative to the reference time point data calculated. Data analysis and drawing graphs were performed in R (R Core Team, 2016).
RNAseq Library Preparation
RNAseq libraries were prepared as described (Kumar et al., 2012) with 96 barcodes. Single-end 50-bp sequencing was performed by the University of California, Berkeley/QB3 Vincent J. Coates Genomic Sequencing Laboratory. Reads were sorted by barcode and filtered to remove adaptor contamination as described (Devisetty et al., 2014). Reads were mapped using BWA (Li and Durbin, 2009) to the Arabidopsis TAIR10 reference cDNA. Library genotypes were validated with multiple points: visual inspection of causal single-nucleotide polymorphism of mutants, truncated transcripts due to T-DNA insertion in a causal gene, and lower expression levels of causal genes. Libraries that did not pass the genotyping were eliminated from our analysis. Statistics of each library were summarized in R scripts.
RNAseq Data Analysis
Differentially expressed gene analysis was done using edgeR with glm function at FDR < 0.05 for treatment-affected genes and FDR < 0.001 for mutant-misexpressed genes (Robinson and Smyth, 2007; Robinson et al., 2010; Robinson and Oshlack, 2010; McCarthy et al., 2012). Two separate glm models were used: (1) time + trt + time:trt + batch for differentially expressed genes between high R/FR and low R/FR in a given genotype, and (2) genotype + trt + genotype:trt + batch for differentially expressed genes between Col and a given genotype under high or low R/FR condition, where time is duration of light treatment, trt is light treatment, and batch is batch effects of experimental sets. We were not able to fit a time*trt*genotype interaction model (time + trt + genotype + time:trt + time:genotype + trt:genotype + time:trt:genotype) due to computational constraints. For coexpression analysis, read counts were transformed by variance stabilization transformation using DEseq2 (Love et al., 2014). Coexpression analysis of low-R/FR-responsive genes was done by the Barnes-Hut t-SNE (van der Maaten and Hinton, 2008; van der Maaten, 2014) method as described (Ranjan et al., 2016) with modification. Instead of scaled data as used in the original method, we used unscaled fold change of responsiveness to low R/FR. Subsequently, the DBscan algorithm (implemented in the fpc R package) was used for clustering with the following parameters: minpts between 13 and 18 were used to capture smaller modules on the periphery, and epsilons between 3.1 and 3.35 were used to avoid the overlapping of internal and closely spaced modules. For representative figures, a minpts of 13 and an epsilon of 3.3 were used. Time-course changes of shade responsiveness in each cluster were visualized in Supplemental Figure S2 for representative parameters and in Supplemental Data Set S1 for all parameters. Coexpression analysis with absolute expression levels across all combinations of light condition and time points was done by the WGCNA package in R/bioconductor (Langfelder and Horvath, 2008). Overrepresentation analysis of differentially expressed genes and WGCNA coexpression modules was done by the overlapTable function in the WGCNA package. For over-representation analysis (ORA) of hormone-responsive genes, custom categories were defined using the preexisting data sets described in Supplemental Data Set S3.
GO category enrichment analysis of t-SNE clusters and WGCNA modules was performed using the GOseq R package with 10,000 permutations (Young et al., 2010) and adjusted by Benjamini and Hochberg correction (Benjamini and Hochberg, 1995). Enrichment of misexpressed genes in mutants was visualized as heat maps where the P value was indicated by color, and overlapping gene numbers between WGCNA modules and misexpressed genes were calculated using a modified version of the overlapTable function in the WGCNA package, where Fisher’s exact test was replaced with the enrichment test from GOseq, using sampling with 2,000 permutations. Differentially expressed genes between Col and phyA/B/D/E mutants grown under continuous red light were analyzed by Rank Product from publicly available data (Gene Expression Omnibus database, accession no. GSE31587; Hu et al., 2013). GO category enrichment analysis of misexpressed genes in light-grown phyA/B/DE mutant seedlings was done using amiGO version 2.2.0 (http://amigo.geneontology.org/amigo; Carbon et al., 2009).
Time-course shade-responsive genes in custom categories were visualized (Fig. 2; Supplemental Data Set S7) by the following procedures. For each custom category, all genes enriched in t-SNE clusters (P < 0.05 in Supplemental Fig. S3 [representative] and Supplemental Data Set S5 [all parameters]) were selected (Supplemental Data Set S6 for all parameters), and their mean shade responsiveness in each genotype was visualized in a heat map (log2 fold change; magenta and green represent induced or repressed, respectively; Fig. 2 [representative] and Supplemental Data Set S7 [all parameters]). To determine if a genotype had a defect in the response of a particular category, a mixed-effects model was used with gene as a random effect and genotype as a fixed effect.
Shade Avoidance Phenotype Measurement and Analysis
The shade avoidance phenotype in leaves was measured as described (Maloof et al., 2013). Data from leaf 3 to leaf 6 were fitted using a linear mixed-effects model as described (Nozue et al., 2015). Briefly, genotype, treatment, and the genotype-by-treatment interaction were used as fixed effects and experimental set was used as a random effect. Mixed-effects models were fit using the lme4 package (Bates et al., 2015) in R (R Core Team, 2016).
Salicylic Acid Measurement and Analysis
Salicylic acid in 18-d-old plants was measured with a liquid chromatography-mass spectrometry system. Leaves were weighed, flash frozen, and immediately homogenized. Following homogenization, frozen tissue was extracted and analyzed using a Thermo Electron LTQ-Orbitrap XL Hybrid mass spectrometer as described previously (Casteel et al., 2015). For analysis, 5 µL of each extract was separated on a Zorbax Extend-C18 HPLC column (Agilent; 3.5 µm, 150 × 3 mm) using 0.1% (v/v) formic acid in water and 0.1% (v/v) formic acid in acetonitrile. Data were fitted using a linear mixed-effects model:
where plant is a mutant, treatment is low or high R/FR condition, plant:treatment is the interaction of plant and treatment, and ε is the error. The model was applied to calculate coefficient (sun value). Mutants were considered to have a defect in SAS when the plant:treatment term was significant (P < 0.05), indicating that the genotype of the plant (mutant versus wild type) affected the response to shade.
Accession Numbers
The reads were submitted to the National Center for Biotechnology Information Sequence Read Archive with accession number PRJNA343549.
All R scripts for this article and raw data and supplemental data are available at https://github.com/MaloofLab/Nozue2018_SAStranscriptome_scripts.
Mutants used for RNAseq are given in Table 1. Additionally, the following genes and data sets are of significance to the work described in this article: ACD6, AT4G14400; CBP60g, AT5G26920; EDS16/ICS1, AT1G74710; MYC2, AT1G32640; MYC3, AT5G46760; MYC4, AT4G17880; NPR1, AT1G64280; NPR3, AT5G45110; NPR4, AT4G19660; PCC1, AT3G22231; PHYA, AT1G09570; PHYC, AT5G35840; PHYE, AT4G18130; EDS5/SID1, AT4G39030; EDS16/SID2, AT1G74710; WRKY25, AT2G30250; WRKY33, AT2G38470; WRKY46, AT2G46400; WRKY54, AT2G40750; WRKY70, AT3G56400; phyA/B/D/E transcriptome, National Center for Biotechnology Information Gene Expression Omnibus accession number GSE31587.
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. Characterization of phytochrome-regulated genes in three different situations.
Supplemental Figure S2. Coexpression clusters of shade-responsive genes (representative).
Supplemental Figure S3. Overrepresentation analysis of t-SNE gene clusters.
Supplemental Figure S4. YUC9 promoter activity was induced mainly in leaves.
Supplemental Figure S5. Response of target genes at 1 h.
Supplemental Figure S6. Overrepresentation analysis of shade-induced genes and phyA/B/D/E misregulated genes with WGCNA modules.
Supplemental Figure S7. Salicylic acid levels were not influenced upon 4 d of treatment with low R/FR.
Supplemental Data Set S1. Complete set of t-SNE clusters across different parameter values (combined PDF files). Figure 2 is a representative example.
Supplemental Data Set S2. GO enrichment analysis of coexpressed t-SNE clusters of shade-responsive genes.
Supplemental Data Set S3. Custom categories used in this study.
Supplemental Data Set S4. Shade-responsive genes in custom categories used in Figure 2.
Supplemental Data Set S5. Overrepresentation analysis of t-SNE gene clusters with all given parameters.
Supplemental Data Set S6. Complete list of shade-responsive genes at least in one genotype and t-SNE cluster in each mutant and custom enrichment analysis of coexpressed t-SNE clusters of shade-responsive genes.
Supplemental Data Set S7. Time-course visualization of shade-responsive genes in custom categories with all given parameters.
Supplemental Data Set S8. GO enrichment analysis of WGCNA gene coexpression modules.
Supplemental Data Set S9. Custom category enrichment analysis of WGCNA gene coexpression modules.
Supplemental Data Set S10. Complete gene list of genes misexpressed in at least one mutant and found in WGCNA coexpression modules (visualized in Fig. 4).
Acknowledgments
We thank Maxwell R. Mumbach (University of California, Davis) and Christine Palmer (University of California, Davis) for collecting tissue. We thank Yasunori Ichihashi (University of California, Davis) for helping with coexpression analysis. We thank Cody Markelz, Ruijuan Li, and Leonela Carriedo for critical reading of the article.
Footnotes
This work was supported by the National Science Foundation Integrative Organismal Systems (IOS-0923752), the U.S. Department of Agriculture NIFA project (CA-D-PLB-7226-H), and funds from the University of California, Davis.
Articles can be viewed without a subscription.
References
- Andreasson E, Jenkins T, Brodersen P, Thorgrimsen S, Petersen NH, Zhu S, Qiu JL, Micheelsen P, Rocher A, Petersen M, et al. (2005) The MAP kinase substrate MKS1 is a regulator of plant defense responses. EMBO J 24: 2579–2589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balasubramanian S, Sureshkumar S, Lempe J, Weigel D (2006) Potent induction of Arabidopsis thaliana flowering by elevated growth temperature. PLoS Genet 2: e106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballaré CL. (2014) Light regulation of plant defense. Annu Rev Plant Biol 65: 335–363 [DOI] [PubMed] [Google Scholar]
- Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67: 1–48 [Google Scholar]
- Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: A practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57: 289–300 [Google Scholar]
- Birkenbihl RP, Diezel C, Somssich IE (2012) Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol 159: 266–285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boccalandro HE, Ploschuk EL, Yanovsky MJ, Sánchez RA, Gatz C, Casal JJ (2003) Increased phytochrome B alleviates density effects on tuber yield of field potato crops. Plant Physiol 133: 1539–1546 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao H, Bowling SA, Gordon AS, Dong X (1994) Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6: 1583–1592 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao H, Glazebrook J, Clarke JD, Volko S, Dong X (1997) The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 88: 57–63 [DOI] [PubMed] [Google Scholar]
- Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S (2009) AmiGO: Online access to ontology and annotation data. Bioinformatics 25: 288–289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casal JJ. (2013) Photoreceptor signaling networks in plant responses to shade. Annu Rev Plant Biol 64: 403–427 [DOI] [PubMed] [Google Scholar]
- Casteel CL, De Alwis M, Bak A, Dong H, Whitham SA, Jander G (2015) Disruption of ethylene responses by Turnip mosaic virus mediates suppression of plant defense against the green peach aphid vector. Plant Physiol 169: 209–218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cerrudo I, Keller MM, Cargnel MD, Demkura PV, de Wit M, Patitucci MS, Pierik R, Pieterse CMJ, Ballaré CL (2012) Low red/far-red ratios reduce Arabidopsis resistance to Botrytis cinerea and jasmonate responses via a COI1-JAZ10-dependent, salicylic acid-independent mechanism. Plant Physiol 158: 2042–2052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J, Nolan TM, Ye H, Zhang M, Tong H, Xin P, Chu J, Chu C, Li Z, Yin Y (2017) Arabidopsis WRKY46, WRKY54, and WRKY70 transcription factors are involved in brassinosteroid-regulated plant growth and drought responses. Plant Cell 29: 1425–1439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chincinska IA, Liesche J, Krügel U, Michalska J, Geigenberger P, Grimm B, Kühn C (2008) Sucrose transporter StSUT4 from potato affects flowering, tuberization, and shade avoidance response. Plant Physiol 146: 515–528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciolfi A, Sessa G, Sassi M, Possenti M, Salvucci S, Carabelli M, Morelli G, Ruberti I (2013) Dynamics of the shade-avoidance response in Arabidopsis. Plant Physiol 163: 331–353 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cole B, Kay SA, Chory J (2011) Automated analysis of hypocotyl growth dynamics during shade avoidance in Arabidopsis. Plant J 65: 991–1000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devisetty UK, Covington MF, Tat AV, Lekkala S, Maloof JN (2014) Polymorphism identification and improved genome annotation of Brassica rapa through Deep RNA sequencing. G3 (Bethesda) 4: 2065–2078 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Wit M, Spoel SH, Sanchez-Perez GF, Gommers CMM, Pieterse CMJ, Voesenek LACJ, Pierik R (2013) Perception of low red:far-red ratio compromises both salicylic acid- and jasmonic acid-dependent pathogen defences in Arabidopsis. Plant J 75: 90–103 [DOI] [PubMed] [Google Scholar]
- de Wit M, Ljung K, Fankhauser C (2015) Contrasting growth responses in lamina and petiole during neighbor detection depend on differential auxin responsiveness rather than different auxin levels. New Phytol 208: 198–209 [DOI] [PubMed] [Google Scholar]
- Ding Y, Sun T, Ao K, Peng Y, Zhang Y, Li X, Zhang Y (2018) Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 173: 1454–1467.e15 [DOI] [PubMed] [Google Scholar]
- Franklin KA, Praekelt U, Stoddart WM, Billingham OE, Halliday KJ, Whitelam GC (2003) Phytochromes B, D, and E act redundantly to control multiple physiological responses in Arabidopsis. Plant Physiol 131: 1340–1346 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu ZQ, Yan S, Saleh A, Wang W, Ruble J, Oka N, Mohan R, Spoel SH, Tada Y, Zheng N, et al. (2012) NPR3 and NPR4 are receptors for the immune signal salicylic acid in plants. Nature 486: 228–232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaffney T, Friedrich L, Vernooij B, Negrotto D, Nye G, Uknes S, Ward E, Kessmann H, Ryals J (1993) Requirement of salicylic acid for the induction of systemic acquired resistance. Science 261: 754–756 [DOI] [PubMed] [Google Scholar]
- Gao QM, Venugopal S, Navarre D, Kachroo A (2011) Low oleic acid-derived repression of jasmonic acid-inducible defense responses requires the WRKY50 and WRKY51 proteins. Plant Physiol 155: 464–476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcion C, Lohmann A, Lamodière E, Catinot J, Buchala A, Doermann P, Métraux JP (2008) Characterization and biological function of the ISOCHORISMATE SYNTHASE2 gene of Arabidopsis. Plant Physiol 147: 1279–1287 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genoud T, Buchala AJ, Chua NH, Métraux JP (2002) Phytochrome signalling modulates the SA-perceptive pathway in Arabidopsis. Plant J 31: 87–95 [DOI] [PubMed] [Google Scholar]
- Guranowski A, Miersch O, Staswick PE, Suza W, Wasternack C (2007) Substrate specificity and products of side-reactions catalyzed by jasmonate:amino acid synthetase (JAR1). FEBS Lett 581: 815–820 [DOI] [PubMed] [Google Scholar]
- Hersch M, Lorrain S, de Wit M, Trevisan M, Ljung K, Bergmann S, Fankhauser C (2014) Light intensity modulates the regulatory network of the shade avoidance response in Arabidopsis. Proc Natl Acad Sci USA 111: 6515–6520 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holm M, Ma LG, Qu LJ, Deng XW (2002) Two interacting bZIP proteins are direct targets of COP1-mediated control of light-dependent gene expression in Arabidopsis. Genes Dev 16: 1247–1259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu W, Franklin KA, Sharrock RA, Jones MA, Harmer SL, Lagarias JC (2013) Unanticipated regulatory roles for Arabidopsis phytochromes revealed by null mutant analysis. Proc Natl Acad Sci USA 110: 1542–1547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ichihashi Y, Horiguchi G, Gleissberg S, Tsukaya H (2010) The bHLH transcription factor SPATULA controls final leaf size in Arabidopsis thaliana. Plant Cell Physiol 51: 252–261 [DOI] [PubMed] [Google Scholar]
- Izaguirre MM, Mazza CA, Biondini M, Baldwin IT, Ballaré CL (2006) Remote sensing of future competitors: Impacts on plant defenses. Proc Natl Acad Sci USA 103: 7170–7174 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazan K, Manners JM (2013) MYC2: The master in action. Mol Plant 6: 686–703 [DOI] [PubMed] [Google Scholar]
- Kim J, Yi H, Choi G, Shin B, Song PS, Choi G (2003) Functional characterization of phytochrome interacting factor 3 in phytochrome-mediated light signal transduction. Plant Cell 15: 2399–2407 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim SY, Yu X, Michaels SD (2008) Regulation of CONSTANS and FLOWERING LOCUS T expression in response to changing light quality. Plant Physiol 148: 269–279 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozuka T, Kobayashi J, Horiguchi G, Demura T, Sakakibara H, Tsukaya H, Nagatani A (2010) Involvement of auxin and brassinosteroid in the regulation of petiole elongation under the shade. Plant Physiol 153: 1608–1618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar R, Ichihashi Y, Kimura S, Chitwood DH, Headland LR, Peng J, Maloof JN, Sinha NR (2012) A high-throughput method for Illumina RNA-Seq library preparation. Front Plant Sci 3: 202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langfelder P, Horvath S (2008) WGCNA: An R package for weighted correlation network analysis. BMC Bioinformatics 9: 559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laudert D, Weiler EW (1998) Allene oxide synthase: A major control point in Arabidopsis thaliana octadecanoid signalling. Plant J 15: 675–684 [DOI] [PubMed] [Google Scholar]
- Leone M, Keller MM, Cerrudo I, Ballaré CL (2014) To grow or defend? Low red:far-red ratios reduce jasmonate sensitivity in Arabidopsis seedlings by promoting DELLA degradation and increasing JAZ10 stability. New Phytol 204: 355–367 [DOI] [PubMed] [Google Scholar]
- Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25: 1754–1760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L, Ljung K, Breton G, Schmitz RJ, Pruneda-Paz J, Cowing-Zitron C, Cole BJ, Ivans LJ, Pedmale UV, Jung HS, et al. (2012) Linking photoreceptor excitation to changes in plant architecture. Genes Dev 26: 785–790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S, Fu Q, Chen L, Huang W, Yu D (2011) Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta 233: 1237–1252 [DOI] [PubMed] [Google Scholar]
- Lorrain S, Allen T, Duek PD, Whitelam GC, Fankhauser C (2008) Phytochrome-mediated inhibition of shade avoidance involves degradation of growth-promoting bHLH transcription factors. Plant J 53: 312–323 [DOI] [PubMed] [Google Scholar]
- Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15: 550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu H, Rate DN, Song JT, Greenberg JT (2003) ACD6, a novel ankyrin protein, is a regulator and an effector of salicylic acid signaling in the Arabidopsis defense response. Plant Cell 15: 2408–2420 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maloof JN, Nozue K, Mumbach MR, Palmer CM (2013) LeafJ: An ImageJ plugin for semi-automated leaf shape measurement. J Vis Exp 71: e50028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manohar M, Tian M, Moreau M, Park SW, Choi HW, Fei Z, Friso G, Asif M, Manosalva P, von Dahl CC, et al. (2015) Identification of multiple salicylic acid-binding proteins using two high throughput screens. Front Plant Sci 5: 777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy DJ, Chen Y, Smyth GK (2012) Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res 40: 4288–4297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mir R, León J (2014) Pathogen and circadian controlled 1 (PCC1) protein is anchored to the plasma membrane and interacts with subunit 5 of COP9 signalosome in Arabidopsis. PLoS ONE 9: e87216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller-Moulé P, Nozue K, Pytlak ML, Palmer CM, Covington MF, Wallace AD, Harmer SL, Maloof JN (2016) YUCCA auxin biosynthetic genes are required for Arabidopsis shade avoidance. PeerJ 4: e2574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nawrath C, Métraux JP (1999) Salicylic acid induction-deficient mutants of Arabidopsis express PR-2 and PR-5 and accumulate high levels of camalexin after pathogen inoculation. Plant Cell 11: 1393–1404 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nawrath C, Heck S, Parinthawong N, Métraux JP (2002) EDS5, an essential component of salicylic acid-dependent signaling for disease resistance in Arabidopsis, is a member of the MATE transporter family. Plant Cell 14: 275–286 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nito K, Kajiyama T, Unten-Kobayashi J, Fujii A, Mochizuki N, Kambara H, Nagatani A (2015) Spatial regulation of the gene expression response to shade in Arabidopsis seedlings. Plant Cell Physiol 56: 1306–1319 [DOI] [PubMed] [Google Scholar]
- Nozue K, Tat AV, Kumar Devisetty U, Robinson M, Mumbach MR, Ichihashi Y, Lekkala S, Maloof JN (2015) Shade avoidance components and pathways in adult plants revealed by phenotypic profiling. PLoS Genet 11: e1004953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park JH, Halitschke R, Kim HB, Baldwin IT, Feldmann KA, Feyereisen R (2002) A knock-out mutation in allene oxide synthase results in male sterility and defective wound signal transduction in Arabidopsis due to a block in jasmonic acid biosynthesis. Plant J 31: 1–12 [DOI] [PubMed] [Google Scholar]
- Pedmale UV, Huang SC, Zander M, Cole BJ, Hetzel J, Ljung K, Reis PAB, Sridevi P, Nito K, Nery JR, et al. (2016) Cryptochromes interact directly with PIFs to control plant growth in limiting blue light. Cell 164: 233–245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phukan UJ, Jeena GS, Shukla RK (2016) WRKY transcription factors: Molecular regulation and stress responses in plants. Front Plant Sci 7: 760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Procko C, Crenshaw CM, Ljung K, Noel JP, Chory J (2014) Cotyledon-generated auxin is required for shade-induced hypocotyl growth in Brassica rapa. Plant Physiol 165: 1285–1301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ranjan A, Budke JM, Rowland SD, Chitwood DH, Kumar R, Carriedo L, Ichihashi Y, Zumstein K, Maloof JN, Sinha NR (2016) eQTL regulating transcript levels associated with diverse biological processes in tomato. Plant Physiol 172: 328–340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rawat R, Schwartz J, Jones MA, Sairanen I, Cheng Y, Andersson CR, Zhao Y, Ljung K, Harmer SL (2009) REVEILLE1, a Myb-like transcription factor, integrates the circadian clock and auxin pathways. Proc Natl Acad Sci USA 106: 16883–16888 [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Core Team (2016) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna [Google Scholar]
- Reddy SK, Holalu SV, Casal JJ, Finlayson SA (2013) Abscisic acid regulates axillary bud outgrowth responses to the ratio of red to far-red light. Plant Physiol 163: 1047–1058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reed JW, Nagpal P, Poole DS, Furuya M, Chory J (1993) Mutations in the gene for the red/far-red light receptor phytochrome B alter cell elongation and physiological responses throughout Arabidopsis development. Plant Cell 5: 147–157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivas-San Vicente M, Plasencia J (2011) Salicylic acid beyond defence: its role in plant growth and development. J Exp Bot 62: 3321–3338 [DOI] [PubMed] [Google Scholar]
- Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11: R25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, Smyth GK (2007) Moderated statistical tests for assessing differences in tag abundance. Bioinformatics 23: 2881–2887 [DOI] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26: 139–140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roig-Villanova I, Bou-Torrent J, Galstyan A, Carretero-Paulet L, Portolés S, Rodríguez-Concepción M, Martínez-García JF (2007) Interaction of shade avoidance and auxin responses: A role for two novel atypical bHLH proteins. EMBO J 26: 4756–4767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rong D, Luo N, Mollet JC, Liu X, Yang Z (2016) Salicylic acid regulates pollen tip growth through an NPR3/NPR4-independent pathway. Mol Plant 9: 1478–1491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauerbrunn N, Schlaich NL (2004) PCC1: A merging point for pathogen defence and circadian signalling in Arabidopsis. Planta 218: 552–561 [DOI] [PubMed] [Google Scholar]
- Segarra S, Mir R, Martínez C, León J (2010) Genome-wide analyses of the transcriptomes of salicylic acid-deficient versus wild-type plants uncover Pathogen and Circadian Controlled 1 (PCC1) as a regulator of flowering time in Arabidopsis. Plant Cell Environ 33: 11–22 [DOI] [PubMed] [Google Scholar]
- Serrano M, Wang B, Aryal B, Garcion C, Abou-Mansour E, Heck S, Geisler M, Mauch F, Nawrath C, Métraux JP (2013) Export of salicylic acid from the chloroplast requires the multidrug and toxin extrusion-like transporter EDS5. Plant Physiol 162: 1815–1821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sessa G, Carabelli M, Sassi M, Ciolfi A, Possenti M, Mittempergher F, Becker J, Morelli G, Ruberti I (2005) A dynamic balance between gene activation and repression regulates the shade avoidance response in Arabidopsis. Genes Dev 19: 2811–2815 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staswick PE, Tiryaki I (2004) The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 16: 2117–2127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strawn MA, Marr SK, Inoue K, Inada N, Zubieta C, Wildermuth MC (2007) Arabidopsis isochorismate synthase functional in pathogen-induced salicylate biosynthesis exhibits properties consistent with a role in diverse stress responses. J Biol Chem 282: 5919–5933 [DOI] [PubMed] [Google Scholar]
- Tao Y, Ferrer JLL, Ljung K, Pojer F, Hong F, Long JA, Li L, Moreno JE, Bowman ME, Ivans LJ, et al. (2008) Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 133: 164–176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulker B, Shahid Mukhtar M, Somssich IE (2007) The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta 226: 125–137 [DOI] [PubMed] [Google Scholar]
- van der Maaten L. (2014) Accelerating t-SNE using tree-based algorithms. J Mach Learn Res 15: 3221–3245 [Google Scholar]
- van der Maaten L, Hinton G (2008) Visualizing data using t-SNE. J Mach Learn Res 9: 2579–2605 [Google Scholar]
- Wang L, Tsuda K, Truman W, Sato M, Nguyen V, Katagiri F, Glazebrook J (2011) CBP60g and SARD1 play partially redundant critical roles in salicylic acid signaling. Plant J 67: 1029–1041 [DOI] [PubMed] [Google Scholar]
- Wang W, Tang W, Ma T, Niu D, Jin JB, Wang H, Lin R (2016) A pair of light signaling factors FHY3 and FAR1 regulates plant immunity by modulating chlorophyll biosynthesis. J Integr Plant Biol 58: 91–103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Gao J, Zhu Z, Dong X, Wang X, Ren G, Zhou X, Kuai B (2015) TCP transcription factors are critical for the coordinated regulation of isochorismate synthase 1 expression in Arabidopsis thaliana. Plant J 82: 151–162 [DOI] [PubMed] [Google Scholar]
- Westfall CS, Zubieta C, Herrmann J, Kapp U, Nanao MH, Jez JM (2012) Structural basis for prereceptor modulation of plant hormones by GH3 proteins. Science 336: 1708–1711 [DOI] [PubMed] [Google Scholar]
- Wildermuth MC, Dewdney J, Wu G, Ausubel FM (2001) Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414: 562–565 [DOI] [PubMed] [Google Scholar]
- Wollenberg AC, Strasser B, Cerdán PD, Amasino RM (2008) Acceleration of flowering during shade avoidance in Arabidopsis alters the balance between FLOWERING LOCUS C-mediated repression and photoperiodic induction of flowering. Plant Physiol 148: 1681–1694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y, Zhang D, Chu JY, Boyle P, Wang Y, Brindle ID, De Luca V, Després C (2012) The Arabidopsis NPR1 protein is a receptor for the plant defense hormone salicylic acid. Cell Rep. 1: 639–647 [DOI] [PubMed] [Google Scholar]
- Xie XZ, Xue YJ, Zhou JJ, Zhang B, Chang H, Takano M (2011) Phytochromes regulate SA and JA signaling pathways in rice and are required for developmentally controlled resistance to Magnaporthe grisea. Mol Plant 4: 688–696 [DOI] [PubMed] [Google Scholar]
- Yamasaki K, Motomura Y, Yagi Y, Nomura H, Kikuchi S, Nakai M, Shiina T (2013) Chloroplast envelope localization of EDS5, an essential factor for salicylic acid biosynthesis in Arabidopsis thaliana. Plant Signal Behav 8: e23603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene Ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol 11: R14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Cheng YT, Qu N, Zhao Q, Bi D, Li X (2006) Negative regulation of defense responses in Arabidopsis by two NPR1 paralogs. Plant J 48: 647–656 [DOI] [PubMed] [Google Scholar]
- Zhang Y, Xu S, Ding P, Wang D, Cheng YT, He J, Gao M, Xu F, Li Y, Zhu Z, et al. (2010) Control of salicylic acid synthesis and systemic acquired resistance by two members of a plant-specific family of transcription factors. Proc Natl Acad Sci USA 107: 18220–18225 [DOI] [PMC free article] [PubMed] [Google Scholar]