Abstract
Mangrove ecosystems are characteristic of the high salinity, limited nutrients and S-richness. Marichromatium gracile YL28 (YL28) isolated from mangrove tolerates the high concentrations of nitrite and sulfur compounds and efficiently eliminates them. However, the molecular mechanisms of nitrite and sulfur compounds utilization and the habitat adaptation remain unclear in YL28. We sequenced YL28 genome and further performed the comparative genome analysis in 36 purple bacteria including purple sulfur bacteria (PSB) and purple non-sulfur bacteria (PNSB). YL28 has 6 nitrogen cycle pathways (up to 40 genes), and possibly removes nitrite by denitrification, complete assimilation nitrate reduction and fermentative nitrate reduction (DNRA). Comparative genome analysis showed that more nitrogen utilization genes were detected in PNSB than those in PSB. The partial denitrification pathway and complete assimilation nitrate reduction were reported in PSB and DNRA was reported in purple bacteria for the first time. The three sulfur metabolism genes such as oxidation of sulfide, reversed dissimilatory sulfite reduction and sox system allowed to eliminate toxic sulfur compounds in the mangrove ecosystem. Several unique stress response genes facilitate to the tolerance of the high salinity environment. The CRISPR systems and the transposon components in genomic islands (GIs) likely contribute to the genome plasticity in purple bacteria.
Introduction
Purple bacteria are anoxygenic and phototrophic bacteria, including purple sulfur bacteria (PSB) and purple non-sulfur bacteria (PNSB). They are ubiquitously distributed in different natural environments with versatile metabolism potentials. Purple bacteria also serve as models for clarifying the biogeochemistry of C, N, S and Fe in the earth evolution1. YL28, a member of PSB, was isolated from mangrove special ecosystem2. It synthesized abundant rhodopin (not spirilloxanthin) carotenoid component under anaerobic in the light condition. It was also capable of using reduced sulfur compounds, nitrogen compounds or molecular hydrogen as electron donors2. Moreover, YL28 utilized ammonium, nitrite or nitrate as the sole nitrogen source for phototrophic growth. It is interesting that it was found to completely remove nitrite (up to 200 mg/L)3,4 and perform simultaneous heterotrophic nitrification and denitrification under anaerobic conditions5.
Nitrogen is one of the most abundant elements on earth. It comprises the majority of earth’s atmosphere and functions as one of the primary nutrients6. Nitrogen cycle is the most complex biogeochemical one in the biosphere7,8. However, the nitrogen cycle has been drastically disrupted because of overexploitation and modern agricultural activities9. Nearly half of the nitrogen reaches the coastal ocean via river input and/or atmospheric deposition10. This leads to extensive eutrophication of waters and coastal zones and increases inventories of the potent greenhouse gas (such as nitrous oxide). The elevated concentrations of NH3-N and NO2-N (the major pollutants) are problematic in aquatic ecosystems11.
Up to 10 nitrogen cycle pathways were reported9. For examples, purple bacteria have three nitrogen reduction cycles (reduced nitrogen compounds as electron acceptor, such as nitrogen fixation, nitrate assimilation, denitrification) and one nitrogen oxidation cycle (nitrite oxidation to nitrate as electron donor for photosynthesis)12–15. However, nitrification or dissimilatory nitrate reduction to ammonium (DNRA) pathway has not been reported in purple bacteria yet. Denitrification and assimilation nitrate reduction cycles were only reported in PNSB, but unknown in PSB16. Currently, the molecular mechanism of nitrogen cycles in purple bacteria remain unclear17–19. To elucidate nitrogen cycles in purple bacteria, we sequenced the genome of YL28 as a representative strain, compared the nitrogen cycle genes to other 35 sequenced purple bacteria. The genes involved in the sulfur metabolism, salt tolerance and the stress response were also investigated in details to gain the insights of the surviving mechanisms of YL28 in the highly selective environments.
Results
Genome features and phylogenic inference
The general features of genome of YL28 and the other 35 sequenced purple bacteria were presented in Table 1. Removal of short contigs and sequences with potential contamination resulted in 120 contigs with a N50 of 93,809 bp. The largest contig size was 176,489 bp. The assembled genome comprised 3.8 million nucleotides with a GC content of 68.84%. Gene annotation was carried out by the NCBI Prokaryotic Genome Annotation Pipeline (PGAP). A total of 3,361 genes yielded a coding capacity of 3.3 million nucleotides (genes/genome, 86.4%). There were at least 69 RNA sequences, including 56 tRNAs, 9 rRNAs, and 4 function unknown RNAs in the genome. Among the protein-encoding genes, 1,752 of them could be assigned putative functions while 1,612 were predicted to encode hypothetical proteins. At least 1,671 proteins were assigned to 25 different functional categories with 368 subsystems using SEED subsystems by RAST analysis (Supplementary Fig. SF1). 40 genes involved in nitrogen metabolism were found (Supplementary Fig. SF1). Compared to its relative M. purpuratum 984, YL28 has more genes for nitrogen metabolism, photosynthesis and sulfur metabolism (Supplementary Fig. SF2, Tables 2 and 3).
Table 1.
The genome characteristics of purple bacteria used in this study.
| Organisms | Chromosome | Plasmid | Size (Mb) |
Chr 1 (Mb) |
Chr 2 (Mb) |
G + C % |
Protein | gene | Accession |
|---|---|---|---|---|---|---|---|---|---|
| YL28 | 1 | — | 3.84 | 3.84 | — | 68.9 | 3,188 | 3,360 | LSYU00000000.1 |
| Thiocystis violascens DSM 198 | 1 | — | 5.02 | 5.02 | — | 62.6 | 4,261 | 4,465 | CP003154.1 |
| Allochromatium vinosum DSM 180 | 1 | 2 | 3.67 | 3.53 | — | 64.4 | 3,148 | 3,262 | CP001896.1 |
| Marichromatium purpuratum 984 | 1 | — | 3.78 | 3.78 | — | 67.9 | 3,159 | 3,272 | EU850807.1 |
| Ectothiorhodospira sp. BSL-9 | 1 | — | 3.55 | 3.55 | — | 63 | 3118 | 3,249 | CP011994.1 |
| Thioflavicoccus mobilis 8321 | 1 | 1 | 4.13 | 4.05 | — | 65.5 | 3462 | 3,602 | NR102479.1 |
|
Halorhodospira halophila SL1 extremly halophilic |
1 | — | 2.68 | 2.68 | — | 68 | 2375 | 2,458 | CP000544.1 |
| Rhodopseudomonas palustris CGA009 | 1 | 1 | 5.47 | 5.46 | — | 65 | 4,820 | 4,898 | BX572608.1 |
| Rhodopseudomonas palustris HaA2 | 1 | — | 5.33 | 5.33 | — | 66 | 4,683 | 4,772 | CP000250.1 |
| Rhodopseudomonas palustris BisB18 | 1 | — | 5.51 | 5.51 | — | 65 | 4,886 | 5,016 | CP000301.1 |
| Rhodopseudomonas palustris BisB5 | 1 | — | 4.89 | 4.89 | — | 64.8 | 4,397 | 4,492 | CP000283.1 |
| Rhodopseudomonas palustris BisA53 | 1 | — | 5.51 | 5.51 | — | 64.4 | 4,878 | 4,972 | CP000463.1 |
| Rhodopseudomonas palustris TIE-1 | 1 | — | 5.74 | 5.74 | — | 64.9 | 5,246 | 5,382 | CP001096.1 |
| Rhodopseudomonas palustris DX-1 | 1 | — | 5.4 | 5.4 | — | 65.4 | 4,917 | 5,082 | EU221583.1 |
| Rhodoplanes sp. Z2-YC6860 | 1 | — | 8.19 | 8.19 | — | 63.5 | 7443 | 7,606 | GQ369128.3 |
| Blastochloris viridis | 1 | — | 3.73 | 3.72 | — | 67.9 | 3107 | 3,230 | D14430.2 |
| Blastochloris viridis ATCC19567 | 1 | — | 3.73 | 3.72 | — | 67.9 | 3104 | 3,238 | D2 5314.1 |
| Blastochloris viridis DSM_133 | 1 | — | 3.72 | 3.72 | — | 67.9 | 3114 | 3,236 | HQ009851.1 |
| Rhodospirillum rubrum ATCC 11170 | 1 | 1 | 4.41 | 4.35 | — | 65.3 | 3,838 | 3,917 | CP000230.1 |
| Rhodospirillum rubrum F11 | 1 | — | 4.35 | 4.35 | — | 65.4 | 3,878 | 3,946 | CP003046.1 |
|
Rhodospirillum centenum
SW (Rhodocista centenaria) |
1 | — | 4.36 | 4.36 | — | 70.5 | 4,003 | 4,102 | CP000613.2 |
|
Pararhodospirillum photometricum
DSM 122 |
1 | — | 3.88 | 3.88 | — | 64.7 | 3,117 | 3,434 | HE663493.1 |
|
Rhodomicrobium vannielii
ATCC 17100 |
1 | — | 4.01 | 4.01 | — | 62.2 | 3,565 | 3,793 | CP002292.1 |
| Rhodobacter sphaeroides 2.4.1 | 2 | 5 | 4.6 | 3.19 | 0.94 | 68.8 | 4,242 | 4,370 | CP000144.2 |
| Rhodobacter sphaeroides MBTLJ-8 | 2 | 4 | 4.66 | 3.19 | 0.94 | 68.7 | 4204 | 4,362 | CP012960.1 |
| Rhodobacter sphaeroides MBTLJ-13 | 2 | 5 | 4.65 | 3.19 | 0.94 | 68.8 | 4200 | 4,358 | CP015210.1 |
| Rhodobactersphaeroides ATCC 17029 | 2 | 1 | 4.49 | 3.15 | 1.22 | 69 | 4,131 | 4,254 | CP000577.1 |
| Rhodobacter sphaeroides ATCC 17025 | 1 | 5 | 4.56 | 3.22 | — | 68.2 | 4,333 | 4,466 | CP000661.1 |
| Rhodobacter sphaeroides KD131 | 2 | 2 | 4.71 | 3.15 | 1.3 | 69.1 | 4,569 | 4,638 | CP001151.1 |
| Rhodoferax ferrireducens T118 | 1 | 1 | 4.97 | 4.71 | — | 59.6 | 4,436 | 4,554 | CP000267.1 |
| Rhodobacter capsulatus SB 1003 | 1 | 1 | 3.87 | 3.74 | — | 66.6 | 3,642 | 3,740 | CP001312.1 |
| Rhodobacter sp. LPB0142 | 1 | 3 | 3.78 | 3.46 | — | 66 | 3530 | 3,660 | KX066852.2 |
| Rhodovulum sulfidophilum DSM1374 | 1 | 2 | 4.34 | 4.13 | — | 66.9 | 3876 | 4,002 | CP015418.1 |
| Rhodovulum sulfidophilum DSM2351 | 1 | 3 | 4.73 | 4.45 | — | 66.9 | 4155 | 4,340 | D16430.1 |
| Rhodovulum sulfidophilum SNK001 | 1 | 1 | 4.19 | 4.08 | — | 66.8 | 3746 | 3,873 | CP015421.1 |
| Rubrivivax gelatinosus IL144 | 1 | — | 5.04 | 5.02 | — | 71.2 | 4505 | 4,591 | AB016167.1 |
Table 2.
The key enzymes responsible for different nitrogen metabolism in purple bacteria.
| Species | Fermentative nitrate reduction | Aissimilatory nitrate reduction | Denitrification | Nitrogen fixation | |||||
|---|---|---|---|---|---|---|---|---|---|
| narGHIJ/napAB | nirBD/nrfAH | narB/NR/nasAB | NIT-6/nirA | narGHIJ/napAB | nirK/nirS | norBC | nosZ | nifDKH/anfG | |
| R. palustris CGA009 | nirA | nirK | norBC | nosZ | nifDKH/anfG | ||||
| R. palustris HaA2 | nirBD | nasAB | nirA | nifDKH | |||||
| R. palustris BisB18 | napAB | napAB | nosZ | nifDKH/anfG | |||||
| R. palustris BisB5 | nifDKH/anfG | ||||||||
| R. palustris BisA53 | nirBD | nasAB | nirA | nirK | norBC | nosZ | |||
| R. palustris TIE-1 | nirA | nirK | norBC | nosZ | |||||
| R. palustris DX-1 | nirA | nirK | norBC | nosZ | nifDKH/anfG | ||||
| Rhodoplanes sp. Z2-YC6860 | narGHIJ | nirBD | nasAB | ||||||
| B. viridis | nifDKH/anfG | ||||||||
| B. viridis ATCC19567 | nifDKH/anfG | ||||||||
| B. viridis DSM_133 | nifDKH/anfG | ||||||||
| R. vannielii ATCC 17100 | napAB | napAB | norBC | nifDKH | |||||
| R. sphaeroides 2.4.1 | napAB | napAB | nirK | norBC | nifDKH | ||||
| R. sphaeroides ATCC17029 | norBC | nifDKH | |||||||
| R. sphaeroides ATCC17025 | napAB | nirK | norBC | nosZ | nifDKH | ||||
| R. sphaeroides KD131 | nirK | norBC | nosZ | ||||||
| R. capsulatus SB1003 | nosZ | nifDKH/anfG | |||||||
| R. sphaeroides MBTLJ-13 | napAB | napAB | norBC | nifDKH | |||||
| R. sphaeroides MBTLJ-8 | napAB | napAB | norBC | nifDKH | |||||
| R. rubrum ATCC11170 | nifDKH/anfG | ||||||||
| R. rubrum F11 | nifDKH/anfG | ||||||||
| R. centenum SW | napAB | napAB | norBC | nosZ | nifDKH | ||||
| P. photometricum DSM 122 | nifDKH/anfG | ||||||||
| R. vannielii ATCC17100 | norBC | nifDKH/anfG | |||||||
| R. ferrireducens T118 | narGHIJ | nirBD | nasAB | narGHIJ | norBC | nosZ | |||
| R. sulfidophilum DSM1374 | nifDKH | ||||||||
| R. sulfidophilum DSM2351 | nifDKH | ||||||||
| R. sulfidophilum SNK001 | nifDKH | ||||||||
| R. gelatinosus IL144 | nirK | norBC | nosZ | nifDKH | |||||
| T. violascens DSM198 | napAB | napAB | nifDKH | ||||||
| A. vinosum DSM180 | nirA | norBC | nifDKH | ||||||
| M. purpuratum 984 | otr | nasAB | nirK | norBC | nifDKH | ||||
| YL28 | nrfAH | nasAB | nirA | nirS | norBC | nifDKH | |||
| H. halophila SL1 | narGHIJ | nrfAH | narGHIJ | norBC | nifDKH | ||||
| Ectothiorhodospira sp. BSL-9 | nrfAH | nifDKH | |||||||
| T. mobilis 8321 | nifDKH | ||||||||
Table 3.
The key enzymes responsible for different sulfur metabolism in purple sulfur bacteria (PSB).
| Species | Oxidation of sulfide | Reversed dissimilatory sulfite reductase | Sox system | |||||
|---|---|---|---|---|---|---|---|---|
| FCC/SQR | DsrAB | AprAB | Sat | SoxB | SoxY/SoxZ | SoxA/SoxX | SoxC/SoxD | |
| A. vinosum DSM180 | SQR | DsrAB | AprAB | Sat | SoxB | SoxY/SoxZ | SoxA/SoxX | — |
| Ectothiorhodospira sp. BSL-9 | FCC | — | — | Sat | — | — | — | — |
| H. halophila SL1 | SQR | DsrAB | — | — | SoxB | SoxY/SoxZ | SoxA | — |
| YL28 | SQR | DsrAB | AprAB | Sat | SoxB | SoxY/SoxZ | SoxA/SoxX | SoxD |
| M. purpuratum 984 | SQR | DsrAB | — | — | SoxB | SoxY/SoxZ | SoxA/SoxX | SoxD |
| T. violascens DSM198 | FCC | DsrAB | AprAB | Sat | SoxB | SoxY/SoxZ | SoxA/SoxX | — |
| T. mobilis 8321 | FCC | DsrAB | AprAB | Sat | — | — | — | — |
No plasmid sequences were discovered in YL28. However, at least one was found in Rhodovulum spp, Rhodobacter spp, Allochromatium spp, R. rubrum ATCC 11170, T. mobilis 8321 and R. palustris CGA009 (Table 1). Moreover, most of the Rhodobacter strains possesses up to two chromosomes. The genomic size of the purple bacteria ranges from 2.6 to 5.4 Mb. Except for T. mobilis 8321 and T. violascens DSM 198, the genome size in PSB is smaller than that in PNSB (Table 1). Overall, genome size in most of PSB is smaller than 4.0 MB, with the smallest one (2.68 Mb) in Halorhodospira halophila SL1. Except for Blastochloris, genome size of PNSB is generally larger than 4.0 MB; among them, Rhodoplanes sp. Z2-YC6860 has the largest genome size (8.19 Mb). Collectively, a high GC content (>55%) is often observed in purple bacteria.
Phylogenic inference
The phylogenetic analysis based on 16S rDNA sequence (Fig. 1A) showed that PSB and PNSB derived from a common ancestor and formed two clades, which was consistent with the previous studies20. 28 PNSB strains were clustered into three subdivisions: the first one consisted of Rhodopseudomonas, Blastochloris, Rhodoplanes and Rhodomicrobium vannielii ATCC 17100; the second one included Rhodobacter (freshwater species) and Rhodovulum (marine species); the third one had Rhodospirillum and Rhodocista. 7 PSB species were clustered together on the same clade. Both the phylogenetic trees based on whole genome or core genome (Fig. 1B,C) showed that some PSB species were included in PNSB. Interestingly, Rubrivivax gelatinosus IL144 (a member of PNSB) was the closest phylogenetic neighbors of PSB based on 16S rDNA sequence and core genome.
Figure 1.
Phylogenetic analyses between PSB and PNSB. The NJ algorithm tree of 16S rRNA genes of 36 strains from purple bacteria by the MEGA 6.06 (A); the Mauve guide tree of the 35 strains based on whole-genomic similarity at the nucleotide level through multiple genome comparison tool Mauve (B); the NJ tree of 77 conserved proteins shared among the 36 strains by the BPGA (C).
Gene repertoire of PSB
The power trend line has not reached the plateau (Fig. 2A), demonstrating that PSB displays an open accessory-genome. The core genome analysis of PSB shows that the numbers of shared genes decreases with the addition of the input genomes and is predicted to converge against 550 (Fig. 2A). Singleton development plot data indicates that up to 1594 new genes are expected with each newly added genome (Fig. 2B). The core genomes for the seven PSB species contain at least 539 CDSs per genome. The current core genome represents the PSB quite well. YL28 and M. purpuratum 984 share at least 2869 genes (data not shown); only 241 and 210 genes uniquely occur in YL28 and 984, respectively. The shared gene numbers in genome between YL28 and other six PSB species range from 732 to 2869 genes (data not shown).
Figure 2.
Pan, core and singleton genome evolution according to the number of selected purple bacteria genomes. (A) Number of genes (core genome and Pan-genome) for the selected purple bacteria genomes sequentially added. (B) Number of unique genes for the selected purple bacteria genomes sequentially added.
Nitrogen cycle genes in YL28 and purple bacteria
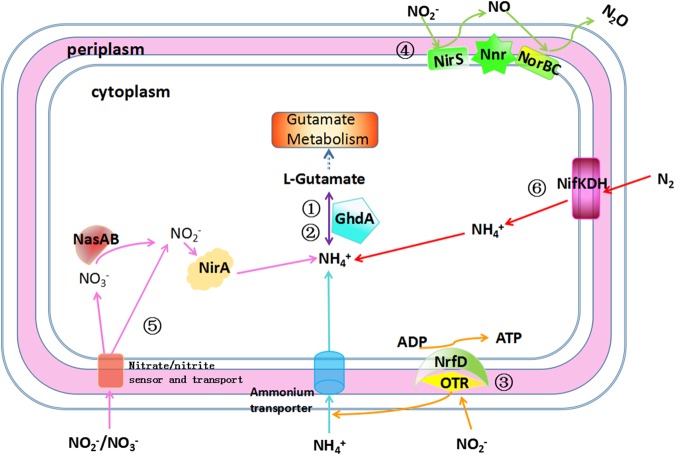
YL28 possesses six nitrogen cycle pathways (Fig. 3 and Table 2). Two genes encoding NirS and NorBC (the key enzymes for nitrite reduction) exists in YL28. These two enzymes in YL28 possibly contribute to transform toxic nitrite into lower cytotoxic N2O by denitification5. The genes encoding NasAB and NirA (the key enzymes for assimilation nitrite reduction) were also detected in YL28, suggesting that it had the complete nitrate reduction ability. YL28 has the genes encoding Nrf which is critical in fermentative nitrate reduction (DNRA). However, the DNRA pathway in YL28 may be incomplete due to the lack of entire complement enzymes. Ammonium assimilation and ammonification related genes were detected in all purple bacteria. Additional four nitrogen cycle pathways display diverse metabolism traits in examined strains (Table 2). The presence of genes encoding NifDKH key enzymes in all purple bacteria shows this group of bacteria has nitrogen fixation potentials. Interestingly, the alternative nitrogenase (AnfG) from Methanosarcina was only found in PNSB but not in PSB21.
Figure 3.
The model of nitrogen metabolism in M. gracile YL28. There are six nitrogen metabolic pathways in YL28 whose key enzyme confirmed by CDART and CDD. (A) Ammonification pathway; (B) ammonium assimilation pathway; (C) fermentative nitrate reduction; (D) denitrification; (E) assimilation nitrate reduction; (F) nitrogen fixation.
Sulfur metabolism genes in YL28
YL28 has genes invovled in at least three sulfur metabolism pathways such as oxidation of sulfide, reversed dissimilatory sulfite reduction and sox system (Table 3). The sqr and the dsr gene cluters (dsrABEFHCMKLJOPNR) are the key enzyme genes for the oxidation of sulfide and the reversed dissimilatory sulfite reduction. These process oxidize toxic sulfide into S0 or sulfate20. In addition, sox gene clusters in YL28 shows that thiosulfate is possbly converted into S0 or sulfate by the truncated Sox system. Moreover, the presence of aprAB and sat genes (encoding adenylylsulfate reductase, sulfate adenylyltransferase, respectively) suggests that YL28 possibly possesses an alternative sulfite oxiditon pathway (converting toxic sulfite into sulfate).
Halo-tolerance
YL28 possessed a gene cluster involved in the salt-alkali tolerance (nhaABCDEFG, encoding Na+/H+ antiporter). Moreover, a novel putative Na+/H+ antiporter gene (duf 2062) exsit in YL28. The unique stress response subsystem in YL28 was listed in Table 4. There are six unique proteins participating in choline and betaine uptake and betaine biosynthesis (betaine aldehyde dehydrogenase, choline dehydrogenase), synthesis of osmoregulated periplasmic glucans (phosphoglycerol transferase I, cation channel protein), heavy metal resistence (DNA-binding heavy metal response regulator, heavy metal sensor histidine kinase).
Table 4.
The unique stress response protein in YL28.
| Category | Subcategory | Subsystem | Role |
|---|---|---|---|
| Virulence, Disease and Defense | Resistance to antibiotics and toxic compounds | Cobalt-zinc-cadmium resistance | DNA-binding heavy metal response regulator |
| Resistance to antibiotics and toxic compounds | Cobalt-zinc-cadmium resistance | Heavy metal sensor histidine kinase | |
| Resistance to antibiotics and toxic compounds | Cobalt-zinc-cadmium resistance | cation channel protein | |
| Stress Response | Osmotic stress | Choline and Betaine Uptake and Betaine Biosynthesis | Betaine aldehyde dehydrogenase (EC 1.2.1.8) |
| Osmotic stress | Choline and Betaine Uptake and Betaine Biosynthesis | Choline dehydrogenase (EC 1.1.99.1) | |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | Phosphoglycerol transferase I (EC 2.7.8.20) |
Genome islands (GIs) of YL28
At least 9 GIs were identified with YL28 (Fig. 4A,B and Supplemental Table ST1) by both IslandPick and IslandPath-DIMOB methods. The GI size ranges from 8 to 31.3 Kb (Supplemental Table ST1). These GIs were noted as follows: the mobile element proteins, transposase, phage structure proteins, integration host factor, proteins involved in CRISPR system, 5-methylcytosine-specific restriction related enzyme, cephalosporin hydroxylase, flagellum synthesis component (CheABR), carbohydrate/nitrogen metabolism proteins (phenylalanyl-tRNA metabolism, Threonyl-tRNA biosynthesis, amylomaltase, asparagine synthetase, glycosyl transferase, aspartate aminotransferase), DNA replication and proofreading systems (DNA recombination, repair DNA recombination and repair protein RecF Type II restriction enzyme, ATP-dependent endonuclease of the OLD family, ATP-dependent DNA helicase pcrA, DNA processing chain A) (Fig. 4A,B and Supplemental Table ST1).
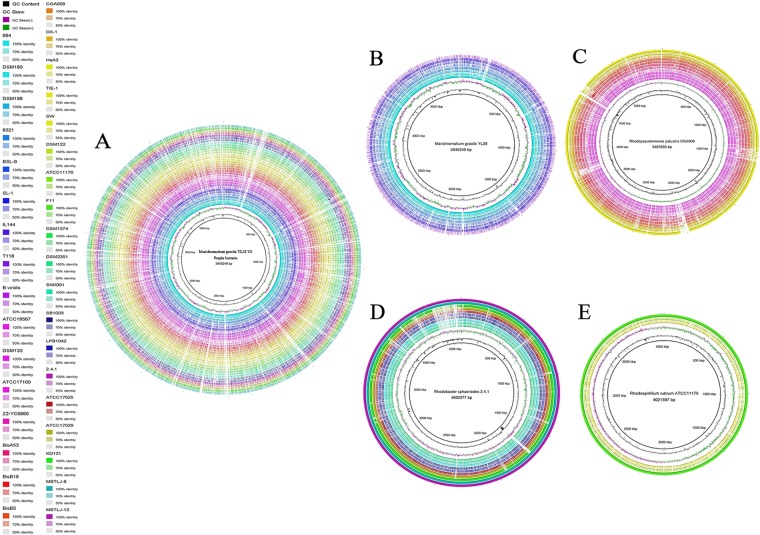
Figure 4.
Whole-genome comparisons in three genera of purple bacteria. The color intensity in each ring represents the BLAST match identity. (A) Whole-genome comparison of all the strains considered in this work; (B) Whole-genome comparisons of strains of purple sulfur bacteria and T118, IL144. The color intensity in each ring represents the BLAST match identity. from inner to outer ring: M. purpuratum 984, A. vinosum DSM 180, T. violascens DSM 198, T. mobilis 8321, Ectothiorhodospira sp. BSL-9, H. halophila SL1, R. gelatinosus IL144, R. ferrireducens T118. (C) Whole-genome comparisons of 12 strains of purple non-sulfur bacteria. The color intensity in each ring represents the BLAST match identity. from inner to outer ring: R. palustris BisA53, BisB18, BisB5, DX-1, HaA2, TIE-1, B. viridis, B. viridis ATCC19567, B. viridis DSM133, Rhodoplanes sp. Z2-YC6860, R. vannielii ATCC 17100. (D) Whole-genome comparisons 11 strains of two gena in Rhodobacter and Rhodovulum in purple non-sulfur bacteria. The color intensity in each ring represents the BLAST match identity from inner to outer ring: R. capsulatus SB 1003, Rhodobacter sp. LPB0142, R. sphaeroides ATCC 17025, R. sphaeroides ATCC 17029, R. sphaeroides KD131, R. sphaeroides MBTLJ-13, R. sphaeroides MBTLJ-8, R. sulfidophilum DSM1374, R. sulfidophilum DSM2351, R. sulfidophilum SNK001; (E) whole-genome comparisons in Rhodobacter, from inner to outer ring: R. rubrum F11, R. centenum SW, P. photometricum DSM 122.
Among these predicted GIs, GI-IX has the largest size (31.3 kb) in YL28. Notably, GI-IX has many DNA metabolic related genes including DNA recombination and repair protein RecF, ATP-dependent endonuclease of the OLD family, ATP-dependent DNA helicase pcrA, ATP-dependent DNA helicase RecQ, DNA processing chain A and type II restriction enzyme. It also contains some genes encoding diverse enzymes such as aspartate aminotransferase, methylase and ATP-dependent protease. The second largest GI (GI-II) carries a chemoreceptor gene cluster including the cheA, cheB, cheR and a gene encoding the methyl-accepting chemotaxis protein. Moreover, GI-II has the predicted two-component hybrid sensor and regulator, 4-alpha-glucanotransferase (amylomaltase), and mobile element protein (Supplemental Table ST1). Immediately downstream of GI-II, there are genes involved in the DNA metabolism such as 5-methylcytosine-specific restriction related enzyme, DNA helicase, phosphatase 2C homolog and 5-methylcytosine-specific restriction related enzyme. The size of other GIs is smaller than GI-IX.
There are five GIs in Rhodopseudomonas, Blastochloris and Rhodoplanes (Supplemental Table ST2). These GIs carry nitrogen-fixing genes (nitrogenase gene and alternative nitrogenase gene), LysR family of proteins transcriptional regulation, glycosyltransferase protein family and arsenic resistance (ArsH). There is an alternative nitrogenase (AnfG) in a GI region of R. palustris, which replaces nitrogenase (NifDKH) for nitrogen fixation22. At least 10 GIs are predicted in Rhodobacter (Supplemental Table ST3). Among these GIs, there are many genes involved in nitrogen metabolism including a nitrogen-fixing island (nitrogenase genes, oxidoreductase/nitrogenase component), a nitrogen-rich23 island (nitrate/sulfonate/bicarbonate ABC transporter, nitrogen regulatory protein P-II), and a solid-island24 (metallophosphoesterase). Moreover, some GIs were found to have several genes that involved in sulfur metabolism, biosynthesis of flagella, phage infection area and BadM/Rrf2 family of transcriptional regulatory protein. Rhodobacter possibly acquires nitrogen-fixation and the sulfur metabolism by the horizontal gene transfer (HGT). At least 11 GIs were predicted in genus Rhodospirillum (Supplemental Table ST4). Among those GIs, Rhodospirillum has gene elements including CRISPR, sulfide metabolism, arsenic resistance and the DNA and ribosomal protein synthesis, allowing this bacterium to survive in toxic environments under certain concentration of arsenic and sulfide.
Synteny analysis
A good collinearity relationship was shown in PSB using the synteny plots analysis (Supplementary Fig. SF3, YL28 as the reference genome). YL28, T. violascens DSM 198, A. vinosum DSM 180, T. mobilis 8312 and M. purpuratum 984 have the closer relationship than do Ectothiorhodospira sp. BSL-9 and H. halophila SL1. When R. palustris CGA009, R. sphaeroides 2.4.1 or R. rubrum ATCC 11170 was selected as a reference sequence, PNSB members showed a diverse collinearity relationship (Supplementary Figs SF4–SF6). Three groups are divided: the first one consists of the 4 genera including Rhodopseudomonas, Blastochloris, Rhodoplanes, and Rhodomicrobium (Fig. SF4); the second one consists of the 2 genera Rhodobacter and Rhodovillum (Fig. SF5); the third one only contains Rhodospirillum (except R. centenum SW) (Fig. SF6). Interestingly, a good collinearity was found between R. ferrireducens T118 (a non-phototrophic member) and PSB. Instead, a relatively poor collinearity was presented among PSB, R. gelatinosus IL144 and R. centenum SW. Collectively, the synteny pattern is consistent with the whole genome phylogenetic tree (Fig. 1B). The poorest collinearity relationship was shown between YL28 and R. gelatinosus IL144.
Discussion
The interfaces of mangrove locate between land and sea in the tropical and the sub-tropical latitudes25. PSB and PNSB are predominantly detected in the mangrove ecosystems. The two bacterial groups significantly contribute to the primary productivity of coastal seaboards and the food-web dynamics of various tropical coastline ecosystems26,27. The high salinity, limited nutrients and S-richness sulfate concentrations often occur in the niche25. However, it remains unclear about how the mangrove-associated microorganisms survive in the mangrove ecosystems and contribute to the host physiology. Particularly, the nitrogen utilization mechanisms in PSB and PNSB have not well studied. In this study, we sequenced the genome of YL28, investigated the nitrogen cycle pathways and compared them to other purple bacteria. Our study will contribute to elucidate bacterial surviving mechanisms in the special mangrove ecosystem.
YL28 utilized ammonium, nitrite or nitrate as the sole nitrogen source for phototrophic growth3–5. In the present study, our results reveal that the strong ability for nitrate and nitrite utilization may be due to the six nitrogen metabolic pathways. Ammonium is preferred nitrogen sources by all species of purple bacteria28. YL28 may grow on ammonium by the ammonium assimilation pathway, and converted organic nitrogen into ammonium by ammonification. Moreover, YL28 utilized N2 as nitrogen source to support cell growth by nitrogen fixation when the absence of ammonium happened. It is interesting that YL28 grew chemoheterotrophically using the nitrate and/or nitrite as electron acceptor. The denitrification and/or fermentative nitrate reduction and/or assimilation nitrate reduction pathways may contribute to this process under the anoxic condition. Diverse nitrogen utilization pathways allow YL28 to grow on the different nitrogen compounds. YL28 can convert nitrite (toxic compounds in the mangrove ecosystems) into non-toxicity or low-toxicity products by fermentative nitrate reduction or assimilation nitrate reduction pathways. Denitrification may also contribute to this process5.
In this study, our bioinformatics studies further show that purple bacteria have 6 nitrogen cycle pathways (Table 3). The alternative nitrogenase genes are frequently observed in purple bacteria. The genes (encoding two nitrogenase subunits) are highly conserved in many phyla of bacteria and archaea, suggesting that nitrogen fixation genes evolve once and subsequently spread by vertical in heritance or by HGT9. Nitrate/nitrite reduction-related genes were found only in the accessory genomes. The observation agrees with the previous studies that show that nitrate/nitrite is not preferred nitrogen sources29. The partial denitrification pathway was widely found in PNSB30,31. To best of our knowledge, our results first demonstrated that the partial denitrification pathway existed in PSB, suggesting that some PSB possibly also played important roles in the nitrogen cycle (e.g., Rhodopseudomonas and Rhodobacter). The partial denitrification pathway contributes to remove excess redox and mitigates the toxicity from the certain nitrogen oxide intermediates20. The fermentative nitrate reduction is less known in purple bacteria and assimilation nitrate reduction is only limited to a member of PNSB (Rhodobacter capsulatus E1F1). However, our results showed the fermentation nitrate reduction (DNRA) was possible occurred in purple bacteria, and assimilation nitrate reduction was in PSB.
The use of reduced sulfur compounds as electron donors for anoxygenic photosynthesis has been found in all groups of purple bacteria (PSB and PNSB). Remarkably, PSB utilize higher concentration reduced sulfur compounds and accumulated element sulfur inside the cells than PNSB20,32,33. Our previous studies showed that YL28 was capable of utilizing diverse reduce sulfur compound and depositing sulfur granules inside the cells. It demonstrated a higher tolerance (3.6 mM) to sulfide than PNSB (0.5–2 mM)1,2. YL28 has the diverse sulfur metabolism-related genes such as sqr, dsr, sox, apr and sat. The oxidation of sulfide and the reversed dissimilatory sulfite reduction (key enzyme gene sqr and dsr) possibly contribute to detoxification of toxic sulfide. Thiosulfate may be converted into S0 or sulfate by truncated Sox system. Moreover, an alternative pathway (key genes apr and sat) also allows to detoxifing the toxic sulfite. However, sulfite may not be directly converted into sulfate via a two-electron transfer because purple bacteria lack of the sulfite:cytochrome c oxidoreductase33. Without soxC/soxD, sulfane sulfur atom cannot be subsequently oxidized34. However, YL28 has soxD. The study on the complete sulfur oxidation in YL28 by soxD needs to be investigated. Some genes involved in classical sulfite metabolism were not detected in most purple bacteria34 (except for some freshwater PSB species such as A. vinosum DSM180)35. However, YL28 has both sulfide: quinone reductase gene (sqr, YL28 could directly oxidize S2− to S0) and APS reductase gene (apr). This suggests that YL28 may oxidize SO32− to SO42− and reduce the sulfite toxicity to cells. Sulfite reductase gene (dsr) was observed in all of the selected PSB (not PNSB). Gene dsr may be a useful taxonomic or systematic marker for PSB phylogeny.
Our studies revealed that there was a distinctive difference in halo-tolerance traits between freshwater and salt-dependent species by the 16S rDNA, core or whole genome sequence analysis. The salt-dependent or salt-requiring species showed different halo-tolerance. For example, YL28 possesses unique stress response genes (betA, duf 2062, mrp, glucan and heavy metal response genes etc.). These genes possibly contribute to the tolerance to in high salinity environment36,37.
The size of genome, the numbers and size of GIs in PSB were smaller than those in PNSB, implying that PSB had more flexible response to environment change. A gene transfer agent (GTA) is an unusual bacteriophage-like element which transfers a random host genomic DNA fragments (4–14 kb in size) between closely related bacteria38–40. Genes involved in the photosynthesis may be horizontally transferred between the same phyla by GTA41. Our results revealed that GTA gene clusters including ICEs (the self-transmissible mobile genetic elements, integrative and conjugative elements) were presented in all examined genomes. PNSB may acquire more genes by HGT to survive in the niches. The flagellum biosynthesis and methyl accepting chemotaxis genes (such as cheA, cheB and cheR) were found in the larger GIs, implying that purple bacteria possibly employ a complex set of chemosensory pathways to swim towards carbon and nitrogen sources, light and/or oxygen42.
Conclusion
This study provided a novel insight into the mechanisms of diverse nitrogen cycle, habitat-specificity and toxic nitrite utilization in YL28 and purple bacteria (including PSB and PNSB). Purple bacteria possess 6 nitrogen cycle pathways. The denitrification, complete assimilation nitrate reduction and fermentative nitrate reduction were first demonstrated in PSB. The fermentative nitrate reduction was possibly widely occurred in purple bacteria. YL28 possesses good ability to utilize toxic nitrite which is possibly linked to the combination of three nitrogen cycle pathways (denitrification, fermentative nitrate reduction and complete assimilation nitrate reduction). Collectively, the genes involved in diverse nitrogen cycles (6 pathways), sulfur cycles (3 pathways), unique salt-alkali tolerance, stress response as well as other traits contribute to bacterial adaptation to the mangrove habitat.
Materials and Methods
DNA extraction, genome sequencing and annotation
YL28 was isolated from an intertidal sediment of the inshore mangrove in Fujian, China2. Genomic DNA of YL28 was extracted using an TIANamp Bacterial DNA kit following the protocols recommended by the manufacturer. Genome sequencing was performed by Shanghai Majorbio Bio-Pharm Technology Co. (China) using the Illumina HiSeq2000 sequencer system with a 500 bp pair-end library. The reads were assembled using SOAPdenovo v2.04. Putative protein-encoding genes were identified using Glimmer 3.0243. Annotation was performed by BLAST +2.2.24 searching against databases, including the National Center for Biotechnology Information (NCBI), Clusters of Orthologous Groups of Proteins (COG), the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene ontology (GO). The genome sequence of M. gracile YL-28 was deposited in the GenBank database under the accession number LSYU0000000044.
Acquisition and re-annotation of the selected genome sequences
The genome sequences of other 35 sequenced purple bacteria were obtained from NCBI database which assessed level was complete. To avoid the possible deviations due to different annotation methods, we used Rapid Annotation using Subsystem Technology (RAST) server for reannotation45 of the selected genomes. Glimmer algorithm was used for gene calling.
Phylogeny
Three different methods were used for constructing the phylogeny trees. The 16S rDNA sequences of the selected microorganisms were first used to infer phylogenies using the Neighbor-Joining (NJ) method of MEGA 6.0.646. The core genomes for these selected genomes were next clustered by USEARCH and the phylogeny trees of core and accessory genomes were constructed by Neighbor-Joining algorithm using the BPGA-1.347. The 36 genome sequences were aligned by the progressive MAUVE to generate a phylogenetic guide tree48.
Comparisons of conserved and variable regions
The genome of YL28 was used as a reference sequence to align all other genomes of 35 purple bacteria. Similarly, the genome of R. palustris CGA009 was selected as a reference to align those in Rhodopseudomonas, Blastochloris and Rhodoplanes; the genome of R. sphaeroides 2.4.1 was used as a reference to align the genomes of two genera Rhodobacter and Rhodovulum; R. rubrum ATCC11170 was selected as a reference to align the genomes of Rhodospirillum genus. The multiple whole–genome alignment was conducted using the progressive alignment algorithm implemented in MAUVE. Syntheny plots were generated by aligning regions of the predicted and reference genomes that differed by default parameter. All regions were aligned and displayed in MAUVE version 2.3.148. The circular genomic map was generated by the BLAST Ring Image Generator (BRIG, version 0.95)49 using alignment reference genome on a local BLAST + basis, with standard parameters (50% lower – 70% upper cut-off for identity and E-value of 10). The ring color gradients correspond to varying degrees of identity of BLAST matches. Circular genomic maps also include the information on GC skew and GC content.
Comparative analyses of core genome, accessory genome and unique genome
In order to depict the core and accessory genome in each genus, a reciprocal best hit search using the BPGA software was performed47. Orthologous clusters (OCs) were assigned by grouping all protein sequences in the 36 genomes using USEARCH based on their sequence similarity (E-value < 10−5, >50% coverage). A series of built-in scripts were used to (i) parse, (ii) upload to the MySQL relational database, (iii) perform a reciprocal best hit analysis to form pairs of sequences, and (iv) normalize the E-values for all the pairs formed. Normalization of E-values was done by averaging all recent ortholog (in paralogs) and dividing each pair of ortholog by the average. Pan-core plot against combinations will give core and pan genome boxplot and dot plot generated using desired number of unique combinations of genomes. Atypical GC analysis will give sequences of core, accessory and unique genes with atypical (extreme) GC content. COG and KEGG distributions of the core, accessory and unique gene families were calculated based on representative sequences. Keywords were used to query for nitrogen metabolism functional in the orthologous families and to calculate the number of matches of those functions, using custom bash commands.
Genomic island prediction
Genomic islands (GIs) were predicted by using IslandViewer350 including IslandPick, IslandPath-DIMOB and SIGI-HMM.
Nitrogen metabolism, sulfur metabolism and stress response analysis
We chose three RAST subsystems for further analyze the nitrogen metabolism, sulfur metabolism and stress response pathways. The key protein sequences51–53 responsible for nitrogen metabolism, sulfur metabolism was used as reference sequences to search against the 36 genomes of purple bacteria by BlastP. The obtained sequence by BLAST was further evaluated by the combination of KEGG, RAST, CDART (conserved domain architecture retrieval tool54 and CDD (the conserved domain database)55.
Electronic supplementary material
Acknowledgements
This work was supported by National Marine Public Industry Research (201505026); by Natural Science Foundation of Fujian Province (Fujian Provincial Natural Science Foundation) (2018J01049); by Subsidized Project for Cultivating Postgraduates Innovative Ability in Scientific Research of Huaqiao University.
Author Contributions
B.Z., X.Z., C.Z., S.Y. and S.C. conceived the study, analyzed the data and wrote the paper.
Competing Interests
The authors declare no competing interests.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Shicheng Chen, Email: shicheng@msu.edu.
Suping Yang, Email: yangsuping@hqu.edu.cn.
Electronic supplementary material
Supplementary information accompanies this paper at 10.1038/s41598-018-36160-2.
References
- 1.Trüper, H. G. & Pfennig, N. Characterization and identification of the anoxygenic phototrophic bacteria, 1. [Starr, M. P., Stolp, H., Trüper, H. G., Balows, A. & Schlegel, H. G. (eds)] The prokaryotes18, 299–312. (Springer, Berlin, Heidelberg, 1981).
- 2.Zhao JY, et al. Identification and characterization of a purple sulfur bacterium from mangrove with rhodopin as predominant carotenoid. Acta Microbiologica Sinica. 2011;51(10):1318–1325. [PubMed] [Google Scholar]
- 3.Jiang P, Hong X, Zhao CG, Yang SP. Reciprocal transformation of Inorganic nitrogen by Resting Cells of Marichromatium gracile YL28. Journal of Huaqiao University. 2015;36(2):45–52. [Google Scholar]
- 4.Jiang P, Zhao CG, Jia YQ, Yang SP. Inorganic nitrogen removal by a marine purple sulfur bacterium capable of growth on nitrite as sole nitrogen source. Microbiology China. 2014;41(5):824–831. [Google Scholar]
- 5.Xuan H, Chen ZW, Zhao CG, Yang SP. Nitrogen transformation under different dissolved oxygen levels by the anoxygenic phototrophic bacterium Marichromatium gracile. World Journal of Microbiology and Biotechnology. 2017;33:113. doi: 10.1007/s11274-017-2280-z. [DOI] [PubMed] [Google Scholar]
- 6.Hungate BA, Dukes JS, Shaw MR, Luo Y, Field CB. Nitrogen and climate change. Science. 2003;302(5650):1512–1513. doi: 10.1126/science.1091390. [DOI] [PubMed] [Google Scholar]
- 7.Stein LY, Klotz MG. The nitrogen cycle. Current Biology. 2016;26(3):94–98. doi: 10.1016/j.cub.2015.12.021. [DOI] [PubMed] [Google Scholar]
- 8.Gruber N, Galloway JN. An earth-system perspective of the global nitrogen cycle. Nature. 2008;451(7176):293. doi: 10.1038/nature06592. [DOI] [PubMed] [Google Scholar]
- 9.Canfield DE, Glazer AN, Falkowski PG. The evolution and future of Earth’s nitrogen cycle. Science. 2010;330(6001):192–196. doi: 10.1126/science.1186120. [DOI] [PubMed] [Google Scholar]
- 10.Duce RA, et al. Impacts of atmospheric anthropogenic nitrogen on the open ocean. Science. 2008;320(5878):893–897. doi: 10.1126/science.1150369. [DOI] [PubMed] [Google Scholar]
- 11.Luo S, Wu BL, Xiong XQ, Wang JW. Short‐term toxicity of ammonia, nitrite, and nitrate to early life stages of the rare minnow (Gobiocypris rarus) Environmental Toxicology and Chemistry. 2015;35(6):1422–1427. doi: 10.1002/etc.3283. [DOI] [PubMed] [Google Scholar]
- 12.Dixon R, Kahn D. Genetic regulation of biological nitrogen fixation. Nature Reviews Microbiology. 1998;2(8):220–229. doi: 10.1038/nrmicro954. [DOI] [PubMed] [Google Scholar]
- 13.Dubbs JM, Tabita FR. Regulators of nonsulfur purple phototrophic bacteria and the interactive control of CO2 assimilation, nitrogen fixation, hydrogen metabolism and energy generation. FEMS Microbiology Reviews. 2004;28(3):353–376. doi: 10.1016/j.femsre.2004.01.002. [DOI] [PubMed] [Google Scholar]
- 14.Griffin BM, Schott J, Schink B. Nitrite, an electron donor for anoxygenic photosynthesis. Science. 2007;316(5833):1870–1870. doi: 10.1126/science.1139478. [DOI] [PubMed] [Google Scholar]
- 15.Hülsen T, et al. Domestic wastewater treatment with purple phototrophic bacteria using a novel continuous photo anaerobic membrane bioreactor. Water Research. 2016;100:486–495. doi: 10.1016/j.watres.2016.04.061. [DOI] [PubMed] [Google Scholar]
- 16.Imhoff, J. F. Taxonomy, phylogeny, and general ecology of anoxygenic phototrophic bacteria, 1. [Mann, Nicholas H. & Noel G. (eds)] Photosynthetic Prokaryotes2, 53–92. (Springer Science and Business Media, 2012).
- 17.Lee TI, et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science. 2002;298:799–804. doi: 10.1126/science.1075090. [DOI] [PubMed] [Google Scholar]
- 18.Pino C, et al. The assimilatory nitrate reduction system of the phototrophic bacterium Rhodobacter capsulatus E1F1. Biochemical Society Transactions. 2006;34(5594):127–129. doi: 10.1042/BST0340127. [DOI] [PubMed] [Google Scholar]
- 19.Olmo-Mira MF, et al. Expression and characterization of the assimilatory NADH-nitrite reductase from the phototrophic bacterium Rhodobacter capsulatus E1F1. Archives of Microbiology. 2006;186(4):339–344. doi: 10.1007/s00203-006-0149-x. [DOI] [PubMed] [Google Scholar]
- 20.Swingley, W. D., Blankenship R. E. & Raymond, J. Evolutionary relationships among purple photosynthetic bacteria and the origin of proteobacterial photosynthetic systems, 1. [Hunter, C. N., Daldal, F., Thurnauer, M. C. & Beatty, J. T. (eds)] The purple phototrophic bacteria2, 17–29. (Springer Netherlands, 2008).
- 21.Young, J. P. W. The phylogeny and evolution of nitrogenases, 1. [Palacios R. & Newton W. E. (eds)] Genomes and genomics of nitrogen-fixing organisms 14, 221–241. (Springer Netherlands, 2005).
- 22.Bishop PE, Joerger RD. Genetics and molecular biology of alternative nitrogen fixation systems. Annual review of plant biology. 1990;41(1):109–125. doi: 10.1146/annurev.pp.41.060190.000545. [DOI] [Google Scholar]
- 23.Yan Y, et al. Nitrogen fixation island and rhizosphere competence traits in the fenome of root-associated Pseudomonas stutzeri A1501. Proceedings of the National Academy of Sciences of the United States of America. 2008;105(21):7564. doi: 10.1073/pnas.0801093105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ramachandran A, Yun H, Wen-Mei WH, Ma J, Chen D. FPGA accelerated DNA error correction. Design, Automation and Test in Europe Conference and Exhibition. 2015;1:1371–1376. doi: 10.7873/DATE.2015.0983. [DOI] [Google Scholar]
- 25.Lin, X. L. et al. Mangrove sediment microbiome: adaptive microbial assemblages and their routed biogeochemical processes in Yunxiao mangrove national nature reserve, China. Microbial Ecology, 10.1007/s00248-018-1261-6 (2018). [DOI] [PubMed]
- 26.Kobayashi M, Fujii K, Shimamoto I, Maki T. Treatment and reuse of industrial waste water by phototrophic bacteria. Progress in Water Technology. 1978;11(1):279–284. [Google Scholar]
- 27.Vethanayagam RR. Purple photosynthetic bacteria from a tropical mangrove environment. Marine Biology. 1991;110(1):161–163. doi: 10.1007/BF01313103. [DOI] [Google Scholar]
- 28.Masepohl, B. & Hallenbeck, P. C. Nitrogen and molybdenum control of nitrogen fixation in the phototrophic bacterium Rhodobacter capsulatus, 1. [Hallenbeck, P. C. (ed.)] Recent advances in phototrophic prokaryotes4, 49–70. (Springer New York, 2010). [DOI] [PubMed]
- 29.Sun Y, et al. Ammonium assimilation: An important accessory during aerobic denitrification of Pseudomonas stutzeri T13. Bioresource Technology. 2017;234:264–272. doi: 10.1016/j.biortech.2017.03.053. [DOI] [PubMed] [Google Scholar]
- 30.Shen JY, et al. Improvement of fisheries water quality and fish growth of photosynthetic bacteria HZPSB. Bulletin of Science and Technology. 2004;15:16–31. [Google Scholar]
- 31.Qiu HD, Shi XA, Guo YH, Fen-Wu WU, Zhao JP. The breeding and application of salt-resistant and high NH4+ or NO2− reducible bacteria of Rhodospirillace. Journal of Fisheries of China. 1999;23:92–96. [Google Scholar]
- 32.Dahl, C. Sulfur metabolism in phototrophic bacteria, 1. [Hallenbeck, P. C. (ed.)] Modern Topics in the Phototrophic Prokaryotes2, 27–66. (Springer International, 2017).
- 33.Kappler U, Dahl C. Enzymology and molecular biology of prokaryotic sulfite oxidation. FEMS Microbiology Letters. 2001;203(1):1–9. doi: 10.1111/j.1574-6968.2001.tb10813.x. [DOI] [PubMed] [Google Scholar]
- 34.Frigaard NU, Dahl C. Sulfur metabolism in phototrophic sulfur bacteria. Advances in Microbial Physiology. 2008;54:103–200. doi: 10.1016/S0065-2911(08)00002-7. [DOI] [PubMed] [Google Scholar]
- 35.Dahl C. Insertional gene inactivation in a phototrophic sulphur bacterium: APS-reductase-deficient mutants of Chromatium vinosum. Microbiology. 1996;142(12):3363–3372. doi: 10.1099/13500872-142-12-3363. [DOI] [PubMed] [Google Scholar]
- 36.Jiang J, et al. Halomonas zhaodongensis sp. nov., a slightly halophilic bacterium isolated from saline-alkaline soils in Zhaodong, China. Antonie Van Leeuwenhoek. 2013;104(5):685–694. doi: 10.1007/s10482-013-9976-3. [DOI] [PubMed] [Google Scholar]
- 37.Meng L, et al. Cloning and identification of group 1 mrp operon encoding a novel monovalent cation/proton antiporter system from the moderate halophile Halomonas zhaodongensis. Extremophiles. 2014;18(6):963–972. doi: 10.1007/s00792-014-0666-5. [DOI] [PubMed] [Google Scholar]
- 38.Marrs B. Genetic recombination in Rhodopseudomonas capsulata. Proceedings of the National Academy of Sciences of the United States of America. 1974;71(3):971–973. doi: 10.1073/pnas.71.3.971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lang AS, Beatty JT. Importance of widespread gene transfer agent genes in alpha-proteobacteria. Trends in Microbiology. 2007;15(2):54–62. doi: 10.1016/j.tim.2006.12.001. [DOI] [PubMed] [Google Scholar]
- 40.Zheng Q, Jiao NZ, Zhang R, Wei JJ, Zhang F. The evolutionary divergence of psbA gene in synechococcus and their myoviruses in the east China Sea. Plos One. 2014;9(1):e86644. doi: 10.1371/journal.pone.0086644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Raymond, J. The role of horizontal gene transfer in photosynthesis, oxygen production, and oxygen tolerance, 1. [Gogarten, M. B., Gogarten, J. P. & Olendzenski, L. C. (eds)] Horizontal Gene Transfer19, 323–338. (Humana Press, 2009). [DOI] [PubMed]
- 42.Armitage, J. P. Swimming and behavior in purple non-sulfur bacteria, 1. [Hunter, C. N., Daldal, F., Thurnauer, M. C. & Beatty, J. T. (eds)] The purple phototrophic bacteria32, 643–654. (Springer Netherlands, 2009).
- 43.Silva ACRD, et al. Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature. 2002;417(6887):459–463. doi: 10.1038/417459a. [DOI] [PubMed] [Google Scholar]
- 44.Zhang XB, Zhao CG, Hong X, Chen SC, Yang SP. Genome sequence of Marichromatium gracile YL-28, a purple sulfur bacterium with bioremediation potential. Genome Announcements. 2016;4(3):e00288–00216. doi: 10.1128/genomeA.00288-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Aziz RK, et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9(1):75–75. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis Version 6.0. Molecular Biology and Evolution. 2013;30(12):2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Chaudhari NM, Kumar GV, Chitra D. BPGA an ultra-fast pan-genome analysis pipeline. Scientific reports. 2016;6:24373. doi: 10.1038/srep24373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Darling AE, Mau B, Perna NT. Progressivemauve: multiple genome alignment with gene gain, loss and rearrangement. Plos One. 2010;5(6):e11147. doi: 10.1371/journal.pone.0011147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Beatson SA, Ben ZNL, Petty NK, Nabil-Fareed A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics. 2011;12(1):1. doi: 10.1186/1471-2164-12-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dhillon BK, et al. IslandViewer 3: more flexible, interactive genomic island discovery, visualization and analysis. Nucleic Acids Research. 2015;43(1):104–108. doi: 10.1093/nar/gkv401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kranz RG, Haselkorn R. Anaerobic regulation of nitrogen-fixation genes in Rhodopseudomonas capsulata. Proceedings of the National Academy of Sciences of the United States of America. 1986;83(18):6805–6809. doi: 10.1073/pnas.83.18.6805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Cabello P, et al. Hydroxylamine assimilation by Rhodobacter capsulatus E1F1 requirement of the hcp gene (hybrid cluster protein) located in the nitrate assimilation nas gene region for hydroxylamine reduction. Journal of Biological Chemistry. 2004;279(44):45485–45494. doi: 10.1074/jbc.M404417200. [DOI] [PubMed] [Google Scholar]
- 53.Bartnikas TB, et al. Characterization of a member of the NnrR regulon in Rhodobacter sphaeroides 2.4.3 encoding a haem-copper protein. Microbiology. 2002;148(3):825. doi: 10.1099/00221287-148-3-825. [DOI] [PubMed] [Google Scholar]
- 54.Geer LY, Domrachev M, Lipman DJ, Bryant SH. CDART: Protein homology by domain architecture. Genome Research. 2002;12(10):1619–1623. doi: 10.1101/gr.278202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Marchlerbauer A, et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Research. 2017;45(1):200–203. doi: 10.1093/nar/gkw1129. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.