Abstract
The most well-understood circadian clock at the level of molecular mechanisms is that of cyanobacteria. This overview is on how solution-state nuclear magnetic resonance (NMR) spectroscopy has contributed to this understanding. By exciting atomic spin-½ nuclei in a strong magnetic field, NMR obtains information on their chemical environments, inter-nuclear distances, orientations, and motions. NMR protein samples are typically aqueous, often at near-physiological pH, ionic strength, and temperature. The level of information obtainable by NMR depends on the quality of the NMR sample, by which we mean the solubility and stability of proteins. Here, we use examples from our laboratory to illustrate the advantages and limitations of the technique.
Introduction
The metabolism, physiology, and behavior of most organisms display circadian rhythms that prepare organisms for the predictable, yet dramatic, swings in ambient light and temperature each day and night (Merrow et al. 2010; Hogenesch and Ueda 2011; Bass 2012; Lear and Allada 2012). The rhythms originate from oscillator components of intracellular circadian clocks (Bell-Pedersen et al. 2005; Brown et al. 2012). Since they are profoundly important for health and reproductive fitness, there has been much interest in the molecular mechanisms of the oscillators. The most progress on this front has been made on the oscillator of cyanobacteria (Ditty et al. 2009; Johnson et al. 2011). It is composed of three proteins, KaiA, KaiB, and KaiC, which together generate an autonomous circadian rhythm of KaiC (auto)phosphorylation and (auto)dephosphorylation (Nakajima et al. 2005). During the subjective day, KaiC phosphorylates through an autokinase activity when it is stimulated through direct interactions with KaiA (Iwasaki et al. 2002; Williams et al. 2002). When KaiB prevents stimulation by KaiA during the subjective night, KaiC undergoes dephosphorylation (Kitayama et al. 2003; Egli et al. 2012; Nishiwaki and Kondo 2012). KaiA–KaiC interactions are necessary for the phosphorylation phase, whereas sequestration of KaiA in a KaiABC complex is essential for the dephosphorylation phase (Hayashi et al. 2004; Kageyama et al. 2006; Mori et al. 2007; Nishiwaki et al. 2007; Rust et al. 2007). The goal of our laboratory is to develop a comprehensive understanding of the mechanism of this three-protein oscillator at the molecular level. Here, we describe the role nuclear magnetic resonance (NMR) played in this research.
This review picks up where an earlier one left off (Vakonakis and LiWang 2009). In that review, we discussed how, before the structures of the full-length clock proteins were solved, NMR provided significant insights into the structural biology of clock proteins using a “bottom-up” approach, that is, solving the structures of proteins one domain at a time. In 2004, the X-ray crystal structures of KaiA (Ye et al. 2004), KaiB (Garces et al. 2004), and KaiC (Pattanayek et al. 2004) all became available, providing a static snapshot of each gear in the cyanobacterial clockwork. Once all three structures were available, the race was on to determine the mechanism whereby their assemblages reorganized according to time of day. During this period, we discovered that relative to the clock proteins from the mesophile, Synechococcus elongatus, those from the thermophile, Thermosynechococcus elongatus, were superior for NMR (Vakonakis et al. 2004). Thus, the NMR studies described in this perspective were all performed on proteins from T. elongatus.
KaiC has two RecA-like domains connected head-to-tail (Leipe et al. 2000). The N-terminal and C-terminal domains are called CI and CII, respectively. The CI and CII domains self-associate into hexameric rings, so that KaiC has an appearance like two donuts stacked together (Hayashi et al. 2003, 2006). In the presence of KaiA and KaiB, two residues in each CII domain, S431 and T432, execute a specific phosphorylation pattern: ST → SpT → pSpT → pST → ST →…, where pS and pT refer to the phosphorylated forms of S431 and T432, respectively (Nishiwaki et al. 2007; Rust et al. 2007). These phosphoforms are functionally distinct. For example, KaiB binds to KaiC only when S431 is phosphorylated (Nishiwaki et al. 2007) and KaiA has a greater affinity for hypophosphorylated KaiC than for hyperphosphorylated KaiC (Hayashi et al. 2004; Ma and Ranganathan 2012). It was long presumed that the temporal assembly and disassembly of KaiA–KaiC, KaiB–KaiC, and KaiA–KaiB–KaiC complexes were dictated by phosphorylation-driven conformational changes in KaiC (Mori et al. 2007; Nishiwaki et al. 2007; Rust et al. 2007; Akiyama et al. 2008; Qin et al. 2010; Murayama et al. 2011; Goda et al. 2012; Phong et al. 2013). Thus, it came as a surprise that the X-ray crystal structures of different phosphoforms were found to have virtually identical structures (Pattanayek et al. 2009). It became clear that the mechanism of this oscillator, that is, how it transitions from one state to another, was not due to KaiC clicking from one rigid phosphoform to the next. At this point, we sought to elucidate what was changing in KaiC which afforded its phosphoforms different functions. To that end, we chose NMR as the primary tool, as it can inform on the structure and dynamics of proteins in solution at near-physiological conditions (Clore and Potts 2012; Barrett et al. 2013).
A drawback of conventional NMR is that proteins larger than ∼50 kDa produce weak signals, and KaiC weighs in at 345 kDa. Thus, KaiC would be invisible by conventional NMR. However, a major advance in isotopic labeling and NMR methodology made the study of supramolecular proteins like KaiC feasible (Goto et al. 1999; Isaacson et al. 2007; Ayala et al. 2009; Kay 2011; Larda et al. 2012). It became possible to produce proteins with 1H and 13C incorporated at the methyl groups of the side chains of alanine, methionine, isoleucine, leucine, valine, and methylated lysine, with 2H and 12C everywhere else. Combining this labeling technique with an NMR experiment specifically optimized for large proteins, called methyl-transverse relaxation-optimized spectroscopy (methyl TROSY), allowed the collection of high-quality NMR spectra on proteins of several hundred kilodaltons (Gelis et al. 2007; Sprangers and Kay 2007; Kay 2011; Tzeng et al. 2012). However, perdeuterated protein samples 13C, 1H-labeled specifically on the methyl groups of amino-acyl side chains are costly. An Ile-δ1-[13C, 1H]-labeled sample from a 200-ml culture typically costs about $300, regardless of yield. The growth medium is D2O based, which lowers the yield relative to H2O-based media. Thus, it was especially important for us to optimize the solubility and stability of KaiC before labeling it with isotopes. NMR is a relatively insensitive technique, requiring samples to be at least hundreds of micromolar. For proteins with limited solubility (∼100 µM), it is typically necessary to average the signal several hours in order to record a two-dimensional spectrum. Even with the development of cryogenic NMR probes, which boosts signal-to-noise by up to ∼four-fold, KaiC was not sufficiently soluble for routine NMR studies. Thus, it was especially important to optimize the solubility of KaiC before isotopic labeling. It should be noted that preparation of NMR samples is often, if not always, the most challenging step for NMR studies. A serious problem we faced was the poor solubility and stability of KaiC. Wild-type KaiC was not soluble above ∼10 µM (monomer concentration), and it precipitated within a few days at that concentration. We endeavored to design kaiC constructs in overexpression systems of Escherichia coli that produced stable and soluble proteins at high yield. We first tried fusing glutathione S-transferase (GST) to KaiC (Harper and Speicher 2011). However, KaiC aggregated badly after cleavage of the GST tag. The final yield was lower than 5 µM in 400 uL from a 200-ml culture. We then replaced the GST with thioredoxin (Malhotra 2009), but encountered the same problem of aggregation of KaiC after cleavage. We reasoned that it was necessary to maintain a solubility tag on KaiC to enhance solubility. However, both GST (220 residues) and thioredoxin (109 residues) would add too many signals to NMR spectra of KaiC.
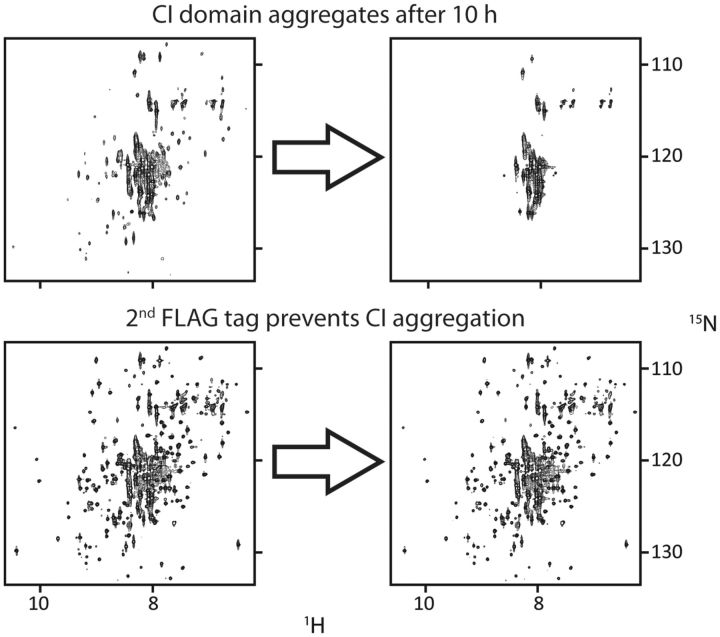
While still a graduate student, the first author of this review noticed that the FLAG tag “DYKDDDDK” (Einhauer and Jungbauer 2001) he was using for detection by antibody dramatically increased the solubility of the ubiquitin-like protein ISG15 (Chang et al. 2008). Its small size is advantageous, because it could be retained on proteins without significantly increasing the complexity of NMR spectra. Indeed, we found that it had a dramatic effect on the solubility of KaiC. Addition of a single FLAG tag to the N-terminus was enough to dramatically increase the solubility of KaiC by 10-fold (from 5 to 50 µM). Furthermore, the stability of KaiC increased from just over a day to a week. However, even with an N-terminal FLAG tag, monomeric CI domain of KaiC, CI*, aggregated within 10 h (Fig. 1). By contrast, adding a second FLAG tag to the C-terminus of CI* yielded a significantly more stable sample, CI*FLAG. Other clock proteins also benefitted from the FLAG tag. With a FLAG tag at each terminus, the solubility of the N-terminal domain of the clock-output protein, SasA, increased from 50 to 800 µM. A dimeric mutant of KaiB without FLAG tags was not soluble above 100 µM at 50 mM salt. However, with a single FLAG tag at the N-terminus, the tagged variant KaiB* was soluble up to 1 mM under low salt. Thus, the FLAG tag was critical for the large contribution by NMR to our understanding of the circadian oscillator of cyanobacteria. (Notice that the superscript FLAG is only used when a protein has two FLAG tags, as for CI*FLAG but not for KaiB*, which has a single tag.)
Fig. 1.
FLAG tags increase stability of the KaiC CI domain. Top panels show 15N, 1H-TROSY spectra of a monomeric variant of the CI domain with a single FLAG tag at the N-terminus before (left panel) and after (right panel) 10-h incubation at 30°C. Bottom panels show spectra of a CI domain having two FLAG tags, one at each terminus. Although a single tag stabilizes CI to a certain extent, two tags had a dramatic effect on long-term7 stability.
Once we were able to reliably produce relatively stable and soluble samples of KaiC, we wanted to see whether NMR could detect something missed by X-ray crystallography. However, the phosphorylated states of KaiC cannot be isolated, and they dephosphorylate within several hours anyway, which means they are impossible to study by NMR. Fortunately, others found that substituting residues S431 and T432 with alanyl/glutamyl residues mimicked their unphosphorylated/phosphorylated states reasonably well (Nishiwaki et al. 2007; Terauchi et al. 2007; Dong et al. 2010). Thus, instead of the unstable SpT-KaiC, pSpT-KaiC, and pST-KaiC phosphoforms, for NMR studies we, respectively, used SE-KaiC, EE-KaiC, and ET-KaiC mutants as their mimics (for the ST-KaiC phosphoform, we simply used dephosphorylated wild-type KaiC). In order to extract the maximum information from methyl-TROSY spectra, though, it is necessary to assign each methyl correlation to a specific amino acyl residue of KaiC.
There are two common ways to assign the methyl groups of supramolecular proteins (>100 kDa), known as the divide-and-conquer and assignment-by-mutagenesis approaches (Plevin and Boisbouvier 2012). The divide-and-conquer approach can be highly successful for multidomain and/or multimeric proteins. Through mutagenesis, the individual domains are isolated and modified to prevent self-association. If this step is successful, chemical shifts of the domains can be assigned using conventional NMR experiments. These assignments are then mapped onto the spectrum of the full-length protein (Gelis et al. 2007; Sprangers and Kay 2007). In the assignment-by-mutagenesis approach, methyl-bearing residues of the protein are replaced through site-directed mutagenesis. Comparisons of methyl TROSY spectra of the wild-type protein against each mutant can lead to the assignments of most chemical shifts (Sprangers et al. 2005; Ren et al. 2010; Yang et al. 2010; Amero et al. 2011). However, with 45, 41, and 31 isoleucinyl, leucinyl, and valinyl residues, respectively, it was prohibitively costly with our limited budget to assign these methyl groups of KaiC. Thus, the residue-specific assignment of methyl groups of KaiC has not been made. However, by following the divide-and-conquer method, we made a few domain-specific assignments that yielded important, albeit qualitative, insights into the function of KaiC, as will be described below.
Following published protocols (Goto et al. 1999; Sprangers and Kay 2007), we produced samples of U-[15N, 2H]-Ile-δ1-[13C, 1H]-labeled KaiC and its phosphomimics. Methyl-TROSY spectra were carried out on a 600-MHz (proton frequency) Bruker Avance III equipped with a cryogenic probe and pulsed-field gradients. Sample concentrations were typically 15 µM, and each two-dimensional spectrum was acquired in ∼20 h. The concentrations of the NMR samples were kept low because of cost. Comparisons of the spectra side-by-side quickly revealed some striking differences (Chang et al. 2011). The line shapes of several methyl correlations of ST-KaiC and SE-KaiC were broad and weak (Fig. 2A–C). In contrast, the NMR peaks of EE-KaiC and ET-KaiC were generally sharper and stronger. This comparison suggested that ST-KaiC and SpT-KaiC had significant breathing motions on the micro-to-millisecond time scale, whereas those motions were largely absent in pSpT-KaiC and pST-KaiC. A comparison of the methyl-TROSY spectra of KaiC against those of its individual domains allowed us to assign the breathing motions to the CII ring.
Fig. 2.
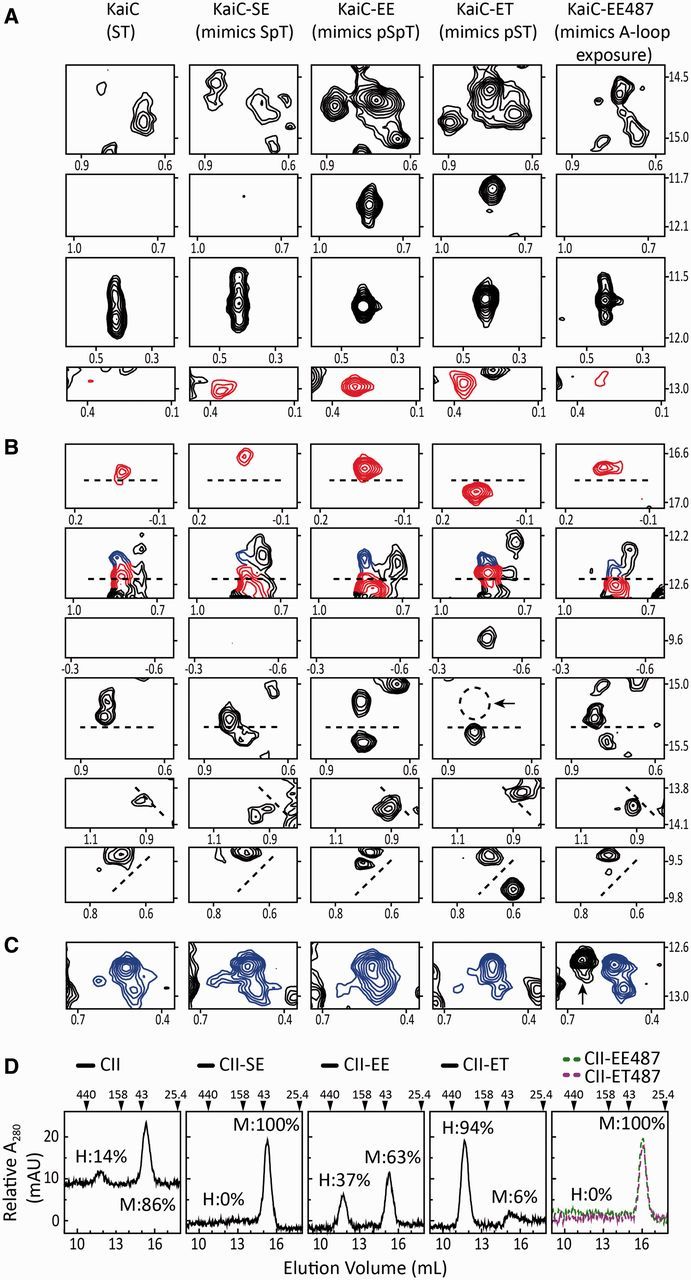
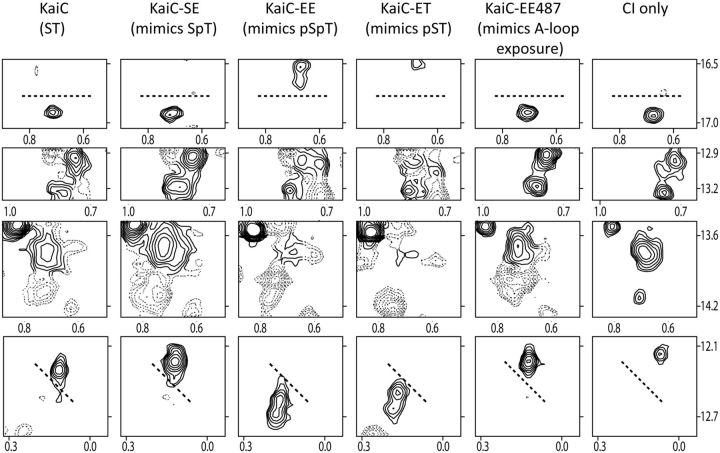
Flexibility of the CII ring depends on the state of phosphorylation at residues S431 and T432 and A-loop position. (A–C) Selected regions from methyl-TROSY spectra of U-[15N, 2H]-Ile-δ1-[13C, 1H]-labeled KaiC (panel 1) and phosphomimics of SpT-KaiC (panel 2), pSpT-KaiC (panel 3), pST-KaiC (panel 4), and pSpT-KaiC487 (panel 5). Horizontal and vertical axes are 1H and 13C chemical shifts in ppm, respectively. Red and blue contours indicate peaks assigned to Ile residues of the CII and CI domains of KaiC, respectively, whereas black contours indicate unassigned peaks. It should be noted that SE-KaiC is ∼30% phosphorylated at residue S431. (D) Gel-filtration profiles of isolated domains of CII (panel 1) and phosphomimics of SpT-CII (panel 2), pSpT-CII (panel 3), pST-CII (panel 4), and pSpT-CII487 and pST-CII487 (panel 5). The hexameric forms of the CI and CII domains eluted at ∼11.7 ml, whereas the monomeric forms eluted at ∼15.4 ml. In panel 5, EE-CII487 and ET-CII487, two KaiC CII domains that were truncated just prior to the A loop at residue 488, eluted as monomers at 16 ml. The percentages of hexameric (H) and monomeric forms (M) are provided in each panel. Adapted from Chang et al. (2011).
The notion of phosphorylation-dependent CII-ring motions was corroborated by gel-filtration chromatography experiments. As show in Fig. 2D, the tendency of isolated CII domains to hexamerize depended on the state of residues at positions 431 and 432. The S431E substitution mimicking phosphoryl-Ser431 greatly enhanced the extent of hexamerization, whereas the T432E substitution somewhat negated the effect of S431E. These data, like the NMR experiments, suggested that the ST and SpT states have loose CII rings, whereas the pSpT state has a tighter CII ring. The tightest CII ring, though, was found in the pST phosphomimic. This raised the question as to whether there is any functional relevance to CII ring dynamics?
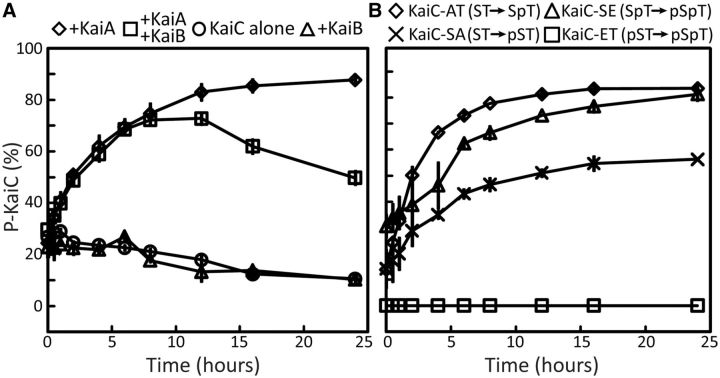
Insight into the functional relevance of CII ring dynamics was gained by monitoring KaiA-stimulated phosphorylation of KaiC variants. Phosphorylation of AT-KaiC, SE-KaiC, SA-KaiC, and ET-KaiC, respectively, mimics the ST → SpT, SpT → pSpT, ST → pST, and pST → pSpT phosphorylation steps. The data on phosphorylation kinetics showed that phosphoryl-S431 inhibited KaiC phosphorylation, whereas phosphoryl-T432 offset the inhibition somewhat. Thus, it appeared that the tightness of the CII ring inhibited the extent to which KaiA could induce KaiC to phosphorylate (Fig. 3). As a result, we were able to conclude that the natural clockwise direction of phosphorylation, ST (loose CII ring) → SpT (loose CII ring) → pSpT (semi-tight CII ring), is promoted because the CII ring is sufficiently loose. In contrast, the counterclockwise direction, ST (loose CII ring) → pST (tight CII ring) → pSpT (semi-tight CII ring), is prohibited because the CII ring becomes prohibitively tight on the first step.
Fig. 3.
Profiles of phosphorylation kinetics of KaiC and KaiC phosphomimics. Each data point represents the average of two experiments. The error bars represent the standard error (SEM). (A) Phosphorylation of KaiC in the presence and/or absence of KaiA and KaiB. KaiC alone (circle); KaiC + KaiA (diamond); KaiC + KaiB (triangle); KaiC + KaiA + KaiB (square). (B) Phosphorylation of KaiC variants AT-KaiC (diamond), SE-KaiC (triangle), SA-KaiC (cross), and ET-KaiC (square) in the presence of KaiA. These profiles were obtained by densitometric analysis of SDS–PAGE gels. Adapted from Chang et al. (2011).
How CII ring motions regulate KaiA stimulation of KaiC can be understood by examining the KaiA-binding site, which is on the CII ring. KaiA binds to the C-terminal segments of KaiC, residues 487–518, which include the so-called A loops (residues 487–497) (Vakonakis and LiWang 2004). The A loops switch KaiC between phosphorylation and dephosphorylation, depending upon their conformation (Kim et al. 2008). In the crystal structures of KaiC, the A loops are buried and inaccessible to KaiA. Thus, they have to be in exchange with an exposed state that can be selectively captured by KaiA. We found that when the A loops are buried, KaiC dephosphorylates. When they are exposed, KaiC phosphorylates. In the buried position, the A loops make extensive intramolecular interactions with the CII domains, as well as with each other, forming a ring of hydrogen bonds that stabilize the CII ring. Thus, a tight CII ring would be expected to have tightly buried A loops, giving little access to KaiA. On the other hand, a loose and breathing CII ring should have A loops that make excursions to the exposed state that can be captured and held there by KaiA. According to this idea, the observed clockwise direction of phosphorylation is that which allows transient exposure of the KaiA-binding site. In contrast, the counterclockwise direction is prohibited in part because it hides the KaiA-binding site. This idea explains earlier suggestions that the KaiA–KaiC interaction weakens when KaiC phosphorylates (Hayashi et al. 2004; Ma and Ranganathan 2012).
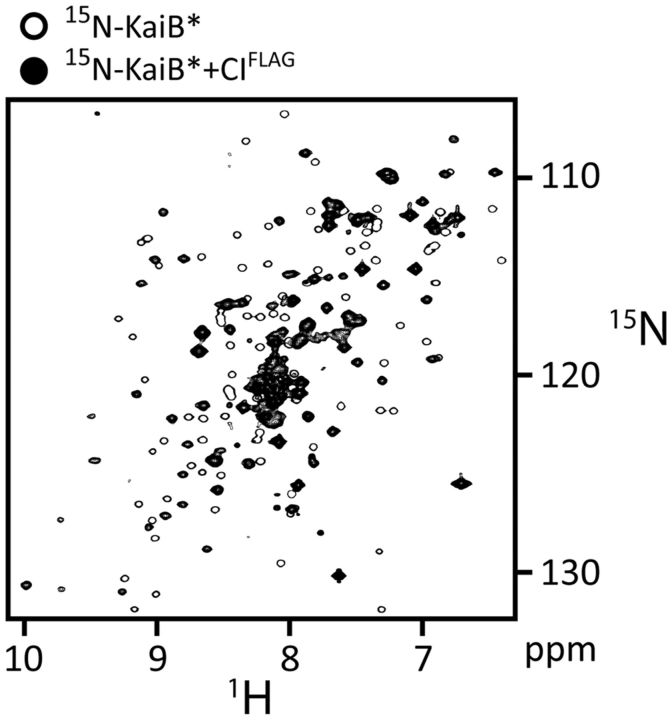
Once the S431 residues of KaiC are phosphorylated, KaiB inhibits KaiA by sequestering it in a KaiA–KaiB–KaiC complex (Nishiwaki et al. 2007; Rust et al. 2007). This action by KaiB is necessary for the dephosphorylation phase of the oscillator. An interesting question has been the mechanism whereby KaiB recognizes KaiC. Free KaiB exists in a tetrameric form (Iwase et al. 2005; Pattanayek et al. 2008; Mutoh et al. 2011; Murakami et al. 2012). It was thought that KaiB binds to the CII ring as a dimer (Akiyama et al. 2008; Pattanayek et al. 2008). The electron microscopy data upon which the KaiB-CII model was based were low so that distinguishing CI from CII was difficult. (A follow-up study on KaiB-CII binding, also by EM, was similarly low in resolution [Pattanayek et al. 2013].) We thought that the data were ambiguous, and wanted, therefore, to reinvestigate the KaiB-binding site on KaiC. In order to circumvent the difficulty of distinguishing the CI and CII domains in full-length KaiC, we tested binding of a dimeric KaiB mutant, KaiB*, to isolated CI and CII domains. NMR showed that KaiB* did not bind to isolated CII domains, but to a monomeric variant of CI (Fig. 4). Virtually every peak in the NMR spectrum of KaiB shifted substantially in the presence of CI, indicating that it changed conformation on a global scale. In this type of experiment, called a 15N-1H heteronuclear single quantum correlation (HSQC) experiment, every 15N-1H bonded pair yields a peak in the spectrum at a position that is dependent on its local environment. Thus, except for prolyl residues, there should be one peak for every amino acyl residue in the protein, approximately. KaiB has 87 non-prolyl residues. The HSQC spectrum of KaiB had 85 backbone correlations (i.e., not arising from side chain 15N-1H2 groups on Asn and Gln). Upon adding unlabeled monomeric CI, approximately 80 correlations were observed. If KaiB bound as a dimer, the symmetry would break upon binding to produce a doubling of peaks (i.e., 160–170 peaks). Since this was not observed, the NMR data indicated that KaiB bound to CI as a monomer. Thus, we concluded that the normally tetrameric KaiB has to disassociate all the way to a monomer, and then make an excursion to a highly different conformation in order to bind the CI domain of KaiC. We think that these thermodynamic barriers set the slow pace of KaiB–KaiC binding (Qin et al. 2010; Chang et al. 2012), and thereby play a major role in the mechanism of the cyanobacterial oscillator.
Fig. 4.
KaiB binds to the CI domain of KaiC. 15N, 1H-HSQC spectra of 15N-labeled KaiB* free (single-contour peaks) and in the presence of CIFLAG (multiple-contour peaks). Adapted from Chang et al. (2012).
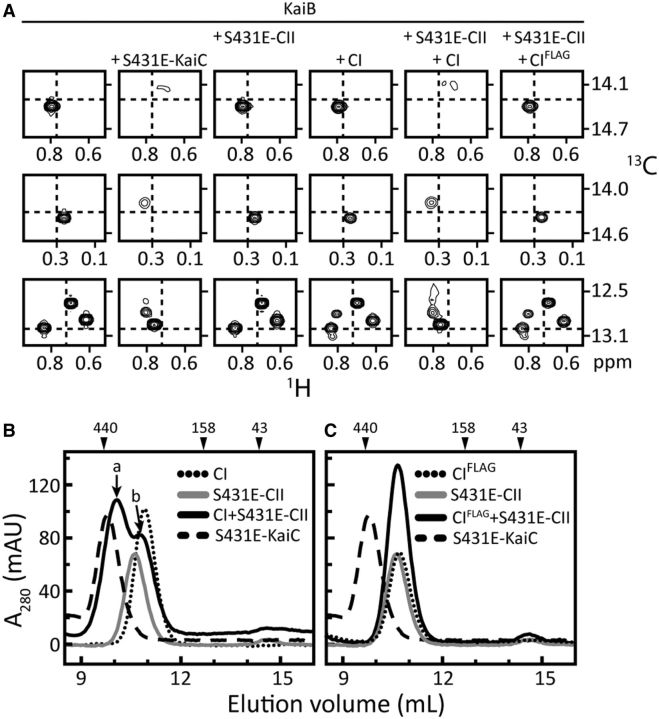
Since KaiB binds to the CI domain of KaiC, but only upon phosphorylation events on the CII side, it became obvious the two rings must communicate. We therefore sought to better understand how CI–CII communication regulated KaiB binding. We used ET-KaiC, that is, S431E-KaiC, and S431E-CII, which have the tightest CII rings, to probe KaiB binding. Figure 5 shows that CI–CII ring-stacking interactions were essential for KaiB binding. Disruption of stacking abolished KaiB binding. As KaiB did not bind to isolated intact CI rings but to monomeric CI subunits, we inferred that ring stacking partially opens/twists the CI ring to expose an important part of the KaiB-binding site. We also compared methyl-TROSY spectra of the isolated CI ring and full-length KaiC at different phosphorylation states. Interestingly, for ST-KaiC and SE-KaiC, CI peaks were in the same positions as in the isolated CI ring (Fig. 6). However, these peaks shifted significantly in EE-KaiC and ET-KaiC. These observations suggested that the CI and CII rings do not stack when the S431 residues are not phosphorylated. Upon S431 phosphorylation, however, the two rings stack together. If the ATPase of CI is activated upon ring stacking, it may provide energy to expose the KaiB-binding site. Indeed, when the ATPase of CI is inhibited, KaiB–KaiC binding is abolished (Phong et al. 2013). The clock output protein SasA (Iwasaki et al. 2000; Takai et al. 2006; Dong et al. 2010) also binds to the CI side of KaiC (Chang et al. 2011) and competes with KaiB for binding (Pattanayek et al. 2011; Murakami et al. 2012). Thus, ring stacking probably also regulates the clock’s output.
Fig. 5.
Stacking of CI and CII rings of KaiC is required for binding of KaiC–KaiB. (A) Selected regions from methyl-TROSY spectra of U-[15N, 2H]-Ile-δ1-[13C, 1H]-labeled KaiB alone (panels in column 1) or in the presence of S431E-KaiC (panels in column 2), S431E-CII (panels in column 3), CI (panels in column 4), S431E-CII + CI (panels in column 5), or S431E-CII + CIFLAG (panels in column 6). (B and C) Gel-filtration profiles of mixtures of CI (B) or CIFLAG (C) with S431E-CII. Full-length S431E-KaiC was run as a control for the expected elution position of stacked rings. Compared with CI, CIFLAG has an extra FLAG at its C-terminus, which was engineered for introducing potential repulsive interactions with the N-terminal FLAG tag on isolated S431E-CII domains. Note that the FLAG tag is highly negatively charged “DYKDDDDK”. Adapted from Chang et al. (2012).
Fig. 6.
CII–CI interactions depend on the state of phosphorylation at residues S431 and T432. Selected regions of methyl-TROSY spectra of U-[15N, 2H]-Ile-δ1-[13C, 1H]-labeled KaiC (panel 1) and the phosphomimics of SpT-KaiC (panel 2), pSpT-KaiC (panel 3), pST-KaiC (panel 4), and pSpT-KaiC-487 (panel 5). Horizontal and vertical axes are 1H and 13C chemical shifts in ppm, respectively. All panels were plotted at the same relative contour level. Solid contours are resonances assigned to the CI domain, whereas dashed contours are those unassigned. Selected regions of methyl-TROSY spectra of free CI are shown in panel 6. Adapted from Chang et al. (2011).
We hope that we have provided helpful examples of how NMR, even with only qualitative analysis, can yield important insights into the mechanisms of protein machinery. The big questions to answer next, regarding the cyanobacterial clock, include how CII ring motions affects KaiC kinase and phosphatase activities; how tightening of the CII ring promotes ring stacking; and how ring stacking regulates the CI ATPase, promotes binding of KaiB, and regulates clock output. Structures of the KaiA–KaiB, KaiB–KaiC, and KaiABC complexes also will need to be determined in order to achieve a comprehensive understanding. To address these problems, we will need to carry out quantitative studies of the system, which will require residue-specific assignments of the chemical shifts. We will start by assigning strategically located methyl-bearing residues by mutagenesis and use them as probes of their local environments. Thus, it is our expectation that NMR will continue to play a major role in advancing knowledge about this circadian clock, even given the high-molecular weights of the clock’s protein complexes.
Funding
This work was supported by a grant from the US Army Research Office (W911NF-10-1-0090). R.T. was supported by an NSF Graduate Research Fellowship.
References
- Akiyama S, Nohara A, Ito K, Maeda Y. Assembly and disassembly dynamics of the cyanobacterial periodosome. Mol Cell. 2008;29:703–16. doi: 10.1016/j.molcel.2008.01.015. [DOI] [PubMed] [Google Scholar]
- Amero C, Asuncion Dura M, Noirclerc-Savoye M, Perollier A, Gallet B, Plevin M, Vernet T, Franzetti B, Boisbouvier J. A systematic mutagenesis-driven strategy for site-resolved NMR studies of supramolecular assemblies. J Biomol NMR. 2011;50:229–36. doi: 10.1007/s10858-011-9513-5. [DOI] [PubMed] [Google Scholar]
- Ayala I, Sounier R, Use N, Gans P, Boisbouvier J. An efficient protocol for the complete incorporation of methyl-protonated alanine in perdeuterated protein. J Biomol NMR. 2009;43:111–9. doi: 10.1007/s10858-008-9294-7. [DOI] [PubMed] [Google Scholar]
- Barrett PJ, Chen J, Cho M, Kim J, Lu Z, Mathew S, Peng D, Song Y, Van Horn WD, Zhuang T, et al. The quiet renaissance of protein nuclear magnetic resonance. Biochemistry. 2013;52:1303–20. doi: 10.1021/bi4000436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bass J. Circadian topology of metabolism. Nature. 2012;491:348–56. doi: 10.1038/nature11704. [DOI] [PubMed] [Google Scholar]
- Bell-Pedersen D, Cassone VM, Earnest DJ, Golden SS, Hardin PE, Thomas TL, Zoran MJ. Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat Rev Genet. 2005;6:544–56. doi: 10.1038/nrg1633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown S, Kowalska E, Dallmann R. (Re)inventing the circadian feedback loop. Dev Cell. 2012;22:477–87. doi: 10.1016/j.devcel.2012.02.007. [DOI] [PubMed] [Google Scholar]
- Chang Y, Kuo N, Tseng R, LiWang A. Flexibility of the C-terminal, or CII, ring of KaiC governs the rhythm of the circadian clock of cyanobacteria. Proc Natl Acad Sci USA. 2011;108:14431–6. doi: 10.1073/pnas.1104221108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang Y, Tseng R, Kuo N, LiWang A. Rhythmic ring-ring stacking drives the circadian oscillator clockwise. Proc Natl Acad Sci USA. 2012;109:16847–51. doi: 10.1073/pnas.1211508109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang Y, Yan X, Xie Y, Gao X, Song A, Zhang D, Hu H. Different roles for two ubiquitin-like domains of ISG15 in protein modification. J Biol Chem. 2008;283:13370–7. doi: 10.1074/jbc.M800162200. [DOI] [PubMed] [Google Scholar]
- Clore GM, Potts J. Cambridge: Royal Society of Chemistry Publishing; 2012. Recent developments in biomolecular NMR, RSC biomolecular sciences series. (doi:10.1039/9781849735391) [Google Scholar]
- Ditty JL, Mackey SR, Johnson CH. Bacterial circadian programs. New York: Springer-Verlag; 2009. [Google Scholar]
- Dong G, Yang Q, Wang Q, Kim Y, Wood TL, Osteryoung KW, van Oudenaarden A, Golden SS. Elevated ATPase activity of KaiC applies a circadian checkpoint on cell division in Synechococcus elongatus. Cell. 2010;140:529–39. doi: 10.1016/j.cell.2009.12.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Egli M, Mori T, Pattanayek R, Xu Y, Qin X, Johnson CH. Dephosphorylation of the core clock protein KaiC in the cyanobacterial KaiABC circadian oscillator proceeds via an ATP synthase mechanism. Biochemistry. 2012;51:1547–58. doi: 10.1021/bi201525n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Einhauer A, Jungbauer A. The FLAGTM peptide, a versatile fusion tag for the purification of recombinant proteins. J Biochem Bioph Meth. 2001;49:455–65. doi: 10.1016/s0165-022x(01)00213-5. [DOI] [PubMed] [Google Scholar]
- Garces RG, Wu N, Gillon W, Pai EF. Anabaena circadian clock proteins KaiA and KaiB reveal a potential common binding site to their partner KaiC. EMBO J. 2004;23:1688–98. doi: 10.1038/sj.emboj.7600190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gelis I, Bonvin AMJJ, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell. 2007;131:756–69. doi: 10.1016/j.cell.2007.09.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goda K, Ito H, Kondo T, Oyama T. Fluorescence correlation spectroscopy to monitor kai protein-based circadian oscillations in real time. J Biol Chem. 2012;287:3241–8. doi: 10.1074/jbc.M111.265777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goto NK, Gardner KH, Mueller GA, Willis RC, Kay LE. A robust and cost-effective method for the production of Val, Leu, Ile (δ1) methyl protonated 15N-, 13C-, 2H-labeled proteins. J Biomol NMR. 1999;13:369–74. doi: 10.1023/a:1008393201236. [DOI] [PubMed] [Google Scholar]
- Harper S, Speicher D. Purification of proteins fused to glutathione S-transferase. Methods Mol Biol. 2011;681:259–80. doi: 10.1007/978-1-60761-913-0_14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi F, Ito H, Fujita M, Iwase R, Uzumaki T, Ishiura M. Stoichiometric interactions between cyanobacterial clock proteins KaiA and KaiC. Biochem Biophys Res Commun. 2004;316:195–202. doi: 10.1016/j.bbrc.2004.02.034. [DOI] [PubMed] [Google Scholar]
- Hayashi F, Iwase R, Uzumaki T, Ishiura M. Hexamerization by the N-terminal domain and intersubunit phosphorylation by the C-terminal domain of cyanobacterial circadian clock protein KaiC. Biochem Biophys Res Commun. 2006;348:864–72. doi: 10.1016/j.bbrc.2006.07.143. [DOI] [PubMed] [Google Scholar]
- Hayashi F, Suzuki H, Iwase R, Uzumaki T, Miyake A, Shen J, Imada K, Furukawa Y, Yonekura K, Namba K, et al. ATP-induced hexameric ring structure of the cyanobacterial circadian clock protein KaiC. Genes Cells. 2003;8:287–96. doi: 10.1046/j.1365-2443.2003.00633.x. [DOI] [PubMed] [Google Scholar]
- Hogenesch JB, Ueda HR. Understanding systems-level properties: timely stories from the study of clocks. Nat Rev Genet. 2011;12:407–16. doi: 10.1038/nrg2972. [DOI] [PubMed] [Google Scholar]
- Isaacson RL, Simpson PJ, Liu M, Cota E, Zhang X, Freemont P, Matthews S. A new labeling method for methyl transverse relaxation-optimized spectroscopy NMR spectra of alanine residues. J Am Chem Soc. 2007;129:15428–9. doi: 10.1021/ja0761784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki H, Nishiwaki T, Kitayama Y, Nakajima M, Kondo T. KaiA-stimulated KaiC phosphorylation in circadian timing loops in cyanobacteria. Proc Natl Acad Sci USA. 2002;99:15788–93. doi: 10.1073/pnas.222467299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki H, Williams SB, Kitayama Y, Ishiura M, Golden SS, Kondo T. A KaiC-interacting sensory histidine kinase, SasA, necessary to sustain robust circadian oscillation in cyanobacteria. Cell. 2000;101:223–33. doi: 10.1016/S0092-8674(00)80832-6. [DOI] [PubMed] [Google Scholar]
- Iwase R, Imada K, Hayashi F, Uzumaki T, Morishita M, Onai K, Furukawa Y, Namba K, Ishiura M. Functionally important substructures of circadian clock protein KaiB in a unique tetramer complex. J Biol Chem. 2005;280:43141–9. doi: 10.1074/jbc.M503360200. [DOI] [PubMed] [Google Scholar]
- Johnson CH, Stewart PL, Egli M. The cyanobacterial circadian system: from biophysics to bioevolution. Annu Rev Biophys. 2011;40:143–67. doi: 10.1146/annurev-biophys-042910-155317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kageyama H, Nishiwaki T, Nakajima M, Iwasaki H, Oyama T, Kondo T. Cyanobacterial circadian pacemaker: Kai protein complex dynamics in the KaiC phosphorylation cycle in vitro. Mol Cell. 2006;23:161–71. doi: 10.1016/j.molcel.2006.05.039. [DOI] [PubMed] [Google Scholar]
- Kay LE. Solution NMR spectroscopy of supra-molecular systems, why bother? A methyl-TROSY view. J Magn Reson. 2011;210:159–70. doi: 10.1016/j.jmr.2011.03.008. [DOI] [PubMed] [Google Scholar]
- Kim Y, Dong G, Carruthers CW, Golden SS, LiWang A. The day/night switch in KaiC, a central oscillator component of the circadian clock of cyanobacteria. Proc Natl Acad Sci USA. 2008;105:12825–30. doi: 10.1073/pnas.0800526105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitayama Y, Iwasaki H, Nishiwaki T, Kondo T. KaiB functions as an attenuator of KaiC phosphorylation in the cyanobacterial circadian clock system. EMBO J. 2003;22:2127–34. doi: 10.1093/emboj/cdg212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larda S, Bokoch M, Evanics F, Prosser RS. Lysine methylation strategies for characterizing protein conformations by NMR. J Biomol NMR. 2012;54:199–209. doi: 10.1007/s10858-012-9664-z. [DOI] [PubMed] [Google Scholar]
- Lear BC, Allada R. In: eLS. Chichester: John Wiley & Sons, Ltd; 2012. Circadian rhythms. (doi: 10.1002/9780470015902.a0000040.pub2) [Google Scholar]
- Leipe DD, Aravind L, Grishin NV, Koonin EV. The bacterial replicative helicase DnaB evolved from a RecA duplication. Genome Res. 2000;10:5–16. [PubMed] [Google Scholar]
- Ma L, Ranganathan R. Quantifying the rhythm of KaiB-C interaction for in vitro cyanobacterial circadian clock. PLoS One. 2012;7:e42581. doi: 10.1371/journal.pone.0042581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malhotra A. Tagging for protein expression. Methods Enzymol. 2009;463:239–58. doi: 10.1016/S0076-6879(09)63016-0. [DOI] [PubMed] [Google Scholar]
- Merrow M, Lenssen D, Roenneberg T. Comparative clocks. The circadian clock. 2010;Vol. 12:157–77. [Google Scholar]
- Mori T, Williams DR, Byrne MO, Qin X, Egli M, McHaourab HS, Stewart PL, Johnson CH. Elucidating the ticking of an in vitro circadian clockwork. PLoS Biol. 2007;5:841–53. doi: 10.1371/journal.pbio.0050093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murakami R, Mutoh R, Iwase R, Furukawa Y, Imada K, Onai K, Morishita M, Yasui S, Ishii K, Valencia S, et al. The roles of the dimeric and tetrameric structures of the clock protein KaiB in the generation of circadian oscillations in cyanobacteria. J Biol Chem. 2012;287:29506–15. doi: 10.1074/jbc.M112.349092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murayama Y, Mukaiyama A, Imai K, Onoue Y, Tsunoda A, Nohara A, Ishida T, Maeda Y, Terauchi K, Kondo T, et al. Tracking and visualizing the circadian ticking of the cyanobacterial clock protein KaiC in solution. EMBO J. 2011;30:68–78. doi: 10.1038/emboj.2010.298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mutoh R, Mino H, Murakami R, Uzumaki T, Ishiura M. Thermodynamically induced conformational changes of the cyanobacterial circadian clock protein KaiB. Appl Magn Reson. 2011;40:525–34. [Google Scholar]
- Nakajima M, Imai K, Ito H, Nishiwaki T, Murayama Y, Iwasaki H, Oyama T, Kondo T. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science. 2005;308:414–5. doi: 10.1126/science.1108451. [DOI] [PubMed] [Google Scholar]
- Nishiwaki T, Kondo T. The circadian autodephosphorylation of cyanobacterial clock protein KaiC occurs via the formation of ATP as an intermediate. J Biol Chem. 2012;287:18030–5. doi: 10.1074/jbc.M112.350660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishiwaki T, Satomi Y, Kitayama Y, Terauchi K, Kiyohara R, Takao T, Kondo T. A sequential program of dual phosphorylation of KaiC as a basis for circadian rhythm in cyanobacteria. EMBO J. 2007;26:4029–37. doi: 10.1038/sj.emboj.7601832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pattanayek R, Mori T, Xu Y, Pattanayek S, Johnson CH, Egli M. Structures of KaiC circadian clock mutant proteins: a new phosphorylation site at T426 and mechanisms of kinase, ATPase and phosphatase. PLoS One. 2009;4:e7529. doi: 10.1371/journal.pone.0007529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pattanayek R, Wang J, Mori T, Xu Y, Johnson CH, Egli M. Visualizing a circadian clock protein: crystal structure of KaiC and functional insights. Mol Cell. 2004;15:375–88. doi: 10.1016/j.molcel.2004.07.013. [DOI] [PubMed] [Google Scholar]
- Pattanayek R, Williams DR, Pattanayek S, Mori T, Johnson CH, Stewart PL, Egli M. Structural model of the circadian clock KaiB–KaiC complex and mechanism for modulation of KaiC phosphorylation. EMBO J. 2008;27:1767–78. doi: 10.1038/emboj.2008.104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pattanayek R, Williams DR, Rossi G, Weigand S, Mori T, Johnson CH, Stewart PL, Egli M. Combined SAXS/EM based models of the S. elongatus post-translational circadian oscillator and its interactions with the output His-Kinase SasA. PLoS One. 2011;6:e23697. doi: 10.1371/journal.pone.0023697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pattanayek R, Yadagirl KK, Ohi MD, Egli M. Nature of KaiB–KaiC binding in the cyanobacterial circadian oscillator. Cell Cycle. 2013;12:810–7. doi: 10.4161/cc.23757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phong C, Markson JS, Wilhoite CM, Rust MJ. Robust and tunable circadian rhythms from differentially sensitive catalytic domains. Proc Natl Acad Sci USA. 2013;110:1124–9. doi: 10.1073/pnas.1212113110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plevin MJ, Boisbouvier J. Isotope-labelling of methyl groups for NMR studies of large proteins. In: Clore M, Potts J, editors. Recent developments in biomolecular NMR. Cambridge: RSC Publishing; 2012. pp. 1–24. [Google Scholar]
- Qin X, Byrne M, Mori T, Zou P, Williams DR, Mchaourab H, Johnson CH. Intermolecular associations determine the dynamics of the circadian KaiABC oscillator. Proc Natl Acad Sci USA. 2010;107:14805–10. doi: 10.1073/pnas.1002119107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren Z, Wang H, Ghose R. Dynamics on multiple timescales in the RNA-directed RNA polymerase from the cystovirus ϕ6. Nucleic Acids Res. 2010;38:5105–18. doi: 10.1093/nar/gkq210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rust MJ, Markson JS, Lane WS, Fisher DS, O’Shea EK. Ordered phosphorylation governs oscillation of a three-protein circadian clock. Science. 2007;318:809–12. doi: 10.1126/science.1148596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sprangers R, Gribun A, Hwang PM, Houry WA, Kay LE. Quantitative NMR spectroscopy of supramolecular complexes: dynamic side pores in ClpP are important for product release. Proc Natl Acad Sci USA. 2005;102:16678–83. doi: 10.1073/pnas.0507370102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sprangers R, Kay LE. Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature. 2007;445:618–22. doi: 10.1038/nature05512. [DOI] [PubMed] [Google Scholar]
- Takai N, Nakajima M, Oyama T, Kito R, Sugita C, Sugita M, Kondo T, Iwasaki H. A KaiC-associating SasA–RpaA two-component regulatory system as a major circadian timing mediator in cyanobacteria. Proc Natl Acad Sci USA. 2006;103:12109–14. doi: 10.1073/pnas.0602955103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terauchi K, Kitayama Y, Nishiwaki T, Miwa K, Murayama Y, Oyama T, Kondo T. ATPase activity of KaiC determines the basic timing for circadian clock of cyanobacteria. Proc Natl Acad Sci USA. 2007;104:16377–81. doi: 10.1073/pnas.0706292104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tzeng SR, Pai MT, Kalodimos CG. NMR studies of large protein systems. Methods Mol Biol. 2012;831:133–40. doi: 10.1007/978-1-61779-480-3_8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vakonakis I, LiWang A. NMR studies of a timekeeping system. In: Ditty JL, Mackey SR, Johnson CH, editors. Bacterial circadian programs. Berlin, Germany: Springer; 2009. pp. 103–20. [Google Scholar]
- Vakonakis I, LiWang A. Structure of the C-terminal domain of the clock protein KaiA in complex with a KaiC-derived peptide: implications for KaiC regulation. Proc Natl Acad Sci USA. 2004;101:10925–30. doi: 10.1073/pnas.0403037101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vakonakis I, Sun J, Wu T, Holzenburg A, Golden SS, LiWang A. NMR structure of the KaiC-interacting C-terminal domain of KaiA, a circadian clock protein: implications for the KaiA–KaiC interaction. Proc Natl Acad Sci USA. 2004;101:1479–84. doi: 10.1073/pnas.0305516101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams SB, Vakonakis I, Golden SS, LiWang A. Structure and function from the circadian clock protein KaiA of Synechococcus elongatus: a potential clock input mechanism. Proc Natl Acad Sci USA. 2002;99:15357–62. doi: 10.1073/pnas.232517099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X, Welch JL, Arnold JJ, Boehr DD. Long-range interaction networks in the function and fidelity of poliovirus RNA-dependent RNA polymerase studied by nuclear magnetic resonance. Biochemistry. 2010;49:9361–71. doi: 10.1021/bi100833r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye S, Vakonakis I, Ioerger TR, LiWang A, Sacchettini JC. Crystal structure of circadian clock protein KaiA from Synechococcus elongatus. J Biol Chem. 2004;279:20511–8. doi: 10.1074/jbc.M400077200. [DOI] [PubMed] [Google Scholar]