Abstract
Multidrug-resistant pathogens, such as methicillin-resistant Staphylococcus aureus (MRSA) pose an increasing health burden and demand alternative antimicrobials to treat bacterial infections. The surface coating AGXX® is a novel broad-spectrum antimicrobial composed of two transition metals, silver and ruthenium that can be electroplated on various surfaces, such as medical devices and implants. AGXX® has been shown to kill nosocomial and waterborne pathogens by production of reactive oxygen species (ROS), but the effect of AGXX® on the bacterial redox balance has not been demonstrated. Since treatment options for MRSA infections are limited, ROS-producing agents are attractive alternatives to combat multi-resistant strains. In this work, we used RNA-seq transcriptomics, redox biosensor measurements and phenotype analyses to study the mode of action of AGXX® microparticles in S. aureus USA300. Using growth and survival assays, the growth-inhibitory amount of AGXX® microparticles was determined as 5 μg/ml. In the RNA-seq transcriptome, AGXX® caused a strong thiol-specific oxidative stress response and protein damage as revealed by the induction of the PerR, HypR, QsrR, MhqR, CstR, CtsR, and HrcA regulons. The derepression of the Fur, Zur, and CsoR regulons indicates that AGXX® also interferes with the metal ion homeostasis inducing Fe2+- and Zn2+-starvation responses as well as export systems for toxic Ag+ ions. The induction of the SigB and GraRS regulons reveals also cell wall and general stress responses. AGXX® stress was further shown to cause protein S-bacillithiolation, protein aggregation and an oxidative shift in the bacillithiol (BSH) redox potential. In phenotype assays, BSH and the HypR-controlled disulfide reductase MerA were required for protection against ROS produced under AGXX® stress in S. aureus. Altogether, our study revealed a strong thiol-reactive mode of action of AGXX® in S. aureus USA300 resulting in an increased BSH redox potential and protein S-bacillithiolation.
Keywords: Staphylococcus aureus, AGXX®, transcriptome, bacillithiol, protein S-bacillithiolation
Introduction
Staphylococcus aureus inhabitants the skin and the nose of one quarter of the human population, but can also lead to serious live-threatening infections when the pathogen enters the bloodstream (Foster, 2004). S. aureus causes various diseases ranging from local skin abscesses to systemic and chronic infections, such as septicemia, endocarditis, pneumonia and osteomyelitis. S. aureus cells can also form biofilms on medical devices, such as catheters or implants (Archer, 1998; Lowy, 1998; Boucher and Corey, 2008). Multiple antibiotic resistant S. aureus isolates are an increasing health-burden that arise in hospitals and in the community, including methicillin-resistant S. aureus (MRSA) isolates (Livermore, 2000). S. aureus belongs to the most serious pathogens worldwide and is classified together with six Gram-negative bacteria as ESKAPE pathogen by the “European Center of Disease Prevention and Control” (Pendleton et al., 2013). Since treatment option for multiple antibiotic resistant S. aureus isolates are limited, new targets for antibiotics need to be identified and alternative antimicrobial approaches developed to combat the increasing problem of antimicrobial resistance. Among these strategies, reactive oxygen species (ROS)-producing antibiotics and inhibitors of antioxidant responses that affect the bacterial redox balance are attractive alternatives as previously shown for other human pathogens (Padiadpu et al., 2016; Tung et al., 2018).
Metals, like silver, copper, and iron are known to cause reactive oxygen species (ROS), such as the highly reactive hydroxyl radical which leads to oxidation of proteins, DNA bases and lipids explaining some antimicrobial effects of metal ions (Grass et al., 2011; Maillard and Hartemann, 2013). Moreover, silver ions can bind to thiol groups of metal-containing proteins, such as copper transporters, metal-sensing regulators or respiratory chain complexes, leading to loss of protein functions. The inhibition of respiratory enzymes in the electron chain might further promote ROS production, such as superoxide anions. The bactericidal effect of metallic copper and silver surfaces was also termed as “contact killing” (Grass et al., 2011; Maillard and Hartemann, 2013; Villapún et al., 2016). These metals are used as surface coating of medical devices, for wound healing and in water pipelines to clean drinking water (Lansdown, 2010). Especially silver has been applied for a long time as medicinal product against wound infections until the evolution of silver resistant bacteria in 1975 (Gupta et al., 1999).
The novel antimicrobial coating AGXX® (Largetec GmbH, Berlin) was recently developed as promising broad-spectrum antimicrobial that can be electroplated on various surfaces, including steel, ceramics, glass and organic polymers (Guridi et al., 2015; Clauss-Lendzian et al., 2018; Vaishampayan et al., 2018). AGXX® strongly inhibited the growth and biofilm formation of many antibiotic resistant nosocomial pathogens, such as S. aureus, Staphylococcus epidermidis, Enterococcus faecalis, and Enterococcus faecium (Guridi et al., 2015). Furthermore, AGXX® had a strong killing effect on Legionella erythra in water pipelines and Escherichia coli in urine samples. This antimicrobial effect of AGXX® was much more efficient compared to classical silver (Guridi et al., 2015). AGXX® is composed of two transition metals, silver (Ag) and ruthenium (Ru) that are conditioned with ascorbic acid. The Ag+ and Ru+ metal ions form a micro-galvanic cell and lead to ROS formation, such as H2O2 and the strong oxidant hydroxyl radical, which are proposed as main antimicrobial mechanism (Gupta et al., 1999; Guridi et al., 2015; Heiss et al., 2017; Clauss-Lendzian et al., 2018). Using RNA-seq transcriptomics, the antimicrobial mode of action of AGXX® has been recently studied in the MRSA isolate 04-02981 (Vaishampayan et al., 2018) and in a clinical E. faecalis isolate (Clauss-Lendzian et al., 2018). Gene expression profiling revealed repression of genes for biofilm formation and virulence factors as well as the agr quorum sensing system in S. aureus. The induction of the heat-shock regulons HrcA and CtsR was indicative of protein damage in both bacteria (Clauss-Lendzian et al., 2018; Vaishampayan et al., 2018). However, it is not known whether AGXX® stress causes an oxidative stress response and affects the redox state in S. aureus due to the production of ROS and if there are potential protection mechanisms against the antimicrobial AGXX®.
Staphylococcus aureus is frequently exposed to ROS, such as H2O2 and the strong oxidant hypochlorous acid (HOCl) during infection by activated macrophages and neutrophils (Winterbourn and Kettle, 2013; Hillion and Antelmann, 2015; Beavers and Skaar, 2016; Winterbourn et al., 2016). Several protection mechanisms are utilized by S. aureus to cope with ROS and HOCl, including the low molecular weight (LMW) thiol bacillithiol (BSH) which is important under macrophage infections (Newton et al., 2012; Pöther et al., 2013; Posada et al., 2014; Chandrangsu et al., 2018). BSH is involved in detoxification of various reactive species, ROS, HOCl, electrophiles, toxins and antibiotics and serves as thiol-cofactor for many redox enzymes, such as thiol-S-transferases, glyoxalases and antioxidant enzymes (Chandrangsu et al., 2018). Under conditions of HOCl stress, BSH forms mixed disulfides with proteins, termed as protein S-bacillithiolation (Chi et al., 2011, 2013; Imber et al., 2018a). Protein S-bacillithiolations are widespread redox-modifications, which function in thiol-protection and redox regulation of redox-sensing transcription factors and metabolic enzymes, such as the glycolytic glyceraldehyde-3-phosphate dehydrogenase (GapDH) and the aldehyde dehydrogenase AldA (Chi et al., 2011, 2013; Loi et al., 2015; Imber et al., 2018a,b,c). GapDH was identified as most abundant S-bacillithiolated protein in the S. aureus redox proteome. GapDH activity was inhibited by S-bacillithiolation under H2O2 and NaOCl stress in vitro which was faster compared to its irreversible inactivation by overoxidation (Imber et al., 2018a). Thus, our data confirmed that S-bacillithiolation efficiently protects the active sites of redox-sensitive enzymes against overoxidation to Cys sulfonic acids (Loi et al., 2015; Imber et al., 2018c).
Further protection mechanisms against oxidative stress are regulated by the SarA/MarR-family redox-sensing regulators (SarA, MgrA and SarZ), by the HOCl-inducible Rrf2-family regulater HypR and by the peroxide regulon repressor PerR (Horsburgh et al., 2001; Chen et al., 2006, 2009, 2011; Hillion and Antelmann, 2015; Ji et al., 2015; Chandrangsu et al., 2018). The SarZ-controlled OhrA peroxiredoxin is involved in detoxification of organic hydroperoxides that are derived from fatty acid oxidation (Dubbs and Mongkolsuk, 2007). The OhrR-homologs SarZ and MgrA can be regulated by S-thiolation with a synthetic LMW thiol, resulting in derepression of transcription of their target genes under oxidative stress (Abomoelak et al., 2009; Poor et al., 2009; Chen et al., 2011; Sun et al., 2012; Hillion and Antelmann, 2015). Recent OxICAT analyses revealed increased oxidation of the redox-sensing Cys residues of MgrA and SarZ under HOCl stress (Imber et al., 2018a). This indicates that both regulators could be regulated by S-bacillithiolation in S. aureus under HOCl stress.
The novel redox-sensing regulator HypR was shown to control the pyridine nucleotide disulfide reductase MerA which provides specific protection under HOCl stress and macrophage infections (Loi et al., 2018). HypR belong to the Rrf2-family that senses and responds to HOCl stress by a thiol-redox switch involving Cys33-Cys99′ intersubunit disulfide formation between adjacent subunits of the HypR dimer (Loi et al., 2018). In addition, the H2O2 response is controlled by the peroxide repressor PerR in S. aureus. PerR uses metal-catalyzed histidine oxidation to sense peroxide stress (Horsburgh et al., 2001; Lee and Helmann, 2006; Ji et al., 2015; Pinochet-Barros and Helmann, 2018). PerR regulates genes encoding catalases (KatA), peroxiredoxins (AhpCF, Bcp, Tpx), the miniferritin (Dps) and the iron-sulfur cluster SUF machinery under H2O2 stress in S. aureus (Horsburgh et al., 2001; Ji et al., 2015; Pinochet-Barros and Helmann, 2018).
In this study, we applied RNA-seq transcriptomics to study the mode of action of the AGXX® antimicrobial coating in S. aureus USA300. Our results revealed that AGXX® causes a thiol-specific oxidative stress response, as shown by the strong up-regulation of the PerR, HypR, QsrR, MhqR, and CstR regulons. The up-regulation of the CtsR and HrcA heat shock regulons is caused by oxidative protein damage and aggregation. The induction of the GraRS and SigmaB regulons further indicates cell wall and general stress responses under AGXX® exposure. Finally, AGXX® stress interferes with metal homeostasis as revealed by the increased transcription of the CsoR, Fur, and Zur regulons. AGXX® further leads to oxidative shift in the BSH redox potential, enhanced protein S-bacillithiolation and protein aggregation, supporting its thiol-reactive mode of action. Phenotype assays revealed that the LMW thiol BSH and the HypR-controlled disulfide reductase MerA provide protection under AGXX® stress in S. aureus.
Materials and Methods
Preparation of AGXX® Microparticles
AGXX® microparticles of charge 373 were kindly provided by Largentec GmbH, Berlin, Germany. The AGXX® microparticles consist of a powder of silver with particle size of 1.5–2.5 μm (MaTecK, Germany), which was chemically coated with ruthenium (Guridi et al., 2015; Heiss et al., 2017). The coating was performed in alkaline medium where the Ru(III) ions were first oxidized by NaOCl to RuO4. Reduction of RuO4 to Ru or RuOX was performed by addition of NaNO2 as described previously (Chen et al., 2010). The black AGXX® powder was further conditioned with 50 mM ascorbic acid for 2 h, followed by filtration, washing with deionized water and drying.
Bacterial Strains, Growth and Cultivation
The S. aureus strains used for the experiments with AGXX® include the highly virulent community-acquired (CA) MRSA strain S. aureus USA300 and the hospital-acquired (HA) MRSA strain S. aureus COL. For phenotype analysis we further compared the S. aureus USA300 ΔbshA mutant (Posada et al., 2014), the S. aureus COL ΔmerA deletion mutant and its complemented strain as described previously (Loi et al., 2018). S. aureus strains were cultivated in RPMI medium as described (Imber et al., 2018a). For survival phenotype assays under AGXX® stress, S. aureus USA300 was grown in RPMI medium to an OD500 of 0.5 and exposed to different amounts of 3–10 μg/ml AGXX® microparticles. The culture was diluted and 10 μl of serial dilutions were spotted onto LB agar plates. Colony forming units (CFU) were observed after incubation of the LB plates for 24 h.
Live/Dead Viability Assays
The viability assay of S. aureus USA300 was performed after treatment with sub-lethal and lethal amounts (4 and 10 μg/ml) of AGXX® at an OD500 of 0.5 using the LIVE/DEAD™ BacLight™ Bacterial Viability Kit (Thermo Fisher) according to the instructions of the manufacturers. S. aureus strains were stained with SYTO9 and propidium iodide for each 15 min. Stained cells were washed in phosphate buffer and live or dead cells were visualized using a fluorescence microscope (Nikon, Eclipse, Ti2) (SYTO9 Ex: 488 nm, propidium iodide Ex: 555 nm). Fluorescence intensity was measured after excitation at 488 and 555 nm and false-colored in green or red for live or dead cells, respectively.
RNA Isolation, Library Preparation, and Next Generation cDNA Sequencing
Staphylococcus aureus USA300 was cultivated in RPMI medium in 3 biological replicates and treated with 4.5 μg/ml AGXX® stress as described previously (Loi et al., 2017). S. aureus cells were harvested before (as untreated control) and 30 min after exposure to 4.5 μg/ml AGXX® and disrupted in 3 mM EDTA/ 200 mM NaCl lysis buffer with a Precellys24 ribolyzer. RNA isolation was performed using the acid phenol extraction protocol as described (Chi et al., 2014). The RNA quality was checked by Trinean Xpose (Gentbrugge, Belgium) and the Agilent RNA Nano 6,000 kit using an Agilent 2,100 Bioanalyzer (Agilent Technologies, Böblingen, Germany). Ribo-Zero rRNA Removal Kit (Bacteria) from Illumina (San Diego, CA, USA) was used to remove the rRNA. TruSeq Stranded mRNA Library Prep Kit from Illumina (San Diego, CA, United States) was applied to prepare the cDNA libraries. The cDNAs were sequenced paired end on an Illumina HiSeq 1,500 (San Diego, CA, United States) using 70 and 75 bp read length and a minimum sequencing depth of 10 million reads per library. The transcriptome sequencing raw data files are available in the ArrayExpress database (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-7074.
Bioinformatics Data Analysis, Read Mapping, Data Visualization, and Analysis of Differential Gene Expression
The paired end cDNA reads were mapped to the S. aureus USA300_TCH1516 genome sequence (accession number CP000730) (Highlander et al., 2007) using bowtie2 v2.2.7 (Langmead and Salzberg, 2012) with default settings for paired-end read mapping. All mapped sequence data were converted from SAM to BAM format with SAMtools v1.3 (Li et al., 2009) and imported to the software ReadXplorer v.2.2 (Hilker et al., 2016).
Differential gene expression analysis of triplicates including normalization was performed using Bioconductor package DESeq2 (Love et al., 2014) included in the ReadXplorer v2.2 software (Hilker et al., 2016). The signal intensity value (A-value) was calculated by log2 mean of normalized read counts and the signal intensity ratio (M-value) by log2 fold-change. The evaluation of the differential RNA-seq data was performed using an adjusted p-value cut-off of P ≤ 0.05 and a signal intensity ratio (M-value) cut-off of ≥ 0.6 or ≤ −0.6 (fold-change of ± 1.5). The cut-off was determined as ± 0.6 since the majority of differentially transcribed regulons fall in this range that are related to the redox-active mode of action of AGXX®.
Construction of the Voronoi Transcriptome Treemap
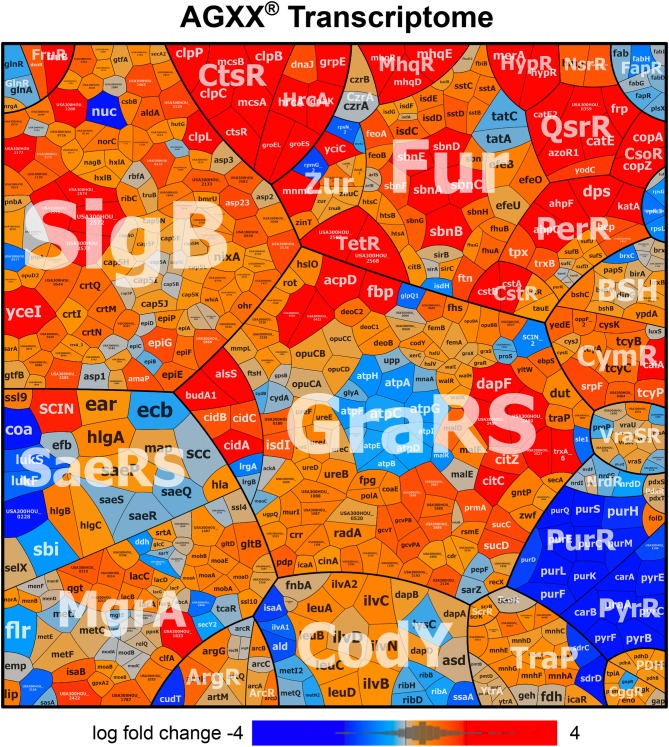
The Paver software (DECODON GmbH, Greifswald, Germany) was used to generate the AGXX® transcriptome treemap as previously described (Mehlan et al., 2013), showing log2-fold changes (M-values) of selected genes, operons and regulons that are up-regulated under AGXX® stress. The treemap is structured into regulons (white labels), genes and operons (smaller labels). The cell sizes are defined as absolute log2-fold changes of expression levels after AGXX® stress divided by the untreated control. The log2-fold ratios of AGXX® vs. the untreated controls are displayed by a blue-red color gradient where blue indicates repression and red induction under AGXX ® stress.
Northern Blot Experiments
Northern blot analysis was performed as described previously (Wetzstein et al., 1992) using RNA isolated from S. aureus USA300 that was exposed to 4.5 μg/ml AGXX® for 30 min as indicated. Hybridizations were performed using the digoxigenin-labeled merA-specific antisense RNA synthesized in vitro using T7 RNA polymerase as described (Tam Le et al., 2006; Loi et al., 2018).
Western Blot Analysis
Staphylococcus aureus USA300 cells were grown in LB until an OD540 of 2 and harvested by centrifugation. Cells were transferred to Belitsky minimal medium (BMM) for 1 h and treated with increasing amounts of 3–10 μg/ml AGXX® microparticles for 30 min. Cells were harvested and washed in TE-buffer (pH 8.0) with 50 mM NEM, disrupted using the ribolyzer and the protein extract was cleared from cell debris by centrifugation. Protein amounts of 25 μg were separated using 12% SDS-PAGE and subjected to BSH-specific Western blot analysis using the polyclonal rabbit anti-BSH antiserum as described previously (Chi et al., 2013).
Measurement of the BSH Redox Potential in S. aureus COL Under AGXX® Stress Using the Brx-roGFP2 Biosensor
For BSH redox potential measurements under AGXX® stress using the Brx-roGFP2 biosensor, we applied S. aureus COL since biosensor expression and fluorescence intensity was higher in S. aureus COL compared to S. aureus USA300 as revealed previously (Loi et al., 2017). S. aureus COL expressing the Brx-roGFP2 plasmid (Loi et al., 2017) was grown in RPMI medium to OD500 of 0.5 and exposed to 4 μg/ml of AGXX® for different times. The Brx-roGFP2 redox state was blocked with 10 mM N-ethylmaleimide (NEM) and the S. aureus cells transferred to the microplate wells for fluorescence measurements as described (Loi et al., 2017). Samples for fully reduced and oxidized controls were treated for 20 min with 10 mM DTT and 5 mM diamide, respectively. The Brx-roGFP2 biosensor fluorescence emission was measured at 510 nm after excitation at 405 and 488 nm using the CLARIOstar microplate reader (BMG Labtech). The OxD of the Brx-roGFP2 biosensor was determined for each sample and normalized to fully reduced (DTT-treated) and oxidized (diamide-treated) controls as described (Loi et al., 2017) based to the following equation (1):
| (1) |
The values of I405sample and I488sample are the observed fluorescence excitation intensities at 405 and 488 nm, respectively. The values of I405red, I488red, I405ox, and I488ox represent the fluorescence intensities of fully reduced and oxidized controls, respectively.
Based on the OxD and = – 280 mV (Dooley et al., 2004), the BSH redox potential can be calculated according to the Nernst equation (2):
| (2) |
Analysis of Protein Aggregates After AGXX® Stress
The fractions with intracellular protein aggregates were isolated as described previously with some modifications (Tomoyasu et al., 2001; Groitl et al., 2017). S. aureus USA300 was grown in RPMI medium until an OD500 of 0.5 and exposed to 4, 8, or 10 μg/ml AGXX® for 30 min. The S. aureus samples were adjusted to 8 ml of an OD500 of 1 and harvested by centrifugation. The cell pellets were resuspended in 40 μl buffer A (10 mM potassium phosphate pH 6.5, 1 mM EDTA, 20% sucrose, 5 μg/ml lysostaphin) and incubated on ice for 30 min. The cells were further diluted in 360 μl buffer B (10 mM potassium phosphate pH 6.5, 1 mM EDTA) and disrupted with glass beads using the PrecellysEvolution ribolyzer (Bertin technologies). Intact cells were removed by centrifugation at 2,000 rpm and the insoluble protein fraction was collected by a second centrifugation step (14,000 rpm, 45 min). Pellets were frozen at−20°C for 30 min and resuspended in 400 μl of buffer B followed by brief sonication. Membrane proteins were dissolved with 160 μl buffer C (buffer B plus 2% Nonidet P-40) and the protein aggregates collected by centrifugation. The aggregated proteins were washed with 400 μl buffer B and separated using SDS-PAGE analysis.
Results
The Novel Antimicrobial Coating AGXX® Efficiently Inhibits the Growth and Viability of S. aureus USA300
To investigate the effective antimicrobial amount of AGXX® microparticles that inhibit the growth of S. aureus USA300, we performed growth curves and live/dead assays. S. aureus USA300 was grown in RPMI medium and exposed to different amounts of AGXX® microparticles (3–5 μg/ml) during the exponential growth phase. The growth of S. aureus USA300 was not affected by 3–4 μg/ml AGXX®, but 4.5–5 μg/ml AGXX® significantly reduced the growth rate (Figure 1A). Live/dead staining with the LIVE/DEAD™ Bacterial Viability Kit was used to analyze bacterial survival after exposure of S. aureus to sub-lethal (4 μg/ml) and lethal amounts (10 μg/ml) of AGXX®. The results revealed that 4 μg/ml is sub-lethal for S. aureus while 10 μg/ml has a strong killing effect on the majority of S. aureus cells in RPMI medium (Figure 1B). These results of live/dead assays were confirmed by analysis of colony forming units (CFUs) as shown in the next sections for phenotype analyses of different mutants. To analyze the physiological effect and stress response caused by AGXX® in the transcriptome of S. aureus USA300, we have chosen sub-lethal amounts of 4.5 μg/ml AGXX® which slightly affected the growth rate (Figure 1A).
Figure 1.
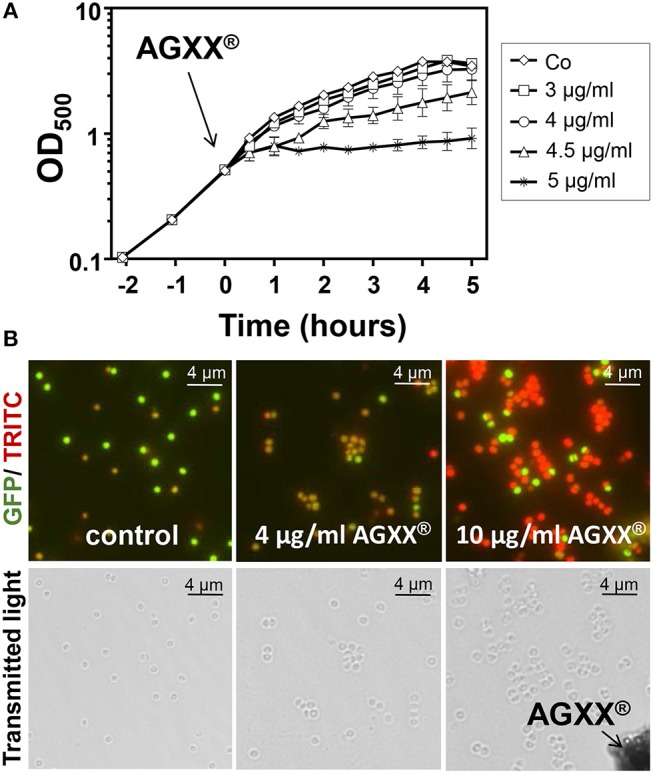
AGXX® has a strong antimicrobial effect on S. aureus USA300. (A) Growth curves of S. aureus USA300 were performed in RPMI medium which was exposed to 3–5 μg/ml AGXX® stress at an OD500 of 0.5. The results are from 3 biological replicate experiments. Error bars represent the standard deviation (SD). (B) Viability of S. aureus USA300 wild type was analyzed before and 30 min after exposure to sub-lethal (4 μg/ml) or lethal amounts (10 μg/ml) of AGXX® at OD500 of 0.5 using the LIVE/DEAD™ BacLight™ Bacterial Viability Kit. Live and dead S. aureus USA30 cells were visualized with a fluorescence microscope (Nikon, Eclipse, Ti2). S. aureus with an intact cytoplasmic membrane shows green fluorescence and dead cells with damaged membranes exhibit red fluorescence. The black arrow denotes AGXX® microparticles.
AGXX® Stress Causes a Thiol-Specific Oxidative and Metal Stress Response in the S. aureus Transcriptome
To investigate the stress response caused by AGXX® in S. aureus USA300, we studied the changes in the RNA-seq transcriptome after exposure to sub-lethal amounts of 4.5 μg/ml AGXX® stress. The RNA samples were isolated from 3 biological replicates before (control) and 30 min after AGXX® stress and analyzed by RNA-seq transcriptomics as in previous studies (Hillion et al., 2017; Loi et al., 2018). For significant expression changes, the M-value cut-off (log2-fold change AGXX®/control) of ≥0.6 and ≤ −0.6 (fold-change of ± 1.5, P ≤ 0.05) was chosen since the majority of most strongly induced regulons fall in this range. In total, 925 transcripts were significantly >1.5-fold up-regulated and 896 were <-1.5-fold down-regulated in the transcriptome of S. aureus under AGXX® stress (Figure 2 and Tables S1, S2). Among the up-regulated genes are 149 with fold-changes of >10-fold that could be classified into the PerR, HypR, QsrR, MhqR, CtsR, HrcA, CidR, CymR, CstR, CsoR, Fur, Zur, SigB, and GraRS regulons. The most strongly induced regulons in the AGXX® transcriptome are shown in the ratio/intensity scatter plot (M/A-plot) and visualized using the Voronoi transcriptomics treemap by red-blue color codes (Figures 2, 3 and Table S2).
Figure 2.
RNA-seq transcriptomics of S. aureus USA300 after 30 min of AGXX® stress. For RNA-seq transcriptomics, S. aureus USA300 was grown in RPMI medium and treated with 4.5 μg/ml AGXX® for 30 min. The gene expression profile under AGXX® stress is shown as ratio/intensity scatter plot (M/A-plot) which is based on differential gene expression analysis using DeSeq2. Colored symbols indicate significantly induced (red, orange, brown, blue, cyan, violet, green, yellow) or repressed (dark gray) transcripts (M ≥0.6 or ≤ -0.6; P ≤ 0.05). Light gray symbols denote transcripts with no fold-changes after AGXX® stress (P > 0.05). The PerR, TetR, HypR, QsrR, MhqR, CtsR, HrcA, CidR, CymR, Fur, CstR, CsoR, and Zur regulons are most strongly up-regulated under AGXX® stress. The transcriptome analysis was performed from three biological replicates. The RNA-seq expression data of all genes after AGXX® stress and their regulon classifications are listed in Tables S1, S2.
Figure 3.
AGXX® induces a strong thiol-specific oxidative and metal stress response in the RNA-seq transcriptome of S. aureus USA300. The treemap shows the log2-fold changes (M-values) for the induction and repression of selected regulons after exposure to 4.5 μg/ml AGXX® stress in the RNA-seq transcriptome of S. aureus USA300. The genes are classified into operons and regulons based on the RegPrecise database (http://regprecise.lbl.gov/RegPrecise/index.jsp) and previous studies (Mäder et al., 2016). Log2-fold changes in gene expression are visualized using a red-blue color code where red indicates log2-fold induction and blue repression of transcription under AGXX® exposure. AGXX® caused a strong thiol-specific oxidative and metal stress response and protein damage as shown by the strong induction of the PerR, HypR, QsrR, MhqR, CtsR, HrcA, and CstR regulons in S. aureus USA300. The induction of the CsoR, Fur, Zur, SigmaB, and GraRS regulons further indicates metal stress, cell wall and general stress responses in S. aureus. The detailed transcriptome data of all differentially expressed genes in response to AGXX® stress are presented in Tables S1, S2.
We have previously used RNA-seq transcriptomics to analyze the response of S. aureus USA300 to the strong oxidant NaOCl (Loi et al., 2018). NaOCl stress caused a thiol-specific oxidative stress signature and protein damage in S. aureus USA300 as revealed by the induction of the HypR, PerR, QsrR, HrcA, and CtsR regulons (Loi et al., 2018). NaOCl stress further provoked the weak induction of the Fur and Zur regulons in S. aureus indicative of metal starvation. Treatment of S. aureus USA300 with 4.5 μg/ml AGXX® stress resulted in similar changes in gene expression, with increased expression of the PerR, HypR and QsrR regulons that are related to the thiol-specific oxidative and electrophile stress response (Figures 2, 3). The HypR-controlled hypR-merA operon was among the most strongly up-regulated hits (fold-changes of 320-372). The USA300HOU_2567-68 operon of unknown function was even 977-1117-fold induced under AGXX® stress. The USA300HOU_2567-68 operon is regulated by the upstream located USA300HOU_2566 gene that encodes for a TetR-family regulator (our unpublished data). Thus, the USA300HOU_2567-68 operon was renamed as TetR regulon.
In addition, the CtsR and HrcA regulons were highly up-regulated under AGXX® with fold-changes of >119 for the ctsR-mcsA-mcsB-clpC operon and of >30 for the hrcA-grpE-dnaKJ operon. The HrcA and CtsR repressors control the transcription of proteases and chaperones of the quality control machinery (Frees et al., 2007, 2014), indicating that AGXX® stress might cause strong protein damage and aggregation. The increased transcription of the Fur and Zur regulons under AGXX® stress further indicates iron and zinc depletion under AGXX® stress, confirming its impact on metal homeostasis. Moreover, in agreement with previous studies, the Cu+ sensing CsoR-regulated copZA operon was highly upregulated (48–56-fold) under AGXX® stress in S. aureus (Grossoehme et al., 2011). In addition, the CsoR-like persulfide-specific CstR regulon which includes the cstAB-sqr and cstR-tauE operons was strongly induced under AGXX® stress in S. aureus (fold-changes of >30 and 2–3, respectively). The CstR regulon responds to hydrogen sulfide (H2S) and nitroxyl (HNO) (Luebke et al., 2013, 2014; Peng et al., 2017). AGXX® might cause thiol-oxidation of the metal binding Cys ligands of CsoR, CstR, Fur and Zur that could trigger the inactivation of the repressors and derepression of the corresponding regulons. We further noted the up-regulation of the CymR regulon and genes related to bacillithiol (BSH) biosynthesis and recycling (e.g., bshA operon, bshB, bshC, brxB, and ypdA) under AGXX® stress in S. aureus. This supports the notion that AGXX® leads to increased thiol-oxidation in proteins and depletion of the LMW thiol BSH.
The SigmaB regulon was also induced under AGXX® stress in S. aureus, which was required for intracellular growth during cell culture infection assays (Pförtner et al., 2014). The cell wall stress responsive GraRS is further induced under AGXX® stress, which responds to cell-wall active antibiotics and antimicrobial peptides and confers resistance to oxidative stress in S. aureus (Falord et al., 2011). However, in contrast to previous studies with AGXX® surface coating, we did not find significant changes in expression of the AgrAC two-component quorum-sensing system controlling virulence factors, such as hemolysins and surface factors (Novick, 2003; Queck et al., 2008). In our RNA-seq analysis, sub-lethal amounts of AGXX® microparticles caused a strong thiol-specific oxidative and metal stress response as well as protein damage as major transcriptome signature.
AGXX® Stress Causes Increased Protein S-bacillithiolation, Protein Aggregation and an Oxidative Shift in the BSH Redox Potential in S. aureus
NaOCl stress has been previously shown to cause increased thiol-oxidation of 58 Cys residues in the redox proteome of S. aureus USA300 (Imber et al., 2018a). Among those NaOCl-sensitive proteins are five S-bacillithiolated proteins including the glycolytic GapDH as major target for S-bacillithiolation (Imber et al., 2018a). Since the transcriptome signature overlaps between NaOCl and AGXX® stress, we hypothesized that AGXX® could also lead to increased S-bacillithiolation in S. aureus USA300. Using BSH-specific non-reducing Western blots, we monitored the level of S-bacillithiolated GapDH after exposure of S. aureus USA300 to different amounts of 3–10 μg/ml AGXX® (Figure 4A). As control, the bshA mutant was exposed to 10 μg/ml AGXX® which should not show any BSH-mixed disulfides in the BSH-Western blot. The treatment of S. aureus USA300 with sub-lethal 3–4 μg/ml AGXX® resulted in increased levels of the S-bacillithiolated GapDH (GapDH-SSB). The level of GapDH-SSB gradually increased with higher AGXX® amounts and bands of other S-bacillithiolated proteins appeared with 10 μg/ml AGXX® (Figure 4A). This confirms a similar thiol-reactive nature of AGXX® and NaOCl, which both induce protein S-bacillithiolation in S. aureus USA300.
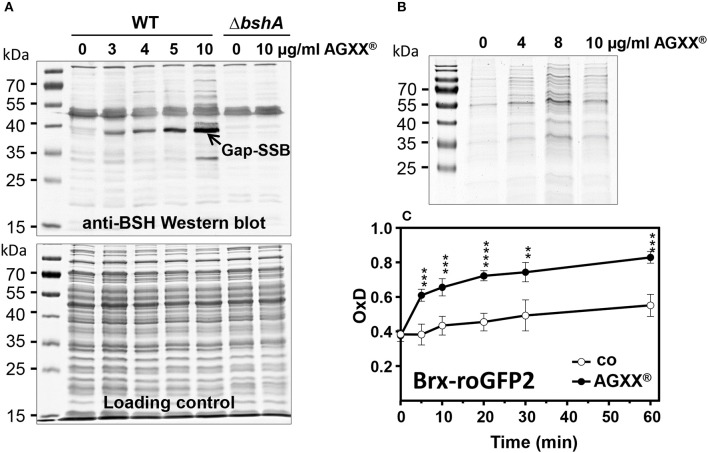
Figure 4.
AGXX® causes protein S-bacillithiolation, protein aggregation and an oxidative shift in the BSH redox potential in S. aureus. (A) Non-reducing BSH-specific Western blot analysis showed increasing of S-bacillithiolated GapDH protein in S. aureus USA300 strain under AGXX® stress. The Coomassie-stained SDS-PAGE is shown as loading control below the BSH Western blot. (B) To analyze protein aggregates, S. aureus cells were grown in RPMI medium to OD500 of 0.5 and treated with 4–10 μg/ml AGXX® for 30 min. The insoluble protein fractions of protein aggregates were obtained by centrifugation of the cell lysates and repeated centrifugation of the NP-40-dissolved pellet as described in the Methods section. The insoluble protein fractions were analyzed by SDS-PAGE and visualized with Coomassie staining (C) S. aureus COL expressing Brx-roGFP2 was treated with the sub-lethal amount of 4 μg/ml AGXX® at OD500 of 0.5 in RPMI medium and the biosensor oxidation degree (OxD) was measured at different times after stress. The Brx-roGFP2 biosensor is rapidly oxidized by AGXX®. The OxD values of the Brx-roGFP2 biosensor measurements with and without AGXX® were calibrated to fully reduced (DTT-treated) and oxidized (diamide-treated) control samples. The OxD values of fully reduced and oxidized controls were defined as “0” and “1,” respectively. The data were calculated as mean values from 3 biological replicate experiments. Error bars represent the SD and p-values are calculated using a Student's unpaired two-tailed t-test by the graph prism software. Symbols are defined as follows: **p ≤ 0.01; ***p ≤ 0.001 and ****p ≤ 0.0001 (p < 0.0003 for control/AGXX stress at 5 min; p = 0.0002 at 10 min; p < 0.0001 at 20 min; p = 0.0013 at 30 min; p = 0.0001 at 60 min).
The strong induction of the HrcA and CtsR regulons further indicates oxidative protein unfolding and aggregation which has been shown under HOCl stress previously (Groitl et al., 2017). Thus, we isolated the insoluble protein fraction that includes protein aggregates after exposure of S. aureus USA300 to AGXX® stress. Increasing levels of 4–10 μg/ml AGXX® microparticles resulted in elevated amounts of aggregated proteins as shown in the SDS-PAGE of the insoluble protein fractions (Figure 4B). Thus, AGXX® clearly caused oxidative protein unfolding and aggregation in S. aureus USA300.
The increased level of protein S-bacillithiolation of GapDH under AGXX® stress points to an oxidative shift in the BSH redox potential (EBSH) in S. aureus USA300. We have recently constructed a genetically encoded Brx-roGFP2 biosensor to measure dynamic EBSH changes in S. aureus (Loi et al., 2017). Using the Brx-roGFP2 biosensor, we analyzed the changes in EBSH in S. aureus COL after different times of treatment with sub-lethal 4 μg/ml AGXX® (Figure 4C). The oxidation degree (OxD) of the biosensor was around 0.4 in S. aureus control cells at an OD500 of 0.5 and increased to an OxD of ~0.5 after 60 min. In AGXX®-treated cells, the OxD of the Brx-roGFP2 biosensor increased to 0.6–0.7 after 5–20 min and reached nearly the fully oxidized state of 0.8 after 60 min of stress (Figure 4C). This confirms the thiol-reactive mode of action of AGXX® which leads to an oxidative shift of EBSH due to increased ROS formation as revealed previously (Clauss-Lendzian et al., 2018).
Bacillithiol and the HypR-controlled Disulfide Reductase MerA Provide Protection Under Oxidative Stress Generated During AGXX® Stress in S. aureus USA300
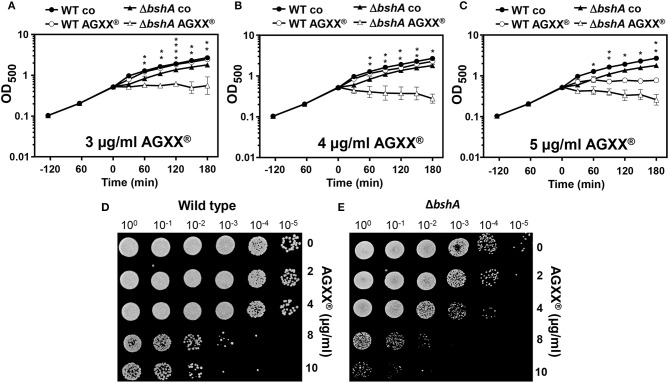
The RNA-seq transcriptome results and redox biosensor measurements revealed the thiol-reactive mode of action of AGXX® in S. aureus. Next, we studied the impact of the LMW thiol BSH in protection against ROS produced under AGXX® stress in S. aureus. We compared the growth and survival of the S. aureus USA300 wild type and the bshA mutant after treatment with increasing doses of AGXX®. While the growth of the wild type was not affected with 3–4 μg/ml AGXX®, the bshA mutant was very sensitive and showed a growth defect with 3 μg/ml AGXX® (Figures 5A–C). The survival of the bshA mutant was also strongly decreased after exposure to 4 μg/ml which is sub-lethal for the wild type. In addition, lethal doses of 8–10 μg/ml AGXX® have a stronger killing effect for the bshA mutant compared to the wild type (Figures 5D,E). These phenotype results indicate that BSH provides protection under AGXX® stress and is most likely involved in detoxification of ROS, such as H2O2 and hydroxyl radical generated during AGXX® exposure in S. aureus.
Figure 5.
BSH protects S. aureus against AGXX® stress. Growth phenotype analyses (A–C) and survival assays (D–E) were performed for the S. aureus USA300 wild type (WT) and the bshA mutant before and after exposure to AGXX® stress. (A–C) For growth comparisons, strains were exposed to 3, 4, and 5 μg/ml AGXX® stress at an OD500 of 0.5. The results are from 3 biological replicate experiments. Error bars represent the SD and the statistics was calculated using a Student's unpaired two-tailed t-test by the graph prism software. Symbols are defined as follows:*p ≤ 0.05; **p ≤ 0.01 and ****p ≤ 0.0001. (D–E) For survival assays, S. aureus USA300 wild-type (D) and the bshA mutant (E) were grown in RPMI until an OD500 of 0.5 and treated with 2–10 μg/ml AGXX®. Survival assays were performed by spotting 10 μl of serial dilutions after 2 and 3 h of AGXX® exposure onto LB agar plates to observe CFUs after 24 h incubation.
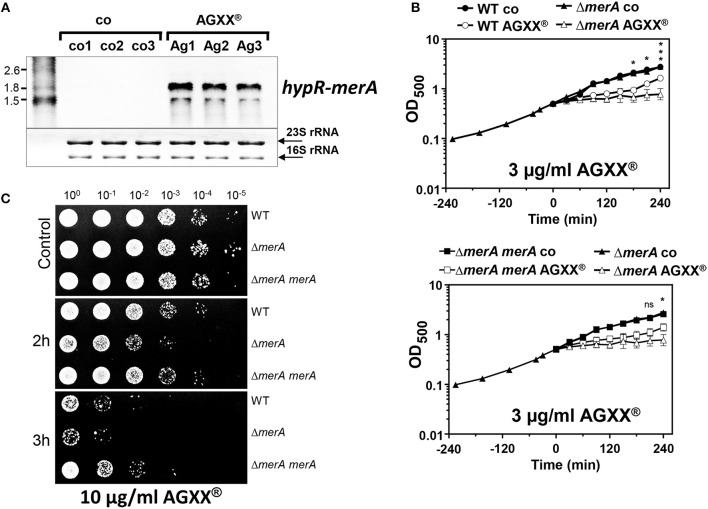
In addition, the hypR-merA operon was most strongly induced under AGXX® stress (fold-changes of 320–372) in the RNA-seq transcriptome, which was previously characterized as defense mechanism under HOCl stress in S. aureus (Loi et al., 2018). The Rrf2 family redox regulator HypR was shown to sense HOCl stress by a thiol-disulfide switch leading to repressor inactivation and induction of the NADPH-dependent flavin disulfide reductase MerA. MerA provides protection under HOCl stress and infection conditions in S. aureus COL (Loi et al., 2018). The strong induction of hypR-merA operon in S. aureus by sub-lethal 4.5 μg/ml AGXX® was confirmed in Northern blot analysis (Figure 6A). Thus, we studied the role of MerA in protection under AGXX® stress in S. aureus COL using growth and survival phenotype assays. For phenotype analyses, we have chosen S. aureus COL as wild type, since all previous phenotype analysis of the merA mutant and its complemented strain under NaOCl stress were performed in S. aureus COL background (Loi et al., 2018). First, we noted that the growth of S. aureus COL is slightly impaired with 3 μg/ml AGXX® indicating that S. aureus COL is more sensitive to 3 μg/ml AGXX® compared to S. aureus USA300 (Figures 5A, 6B). The merA mutant showed a stronger growth inhibition with 3 μg/ml AGXX® than the wild type. Complementation of the merA mutant with plasmid-encoded merA restored the growth to wild type level (Figure 6B). The merA mutant was also more sensitive to killing by the lethal dose of 10 μg/ml AGXX® stress compared to the wild type (Figure 6C). The survival phenotype of the merA mutant could be restored back to wild type level after complementation with plasmid-encoded merA. This indicates that MerA provides protection against oxidative stress that is encountered during AGXX® treatment in S. aureus. MerA might help to reduce oxidized protein disulfides or LMW thiol disulfides, which remains to be elucidated. These phenotype results identified important roles of the LMW thiol BSH and the disulfide reductase MerA in the defense against strong disulfide stress encountered from ROS generation during AGXX® exposure in S. aureus USA300.
Figure 6.
The NADPH-dependent disulfide reductase MerA is strongly induced and required for growth and survival under AGXX® stress in S. aureus. (A) Northern blot analysis was performed for hypR-merA operon transcription in S. aureus USA300 wild type before (Co1-3) and 30 min after exposure to 4.5 μg/ml AGXX® (Ag1-3). The RNA samples 1–3 correspond to 3 biological replicates. The methylene blue bands denote the 16S and 23S rRNAs as RNA loading controls below the Northern blots. (B) The growth curves were compared for the S. aureus COL wild type (WT), the merA deletion mutant and the merA complemented strain in RPMI medium before (control) and after treatment with 3 μg/ml AGXX®. The time point before AGXX® exposure was set to “0.” Results of three biological replicates are presented as mean values. Error bars represent the SD and the statistics was calculated using a Student's unpaired two-tailed t-test by the graph prism software. Symbols are defined as follows: nsp > 0.05; *p ≤ 0.05 and ***p ≤ 0.001. (C) Survival was analyzed for S. aureus COL wild-type (WT), the merA mutant and the complemented strain after exposure to 10 μg/ml AGXX® in RPMI medium. Survival assays were performed by spotting 10 μl of serial dilutions after 2 and 3 h of AGXX® exposure onto LB agar plates. Three biological replicate experiments were performed and one representative survival experiment is shown.
Discussion
The novel antimicrobial coating AGXX® has a strong killing effect on a broad range of Gram-positive and Gram-negative bacteria and was shown to inhibit biofilm formation in the MRSA strain S. aureus 04-02981 and in a clinical E. faecalis isolate (Guridi et al., 2015; Clauss-Lendzian et al., 2018; Vaishampayan et al., 2018). Previous RNA-seq transcriptomics analysis in S. aureus revealed the downregulation of genes encoding virulence factors regulated by the Agr system (Vaishampayan et al., 2018). It was postulated that AGXX® inhibits the Agr system and thereby acts as potential biofilm inhibitor as main killing mode (Vaishampayan et al., 2018). In our RNA-seq experiments, we could not confirm the significant downregulation of the agrBDCA operon upon AGXX® stress and observed rather a 2–3-fold up-regulation of the α- and γ-hemolysins (hla, hlgABC) and significantly down-regulation of surface proteins spa, sdrC, sdrD, sle1 in the transcriptome. Thus, our analysis suggests rather a slightly increased Agr activity under sub-lethal AGXX® stress in S. aureus. The Agr system is activated by autoinducer peptides that are induced by high cell density after the entry into the stationary phase (Novick, 2003; Queck et al., 2008). Since AGXX® exposure caused a growth defect, this could lead to activation of the Agr system in our study resulting in increased toxin and decreased surface protein expression. The discrepancies between the RNAseq results of the previous study (Vaishampayan et al., 2018) and our work might be related to the different AGXX® coating surfaces. We used small AGXX® microparticles (1.5–2.5 μM), which have a large volume/surface area, while larger AGXX® coated metal sheets (24 cm2) were used in the previous study (Vaishampayan et al., 2018). Thus, bacteria might be exposed more equally to the small microparticles in the shaking bacterial cultures, while metal sheets may not contact all bacteria in the shaking culture simultaneously.
However, previous analyses in E. faecalis revealed also the formation of ROS, such as H2O2 and hydroxyl radicals at the surface of AGXX® coatings as measured using spectroscopic methods (Clauss-Lendzian et al., 2018). These data point to a major role of ROS in the killing mode of AGXX® in bacteria. It is postulated that an electric field is formed between the transition metals silver and ruthenium leading to step-wise electron transfer to molecular oxygen and subsequent ROS formation (Guridi et al., 2015). Transcriptome in E. faecalis supported the oxidative stress mode of action of AGXX® since expression of genes for thioredoxins, catalase, superoxide dismutase and glutathione synthetase was highly upregulated (Clauss-Lendzian et al., 2018). In this study, we used a combination of RNA-seq transcriptomics, analyses for protein S-bacillthiolations and protein aggregations, redox biosensor measurements and mutant phenotype analyses to elucidate the thiol-reactive mode of action of AGXX® coating in S. aureus USA300.
The transcriptome results identified a striking overlap of the AGXX® response with the RNA-seq signature caused by the strong oxidant and microbicidal agent HOCl (Loi et al., 2018). Among the most strongly up-regulated oxidative stress specific regulons are the HypR, PerR, and QsrR regulons that are induced by H2O2, HOCl, and quinones in S. aureus (Horsburgh et al., 2001; Loi et al., 2018). The homologous HypR, PerR, and YodB regulons were also upregulated by HOCl stress in Bacillus subtilis (Chi et al., 2011). This confirms the generation of various forms of ROS, such as the less reactive H2O2, the strong oxidant hydroxyl radical and possibly quinones as the killing mode of AGXX® in S. aureus USA300. The formation of quinones is supported by the strong induction of the mhqRDE operon (Figures 2, 3). The mhqRED operon encodes for homologs of the quinone sensing MhqR repressor in B. subtilis, the phospholipase/carboxylesterase MhqE and the ring-cleavage dioxygenase MhqD, which specifically respond to quinones in B. subtilis (Töwe et al., 2007). Based on this transcriptome signature, we confirmed the generation of highly reactive ROS and quinones at the AGXX® coating in S. aureus as the main microbicidal mode of action.
In addition to ROS generation, previous studies suggested that free Ag+ ions could be released by AGXX® that might affect the metal homeostasis in bacteria (Guridi et al., 2015; Clauss-Lendzian et al., 2018). Transcriptome studies of the AGXX® response in E. faecalis showed the strong induction of the copZA operon, which also responds to elevated levels of Cu+ and Ag+ (Clauss-Lendzian et al., 2018). The activation of copper signaling and induction of the oxidative stress response by Ag+ was shown also in E. coli using a transcriptome study (McQuillan and Shaw, 2014). In S. aureus, the copZA operon is regulated by the Cu+ sensing CsoR repressor (Grossoehme et al., 2011) and is 48–56-fold induced by AGXX®. The copZA operon encodes a copper-binding chaperone CopZ and copper-translocating P-type ATPase CopA, that could be involved in transport and export of excess Ag+ to avoid silver intoxification. Moreover, we found strong induction of the CstR regulon which responds to hydrogen sulfide and nitroxyl (HNO) in S. aureus (Luebke et al., 2013, 2014; Peng et al., 2017). AGXX® might cause thiol-oxidation of the Cys residues of CstR resulting in inactivation of CstR and derepression of the cstAB operon encoding a thiosulfate sulfurtransferase and persulfide dioxygenase-sulfurtransferase (Higgins et al., 2015; Shen et al., 2015). Since CstR is a close homolog to CsoR (Grossoehme et al., 2011), inactivation of CsoR could similarly occur by thiol-oxidation of its Cu+-binding Cys residues leading to copZA derepression. In addition, our results revealed the strong induction of the Fur and Zur regulons under AGXX® that are indicative for iron and zinc starvation (Moore and Helmann, 2005; Chandrangsu et al., 2017). The Fur and Zur repressors harbor structural Zn-redox switch motifs that are susceptible for oxidation by the strong oxidant HOCl as shown in several redox proteomics studies in E. coli (Ilbert et al., 2006; Winter et al., 2008), S. aureus (Imber et al., 2018a) and Mycobacterium smegmatis (Hillion et al., 2017). Thus, oxidation of metal binding Cys ligands in Fur and Zur could trigger zinc release and the subsequent zinc and iron starvation responses.
Highly up-regulated under AGXX® stress are further the CtsR and HrcA-controlled chaperones and proteases of the protein quality control machinery involved in repair and degradation of oxidatively damaged and unfolded proteins (Frees et al., 2003, 2004, 2007, 2014). Among the Clp proteolytic and ATPase subunits, we identified clpP, clpB and the mcsAB-clpC-ctsR operon as ~50–150-fold up-regulated under AGXX® exposure. These results are in agreement with previous RNA-seq studies, showing the strong induction of clpB, clpC and ctsR genes in S. aureus and of clpB and clpE in E. faecalis (Clauss-Lendzian et al., 2018; Vaishampayan et al., 2018). The HrcA-dependent groESL operon was also highly induced in our and previous RNA-seq studies (Clauss-Lendzian et al., 2018; Vaishampayan et al., 2018), indicating that proteins might undergo thiol-oxidations and aggregation under AGXX® stress. Using a protein aggregation assay, we found higher amounts of aggregated proteins in the AGXX® treated proteome samples in agreement to the induction of the CtsR and HrcA regulons.
We have previously shown that HOCl causes >10% increased protein thiol-oxidation for 58 proteins in the redox proteome of S. aureus which includes five proteins that are modified by protein S-bacillithiolation (Chandrangsu et al., 2018; Imber et al., 2018a). The glycolytic GapDH was identified as the most abundant S-bacillithiolated protein under HOCl stress in the proteome. GapDH activity was inhibited by S-bacillithiolation to prevent its overoxidation to irreversible Cys sulfonic acid (Chandrangsu et al., 2018; Imber et al., 2018a). Here, we showed that protein S-bacillithiolation of GapDH is similar strongly increased under AGXX® treatment in S. aureus USA300. This further confirms the thiol-reactive mode of action of AGXX® in S. aureus. The increased formation of BSH mixed protein disulfides is accompanied by an oxidized shift in EBSH which was measured using our Brx-roGFP2 biosensor (Loi et al., 2017). Brx-roGFP2 was previously shown to respond specifically to low levels of BSSB in vitro and was strongly oxidized by 50–100 μM HOCl in S. aureus cells in vivo. The Brx-roGFP2 biosensor was nearly fully oxidized after 60 min of sub-lethal 4 μg/ml AGXX® treatment in S. aureus. This clearly indicates that AGXX® affects the redox balance in S. aureus.
Since BSH is involved in protein S-bacillithiolation under AGXX® stress, we performed phenotype assays of the bshA mutant to investigate if BSH protects S. aureus from ROS encountered during AGXX® exposure. Our results revealed a strong growth and survival defect of the bshA mutant under sub-lethal and lethal doses of AGXX®. Thus, the LMW thiol BSH plays an important role in protection against microbicidal ROS produced by AGXX® exposure in S. aureus. Apart from BSH, the HypR-controlled disulfide reductase MerA (Loi et al., 2018) is involved in the defense against ROS produced by AGXX®. This supports the increased formation of protein disulfides and LMW thiol disulfides as potential substrates for MerA that remain to be elucidated (Loi et al., 2018).
Altogether, we have shown here that AGXX® causes a strong thiol-specific oxidative and metal stress response as well as strong protein damage as comprehensive mode of action in S. aureus USA300. The derepression of the Fur, Zur, CsoR and CstR regulons suggests that AGXX® impairs metal ion homeostasis by induction of Fe2+- and Zn2+-starvation responses as well as export systems for toxic Ag+ ions. The oxidative mode of action in the transcriptome was supported by increased protein S-bacillithiolation and protein aggregation as well as by an oxidative shift of the BSH redox potential as revealed using the Brx-roGFP2 biosensor. In addition, we showed that BSH and the disulfide reductase MerA play major roles in the defense against ROS produced under AGXX® stress.
Thus, the ROS-producing antimicrobial AGXX® has multiple modes of actions to kill pathogens that exclude the development of resistance mechanisms. The thiol-reactive AGXX® antimicrobial coating is also environmentally friendly, sustainable and can be applied on various medical devices, wound dressings, water pipelines and in the food industry and provides a promising strategy to efficiently combat multiple antibiotic resistant pathogens.
Author Contributions
VL and HA conceived the project. VL, TB, and HA designed the experiments of the project. TB and JK contributed with RNA-seq transcriptomics analyses to the project. VL and TP performed the growth, survival, biosensor experiments and protein S-bacillithiolation assays. The AGXX® transcriptome treemap was constructed by JB. HA wrote the manuscript. All authors read and approved the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We are thankful to C. Meyer and U. Landow (Largentec GmbH, Berlin, Germany) for providing AGXX® microparticles of charge 373. This work was supported by an European Research Council (ERC) Consolidator grant (GA 615585) MYCOTHIOLOME and grants from the Deutsche Forschungsgemeinschaft, Germany (AN746/4-1 and AN746/4-2) within the SPP1710 on Thiol-based Redox switches, by the Research Training Group GRK1947 (project C01), by the SFB973 (project C08) and TR84 (project B06) to HA.
Glossary
Abbreviations
- Ag
silver
- AGXX®
antimicrobial coating of Ag+ and Ru+ ions
- BSH
bacillithiol
- BSSB
oxidized bacillithiol disulfide
- Brx
bacilliredoxins
- Brx-roGFP2
Brx fused to roGFP2
- CFU
colony-forming unit
- DTT
dithiothreitol
- EBSH
bacillithiol redox potential
- EDTA
ethylenediaminetetraacetic acid
- GapDH
glyceraldehyde-3-phosphate dehydrogenase
- H2O2
hydrogen peroxide
- HOCl
hypochloric acid
- LB
Luria Bertani
- LMW thiol
low molecular weight thiol
- MRSA
methicillin-resistant Staphylococcus aureus
- NADPH
nicotinamide adenine dinucleotide phosphate
- NaOCl
sodium hypochlorite
- NEM
N-ethylmaleimide
- NP-40
Nonidet P-40
- OD500
optical density at 500 nm
- OxD
oxidation degree of the Brx-roGFP2 biosensor
- roGFP2
redox-sensitive GFP2
- ROS
reactive oxygen species
- Ru
ruthenium.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2018.03037/full#supplementary-material
References
- Abomoelak B., Hoye E. A., Chi J., Marcus S. A., Laval F., Bannantine J. P., et al. (2009). mosR, a novel transcriptional regulator of hypoxia and virulence in Mycobacterium tuberculosis. J. Bacteriol. 191, 5941–5952. 10.1128/JB.00778-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Archer G. L. (1998). Staphylococcus aureus: a well-armed pathogen. Clin. Infect. Dis. 26, 1179–1181. 10.1086/520289 [DOI] [PubMed] [Google Scholar]
- Beavers W. N., Skaar E. P. (2016). Neutrophil-generated oxidative stress and protein damage in Staphylococcus aureus. Pathog. Dis. 74:ftw060. 10.1093/femspd/ftw060 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boucher H. W., Corey G. R. (2008). Epidemiology of methicillin-resistant Staphylococcus aureus. Clin. Infect. Dis. 46 (Suppl. 5), S344–S349. 10.1086/533590 [DOI] [PubMed] [Google Scholar]
- Chandrangsu P., Loi V. V., Antelmann H., Helmann J. D. (2018). The role of bacillithiol in Gram-positive Firmicutes. Antioxid. Redox Signal. 28, 445–462. 10.1089/ars.2017.7057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandrangsu P., Rensing C., Helmann J. D. (2017). Metal homeostasis and resistance in bacteria. Nat. Rev. Microbiol. 15, 338–350. 10.1038/nrmicro.2017.15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J.-Y., Wang L.-Y., Wu P.-W. (2010). Preparation and characterization of ruthenium films via an electroless deposition route. Thin Solid Films 518, 7245–7248. 10.1016/j.tsf.2010.04.086 [DOI] [Google Scholar]
- Chen P. R., Bae T., Williams W. A., Duguid E. M., Rice P. A., Schneewind O., et al. (2006). An oxidation-sensing mechanism is used by the global regulator MgrA in Staphylococcus aureus. Nat. Chem. Biol. 2, 591–595. 10.1038/nchembio820 [DOI] [PubMed] [Google Scholar]
- Chen P. R., Brugarolas P., He C. (2011). Redox signaling in human pathogens. Antioxid. Redox Signal. 14, 1107–1118. 10.1089/ars.2010.3374 [DOI] [PubMed] [Google Scholar]
- Chen P. R., Nishida S., Poor C. B., Cheng A., Bae T., Kuechenmeister L., et al. (2009). A new oxidative sensing and regulation pathway mediated by the MgrA homologue SarZ in Staphylococcus aureus. Mol. Microbiol. 71, 198–211. 10.1111/j.1365-2958.2008.06518.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi B. K., Busche T., Van Laer K., Bäsell K., Becher D., Clermont L., et al. (2014). Protein S-mycothiolation functions as redox-switch and thiol protection mechanism in Corynebacterium glutamicum under hypochlorite stress. Antioxid. Redox Signal. 20, 589–605. 10.1089/ars.2013.5423 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi B. K., Gronau K., Mäder U., Hessling B., Becher D., Antelmann H. (2011). S-bacillithiolation protects against hypochlorite stress in Bacillus subtilis as revealed by transcriptomics and redox proteomics. Mol. Cell. Proteomics 10:M111009506. 10.1074/mcp.M111.009506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi B. K., Roberts A. A., Huyen T. T., Bäsell K., Becher D., Albrecht D., et al. (2013). S-bacillithiolation protects conserved and essential proteins against hypochlorite stress in Firmicutes bacteria. Antioxid. Redox Signal. 18, 1273–1295. 10.1089/ars.2012.4686 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clauss-Lendzian E., Vaishampayan A., De Jong A., Landau U., Meyer C., Kok J., et al. (2018). Stress response of a clinical Enterococcus faecalis isolate subjected to a novel antimicrobial surface coating. Microbiol. Res. 207, 53–64. 10.1016/j.micres.2017.11.006 [DOI] [PubMed] [Google Scholar]
- Dooley C. T., Dore T. M., Hanson G. T., Jackson W. C., Remington S. J., Tsien R. Y. (2004). Imaging dynamic redox changes in mammalian cells with green fluorescent protein indicators. J. Biol. Chem. 279, 22284–22293. 10.1074/jbc.M312847200 [DOI] [PubMed] [Google Scholar]
- Dubbs J. M., Mongkolsuk S. (2007). Peroxiredoxins in bacterial antioxidant defense. Subcell. Biochem. 44, 143–193. 10.1007/978-1-4020-6051-9_7 [DOI] [PubMed] [Google Scholar]
- Falord M., Mäder U., Hiron A., Débarbouillé M., Msadek T. (2011). Investigation of the Staphylococcus aureus GraSR regulon reveals novel links to virulence, stress response and cell wall signal transduction pathways. PLoS ONE 6:e21323. 10.1371/journal.pone.0021323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foster T. J. (2004). The Staphylococcus aureus “superbug”. J. Clin. Invest. 114, 1693–1696. 10.1172/JCI200423825 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frees D., Chastanet A., Qazi S., Sørensen K., Hill P., Msadek T., et al. (2004). Clp ATPases are required for stress tolerance, intracellular replication and biofilm formation in Staphylococcus aureus. Mol. Microbiol. 54, 1445–1462. 10.1111/j.1365-2958.2004.04368.x [DOI] [PubMed] [Google Scholar]
- Frees D., Gerth U., Ingmer H. (2014). Clp chaperones and proteases are central in stress survival, virulence and antibiotic resistance of Staphylococcus aureus. Int. J. Med. Microbiol. 304, 142–149. 10.1016/j.ijmm.2013.11.009 [DOI] [PubMed] [Google Scholar]
- Frees D., Qazi S. N., Hill P. J., Ingmer H. (2003). Alternative roles of ClpX and ClpP in Staphylococcus aureus stress tolerance and virulence. Mol. Microbiol. 48, 1565–1578. 10.1046/j.1365-2958.2003.03524.x [DOI] [PubMed] [Google Scholar]
- Frees D., Savijoki K., Varmanen P., Ingmer H. (2007). Clp ATPases and ClpP proteolytic complexes regulate vital biological processes in low GC, Gram-positive bacteria. Mol. Microbiol. 63, 1285–1295. 10.1111/j.1365-2958.2007.05598.x [DOI] [PubMed] [Google Scholar]
- Grass G., Rensing C., Solioz M. (2011). Metallic copper as an antimicrobial surface. Appl. Environ. Microbiol. 77, 1541–1547. 10.1128/AEM.02766-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groitl B., Dahl J. U., Schroeder J. W., Jakob U. (2017). Pseudomonas aeruginosa defense systems against microbicidal oxidants. Mol. Microbiol. 106, 335–350. 10.1111/mmi.13768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grossoehme N., Kehl-Fie T. E., Ma Z., Adams K. W., Cowart D. M., Scott R. A., et al. (2011). Control of copper resistance and inorganic sulfur metabolism by paralogous regulators in Staphylococcus aureus. J. Biol. Chem. 286, 13522–13531. 10.1074/jbc.M111.220012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta A., Matsui K., Lo J. F., Silver S. (1999). Molecular basis for resistance to silver cations in Salmonella. Nat. Med. 5, 183–188. 10.1038/5545 [DOI] [PubMed] [Google Scholar]
- Guridi A., Diederich A. K., Aguila-Arcos S., Garcia-Moreno M., Blasi R., Broszat M., et al. (2015). New antimicrobial contact catalyst killing antibiotic resistant clinical and waterborne pathogens. Mater. Sci. Eng. C Mater. Biol. Appl. 50, 1–11. 10.1016/j.msec.2015.01.080 [DOI] [PubMed] [Google Scholar]
- Heiss A., Freisinger B., Held-Föhn E. (2017). Enhanced antibacterial activity of silver-ruthenium coated hollow microparticles. Biointerphases 12:05G608. 10.1116/1.5003803 [DOI] [PubMed] [Google Scholar]
- Higgins K. A., Peng H., Luebke J. L., Chang F. M., Giedroc D. P. (2015). Conformational analysis and chemical reactivity of the multidomain sulfurtransferase, Staphylococcus aureus CstA. Biochemistry 54, 2385–2398. 10.1021/acs.biochem.5b00056 [DOI] [PubMed] [Google Scholar]
- Highlander S. K., Hultén K. G., Qin X., Jiang H., Yerrapragada S., Mason E. O., Jr., et al. (2007). Subtle genetic changes enhance virulence of methicillin resistant and sensitive Staphylococcus aureus. BMC Microbiol. 7:99. 10.1186/1471-2180-7-99 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hilker R., Stadermann K. B., Schwengers O., Anisiforov E., Jaenicke S., Weisshaar B., et al. (2016). ReadXplorer 2-detailed read mapping analysis and visualization from one single source. Bioinformatics 32, 3702–3708. 10.1093/bioinformatics/btw541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hillion M., Antelmann H. (2015). Thiol-based redox switches in prokaryotes. Biol. Chem. 396, 415–444. 10.1515/hsz-2015-0102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hillion M., Bernhardt J., Busche T., Rossius M., Maaß S., Becher D., et al. (2017). Monitoring global protein thiol-oxidation and protein S-mycothiolation in Mycobacterium smegmatis under hypochlorite stress. Sci. Rep. 7:1195. 10.1038/s41598-017-01179-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horsburgh M. J., Clements M. O., Crossley H., Ingham E., Foster S. J. (2001). PerR controls oxidative stress resistance and iron storage proteins and is required for virulence in Staphylococcus aureus. Infect. Immun. 69, 3744–3754. 10.1128/IAI.69.6.3744-3754.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ilbert M., Graf P. C., Jakob U. (2006). Zinc center as redox switch–new function for an old motif. Antioxid. Redox Signal. 8, 835–846. 10.1089/ars.2006.8.835 [DOI] [PubMed] [Google Scholar]
- Imber M., Huyen N. T. T., Pietrzyk-Brzezinska A. J., Loi V. V., Hillion M., Bernhardt J., et al. (2018a). Protein S-Bacillithiolation functions in thiol protection and redox regulation of the glyceraldehyde-3-phosphate dehydrogenase Gap in Staphylococcus aureus under hypochlorite stress. Antioxid. Redox Signal. 28, 410–430. 10.1089/ars.2016.6897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imber M., Loi V. V., Reznikov S., Fritsch V. N., Pietrzyk-Brzezinska A. J., Prehn J., et al. (2018b). The aldehyde dehydrogenase AldA contributes to the hypochlorite defense and is redox-controlled by protein S-bacillithiolation in Staphylococcus aureus. Redox Biol. 15, 557–568. 10.1016/j.redox.2018.02.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imber M., Pietrzyk-Brzezinska A. J., Antelmann H. (2018c). Redox regulation by reversible protein S-thiolation in Gram-positive bacteria. Redox Biol. 20, 130–145. 10.1016/j.redox.2018.08.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji C. J., Kim J. H., Won Y. B., Lee Y. E., Choi T. W., Ju S. Y., et al. (2015). Staphylococcus aureus PerR is a hypersensitive hydrogen peroxide sensor using iron-mediated histidine oxidation. J. Biol. Chem. 290, 20374–20386. 10.1074/jbc.M115.664961 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B., Salzberg S. L. (2012). Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359. 10.1038/nmeth.1923 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lansdown A. B. (2010). A pharmacological and toxicological profile of silver as an antimicrobial agent in medical devices. Adv. Pharmacol. Sci. 2010:910686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J. W., Helmann J. D. (2006). The PerR transcription factor senses H2O2 by metal-catalysed histidine oxidation. Nature 440, 363–367. 10.1038/nature04537 [DOI] [PubMed] [Google Scholar]
- Li H., Handsaker B., Wysoker A., Fennell T., Ruan J., Homer N., et al. (2009). The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079. 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livermore D. M. (2000). Antibiotic resistance in staphylococci. Int. J. Antimicrob. Agents 16 (Suppl. 1), S3–10. 10.1016/S0924-8579(00)00299-5 [DOI] [PubMed] [Google Scholar]
- Loi V. V., Busche T., Tedin K., Bernhardt J., Wollenhaupt J., Huyen N. T. T., et al. (2018). Redox-sensing under hypochlorite stress and infection conditions by the Rrf2-family repressor HypR in Staphylococcus aureus. Antioxid. Redox Signal. 29, 615–636. 10.1089/ars.2017.7354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loi V. V., Harms M., Müller M., Huyen N. T. T., Hamilton C. J., Hochgräfe F., et al. (2017). Real-time imaging of the bacillithiol redox potential in the human pathogen Staphylococcus aureus using a genetically encoded bacilliredoxin-fused redox biosensor. Antioxid. Redox Signal. 26, 835–848. 10.1089/ars.2016.6733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loi V. V., Rossius M., Antelmann H. (2015). Redox regulation by reversible protein S-thiolation in bacteria. Front. Microbiol. 6:187. 10.3389/fmicb.2015.00187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love M. I., Huber W., Anders S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15:550. 10.1186/s13059-014-0550-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowy F. D. (1998). Staphylococcus aureus infections. N. Engl. J. Med. 339, 520–532. 10.1056/NEJM199808203390806 [DOI] [PubMed] [Google Scholar]
- Luebke J. L., Arnold R. J., Giedroc D. P. (2013). Selenite and tellurite form mixed seleno- and tellurotrisulfides with CstR from Staphylococcus aureus. Metallomics 5, 335–342. 10.1039/c3mt20205d [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luebke J. L., Shen J., Bruce K. E., Kehl-Fie T. E., Peng H., Skaar E. P., et al. (2014). The CsoR-like sulfurtransferase repressor (CstR) is a persulfide sensor in Staphylococcus aureus. Mol. Microbiol. 94, 1343–1360. 10.1111/mmi.12835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mäder U., Nicolas P., Depke M., Pané-Farré J., Debarbouille M., Van Der Kooi-Pol M. M., et al. (2016). Staphylococcus aureus transcriptome architecture: from laboratory to infection-mimicking conditions. PLoS Genet. 12:e1005962. 10.1371/journal.pgen.1005962 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maillard J. Y., Hartemann P. (2013). Silver as an antimicrobial: facts and gaps in knowledge. Crit. Rev. Microbiol. 39, 373–383. 10.3109/1040841X.2012.713323 [DOI] [PubMed] [Google Scholar]
- McQuillan J. S., Shaw A. M. (2014). Differential gene regulation in the Ag nanoparticle and Ag(+)-induced silver stress response in Escherichia coli: a full transcriptomic profile. Nanotoxicology 8 (Suppl. 1), 177–184. 10.3109/17435390.2013.870243 [DOI] [PubMed] [Google Scholar]
- Mehlan H., Schüler F., Weiss S., Schuler J., Fuchs S., Riedel K., et al. (2013). Data visualization in environmental proteomics. Proteomics 13, 2805–2821. 10.1002/pmic.201300167 [DOI] [PubMed] [Google Scholar]
- Moore C. M., Helmann J. D. (2005). Metal ion homeostasis in Bacillus subtilis. Curr. Opin. Microbiol. 8, 188–195. 10.1016/j.mib.2005.02.007 [DOI] [PubMed] [Google Scholar]
- Newton G. L., Fahey R. C., Rawat M. (2012). Detoxification of toxins by bacillithiol in Staphylococcus aureus. Microbiology 158, 1117–1126. 10.1099/mic.0.055715-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novick R. P. (2003). Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol. Microbiol. 48, 1429–1449. 10.1046/j.1365-2958.2003.03526.x [DOI] [PubMed] [Google Scholar]
- Padiadpu J., Baloni P., Anand K., Munshi M., Thakur C., Mohan A., et al. (2016). Identifying and tackling emergent vulnerability in drug-resistant Mycobacteria. ACS Infect Dis. 2, 592–607. 10.1021/acsinfecdis.6b00004 [DOI] [PubMed] [Google Scholar]
- Pendleton J. N., Gorman S. P., Gilmore B. F. (2013). Clinical relevance of the ESKAPE pathogens. Expert Rev. Anti Infect. Ther. 11, 297–308. 10.1586/eri.13.12 [DOI] [PubMed] [Google Scholar]
- Peng H., Shen J., Edmonds K. A., Luebke J. L., Hickey A. K., Palmer L. D., et al. (2017). Sulfide homeostasis and nitroxyl intersect via formation of reactive sulfur species in Staphylococcus aureus. mSphere 2, e00082–17. 10.1128/mSphere.00082-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pförtner H., Burian M. S., Michalik S., Depke M., Hildebrandt P., Dhople V. M., et al. (2014). Activation of the alternative sigma factor SigB of Staphylococcus aureus following internalization by epithelial cells - an in vivo proteomics perspective. Int. J. Med. Microbiol. 304, 177–187. 10.1016/j.ijmm.2013.11.014 [DOI] [PubMed] [Google Scholar]
- Pinochet-Barros A., Helmann J. D. (2018). Redox sensing by Fe2+ in bacterial fur family metalloregulators. Antioxid. Redox Signal. 29, 1858–1871. 10.1089/ars.2017.7359 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poor C. B., Chen P. R., Duguid E., Rice P. A., He C. (2009). Crystal structures of the reduced, sulfenic acid, and mixed disulfide forms of SarZ, a redox active global regulator in Staphylococcus aureus. J. Biol. Chem. 284, 23517–23524. 10.1074/jbc.M109.015826 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Posada A. C., Kolar S. L., Dusi R. G., Francois P., Roberts A. A., Hamilton C. J., et al. (2014). Importance of bacillithiol in the oxidative stress response of Staphylococcus aureus. Infect. Immun. 82, 316–332. 10.1128/IAI.01074-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pöther D. C., Gierok P., Harms M., Mostertz J., Hochgräfe F., Antelmann H., et al. (2013). Distribution and infection-related functions of bacillithiol in Staphylococcus aureus. Int. J. Med. Microbiol. 303, 114–123. 10.1016/j.ijmm.2013.01.003 [DOI] [PubMed] [Google Scholar]
- Queck S. Y., Jameson-Lee M., Villaruz A. E., Bach T. H., Khan B. A., Sturdevant D. E., et al. (2008). RNAIII-independent target gene control by the agr quorum-sensing system: insight into the evolution of virulence regulation in Staphylococcus aureus. Mol. Cell 32, 150–158. 10.1016/j.molcel.2008.08.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen J., Keithly M. E., Armstrong R. N., Higgins K. A., Edmonds K. A., Giedroc D. P. (2015). Staphylococcus aureus CstB is a novel multidomain persulfide dioxygenase-sulfurtransferase involved in hydrogen sulfide detoxification. Biochemistry 54, 4542–4554. 10.1021/acs.biochem.5b00584 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun F., Ding Y., Ji Q., Liang Z., Deng X., Wong C. C., et al. (2012). Protein cysteine phosphorylation of SarA/MgrA family transcriptional regulators mediates bacterial virulence and antibiotic resistance. Proc. Natl. Acad. Sci. U.S.A. 109, 15461–15466. 10.1073/pnas.1205952109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tam Le T., Eymann C., Albrecht D., Sietmann R., Schauer F., Hecker M., et al. (2006). Differential gene expression in response to phenol and catechol reveals different metabolic activities for the degradation of aromatic compounds in Bacillus subtilis. Environ. Microbiol. 8, 1408–1427. 10.1111/j.1462-2920.2006.01034.x [DOI] [PubMed] [Google Scholar]
- Tomoyasu T., Arsene F., Ogura T., Bukau B. (2001). The C terminus of sigma(32) is not essential for degradation by FtsH. J. Bacteriol. 183, 5911–5917. 10.1128/JB.183.20.5911-5917.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Töwe S., Leelakriangsak M., Kobayashi K., Van Duy N., Hecker M., Zuber P., et al. (2007). The MarR-type repressor MhqR (YkvE) regulates multiple dioxygenases/glyoxalases and an azoreductase which confer resistance to 2-methylhydroquinone and catechol in Bacillus subtilis. Mol. Microbiol. 66, 40–54. 10.1111/j.1365-2958.2007.05891.x [DOI] [PubMed] [Google Scholar]
- Tung Q. N., Linzner N., Loi V. V., Antelmann H. (2018). Application of genetically encoded redox biosensors to measure dynamic changes in the glutathione, bacillithiol and mycothiol redox potentials in pathogenic bacteria. Free Radic. Biol. Med. 128, 84–96. 10.1016/j.freeradbiomed.2018.02.018 [DOI] [PubMed] [Google Scholar]
- Vaishampayan A., De Jong A., Wight D. J., Kok J., Grohmann E. (2018). A novel antimicrobial coating represses biofilm and virulence-related genes in methicillin-resistant Staphylococcus aureus. Front. Microbiol. 9:221. 10.3389/fmicb.2018.00221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villapún V. M., Dover L. G., Cross A., González S. (2016). Antibacterial metallic touch surfaces. Materials 9:736. 10.3390/ma9090736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wetzstein M., Völker U., Dedio J., Löbau S., Zuber U., Schiesswohl M., et al. (1992). Cloning, sequencing, and molecular analysis of the dnaK locus from Bacillus subtilis. J. Bacteriol. 174, 3300–3310. 10.1128/jb.174.10.3300-3310.1992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winter J., Ilbert M., Graf P. C., Ozcelik D., Jakob U. (2008). Bleach activates a redox-regulated chaperone by oxidative protein unfolding. Cell 135, 691–701. 10.1016/j.cell.2008.09.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winterbourn C. C., Kettle A. J. (2013). Redox reactions and microbial killing in the neutrophil phagosome. Antioxid. Redox Signal. 18, 642–660. 10.1089/ars.2012.4827 [DOI] [PubMed] [Google Scholar]
- Winterbourn C. C., Kettle A. J., Hampton M. B. (2016). Reactive oxygen species and neutrophil function. Annu. Rev. Biochem. 85, 765–792. 10.1146/annurev-biochem-060815-014442 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.