Abstract
DNA-PKcs deficiency has been studied in numerous animal models and cell culture systems. In previous studies of kinase inactivating mutations in cell culture systems, ablation of DNA-PK’s catalytic activity results in a cell phenotype that is virtually indistinguishable from that ascribed to complete loss of the enzyme. However, a recent compelling study demonstrates a remarkably more severe phenotype in mice harboring a targeted disruption of DNA-PK’s ATP binding site as compared to DNA-PKcs deficient mice. Here we investigate the mechanism for these divergent results. We find that kinase inactivating DNA-PKcs mutants markedly radiosensitize immortalized DNA-PKcs deficient cells, but have no substantial effects on transformed DNA-PKcs deficient cells. Since the non-homologous end joining mechanism likely functions similarly in all of these cell strains, it seems unlikely that kinase inactive DNA-PK could impair the end joining mechanism in some cell types, but not in others. In fact, we observed no significant differences in either episomal or chromosomal end joining assays in cells expressing kinase inactivated DNA-PKcs versus no DNA-PKcs. Several potential explanations could explain these data including a non-catalytic role for DNA-PKcs in promoting cell death, or alteration of gene expression by loss of DNA-PKcs as opposed to inhibition of its catalytic activity.
Finally, controversy exists as to whether DNA-PKcs autophosphorylates or is the target of other PIKKs; we present data demonstrating that DNA-PK primarily autophosphorylates.
Introduction
DNA-PKcs deficiency has been exhaustively studied in five different animal models, and in numerous cell culture systems derived from several different species. In most of these studies, functionally null alleles of DNA-PKcs were examined [1-10]. Although three spontaneous, null DNA-PKcs mutations have been characterized in animals, none have been observed in humans. In all of these animal models, the predominant phenotype is radiosensitive severe combined immunodeficiency (RS-SCID). Recently, two hypomorphic human DNA-PKcs mutations were reported [11, 12]; one of these mutations severely impairs DNA-PK’s catalytic function and expression [13]. In the patient with this mutation, a fatal neurologic phenotype occurred providing evidence for additional function(s) for DNA-PK in humans.
A number of studies (including data presented here) address potential non-catalytic functions of DNA-PK utilizing kinase inactivating mutants of DNA-PKcs in cell culture models [14-19]. In general, cell lines expressing kinase inactive DNA-PKcs mutants impart cellular phenotypes that are extremely similar, if not indistinguishable from those observed with complete DNA-PKcs deficiency. Thus, the compelling study from Zha and colleagues, describing a severe growth retardation phenotype in mice expressing a catalytically inactive DNA-PKcs mutant that suggests that catalytically inactive protein suppresses end joining was of considerable interest [20]. Here we demonstrate in cell culture models that none of three different catalytically inactive mutants of either human or murine DNA-PKcs tested potentiate cellular radiosensitivity or suppress end joining in transformed DNA-PKcs deficient human or rodent cell strains; thus, the discrepancies between cell culture models and the mouse model cannot be explained by species-specific differences in the DNA-PKcs enzyme itself. In contrast, expression of the DNA-PKcs kinase inactive mutants in DNA-PKcs deficient, non-transformed, immortalized DNA-PKcs deficient mouse fibroblasts results in markedly increased radio-sensitization as compared to cells expressing no DNA-PKcs. Although non-transformed SCID fibroblasts expressing kinase inactive DNA-PKcs are more radiosensitive than cells lacking DNA-PKcs, DNA end joining (in both episomal and chromosomal assays) is similarly impaired in both, displaying characteristics of alternative non-homologous end joining (a-NHEJ). A potential straightforward explanation for these data is the fact that loss of DNA-PKcs [but not gene targeted or chemical ablation of its catalytic activity] results in substantial loss of ATM expression [21-24]. For example, the severe phenotype observed with DNA ligase IV deficiency can be attenuated by deletion of just one ATM allele [25]. We suggest that retention of ATM expression in mice with targeted disruption of DNA-PK enzymatic activity may explain (at least in part) differences between DNA-PKcs deficient mice versus mice with a DNA-PK kinase inactivating mutation; diminished ATM expression in DNA-PKcs deficient mice may attenuate the classical non-homologous end joining (c-NHEJ) defective phenotype, just as targeted deletion of ATM partially rescues embryonic lethality in ligase IV deficient mice.
However, in cell culture experiments, ATM expression is not restored in DNA-PKcs deficient cells by either stable or transient complementation with either wild type or mutant DNA-PKcs. Thus, the increased radiosensitivity in non-transformed cell strains expressing catalytically inactive DNA-PKcs is not dependent on ATM expression levels and must result from another mechanism. Several potential explanations could explain these data including a non-catalytic role for DNA-PKcs in promoting cell death, cell type or species-specific differences, or alteration of gene expression by loss of DNA-PKcs as opposed to inhibition of its catalytic activity.
Finally, discrepancies also exist in the literature as to whether DNA-PK primarily autophosphorylates (or is phosphorylated by ATM and/or ATR) on sites within two major phosphorylation clusters. Previously we and others have concluded that phosphorylation within the ABCDE and PQR clusters is largely autophosphorylation, although other kinases may contribute to a substantially lesser extent [26-28]. In contrast, other studies have concluded that ATM contributes to ABCDE phosphorylation [29] or exclusively [20] phosphorylates the ABCDE cluster, and that ATR primarily phosphorylates this cluster during replication stress [30]. To clarify how DNA-PKcs is regulated, we examine and validate a number of commercial antibodies. Many commercially available and widely utilized reagents for detection of phosphorylated residues within DNA-PKcs do not recognize the appropriate sites, and in some cases detect damage induced phosphorylation of other proteins, but not DNA-PKcs. Using antibodies validated with specific phospho-site ablated DNA-PKcs mutants, we show that DSB-induced phosphorylations within both the ABCDE and PQR clusters occur primarily by autophosphorylation, and do not require the ATM kinase.
Methods
Plasmids, cell culture and cell strains.
Expression plasmids for human [31, 32] and murine [33] wild type and kinase inactivating mutants have been described previously. 293T, and Sf19 cells were cultured in Dulbecco’s Modified Eagle Medium (Life Technologies) supplemented with 10% fetal bovine serum (Atlanta Biologicals, GA), 2 mM L-glutamine, 0.1 mM non-essential amino acids, 1 mM sodium pyruvate, 100 U/ml penicillin, 100 µg/ml streptomycin (Life Technologies) and 10 μg/ml ciprofloxacin. V3 cells were cultured in alpha-MEM with the same supplements. Cells were maintained at 37°C with 5% CO2. Methods to derive V3 and Sf19 transfectants have been previously described [15, 31]. Briefly, each parental cell strain was co-transfected with the indicated DNA-PKcs expression plasmid and the pSuper-puro plasmid, to confer puromycin resistance, using FuGENE 6 (Roche) according to the manufacturer’s instructions. Independent, stable transfectants were selected and maintained in complete medium containing 10 μg/ml puromycin. At least two independently derived clones expressing each mutant were studied.
Immunoblot Analyses.
Immunoblotting was performed as described [34]. Antibodies used in this study are rabbit polyclonal anti-XRCC4 (Abcam, 213729), rabbit anti-ATM (Abcam, 2C1), rabbit anti-Tel2 (Abcam, 15975), rabbit anti-Tti2 (Bethyl, A303-476A), rabbit anti-Tti1 (Bethyl, A303-451A), mouse anti-p53 (Santa Cruz, sc-98), and mouse anti-phospho-γh2AX (Millipore, 05-636). The DNA-PKcs antibody (42-27) was the generous gift of Tim Carter. DNA-PKcs phospho-specific antibodies utilized in this study include:
anti-phospho-T2609: Abcam 18356, Abcam 97611, Abcam 4194, and Sigma SAB4300137. In addition, a rabbit anti-phosphoT2609 reagent was the generous gift of Dale Ramsden.
anti-phospho-S2056: Abcam 18192, Abcam 124918.
anti-phospho-S2612: Abcam 133441.
Cas9-mediated gene disruption.
Cas9-targeted gene disruption was performed using methods similar to those reported by Mali et al. [35]. Briefly, a gRNA specific for a PAM site adjacent to K3752 was synthesized as a 455 bp fragment (Integrated DNA Technologies). The synthesized fragment was cloned into pCR2.1 using a TOPO TA cloning kit according to the manufacturers’ instructions (Life Technologies). Cells were transfected with 1 μg gRNA plasmid, 1 μg Cas9 expression plasmid (Addgene), and 0.2 μg of pSuper-Puro (to confer puromycin resistance) in 200ul Optimem (Life Technologies) and 4ul PEI polyethylenimine (PEI, 1 ug/ml, Polysciences). To promote homology directed repair (to generate the K3752R mutation) a 120bp oligonucleotide encoding the mutation as well as a silent mutation generating a novel restriction site (SacII) was co-transfected. 48 hours after transfection, cells were replated at cloning densities in media containing puromycin (1ug/ml). Puromycin was removed after 72 hours. Isolated clones were selected, and DNA isolated with DNAzol (Sigma) according to the manufacturer’s protocol. Restriction digestion of PCR products was utilized to detect the K3752>A mutation. Western blotting was used to confirm expression. In all cases, mutations were confirmed by TA cloning (Invitrogen) and sequencing of PCR products. The sequence of the 19 mer specific for the K3752 PAM site is: ACCCTTTCCTGGTGAAGGG
The HDR oligonucleotide and oligonucleotides utilized for TA cloning is: K3752>R HDR:
GAAGGCCCAAGCGCATCATCATCCGTGGCCATGACGAGAGGGAACACCCTTTCCTGGTCCGCGGTGGCGAGGACCTGCGG
CAGGACCAGCGCGTGGAGCAGCTCTTCCAGGTCATGAATG
5’K3752: TGTGCCCTGAGATGACATTCTGATT
3’K3752: ACCTGGAGGTCATGGGCACAACGCT
Survival assays.
Clonogenic survival assays were performed for V3 cells and SF19 cells. Briefly, a hundred cells were plated for each transfectant into complete medium containing the indicated dose of zeocin in 60 mm diameter tissue culture dishes. After 7 to 10 days, cell colonies were stained with 1% (w/v) crystal violet in ethanol to measure relative survival. MTT staining was performed to assess cell viability for 293T cells. 30,000 to 50,000 cells were plated in each well of a 24-well plate, containing medium with varying concentrations of zeocin. After 5 to 7 days of zeocin treatment, cells were treated with 1 mg/ml MTT (Sigma) solution for 1 hr. Medium containing MTT was then removed and formazan crystals thus produced were solubilized in acidic isopropanol. Absorbance was read at 570 nm to determine relative survival.
Episomal end joining assays
The fluorescent ISce-1 substrate has been described [33] and is derived from pECFP-N1 (Clontech). Briefly, extrachromosomal fluorescent joining assays were performed on cells plated at 20-40% confluency into 24-well plates in complete medium. Cells were transfected with 0.125 μg substrate, 0.25 I-Sce1 per well using polyethylenimine (PEI, 1 ug/mL, Polysciences) at 2 μL/1 μg DNA. Cells were harvested 72 hours after transfection and analyzed for CFP and RFP expression by flow cytometry. The percentage of recombination was calculated as the percentage of live cells expressing CFP divided by the percentage expressing RFP. Data presented represents at least three independent experiments, which each includes triplicate transfections.
To assess hairpin joining of plasmid substrates, the target site for the TelN nuclease was cloned into the Nhe1 and BamH1 sites of peCFP-N1 using the following oligonucleotides:
Top:CTAGTATCAGCACACAATTGCCCATTATACGCGCGTATAATGGACTATTGTGTGCTGATA
Bottom:GATCTATCAGCACACAATAGTCCATTATACGCGCGTATAATGGGCAATTGTGTGCTGATA
To compare joining of hairpin termini, 2ug of TelN restricted TelN substrate and 2ug dsRED were co-transfected into each cell type. CFP/RFP fluorescence provides an assessment of hairpin joining.
Chromosomal end joining assays
For chromosomal joining assays, the plasmid pSpCas9(BB)-2A-Puro (Addgene), which provides co-expression of a gRNA, Cas9, and the puromycin resistance gene was utilized. gRNAs specific for two sites within the gene encoding murine activation induced cytidine deaminase were utilized. The 20mers inserted into pSpCas9(BB)-2A-Puro were:
Target 1: TTGTCATCGACTTCGTACAA
Target 2: GGATTTTGAAAGCAACCTCC
SF19 cells expressing WT, D3922>A or vector alone were transfected with 2ug of the two gRNA plasmids and .5ug pcDNA6 to confer blasticidin resistance. Cells were plated onto blasticidin after 48 hours. Cells were harvested 96 hours after transfection and genomic DNA extracted using DNAzol (Thermo-Fisher) according to the manufacturer's recommendations. PCR amplification was performed using Go-Taq (Thermo-Fisher) with the following amplification primers:
Primer 1: TTTGCTAACCTCCCCTCAGGTG
Primer 2: TTTGCTAACCTCCCCTCAGGTG
Products consistent with deletions between the target sites were gel purified, TA cloned and sequenced.
Results
ATM expression is substantially reduced in cultured cells ablated for DNA-PKcs.
It has been shown that ablation of DNA-PKcs (but not chemical inhibition of its enzymatic activity) results in loss of ATM expression in both DNA-PKcs deficient animals and DNA-PKcs deficient cell strains [21-24]. A reasonable explanation for the more severe phenotype in DNA-PKcskd mice is that ATM expression is retained whereas ATM expression is reduced in SCID mice; thus, we first considered that loss of ATM attenuates the severity of DNA-PKcs deficiency. To address whether DNA-PKcs ablation universally suppresses ATM expression, ATM expression was examined in a variety of cell strains. As can be seen, ATM is markedly lost in untransformed fibroblasts from both SCID mice and a human patient with a severe (although not completely null) DNA-PKcs defect [12] (Fig. 1A). The magnitude of ATM loss was assessed in three different DNA-PKcs deficient human cell strains. DNA-PKcs deficient MO59J cells express ~4 fold less ATM than MO59K, DNA-PKcs proficient cells (Fig. 1B), whereas loss of ATM expression in DNA-PKcs deficient HCT116 and 293T cells strains is substantially more than 4 fold. Loss of ATM expression is specific for DNA-PKcs ablation; 293T cells deficient in XRCC4, XLF, or PAXX retain wild type levels of ATM (Fig. 1C). Moreover, loss of ATM does not impact the expression level of DNA-PKcs (Fig.1C). We also performed a targeted kinase inactivating mutation of the DNA-PKcs gene (K3752>R) in human HEK293T cells utilizing a Crispr/Cas9 strategy screened initially by PCR (Sup. Fig.1), and then assessed ATM expression. As can be seen, whereas loss of DNA-PKcs in two different clones results in dramatic reduction of ATM expression, two clones with the targeted kinase inactivating mutation maintain wild type levels of ATM levels (Fig. 1D). De Lange and colleagues have reported that a chaperone complex (TTT complex) including Tel2, Tti-1, and Tti-2 is required for stability of PIKK kinases [36]. However, expression levels of the three components of the TTT complex are expressed similarly in DNA-PKcs deficient and wild type 293T cells (Fig.1D). We have shown previously that restoring DNA-PKcs expression in DNA-PKcs deficient V3 cells does not restore ATM expression [15]. We also attempted to reverse the loss of ATM expression in DNA-PKcs deficient 293T cells by re-introducing DNA-PKcs. Although it is impossible to completely replace the high levels of DNA-PKcs in human cells by transient transfection, ATM levels are not altered after 72 hours of transient DNA-PKcs expression (Fig 1E). Similar results were obtained using a variety of DNA-PKcs expression constructs (data not shown). The inability to restore ATM levels is not due to an intrinsic instability of ATM in DNA-PKcs deficient cells because it is expressed ectopically, equivalently in DNA-PKcs deficient and proficient cells, either transiently (Fig 1F), or in stable transfectants (not shown). In sum, loss of DNA-PKcs consistently reduces ATM expression in a variety of cell types from both rodents and humans.
Figure 1. ATM expression is substantially reduced in cultured cells ablated for DNA-PKcs.
(A). DNA-PKcs and ATM protein expression levels were examined by western blotting of whole cell extracts obtained from wild type or SCID ear fibroblasts (left panel), and patient derived fibroblasts (right panel). A non-specific band detected by the ATM antibody (left panel) or Ku86 (right panel) serve as loading controls. (B) ATM protein expression levels were assessed in wild type and DNA-PKcs deficient 293T cells, HCT116 cells, and MO59K/J cells. Protein levels from indicated amounts of whole cell extract were assessed by immunoblotting. A non-specific band detected by the ATM antibody serves as a loading control. (C) ATM expression was assessed from 293T cells ablated for other c-NHEH components as indicated.(D) Expression of the components of the TTT complex were assessed by immunoblotting in wild type 293T cells, or in cells lacking DNA-PKcs, or harboring a kinase inactivating mutation of DNA-PKcs as indicated. White line indicate removal of lanes not presented; figure is derived from a single blot. (E) DNA-PKcs and ATM expression in wild type or DNA-PKcs deficient 293T cells 72 hours after transfection with a DNA-PKcs expression construct or vector alone. A non-specific band detected by the ATM antibody serves as a loading control. (F) GFP and ATM expression levels in wild type or DNA-PKcs deficient 293T cells transfected with either or both GFP-DNA-PKcs or GFP-ATM as indicated.
Human and murine DNA-PKcs complement radiomimetic drug sensitivity in DNA-PKcs deficient hamster cells to a similar extent; kinase-inactivating mutations impart similar radiosensitivity as DNA-PKcs deficiency in both V3 cells and 293T cells.
The current literature is muddled with conflicting reports that ascertain: 1) the necessity for DNA-PKcs for VDJ signal joining [2, 5, 14, 31, 37, 38]; 2) the requirement for DNA-PK’s catalytic activity for VDJ recombination [31, 39]; 3) the ability of catalytically inactive DNA-PKcs to inhibit other DSBR pathways and the more specific repair of VDJ recombination intermediates [18, 19]; and whether phosphorylation of DNA-PKcs is predominately autophosphorylation or is primarily mediated by ATM and/or ATR [26, 29, 30]. Discrepancies in conclusions from these many well-designed studies likely represent differences in experimental systems, and perhaps species-specific, or cell-type specific differences that might be accounted for by expression of compensatory factors, or species-specific differences in DNA-PK itself.
Our recent studies provide strong evidence that differential loss of ATM expression dramatically alters the proficiency of signal joining in DNA-PKcs deficient cells providing a good explanation for well-documented discrepancies in signal joining in different DNA-PKcs deficient animals [33, 40]. These same publications extend previous conclusions that DNA-PK’s catalytic activity is required for VDJ joining. Here we focus on clarifying discordant results of studies utilizing kinase-inactivating mutations of DNA-PKcs.
We first assessed whether the discrepancies could be ascribed to functional differences between human and mouse DNA-PKcs by determining whether murine and human DNA-PKcs proteins function equivalently in cellular assays. Previous cellular studies have relied heavily on the DNA-PKcs deficient CHO cell strain, V3. The DNA-PKcs defect in V3 cells results from a premature termination codon in one allele at amino acid 4024; this is the only DNA-PKcs allele expressed in this cell strain. Although this mutation is C-terminal to the kinase domain, there is no detectable DNA-PKcs expression or enzymatic activity in V3 cells [41]. Expression constructs encoding both wild type and catalytically inactive human and murine DNA-PKcs were prepared. We have studied numerous kinase inactivating human DNA-PKcs mutations in V3 cells that consistently behave similar to vector only controls [15, 31, 32, 42]. Here, two different kinase-inactivating mutants in human DNA-PKcs (K3752>R and D3921>A) and one kinase-inactivating mutant in murine DNA-PKcs (D3922>A) were tested (Fig. 2A). Several investigators have extensively studied these two mutations, and previous studies have concluded that both behave equivalently in cellular assays [15, 17, 31]. At least two different clones for each mutant were tested; DNA-PKcs expression levels in these clones varied substantially. The zeocin sensitive phenotype of V3 cells is similarly and substantially reversed by both human and murine wild type DNA-PKcs, even in the cell clone expressing low levels of DNA-PKcs (hWT59). As expected, none of the enzymatically inactive DNA-PKcs mutants reverse the zeocin sensitivity of V3 cells; however, none of the enzymatically inactive mutants potentiate zeocin sensitivity of V3 cells, even cell clones expressing high levels of DNA-PKcs (mD3922A-13, mD3922A-19, hK3752R-5). In total, we have previously studied radioresistance in V3 cells with three different DNA-PKcs mutations that totally abolish catalytic activity [31, 32], and three mutations that substantially abolish catalytic activity [12, 15, 42]. All behave indistinguishably from each other and to vector only controls with regards to radiosensitivity. As in our previous study [15], ATM levels vary in the V3 DNA-PKcs clones studied here; however, vector only or kinase inactive expressing clones that have markedly different ATM levels (for example D3922>A-13 and K3752>R-5) do not display differential sensitivity to zeocin sensitivity.
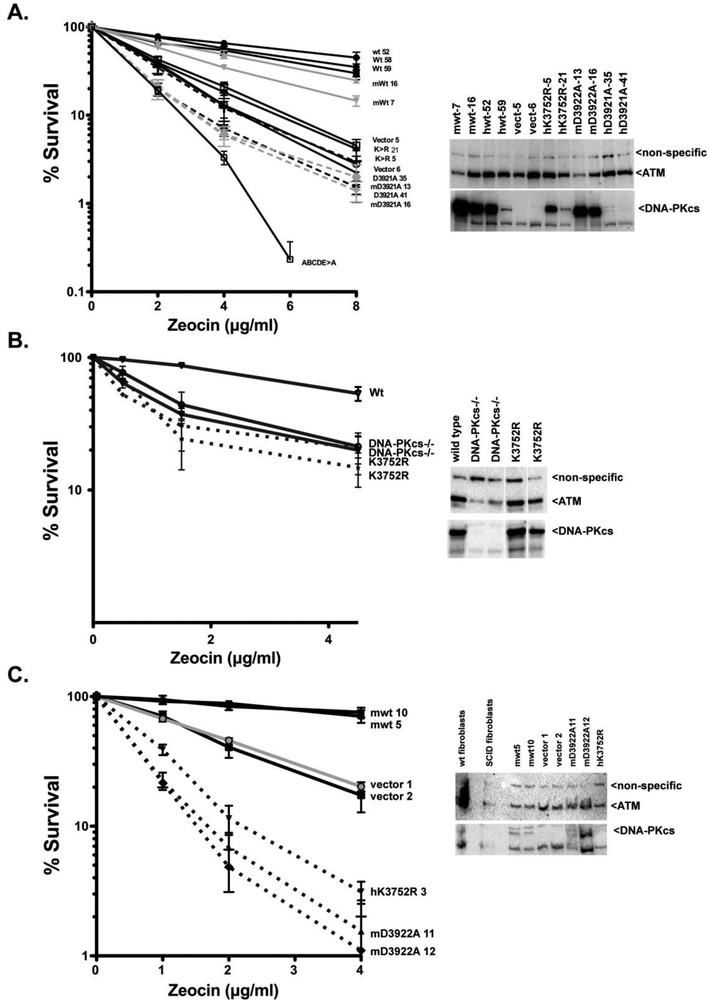
Figure 2. Human and murine DNA-PKcs complement radiomimetic drug sensitivity in DNA-PKcs deficient hamster cells to a similar extent, and kinase-inactivating mutations impart similar radiosensitivity as DNA-PKcs deficiency in both V3 cells and 293T cells; non-transformed SCID fibroblasts expressing kinase-inactivating DNA-PKcs mutants are substantially more radiosensitive than DNA-PKcs deficient cells.
(A) Left, V3 clonal transfectants expressing wild type human DNA-PKcs (hWt), wild type murine DNA-PKcs (mWt), the catalytically inactive K3753>R mutant of human DNA-PKcs (hK>R), the catalytically inactive D3921>A mutant of human DNA-PKcs (hD>A), the catalytically inactive D3922>A mutant of murine DNA-PKcs (mD>A), the ABCDE>ala cluster mutant (ABCDE>A), or no DNA-PKcs (vector) were plated at cloning densities into complete medium with increasing doses of Zeocin. Colonies were stained after seven days, and percent survival was calculated. Error bars represent the standard error of the means for six independent experiments. Right, immunoblot analyses of 50ug whole cell extracts assessed first for ATM and then for DNA-PKcs expression as indicated; non-specific band serves as a loading control. (B) Left; Wild type 293T cells, or 293T cells with targeted ablation of DNA-PKcs, or a kinase inactivating targeted mutation resulting in the K3752>R mutation were assessed for Zeocin sensitivity by MTS staining. Error bars represent the standard error of the means for six independent experiments Right, immunoblot analyses of 50ug whole cell extracts assessed first for ATM and then for DNA-PKcs as indicated. White line indicate removal of lanes not presented; figure is derived from a single blot. (C) Left, Sf19 transfectants expressing DNA-PKcs constructs described in A. were analyzed for Zeocin sensitivity as described in A. Error bars represent the standard error of the means for four independent experiments. Right, immunoblot analyses of 50ug whole cell extracts assessed first for ATM and then for DNA-PKcs expression as indicated; non-specific band serves as a loading control.
In this experiment, we included for comparison, a human DNA-PKcs construct that includes alanine substitutions of six phosphorylation sites between residues 2609 and 2647 (ABCDE cluster; ABCDE>ala); this mutant retains full catalytic activity, but renders cells substantially more radiosensitive than cells that lack DNA-PKcs or that express kinase inactive DNA-PKcs [15, 43, 44]. Consistent with previous studies, the ABCDE>ala mutant renders cells substantially more zeocin sensitive than cells lacking DNA-PKcs or cells expressing kinase inactive DNA-PKcs. We have dissected the basis for the ABCDE>ala dominant negative effect and have shown that inhibition requires enzymatic activity and likely requires a phosphorylation-dependent initiation of c-NHEJ coupled with impaired progress through c-NHEJ [16]. Since the enzymatically inactive mutants cannot initiate the phosphorylation dependent step, increased radiosensitivity is not observed.
To extend these findings further, we performed a targeted kinase inactivating mutation of the DNA-PKcs gene (K3752>R) in human HEK293T cells. Zeocin sensitivity of two independent clones expressing kinase dead DNA-PKcs to two previously established DNA-PKcs deficient 293T cell clones was assessed (Fig. 1B) [33]. The strategy used to ablate DNA-PKcs in 293T cells targets exon 6 encoding the extreme N terminus and resulted in frame shift mutations on two alleles; no untargeted alleles were detectable, and no DNA-PKcs protein is expressed. The kinase-inactivating mutant human 293T clones behave similarly to DNA-PKcs deficient 293T cells in zeocin sensitivity assays even though both express substantially more ATM than DNA-PKcs deficient cells. We conclude that kinase inactivating DNA-PKcs mutants do not potentiate radiosensitivity as compared to complete loss of DNA-PKcs in 293T or V3 cells.
SCID fibroblasts expressing kinase-inactivating DNA-PKcs mutants are substantially more radiosensitive than DNA-PKcs deficient cells.
Jiang et al demonstrated that ablation of just one copy of p53 in the DNA-PKcskd mouse rescued embryonic lethality. Since p53 function is abnormal in both 293T cells and CHO V3 cells [45, 46], we next assessed the function of these DNA-PKcs mutants in the non-transformed, immortalized SCID fibroblast cell strain, Sf19 [1, 47]. The mutation in SCID mice is the result of a premature termination codon at position 4045; although this mutation is C-terminal to the kinase domain, there is no detectable DNA-PKcs expression or enzymatic activity in cells derived from SCID mice [10]. As can be seen, whereas wild type DNA-PKcs substantially reverses zeocin sensitivity of Sf19 cells, two clones expressing the mD3922>A kinase-inactivating mutation and one clone expressing hK3752>R are substantially more zeocin sensitive than vector only controls (Fig. 1C). An additional three hK3752>R clones that expressed high levels of DNA-PKcs were generated; these were also substantially more zeocin sensitive than vector only controls (data not shown). Similar results were obtained in γ-irradiation assays (data not shown). As with V3 transfectants, ATM expression in DNA-PKcs complemented Sf19 cells varies; however, ATM levels never approach levels observed in wild type mouse fibroblasts. Moreover, ATM expression does not correlate with zeocin sensitivity (for example, compare mD3922>A-12 to hK3752>R-5). .
One cell clone studied in this series of experiments that expressed lower levels of the hK3752>R mutant displayed zeocin sensitivity comparable to DNA-PKcs deficient cells (not shown). We previously studied an Sf19 clone expressing a kinase-inactivating mutation of DNA-PKcs and found similar radiosensitivity as DNA-PKcs deficient cells [31]; however, the DNA-PKcs expression level in that cell clone was substantially less than in the wild type complemented cell control. Thus, we suggest that expression levels may contribute to the increased radiosensitivity imparted in mouse SCID fibroblasts expressing kinase inactive DNA-PKcs.
We conclude that expression of kinase inactive DNA-PKcs potentiates radiosensitivity in immortalized SCID mouse fibroblasts but not in transformed V3 or 293T cells. However, the differences in radioresistance in transformed versus non-transformed cell strains expressing kinase inactivating DNA-PKcs mutants cannot be attributed to differential ATM expression.
Differential p53 function would be a logical explanation for the observed cellular phenotypic differences. Sf19 cells were derived from SCID mice by Bosma and colleagues and were isolated by a passage and cell density 3T3 protocol [47]; however, it is well-established that this protocol can compromise p53 function [48]. Thus, we assessed p53 expression before and after exposure to 200nM calicheamicin, that potently induces DNA DSBs. As expected, phosphorylated γH2AX is not detected in untreated cells but is robustly induced after 1 hour exposure to calicheamicin. In contrast, p53 is present in both treated and untreated cells at similar levels. Similar results were observed after 3 hours or 24 hours. We conclude that p53 is dysregulated in the Sf19 SCID fibroblast cell strain and that p53 effects likely do not explain why kinase inactive DNA-PKcs potentiates radiosensitivity in DNA-PKcs deficient mouse fibroblasts but not in DNA-PKcs deficient 293T cells or CHO cells.
Kinase inactive DNA-PKcs does not alter end-joining deficits in DNA-PKcs deficient cells.
We considered that the distinct effects of kinase inactivating DNA-PKcs mutants observed in 293T and V3 cells as opposed to SCID fibroblasts and living mice, might be explained either by potentiation of the end joining deficit in DNA-PK deficient cells, or 2) altering induction of cell death. This possibility is complicated by the fact that the loss of DNA-PKcs (but not kinase inactivation) dramatically reduces ATM expression in an irreversible manner (as discussed above). This is particularly relevant for end joining that occurs during VDJ recombination. Recent studies [49] (including studies from our lab) [33, 40] have documented substantial effects of ATM loss on resolution of RAG-induced breaks because of loss of stability of the RAG post cleavage complex. Jiang et al proposed that the kinase inactivating mutation inhibited end joining in vivo [20]. However, much of their data in support of reduced end joining was derived by studying resolution of VDJ-associated DSBs. We have shown that restoring ATM in DNA-PKcs deficient cells substantially blocks signal end joining; we considered that the more severe defect in signal joining in mice harboring a kinase inactivating mutation of DNA-PKcs as compared to DNA-PKcs deficient mice may be attributable to maintenance of normal ATM levels in these mice, and not a more severe deficit in DNA end joining. Thus, we next assessed end joining of non-RAG induced DSBs in isogenic cell strains utilizing both episomal and chromosomal end-joining assays.
Episomal end joining assays were performed using a substrate (depicted in 3A, left panel) that measures joining of I-Sce1 induced DSBs. As can be seen, joining of I-Sce1 induced DSBs in cells expressing kinase inactive or no DNA-PKcs are each reduced to a similar level as compared to wild type DNA-PKcs. We assume that much of the end joining in cells lacking functional DNA-PKcs is mediated via the alternative end-joining pathway; however, there is no evidence that expression of the kinase inactive DNA-PKcs blocks end joining of episomal substrates by a-NHEJ.
Jiang et al. reported that kinase inactive DNA-PK could facilitate hairpin opening of coding intermediates via ATM phosphorylation of the ABCDE cluster in DNA-PKcs. We next examined the ability of DNA-PK deficient cells to join hairpinned ends, not associated with the RAG post cleavage complex. The TelN protelomerase targets a 56 bp recognition site, generating a double strand break, leaving both termini as covalently sealed hairpinned ends. A TelN site was inserted between the promoter and the CFP structural gene in peCFPN1. As can be seen, cells lacking DNA-PKcs or expressing kinase inactive DNA-PKcs can similarly join hairpinned termini; this joining is only slightly reduced from cells expressing wild type DNA-PKcs. We conclude that a-NHEJ can efficiently resolve DNA DSBs with hairpinned termini.
To assess chromosomal end joining in the isogenic Sf19 cell strains expressing wild type DNA-PKcs, the D3922>A mutant, or vector only, we took advantage of a Cas9/CRISPR targeting strategy and targeted two sites within the gene encoding Activation Induced Cytidine Deaminase (Figure 3B). As can be seen, chromosomal deletions are readily detected in all three cell strains. By sequence analyses, joints recovered from cells expressing wild type DNA-PKcs displayed less nucleotide loss and joining via microhomology. Nucleotide loss was increased in cells either lacking DNA-PKcs or expressing kinase inactive DNA-PKcs. However, the extent of nucleotide loss was not statistically different in joints recovered from cells expressing kinase inactive DNA-PKcs versus cells lacking DNA-PKcs. We conclude that murine SCID fibroblasts expressing kinase inactive DNA-PKcs have similar end joining deficits as cells lacking DNA-PKcs.
Figure 3. Kinase inactive DNA-PKcs does not alter end-joining deficits in DNA-PKcs deficient cells.
(A) The fluorescent substrate ISceRFP/CFP (left panel) was utilized to detect joining of I-Sce1 induced DSBs. Percent recombination of episomal fluorescent I-Sce1 end joining substrate in Sf19 transfectants expressing wild type DNA-PKcs, vector alone, or the D3922>A mutation. The fluorescent substrate telN-CFP (right panel) was utilized to detect CFP expression from uncut (U) or cut substrate (C), co-transfected with dsRED to assess transfection efficiency. TelN cleavage generates a DNA break with hairpin termini. Percent CFP/RFP is presented. Error bars indicate SEM from three independent experiments. Ns, P value, not significant, in two-tailed unpaired t test.
(B) Diagram of region of murine AICDA locus targeted by two gRNAs and position of primers utilized to detect chromosomal deletions. Ethidium bromide staining of PCR amplifications of DNA isolated from Sf19 transfectants expressing wild type DNA-PKcs, vector only, or D3922A mutant DNA-PKcs as indicated. (C) Summary of sequenced joints. P= 0.1916; (not significant) in two-tailed unpaired t test comparing nucleotide loss between sequences from cells expressing vector only versus the D3922>A mutant.
DNA-PKcs primarily autophosphorylates the ABCDE and PQR clusters.
In previous studies, we concluded that phosphorylation within the ABCDE and PQR clusters occurred primarily by autophosphorylation although very low levels of phosphorylation in both clusters of kinase inactive DNA-PKcs suggested that another kinase(s) can minimally target these sites [26]. These studies utilized both in vitro and cellular methods and used both kinase inactivating mutations and cellular inhibitors of both ATM and DNA-PKcs. In contrast, other studies have concluded that ATM and ATR preferentially or exclusively target the ABCDE sites (but not the PQR sites); these studies used both ATM deficient cell strains and kinase inhibitors [20, 28-30, 50, 51]. Because ABCDE phosphorylation is both required for c-NHEJ and likely required to prevent “toxic c-NHEJ” [16, 52], it is critical to clarify what kinase(s) phosphorylate these sites.
We considered that non-specific reagents might be responsible for conflicting results. Of note, in our study, we validated the specificity of the phospho-specific antibodies using full-length DNA-PKcs molecules with targeted alanine substitutions of phosphorylation sites. In studies from other laboratories, this stringent validation was only utilized in studies of S2056 phosphorylation [51]; this validation was not reported in studies assessing phosphorylation of sites in the ABCDE cluster [20, 28-30]. There are numerous commercially available phosphospecific DNA-PKcs antibodies. We acquired many of these and assessed their specificity. Two phospho-S2056 reagents were tested [Abcam 18192, Abcam 124918]; both detect robust phosphorylation after treatment with zeocin and okadaic acid in wild type DNA-PKcs V3 transfectants, and ABCDE>ala phospho-blocked DNA-PKcs, but not in PQR>ala blocked transfectants (Fig. 4A). In human 293T cells, robust S2056 phosphorylation is detected in wild type, and ATM deficient, but not in DNA-PKcs deficient cells. A minimal level of zeocin-induced S2056 phosphorylation is observed in the cell strain expressing kinase inactive DNA-PKcs suggesting that another kinase contributes in a minor capacity to 2056 phosphorylation (Fig. 4B). Pre-treatment of cells with an ATM inhibitor partially ablates phosphorylation in the kinase dead cell strain, but does not substantially affect 2056 phosphorylation in wild type cells or ATM deficient cells (Fig. 4c). We have previously shown that minimal DNA-PKcs S2056 phosphorylation also occurs during replication stress in V3 transfectants expressing kinase inactive DNA-PKcs [27]. Thus, these results are consistent with both our previous studies and those from Chen and colleagues [51].
Figure 4. DNA-PKcs primarily autophosphorylates the ABCDE and PQR clusters.
(A) Left, V3 transfectants expressing wild type, vector alone, ABCDE>Ala, or PQR>Ala DNA-PKcs were treated or not with zeocin and okadaic acid for one hour. 50ug whole cell extracts were analyzed by immunoblotting (left panel) with the S2056 phospho-specific antibody (top panel) or a DNA-PKcs specific antibody (second panel). On a separate gel (right panel), 50ug whole cell extracts were analyzed by immunoblotting with the T2609 phospho-specific antibody (top panel) or a DNA-PKcs specific antibody (second panel). On a third gel, 10ug whole cell extracts were analyze for phosphorylated γH2AX or actin. (B) 293T cells of the indicated genotypes were treated or not with zeocin for one hour. 50ug whole cell extracts were analyzed by immunoblotting with the S2056 phospho-specific antibody (top panel) or a T2609 phospho-specific antibody (second panel), or a DNA-PKcs specific antibody (third panel). 10ug whole cell extracts were analyze for phosphorylated γH2AX (forth panel) or actin (bottom panel). (C) 293T cells of the indicated genotypes were treated or not with zeocin with or without the ATM inhibitor KU55933 as indicated for one hour. Whole cell extracts were analyzed by immunoblotting with the S2056 phospho-specific antibody or a DNA-PKcs specific antibody. (D) Wild type 293T cells were treated or not with zeocin and Okadaic acid for one hour. Whole cell extracts were analyzed by immunoblotting with the S2056 phospho-specific antibody, T2609 phospho-specific antibody, or a DNA-PKcs specific antibody.
Several commercially available ABCDE phospho-specific antibodies were tested [phospho-T2609: Abcam 18356 (mouse monoclonal 10B1), Abcam 97611 (rabbit polyclonal), Abcam 4194 (rabbit polyclonal), and Sigma SAB4300137 (rabbit polyclonal); phospho-S2612: Abcam 133441 (rabbit monoclonal)]. None of these reagents are specific for phosphorylations in the ABCDE cluster (Sup. Fig. 3–6). In our previous study we utilized an earlier lot number of Abcam 97611; this reagent is no longer available. In our previous study, we also utilized a phospho-specific T2609 reagent provided by Dale Ramsden [26]. To demonstrate specificity of this reagent, V3 transfectants were treated with zeocin and OA; as can be seen phospho-T2609 is detected in cells expressing wild type DNA-PKcs and the PQR>ala mutant but not in cells expressing the ABCDE>Ala mutant. In human 293T cells, robust T2609 phosphorylation is detected in wild type and ATM deficient cells, but not in DNA-PKcs deficient cells (Fig. 4B). Of note, another zeocin-induced species is detected with this reagent; however, this species (that is reduced in both ATM deficient cells and in DNA-PK deficient cells) has distinct electrophoretic mobility in this 5% gel. With 8% gels, this band appears to co-migrate with DNA-PKcs, and might be mistaken for DNA-PKcs. Of note, similar DSB-induced phospho-proteins are detected by Abcam antibody, 18356 (mouse mAB 10B1, Sup. Fig. 3). Unlike S2056 phosphorylation, no zeocin-induced T2609 phosphorylation is observed in the cell strain expressing kinase inactive DNA-PKcs suggesting that DNA-PK predominately autophosphorylates T2609 in 293T cells. In our previous study, minimal IR-induced T2609 phosphorylation was detected in living cells; however, blocking cellular phosphatases with okadaic acid (OA) greatly enhanced levels of phospho-T2609. We assessed the affect of blocking cellular phosphatases in human 293T cells. Whereas OA greatly enhances detection of phospho-T2609, it does not alter levels of phospho-S2056. We conclude that T2609 is primarily autophosphorylated; clearly ATM does not substantially contribute to this phosphorylation in response to DSBs in human 293T cells. Conclusions based on the use of currently available commercially available reagents to detect DNA-PKcs phosphorylations should be re-evaluated.
Discussion
DNA-PK regulates DNA end access by preferential phosphorylation of the ABCDE and PQR clusters; whereas ABCDE phosphorylation promotes end processing, PQR phosphorylation blocks end processing [43, 44, 53, 54]. Although there is good consensus that phosphorylations within the PQR cluster are primarily autophosphorylations, there is some controversy as to whether phosphorylations within the ABCDE cluster occur primarily by autophosphorylation or by ATM or ATR [20, 26, 28-30, 50, 51, 55]. Here we demonstrate that commercially available reagents that are purported to detect phosphorylations within the ABCDE cluster, in fact do not. Here we show that DSB-induced T2609 phosphorylation of kinase inactivated DNA-PKcs is not detectable in human 293T cells. Paull and colleagues have recently shown that DNA-PK inhibits ATM activity [56]; thus, we would anticipate hyper ATM activity in cells expressing kinase inactive DNA-PKcs. Still, phosphorylation of T2609 on kinase inactivated DNA-PKcs, in cells that maintain wild type levels of ATM, is not detectable. We conclude that DSB-induced phosphorylation of the ABCDE cluster occurs primarily by autophosphorylation.
Data presented here substantiate previous reports that document loss of ATM in DNA-PKcs deficient (but not DNA-PKcs inactive) cell strains. Although differences in ATM expression do not explain the different effect of kinase ablation versus DNA-PKcs loss in cell culture models, there is incontrovertible evidence that loss of even one ATM allele dramatically affects the phenotype of c-NHEJ deficient mice [25]. It seems likely that retention of ATM expression in mice with targeted disruption of DNA-PK enzymatic activity may contribute to the severity of the phenotype in these animals as compared to DNA-PKcs deficient mice. Moreover, loss of ATM expression in DNA-PKcs deficient mice may also contribute to the observed synthetic lethality between XLF and DNA-PKcs [57] since it is well documented that XLF and ATM are functionally redundant [58-60].
The report from Jeggo and colleagues describing a DNA-PKcs deficient patient with profound neurologic pathology suggests that DNA-PK has an essential role in human neural development [12]. Here we show substantial loss of ATM in cells from that patient. Although patients afflicted with ataxia telangiectasia have a much different neurologic deficit than this DNA-PKcs deficient patient, it is possible that loss of ATM expression may have contributed to the neurologic deficits.
Here we demonstrate dramatically different effects of DNA-PKcs kinase inactivation in transformed versus non-transformed cells. Although catalytically inactive DNA-PKcs can effect VDJ joining [20, 33, 61], we find no evidence of substantial differences in end-joining deficits (of DSBs not initiated by the RAG endonuclease) in either transformed or non-transformed cells expressing kinase inactive DNA-PKcs versus no DNA-PKcs. Several potential explanations could explain these data including a non-catalytic role for DNA-PKcs in promoting cell death, or alteration of gene expression by loss of DNA-PKcs as opposed to inhibition of its catalytic activity. The preferential sensitivity of untransformed cells as compared to transformed cells to DNA-PK inhibition may impact safety of therapeutic strategies to enhance effects of cancer therapies by DNA-PK inhibition.
Supplementary Material
Acknowledgments
This work was supported by Public Health Service grant AI048758 (KM).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
DNA-PKcs deficiency has been studied exhaustively. In cell culture models, ablation of DNA-PK’s catalytic activity results in a cell phenotype that is virtually indistinguishable from that ascribed to complete loss of the enzyme. But, the same kinase inactivating mutation in living mice results in a markedly more severe phenotype. Here, we investigate potential explanations for these and other controversial results in studies of this important enzyme.
References
- [1].Bosma GC, Custer RP, Bosma MJ, A severe combined immunodeficiency mutation in the mouse, Nature, 301 (1983) 527–530. [DOI] [PubMed] [Google Scholar]
- [2].Meek K, Kienker L, Dallas C, Wang W, Dark MJ, Venta PJ, Huie ML, Hirschhorn R, Bell T, SCID in Jack Russell terriers: a new animal model of DNA-PKcs deficiency, Journal of immunology, 167 (2001) 2142–2150. [DOI] [PubMed] [Google Scholar]
- [3].Wiler R, Leber R, Moore BB, VanDyk LF, Perryman LE, Meek K, Equine severe combined immunodeficiency: a defect in V(D)J recombination and DNA-dependent protein kinase activity, Proceedings of the National Academy of Sciences of the United States of America, 92 (1995) 11485–11489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].Moore JC, Tang Q, Yordan NT, Moore FE, Garcia EG, Lobbardi R, Ramakrishnan A, Marvin DL, Anselmo A, Sadreyev RI, Langenau DM, Single-cell imaging of normal and malignant cell engraftment into optically clear prkdc-null SCID zebrafish, The Journal of experimental medicine, 213 (2016) 2575–2589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Bogue MA, Jhappan C, Roth DB, Analysis of variable (diversity) joining recombination in DNAdependent protein kinase (DNA-PK)-deficient mice reveals DNA-PK-independent pathways for both signal and coding joint formation, Proceedings of the National Academy of Sciences of the United States of America, 95 (1998) 15559–15564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Taccioli GE, Amatucci AG, Beamish HJ, Gell D, Xiang XH, Torres Arzayus MI, Priestley A, Jackson SP, Marshak Rothstein A, Jeggo PA, Herrera VL, Targeted disruption of the catalytic subunit of the DNA-PK gene in mice confers severe combined immunodeficiency and radiosensitivity, Immunity, 9 (1998) 355–366. [DOI] [PubMed] [Google Scholar]
- [7].Lieber MR, Hesse JE, Lewis S, Bosma GC, Rosenberg N, Mizuuchi K, Bosma MJ, Gellert M, The defect in murine severe combined immune deficiency: joining of signal sequences but not coding segments in V(D)J recombination, Cell, 55 (1988) 7–16. [DOI] [PubMed] [Google Scholar]
- [8].Gao Y, Chaudhuri J, Zhu C, Davidson L, Weaver DT, Alt FW, A targeted DNA-PKcs-null mutation reveals DNA-PK-independent functions for KU in V(D)J recombination, Immunity, 9 (1998) 367–376. [DOI] [PubMed] [Google Scholar]
- [9].Errami A, He DM, Friedl AA, Overkamp WJ, Morolli B, Hendrickson EA, Eckardt-Schupp F, Oshimura M, Lohman PH, Jackson SP, Zdzienicka MZ, XR-C1, a new CHO cell mutant which is defective in DNA-PKcs, is impaired in both V(D)J coding and signal joint formation, Nucleic acids research, 26 (1998) 3146–3153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Blunt T, Gell D, Fox M, Taccioli GE, Lehmann AR, Jackson SP, Jeggo PA, Identification of a nonsense mutation in the carboxyl-terminal region of DNA-dependent protein kinase catalytic subunit in the scid mouse, Proceedings of the National Academy of Sciences of the United States of America, 93 (1996) 10285–10290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [11].van der Burg M, Ijspeert H, Verkaik NS, Turul T, Wiegant WW, Morotomi-Yano K, Mari PO, Tezcan I, Chen DJ, Zdzienicka MZ, van Dongen JJ, van Gent DC, A DNA-PKcs mutation in a radiosensitive T-B-SCID patient inhibits Artemis activation and nonhomologous end-joining, The Journal of clinical investigation, 119 (2009) 91–98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Woodbine L, Neal JA, Sasi NK, Shimada M, Deem K, Coleman H, Dobyns WB, Ogi T, Meek K, Davies EG, Jeggo PA, PRKDC mutations in a SCID patient with profound neurological abnormalities, The Journal of clinical investigation, 123 (2013) 2969–2980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Woodbine L, Haines J, Coster M, Barazzuol L, Ainsbury E, Sienkiewicz Z, Jeggo P, The rate of X-ray induced DNA double strand break repair in the embryonic mouse brain is unaffected by exposure to 50 Hz magnetic fields, International journal of radiation biology, (2015) 1–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Shin EK, Perryman LE, Meek K, A kinase-negative mutation of DNA-PK(CS) in equine SCID results in defective coding and signal joint formation, Journal of immunology, 158 (1997) 3565–3569. [PubMed] [Google Scholar]
- [15].Neal JA, Dang V, Douglas P, Wold MS, Lees-Miller SP, Meek K, Inhibition of homologous recombination by DNA-dependent protein kinase requires kinase activity, is titratable, and is modulated by autophosphorylation, Molecular and cellular biology, 31 (2011) 1719–1733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Neal JA, Sugiman-Marangos S, VanderVere-Carozza P, Wagner M, Turchi J, Lees-Miller SP, Junop MS, Meek K, Unraveling the complexities of DNA-dependent protein kinase autophosphorylation, Molecular and cellular biology, 34 (2014) 2162–2175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Kurimasa A, Kumano S, Boubnov NV, Story MD, Tung CS, Peterson SR, Chen DJ, Requirement for the kinase activity of human DNA-dependent protein kinase catalytic subunit in DNA strand break rejoining, Molecular and cellular biology, 19 (1999) 3877–3884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Shrivastav M, Miller CA, De Haro LP, Durant ST, Chen BP, Chen DJ, Nickoloff JA, DNA-PKcs and ATM co-regulate DNA double-strand break repair, DNA repair, 8 (2009) 920–929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [19].Uematsu N, Weterings E, Yano K, Morotomi-Yano K, Jakob B, Taucher-Scholz G, Mari PO, van Gent DC, Chen BP, Chen DJ, Autophosphorylation of DNA-PKCS regulates its dynamics at DNA double-strand breaks, The Journal of cell biology, 177 (2007) 219–229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Jiang W, Crowe JL, Liu X, Nakajima S, Wang Y, Li C, Lee BJ, Dubois RL, Liu C, Yu X, Lan L, Zha S, Differential phosphorylation of DNA-PKcs regulates the interplay between end-processing and end-ligation during nonhomologous end-joining, Molecular cell, 58 (2015) 172–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Peng Y, Woods RG, Beamish H, Ye R, Lees-Miller SP, Lavin MF, Bedford JS, Deficiency in the catalytic subunit of DNA-dependent protein kinase causes down-regulation of ATM, Cancer research, 65 (2005) 1670–1677. [DOI] [PubMed] [Google Scholar]
- [22].Yan D, Ng WL, Zhang X, Wang P, Zhang Z, Mo YY, Mao H, Hao C, Olson JJ, Curran WJ, Wang Y, Targeting DNA-PKcs and ATM with miR-101 sensitizes tumors to radiation, PloS one, 5 (2010) e11397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Gately DP, Hittle JC, Chan GK, Yen TJ, Characterization of ATM expression, localization, and associated DNA-dependent protein kinase activity, Molecular biology of the cell, 9 (1998) 2361–2374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Lees-Miller SP, Godbout R, Chan DW, Weinfeld M, Day RS 3rd, Barron GM, Allalunis-Turner J, Absence of p350 subunit of DNA-activated protein kinase from a radiosensitive human cell line, Science, 267 (1995) 1183–1185. [DOI] [PubMed] [Google Scholar]
- [25].Sekiguchi J, Ferguson DO, Chen HT, Yang EM, Earle J, Frank K, Whitlow S, Gu Y, Xu Y, Nussenzweig A, Alt FW, Genetic interactions between ATM and the nonhomologous end-joining factors in genomic stability and development, Proceedings of the National Academy of Sciences of the United States of America, 98 (2001) 3243–3248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Meek K, Douglas P, Cui X, Ding Q, Lees-Miller SP, trans Autophosphorylation at DNA-dependent protein kinase's two major autophosphorylation site clusters facilitates end processing but not end joining, Molecular and cellular biology, 27 (2007) 3881–3890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Ying S, Chen Z, Medhurst AL, Neal JA, Bao Z, Mortusewicz O, McGouran J, Song X, Shen H, Hamdy FC, Kessler BM, Meek K, Helleday T, DNA-PKcs and PARP1 Bind to Unresected Stalled DNA Replication Forks Where They Recruit XRCC1 to Mediate Repair, Cancer research, 76 (2016) 1078–1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].Chan DW, Chen BP, Prithivirajsingh S, Kurimasa A, Story MD, Qin J, Chen DJ, Autophosphorylation of the DNA-dependent protein kinase catalytic subunit is required for rejoining of DNA double-strand breaks, Genes & development, 16 (2002) 2333–2338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Chen BP, Uematsu N, Kobayashi J, Lerenthal Y, Krempler A, Yajima H, Lobrich M, Shiloh Y, Chen DJ, Ataxia telangiectasia mutated (ATM) is essential for DNA-PKcs phosphorylations at the Thr-2609 cluster upon DNA double strand break, The Journal of biological chemistry, 282 (2007) 6582–6587. [DOI] [PubMed] [Google Scholar]
- [30].Yajima H, Lee KJ, Chen BP, ATR-dependent phosphorylation of DNA-dependent protein kinase catalytic subunit in response to UV-induced replication stress, Molecular and cellular biology, 26 (2006) 7520–7528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Kienker LJ, Shin EK, Meek K, Both V(D)J recombination and radioresistance require DNA-PK kinase activity, though minimal levels suffice for V(D)J recombination, Nucleic acids research, 28 (2000) 2752–2761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Douglas P, Cui X, Block WD, Yu Y, Gupta S, Ding Q, Ye R, Morrice N, Lees-Miller SP, Meek K, The DNA-dependent protein kinase catalytic subunit is phosphorylated in vivo on threonine 3950, a highly conserved amino acid in the protein kinase domain, Molecular and cellular biology, 27 (2007) 1581–1591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Neal JA, Xu Y, Abe M, Hendrickson E, Meek K, Restoration of ATM Expression in DNA-PKcs-Deficient Cells Inhibits Signal End Joining, Journal of immunology, 196 (2016) 3032–3042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Roy S, Andres SN, Vergnes A, Neal JA, Xu Y, Yu Y, Lees-Miller SP, Junop M, Modesti M, Meek K, XRCC4's interaction with XLF is required for coding (but not signal) end joining, Nucleic acids research, 40 (2012) 1684–1694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM, RNA-guided human genome engineering via Cas9, Science, 339 (2013) 823–826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Takai H, Wang RC, Takai KK, Yang H, de Lange T, Tel2 regulates the stability of PI3K-related protein kinases, Cell, 131 (2007) 1248–1259. [DOI] [PubMed] [Google Scholar]
- [37].Kulesza P, Lieber MR, DNA-PK is essential only for coding joint formation in V(D)J recombination, Nucleic acids research, 26 (1998) 3944–3948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Shin EK, Rijkers T, Pastink A, Meek K, Analyses of TCRB rearrangements substantiate a profound deficit in recombination signal sequence joining in SCID foals: implications for the role of DNA-dependent protein kinase in V(D)J recombination, Journal of immunology, 164 (2000) 1416–1424. [DOI] [PubMed] [Google Scholar]
- [39].Lee BS, Gapud EJ, Zhang S, Dorsett Y, Bredemeyer A, George R, Callen E, Daniel JA, Osipovich O, Oltz EM, Bassing CH, Nussenzweig A, Lees-Miller S, Hammel M, Chen BP, Sleckman BP, Functional intersection of ATM and DNA-dependent protein kinase catalytic subunit in coding end joining during V(D)J recombination, Molecular and cellular biology, 33 (2013) 3568–3579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Meek K, Xu Y, Bailie C, Yu K, Neal JA, The ATM Kinase Restrains Joining of Both VDJ Signal and Coding Ends, Journal of immunology, 197 (2016) 3165–3174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Priestley A, Beamish HJ, Gell D, Amatucci AG, Muhlmann-Diaz MC, Singleton BK, Smith GC, Blunt T, Schalkwyk LC, Bedford JS, Jackson SP, Jeggo PA, Taccioli GE, Molecular and biochemical characterisation of DNA-dependent protein kinase-defective rodent mutant irs-20, Nucleic acids research, 26 (1998) 1965–1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Woods T, Wang W, Convery E, Errami A, Zdzienicka MZ, Meek K, A single amino acid substitution in DNA-PKcs explains the novel phenotype of the CHO mutant, XR-C2, Nucleic acids research, 30 (2002) 5120–5128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Cui X, Yu Y, Gupta S, Cho YM, Lees-Miller SP, Meek K, Autophosphorylation of DNA-dependent protein kinase regulates DNA end processing and may also alter double-strand break repair pathway choice, Molecular and cellular biology, 25 (2005) 10842–10852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [44].Ding Q, Reddy YV, Wang W, Woods T, Douglas P, Ramsden DA, Lees-Miller SP, Meek K, Autophosphorylation of the catalytic subunit of the DNA-dependent protein kinase is required for efficient end processing during DNA double-strand break repair, Molecular and cellular biology, 23 (2003) 5836–5848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Hu T, Miller CM, Ridder GM, Aardema MJ, Characterization of p53 in Chinese hamster cell lines CHO-K1, CHO-WBL, and CHL: implications for genotoxicity testing, Mutation research, 426 (1999) 51–62. [DOI] [PubMed] [Google Scholar]
- [46].Moro F, Ottaggio L, Bonatti S, Simili M, Miele M, Bozzo S, Abbondandolo A, p53 expression in normal versus transformed mammalian cells, Carcinogenesis, 16 (1995) 2435–2440. [DOI] [PubMed] [Google Scholar]
- [47].Harrington J, Hsieh CL, Gerton J, Bosma G, Lieber MR, Analysis of the defect in DNA end joining in the murine scid mutation, Molecular and cellular biology, 12 (1992) 4758–4768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Harvey DM, Levine AJ, p53 alteration is a common event in the spontaneous immortalization of primary BALB/c murine embryo fibroblasts, Genes & development, 5 (1991) 2375–2385. [DOI] [PubMed] [Google Scholar]
- [49].Canela A, Sridharan S, Sciascia N, Tubbs A, Meltzer P, Sleckman BP, Nussenzweig A, DNA breaks and end resection measured genome-wide by end sequencing., Mol. Cell, 63 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Yajima H, Lee KJ, Zhang S, Kobayashi J, Chen BP, DNA double-strand break formation upon UV-induced replication stress activates ATM and DNA-PKcs kinases, Journal of molecular biology, 385 (2009) 800–810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [51].Chen BP, Chan DW, Kobayashi J, Burma S, Asaithamby A, Morotomi-Yano K, Botvinick E, Qin J, Chen DJ, Cell cycle dependence of DNA-dependent protein kinase phosphorylation in response to DNA double strand breaks, The Journal of biological chemistry, 280 (2005) 14709–14715. [DOI] [PubMed] [Google Scholar]
- [52].Zhang S, Yajima H, Huynh H, Zheng J, Callen E, Chen HT, Wong N, Bunting S, Lin YF, Li M, Lee KJ, Story M, Gapud E, Sleckman BP, Nussenzweig A, Zhang CC, Chen DJ, Chen BP, Congenital bone marrow failure in DNA-PKcs mutant mice associated with deficiencies in DNA repair, The Journal of cell biology, 193 (2011) 295–305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [53].Meek K, Gupta S, Ramsden DA, Lees-Miller SP, The DNA-dependent protein kinase: the director at the end, Immunological reviews, 200 (2004) 132–141. [DOI] [PubMed] [Google Scholar]
- [54].Reddy YV, Ding Q, Lees-Miller SP, Meek K, Ramsden DA, Non-homologous end joining requires that the DNA-PK complex undergo an autophosphorylation-dependent rearrangement at DNA ends, The Journal of biological chemistry, 279 (2004) 39408–39413. [DOI] [PubMed] [Google Scholar]
- [55].Blackford AN, Jackson SP, ATM ATR, and DNA-PK: The Trinity at the Heart of the DNA Damage Response, Molecular cell, 66 (2017) 801–817. [DOI] [PubMed] [Google Scholar]
- [56].Zhou Y, Lee JH, Jiang W, Crowe JL, Zha S, Paull TT, Regulation of the DNA Damage Response by DNA-PKcs Inhibitory Phosphorylation of ATM, Molecular cell, 65 (2017) 91–104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Oksenych V, Kumar V, Liu X, Guo C, Schwer B, Zha S, Alt FW, Functional redundancy between the XLF and DNA-PKcs DNA repair factors in V(D)J recombination and nonhomologous DNA end joining, Proceedings of the National Academy of Sciences of the United States of America, 110 (2013) 2234–2239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [58].Zha S, Guo C, Boboila C, Oksenych V, Cheng HL, Zhang Y, Wesemann DR, Yuen G, Patel H, Goff PH, Dubois RL, Alt FW, ATM damage response and XLF repair factor are functionally redundant in joining DNA breaks, Nature, 469 (2011) 250–254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Roy S, de Melo AJ, Xu Y, Tadi SK, Negrel A, Hendrickson E, Modesti M, Meek K, XRCC4/XLF Interaction Is Variably Required for DNA Repair and Is Not Required for Ligase IV Stimulation, Molecular and cellular biology, 35 (2015) 3017–3028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [60].Kumar V, Alt FW, Oksenych V, Reprint of “Functional overlaps between XLF and the ATM-dependent DNA double strand break response”, DNA repair, 17 (2014) 52–63. [DOI] [PubMed] [Google Scholar]
- [61].Cui X, Meek K, Linking double-stranded DNA breaks to the recombination activating gene complex directs repair to the nonhomologous end-joining pathway, Proceedings of the National Academy of Sciences of the United States of America, 104 (2007) 17046–17051. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.