Abstract
Cannabis cultivation is an emerging industry within the United States. Organic dust derived in part from naturally occurring microorganisms is known to cause byssinosis in the hemp industry. In this pilot study, bacteria and fungi encountered by workers at an outdoor cannabis farm that utilized organic practices were elucidated by 16S ribosomal RNA (rRNA) and Internal Transcribed Spacer (ITS) region sequencing, respectively. Area (n=14) and personal air samples (n=12) were collected during harvesting and processing activities. 16S rRNA and ITS regions of extracted bacterial and fungal genomic DNA were amplified and sequenced using Sanger sequencing. Bacterial sequencing resolved 1077 sequences that were clustered into 639 operational taxonomic units (OTUs) and predominantly placed in the phylum, Actinobacteria (46%). Personal air samples revealed higher bacterial and Actinobacteria diversity compared to outdoor area samples collected within the facility (p<0.05). A high degree of dissimilarity between bacteria was identified within and between samples. Fungal sequences (n=985) were identified and predominantly clustered in the phylum Ascomycota (53%). Of the 216 fungal OTUs elucidated, the cannabis plant pathogenic species, Botrytis cinerea, was the most prevalent and accounted for 34% of all fungal sequences. The relative abundance of B. cinerea was highest in personal air samples (59%) compared to area samples collected in the drying room (19%), greenhouse (18%) and outdoor environment (6%). There was 49% sample similarity between fungi identified within personal air samples, but higher dissimilarity coefficients were observed within and between greenhouse, drying room, and outdoor area air samples. The results of this pilot study suggest that the cannabis farm workers are potentially exposed to Actinobacteria as well as the cannabis plant pathogen, B. cinerea during harvesting, bud stripping, and hand trimming processes.
Keywords: Exposure assessment, fungus, bacteria, gene sequencing, occupational hazards, cannabis
INTRODUCTION
Recreational and medicinal cannabis use has been legalized in Alaska, Colorado, Oregon, Washington, the District of Columbia and most recently in California, Maine, Massachusetts, and Nevada. An additional 20 states have legalized medicinal cannabis use with Arkansas, Florida, and North Dakota approving initiatives in the 2016 election. Cannabis legalization in the United States has resulted in the emergence of a cultivation industry that operates under a state regulatory environment.(1) Cultivating operations include outdoor, indoor or a combination of indoor/outdoor farms that employ a workforce that maintains facilities, harvests, and processes cannabis for commercial distribution.
The acute and chronic health effects associated with consumption of the psychoactive compound delta 9-tetrahydrocannabinol (Δ9-THC) derived from C. sativa and C. indica are well described.(2) However, the potential safety and occupational health hazards encountered by workers that harvest and process cannabis in either outdoor processing environments or within greenhouses have not been well studied. Outdoor farms utilizing organic practices are a subset of cannabis growing operations that do not utilize broad-spectrum pesticides. These occupational environments are susceptible to naturally occurring microorganisms and plant pathogens. Concentrations can exceed one million colony-forming units (CFU) per m3 of air in areas where the plant is manipulated.(3, 4) Occupational health studies of forensic laboratorians have described dermal (contact urticaria) and respiratory immunoglobulin E (IgE)-mediated hypersensitivity reactions following direct contact with cannabis.(5–7) Recent outbreaks of hypersensitivity reactions to cannabis have also been recently reported among Canadian recreational users.(8) Workers in hemp processing facilities can also be exposed to organic dust that contains respirable and inhalable particles derived from microbial contaminants.(3, 9) Hemp workers with exposure to organic aerosols have higher IgE titers and chronic respiratory symptoms compared to non-exposed hemp workers(10) and a high prevalence of work-related respiratory conditions, such as byssinosis.(3, 11–14) Like hemp workers, cannabis industry workers are expected to be exposed to products, such as leaves, buds, sap/exudate, flowers, and pollen and their associated plant pathogens, including bacteria and fungi, when handling the plant during cultivation and processing procedures.
In 2015, the National Institute for Occupational Safety and Health (NIOSH) Health Hazard Evaluation (HHE) program received a request to evaluate the potential ergonomic, chemical, and biological hazards associated with cannabis harvesting and processing operations at one outdoor cannabis farm. The farm produced and harvested C. sativa and C. indica for medicinal use. Specific work-related tasks evaluated included big leafing, destemming, and hand trimming. Based on the occurrence of exposure to organic dust within the similar hemp industry, the objectives of this pilot study was to use 16S ribosomal RNA (rRNA) and Internal Transcribed Spacer (ITS) region sequencing to characterize the microbial hazards that can be encountered within an outdoor U.S. cannabis production facility.
METHODS
Facility and process description
The site of the HHE was a five-acre outdoor farm in Washington, a state that has legalized the medicinal and recreational use of cannabis.(1) The farm cultivated C. sativa, C. indica, as well as a C. sativa/indica hybrid for medicinal use. In addition to cannabis, the farm also produced fruits and vegetables utilizing organic practices. Pesticides were not used at the farm.
NIOSH visited the farm in August and October of 2015. During the October site evaluation, the farm’s workforce consisted of the owner and three employees. The farm cultivated approximately forty cannabis plants at a time from seeds. The plants were initially contained within a greenhouse and then transferred to hoop houses. Harvesting the plants included using hand pruners to cut the cola, a large stem with attached flowers/buds that emerge from the main trunk. The colas were then transferred to an area within the hoop house referred to as the big leafing area, where larger leaves were removed by employees. Colas were then dried in a drying area that was housed in a separate enclosed structure within the facility for drying times that were subjectively judged. The dried flowers/buds remaining on the dried colas were then removed through a process termed destemming. Flowers/buds were then individually trimmed and prepared for consumption.
Air sampling
Occupational tasks were observed, and personal breathing zone and area air samples were collected in the harvesting, big leafing, drying, destemming and hand trimming work locations. Air samples were collected at each site using a NIOSH BC251 bioaerosol air sampler for gene sequencing analysis and an AirCheck2000® personal air sampling pump calibrated at 2 liters per minute. Air samplers were placed outdoors (n=3), in various areas of the greenhouse (n=3) and drying room (n=8), and on farm workers (n=12) for eight hours. The NIOSH BC251 bioaerosol air sampler consists of two cyclones that deposit aerosols into standard centrifuge tubes and onto a 2 μm pore PTFE filter, allowing for the collection of particles across three size fractions: ≥ 4.9 μm; 1.7–4.9 μm; and < 1.7 μm aerodynamic diameter. Following collection, area and personal air samples were packaged and shipped overnight at room temperature to NIOSH.
Genomic DNA extraction from air samples
Air samples (n=26), field blanks (n=5) and media blanks (n=5) were processed separately for bacterial and fungal DNA extraction using the High Pure PCR Template kit (Roche, Basel, Switzerland) as previously described.(15, 16) For air samples, including field controls, each stage from the NIOSH BC251 air sampler was combined and processed sequentially with 650 μL of the High Pure PCR Template kit lysis buffer as previously described.(15) The samples were centrifuged at 20,000 × g for 1 minute, transferred to new microcentrifuge tubes, and incubated with 40 μL CelLytic B lysis reagent (Sigma-Aldrich) for 15 min at 37°C. The kit’s binding buffer (200 μL) and proteinase K (40 μL) was mixed with the samples and then incubated at 70°C for 10 minutes. The samples were mixed with 100 μL of isopropanol, washed, and eluted as recommended by the manufacturer (Roche). The eluate was aliquoted into a 1.5 mL Eppendorf tube and stored at −20°C. Extraction reagent blanks (n=5), referred to as media blanks, were additionally included in the analysis as a control.
Bacterial 16S rRNA and fungal ITS region amplification, cloning, and Sanger sequencing
Extracted bacterial (16S rRNA) and fungal (ITS) genomic DNA were targeted for PCR amplification as previously described.(15, 17, 18) Briefly, bacterial 16S rRNA genes were amplified using Platinum Taq DNA polymerase (Invitrogen, Carlsbad, CA) and the highly conserved primer pair p8FPL (AGTTTGATCCTGGCTCAG) and p806R (GGACTACCAGGGTATCTAAT) using a modified method of McCabe et al.(19) The PCR conditions included initial denaturation at 95 °C for 4 minutes, followed by 33 cycles of denaturation at 94 °C for 1 minute, annealing at 55 °C for 1 minute, and extension at 72 °C for 2 minutes, and completed with a final extension at 72 °C for 10 minutes. Fungal ITS1 and ITS2 regions were amplified using the primer pair Fun18Sf (forward, 5’-TTGCTCTTCAACGAGGAAT) and ITS4 (reverse, 5’- TCCTCCGCTTATTGATATGC) and Platinum Taq DNA polymerase (Invitrogen, Carlsbad, CA, USA) according to the methods previously described.(18) For bacterial and fungal amplification, three replicate PCR reactions (50 μL) were run for each sample using 5 μL of DNA template. The triplicate PCR reactions were then combined, and the bacterial and fungal amplicons were purified using a QIAquick PCR purification kit following the methods recommended by the manufacturer (Qiagen, Valencia, CA). Purified product (8 μL) was then resolved on a 1% agarose gel containing 1 μg/mL ethidium bromide and examined for amplicons using ultraviolet light.
Bacterial and fungal amplicons were separately cloned into the pDRIVE vector using a PCR cloning kit (Qiagen) as previously described.(18) Clone libraries were produced by transforming cloned plasmids into chemically competent Escherichia coli cells as previously described.(18) Forty-eight positive colonies per sample (as determined colorimetrically by the inactivation of the lacZ gene) were randomly selected and cultured for 16 hours at 37°C in liquid Luria-Bertani media containing 100 μg/mL ampicillin. Resultant cells were centrifuged at 1800 × g and the pellet resuspended in 200 μL of 15% glycerol in a 96-well plate. The glycerol stocks were then submitted for Sanger sequencing of the bacterial 16S rRNA or fungal ITS insert (Genewiz, Inc., South Plainfield, NJ, USA).
Plasmid DNA containing bacterial 16S rRNA and fungal ITS regions were sequenced in both directions using primers T7 and SP6. Downloaded sequences were then trimmed, and forward and reverse reads assembled using Geneious R7 Software (Biomatters Ltd, Auckland, New Zealand). Assembled sequences were initially filtered for the presence of primers, low-quality sequences, chimeras, or short nucleotide read lengths as previously described.(18) Sequences that could not assemble or be trimmed due to poor sequence quality did not pass filtering and were omitted from the analysis.
Operational taxonomic units and diversity analysis
Sequence data that passed quality filtering were then clustered into operational taxonomic units (OTUs) using MOTHUR software version 1.32.1 and the ClustalW Alignment algorithm with a 97% similarity cutoff as previously described.(18) Sequences representative of each OTU were then queried against the National Center for Biotechnology Information (NCBI) using the Basic Local Alignment Search Tool (BLAST). OTU identification and abundance data were then prepared for each sample as previously described.(18)
OTU abundance datasets were used to calculate richness and diversity indices for samples derived from personal, greenhouse, drying room, and outdoor air samples. Richness and diversity indices including Chao2, the Shannon diversity index, and the Simpson reciprocal index of diversity were calculated for each sample according to the calculations presented by Magurran.(20) Beta diversity was additionally analyzed by calculating the Bray-Curtis dissimilarity coefficient, a value between 0 (similar) and 1 (dissimilar) to evaluate sample dissimilarity within and between collected samples as previously described.(18)
Statistical analysis
Graphical presentation (relative abundance of fungal community distribution) and statistical analysis of OTUs captured from the evaluation of personal air sampler, greenhouse, drying room and outdoor air samples was completed using Sigmaplot version 12.5 (Systat Software Inc, San Jose, CA). The differences in fungal and bacterial OTUs, Actinobacteria, and Botrytis cinerea values among personal and area sampling groups were evaluated using the nonparametric rank-based, Kruskal-Wallis test to measure statistical significance at α=0.05. A Mann-Whitney U test was then utilized to compare personal sampler OTUs, Actinobacteria, and B. cinerea values to area sampler groups to measure statistical significance at α=0.05.
RESULTS
16S rRNA sequencing analysis of air samples elucidated 1,077 bacterial clones clustered into 639 OTUs. Bacterial phyla with the highest relative abundance included the Actinobacteria (46%), Proteobacteria (26%), Firmicutes (15%), and Bacteroidetes (9%) (Figure 1A). An additional eleven bacterial phyla were identified in personal and area samples and accounted for 4% of bacterial clones (Figure 1A).
Figure 1:
Relative abundance of the bacterial phyla, classes and species identified in the analysis of personal, greenhouse, drying room and outdoor air samples. (A) Identified bacterial phyla. Y-axes represent the rank order of relative abundance of bacterial phyla sequences. (B) Rank order of the most abundant bacterial classes. Y-axes represent the rank order of relative abundance of sequences placed in the four most abundant bacterial phyla, including Actinobacteria, Proteobacteria, Firmicutes, and Bacteroidetes. (C) The relative abundance of the ten most frequently detected bacterial taxa (sequences with 97% similarity) out of 639 identified bacterial OTUs. The x-axis represents the percentage of 1077 bacterial sequences.
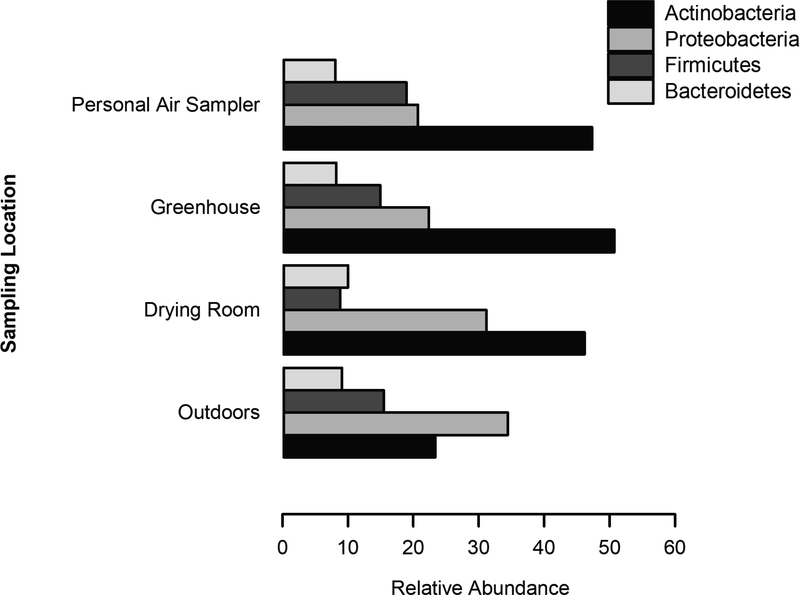
The relative abundance of the four most prominent bacterial classes is shown in Figure 1B. Bacterial classes with over 10% relative abundance included the Actinobacteria (43%), Alphaproteobacteria (16%), and Bacilli (13%; Figure 1B). In the area and personal samples, the most abundant bacterial genera included Arthrobacter spp. (2.5%), Nocardioides spp. (2.5%), and Bacillus spp. (2.1%; Figure 1C). A complete list of identified bacterial taxa are included in Supplementary Table 1. Gram-positive bacteria placed in the Actinobacteria comprised 47%, 51%, 46% and 23% in the personal air samples, greenhouse, drying room and outdoor area samples, respectively (Table 1 and Figure 2). In addition to gram-positive Actinobacteria, approximately 40% of bacterial phyla were endotoxin-producing gram-negative bacteria primarily placed in the phyla, Firmicutes and Bacteroidetes (Figure 1A and 2). In some field and media negative controls, bacterial DNA derived from species such as Bradyrhizobium elkanii were identified and subtracted from the personal and area air sampling results.
Table 1.
Bacteria and Actinobacteria operational taxonomic units by sampling location. Values presented included mean and range for OTUs, sum of Actinobacteria clone libraries (range) and the prevalence of Actinobacteria to total bacteria (range).
| Sampling Location | Sample Size | Bacterial OTU† | Actinobacteria OTU | Actinobacteria Clone Libraries | Actinobacteria % of Total Bacteria |
|---|---|---|---|---|---|
| Personal Air Samples | 11‡ | 43.1 (38–47) | 20.1 (12–28) | 21.8 (13–29) | 47.3% (28.8–60.4) |
| Harvesting | 3 | 43.7 (41–47) | 24 (20–28) | 25.3 (22–29) | 55.1% (46.8–60.4) |
| Bud Stripping | 3 | 42.3 (40–44) | 18.7 (13–24) | 21.3 (16–28) | 46.7% (36.4–58.3) |
| Trimming | 4 | 43 (38–47) | 17.8 (12–23) | 18.5 (13–24) | 40.2% (28.8–52.2) |
| Greenhouse | 3 | 40.3 (38–42) | 19.7 (16–22) | 22.7 (18–28) | 50.8% (40.9–62.2) |
| Drying Room | 8 | 37.2 (14–45) | 16.3 (6–23) | 19.2 (8–26) | 46.2% (36.2–55.3) |
| Outdoor | 3 | 26 (15–36)* | 8.3 (1–17)* | 9.7 (1–19)* | 26.4% (5.9–42.2) |
Statistically significant differences between the 4 sampling groups were observed in bacterial OTU (p = 0.034) but not Actinobacteria OTU (p = 0.125) or Actinobacteria clone library (p = 0.172) treatment groups.
In addition to the personal air samples that were collected from workers that harvested, bud stripped, and trimmed cannabis, one personal air sample was additionally collected from a site manager that visited different locations throughout the facility during the eight-hour sampling interval. One additional personal air sample was collected from a harvesting worker but a technical issue during the sampling interval resulted in the removal of this sample from the analysis.
The Mann Whitney U test, used to compare area sampling groups to the personal sampler values, revealed statistical significant differences between the personal and outdoor air sample values for bacterial OTU (p=0.012), Actinobacteria OTU (=0.043), and Actinobacteria clone libraries (p=0.043).
Figure 2:
Relative abundance of the four most abundant bacterial phyla, including Actinobacteria, Proteobacteria, Firmicutes, and Bacteroidetes, identified in personal (n=11), greenhouse (n=3), drying room (n=8), and outdoor air samples (n=3).
The mean values of bacterial OTUs were identified to be statistically different between the cannabis farm sampling locations (p=0.034; Table 1). Pairwise comparison of sampling groups also revealed a statistically significant difference between the mean number of bacterial OTUs elucidated in personal compared to outdoor air samples (p=0.012; Table 1). A significantly higher number of Actinobacteria OTUs and clone libraries were also identified in the personal air samples compared to outdoor air samples (p<0.05). No statistically significant differences were identified between personal, greenhouse or drying room samples (Table 1). Analysis of bacterial richness and diversity indices confirmed the results of the OTU statistical analysis and showed that all indices were higher in personal air samples compared to the greenhouse, drying room, or outdoor area air samples (Table 2). The Bray-Curtis dissimilarity coefficient also demonstrated a high degree of compositional dissimilarity within and between sampling groups (Table 2).
Table 2.
Bacteria species richness and diversity indices calculated for personal, greenhouse, drying room and outdoor air samples.
| Sample Size | Chao-2 | Shannon Diversity Index* | Simpson Reciprocal Index† | Bray-Curtis Dissimilarity Coefficient‡ | Bray-Curtis Dissimilarity Coefficient‡ Personal versus Area Sample | ||
|---|---|---|---|---|---|---|---|
| Bacteria | Personal Air Samples | 11 | 567.77 | 3.63 (0.01) | 165.65 (61.17) | 0.84 (0.05) | - |
| Greenhouse | 3 | 225.93 | 3.55 (0.15) | 155.68 (92.82) | 0.81 (0.03) | 0.84 (0.06) | |
| Drying Room | 8 | 362.83 | 3.45 (0.37) | 102.01 (45.11) | 0.86 (0.06) | 0.88 (0.05) | |
| Outdoor | 3 | 232.76 | 3.06 (0.39) | 51.33 (17.05) | 0.85 (0.09) | 0.92 (0.06) | |
Shannon diversity index represents the mean and standard deviation in brackets for each sample group.
Simpson reciprocal index of diversity represents the mean and standard deviation in brackets for each sample group.
Bray-Curtis dissimilarity coefficient represents the mean and standard deviation calculated for within and between samples.
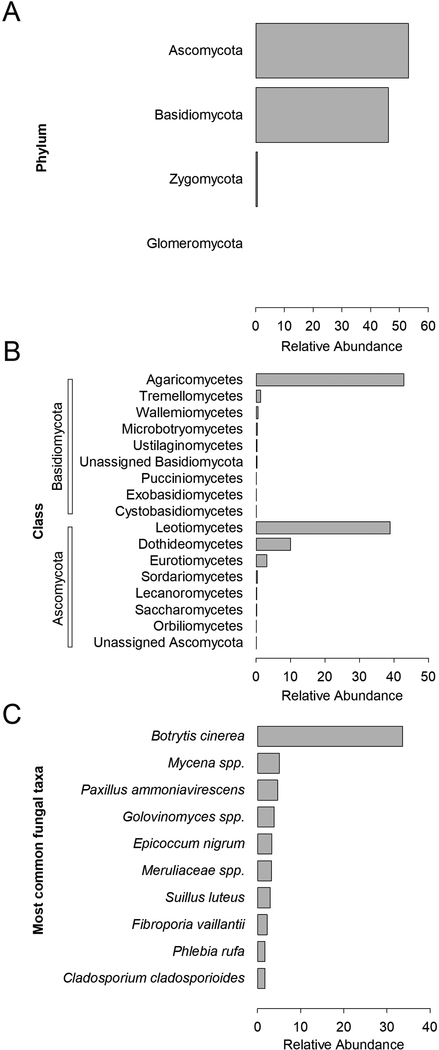
Fungal ITS region sequencing elucidated 985 clones clustered into 216 fungal OTUs. Identified sequences were placed into the fungal phyla Ascomycota (53%), Basidiomycota (46%), Zygomycota (1.2%), and Glomeromycota (0.5%; Figure 3A). The relative abundance of Ascomycota and Basidiomycota classes are shown in Figure 3B. The Agaricomycetes (Basidiomycota) and the Leotiomycetes (Ascomycota) were the most abundant classes and accounted for 42% and 38% of fungal clones, respectively. Analysis of area samples showed Basidiomycota sequences were predominant in outdoor samples from the farm (91%) and within the drying room (70%; Figure 4). In contrast, greenhouse samples included a similar relative abundance of Ascomycota (49%) and Basidiomycota (47%), as well as Zygomycota (2.7%; Figure 4).
Figure 3:
Relative abundance of the fungal phyla, classes and species identified in the analysis of personal, greenhouse, drying room and outdoor air samples. (A) Identified fungal phyla. Y-axes represent the rank order of relative abundance of fungal phyla sequences. (B) Rank order of the most abundant fungal classes. Y-axes represent the rank order of relative abundance of sequences placed in the most abundant fungal phyla, including Ascomycota and Basidiomycota. (C) The relative abundance of the ten most frequently detected fungal taxa (sequences with 97% similarity) out of 216 identified fungal OTUs. The x-axis represents the percentage of 985 bacterial sequences.
Figure 4:
Relative abundance of the fungal phyla identified in personal (n=11), greenhouse (n=3), drying room (n=8), and outdoor air samples (n=3).
Personal air samples were dominated by sequences placed in the Ascomycota (87%, Figure 4). Botrytis cinerea, a plant pathogen of cannabis that causes gray mold, was identified to be the most common fungus in the analysis of personal and area samples and accounted for 34% of fungal sequences (Figure 3C). A complete list of identified fungal taxa are included in Supplementary Table 2. Other cannabis plant pathogens placed in the class Leotiomycetes including Golovinomyces and Sclerotinia sclerotiorum were also identified in the analysis and accounted for 4% of ITS sequences. (Figure 3B and 3C). B. cinerea was the most common species accounting for approximately 59% of the fungi detected in personal air samples, 19% of the drying room air samples, and 18% of the greenhouse air samples (Table 3).
Table 3.
Mean fungal operational taxonomic units and Botrytis cinerea clone library number and prevalence by sampling location. Values presented included mean and range for OTUs, sum of Botrytis cinerea clone libraries (range) and the prevalence of Botrytis cinerea to total fungal sequences (range).
| Location | Sample Size | Fungal OTU† | Botrytis cinerea Clone Libraries† | Botrytis cinerea % of Total Fungi |
|---|---|---|---|---|
| Personal Air Samples | 11‡ | 13.1 (5–24) | 22 (3–40) | 59.2% (6.7–87) |
| Harvesting | 3 | 11.7 (5–16) | 31 (23–40) | 68.4% (49–87) |
| Bud Stripping | 3 | 11 (6–14) | 15 (11–21) | 54.2% (24–60) |
| Trimming | 4 | 13.5 (7–24) | 24.3 (3–36) | 61.4% (6.7–85.7) |
| Greenhouse | 3 | 22.7 (18–28) | 8 (1–17) | 17.7% (2.1–35.4) |
| Drying Room | 8 | 22.5 (15–33)* | 7 (1–14)* | 19.3% (2.2–29.2) |
| Outdoor | 3 | 30 (26–34)* | 3 (0–8)* | 6 % (0–13.6) |
Statistically significant differences between the 4 sampling groups were observed for fungal OTU (p <0.003), and B. cinerea clone library (p = 0.013) treatment groups.
In addition to the personal air samples that were collected from workers that harvested, bud stripped, and trimmed cannabis, one personal air sample was additionally collected from a site manager that visited different locations throughout the facility during the eight-hour sampling interval. One additional personal air sample was collected from a harvesting worker but a technical issue during the sampling interval resulted in the removal of this sample from the analysis.
The Mann Whitney U test, used to compare area sampling groups to the personal sampler values, revealed statistical significant differences between the personal air samples and the drying room values for fungal OTU (p=0.006), and B. cinerea clone libraries (p=0.012). Statistical significant differences were additionally observed between the personal air samples and the outdoor air sample values for fungal OTU (p=0.013), and B. cinerea clone libraries (p=0.029).
The mean numbers of fungal OTUs were significantly different among the fungal sampling groups (p=0.003; Table 3). Pairwise analysis of sampling groups showed that personal air samples had significantly lower OTUs compared to drying room (p=0.006) and outdoor air samples (p=0.013; Table 3). There were no statistically significant differences between fungal OTUs by occupational tasks including harvesting, bud stripping, or hand trimming (Table 3). Analysis of B. cinerea clone libraries showed that the highest number of clone libraries were resolved in personal air samples, in particular among workers performing harvesting and trimming occupational tasks (Table 3). Personal B. cinerea clone libraries were significantly higher compared to drying room (p=0.012) and outdoor air sampling groups (p=0.029; Table 3). Fungal richness was highest in personal and drying room samples (Table 4). However, richness may have been underestimated as rarefaction curves for each sampling group did not reach a horizontal asymptote (data not shown). In contrast to bacterial diversity results, fungal diversity indices were highest in outdoor and drying room sampling groups compared to personal air samples (Table 4). Bray-Curtis dissimilarity coefficient scores showed 49% sample similarity within personal air samples; however, higher dissimilarity coefficients were observed within and between greenhouse, drying room and outdoor area air samples (Table 4).
Table 4.
Fungal species richness and diversity indices calculated for personal, greenhouse, drying room and outdoor air samples.
| Sample Size | Chao-2 | Shannon Diversity Index* | Simpson Reciprocal Index† | Bray-Curtis Dissimilarity Coefficient‡ | Bray-Curtis Dissimilarity Coefficient‡ Personal versus Area Sample | ||
|---|---|---|---|---|---|---|---|
| Fungi | Personal Air Samples | 11 | 266.05 | 1.52 (0.82) | 5.78 (7.31) | 0.51 (0.19) | - |
| Greenhouse | 3 | 129.22 | 2.64 (0.4) | 14.5 (12.34) | 0.71 (0.05) | 0.73 (0.15) | |
| Drying Room | 8 | 224.44 | 2.71 (0.4) | 22.39 (18.21) | 0.65 (0.09) | 0.78 (0.11) | |
| Outdoor | 3 | 119.67 | 3.01 (0.12) | 18.88 (4.28) | 0.61 (0.06) | 0.91 (0.07) | |
Shannon diversity index represents the mean and standard deviation in brackets for each sample group.
Simpson reciprocal index of diversity represents the mean and standard deviation in brackets for each sample group.
Bray-Curtis dissimilarity coefficient represents the mean and standard deviation calculated for within and between samples.
DISCUSSION
Sequencing of 16S rRNA and ITS regions is a method increasingly used in occupational health studies to assess the diversity of bacteria and fungi in air and dust samples.(17, 18, 21) This methodological approach overcomes some of the limitations associated with traditional methods of microbial exposure assessment, such as elucidating non-culturable fungi from viable fungi or microscopically differentiating fungal spores that share the same morphology.(15) In the present study, analysis of personal and area air samples derived from an outdoor cannabis farm that utilized organic practices revealed an expected broad diversity of bacterial and fungal sequences. In addition to exposure to cannabis plant components, the results of this study further elucidated Actinobacteria as well as the fungal species B. cinerea as additional microbial exposures for workers that performed harvesting, bud stripping, and hand trimming tasks. These findings build on recent research findings that found culturable fungi in thirty U.S. indoor cannabis grow operations.(4)
Actinobacteria are a diverse and abundant phylum of gram-positive bacteria (22) that can be aerosolized into the breathing zone of a worker.(23, 24) Occupational environments that produce or use organic materials such as mushroom compost, potato processing, or swine confinement facilities and even water damaged building materials can support the growth and proliferation of Actinobacteria species.(23–27) The Actinobacteria were the most abundant bacterial phyla identified in the drying room, greenhouse and personal air samples collected at the organic cannabis farm. Areas, where workers processed and handled cannabis, had the highest relative abundance of Actinobacteria that was dominated by the genera Arthrobacter and Nocardioides. Members of the genus Arthrobacter have been previously identified in other occupational environments such as sawmills(28) and accounted for as much as 81% of the total bacterial count in a potato processing environment where 46% of workers reported work-related respiratory symptoms.(25) In contrast, Nocardioides is a soil-borne genus that produces hyphae that fragment into rod-coccoid elements.(29) The prevalence of Nocardioides in occupational environments has not been widely reported, although this genus was recently described in low abundance in a study that evaluated the bacterial microbiome of indoor dust and outdoor air samples collected as part of a Boston area birth cohort and school exposure study.(30) In addition to Actinobacteria, the bacterial diversity indices demonstrated a higher richness and diversity in personal compared to outdoor air samples. Approximately 40% of bacterial phyla identified in the analysis consisted of endotoxin-producing gram-negative bacteria and included Firmicutes genera such as Bacillus. Occupational exposure to gram-negative endotoxin-producing bacteria is well characterized and can elicit adverse respiratory health effects, such as hypersensitivity pneumonitis, chronic bronchitis, organic dust toxic syndrome, asthma and allergic sensitization.(31–34) In the present study, the bacterial sequencing datasets suggest that workers that handle and trim organic cannabis can be exposed to a broad diversity of gram-positive and gram-negative bacterial taxa.
Although unable to quantify biomass, Sanger sequencing of fungal ITS regions revealed a diversity of fungi that was predominantly placed in the phyla Basidiomycota and Ascomycota. Outdoor samples were primarily composed of Basidiomycota species placed in the class, Agaricomycetes. Common sequences identified in area samples included Mycena, Paxillus ammoniavirescens, and Meruliaceae species. The Agaricomycetes is a diverse class that accounts for approximately twenty percent of all fungi and includes species that decay wood(35) and produce fruiting structures termed basidiocarps (mushrooms) that are either edible or contain toxins.(35, 36) This fungal class was prominent in outdoor and the drying room samples of the organic cannabis farm and has been previously identified in sequencing studies of occupational(18) and residential environments.(15, 37, 38) Members of this class include most commercially produced edible mushroom species such as Lentinus edodes (Shiitake) and Pleurotus ostreatus (Oyster). Worker exposure to the basidiospores in cultivation and packing facilities can result in the development of hypersensitivity pneumonitis.(39–41) In the present study, we hypothesize that workers may additionally be exposed to Agaricomycete spores that are produced by species growing on surrounding trees or breaking down decomposing plant material in the humus soil. Many of the Agaricomycete sequences identified in this analysis have not been evaluated in exposure assessment studies, and to date, little is known about potential adverse health effects associated with inhalation exposures.
Personal air samples derived from workers conducting harvesting, bud stripping and hand trimming tasks revealed B. cinerea to be the most common fungal sequence detected in the ITS sequence analysis accounting for 34% of all fungal sequences. B. cinerea is placed in the phylum Ascomycota and is a well-characterized plant pathogen that causes gray mold in over 200 plant species.(42) B. cinerea is considered the most significant fungal pathogen of cannabis and causes bud rot that affects the seedlings, stems, and buds.(43, 44) B. cinerea has been found to be among the most common fungi identified in other greenhouse environments(45, 46) and disturbance associated with occupational tasks has been shown to raise airborne B. cinerea conidia levels.(47, 48) In general, the prevalence of Botrytis in the outdoor air is low with values as low as 1.1% calculated in a recent review.(49) In the present study, the species accounted for 6% of all outdoor fungal sequences perhaps due to the lack of pesticide use. Other cannabis plant pathogens placed in the class Leotiomycetes were also identified in the analysis, including Golovinomyces and Sclerotinia sclerotiorum,(50) and accounted for approximately 4% of all fungal clone libraries. These results further suggest that workers can be exposed to cannabis plant pathogens that produce respirable size spores. Occupational exposure to spores mainly derived from B. cinerea may elicit allergic sensitization or hypersensitivity pneumonitis as shown in other occupational settings, including bell pepper greenhouses,(51, 52) Chrysanthemum greenhouses,(53) and berry production facilities (raspberry and grape).(54–56)
Additionally, the results of this study build upon a preliminary analysis of 30 indoor cannabis grow operations and demonstrate that workers handling cannabis may be exposed to a broad diversity of plant pathogens or other regionally abundant microorganisms.(4) Although focused on law enforcement worker exposures, Martyny and colleagues(4) showed that removal of cannabis plants resulted in elevated airborne Penicillium levels that were as high as 5.3 ×105 spores per m3. In our study, ITS sequencing of air samples derived from an outdoor organic cannabis farm showed lower fungal diversity in personal air samples dominated by the fungal pathogen B. cinerea as well as fungi placed in the class Agaricomycetes. Differences in the reported species between these studies could be a function of several previously described limiting variables,(18) including discriminating viable from nonviable fungi, differentiating unicellular fungal spores as well as sequencing limitations such as gDNA extraction bias, amplification bias (Cladosporium and Penicillium species), homology within ITS sequences of the selected primer set or variable copy number of DNA, and potentially differences in pesticide use. Although other culture-independent methods such as matrix-assisted laser desorption/ionization time-of-flight mass spectrometry may overcome some of these methodological limitations in future analyses,(21) the sequencing data obtained in this study further highlight that workers handling cannabis may be potentially exposed to respirable spores produced by cannabis plant pathogens such as B. cinerea.
These findings highlight coexposures that may occur in workers that process or handle cannabis. Co-exposure to organic dust that is composed of bacteria (gram-positive and gram-negative), fungi, insects, and protozoa that may predate on C. sativa and C. indica have been previously reported in European hemp processing facilities. Occupational disturbance procedures, such as floor sweeping, result in the generation of organic dust that includes endotoxin and other microorganisms.(3) In one hemp factory, airborne levels of Actinobacteria and fungi exceeded 6 × 105 CFU per m3. Occupational exposure to organic dust has been particularly problematic in the hemp processing industry and is a risk factor for the development of work-related respiratory symptoms.(3, 57) Although bacteria and fungi were not quantified in the present study, the sequence data provide preliminary insight to suggest that workers undertaking various occupational tasks could be exposed to a variety of microbial hazards.
CONCLUSIONS
Sequencing studies have provided unique insight into the diversity of bacteria and fungi in a variety of indoor and occupational environments. Analysis of personal and area samples derived from a U.S. outdoor cannabis farm revealed a broad diversity of bacteria and fungi that included gram-positive Actinobacteria and the cannabis fungal plant pathogen, B. cinerea. These results highlight potential exposure to microbiological hazards in workers that harvest, bud strip or hand trim organically produced cannabis. Based on the literature derived from the hemp industry and given what is known about the microorganisms detected in this study, the relationship between these exposures and health should be examined in this industry, to determine the risk to workers and the need for controls.
Supplementary Material
ACKNOWLEDGEMENTS
The findings and the conclusions in this report are those of the authors and do not necessarily represent the views of the National Institute for Occupational Safety and Health. The authors declare no conflict of interest. This study was supported by internal NIOSH funds and in part by an interagency agreement between NIOSH and NIEHS (AES12007001–1-0–6) as a collaborative National Toxicology Program research activity.
REFERENCES
- 1.Jensen EL, and Roussell A: Field observations of the developing legal recreational cannabis economy in Washington State. Int J Drug Policy 33: 96–101 (2016). [DOI] [PubMed] [Google Scholar]
- 2.Hall W, and Degenhardt L: Adverse health effects of non-medical cannabis use. Lancet 374(9698): 1383–1391 (2009). [DOI] [PubMed] [Google Scholar]
- 3.Fishwick D, Allan LJ, Wright A, and Curran AD: Assessment of exposure to organic dust in a hemp processing plant. Ann Occup Hyg 45(7): 577–583 (2001). [PubMed] [Google Scholar]
- 4.Martyny JW, Serrano KA, Schaeffer JW, and Van Dyke MV: Potential exposures associated with indoor marijuana growing operations. J Occup Environ Hyg 10(11): 622–639 (2013). [DOI] [PubMed] [Google Scholar]
- 5.Herzinger T, Schopf P, Przybilla B, and Rueff F: IgE-mediated hypersensitivity reactions to cannabis in laboratory personnel. Int Arch Allergy Immunol 156(4): 423–426 (2011). [DOI] [PubMed] [Google Scholar]
- 6.Majmudar V, Azam NA, and Finch T: Contact urticaria to Cannabis sativa. Contact Dermatitis 54(2): 127 (2006). [DOI] [PubMed] [Google Scholar]
- 7.Williams C, Thompstone J, and Wilkinson M: Work-related contact urticaria to Cannabis sativa. Contact Dermatitis 58(1): 62–63 (2008). [DOI] [PubMed] [Google Scholar]
- 8.Tessmer A, Berlin N, Sussman G, Leader N, Chung EC, and Beezhold D: Hypersensitivity reactions to marijuana. Ann Allergy Asthma Immunol 108(4): 282–284 (2012). [DOI] [PubMed] [Google Scholar]
- 9.Zuskin E, Mustajbegovic J, and Schachter EN: Follow-up study of respiratory function in hemp workers. Am J Ind Med 26(1): 103–115 (1994). [DOI] [PubMed] [Google Scholar]
- 10.Zuskin E, Kanceljak B, Schachter EN, Witek TJ, Maayani S, Goswami S et al. : Immunological findings in hemp workers. Environ Res 59(2): 350–361 (1992). [DOI] [PubMed] [Google Scholar]
- 11.Er M, Emri SA, Demir AU, Thorne PS, Karakoca Y, Bilir N et al. : Byssinosis and COPD rates among factory workers manufacturing hemp and jute. Int J Occup Med Environ Health 29(1): 55–68 (2016). [DOI] [PubMed] [Google Scholar]
- 12.Valic F, Zuskin E, Walford J, Kersic W, and Paukovic R: Byssinosis, chronic bronchitis, and ventilatory capacities in workers exposed to soft hemp dust. Br J Ind Med 25(3): 176–186 (1968). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Velvart J, and Stavrovska O: Health of workers engaged in the processing of hemp. Prac Lek 15: 153–157 (1963). [PubMed] [Google Scholar]
- 14.Zuskin E, Kanceljak B, Pokrajac D, Schachter EN, and Witek TJ Jr.: Respiratory symptoms and lung function in hemp workers. Br J Ind Med 47(9): 627–632 (1990). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Rittenour WR, Ciaccio CE, Barnes CS, Kashon ML, Lemons AR, Beezhold DH et al. : Internal transcribed spacer rRNA gene sequencing analysis of fungal diversity in Kansas City indoor environments. Environ Sci Process Impacts 16(1): 33–43 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rittenour WR, Park JH, Cox-Ganser JM, Beezhold DH, and Green BJ: Comparison of DNA extraction methodologies used for assessing fungal diversity via ITS sequencing. J Environ Monit 14(3): 766–774 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Broadwater K, de Perio MA, Roberts J, Burton NC, Lemons AR, Green BJ et al. : Investigating a persistent odor at an aircraft seat manufacturer. J Occup Environ Hyg 13(10): D159–165 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Green BJ, Lemons AR, Park Y, Cox-Ganser JM, and Park JH: Assessment of fungal diversity in a water-damaged office building. J Occup Environ Hyg 14(4): 285–293 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.McCabe KM, Khan G, Zhang YH, Mason EO, and McCabe ER: Amplification of bacterial DNA using highly conserved sequences: automated analysis and potential for molecular triage of sepsis. Pediatrics 95(2): 165–169 (1995). [PubMed] [Google Scholar]
- 20.Magurran AE: Measuring biological diversity: Blackwell Publishing, Oxford, 2004. [Google Scholar]
- 21.Madsen AM, Zervas A, Tendal K, and Nielsen JL: Microbial diversity in bioaerosol samples causing ODTS compared to reference bioaerosol samples as measured using Illumina sequencing and MALDI-TOF. Environ Res 140: 255–267 (2015). [DOI] [PubMed] [Google Scholar]
- 22.Ventura M, Canchaya C, Tauch A, Chandra G, Fitzgerald GF, Chater KF et al. : Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol Mol Biol Rev 71(3): 495–548 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dutkiewicz J, Jabloński L, and Olenchock SA: Occupational biohazards: A review. Am J Ind Med 14(5): 605–623 (1988). [DOI] [PubMed] [Google Scholar]
- 24.Rautiala S, Kangas J, Louhelainen K, and Reiman M: Farmers’ exposure to airborne microorganisms in composting swine confinement buildings. AIHA Journal 64(5): 673–677 (2003). [DOI] [PubMed] [Google Scholar]
- 25.Dutkiewicz J: Bacteria, fungi, and endotoxin as potential agents of occupational hazard in a potato processing plant. Am J Ind Med 25(1): 43–46 (1994). [DOI] [PubMed] [Google Scholar]
- 26.Pasciak M, Pawlik K, Gamian A, Szponar B, Skora J, and Gutarowska B: An airborne actinobacteria Nocardiopsis alba isolated from bioaerosol of a mushroom compost facility. Aerobiologia 30(4): 413–422 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Schäfer J, Jäckel U, and Kämpfer P: Analysis of Actinobacteria from mould-colonized water damaged building material. Syst Appl Microbiol 33(5): 260–268 (2010). [DOI] [PubMed] [Google Scholar]
- 28.Dutkiewicz J, Krysińska-Traczyk E, Skórska C, Milanowski J, Sitkowska J, Dutkiewicz E et al. : Microflora of the air in sawmills as a potential occupational hazard: concentration and composition of microflora and immunologic reactivity of workers to microbial aeroallergens. Pneumonol Alergol Pol 64 (Suppl 1): 25–31 (1996). [PubMed] [Google Scholar]
- 29.Prauser H: Nocardioides, a New Genus of the Order Actinomycetales. Int. J. Syst. Evol. Microbiol 26(1): 58–65 (1976). [Google Scholar]
- 30.Hanson B, Zhou Y, Bautista EJ, Urch B, Speck M, Silverman F et al. : Characterization of the bacterial and fungal microbiome in indoor dust and outdoor air samples: a pilot study. Environ Sci Process Impacts (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lacey J, and Crook B: Fungal and actinomycete spores as pollutants of the workplace and occupational allergens. Ann Occup Hyg 32(4): 515–533 (1988). [DOI] [PubMed] [Google Scholar]
- 32.Mackiewicz B, Skórska C, and Dutkiewicz J: Relationship between concentrations of microbiological agents in the air of agricultural settings and occurenceof work-related symptoms in exposed persons. Ann Agric Environ Med 22(3)(2015). [DOI] [PubMed] [Google Scholar]
- 33.Park JH, Cox‐Ganser J, Rao C, and Kreiss K: Fungal and endotoxin measurements in dust associated with respiratory symptoms in a water‐damaged office building. Indoor Air 16(3): 192–203 (2006). [DOI] [PubMed] [Google Scholar]
- 34.Pepys J, Jenkins P, Festenstein G, Lacey M, Gregory P, and Skinner F: Farmer’s lung thermophilic actinomycetes as a source of “farmer’s lung hay” antigen. Lancet 282(7308): 607–611 (1963). [DOI] [PubMed] [Google Scholar]
- 35.Hibbett DS, Bauer R, Binder M, Giachini AJ, Hosaka K, Justo A et al. : Agaricomycetes In Systematics and Evolution: Part A, McLaughlin JD and Spatafora WJ (eds.), pp. 373–429. Berlin, Heidelberg: Springer Berlin Heidelberg, 2014. [Google Scholar]
- 36.Hibbett DS: After the gold rush, or before the flood? Evolutionary morphology of mushroom-forming fungi (Agaricomycetes) in the early 21st century. Mycol Res 111(9): 1001–1018 (2007). [DOI] [PubMed] [Google Scholar]
- 37.Pitkäranta M, Meklin T, Hyvärinen A, Nevalainen A, Paulin L, Auvinen P et al. : Molecular profiling of fungal communities in moisture damaged buildings before and after remediation-a comparison of culture-dependent and culture-independent methods. BMC Microbiol 11(1): 1 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yamamoto N, Nazaroff WW, and Peccia J: Assessing the aerodynamic diameters of taxon-specific fungal bioaerosols by quantitative PCR and next-generation DNA sequencing. J Aerosol Sci 78: 1–10 (2014). [Google Scholar]
- 39.Ampere A, Delhaes L, Soots J, Bart F, and Wallaert B: Hypersensitivity pneumonitis induced by Shiitake mushroom spores. Med Mycol 50(6): 654–657 (2012). [DOI] [PubMed] [Google Scholar]
- 40.Bekci TT, Calik M, Calik SG, and Esme H: Oyster mushrooms (Pleurotus ostreatus) caused hypersensitivity pneumonitis: Mushroom worker’s lung. Eur Respir J 44(Suppl 58): P4023 (2014). [Google Scholar]
- 41.Hodgson MJ, and Flannigan B: Occupational Respiratory Disease: Hypersensitivity pneumonitis and other forms of intersitial lung disease. In Microorganisms in Home and Indoor Work Environments: Diversity, Health Impacts, Investigation and Control, pp. 183, 2016. [Google Scholar]
- 42.Williamson B, Tudzynski B, Tudzynski P, and Van Kan JAL: Botrytis cinerea: the cause of grey mould disease. Mol Plant Pathol 8(5): 561–580 (2007). [DOI] [PubMed] [Google Scholar]
- 43.McPartland JM: A review of Cannabis diseases. J Int Hemp Assoc 3(1): 19–23 (1996). [Google Scholar]
- 44.Rodriguez G, Kibler A, Campbell P, and Punja ZK: “Fungal diseases of Cannabis sativa in British Columbia, Canada.” [Online] Available, 2015).
- 45.Monso E, Magarolas R, Badorrey I, Radon K, Nowak D, and Morera J: Occupational asthma in greenhouse flower and ornamental plant growers. Am J Respir Crit Care Med 165(7): 954–960 (2002). [DOI] [PubMed] [Google Scholar]
- 46.Radon K, Danuser B, Iversen M, Monso E, Weber C, Hartung J et al. : Air contaminants in different European farming environments. Ann Agric Environ Med 9(1): 41–48 (2002). [PubMed] [Google Scholar]
- 47.Hausbeck M, and Pennypacker S: Influence of grower activity and disease incidence on concentrations of airborne conidia of Botrytis cinerea among geranium stock plants. Plant Dis 75(8): 798–803 (1991). [Google Scholar]
- 48.Simeray J, Mandin D, Mercier M, and Chaumont J-P: Survey of viable airborne fungal propagules in French wine cellars. Aerobiologia 17(1): 19–24 (2001). [Google Scholar]
- 49.Jurgensen CW, and Madsen A: Exposure to the airborne mould Botrytis and its health effects. Ann Agric Environ Med 16(2): 183–196 (2009). [PubMed] [Google Scholar]
- 50.Boland G, and Hall R: Index of plant hosts of Sclerotinia sclerotiorum. Can J Plant Pathol 16(2): 93–108 (1994). [Google Scholar]
- 51.Groenewoud GC, de Graaf in ‘t Veld C, vVan Oorschot-van Nes AJ, de Jong NW, Vermeulen AM, van Toorenenbergen AW et al. : Prevalence of sensitization to the predatory mite Amblyseius cucumeris as a new occupational allergen in horticulture. Allergy 57(7): 614–619 (2002). [DOI] [PubMed] [Google Scholar]
- 52.Groenewoud GC, de Jong NW, van Oorschot-van Nes AJ, Vermeulen AM, van Toorenenbergen AW, Mulder PG et al. : Prevalence of occupational allergy to bell pepper pollen in greenhouses in the Netherlands. Clin Exp Allergy 32(3): 434–440 (2002). [DOI] [PubMed] [Google Scholar]
- 53.Groenewoud GC, de Jong NW, Burdorf A, de Groot H, and van Wyk RG: Prevalence of occupational allergy to Chrysanthemum pollen in greenhouses in the Netherlands. Allergy 57(9): 835–840 (2002). [DOI] [PubMed] [Google Scholar]
- 54.Jarvis W: The dispersal of spores of Botrytis cinerea Fr. in a raspberry plantation. Trans Brit Mycol Soc 45(4): 549–559 (1962). [Google Scholar]
- 55.Jeebhay MF, Baatjies R, Chang YS, Kim YK, Kim YY, Major V et al. : Risk factors for allergy due to the two-spotted spider mite (Tetranychus urticae) among table grape farm workers. Int Arch Allergy Immunol 144(2): 143–149 (2007). [DOI] [PubMed] [Google Scholar]
- 56.Popp W, Ritschka L, Zwick H, and Rauscher H: “Berry sorter’s lung” or wine grower’s lung--an exogenous allergic alveolitis caused by Botrytis cinerea spores. Prax Klin Pneumol 41(5): 165–169 (1987). [PubMed] [Google Scholar]
- 57.Fishwick D, Allan LJ, Wright A, Barber CM, and Curran AD: Respiratory symptoms, lung function and cell surface markers in a group of hemp fiber processors. Am J Ind Med 39(4): 419–425 (2001). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.