Abstract
Objective
The tumor susceptibility gene 101 (TSG101) is closely associated with various tumor types, but its role in the pathogenesis of renal cell carcinoma (RCC) is still unknown. This study used RNA interference to silence the expression of TSG101 in RCC cell lines and explore the role of TSG101 in RCC.
Methods
Immunohistochemistry and western blot were performed to detect the expression of TSG101 in 15 paired renal tumor samples. A small interfering RNA (siRNA) targeting TSG101 was transfected into A498 and 786-O cell lines. The Cell Counting Kit-8 (CCK-8) assay and colony formation assay were used to observe the changes in cell proliferation after transfection. Flow cytometry was used to detect the effect on the cell cycle. Western blot was conducted to study the changes of related functional proteins.
Results
The expression of TSG101 was higher in RCC tissues than in adjacent normal tissues. The CCK-8 assay showed that the proliferation and colony formation of the A498 and 786-O cell lines were attenuated after suppression of TSG101. Flow cytometry showed that silencing of TSG101 induced G0/G1 arrest. The western blot results revealed that the levels of cell cycle-related proteins (c-myc, cyclin E1 and cyclin-dependent kinase 2 (CDK2)) were markedly decreased in the siRNA groups.
Conclusions
TSG101 promotes proliferation of RCC cells. This positive effect on tumor growth involves activation of c-myc and cyclin E1/CDK2 and their effect on cell cycle distribution.
Keywords: Renal cell carcinoma, TSG101, Cell proliferation, Cell cycle
Introduction
Renal cell carcinoma (RCC) is one of the most refractory cancers in the world and accounts for about 2 to 3% of adult malignancies [1]. Among RCCs, clear cell RCC (ccRCC) is the most common subtype [2]. In addition to the undetermined pathogenesis, the nonspecific symptoms and metastatic lesions at initial diagnosis result in poor prognosis [3]. Currently, therapies targeting the vascular endothelial growth factor (VEGF) receptors and mammalian target of rapamycin (mTOR) are commonly applied in RCC treatment and have obtained a certain curative effect [4], yet some patients still do not achieve the expected efficacy due to drug resistance. Hence, surgery remains the main treatment of RCC [5, 6] and seeking novel therapeutic strategies and prognostic markers is critical.
The tumor susceptibility gene 101 (TSG101) is located in chromosome 11p15 and encodes a 46 kDa protein of 390 amino acid residues [7]. TSG101 is a multi-functional protein whose functions include the sorting and transport of endosomes [8–10], modulation of protein ubiquitination [11], and participation in p53/MDM2 feedback control loops [12, 13], thereby affecting epithelial cell growth and differentiation [14] and regulation of the cell cycle [15], with significant roles in the maintenance of cell homeostasis. To date, a growing body of evidence has indicated that TSG101 is overexpressed in various tumors [16–20], suggesting that TSG101 contributes to the promotion of cancers. Hence, we specifically down-regulated TSG101 using a small interfering RNA (siRNA) in order to observe its impact on the proliferation and cell cycle of RCC cells.
In this study, TSG101 was proved to be a novel oncogenic gene that facilitated RCC cell cycle progression. In addition, we predicted and verified that the effect of TSG101 on the growth of tumor was related to elevated c-myc protein levels, accompanied by up-regulated cyclin E1/cyclin-dependent kinase 2 (CDK2) complex. These findings may shed some light on the oncogenesis of RCC and provide more valuable strategies for the treatment of patients with RCC.
Materials and methods
Clinical samples
A total of 15 paired tumor tissues were harvested from patients who received partial or radical nephrectomy at the Department of Urology of Shanghai Tenth People’s Hospital. The specimens were freshly frozen in liquid nitrogen until use. Informed consent was obtained from the patients and the study was approved by the ethics committee of the Tenth People’s Hospital of Tongji University (approved on February 23, 2017; approval # SHSY-IEC-KY -4.0/17–86/01), according to the tenets of the Declaration of Helsinki.
Immunohistochemistry
Fresh tissue samples were fixed in 4% paraformaldehyde, dehydrated through a graded series of ethanol solution and embedded in paraffin. Then the sections were deparaffinized in xylene and dehydrated with an ethanol gradient followed by blocking of endogenous peroxidase activity with 0.3% hydrogen peroxide in methanol for 20 min. Nonspecific binding was blocked by incubating the sections with 10% normal goat serum in phosphate buffered saline (PBS) for 1 h at room temperature. The sections were incubated with the primary antibody against TSG101 (1500, ab125011, Abcam, Cambridge, MA, USA) in PBS at 4 °C overnight. Horseradish peroxidase-polymer conjugated anti-rabbit IgG was used as the secondary antibody. The slides were stained with hematoxylin for visualization.
Cell culture
The human RCC cell lines A498 and 786-O were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in Dulbecco’s modified Eagle’s medium (DMEM) and RPMI-1640 (GIBCO, Life Technologies, Grand Island, NY, USA), respectively, supplemented with 10% fetal bovine serum (FBS; GIBCO) and 1% penicillin-streptomycin (Hyclone, Logan, Utah, USA), in an incubator at 37 °C with 5% CO2.
Transfection assay
A498 and 786-O cells (105/well) were cultured in a 6-well plate before transfection. When the confluence reached 30–50%, transfection of the TSG101 siRNA and negative control (NC) siRNA was performed using the Lipofectamine 2000 Transfection kit (Invitrogen, Thermo Fisher Scientific, Inc., USA), according to the manufacturer’s instructions. The amount of siRNAs was 0.1 nmol. After 4–6 h of incubation, the medium was replaced with Opti-MEM (GIBCO, Life Technologies, Grand Island, NY, USA). The cells were used for future analysis 48 h after transfection.
The TSG101 and NC siRNAs were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). The sequence of the TSG101 siRNA was 5′-GCC UAC UAG UUC AAU GAC UTT-3′ (sense) and 5′-AGU CAU UGA ACU AGU AGG CTT-3′ (antisense), while the sequence of the NC siRNA was 5′-UUC UCC GAA CGU GUC ACG UTT-3′ (sense) and 5’-ACG UGA CAC GUU CGG AGA ATT-3′ (antisense).
Cell counting Kit-8 (CCK-8) and colony formation assay
The CCK-8 assay (Dojindo Molecular Technologies, Kimamoto, Japan) was used to assess cell proliferation, following the manufacturer’s instructions. Transfected cells were seeded into 96-well plates at 103 cells/well. The CCK-8 reagent (10 μl) was added to the wells after 24 h and incubated for 2 h, followed by the measurement of optical density (OD) at 450 nm using a microplate spectrophotometer (Bio-Tek, Winooski, VT, USA). The transfected cells were seeded into 6-well plates at 103 cells/well and used to perform the colony formation assay. After 8–10 days, the cells were washed with PBS lightly, fixed in 95% ethanol, and stained with 0.1% crystal violet.
Cell cycle analysis
The transfected cells were detached by EDTA-free trypsin (GIBCO), washed with precooled PBS, and fixed in 75% ethanol at 4 °C overnight. The cells were resuspended in 0.2 mL of PI/RNase Staining Buffer (BD Biosciences, San Jose, CA, USA) and incubated in the dark for 30 min. The cells were analyzed using a flow cytometer (BD Biosciences).
RNA extraction and reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
According to the manufacturer’s protocol, total cellular RNA was extracted using TRIzol (Invitrogen) and stored at − 80 °C. For TSG101 detection, the cDNA was generated using the PrimeScript RT-PCR kit (Takara Bio, Inc., Otsu, Japan), following the manufacturer’s instructions. The RT conditions were 37 °C for 15 min, then 85 °C for 5 s. The SYBR-Green PCR master mix (Takara) was used for qPCR on a 7900HT Fast RT-PCR instrument (Applied Biosystems; Thermo Fisher Scientific). The amplification protocol was: 3 min at 95 °C, followed by 40 cycles at 95 °C for 3 s and 60 °C for 30 s. The TSG101 mRNA levels were normalized to the β-Actin mRNA levels using the 2-ΔΔCt method. The primer sequences were: TSG101: 5′-GCC ACC TCT AGA ATG GCG GTG TCG GAG AGC C-3′ (F) and 5′-GGT GGC GTC GAC TCA GTA GAG GTC ACT GAG ACC-3′ (R); β-actin 5′-CAG AGC CTC GCC TTT GCC-3′ (F) and 5′-GTC GCC CAC ATA GGA ATC-3′ (R).
Western blot assay
Cells were washed with PBS twice and lysed in radioimmunoprecipitation assay lysis buffer (70 μL/well, Beyotime, Jiangsu, China) after 48–72 h of transfection. Protein concentration was determined using the bicinchoninic acid protein assay (Beyotime). Equal amounts of protein (25–50 μg) were loaded in 8% or 10% SDS-PAGE gel for electrophoresis. The proteins were transferred to nitrocellulose membranes (Sangon Biotech, Shanghai, China). The membranes were blocked at room temperature in 5% skimmed milk diluted with PBS plus Tween 20 (PBST) for 1 h and hybridized with the primary antibodies overnight at 4 °C. The membranes were washed with PBST three times and incubated with the appropriate secondary antibodies for 1 h. The membranes were washed with PBST three times and the protein bands were visualized using an Odyssey Scanner (LI-COR Biosciences, Lincoln, NE, USA).
The antibodies were: GAPDH (1:10,000, ab181602, Abcam, Cambridge, MA, USA), TSG01 (1:2000, ab125011, Abcam), c-myc (1:1000, 13,987, Cell Signaling, Danvers, MA, USA), cyclin E1 (1:1000, ab33911, Abcam), and CDK2 (1:1000, ab32147, Abcam).
Statistical analysis
Data were analyzed using SPSS 20.0 (IBM, Armonk, NY, USA) and GraphPad Prism software 5.0 (GraphPad Prism Software Inc., San Diego, CA, USA). All data were from at least three independent experiments and are presented as means ± standard deviation (SD). Student’s t-test was used for comparison and p < 0.05 was considered to be statistically significant.
Results
Expression of TSG101 was higher in RCC tumor tissues compared with adjacent normal tissues
Previous studies have identified TSG101 as an oncogene in various tumor types [19–21]. First, we detected the TSG101 expression in RCC tissues compared with in paracancerous normal tissues. The protein level of TSG101 was examined by immunohistochemistry. Representative staining of TSG101 in tumor tissues compared to adjacent normal tissue of RCC patients is shown in Fig. 1a. As expected, compared with adjacent normal tissues, TSG101 mRNA expression was higher in RCC tumor tissues (p < 0.05) (Fig. 1b). These data hint that high expression of TSG101 is associated with RCC development.
Fig. 1.
a Representative immunohistochemistry of TSG101 in tumor tissues compared to adjacent normal tissue of RCC patients (magnification × 100). b The expression of TSG101 mRNA was determined in tumor tissues and their matched adjacent normal tissues by qRT-PCR
Expression of TSG101 was obviously reduced by siRNA
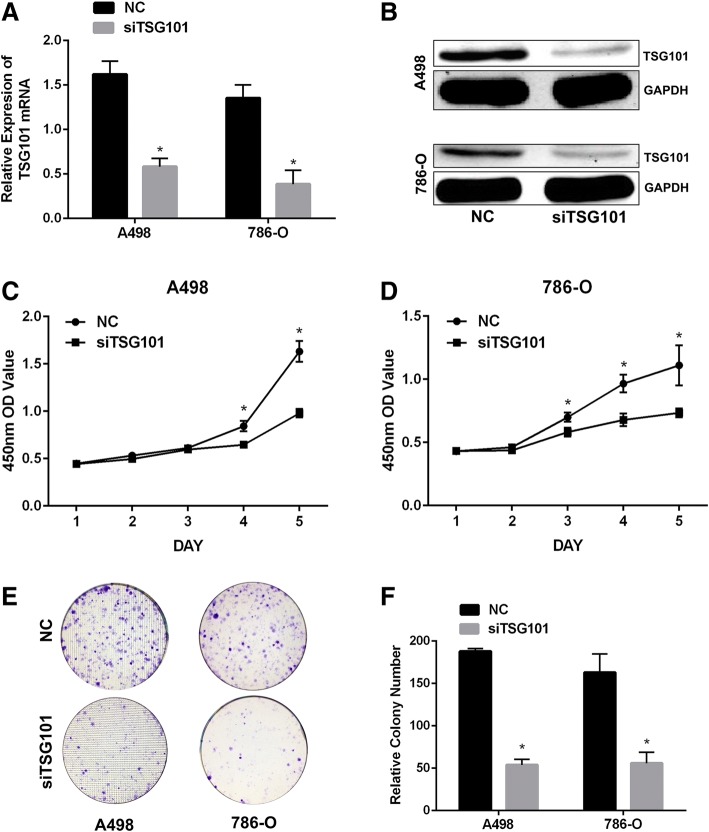
To further investigate the potential function of TSG101, the RCC cell lines A498 and 786-O were selected for the RNA interference experiment and transfected with siRNA targeting human TSG101 (siTSG101) and a NC siRNA. The silencing efficiency of the siRNA on TSG101 expression was detected by western blot and qRT-PCR (p < 0.05) (Fig. 2a and b). The data showed that the siRNA exclusively targeted TSG101. Compared with NC siRNA, siTSG101 reduced endogenous TSG101 expression in A498 and 786-O cells.
Fig. 2.
Knockdown of TSG101 inhibited cell proliferation and colony formation ability in RCC. a qRT-PCR and (b) western blot confirmed that TSG101 could be silenced successfully by siRNA. CCK-8 assays of (c) A498 and (d) 786-O cells transfected with siTSG101 or NC. e and f Colony formation assays of A498 and 786-O cells transfected with siTSG101 or NC. Representative images and average colony numbers are shown. Data are presented as means ± SD. * p < 0.05 vs. NC (n = 3)
TSG101 downregulation inhibited cell proliferation in RCC
We performed CCK-8 and colony formation assays to examine the effect of TSG101 on the proliferation of RCC cells. As determined by the CCK-8 assay, silencing TSG101 slowed cell proliferation in a time-dependent manner in A498 and 786-O cells compared with the cells transfected with NC siRNA (p < 0.05) (Fig. 2c and d). The results of the colony formation assay supported those of the CCK-8 assay (p < 0.05) (Fig. 2e and f). These results, taken together, indicate that TSG101 could enhance RCC cell proliferation.
Knockdown of TSG101 led to RCC cell cycle arrest
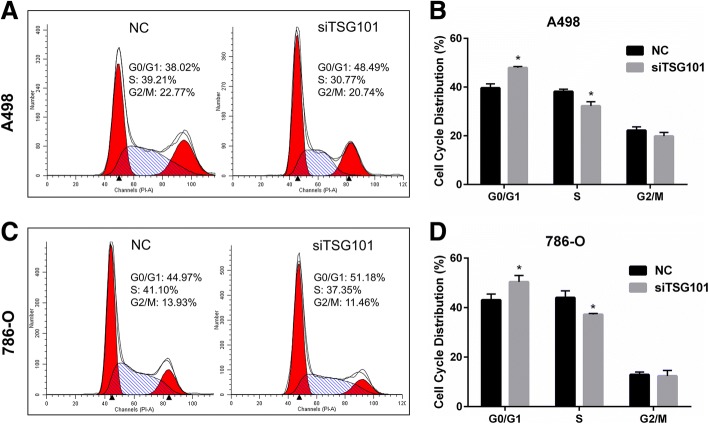
Cell cycle analysis was conducted to investigate the mechanisms by which the silencing of TSG101 blunted cell proliferation. The cell cycle distribution was altered by the inhibition of TSG101. The proportion of G0/G1 phase cells was increased and the proportion of S phase cells was reduced by down-regulated TSG101 in A498 and 786-O cell lines (p < 0.05) (Fig. 3). Based on these data, we hypothesized that downregulation of TSG101 may inhibit proliferation by inducing cell cycle arrest in RCC cells.
Fig. 3.
TSG101 downregulation induced RCC cell cycle arrest. Flow cytometry in (a) A498 and (c) 786-O cells transfected with siTSG101 or NC. b and d The G0/G1, S, and G2/M phase proportions of A498 and 786-O cells transfected with siTSG101 or NC. Data are presented as means ± SD. * p < 0.05 vs. control group (n = 3)
Cell cycle-related proteins levels were suppressed by inhibition of TSG101 in RCC cells
Cyclins are a family of proteins that control the progression of the cell cycle by activating CDK enzymes and their aberrant expression disturbs the cell cycle [22, 23]. In addition, inhibition of c-myc in growing cells leads to G1 arrest with a consequent decrease of cyclin E and CDK2 [24]. As shown in Fig. 4, western blotting showed that compared with those in the NC groups, the expression levels of cell cycle-associated proteins (c-myc, cyclin E1, and CDK2) were markedly reduced in the siRNA group, supporting the results of the cell cycle analysis. These results implied that TSG101 regulated, at least in part, the cell cycle of RCC cells through cyclin E1/CDK2 activity, which has a positive correlation with c-Myc.
Fig. 4.

Cell cycle related proteins were suppressed by inhibition of TSG101 in RCC cells. Western blot analysis of c-myc, CDK2 cyclin E1, and TSG101 protein levels of the siTSG101 group compared with the NC group in (a) A498 and (b) 786-O cells
Discussion
Since the study by Li et al. that reported TSG101 as a novel tumor susceptibility gene [25], a number of studies have demonstrated that high expression of TSG101 in numerous malignant tumors was responsible for cancer cell growth. An early study verified that the upregulation of TSG101 might boost carcinogenesis in papillary thyroid carcinoma [21], followed by a study that proved that its expression might be necessary for processes involved in tumor progression [20]. Moreover, recent studies also showed that positive TSG101 expression was associated with the clinical, pathological, and biological behaviors and with poor prognosis in breast cancer and hepatocellular carcinoma [18, 19]. These findings highlight TSG101 as a cancer-promoting gene, but until now, no report has investigated the role of TSG101 in the development of RCC.
In the present study, siRNA was used to down-regulate endogenous TSG101 to study the role of TSG101 in RCC cell lines and thereby explored the association between TSG101 and tumorigenesis of RCC. The CCK-8 and colony formation assays revealed that depletion of TSG101 reduced cell growth, which was in accordance with previous studies. Furthermore, inhibition of TSG101 promoted RCC cells to be arrested in the G0/G1 phase.
The accurate transition from G1 phase to S phase is crucial for the control of eukaryotic cell proliferation. Cyclin E1/CDK2 kinase activity peaks at the G1/S phase and is required for proper cell cycle progression into the S phase [26, 27]. Beyond question, the dysregulation of cyclin E1/CDK2 kinase activity is involved in oncogenesis in ovarian cancer [28], bladder cancer [29], and breast cancer [30], among others. C-myc also regulates the cell cycle and contributes to the formation of cancers [31–33]. In addition, a previous study constructed a mutant c-myc protein (named MadMyc) that could cause repression of the c-myc target genes; the study showed that the MadMyc-induced G1 arrest was rescued by ectopic expression of cyclin E and CDK2 [24]. So far, several studies have reported that c-myc had a bearing on cyclin E1/CDK2 kinase activity [34–36]. Interestingly, this same situation could be observed in the present study. Indeed, repression of TSG101 reduced the protein levels of c-myc, cyclin E1, and CDK2 in RCC cells.
In conclusion, the present study used siRNA to provide some evidence that TSG101 has a positive influence on RCC cell proliferation and that this effect may rely on proteins that are involved in cell cycle regulation, including c-myc, cyclin E1, and CDK2. These findings might help explore the terra incognita of RCC tumorigenesis and could provide promising strategies for the treatment of RCC.
Acknowledgements
Not applicable.
Funding
This research was sponsored by grants from the National Natural Science Foundation of China (No. 31570775 and 81602216).
Availability of data and materials
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- CCK-8
Cell Counting Kit-8
- CDK2
cyclin-dependent kinase 2
- mTOR
mammalian target of rapamycin
- PBS
phosphate buffered saline
- RCC
renal cell carcinoma
- siRNA
small interfering RNA
- TSG101
tumor susceptibility gene 101
- VEGF
vascular endothelial growth factor
Authors’ contributions
Junhua Zheng conceived and designed the research. Chen Xu performed the experiments and analyzed the data. Junhua Zheng and Chen Xu drafted the manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to participate
Informed consent was obtained from patients and the study was approved by the ethics committee of Tenth People’s Hospital of Tongji University (approved on February 23, 2017; approval # SHSY-IEC-KY -4.0/17–86/01), according to the tenets of the Declaration of Helsinki.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67:7–30. doi: 10.3322/caac.21387. [DOI] [PubMed] [Google Scholar]
- 2.Capitanio U, Montorsi F. Renal cancer. Lancet. 2016;387(10021):894–906. doi: 10.1016/S0140-6736(15)00046-X. [DOI] [PubMed] [Google Scholar]
- 3.Mancuso A, Sternberg CN. New treatments for metastatic kidney cancer. Can J Urol. 2005;12(Suppl 1):66–70. [PubMed] [Google Scholar]
- 4.Calvo E, Schmidinger M, Heng DY, Grunwald V, Escudier B. Improvement in survival end points of patients with metastatic renal cell carcinoma through sequential targeted therapy. Cancer Treat Rev. 2016;50:109–117. doi: 10.1016/j.ctrv.2016.09.002. [DOI] [PubMed] [Google Scholar]
- 5.Motzer RJ, Michaelson MD, Redman BG, Hudes GR, Wilding G, Figlin RA, et al. Activity of SU11248, a multitargeted inhibitor of vascular endothelial growth factor receptor and platelet-derived growth factor receptor, in patients with metastatic renal cell carcinoma. J Clin Oncol. 2006;24(1):16–24. doi: 10.1200/JCO.2005.02.2574. [DOI] [PubMed] [Google Scholar]
- 6.Rini BI, Atkins MB. Resistance to targeted therapy in renal-cell carcinoma. Lancet Oncol. 2009;10(10):992–1000. doi: 10.1016/S1470-2045(09)70240-2. [DOI] [PubMed] [Google Scholar]
- 7.Li L, Li X, Francke U, Cohen SN. The TSG101 tumor susceptibility gene is located in chromosome 11 band p15 and is mutated in human breast cancer. Cell. 1997;88(1):143–154. doi: 10.1016/S0092-8674(00)81866-8. [DOI] [PubMed] [Google Scholar]
- 8.Ren X, Kloer DP, Kim YC, Ghirlando R, Saidi LF, Hummer G, et al. Hybrid structural model of the complete human ESCRT-0 complex. Structure. 2009;17(3):406–416. doi: 10.1016/j.str.2009.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lee HH, Elia N, Ghirlando R, Lippincott-Schwartz J, Hurley JH. Midbody targeting of the ESCRT machinery by a noncanonical coiled coil in CEP55. Science. 2008;322(5901):576–580. doi: 10.1126/science.1162042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Muromoto R, Sugiyama K, Yamamoto T, Oritani K, Shimoda K, Matsuda T. Physical and functional interactions between Daxx and TSG101. Biochem Biophys Res Commun. 2004;316(3):827–833. doi: 10.1016/j.bbrc.2004.02.126. [DOI] [PubMed] [Google Scholar]
- 11.Goff A, Ehrlich LS, Cohen SN, Carter CA. Tsg101 control of human immunodeficiency virus type 1 gag trafficking and release. J Virol. 2003;77(17):9173–9182. doi: 10.1128/JVI.77.17.9173-9182.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cheng TH, Cohen SN. Human MDM2 isoforms translated differentially on constitutive versus p53-regulated transcripts have distinct functions in the p53/MDM2 and TSG101/MDM2 feedback control loops. Mol Cell Biol. 2007;27(1):111–119. doi: 10.1128/MCB.00235-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Szemraj J, Rozponczyk E, Bartkowiak J, Greger J, Oszajca K. Significance of MDM2 protein in the cell cycle. Postepy Biochem. 2005;51(1):44–51. [PubMed] [Google Scholar]
- 14.You HL, Eng HL, Hsu SF, Chen CM, Ye TC, Liao WT, et al. A PKC-Sp1 signaling pathway induces early differentiation of human keratinocytes through upregulation of TSG101. Cell Signal. 2007;19(6):1201–1211. doi: 10.1016/j.cellsig.2007.01.014. [DOI] [PubMed] [Google Scholar]
- 15.Moberg KH, Schelble S, Burdick SK, Hariharan IK. Mutations in erupted, the Drosophila ortholog of mammalian tumor susceptibility gene 101, elicit non-cell-autonomous overgrowth. Dev Cell. 2005;9(5):699–710. doi: 10.1016/j.devcel.2005.09.018. [DOI] [PubMed] [Google Scholar]
- 16.Liu DC, Yang ZL, Jiang S. Identification of PEG10 and TSG101 as carcinogenesis, progression, and poor-prognosis related biomarkers for gallbladder adenocarcinoma. Pathol Oncol Res. 2011;17(4):859–866. doi: 10.1007/s12253-011-9394-7. [DOI] [PubMed] [Google Scholar]
- 17.Liu F, Yu Y, Jin Y, Fu S. TSG101, identified by screening a cancer cDNA library and soft agar assay, promotes cell proliferation in human lung cancer. Mol Biol Rep. 2010;37(6):2829–2838. doi: 10.1007/s11033-009-9835-5. [DOI] [PubMed] [Google Scholar]
- 18.Shao Z, Ji W, Liu A, Qin A, Shen L, Li G, et al. TSG101 silencing suppresses hepatocellular carcinoma cell growth by inducing cell cycle arrest and Autophagic cell death. Med Sci Monit. 2015;21:3371–3379. doi: 10.12659/MSM.894447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang Y, Song M, Cui ZS, Li CY, Xue XX, Yu M, et al. Down-regulation of TSG101 by small interfering RNA inhibits the proliferation of breast cancer cells through the MAPK/ERK signal pathway. Histol Histopathol. 2011;26(1):87–94. doi: 10.14670/HH-26.87. [DOI] [PubMed] [Google Scholar]
- 20.Zhu G, Gilchrist R, Borley N, Chng HW, Morgan M, Marshall JF, et al. Reduction of TSG101 protein has a negative impact on tumor cell growth. Int J Cancer. 2004;109(4):541–547. doi: 10.1002/ijc.20014. [DOI] [PubMed] [Google Scholar]
- 21.Liu RT, Huang CC, You HL, Chou FF, Hu CC, Chao FP, et al. Overexpression of tumor susceptibility gene TSG101 in human papillary thyroid carcinomas. Oncogene. 2002;21(31):4830–4837. doi: 10.1038/sj.onc.1205612. [DOI] [PubMed] [Google Scholar]
- 22.Rahman MM, Kipreos ET. The specific roles of mitotic cyclins revealed. Cell Cycle. 2010;9(1):22–23. doi: 10.4161/cc.9.1.10577. [DOI] [PubMed] [Google Scholar]
- 23.Galderisi U, Jori FP, Giordano A. Cell cycle regulation and neural differentiation. Oncogene. 2003;22(33):5208–5219. doi: 10.1038/sj.onc.1206558. [DOI] [PubMed] [Google Scholar]
- 24.Berns K, Hijmans EM, Bernards R. Repression of c-Myc responsive genes in cycling cells causes G1 arrest through reduction of cyclin E/CDK2 kinase activity. Oncogene. 1997;15(11):1347–1356. doi: 10.1038/sj.onc.1201280. [DOI] [PubMed] [Google Scholar]
- 25.Li L, Cohen SN. Tsg101: a novel tumor susceptibility gene isolated by controlled homozygous functional knockout of allelic loci in mammalian cells. Cell. 1996;85(3):319–329. doi: 10.1016/S0092-8674(00)81111-3. [DOI] [PubMed] [Google Scholar]
- 26.Bertoli C, Skotheim JM, de Bruin RA. Control of cell cycle transcription during G1 and S phases. Nat Rev Mol Cell Biol. 2013;14(8):518–528. doi: 10.1038/nrm3629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Honda R, Lowe ED, Dubinina E, Skamnaki V, Cook A, Brown NR, et al. The structure of cyclin E1/CDK2: implications for CDK2 activation and CDK2-independent roles. EMBO J. 2005;24(3):452–463. doi: 10.1038/sj.emboj.7600554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Marone M, Scambia G, Giannitelli C, Ferrandina G, Masciullo V, Bellacosa A, et al. Analysis of cyclin E and CDK2 in ovarian cancer: gene amplification and RNA overexpression. Int J Cancer. 1998;75(1):34–39. doi: 10.1002/(SICI)1097-0215(19980105)75:1<34::AID-IJC6>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 29.Juan G, Cordon-Cardo C. Intranuclear compartmentalization of cyclin E during the cell cycle: disruption of the nucleoplasm-nucleolar shuttling of cyclin E in bladder cancer. Cancer Res. 2001;61(3):1220–1226. [PubMed] [Google Scholar]
- 30.Lucenay KS, Doostan I, Karakas C, Bui T, Ding Z, Mills GB, et al. Cyclin E associates with the Lipogenic enzyme ATP-citrate Lyase to enable malignant growth of breast Cancer cells. Cancer Res. 2016;76(8):2406–2418. doi: 10.1158/0008-5472.CAN-15-1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Eriksson A, Kalushkova A, Jarvius M, Hilhorst R, Rickardson L, Kultima HG, et al. AKN-028 induces cell cycle arrest, downregulation of Myc associated genes and dose dependent reduction of tyrosine kinase activity in acute myeloid leukemia. Biochem Pharmacol. 2014;87(2):284–291. doi: 10.1016/j.bcp.2013.10.022. [DOI] [PubMed] [Google Scholar]
- 32.Li E, Liu L, Li F, Luo L, Zhao S, Wang J, et al. PSCA promotes prostate cancer proliferation and cell-cycle progression by up-regulating c-Myc. Prostate. 2017;77(16):1563–1572. doi: 10.1002/pros.23432. [DOI] [PubMed] [Google Scholar]
- 33.Wang H, Mannava S, Grachtchouk V, Zhuang D, Soengas MS, Gudkov AV, et al. C-Myc depletion inhibits proliferation of human tumor cells at various stages of the cell cycle. Oncogene. 2008;27(13):1905–1915. doi: 10.1038/sj.onc.1210823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dimberg A, Bahram F, Karlberg I, Larsson LG, Nilsson K, Oberg F. Retinoic acid-induced cell cycle arrest of human myeloid cell lines is associated with sequential down-regulation of c-Myc and cyclin E and posttranscriptional up-regulation of p27(Kip1) Blood. 2002;99(6):2199–2206. doi: 10.1182/blood.V99.6.2199. [DOI] [PubMed] [Google Scholar]
- 35.Santoni-Rugiu E, Falck J, Mailand N, Bartek J, Lukas J. Involvement of Myc activity in a G(1)/S-promoting mechanism parallel to the pRb/E2F pathway. Mol Cell Biol. 2000;20(10):3497–3509. doi: 10.1128/MCB.20.10.3497-3509.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Prall OW, Rogan EM, Musgrove EA, Watts CK, Sutherland RL. C-Myc or cyclin D1 mimics estrogen effects on cyclin E-Cdk2 activation and cell cycle reentry. Mol Cell Biol. 1998;18(8):4499–4508. doi: 10.1128/MCB.18.8.4499. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.