Abstract
The clinical benefit in patients with diverse types of metastatic cancers that is observed upon blockade of the PD-1 – PD-L1 interaction has highlighted the importance of this inhibitory axis in the suppression of tumor-specific T cell responses1–9. In spite of the key role of PD-L1 expression by cells within the tumor micro-environment, our understanding of the regulation of the PD-L1 protein is limited10–15. Using a haploid genetic screen, we here identify CMTM6, a type 3 transmembrane protein of previously unknown function, as a regulator of the PD-L1 protein. Interference with CMTM6 expression results in impaired PD-L1 protein expression in all tumor cell types tested and in primary human dendritic cells. Furthermore, through both a haploid genetic modifier screen in CMTM6 deficient cells and genetic complementation experiments, we demonstrate that this function is shared by its closest family member CMTM4, but not by all other CMTM members tested. Notably, CMTM6 increases the PD-L1 protein pool without affecting PD-L1 transcript levels. Rather, we demonstrate that CMTM6 is present at the cell surface, associates with the PD-L1 protein, reduces its ubiquitination and increases PD-L1 protein half-life. Consistent with its role in PD-L1 protein regulation, T cell inhibitory capacity of PD-L1 expressing tumor cells is enhanced by CMTM6. Collectively, our data reveal that PD-L1 relies on CMTM6/4 to efficiently carry out its inhibitory function, and suggest potential new avenues to block this pathway.
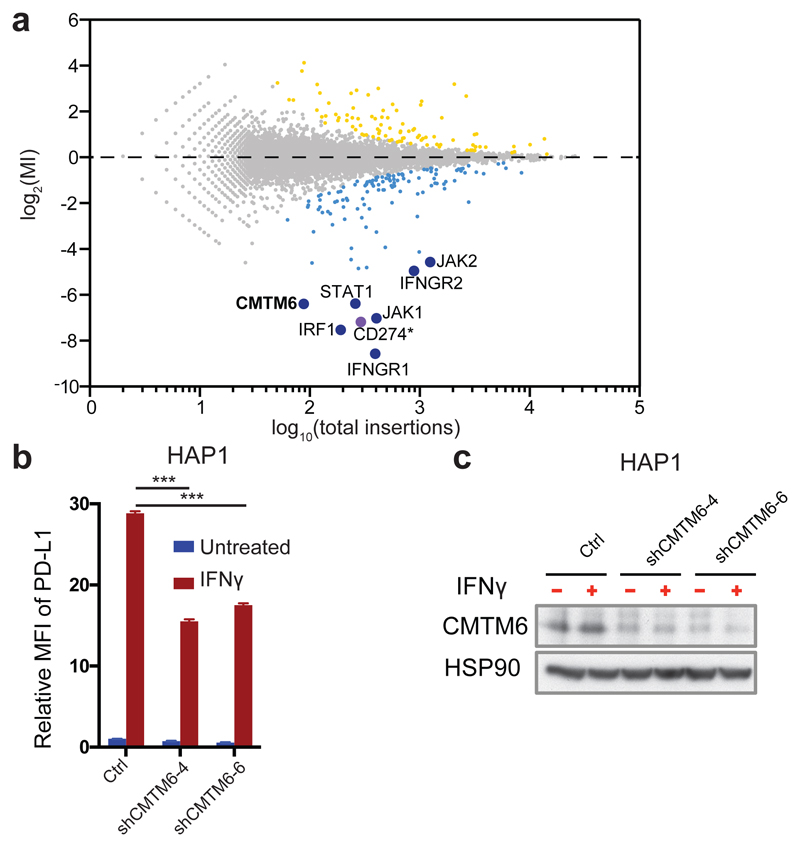
Antibodies that block the PD-1 – PD-L1 axis are currently evaluated in approximately 800 clinical studies and have been approved for 7 different tumor types. In addition, expression of PD-L1 on either tumor cells or on tumor-infiltrating immune cells identifies patients that are more likely to respond to these therapies16,17. In view of the limited understanding of the regulation of PD-L1 expression, we set out to identify PD-L1 protein regulators through genetic screening. Interferon gamma (IFNγ) treated haploid HAP1 cells18,19 express high levels of cell surface PD-L1 (Extended Data Fig. 1a). Based on this observation, we performed a fluorescence activated cell sorting (FACS)-based haploid genetic screen for PD-L1 modulators in IFNγ treated HAP1 (Fig. 1a, experimental outline as in 20). The entire IFNγR signaling pathway21 plus IRF1, a known regulator of PD-L1 upon IFNγ exposure10 were identified as strong hits (Fig. 1a, Supplementary table 1), demonstrating the validity of the screen setup. In addition, the PD-L1 gene itself (CD274) showed a strikingly different integration pattern in PD-L1HI and PD-L1LOW cells. Specifically, whereas PD-L1LOW cells showed the expected enrichment of integrations towards the 5’ end of the gene, a strong enrichment of integrations in intron 5 and 6 was observed in PD-L1HI cells (Extended Data Fig. 1b), fully consistent with the recently described negative regulatory role of the PD-L1 3’ UTR11 (Extended Data Fig. 1c).
Figure 1. Identification of CMTM6 as a modulator of PD-L1 expression.
(a) Flow cytometry-based screen for modulators of PD-L1 cell surface expression in HAP1 cells. Dots represent individual genes, X axis indicates the number of disruptive insertions per gene, Y axis the frequency of independent insertions in the PD-L1HI channel over the frequency of insertions in the PD-L1LOW channel for each gene. Light blue and orange dots indicate genes with significant enrichment of insertions (FDR-corrected P-value, FCPv<10-6)27 within the PD-L1LOW and PD-L1HI population, respectively. Dark blue circles indicate known components of the IFNγR signaling pathway plus IRF1 and CMTM6 (in bold). The purple dot represents PD-L1 (CD274*) when excluding integrations downstream of exon 5 (Refseq identifier NM_014143.3). See https://phenosaurus.nki.nl for interactive graphs. (b) Relative PD-L1 cell surface expression in control or independent CMTM6 knockdown HAP1 cells, either with or without IFNγ exposure. (c) Validation of CMTM6 knockdown by Western Blot. Data are representative of one (a) or at least three (b,c) independent experiments, and were analyzed by unpaired t-test (b). Error bars represent s.d. of triplicates (b). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; MI, mutation index.
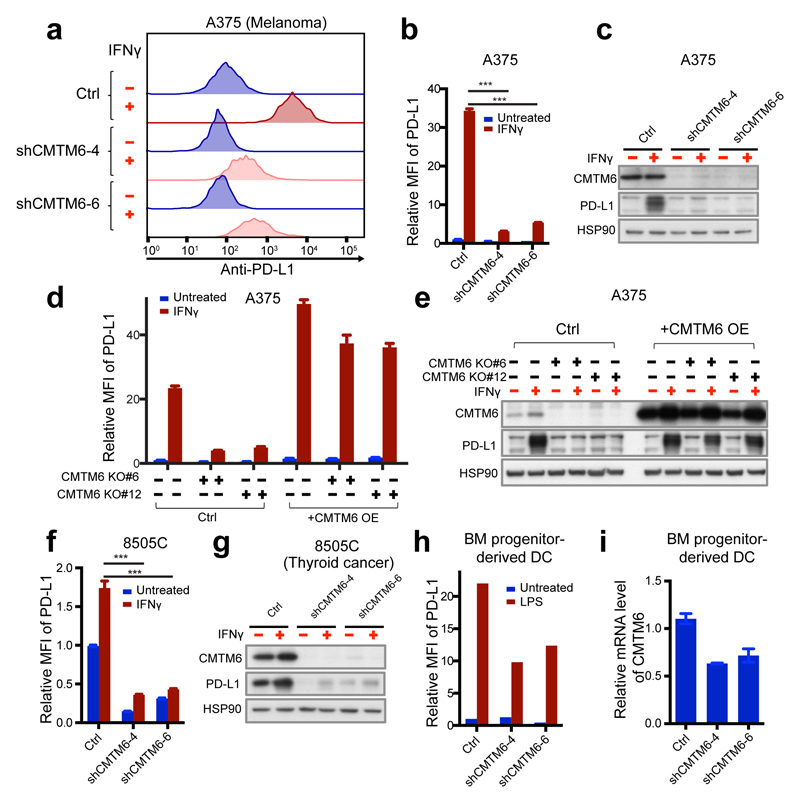
In addition to the above hits, we identified CKLF (Chemokine-like factor)-like MARVEL transmembrane domain containing family member 6 (CMTM6) as one of the most significant hits within PD-L1LOW cells. CMTM6 was not observed in a similar screen for regulators of IRF1 protein levels20, suggesting that its role was independent of the IFNγR pathway. CMTM6 is a ubiquitously expressed transmembrane protein that belongs to a family of 8 MARVEL domain-containing proteins22 for which no clear function has been described. Transcriptome analysis of tumor samples in The Cancer Genome Atlas (TCGA) showed CMTM6 expression in all of the analyzed samples distributed across 30 cancer types, and showed that RNA expression levels of CMTM6 and CD274 are weakly correlated in the majority of tumor types (Extended Data Fig. 2). shRNA mediated knockdown of CMTM6 in HAP1 cells reduced IFNγ-induced PD-L1 expression approximately 2-fold as compared to control (Fig. 1b,c). To assess whether CMTM6 also influences PD-L1 cell surface levels beyond the HAP1 system, we examined the effect of CMTM6 knockdown in a series of tumor lines. In A375 melanoma cells, which only show detectable PD-L1 expression after IFNγ exposure, CMTM6 knockdown prevented IFNγ–induced PD-L1 expression to a large extent (Fig. 2a-c, reduction up to 11 fold). CMTM6-deficient A375 clones generated with CRISPRs/Cas9 likewise showed reduced cell surface and overall PD-L1 protein levels, while lentiviral reconstitution of CMTM6 reverted this phenotype (Fig. 2d,e). In the 8505C thyroid cancer cell line that shows a high basal level of PD-L1 expression, both steady state and IFNγ-induced PD-L1 cell surface and total protein levels were substantially reduced by CMTM6 knockdown (Fig. 2f,g, up to 7- and 5-fold). In total, we assessed the effect of CMTM6 knockdown in 12 human tumor lines, representing melanoma (Fig. 2a-c, Extended Data Fig. 3a-d), thyroid cancer (Fig. 2f,g), colorectal cancer (Extended Data Fig. 3e,f,i,k), lung cancer (Extended Data Fig. 3l,n-p) and CML (Fig. 1b,c), and also in three short term cultures of melanoma xenografts (Extended Data Fig. 3q), and consistently observed diminished expression of PD-L1 (between 2 and 11 fold) upon knockdown of CMTM6. Reduced PD-L1 cell surface levels were likewise observed when cells were stained with recombinant PD-1-Fc protein (Extended Data Fig. 3h,j,m). PD-L1 can both be expressed by cancer cells and by infiltrating immune cells, and PD-L1 expression on immune cells may contribute to T cell inhibition16,17. To assess whether CMTM6 also influences PD-L1 levels in primary human dendritic cells (DCs), we generated DCs from human bone marrow (BM) progenitors23. Comparison of LPS-induced PD-L1 expression in control and CMTM6 knockdown DCs showed that partial knockdown of CMTM6 resulted in partial reduction of PD-L1 cell surface levels (Fig. 2h,i).
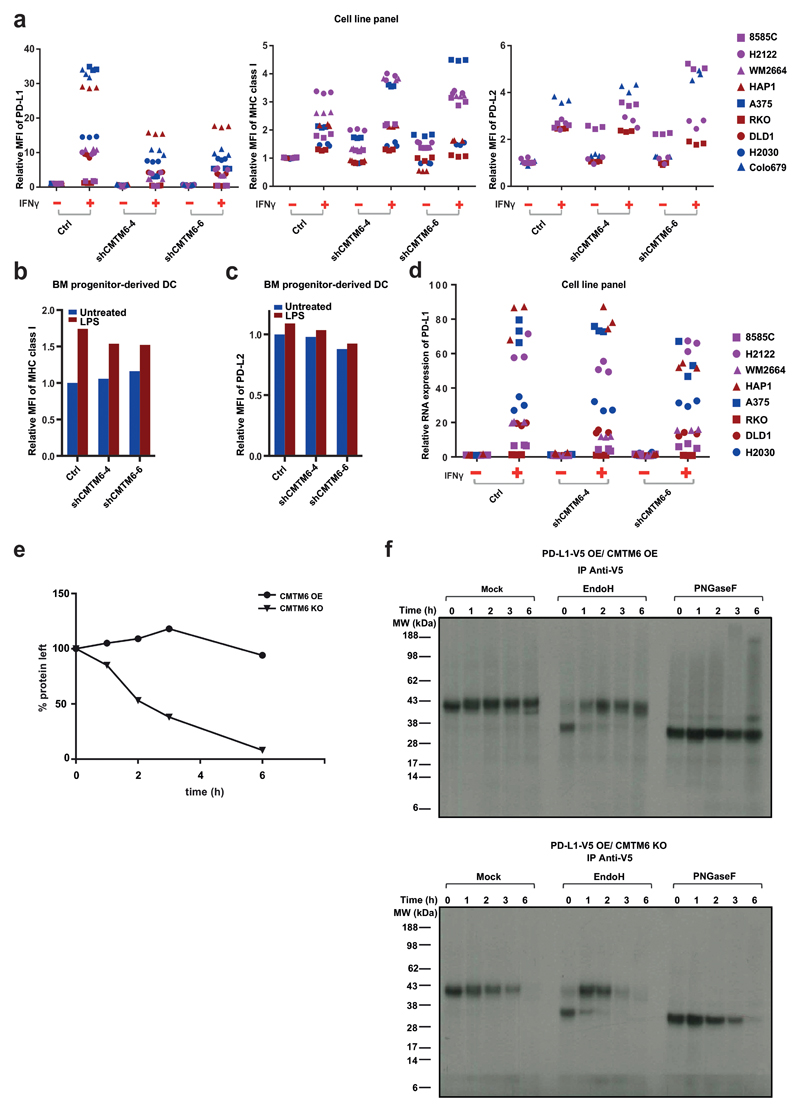
Figure 2. CMTM6 regulates PD-L1 expression in different tumor types and primary dendritic cells.
(a,b,f) Relative PD-L1 cell surface expression in control or independent CMTM6 knockdown A375 melanoma cells (a-b), 8505c thyroid cancer cells (f), either with or without IFNγ exposure. (c,g) Western blot analysis of CMTM6 and PD-L1 expression in control or independent CMTM6 knockdown A375 melanoma cells (c) or 8505c thyroid cancer cells (g), either with or without IFNγ exposure. Each cell line was tested in at least three independent experiments, representative results are shown. (d) Flow cytometric analysis, and (e) western blot analysis of PD-L1 expression in parental, CMTM6 deficient, CMTM6 overexpressing and CMTM6 reconstituted A375 melanoma cells. (l) Flow cytometric analysis of PD-L1 expression in control or independent CMTM6 knockdown primary BM progenitor-derived DCs, either with or without LPS exposure. (m) Knockdown efficiency of CMTM6 in primary BM progenitor-derived DCs. Data are representative of at least three (a-k) or two (l, m) independent experiments and were analyzed by unpaired t-test (b,f,h,j). Error bars represent s.d. of triplicates (b,d,f,h,j,m). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; BM, bone marrow; DC, dendritic cell.
The above data establish CMTM6 as a modulator of PD-L1 protein levels. Whereas in some tumor lines, the effect of CMTM6 knockdown is profound (e.g. A375, 8505c), in others it is moderate (e.g. HAP1), suggesting the possible existence of (an) additional regulator(s). We therefore generated CMTM6-knockout HAP1 cells and performed a modifier screen, with the aim to identify genetic factors that selectively regulate PD-L1 in the absence of CMTM6. Consistent with the primary screen, genes mediating IFNγR signaling were prominent hits. As expected, in this setting, integrations within the CMTM6 locus were no longer enriched within the PD-L1LOW cell population (Fig. 3a, Supplementary table 2). Strikingly, in CMTM6-deficient HAP1 cells, CMTM4, another member of the CMTM family with 55% homology to CMTM6 was identified as a positive regulator of PD-L1 expression (Fig. 3a). Notably, while CMTM4 is a highly significant hit in CMTM6-deficient HAP1 cells (36.7-fold, FCPv≤10-314), it is not in CMTM6 proficient HAP1 cells (0.9-fold, FCPv=0.94789), suggesting that in this system, CMTM4 functions as a back-up regulator of PD-L1 expression.
Figure 3. Identification of CMTM4 as a second PD-L1 regulator.
(a) Haploid genetic screen for modulators of PD-L1 cell surface expression in CMTM6-deficient HAP1 cells. (b) PD-L1 surface expression of parental, CMTM6 knockdown, CMTM4 knockdown, or double knockdown H2030 cells, either with or without IFNγ exposure. (c) Flow cytometric analysis and (d) western blot analysis of PD-L1 expression in parental A375, CMTM6 KO A375, and CMTM6 KO A375 reconstituted with the indicated CMTM family member, either with or without IFNγ exposure. Data are representative of one (a), two (b) or three (c,d) independent experiments and were analyzed by unpaired t-test (b). Error bars represent s.d. of triplicates (b). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; KO, knockout. MI, mutation index.
To validate these data, we transduced H2030 cells, in which CMTM6 depletion modestly suppresses PD-L1 expression and CMTM4 is highly expressed, with shRNAs for CMTM4 and CMTM6, either separately or in combination. In these cells, silencing of CMTM6 led to repression of IFNγ-induced PD-L1 expression that was further enhanced when CMTM4 was simultaneously targeted (Fig. 3b, Extended Data Fig. 4a). More directly, ectopic expression of CMTM4 could fully restore IFNγ-induced PD-L1 expression in CMTM6-knockout cells (Extended Data Fig. 4b,c). To understand whether regulation of PD-L1 expression is a specific property of CMTM6 and 4, we individually introduced FLAG-tagged versions of all CMTM family members into CMTM6-deficient A375 cells. Contrary to what was observed upon CMTM6 and 4 introduction, expression of other CMTM members (detected for CMTM1, 3, 5, and 7, Extended Data Fig. 4d,e) did not induce a substantial increase in PD-L1 expression, as assessed by either flow cytometry or Western blot analysis (Fig. 3c,d).
To study the mechanism by which CMTM6 regulates PD-L1 levels, we first assessed the relationship between CMTM6 expression and PD-L1 mRNA levels. Comparison of PD-L1 mRNA levels and cell surface protein levels at different time points after IFNγ stimulation revealed that while CMTM6 depletion greatly reduced PD-L1 cell surface levels, induction of PD-L1 mRNA by IFNγ was not substantially altered (Fig. 4a,b, Extended Data Fig. 5d). Notably, both in cell lines (Extended Data Fig. 5a) and in primary DCs (Extended Data Fig. 5b,c) levels of MHC-I and PD-L2 protein were not significantly affected by CMTM6 inhibition, indicating that while CMTM6 regulates PD-L1 at the protein level, it is not a general regulator of protein translation or stability.
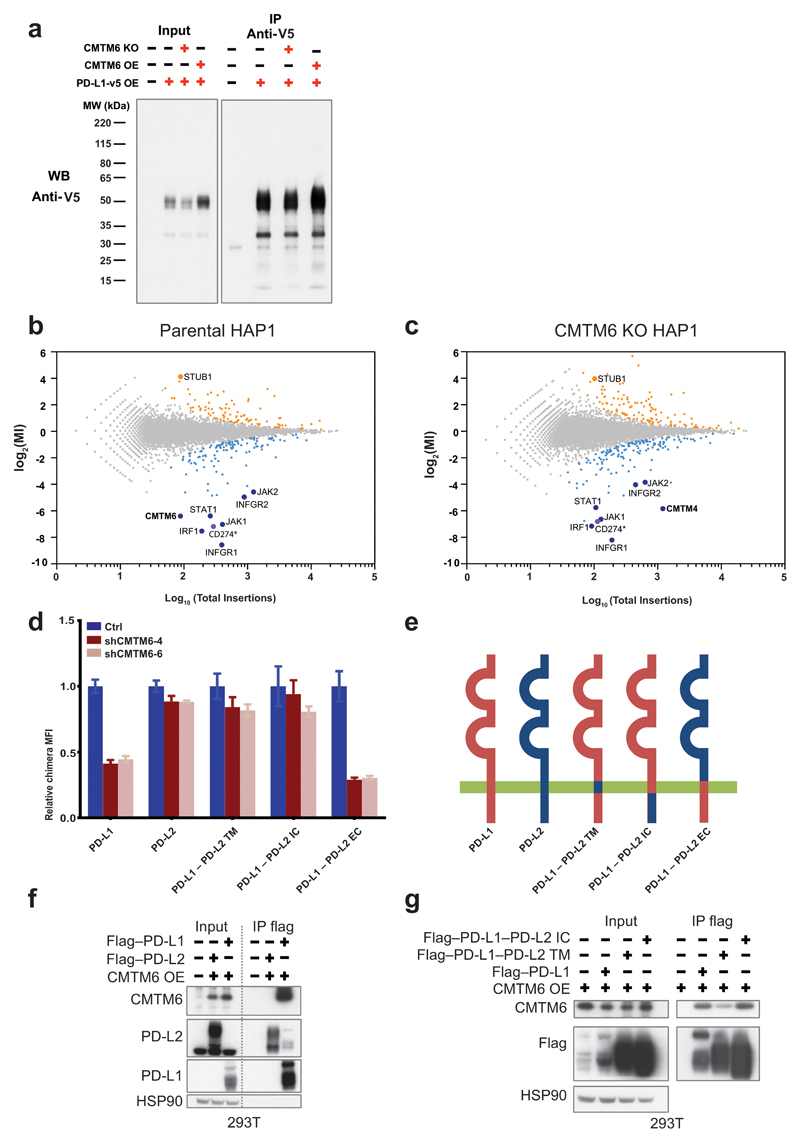
Figure 4. CMTM6 forms a molecular partner of PD-L1 and regulates PD-L1 protein stability.
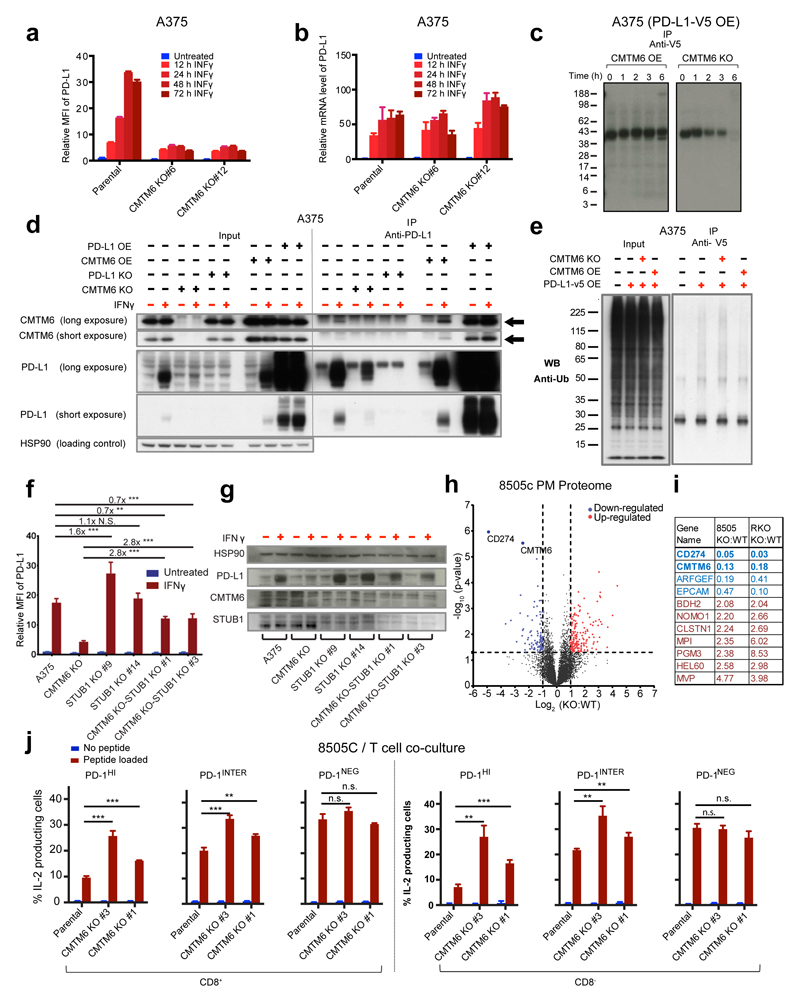
Time course of PD-L1 surface protein (a) or mRNA (b) levels in A375 cells upon IFNγ exposure (c) SDS-PAGE analysis of anti-V5 immunoprecipitates from V5-PD-L1 overexpressing CMTM6 KO or CMTM6 overexpressing A375 cells at different time points after 35S methionine/cysteine labeling. (d) Immunoblots of lysates or anti-PD-L1 immunoprecipitates from the indicated A375 melanoma cells, either with or without IFNγ exposure Arrows indicate CMTM6. (e) Immunoblots of lysates or anti-V5 immunoprecipitates from V5-PD-L1 overexpressing parental, CMTM6 KO or CMTM6 overexpressing A375 cells. PD-L1 surface protein (f) or lysate immunoblots (g) of parental, CMTM6 KO, STUB1 KO or double KO A375 cells. (h) Comparative membrane-fractionated mass spectrometry of 4 CMTM6 proficient and 4 CMTM6 deficient 8505c clones. (i) Overview of proteins consistently found up- or down-regulated upon CMTM6 removal in both 8505c and RKO. (j) IL-2 production by PD-1HI, PD-1INTER and PD-1NEG primary human T cells obtained by transduction with the MART-1 specific 1D3 TCR and PD-1, and co-cultured with unloaded or MART-I peptide-loaded parental or CMTM6 KO 8505C cells. Data are representative of two (a, b), one (c,h) or three (d,e,f,g,j) independent experiments and were analyzed by unpaired t-test (f,j). Error bars represent s.d. of triplicates (a,b,f,j). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; KO, knockout; OE, overexpression.
To determine where in the PD-L1 protein life cycle CMTM6 exerts its effect, CMTM6-deficient and CMTM6-overexpressing A375 cells were transduced with a V5-tagged PD-L1 gene. Immunoprecipitation of PD-L1-V5 at different time points after a 1-h 35S pulse labeling demonstrated a much more rapid decay of PD-L1 in the absence of CMTM6 (fraction PD-L1 remaining at t= 6h; 94% versus 8%, Fig. 4c, Extended Data Fig. 5e). Notably, PD-L1 resistance to deglycosylation by Endoglycosidase H, was equally efficient in both cell populations, indicating that CMTM6 influences PD-L1 protein fate after egress from the endoplasmic reticulum (Extended Data Fig. 5f).
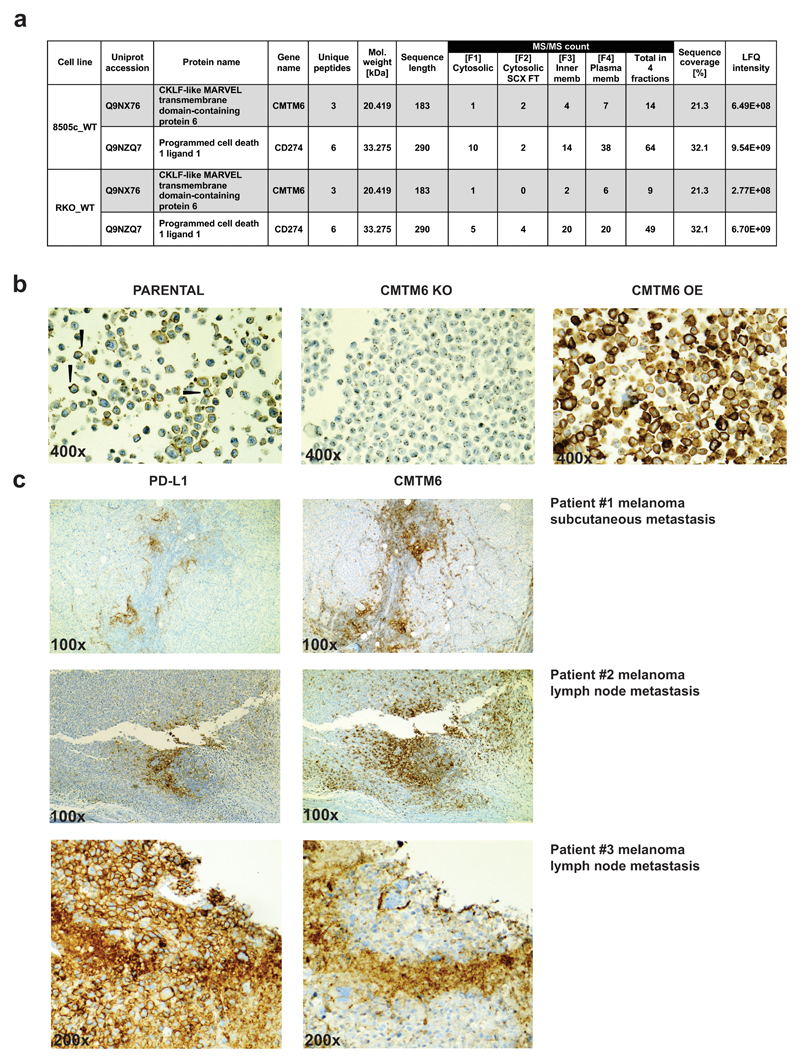
To reveal the cellular localization of endogenous CMTM6, we performed mass spectrometry analysis of different subcellular fractions, demonstrating that endogenous CMTM6 is predominantly present within the plasma membrane fraction (Extended Data Fig. 6a) In addition, immunohistochemical analysis confirmed the presence of CMTM6 at the cell membrane (Extended Data Fig.6b). Furthermore, immunohistochemical analysis of 9 melanomas revealed CMTM6 protein expression in human tumors, and also showed that PD-L1 staining in 8 of these samples was restricted to areas with clear CMTM6 expression (Extended Data Fig. 6c). Similarly, in 3 out of 5 PD-L1 positive lung cancer samples, we observed PD-L1 localization in CMTM6 positive areas.
A hypothesis arising from the above data is that CMTM6 and PD-L1 could interact molecularly. To test this, we performed immunoprecipitations of PD-L1 followed by Western blot analysis of CMTM6, and vice versa. In lysates from A375 cells or 8505C cells, anti-PD-L1 antibody co-immunoprecipitated CMTM6. Likewise, PD-L1 was present in anti-CMTM6 immunoprecipitates. As expected, co-immunoprecipitation of PD-L1 and CMTM6 in A375 was dependent upon PD-L1 induction by IFNγ. As a further control for antibody specificity, co-immunoprecipation of both CMTM6 and PD-L1 was abrogated upon gene inactivation of the partner molecule (Fig. 4d, Extended Data Fig. 7a,b). Co-immunoprecipitation was likewise observed for PD-L1 and CMTM4, and CMTM4 and 6 (Extended data Fig. 7c,d).
To understand how CMTM6 influences PD-L1 degradation, wild type, CMTM6 KO, and CMTM6 overexpressing A375 were transduced with a V5-tagged PD-L1 gene and ubiquitination of PD-L1 was analyzed. In the absence of CMTM6, the amount of ubiquitinated PD-L1 was increased, in spite of the overall lower PD-L1 levels (Fig. 4e, Extended Data Fig.8a), suggesting that CMTM4/6 may protect PD-L1 from ubiquitination. Intriguingly, STUB1, an E3 ubiquitin ligase that has amongst others been implicated in degradation of Foxp3 in regulatory T cells24, was identified as a negative regulator of PD-L1 expression in both haploid genetic screens (Extended Data Fig. 8b,c). To assess whether STUB1 affects PD-L1 degradation, we disrupted STUB1 in either CMTM6 proficient or deficient A375 cells. Deletion of STUB1 resulted in a more profound increase in PD-L1 levels in CMTM6 deficient than in CMTM6 proficient cells, identifying STUB1 as an E3 ligase that causes destabilization of PD-L1 (Fig. 4f,g), either by direct modification of one of the lysines in the PD-L1 cytoplasmic domain or indirectly. Consistent with the model that CMTM6 may protect PD-L1 by preventing ubiquitination, cell surface levels of PD-L1/L2 fusion proteins are only influenced by CMTM6 when carrying the PD-L1 transmembrane and intracellular domain (Extended Data Fig. 8d,e). Exploiting the fact that CMTM6 and PD-L2 do not show detectable association (Extended Data Fig. 8f), we also used PD-L1/L2 fusion proteins to demonstrate that the PD-L1 transmembrane domain is required for efficient interaction (Extended Data Fig. 8g). In line with the role of the PD-L1 transmembrane and intracellular domain in CMTM6-mediated stabilization, orientation mapping of CMTM6 revealed that a large part of the molecule is located within the cytosol and cell membrane (Extended Fig. 9).
In view of the broad RNA expression pattern of CMTM6, we wished to assess the effects of CMTM6 on the membrane proteome in an unbiased manner. Mass spectrometric analysis of a series of independent CMTM6 deficient and proficient clones revealed that, both within the RKO colorectal cancer line and the 8505c thyroid cancer cell line, PD-L1 was the most significantly influenced hit (Fig. 4h,i, Extended Data Fig. 10a,b). Expression of PD-L1 affects T cell responsiveness in a quantitative manner, with higher levels of PD-L1 expression leading to an increased impairment of T cell survival/ activity11,25. To determine whether CMTM6 influences PD-L1 mediated T cell suppression, we incubated mixtures of MART-I TCR transduced T cells that expressed different levels of PD-1 with antigen-loaded CMTM6-deficient or –proficient 8505C or A375 cells. IL-2 production of PD-1INTER and PD-1HI T cells upon encounter of CMTM6-proficient tumor cells was reduced relative to that of PD-1NEG T cells. However, upon CMTM6 disruption in either 8505C or A375 tumor cells, IL-2 production of PD-1 expressing T cells was significantly restored (Fig. 4j, Extended Data Fig. 10c-e).
Recent work has revealed a number of mechanisms of transcriptional and post-transcriptional (dys)regulation of the PD-L1 gene in tumor cells10–15. Here we identify CMTM6 and CMTM4 as regulators of PD-L1 protein stability. Based on the available data we conclude that CMTM6/4, the two most closely related members of the CMTM family (Extended Data Fig. 4f), influence PD-L1 expression across a range of cell types. Furthermore, the observations that I) CMTM6 affects PD-L1 protein stability at late time points after biosynthesis, II) CMTM6, CMTM4 and PD-L1 interact, as shown by co-immunoprecipitation, III) CMTM6 is largely located at the cell surface collectively sketch a model in which CMTM6 interacts with PD-L1 at the tumor cell surface and thereby protects it from degradation. In line with this, CMTM6 influences the levels of PD-L1 ubiquitination and absence of the STUB1 E3 ubiquitin ligase partially reverts the CMTM6 KO phenotype. Intriguingly, for one of the other CMTM family members, CMTM7, cell surface expression has been described in association with the B cell receptor (BCR) complex, where it may contribute to BCR signaling26. It could be speculated that CMTM6 may also fulfill a similar role in the immunological synapse between T cells and tumor cells or antigen presenting cells (APCs). Finally, the co-localization of PD-L1 and CMTM6 in melanoma samples and the observation that CMTM6 depletion ameliorates PD-L1 mediated T cell suppression suggest a potential value of CMTM6/4 as therapeutic targets, either in isolation, or to enhance the effectiveness of the current PD-L1/PD-1 blocking therapies.
Methods
Cell lines
A375, DLD1, RKO, H2030, and H2122 cells were purchased from American Type Culture Collection (ATCC). 8505C was purchased from Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH (DMSZ). WM2664 and COLO679 cells were kind gifts from Rene Bernards (The Netherlands Cancer Institute). Short term cell lines from patient derived melanoma xenografts were generated as described28 and were a kind gift of Daniel Peeper and Kristel Kemper. HAP1 cells have been described previously19. HAP1 cells were cultured in IMDM (ThermoFisher Scientific) supplemented with 10% fetal calf serum (FCS, Sigma), 100U/ml penicillin–streptomycin (ThermoFisher Scientific) and L-glutamine (ThermoFisher Scientific); A375 and short term melanoma xenograft cultures were maintained in DMEM supplemented with 10% FCS (Sigma) and 100U/ml penicillin/streptomycin (ThermoFisher Scientific). All other cell lines were cultured in RPMI supplemented with 10% FCS (Sigma) and 100U/ml penicillin/streptomycin (ThermoFisher Scientific). IFNγ treatment was performed over a period of 48h at a concentration of 25 ng/ml, if not indicated otherwise.
Identification of genetic regulators
The approach as described in Brockmann et al. was followed to identify regulators of PD-L1 abundance. Mutagenized HAP1 libraries (starting with either wild-type cells or CMTM6-deficient HAP1 cells) were expanded to approximately 1.5x109 cells and subsequently treated with 0.5 ng/ml IFNγ (Peprotech) for 24 hours to induce expression of PD-L1. Subsequently, approximately 3x109 cells were dissociated using trypsin-EDTA (Life technologies), washed with PBS and stained with FITC labelled anti-PD-L1 antibody (BD pharmingen) at 1:20 dilution for 30’ at RT in PBS containing 0.5% w/v bovine serum albumin (Sigma) and 0.2% w/v sodium azide (Sigma). Subsequently, cells were washed three times with PBS containing 1% FCS and stained with FITC labeled polyclonal goat anti-mouse Ig (BD pharmingen) at 1:100 dilution for 30’ at RT in PBS containing 0.5% w/v bovine serum albumin (Sigma) and 0.2% w/v sodium azide (Sigma) to allow signal amplification. Following two washes with PBS containing 1% FCS and one wash with PBS, stained cells were passed through a 40 µm strainer (BD FalconTM) and subsequently fixed using BD fix buffer I (BD biosciences) for 10 minutes at 37°C, followed by a wash with PBS containing 1% FCS. Subsequently, cells were permeabilized by suspension in cold (-20°C) BD permeabilization buffer (BD biosciences) while vortexing, and incubated on ice for 30 minutes prior to incubation with 100 µg/ml RNAse A (Qiagen, Germany) and 10 µg/ml propidium iodide (Cayman Chemical) at 37°C temperature for 30 minutes. Alternatively, cells were subjected to treatment with 3 µM 4',6- diamidino-2-phenylindole (DAPI) for 30 minutes. Stainings were concluded by a final wash in PBS 10% FCS. Following staining, cells were sorted on a Biorad S3 Cell sorter (Biorad) or a Moflo Asterios cell sorter (Beckman Coulter) to collect the 1-5% of cells with the highest and lowest PD-L1 staining intensity and 1n DNA content. Sorted cells were used for isolation of genomic DNA and retroviral gene-trap insertion sites were retrieved, mapped and analyzed as described in Brockmann et al. For PD-L1 it has been described that alterations of the 3’ portion of the gene can stabilize the gene product and lead to higher PD-L1 proteins levels11. As this was also recapitulated by our gene-trap insertion method (gene-trap integrations into the 3’ portion of the gene resulting in increased rather than decreased staining intensity for PD-L1), integrations in the portion of the gene that lies downstream of exon 5 (Refseq identifier NM_014143.3) were disregarded where indicated.
Generation of knockout cell lines
Knockout cell lines were generated using the CRISPR/Cas9 system. To generate knockout HAP1 cells, cells were transfected with px330 vector (Addgene #42230) encoding a gRNA for the gene of interest and a vector encoding a gRNA for the zebrafish TIA gene (5'- ggtatgtcgggaacctctcc-3'), as well as a P2A-blasticidin resistance cassette flanked by two TIA target sites. This allows incorporation of the blasticidin resistance gene into the locus of interest, resulting in a stable knockout, essentially as described29. Following blasticidin selection (10 µg/ml), resistant clones were expanded.
To generate knockout A375 and 8505c cells, cells were transfected with pLentiCRISPRv2 vector (Addgene #52961) encoding gRNAs targeting non-overlapping regions of the CMTM6 gene 30. Following puromycin selection (2 µg/ml, for 2 days), single cell clones were expanded and gene disruptions were validated by sequencing and Western blot analysis.
The gRNA sequence CCGGGTCCTCCTCCGTAGTG was used to generate the A375 CMTM6 knockout clone “CMTM6 KO#6” and the 8505C CMTM6 knockout clone “CMTM6 KO#1”, the gRNA sequence TCACAATGTACTTTATGTGG was used to generate the A375 CMTM6 knockout clone “CMTM6 KO#12” and the 8505C CMTM6 knockout clone “CMTM6 KO#3”. The gRNA sequence ACTGCTTGTCCAGATGACTT was used to generate the A375 PD-L1 KO clone and the gRNA sequence GGAGATGGAGAGCTATGATG was used to generate all the STUB1 KO clones.
Immunoprecipitation, SDS-PAGE and Western blot analysis
Cells for Western blot analysis were seeded in 6-well plates and cultured as described in figure legends. To harvest cell lysates, cells were washed with PBS and lysed with RIPA buffer supplemented with freshly added protease inhibitor cocktail (Roche). After incubation on ice for 30 minutes, cell lysates were subjected to centrifugation at 20,000g for 15 minutes at 4°C. Supernatants were subsequently processed using Novex NuPAGE Gel Electrophoresis Systems, according to the manufacturer’s instructions (ThermoFisher Scientific).
Cells for (co)immunoprecipitation experiments were seeded in 15-cm dishes and cultured as described in figure legends, using 5 million cells per immunoprecipitation reaction. Cells were washed with cold PBS buffer and lysed in CHAPS buffer (1% CHAPS, 50 mM TRIS-HCl pH 7.5, 150 mM NaCl). For the detection of protein ubiquitination, cells were lysed in the denaturing buffer (50 mM Tris-HCl, 0.5 mM EDTA and 1% SDS) followed by heating at 95 °C for 10 minutes and then quenched by adding 9 volumes of quenching buffer (0.5% Triton X-100, 20 mM Tris-HCl (pH 8.0), 137 mM NaCl, 10% glycerol, 2 mM EDTA). Protease inhibitor cocktail (Roche) was freshly added to all buffers. Cell lysates were incubated on a rotator for 30 minutes at 4°C, and then centrifuged at 20,000g for 15 minute at 4°C. Supernatants were subsequently processed using Dynabeads® Protein A or Protein G for Immunoprecipitation (ThermoFisher Scientific), and the indicated antibodies. The final elute was processed and Western blot analysis was performed using Novex NuPAGE Gel Electrophoresis Systems, according to the manufacturer’s instructions (ThermoFisher Scientific).
Pulse Chase and EndoH-PNGaseF treatment
V5-tagged PD-L1 transduced CMTM6 overexpressing A375 cells, and V5-tagged PD-L1 transduced CMTM6 knockout A375 cells were cultured in methionine- and cysteine-free medium for 1h at 37°C. Cells were then pulse labeled with 0.5 mCi/ml [35S]Cys/[35S]Met (PerkinElmer) for 1 hour. Cells were washed with PBS to remove residual [35S]Cys/[35S]Met, and then cultured in regular medium with extra ‘cold’ methionine and cysteine for 0, 1, 2, 3 and 6h. Cell samples were lysed and used for immunoprecipitation with anti-V5 antibody (ThermoFisher) immobilized on protein A or protein G coated beads (ThermoFisher). Immunoprecipitates were either left untreated or treated with EndoH or PNGaseF (New England Biolabs), according to the manufacturer’s instructions.
Immunoprecipitates were run on NuPAGE 4-12% gels. Gels were treated with 1M NaSalicylate pH5.6 before drying, and then analysed on Fujifilm BAS-MP phosphor imager screens. Quantification was performed using a Fujifilm FLA-3000 phosphorimager and AIDA image analyzer software. Gels were exposed to film using intensifier screens at -80 C.
Viral vectors
Lentiviral shRNA vectors were retrieved from the arrayed TRC human genome-wide shRNA collection. Additional information is available at http://www.broadinstitute.org/rnai/public/clone/search using the TRCN number. The following lentiviral shRNA vectors were used:
shCMTM6-4: TRCN0000127888
shCMTM6-6: TRCN0000130177
shCMTM4-1: TRCN0000142717
shCMTM4-2: TRCN0000142470
PD-L1, PD-1, PD-L2, CMTM6, CMTM4 and PD-L1 – PD-L2 chimeras expressing lentiviral vectors were generating by insertion of the relevant gblock (IDT) into a pCDH-CMV-MCS-EF1-Puro (CD510B-1, System Bioscience)-derived vector in which the puromycin resistance cassette was substituted with a blasticidin resistance cassette. PD-L1 – PD-L2 chimeras were generated as follow:
PD-L1 – PD-L2 TM: aa 1-18 PD-L1 – DYDDDDKD – aa 19-238 PD-L1 – aa 221-242 PD-L2 – aa 263-290 PD-L1
PD-L1 – PD-L2 IC: aa 1-18 PD-L1 – DYDDDDKD – aa 19-262 PD-L1 – aa 246-273 PD-L2
PD-L1 – PD-L2 EC: aa 1-20 PD-L2 – DYDDDDKD – aa 21-221 PD-L2 – aa 232-290 PD-L1, in which DYDDDDKD refers to the sequence of the FLAG epitope tag. For generation of chimeras isoform NP_054862.1 (PD-L1) and NP_079515.2 (PD-L2) were used.
V5 tagged PD-L1 was retrieved from the CCSB-Broad Lentiviral Expression Library (#ccsbBroad304_15876). CMTM family members were ordered as individual gblocks (IDT) coding for the different family members using Ensemble gold transcripts ENST00000379500.6 (CMTM1), ENST00000268595.2 (CMTM2), ENST00000361909.8 (CMTM3), ENST00000330687.8 (CMTM4), ENST00000339180.8 (CMTM5), ENST00000205636.3 (CMTM6), ENST00000334983.9 (CMTM7), ENST00000307526.3 (CMTM8) c terminally fused with a FLAG tag, preceded by a short AAV-linker and cloned into the pMX-IRES-Blast vector using restriction enzymes BglII and SalI (CMTM1 and CMTM4), EcoRI and NotI (CMTM2) or BamHI and SalI (CMTM3, CMTM5, CMTM6, CMTM7, CMTM8). The retroviral vector pBABE-puro encoding C-terminally FLAG-tagged CMTM6 (pBp-CMTM6-FLAG) was generated by cloning a gblock for CMTM6 (ENST00000205636.3) digested with BamHI and XhoI into pBABE-puro digested with BamHI and SalI.
For production of lentiviral particles, the described plasmids were co-transfected into HEK293T cells along with packaging plasmids (pPAX2, pVSV-G). Two days after transfection, lentiviral supernatant was harvested and used for transduction. Retroviral particles were produced and purified as described for HAP1 mutagenesis, except that multiple harvests and ultracentrifugation were omitted. Two days after transduction, cells were selected by exposing them to blasticidin or puromycin.
Antibodies
The following antibodies were used for Western Blot analyses and immunoprecipitations: anti-HSP90: H114 (SantaCruz), anti-CMTM6: HPA026980 (Atlas) or anti-CMTM6 monoclonal antibody directed against a peptide in the C-terminal domain of CMTM6 generated by Absea, anti-CMTM4: HPA014704 (Atlas), anti-PD-L1 for Western blot analysis: 405.9A11 (Cell Signaling), anti-PD-L1 for immunoprecipitation: E1L3N (Cell Signaling), normal rabbit IgG: #2729 (Cell Signaling), anti-FLAG tag: M2 (Sigma), anti-V5 tag: R960-25 (Thermofisher), anti-STUB1 sc 133066(Santa Cruz), anti-Ubiquitin antibody #3933 (Cell signaling), Goat anti-mouse IgG (H+L)- HRP conjugate (BIO-RAD), and Goat anti-rabbit IgG (H+L)- HRP conjugate (BIO-RAD). The following antibodies were used for flow cytometry: anti-PD-L1: M1H1 (eBioscience), anti-PD-L2: 24F.10C12 (Biolegend), anti-MHC-I: BB7.2 (BD bioscience), anti-murine TCR: H57-597 (BD bioscience), anti-CD8: RPA-T8 (BD bioscience), anti-CD3: SK7 (eBioscience), anti-PD-1: eBioJ105 (eBioscience), anti-IL-2: 554567 (BD bioscience). The following antibodies were used for immunohistochemistry: anti-PD-L1: 22C3 (Dako); anti-CMTM6 monoclonal antibody directed against a peptide in the C-terminal domain of CMTM6, was generated by Absea.
RNA isolation, first strand cDNA synthesis and qRT-PCR
Total RNA was isolated from cells using the RNeasy Mini Kit (Qiagen). cDNA was obtained by reverse transcription using the Maxima First Strand cDNA Synthesis Kit for RT-qPCR (ThermoFisher Scientific), according to the manufacturer’s instructions. SensiFAST SYBR® No-ROX Kit (Bioline) was used for RT-PCR gene expression analysis, carried out on Roche LightCycler® 480 platform. Relative mRNA levels were normalized to GAPDH mRNA levels.
-
Primer sets used were as follows:
CD274-F:ATTTGGAGGATGTGCCAGAG
CD274-R:CCAGCACACTGAGAATCAACA
GAPDH-F:AAGGTGAAGGTCGGAGTCAA
GAPDH-R:AATGAAGGGGTCATTGATGG
IL-2 production assay
Human peripheral blood T cells (Sanquin) were activated and transduced with a retroviral vector encoding the MART-I specific 1D3 TCR as described 31 and with a lentiviral vector encoding PD-1, and MART-I TCR and PD-1 expression was validated by flow cytometry. Both 8505c (parental and CMTM6 KO) and A375 cells (parental, PD-L1 overexpressing, and CMTM6KO PD-L1 overexpressing) were pulsed with 10 ng/ml MART-126-35 peptide for 1 hour at 37°C. Next, 1 × 105 transduced cells were incubated with 1 × 105 peptide-pulsed cells or non-pulsed cells in the presence 1 μL/mL Golgiplug (BD Biosciences). After a 5-hour incubation at 37°C, cells were washed and stained with phycoerythrin (PE)-labeled anti murine TCR beta chain, V500 labeled anti-CD8, PerCP Cy5.5 labeled anti-CD3 and PE Cy7 labeled anti-PD-1, and analyzed for IL-2 production by intracellular cytokine staining. Activity of T cells with different levels of PD-1 expression was analyzed by gating on murine TCR beta chain-positive cells expressing low, intermediate, or high levels of PD-1.
Membrane fractionation and MS analysis
Snap-frozen cell pellets were lysed by gentle homogenization in isotonic buffers supplemented with phosphatase inhibitor (PhosSTOP, Roche) and protease inhibitor (cOmplete mini EDTA-free, Roche). Cellular disruption of >95% was confirmed by microscopy. Plasma membrane (F4), inner membrane (F3) and cytosolic (F1) fractions were prepared by differential centrifugation using a plasma membrane purification kit (Abcam, ab65400).
From fractions F1, F3 and F4, 20 ug of proteins was diluted 20 times in 50mM ammonium bicarbonate, reduced in 4mM dithiothreitol (DTT), alkylated in 8mM iodoacetamide (IAA), and digested sequentially at 37oC with 1:75 Lys C (Wako) and 1:50 trypsin (Sigma-Aldrich) for 4 and 12 hours respectively. Digested peptides were acidified to 0.1% formic acid (FA) and purified by strong cation exchange (SCX) STAGE tips, using loading buffer 80% acetonitrile (ACN), 0.1% FA and elution buffer 0.5M ammonium acetate, 20% ACN, 0.1% FA. Eluted peptides were dried by vacuum and 4μg equivalent of peptides was analyzed in a 3hr reverse-phase separation on the UHPLC 1290 system (Agilent) coupled to an Orbitrap Q Exactive HF mass spectrometer (Thermo Scientific). SCX flowthrough from cytosolic fraction (denoted F2) was analysed separately to increase proteome coverage.
RP-nanoLC-MS/MS
Proteomics data were acquired using an UHPLC 1290 system (Agilent) coupled to an Orbitrap Q Exactive HF spectrometer (Thermo Scientific). Peptides were first trapped on a 2 cm x 100 μm Reprosil C18 pre-column (3 μm) and then separated on a 50 cm x 75 μm Poroshell EC-C18 analytical column (2.7 μm). Trapping was performed for 10 min in 0.1 M acetic acid (Solvent A) and elution with 80% ACN in 0.1M acetic acid (Solvent B) in gradients as follows: 10-40% solvent B in 155 min, 40-100% in 3min and finally 100% for 1min. Flow was passively split to 300 nl/min. MS data were obtained in data-dependent acquisition mode. Full scans were acquired in the m/z range of 375-1600 at the resolution of 35,000 (m/z 400) with AGC target 3E6. Top 15 most intense precursor ions were selected for HCD fragmentation performed at normalized collision energy (NCE) 25%, after accumulation to target value of 5E4. MS/MS acquisition was performed at a resolution of 17,500.
Database search
Raw files were processed using MaxQuant version 1.5.3.30 and searched against the human Swissprot database (version May 2016) using Andromeda.
Cysteine carbamidomethylation was set to fixed modification, while variable modifications of methionine oxidation and protein N-terminal acetylation, as well as up to 2 missed cleavages were allowed. False discovery rate (FDR) was restricted to 1% in both protein and peptide identification. Label-free quantification (LFQ) was performed with “match between runs” enabled.
Analysis of CMTM6 RNA levels and correlation between CMTM6 and CD274 RNA levels in TCGA samples
TCGA RNA sequencing data was downloaded from the Broad TCGA genome data analysis center 2015-11-01 release of the standard runs (http://gdac.broadinstitute.org/runs/stddata). For projects where data from multiple sequencing platforms is available, Illumina HiSeq data was preferentially used. The (RSEM) normalized read count field was multiplied by 10^6 to arrive at the reported TPM expression values. Correlation coefficients and associated unadjusted p-values between CMTM6 and CD274 were computed per TCGA sequencing project with Pearson’s method. Two-dimensional kernel density estimates were computed using the MASS::kde2d() function in version 3.3.1 of the R programming language. We computed the correlations between CMTM6 and 104 randomly selected genes (identical between sequencing projects) to obtain a reference distribution of correlation coefficients for CMTM6, specific for each TCGA project. The reported empirical p-values are defined as one minus the quantile of the CMTM6 and CD274 correlation within this reference distribution.
Immunohistochemistry
Immunohistochemistry of the formalin-fixed paraffin-embedded samples was performed on a BenchMark Ultra autostainer (Ventana Medical Systems). Briefly, 3 µm paraffin serial sections were cut, heated at 75°C for 28 minutes and deparaffinised in the instrument with EZ prep solution (Ventana Medical Systems). Heat-induced antigen retrieval was carried out using Cell Conditioning 1 (CC1, Ventana Medical Systems) for 48’ for PD-L1, and 64’ for CMTM6 antibodies at 95°C.
PD-L1 clone 22C3 (Dako) was used at 1:40 dilution, 1 hour at room temperature and CMTM6 clone 1D6 was used directly from hybridoma supernatant at either 1:500 or 1:1000 dilution for tumor samples and 1:100 dilution for cell lines, 1 hour at room temperature. Bound antibody was detected using the OptiView DAB Detection Kit (Ventana Medical Systems). Slides were counterstained with Hematoxylin and Bluing Reagent (Ventana Medical Systems).
Patient melanoma samples were obtained (following Institutional Review Board approval) from the NKI-AVL pathology archive biobank and selected for PD-L1 expression.
Statistical analysis
All student T tests were two tailed under the assumption of equal variance between samples.
Extended Data
Extended data Figure 1. PD-L1 is regulated by IFNγ and by the UTR in HAP1.
(a) Flow cytometric analysis of PD-L1 expression of untreated HAP1 cells and HAP1 cells treated with the indicated concentrations of IFNγ. An IFNγ concentration of 0.5 ng/ml was chosen for the subsequent genetic screen, to allow identification of gene integrations that either enhance or suppress PD-L1 expression. Data are representative of three independent experiments. (b) Schematic representation of the PD-L1 gene and of the gene trap insertions observed in HAP1 cells sorted on the basis of either low or high PD-L1 expression. Note the bias towards integrations within introns 5 and 6 in the PD-L1 gene in PD-L1HI cells relative to PD-L1LOW cells, consistent with the structural variants beyond exon 4 of PD-L1 that have been shown to result in enhanced PD-L1 expression in a subset of adult T-cell leukaemia, diffuse large B-cell lymphoma, and stomach cancers11. (c) Screen data as depicted in Fig. 1, but now with PD-L1 (CD274) data plotted when either including (CD274) or excluding (CD274*) integrations downstream of exon 5 (Refseq identifier NM_014143.3). MI, mutation index.
Extended data Figure 2. RNA expression of CMTM6 in human cancers and correlation with PD-L1 mRNA levels.
Pearson correlation coefficients are shown along with associated unadjusted p-values. As randomly selected genes are on average also weakly positively correlated (not shown), empirical p-values, which represent one minus the quantile of the CMTM6 and CD274 expression correlation coefficient among a reference distribution composed of correlation coefficients between CMTM6 and randomly selected genes, are also depicted. Empirical p-values smaller than .5 denote a stronger correlation between CMTM6 and CD274 than the median observed correlation in the reference distribution. TPM, transcript per million. ACC: adrenocortical carcinoma, BLCA: urothelial bladder carcinoma, BRCA: breast cancer, CESC: cervical squamous cell carcinoma, CHOL: cholangiocarcinoma, COAD: colorectal adenocarcinoma, DLBC: diffuse large B-cell lymphoma, ESCA: esophageal cancer, GBM: glioblastoma multiforme, HNSC: head and neck squamous, KICH: chromophobe renal cell carcinoma, KIRC: clear cell kidney carcinoma, KIRP: papillary kidney carcinoma, LAML: acute myeloid leukemia, LGG: lower grade glioma, LIHC: liver hepatocellular carcinoma, LUAD: lung adenocarcinoma, LUSC: lung squamous cell carcinoma, OV: ovarian serous cystadenocarcinoma, PAAD: pancreatic ductal adenocarcinoma, PCPG: pheochromocytoma and paraganglioma, PRAD: prostate adenocarcinoma, READ: rectum adenocarcinoma, SKCM: cutaneous melanoma, STAD: stomach cancer, TGCT: testicular germ cell cancer, THCA: papillary thyroid carcinoma, UCEC: uterine corpus endometrial carcinoma, UCS: uterine carcinosarcoma, UVM: uveal melanoma
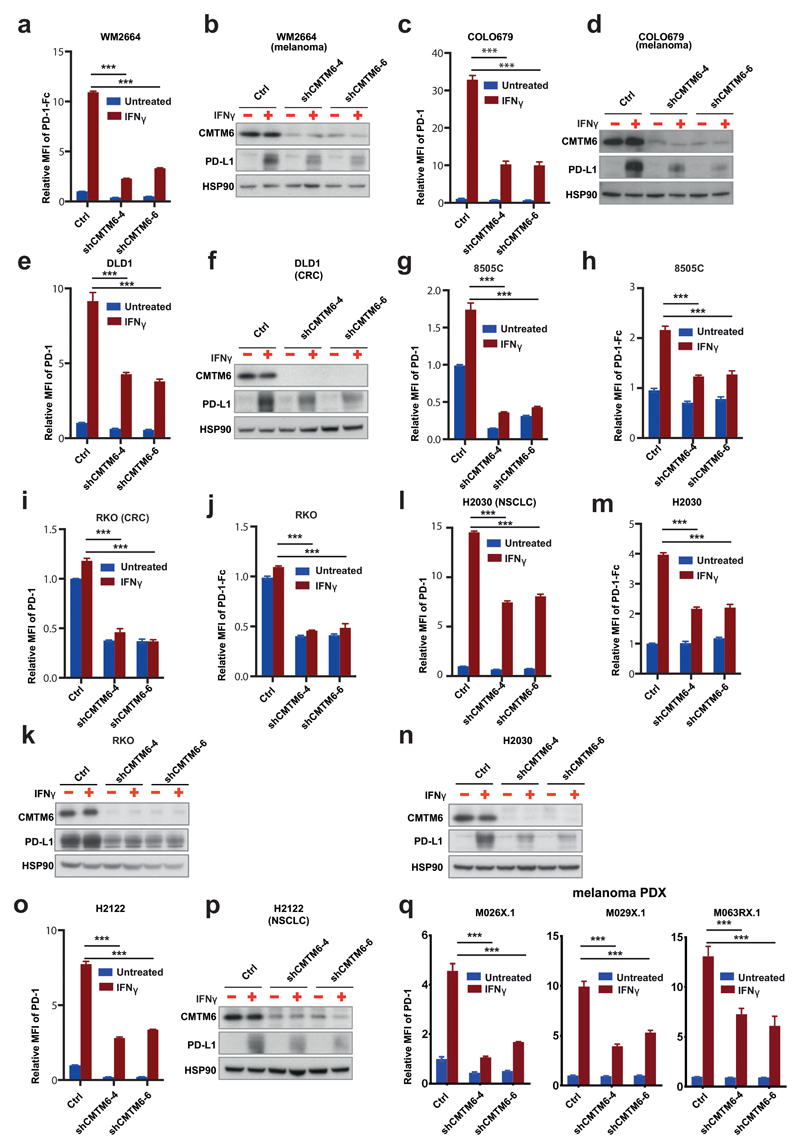
Extended data Figure 3. Regulation of PD-L1 by CMTM6 in different tumor types.
Flow cytometric analysis of PD-L1 expression of WM2664 melanoma (a), COLO679 melanoma (c), DLD1 colorectal cancer (e), H2122 non-small lung cancer (o) cells and three short term cultures of melanoma xenografts (q), in which cells transduced independently with two vectors expressing different CMTM6 shRNAs are compared with cells transduced with control vector. Western blot analysis of CMTM6 and PD-L1 expression in WM2664 melanoma (b), COLO679 melanoma (d), DLD1 colorectal cancer (f), H2122 non-small lung cancer (p) cells, in which cells transduced independently with two vectors expressing different CMTM6 shRNAs are compared with cells transduced with control vector. Western blot analysis of CMTM6 and PD-L1 expression in WM2664 melanoma (b), COLO679 melanoma (d), DLD1 colorectal cancer (f) and H2122 non-small lung cancer (p) cells in which cells transduced independently with two vectors expressing different CMTM6 shRNAs are compared with cells transduced with control vector. Flow cytometric analysis of PD-L1 expression as upon staining with anti-PD-L1 antibody or with PD-1-Fc in 8505c thyroid cancer (g,h), RKO colorectal cancer (i,j) and H2030 non-small lung cancer (l,m) cells and western blot analysis of CMTM6 and PD-L1 expression in RKO colorectal cancer (k) and H2030 non-small cell lung cancer in which cells transduced independently with two vectors expressing different CMTM6 shRNAs are compared with cells transduced with control vector. In all cases, cells treated with IFNγ (25 ng/ml) or left untreated were compared. Panel g is identical to the one depicted in Fig. 2, shown again here to facilitate comparison. , Data are representative of three (c-n), two (a,b,o,p) or one (q) independent experiments and were analyzed by unpaired t-test (a,c,e,g-m,o,q). Error bars represent s.d. of triplicates (a,c,e,g-m,o,q). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; PDX, patient derived xenograft; NSCLC, non-small cell lung cancer; CRC, colorectal cancer.
Extended data Figure 4. CMTM4 and CMTM6, but not other CMTM family members, are regulators of PD-L1.
(a) Validation of CMTM6 and CMTM4 downregulation by Western blot analysis of cells shown in Fig. 3b. (b,c) Ectopic expression of CMTM4 restores IFNγ-induced PD-L1 expression in CMTM6-deficient cells. Two clones of CMTM6-deficient A375 cells (‘CMTM6 KO#6’ and ‘CMTM6 KO#12’) were transduced with retroviral vectors encoding CMTM4 (‘CMTM4 OE’) or CMTM6 (‘CMTM6 OE’) individually. After blasticidin selection, cells were cultured in the absence (‘untreated’) or presence of 25ng/ml IFNγ for 72 hours before analysis by flow cytometry (b), and Western blot analysis (c). Untransduced A375 parental cells served as controls. (d,e) Western blot analysis of expression of the indicated CMTM family members, as determined by staining with an anti-FLAG antibody. Two exposures of the same gel are shown. Expression of CMTM2 and 8 is not detected, and CMTM5 expression is low as compared to that of other CMTM family members. (f) Phylogenetic analysis of the CMTM family by CLUSTALW. CMTM6 and 4 form the two most closely related members. In view of the lack of detectable expression/ low expression observed for CMTM2, 8 and 5, an effect of these CMTM members on PD-L1 protein fate cannot be excluded. However, the observation that CMTM family members 7 and 3 that are more closely related to CMTM4 and 6 do not influence PD-L1 expression makes this unlikely. (g) Results of the flow cytometry based screen as shown in Fig. 1a, with the position of all CMTM family members indicated. Data are representative of two independent experiments (a-e). Error bars represent s.d. of triplicates (b). *P<0.05; **P<0.01; ***P<0.001. MFI, median fluorescence intensity; KO, knockout; OE, overexpression. MI, mutation index.
Extended data Figure 5. CMTM6 downregulation does not affect MHC class I and PD-L2 cell surface levels or PD-L1 mRNA levels and regulates PD-L1 stability after egress from the endoplasmic reticulum.
Flow cytometric analysis of MHC class I and PD-L2 expression in the panel of cell lines tested in Fig. 2 and Extended Data Fig. 3 (a), and in BM progenitor-derived DCs (b,c) in which cells transduced with control vector are compared with cells transduced independently with two vectors expressing different shRNA directed against CMTM6. For cell lines, cells treated with IFNγ (as indicated in the other figure legends) or left untreated were compared, for BM progenitor-derived DCs, cells treated with 500 ng/ml LPS or left untreated were compared. (d) qPCR analyses were performed to quantify relative mRNA levels of PD-L1 in the abovementioned tumor lines. (e) Quantification of the experiment shown in Fig. 4c (f) Immunoprecipitates of the same samples as used in Fig. 4c were either mock treated, treated with EndoH, or treated with PNGaseF to examine the kinetics of protein maturation. No relevant difference in maturation kinetics were observed between cells overexpressing CMTM6 and CMTM6-deficient cells. Pulse chase experiments were performed three times, once comparing CMTM6 overexpressing and CMTM6-deficient cells (a), and twice comparing wt and CMTM6-deficient cells. Other data are representative of at least two independent experiments. MFI, median fluorescence intensity; BM, bone marrow; DC, dendritic cell; KO, knockout; OE, overexpression; EndoH, endoglycosidase H; PNGaseF, peptide-N-glycosidase F..
Extended data Figure 6. Specificity and membrane localization of CMTM6, and co-localization with PD-L1 in human tumors.
(a), Membrane-fractionated proteome of 8505C and RKO cells. CMTM6 and PD-L1 were detected by LC-MS/MS predominantly from the plasma membrane fractions. Label-free quantification (LFQ) performed by intensity-based normalization of 4 fractions together across different cell lines is depicted. (b) A375 parental cells, CMTM6 KO or CMTM6 overexpressing cells were fixed and formalin embedded, and stained for CMTM6 with a monoclonal antibody (1D6) generated against a peptide from the C-terminal domain of CMTM6. Analysis shows mainly membranous stain, as indicated by the arrowheads. (c) Sequential slides from lymph node and subcutaneous metastases from 3 melanoma patients were stained for PD-L1 (left) or for CMTM6 (right), showing frequent localization of PD-L1 within CMTM6 positive areas. In total, samples from 9 melanoma patients and 5 PD-L1 positive lung cancer samples were analyzed. OE, overexpression; KO, knockout.
Extended data Figure 7. Interactions between CMTM6, PD-L1, and CMTM4, and effect of CMTM6 on PD-L1 stability.
(a) A375 parental cells, CMTM6-deficient cells, PD-L1-deficient cells, and cells ectopically expressing CMTM6 or PD-L1, were cultured in the absence or presence of 25ng/ml IFNγ for 48 hours before preparation of cell lysates. Immunoprecipitation was performed using a CMTM6-specific antibody immobilized on protein A coated beads. Immunoprecipitates and whole cell lysate were subjected to SDS-PAGE and immunoblotted for CMTM6 and PD-L1. Two exposures of the same western blot are shown. Arrows indicate PD-L1 bands. (b) parental 8505C cells and CMTM6-deficient 8505C cells were cultured in the absence or presence of 50ng/ml IFNγ for 72 hours before preparation of cell lysates. Immunoprecipitation was performed using CMTM6- or PD-L1-specific antibodies immobilized on protein A coated beads. Immunoprecipitates and whole cell lysates were subjected to SDS-PAGE and immunoblotted for CMTM6 and PD-L1. Two exposures of the same western blot are shown. Normal IgG served as control. Arrows indicate PD-L1 bands. (c, d) Parental and CMTM6 knockout RKO cells (c) and (d) 8505c cells were lysed and immunoprecipitation was performed using antibodies immobilized on protein G coated beads as indicated. Immunoprecipitates and whole cell lysates were subjected to SDS-PAGE, and Western blot analysis of CMTM4 and PD-L1 was carried out. Two exposures of the same Western blots are shown. Arrows indicate CMTM4. Data are representative of three independent experiments. KO, knockout; OE, overexpression.
Extended data Figure 8. Aspects of PD-L1 regulation by CMTM6 and STUB1.
(a) V5-tagged PD-L1 was introduced into parental, CMTM6-overexpressing and CMTM6-deficient A375 cells. Cell lysates were denatured and then subjected to immunoprecipitation with anti-V5 antibody immobilized on protein G-coated beads. Immunoprecipitates were then analyzed by immunoblotting with anti-V5 antibody as a control for the experiments shown in Fig. 4e. Results of the FACS-based genetic screens in CMTM6 expressing and CMTM6 deficient HAP1 cells as shown in Fig. 1a (b) and in Fig. 3a (c), with the position of STUB1 indicated. (d) Relative expression of PD-L1, PD-L2 and the indicated PD-L1 – PD-L2 chimeric proteins in CMTM6 KD A375 cells as compared to matched control. Chimeras were detected with an anti PD-L1 or an anti PD-L2 antibody. (e) Schematic overview of the chimeric proteins analyzed. (f,g) 293T human embryonic kidney cells were co-transfected with a vector encoding either PD-L1, PD-L2 or the indicated chimeric protein, together with a vector encoding CMTM6. Cell lysates were denatured and subjected to immunoprecipitation with anti-flag antibody immobilized on protein G-coated beads. Lysates and immunoprecipitates were then analyzed by immunoblotting with the indicated antibodies. Data are representative of three (a,d), one (f) or two (g) independent experiments. Error bars represent s.d. of triplicates. MFI, median fluorescence intensity; KO, knockout; OE, overexpression; TM, transmembrane; IC, intracellular; EC, extracellular.
Extended data Figure 9. Orientation mapping of CMTM6.
(a) Predicted domain topology of CMTM6 according to TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM/). (b,c) A375 cells were transduced with C- or N-terminal HA epitope tagged CMTM6. HA staining was performed in both live cells and fixed and permeabilized cells followed by flow cytometry analysis and quantified in (c). MFI, median fluorescence intensity.
Extended data Figure 10. Selectivity of CMTM6 and CMTM6 loss alleviates PD-L1-mediated T cell suppression.
(a) Comparative membrane-fractionated mass spectrometry of CMTM6 proficient or deficient RKO cells. 4 wild type and 4 CMTM6 KO RKO clones were analyzed by LC-MS/MS and differential protein abundance is shown in a volcano plot. (b) Table indicating proteins found up- or down-regulated upon CMTM6 removal in both 8505c and RKO. Flow cytometric (c) and Western blot (d) analysis of CMTM6 and PD-L1 expression in parental A375 or CMTM6 deficient A375 clones in which PD-L1 is ectopically expressed by lentiviral transduction. (e). Primary human T cells were transduced with the MART-1 specific 1D3 TCR31 and PD-1. Transduced T cells were co-cultured with unloaded or MART-1 peptide loaded PD-L1-overexpressing A375 cells (‘Parental + PD-L1 OE’), parental A375 cells (‘Parental’), or CMTM6-deficient A375 cells that overexpressed PD-L1 (‘CMTM6 KO+PD-L1 OE’). IL-2 production in T cells that expressed high, intermediate, or low levels of PD-1 (‘PD-1HI’, ‘PD-1INTER’ or ‘PD-1LOW’) were analyzed by flow cytometry. Untransduced A375 cells (‘Parental’) served as controls. Data are representative of three independent experiments and were analyzed by unpaired t-test (c). Error bars represent s.d. of triplicates. *P<0.05; **P<0.01; ***P<0.001. KO, knockout; OE, overexpression; TM, transmembrane; PM, plasma membrane.
Supplementary Material
Acknowledgements
We would like to thank F. Scheeren, E. Stickel, V. Blomen, M. Brockmann and the other members of the Schumacher and Brummelkamp lab for valuable discussions, and Joanna Grabowska for technical assistance. We would like to thank Kristel Kemper and Daniel Peeper for sharing melanoma PDX models. We would like to acknowledge the NKI- AVL flow facility, the NKI-AVL Core Facility Molecular Pathology & Biobanking (CFMPB) for supplying NKI-AVL Biobank material and /or lab support. This work was supported by The Queen Wilhelmina Cancer Research Award and European Research Council (ERC) Advanced Grant SENSIT (to T.N.M.S.), NWO Vici Grant (016.Vici.170.033), the Cancer Genomics Center (CGC.nl), and Ammodo KNAW Award 2015 for Biomedical Sciences (to T.R.B.), The Cancer Research Institute (CRI) Irvington Postdoctoral Fellowship (to C.S.), The Landsteiner Foundation for Blood Research, grant 1355 (to J.B.), Proteins@Work, a program of the Netherlands Proteomics Centre financed by NWO, the Netherlands Organisation for Scientific Research (to A.J.H.), and The Institute for Chemical Immunology, an NWO Gravitation project (to T.N.M.S. and A.J.H.).
Footnotes
Author Contributions
R.M., C.S. and L.T.J. conceived the project, designed and performed experiments, interpreted data, and co-wrote the paper. R.G.E. designed, performed and interpreted functional assays. W.W. designed, performed and interpreted mass spectrometry analyses, A.J.H. designed and interpreted mass spectrometry analyses, E.d.V. designed, performed and interpreted immunoprecipitation experiments, Y.X. designed, performed and interpreted human DC experiments, M.E.W.L. performed and interpreted melanoma PDX experiments, M.S. performed bio-informatic analyses, L.F.A.W. supervised bio-informatic analysis, E.A.R. and I.H. identified samples and performed IHC analyses, A.B. and H.M.H. supervised and scored IHC analyses, C.U.B. provided and identified samples for IHC analyses, J.B. designed and interpreted immunoprecipitation and human DC experiments, T.R.B. and T.N.M.S. designed experiments, interpreted data, and co-wrote the manuscript.
Competing financial interests
R.M., C.S., L.T.J., T.R.B. and T.N.M.S. are inventors on a patent application covering the use of CMTM6, CMTM4 and STUB1 as therapeutic and diagnostic targets. T.R.B. is co-founder and shareholder of Scenic Biotech. The other authors declare no competing interests.
Data availability
All sequencing datasets have been deposited in the NCBI Sequence Read Archive under accession number SRP108407. In addition, all processed screen results are accessible in an interactive database (https://phenosaurus.nki.nl/). Source Data for the main and Extended Data figures are provided in the online version of the paper.
References
- 1.Topalian SL, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. The New England journal of medicine. 2012;366:2443–2454. doi: 10.1056/NEJMoa1200690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brahmer JR, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. The New England journal of medicine. 2012;366:2455–2465. doi: 10.1056/NEJMoa1200694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ansell SM, et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin's lymphoma. The New England journal of medicine. 2015;372:311–319. doi: 10.1056/NEJMoa1411087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Powles T, et al. MPDL3280A (anti-PD-L1) treatment leads to clinical activity in metastatic bladder cancer. Nature. 2014;515:558–562. doi: 10.1038/nature13904. [DOI] [PubMed] [Google Scholar]
- 5.Garon EB, et al. Pembrolizumab for the treatment of non-small-cell lung cancer. The New England journal of medicine. 2015;372:2018–2028. doi: 10.1056/NEJMoa1501824. [DOI] [PubMed] [Google Scholar]
- 6.Le DT, et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. The New England journal of medicine. 2015;372:2509–2520. doi: 10.1056/NEJMoa1500596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Motzer RJ, et al. Nivolumab versus Everolimus in Advanced Renal-Cell Carcinoma. The New England journal of medicine. 2015;373:1803–1813. doi: 10.1056/NEJMoa1510665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Nghiem PT, et al. PD-1 Blockade with Pembrolizumab in Advanced Merkel-Cell Carcinoma. The New England journal of medicine. 2016;374:2542–2552. doi: 10.1056/NEJMoa1603702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ferris RL, et al. Nivolumab for Recurrent Squamous-Cell Carcinoma of the Head and Neck. The New England journal of medicine. 2016 doi: 10.1056/NEJMoa1602252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lee SJ, et al. Interferon regulatory factor-1 is prerequisite to the constitutive expression and IFN-gamma-induced upregulation of B7-H1 (CD274) FEBS letters. 2006;580:755–762. doi: 10.1016/j.febslet.2005.12.093. [DOI] [PubMed] [Google Scholar]
- 11.Kataoka K, et al. Aberrant PD-L1 expression through 3'-UTR disruption in multiple cancers. Nature. 2016;534:402–406. doi: 10.1038/nature18294. [DOI] [PubMed] [Google Scholar]
- 12.Casey SC, et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science (New York, N.Y.) 2016;352:227–231. doi: 10.1126/science.aac9935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dorand RD, et al. Cdk5 disruption attenuates tumor PD-L1 expression and promotes antitumor immunity. Science (New York, N.Y.) 2016;353:399–403. doi: 10.1126/science.aae0477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li C-W, et al. Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nature communications. 2016;7:12632. doi: 10.1038/ncomms12632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lim SO, et al. Deubiquitination and Stabilization of PD-L1 by CSN5. Cancer cell. 2016 doi: 10.1016/j.ccell.2016.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Herbst RS, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515:563–567. doi: 10.1038/nature14011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ribas A, Hu-Lieskovan S. What does PD-L1 positive or negative mean? The Journal of experimental medicine. 2016 doi: 10.1084/jem.20161462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Carette JE, et al. Haploid genetic screens in human cells identify host factors used by pathogens. Science (New York, N.Y.) 2009;326:1231–1235. doi: 10.1126/science.1178955. [DOI] [PubMed] [Google Scholar]
- 19.Carette JE, et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature. 2011;477:340–343. doi: 10.1038/nature10348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Brockmann M, et al. Genetic wiring maps of single-cell protein states reveal an off-switch for GPCR signalling. Nature. 2017 doi: 10.1038/nature22376. [DOI] [PubMed] [Google Scholar]
- 21.Platanias LC. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nature reviews Immunology. 2005;5:375–386. doi: 10.1038/nri1604. [DOI] [PubMed] [Google Scholar]
- 22.Han W, et al. Identification of eight genes encoding chemokine-like factor superfamily members 1-8 (CKLFSF1-8) by in silico cloning and experimental validation. Genomics. 2003;81:609–617. doi: 10.1016/s0888-7543(03)00095-8. [DOI] [PubMed] [Google Scholar]
- 23.Xiao Y, et al. Identification of the Common Origins of Osteoclasts, Macrophages, and Dendritic Cells in Human Hematopoiesis. Stem cell reports. 2015;4:984–994. doi: 10.1016/j.stemcr.2015.04.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen Z, et al. The ubiquitin ligase Stub1 negatively modulates regulatory T cell suppressive activity by promoting degradation of the transcription factor Foxp3. Immunity. 2013;39:272–285. doi: 10.1016/j.immuni.2013.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wei F, et al. Strength of PD-1 signaling differentially affects T-cell effector functions. Proceedings of the National Academy of Sciences of the United States of America. 2013;110:E2480–2489. doi: 10.1073/pnas.1305394110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Miyazaki A, Yogosawa S, Murakami A, Kitamura D. Identification of CMTM7 as a transmembrane linker of BLNK and the B-cell receptor. PloS one. 2012;7:e31829. doi: 10.1371/journal.pone.0031829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B (Methodological) 1995;57:289–300. [Google Scholar]
- 28.Kemper K, et al. Intra- and inter-tumor heterogeneity in a vemurafenib-resistant melanoma patient and derived xenografts. EMBO molecular medicine. 2015;7:1104–1118. doi: 10.15252/emmm.201404914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lackner DH, et al. A generic strategy for CRISPR-Cas9-mediated gene tagging. Nature communications. 2015;6:10237. doi: 10.1038/ncomms10237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Doench JG, et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nature biotechnology. 2016;34:184–191. doi: 10.1038/nbt.3437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Jorritsma A, et al. Selecting highly affine and well-expressed TCRs for gene therapy of melanoma. Blood. 2007;110:3564–3572. doi: 10.1182/blood-2007-02-075010. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.