Abstract
Epigenetic marks are likely to explain variability of response to antigen in granulomatous lung disease. The objective of this study was to identify DNA methylation and gene expression changes associated with chronic beryllium disease (CBD) and sarcoidosis in lung cells obtained by BAL. BAL cells from CBD (n = 8), beryllium-sensitized (n = 8), sarcoidosis (n = 8), and additional progressive sarcoidosis (n = 9) and remitting (n = 15) sarcoidosis were profiled on the Illumina 450k methylation and Affymetrix/Agilent gene expression microarrays. Statistical analyses were performed to identify DNA methylation and gene expression changes associated with CBD, sarcoidosis, and disease progression in sarcoidosis. DNA methylation array findings were validated by pyrosequencing. We identified 52,860 significant (P < 0.005 and q < 0.05) CpGs associated with CBD; 2,726 CpGs near 1,944 unique genes have greater than 25% methylation change. A total of 69% of differentially methylated genes are significantly (q < 0.05) differentially expressed in CBD, with many canonical inverse relationships of methylation and expression in genes critical to T-helper cell type 1 differentiation, chemokines and their receptors, and other genes involved in immunity. Testing of these CBD-associated CpGs in sarcoidosis reveals that methylation changes only approach significance, but are methylated in the same direction, suggesting similarities between the two diseases with more heterogeneity in sarcoidosis that limits power with the current sample size. Analysis of progressive versus remitting sarcoidosis identified 15,215 CpGs (P < 0.005 and q < 0.05), but only 801 of them have greater than 5% methylation change, demonstrating that DNA methylation marks of disease progression changes are more subtle. Our study highlights the significance of epigenetic marks in lung immune response in granulomatous lung disease.
Keywords: epigenetics, berylliosis, sarcoidosis, genomics, immunology
Clinical Relevance
Granulomatous diseases, including chronic beryllium disease (CBD) and sarcoidosis, are characterized by a highly variable response to environmental antigen, beryllium in CBD, and unknown in sarcoidosis. Genetic risk factors do not fully explain disease susceptibility or clinical variability, strongly implicating the environment and gene × environment interactions, including epigenetic mechanisms. Our study identified DNA methylation changes associated with CBD and sarcoidosis that are also related to changes in gene expression in lung cells obtained by BAL. Importantly, this is the first study of its kind in CBD and sarcoidosis, and it highlights the significance of epigenetic marks in lung immune response in these environmental lung diseases. DNA methylation changes in CBD are prominent, whereas those in sarcoidosis are more subtle and variable, with the variability likely due to disease heterogeneity and/or inciting antigen. Our study adds to the growing body of evidence supporting the notion that CBD and sarcoidosis are similar at the molecular level.
Granulomatous diseases, including those of known etiology, such as chronic beryllium disease (CBD), and unknown etiology, such as sarcoidosis, affect people of both genders and all ages and races (1, 2). It has been estimated that over 1 million individuals have been exposed to beryllium (3). Of those exposed, 2–20% will develop the precursor to disease, beryllium sensitization (BeS) and are an at-risk population, as 50–100% of them will develop CBD (4). Similarly, sarcoidosis affects at least 45–300/100,000 individuals in the United States, and the mortality rate is rising (5). Individuals with sarcoidosis are usually diagnosed between the ages of 20 and 50 and experience a significantly decreased productivity and quality of life (6). These diseases share many clinical, radiographic, pathologic, and immunologic similarities (7).
In these immune-mediated lung diseases, exposure to antigen (beryllium for CBD and unknown for sarcoidosis) results in an antigen-specific T-helper cell type (Th) 1 immune response and subsequent granulomatous inflammation, in some cases resulting in fibrosis and end-stage disease. Genetic factors play a role in the host response to antigen. The immune response to beryllium is strongly associated with a functional human leukocyte antigen (HLA) class II variant, a glutamic acid at amino acid position 69 in HLA-DPB1 (E69) (8, 9). Studies by our group and others have shown that the majority of beryllium (Be)-responsive CD4+ T cells respond to a beryllium antigen (Be-Ag) presented by these E69 variants, that the E69 variants impact cell proliferation and cytokine production, and are critical in CBD pathogenesis (9, 10). Genetic risk factors for sarcoidosis are less well known, although HLA and other genomic loci are associated with disease risk (11, 12) and disease severity (13). However, genetic risk factors do not fully explain CBD and sarcoidosis susceptibility or clinical variability, strongly implicating the environment and gene × environment interactions, including epigenetic mechanisms.
Epigenetic alterations in combination with genetic susceptibility may provide a crucial link between exposure and disease risk. Several lines of evidence support the notion that epigenetics will play a role in CBD and sarcoidosis. Differentiation of naive CD4+ T cells into Th1, Th2, Th17, and T regulatory lineages, critical in CBD and sarcoidosis, is regulated by DNA methylation and histone modifications (14–22). Moreover, other immune-mediated diseases implicate epigenetic regulation of immune function and immunopathogenesis. Furthermore, genes that are important in epigenetic regulation, including HDAC and other histone- and chromatin-modifying genes, are associated with sarcoidosis (23). Finally, sarcoidosis has a variable disease course, with some individuals progressing to fibrotic lung disease and others demonstrating disease remission. Another fibrotic lung disease, idiopathic pulmonary fibrosis, is associated with extensive DNA methylation and gene expression differences in lung tissue (24), suggesting that this is likely going to be the case for progressive forms of sarcoidosis. However, no studies to date have examined DNA methylation or histone modifications in either CBD or sarcoidosis. In this study, we profiled genome-wide DNA methylation changes in BAL from subjects with CBD, sarcoidosis, and BeS. We identified a large number of DNA methylation changes associated with CBD that also appear to affect gene expression. DNA methylation levels in sarcoidosis were present, albeit less pronounced and more variable than in CBD.
Methods
Study Population
Our primary study population consisted of cases of CBD (n = 8), beryllium sensitization with no disease (BeS, n = 8), and sarcoidosis (n = 8). Study subjects were recruited at National Jewish Health using standard case criteria under an approved Institutional Review Board protocol to collect leftover cells from either a clinical or research bronchoscopy (HS-2458). The second population consisted of an additional 24 cases of sarcoidosis (n = 24), including 9 cases with progressive and 15 with remitting pulmonary disease. Inclusion and exclusion criteria are specified in the data supplement.
Genome-Wide DNA Methylation and mRNA Expression Assessment
To measure methylation at approximately 485,000 single CpG sites across the genome, we used Illumina’s Infinium Human Methylation 450k BeadChip on bisulfite-treated samples, as previously reported (25). Genome-wide expression was determined for the first batch of RNA samples (eight subjects with CBD, eight subjects with sarcoidosis, and eight subjects with BeS) using the Human U133 Plus 2.0 Array (Affymetrix). The second batch of additional 24 sarcoidosis samples was profiled on the Agilent G3 SurePrint 8 × 60 human gene expression one-color microarray platform. For additional details, see the data supplement.
Pyrosequencing
Bisulfite-converted DNA was PCR amplified using primers designed in the Pyromark Assay Design Software (Qiagen), using PCR and sequencing primers listed in Table E1 in the data supplement.
Data Quality Controls
We performed several quality control steps that are standard in the analysis of Illumina 450k array data, as previously published (26). We removed probes with high detection P values, probes with fewer than three beads in at least 10% of the samples, and probes with known SNPs within the CpG motif (27). We removed one sample more than 3 SDs away from the mean of the top three principal components in the first batch of samples (Figure E1A) and two in the second batch of samples (Figure E1B). There were no outliers on principal component analysis (PCA) in expression datasets, and we therefore did not remove any samples from further analysis. DNA methylation and gene expression datasets have been deposited to the Gene Expression Omnibus (accession number in progress GSE110780).
Statistical Analysis of DNA Methylation and Gene Expression Data
All statistical analyses of microarray data were performed in the Partek Genomics Suite. Data from the methylation array were normalized using the Subset-quantile within Array Normalization method (SWAN) (28), and the normalized M values were used in downstream statistical analyses due to the more normal distribution (Figure E2), whereas β values, on the scale of 0–100%, are used for tables and figures, because these values are easier to interpret biologically. Differences between groups are reported as percent methylation changes using β values. Gene expression arrays were normalized using quantile normalization and expression levels log2 transformed before the analysis.
PCA was also used to examine technical variables (scan date, array, and position on the array) that may confound statistical analysis and need to be adjusted for as “batch effects” (Figure E1). Mixed-effects linear models were run to identify differentially methylated CpGs or differentially expressed genes among disease groups, while adjusting for the effect of position on the array (methylation) or scan date (expression) as a random effect. Statistical significance was assessed by calculating false discovery rate (FDR)–adjusted P values from linear model P values using the method of Benjamini and Hochberg (29).
Pathway and Network Analysis
We used the Network Analyst tool to generate the protein–protein interactomes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment within the interactomes (30). We used Ingenuity Pathway Analysis (IPA) (31) to identify enriched pathways and upstream regulators.
Results
Genome-Wide Methylation Profiling Demonstrates Enrichment of Immune Genes and Pathways in CBD
We first analyzed Illumina 450k DNA methylation data in our primary cohort (eight CBD, eight BeS, and eight sarcoidosis) to identify DNA methylation changes associated with either CBD or sarcoidosis compared to BeS. Because there were no significant differences in demographics and cigarette smoking among the three groups (Table 1), we did not adjust for these covariates in our analysis. Based on PCA (Figure E1A), we adjusted for the array position (as a random effect) in the mixed-effects linear model. This analysis revealed extensive, genome-wide significant DNA methylation changes in CBD (52,860 CpGs with nominal P < 0.005 and FDR-adjusted q < 0.05; Figure 1A; Tables 2 and 3), but not in sarcoidosis. Of the 52,860 CBD-associated CpGs, 36,816 are near 11,553 unique genes, suggesting substantial genomic alterations in DNA methylation in CBD compared with BeS. Focusing on the largest methylation changes among these, 3,654 CpGs have an absolute change greater than 25%; 2,726 of them are near 1,944 unique genes (Table E2). Although all significant CpGs are enriched for hypomethylation, those with greater than 25% absolute change have a similar number of hypo- and hypermethylated CpGs (Table 2). We validated select genes by pyrosequencing for technical validation of these findings (Figure E3). Given that five out of eight subjects with CBD were treated with immunosuppressants, we examined the effect of treatment on 2,726 CpGs by adjusting for treatment in the statistical model. Treatment with immunosuppressants had no notable impact on CBD-associated DNA methylation; all CpGs remained significant (q < 0.05) after adjustment for treatment. This is not surprising, given previous work showing no effect of steroid treatment on cytokine production and proliferation in CBD (32, 33). We also explored the impact of proportion of lymphocytes in the BAL on these findings by running an additional analysis of 2,726 CpGs, adjusting for lymphocyte percentage in the statistical model. After adjustment for lymphocyte counts, 786 CpGs (29%) remained significant at q less than 0.05, and an additional 965 CpGs (35%) at nominal P less than 0.05 (Table E3). We also considered adjustment for percent alveolar macrophages, but this is the same analysis statistically, because lymphocytes and macrophages add up to greater than 97% in each subject (few other cells are detected in the BAL).
Table 1.
Basic Demographic and Clinical Characteristics of 24 Subjects Included in the Primary Cohort
| |
BeS |
CBD |
Sarcoidosis |
|
|---|---|---|---|---|
| (n = 8) | (n = 8) | (n = 8) | P Value | |
| Age, yr* | 56.8 ± 8.3 | 58.6 ± 6.8 | 56.6 ± 11.5 | 0.89† |
| Male sex, n (%) | 7 (87.5) | 7 (87.5) | 6 (75) | 0.74‡ |
| White race, n (%) | 8 (100) | 8 (100) | 7 (87.5) | 0.35‡ |
| Nonhispanic ethnicity, n (%) | 6 (75) | 7 (87.5) | 8 (100) | 0.32‡ |
| Smoking, n | 0.84‡ | |||
| Never | 5 | 5 | 4 | |
| Former | 3 | 3 | 4 | |
| FEV1, % predicted* | 99.4 ± 14.0 | 89.8 ± 16.7 | 77.6 ± 20.8 | 0.064† |
| FVC, % predicted* | 95.5 ± 11.4 | 86.8 ± 19.4 | 81.4 ± 16.2 | 0.23† |
| TLC, % predicted* | 112.1 ± 16.5 | 98.9 ± 19.2 | 92.3 ± 17.0 | 0.11† |
| DlCO, % predicted* | 96.6 ± 10.0 | 60.7 ± 23.2 | 69.4 ± 12.4 | 0.0014† |
| Immunosuppressive therapy, n (%) | 0 (0) | 4 (50) | 1(12.5) | 0.038‡ |
| Methothrexate, n (%) | 0 (0) | 0 (0) | 1(12.5) | 0.35‡ |
| Lymphocyte, %* | 6.1 ± 4.3 | 46.2 ± 32.1 | 19.8 ± 17.9 | 0.004† |
| Macrophage, %* | 92.9 ± 4.1 | 52.2 ± 31.2 | 78.1 ± 18.6 | 0.003† |
Definition of abbreviations: BeS = beryllium sensitized; CBD = chronic beryllium disease; FEV1 = forced expiratory volume in 1 second; FVC = forced vital capacity; TLC = total lung capacity.
Presented as mean ± SD.
Determined by one-way ANOVA or Student’s t test.
Determined by chi-square test.
Figure 1.
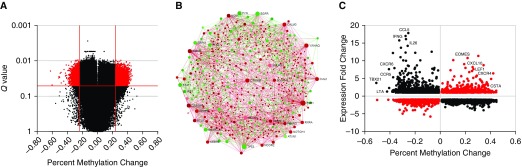
Differentially methylated single-CpG probes in BAL cells associated with chronic beryllium disease (CBD). (A) A volcano plot of false discovery rate (FDR) adjusted q values for disease status (CBD/beryllium-sensitized [BeS]) from the ANOVA model plotted as a function of methylation change (CBD − BeS). Probes with statistically significant change q < 0.05 and methylation change >25% are highlighted in red. (B) Protein–protein interactome analysis of 1,944 genes with significant CBD-associated DNA methylation changes (q < 0.05, P < 0.005, and methylation change >25%). The zero-order interactome was created using NetworkAnalyst (57) and the STRING PPI dataset. The nodes are colored based on their methylation (green are hypomethylated, whereas red are hypermethylated). The sizes of nodes are proportional to their betweenness centrality values. (C) Methylation-expression plot for genes with CBD-associated DNA methylation changes with q < 0.05, P < 0.005; x-axis methylation difference is represented by the mean % methylation difference in CBD compared with BeS subjects; y-axis expression difference is represented by the mean fold change in CBD compared with subjects with BeS. The black solid circles represent hypomethylated genes that were associated with increased gene expression as well as some hypermethylated genes associated with decreased gene expression. The red solid circles represent methylation changes that were not associated with canonical gene expression differences.
Table 2.
Number of CpGs on the Illumina Array Meeting Criteria for Significant Differential Methylation in Chronic Beryllium Disease Compared to Beryllium Sensitization
| |
|
|
|
Benjamini-Hochberg q < 0.05 |
|---|---|---|---|---|
| |
|
Benjamini-Hochberg q < 0.05 |
Benjamini-Hochberg q < 0.05 |
Methylation Change >25% |
| Benjamini-Hochberg q < 0.05 | (Unadjusted P < 0.005) | Methylation Change >25% | (Unadjusted P < 0.005) | |
| All probes | 61,238 | 52,831 | 4,411 | 3,654 |
| Hypomethylated | 45,058 | 40,066 | 1,885 | 1,700 |
| Hypermethylated | 16,180 | 12,765 | 2,526 | 1,954 |
| Probes near genes | 42,747 | 36,792 | 3,271 | 2,726 |
| Hypomethylated | 30,354 | 26,975 | 1,382 | 1,249 |
| Hypermethylated | 12,393 | 9,817 | 1,889 | 1,477 |
| Unique genes | 13,518 | 12,565 | 2,320 | 1,944 |
Table 3.
KEGG Pathways Significantly (q < 0.01) Enriched in Genes That Are Hypomethylated by >25% in Chronic Beryllium Disease versus Beryllium Sensitization at q < 0.05 and Unadjusted P < 0.005
| Pathway | Total | Expected | Hits | P Value | q Value |
|---|---|---|---|---|---|
| Measles | 102 | 3.86 | 16 | 9.82 × 10−07 | 0.000124 |
| Pathways in cancer | 310 | 11.7 | 30 | 1.14 × 10−06 | 0.000124 |
| HTLV-I infection | 199 | 7.53 | 22 | 4.22 × 10−06 | 0.000305 |
| Regulation of actin cytoskeleton | 182 | 6.88 | 20 | 1.31 × 10−05 | 0.00071 |
| Pancreatic cancer | 69 | 2.61 | 11 | 4.47 × 10−05 | 0.00194 |
| FcγR–mediated phagocytosis | 97 | 3.67 | 13 | 6.02 × 10−05 | 0.00199 |
| Cytokine–cytokine receptor interaction | 253 | 9.57 | 23 | 6.40 × 10−05 | 0.00199 |
| Chagas disease | 89 | 3.37 | 12 | 0.00011 | 0.00298 |
| Glioma | 65 | 2.46 | 10 | 0.000136 | 0.00329 |
| Focal adhesion | 200 | 7.56 | 19 | 0.000163 | 0.00354 |
| Chemokine signaling pathway | 189 | 7.15 | 18 | 0.000237 | 0.00467 |
| T cell receptor signaling pathway | 98 | 3.71 | 12 | 0.000279 | 0.00505 |
| Jak–STAT signaling pathway | 99 | 3.74 | 12 | 0.000307 | 0.00513 |
| Natural killer cell–mediated cytotoxicity | 138 | 5.22 | 14 | 0.000634 | 0.00983 |
Definition of abbreviations: HTLV-I = human T-cell leukemia virus type 1; Jak-STAT = Janus kinase–signal transducer of activation; KEGG = Kyoto Encyclopedia of Genes and Genomes.
Network analysis of 1,944 genes with CBD-associated methylation changes greater than 25%, using the STRING interactome data, shows hypomethylation of immune and proliferation genes, such as the transcription factor STAT1, and kinases JAK1, PIK3CD, PTK2, CSK, and SYK (Figure 1B). Similarly, KEGG enrichment analysis of the pathways in the first-order network (Figure E4) shows enrichment of immune pathways, including cytokine–cytokine receptor interactions, T cell receptor signaling, chemokine signaling, and Jak-STAT signaling (Table 3). Pathways related to cancer are also enriched, likely due to large numbers of cell proliferation genes in cancer pathways, as well as FcγR–mediated phagocytosis, potentially related to antigen-processing and presentation genes. Enrichment of pathways involved in immune response, especially T cells, and proliferation among hypomethylated genes were also observed by IPA (Figure E5). No significant KEGG pathway enrichment was observed in hypermethylated genes in the network, and IPA showed some enrichment in pathways related to vitamin D and peroxisome proliferator–activated receptors (PPARs) signaling among hypermethylated genes, although significance of enrichment was much weaker than in hypomethylated genes (Figure E5). Taken together, these data demonstrate that genes and pathways known to be important in CBD appear to be epigenetically regulated.
Gene Expression Changes Are Related to DNA Methylation Changes in CBD
To examine the effect of these DNA methylation changes on gene expression, we analyzed Affymetrix U133 Plus gene expression data in the same samples for the genes with CBD-associated methylation changes greater than 25%. A total of 1,334/1,944 (69%) differentially methylated genes are also significantly (q < 0.05) differentially expressed in CBD compared with BeS (Table E4). Plotting gene expression as a function of DNA methylation across all genes reveals canonical inverse relationships with large effect sizes in genes critical to Th1 cell differentiation (TBX21/TBET, IFNG), chemokines and their receptors (CCR5, CXCR6, CCL5), and other genes involved in immunity (LTA, IL26) (Figure 1C). However, we also observed noncanonical direct correlations of DNA methylation and gene expression in immune genes (CXCR4, CXCL10, CSTA, EOMES). These data suggest that, in general, CBD-associated epigenetic marks affect gene expression in BAL cells.
DNA Methylation Changes in Sarcoidosis Are Present, but Not Significant and Smaller in Magnitude than in CBD due to Disease Heterogeneity
We further explored the data to begin to understand why we were not able to identify genome-wide significant DNA methylation changes in sarcoidosis when compared with BeS. Testing of 2,726 CpGs (1,944 unique genes) that were associated with CBD (P < 0.005, q < 0.05, and methylation change >25%) reveals that the majority has methylation changes that are not statistically significant, but approach significance (q < 0.15) and are methylated in the same direction as CBD, but with a smaller magnitude of change (Table E2). Closer examination of the PCA plot (Figure E1A) and individual data points for some of the genes identified in CBD (Figure 2) suggests that there is more variability in DNA methylation and gene expression in sarcoidosis, and that this may be due to disease heterogeneity, such as disease progression versus remission. These data demonstrate that CBD and sarcoidosis have many similarities in DNA methylation, but that sarcoidosis is more heterogeneous.
Figure 2.
DNA methylation and gene expression are more variable in sarcoidosis (Sarc) than CBD. Representative plots of (A) DNA methylation and (B) relationship of DNA methylation with gene expression of genes from Figure 1; BeS, red; CBD, blue; sarcoidosis, green.
Genome-Wide DNA Methylation Changes Are Associated with Progressive versus Remitting Sarcoidosis, and Are Related to Gene Expression Changes
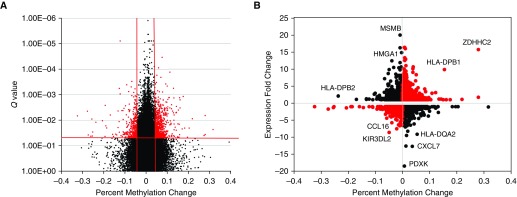
Given the heterogeneity that we observed in sarcoidosis in the original cohort, we performed genome-wide profiling on an additional 9 cases of progressive and 15 of remitting sarcoidosis (Table 4) with the goal of identifying DNA methylation and gene expression changes associated with disease progression. We first examined the 2,726 CpGs (1,944 unique genes) that are significant in CBD (P < 0.005, q < 0.05, and methylation change >25%), and determined that these are not significantly differentially methylated in progressive compared with remitting sarcoidosis (data not shown). We then performed genome-wide analysis to seek novel candidate genes and pathways for disease progression in sarcoidosis. This analysis identified 15,215 CpGs with nominal P value less than 0.005 and FDR-adjusted q less than 0.05. Only 801/15,215 have greater than 5% methylation change in progressive compared with remitting disease, with 516 of them near 420 unique genes (Figure 3A and Table E5); 506/516 remain significant after adjustment for immunosuppressant treatment (Table E5). Differentially methylated and expressed genes in progressive compared with remitting sarcoidosis include genes with function in immunity (HLA, chemokines, MSMB, KIR3DL2), chromatin remodeling (HMGA1), transcription (ZDHHC2), and vitamin B6 metabolism (PDXK), (Figure 3B). These data demonstrate that DNA disease progression methylation changes are much more subtle and are associated with expression of genes in previously identified and novel molecular pathways.
Table 4.
Basic Demographic and Clinical Characteristics of 24 Additional Subjects in the Second Cohort
| |
Remitting Sarcoidosis |
Progressive Sarcoidosis |
|
|---|---|---|---|
| (n = 15) | (n = 9) | P Value | |
| Age, yr* | 54.9 ± 7.7 | 50.3 ± 11.0 | 0.25† |
| Male sex, n (%) | 11 (73.3) | 6 (66.7) | 0.73‡ |
| White race, n (%) | 13 (86.7) | 7 (77.8) | 0.57‡ |
| Nonhispanic ethnicity, n (%) | 15 (100) | 8 (88.9) | 0.19‡ |
| Smoking, n | 0.65‡ | ||
| Never | 9 | 4 | |
| Former | 6 | 4 | |
| FEV1, % predicted* | 88.6 ± 19.8 | 83.3 ± 16.3 | 0.51† |
| FVC, % predicted* | 90.0 ± 16.8 | 84.6 ± 16.8 | 0.44† |
| TLC, % predicted* | 98.7 ± 14.1 | 94.4 ± 23.0 | 0.67† |
| DlCO, % predicted* | 89.9 ± 16.1 | 82.0 ± 21.1 | 0.32† |
| Immunosuppressive therapy, n (%) | 0 (0) | 5 (55) | 0.0012‡ |
| Methothrexate, n (%) | 3 (20) | 3 (33.3) | 0.47‡ |
| Scadding stage, n | 0.16‡ | ||
| 0/1 | 2 | 1 | |
| 2 | 4 | 6 | |
| 3 | 5 | 0 | |
| 4 | 2 | 1 |
Definition of abbreviations: FEV1 = forced expiratory volume in 1 second; FVC = forced vital capacity; TLC = total lung capacity.
Presented as mean ± SD.
Determined by one-way ANOVA or Student’s t test.
Determined by chi-square test.
Figure 3.
Differentially methylated single-CpG probes in BAL cells associated with disease progression in sarcoidosis. (A) A volcano plot of FDR-adjusted q values for disease status (progressive/remitting sarcoidosis) from the ANOVA model plotted as a function of methylation change (progressive − remitting sarcoidosis). Probes with statistically significant change q < 0.05 and methylation change >5% are highlighted in red. (B) Methylation-expression plot for genes with progressive sarcoidosis-associated DNA methylation changes with q < 0.05; x-axis methylation difference is represented by the mean % methylation difference in progressive compared with remitting sarcoidosis subjects; y-axis expression difference is represented by the mean fold change in progressive compared with remitting sarcoidosis subjects. The black solid circles represent hypomethylated genes that were associated with increased gene expression as well as some hypermethylated genes associated with decreased gene expression. The red solid circles represent methylation changes that were not associated with canonical gene expression differences.
Discussion
Our study identified DNA methylation changes associated with granulomatous lung diseases that are also related to changes in gene expression in lung cells obtained by BAL. Importantly, this is the first study of its kind in CBD and sarcoidosis, and highlights the significance of epigenetic marks in lung immune response in these environmental lung diseases. DNA methylation changes in CBD are prominent, whereas those in sarcoidosis are not statistically significant with the current sample size, and are more subtle and variable. This variability is likely due, at least in part, to disease heterogeneity, although we cannot exclude the possibility of a smaller magnitude of change in sarcoidosis due to lower percent lymphocytes in the BAL. Our study adds to the growing body of evidence supporting that notion that CBD and sarcoidosis are similar at the molecular level (7). Some of the variability in sarcoidosis DNA methylation profiles can be attributed to heterogeneity in disease progression; however, it is also likely that multiple antigens are involved in sarcoidosis, and that variability in DNA methylation profiles is also driven by exposure to different antigens and/or antigen persistence, which appears to be the case in CBD.
Our analysis identified specific genes, networks, and pathways that are differentially methylated and expressed in CBD, and, to a lesser extent, in sarcoidosis. Hypomethylation and overexpression of large numbers of genes and pathways involved in immune response, and cell proliferation, are especially prominent in CBD, consistent with recruitment and proliferation of Th1 and Th17 cells as the hallmark of granulomatous lung disease. As an example, we found chemokine signaling pathways associated with CBD, and increased expression and hypomethylation of chemokine genes, such as CCR5, CXCL6, and CCL5 in CBD. These findings are likely functional, as we previously found that genetic variants in CCR5 were associated with BAL lymphocytosis and even more severe disease (34), although this will require future study. The role of Th17 cells in these diseases are implicated by our findings and other studies in sarcoidosis (35), although is not known in CBD. Numerous studies have demonstrated a critical role for immune pathways, including cytokine–cytokine receptor interactions, T cell receptor signaling, and chemokine signaling, in both CBD and sarcoidosis (25, 36, 37). More recently, Jak-STAT signaling has been identified as differentially expressed in both disease (7), and appears to be, at least in part, regulated by miRNAs (38) in sarcoidosis progression.
On the other hand, hypermethylated and downregulated genes are enriched, albeit weakly, in pathways related to vitamin D and PPAR signaling. The role of vitamin D in lung disease is somewhat controversial, especially in immune-mediated diseases, such as asthma (39). Vitamin D has been linked to sarcoidosis as an immunomodulating hormone that inhibits both antigen presentation by cells of the innate immune system and the cytokine release and proliferation of Th1 cells (40). PPARs are a family of ligand-activated nuclear hormone receptors (i.e., PPARα, PPARγ, and PPARδ). There is sufficient evidence that the activation of these receptors induces antiinflammatory and immune-modulatory effects in the lung as well as in other tissues. PPARγ deficiency in the lungs has been observed in the inflammatory diseases, such as asthma, fibrosis, and sarcoidosis. Alveolar macrophages from sarcoidosis are deficient in PPARγ signaling (41). Our data suggest that DNA hypermethylation may potentially contribute to downregulation of this pathway and, in turn, increase the inflammation. In aggregate, our data suggest that genes and pathways known to be important in granulomatous lung disease appear to be epigenetically regulated. Importantly, genes and pathways with no established link to CBD and sarcoidosis are also differentially methylated and expressed in our dataset. In addition to inverse relationships of DNA methylation and gene expression, our analysis also identified some direct correlations; these findings are in line with more recent views of the complexities of the epigenome, and will require further study to delineate (42–44).
The major strengths of our study are: 1) this is the first study of its kind to integrate genome-wide assessment of DNA methylation and transcriptional changes in noninfectious granulomatous disease; and 2) this assessment was undertaken on immune cells obtained from the target organ, the lung. This is of special importance, given that many of the previous studies of DNA methylation in lung disease were conducted in peripheral blood (45, 46). Additional strengths include the clinical data available with follow-up for over 2 years to ensure appropriate phenotyping, that the BeS did not develop CBD, and that the subjects with sarcoidosis were appropriately categorized as progressive versus remitting.
There are also several limitations to our study. First, our study was performed on a mixed population of immune cells. However, we were able to show that many disease-associated CpGs remain significant after adjustment for lymphocyte proportion in the BAL. It is also important to note that both DNA methylation changes that reflect reprogramming of specific cells (inherit methylation differences in specific cell populations) and those that are a result of changes in cell proportions or appearance of new cell populations in disease state are likely to be important in the disease process (47). Future studies in granulomatous lung disease will require isolation of specific T cell populations (Th1, Th17, T regulatory, etc.) or use of single-cell technologies. Second, we did not collect longitudinal samples, and were unable to examine longitudinal variation in DNA methylation due to changes in exposure or disease activity. The relationship between the environment and epigenome, especially in relation to immune-mediated diseases of the lung, remains a particularly important area of future research. Although we noted no significant difference in methylation in treated versus untreated CBD or sarcoidosis in this cross-sectional study, this does not exclude the potential that treatment might have an impact on the methylome within individuals, best addressed with longitudinal study. Third, our small sample size precluded us from assessing methylation marks in relation to disease heterogeneity and disease endotypes; future work in larger cohorts will be required to address this question, and to provide more definitive data on similarities of CBD and sarcoidosis methylomes. The fourth limitation is limited coverage of the epigenome on the Illumina 450k array; more recently implemented technologies, such as Illumina EPIC array and targeted BS-seq, provide better coverage for enhancers and other regulatory features in the genome (48). In addition, we limited focus on DNA methylation when we knew that histone modifications and transcription factors act in concert to regulate chromatin accessibility and gene expression, as our data support. Future studies will need to incorporate multiple technologies to more thoroughly assess chromatin state at the identified loci. Finally, our association analysis is unable to distinguish between DNA methylation marks being the cause or consequence of disease; to overcome this problem, future studies in large cohorts will need to implement mediation and two-step Mendelian randomization (49) analyses to examine causality. If methylation marks are found to be causal, future work will also need to address the effect of genetics and exposures on the methylation marks to determine the source of these epigenetic changes.
Despite these limitations, our study identified substantial and consistent alterations in the DNA methylome of lung immune cells that are associated with CBD and sarcoidosis, and with transcriptional changes in the same cells. These epigenetic signatures need to be further evaluated in larger cohorts for their potential as biomarkers, targets that may be altered with current therapies, and/or potential therapeutic targets. Previous work has shown that peripheral blood transcriptome reflects immune changes observed in the lung in CBD (7) and sarcoidosis (50); this work needs to be extended to DNA methylation as a potential biomarker of disease activity in the lung. DNA methylation marks are also a promising therapeutic target. DNA methyltransferase inhibitors have been approved for the treatment of myelodysplastic syndrome (51, 52), and are in clinical trials for treatment of solid tumors (53, 54). Although currently available DNA methyltransferase inhibitors lack specificity for gene(s) of interest, locus-specific therapies are currently being developed (55, 56).
Acknowledgments
Acknowledgment
The authors thank the University of Colorado Genomics Core staff for collection of Illumina DNA methylation and Affymetrix gene expression data, as well as their patients and workers for participation in this study. They also thank Megan Marchant for excellent administrative support for the manuscript.
Footnotes
This work was supported by National Institutes of Health–National Institute of Environmental Health Sciences R01-ES023826 and National Heath Lung and Blood Institute R01-HL140357 (PIs L.A.M. and I.V.Y.).
Author Contributions: I.V.Y., T.E.F., and L.A.M. designed the study; N.H. and L.A.M. performed clinical phenotyping and bronchoscopies; P.M.M. obtained informed consent and managed the clinical database; K.M., L.L., E.J.D., and M.G. collected laboratory data. I.V.Y., I.K., and L.J.S. analyzed the genomic data; I.V.Y. and L.A.M. wrote the manuscript; all authors read and approved the manuscript.
This article has a data supplement, which is accessible from this issue’s table of contents at www.atsjournals.org.
Originally Published in Press as DOI: 10.1165/rcmb.2018-0177OC on August 24, 2018
Author disclosures are available with the text of this article at www.atsjournals.org.
References
- 1.Mayer AS, Hamzeh N, Maier LA. Sarcoidosis and chronic beryllium disease: similarities and differences. Semin Respir Crit Care Med. 2014;35:316–329. doi: 10.1055/s-0034-1377059. [DOI] [PubMed] [Google Scholar]
- 2.Rybicki BA, Major M, Popovich J, Jr, Maliarik MJ, Iannuzzi MC. Racial differences in sarcoidosis incidence: a 5-year study in a health maintenance organization. Am J Epidemiol. 1997;145:234–241. doi: 10.1093/oxfordjournals.aje.a009096. [DOI] [PubMed] [Google Scholar]
- 3.Infante PF, Newman LS. Beryllium exposure and chronic beryllium disease. Lancet. 2004;363:415–416. doi: 10.1016/S0140-6736(04)15523-2. [DOI] [PubMed] [Google Scholar]
- 4.Newman LS, Mroz MM, Balkissoon R, Maier LA. Beryllium sensitization progresses to chronic beryllium disease: a longitudinal study of disease risk. Am J Respir Crit Care Med. 2005;171:54–60. doi: 10.1164/rccm.200402-190OC. [DOI] [PubMed] [Google Scholar]
- 5.Erdal BS, Clymer BD, Yildiz VO, Julian MW, Crouser ED. Unexpectedly high prevalence of sarcoidosis in a representative U.S. metropolitan population. Respir Med. 2012;106:893–899. doi: 10.1016/j.rmed.2012.02.007. [DOI] [PubMed] [Google Scholar]
- 6.Cox CE, Donohue JF, Brown CD, Kataria YP, Judson MA. The Sarcoidosis Health Questionnaire: a new measure of health-related quality of life. Am J Respir Crit Care Med. 2003;168:323–329. doi: 10.1164/rccm.200211-1343OC. [DOI] [PubMed] [Google Scholar]
- 7.Li L, Silveira LJ, Hamzeh N, Gillespie M, Mroz PM, Mayer AS, et al. Beryllium-induced lung disease exhibits expression profiles similar to sarcoidosis. Eur Respir J. 2016;47:1797–1808. doi: 10.1183/13993003.01469-2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Silveira LJ, McCanlies EC, Fingerlin TE, Van Dyke MV, Mroz MM, Strand M, et al. Chronic beryllium disease, HLA-DPB1, and the DP peptide binding groove. J Immunol. 2012;189:4014–4023. doi: 10.4049/jimmunol.1200798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Maier LA, McGrath DS, Sato H, Lympany P, Welsh K, Du Bois R, et al. Influence of MHC class II in susceptibility to beryllium sensitization and chronic beryllium disease. J Immunol. 2003;171:6910–6918. doi: 10.4049/jimmunol.171.12.6910. [DOI] [PubMed] [Google Scholar]
- 10.Fontenot AP, Canavera SJ, Gharavi L, Newman LS, Kotzin BL. Target organ localization of memory CD4(+) T cells in patients with chronic beryllium disease. J Clin Invest. 2002;110:1473–1482. doi: 10.1172/JCI15846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Grunewald J, Eklund A. Löfgren’s syndrome: human leukocyte antigen strongly influences the disease course. Am J Respir Crit Care Med. 2009;179:307–312. doi: 10.1164/rccm.200807-1082OC. [DOI] [PubMed] [Google Scholar]
- 12.Franke A, Fischer A, Nothnagel M, Becker C, Grabe N, Till A, et al. Genome-wide association analysis in sarcoidosis and Crohn’s disease unravels a common susceptibility locus on 10p12.2. Gastroenterology. 2008;135:1207–1215. doi: 10.1053/j.gastro.2008.07.017. [DOI] [PubMed] [Google Scholar]
- 13.Sato H, Woodhead FA, Ahmad T, Grutters JC, Spagnolo P, van den Bosch JM, et al. Sarcoidosis HLA class II genotyping distinguishes differences of clinical phenotype across ethnic groups. Hum Mol Genet. 2010;19:4100–4111. doi: 10.1093/hmg/ddq325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Huehn J, Polansky JK, Hamann A. Epigenetic control of FOXP3 expression: the key to a stable regulatory T-cell lineage? Nat Rev Immunol. 2009;9:83–89. doi: 10.1038/nri2474. [DOI] [PubMed] [Google Scholar]
- 15.Jones B, Chen J. Inhibition of IFN-gamma transcription by site-specific methylation during T helper cell development. EMBO J. 2006;25:2443–2452. doi: 10.1038/sj.emboj.7601148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lal G, Bromberg JS. Epigenetic mechanisms of regulation of Foxp3 expression. Blood. 2009;114:3727–3735. doi: 10.1182/blood-2009-05-219584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee DU, Agarwal S, Rao A. Th2 lineage commitment and efficient IL-4 production involves extended demethylation of the IL-4 gene. Immunity. 2002;16:649–660. doi: 10.1016/s1074-7613(02)00314-x. [DOI] [PubMed] [Google Scholar]
- 18.Miyara M, Amoura Z, Parizot C, Badoual C, Dorgham K, Trad S, et al. The immune paradox of sarcoidosis and regulatory T cells. J Exp Med. 2006;203:359–370. doi: 10.1084/jem.20050648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tykocinski LO, Hajkova P, Chang HD, Stamm T, Sözeri O, Löhning M, et al. A critical control element for interleukin-4 memory expression in T helper lymphocytes. J Biol Chem. 2005;280:28177–28185. doi: 10.1074/jbc.M502038200. [DOI] [PubMed] [Google Scholar]
- 20.Wei G, Wei L, Zhu J, Zang C, Hu-Li J, Yao Z, et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity. 2009;30:155–167. doi: 10.1016/j.immuni.2008.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Young HA, Ghosh P, Ye J, Lederer J, Lichtman A, Gerard JR, et al. Differentiation of the T helper phenotypes by analysis of the methylation state of the IFN-gamma gene. J Immunol. 1994;153:3603–3610. [PubMed] [Google Scholar]
- 22.Santangelo S, Cousins DJ, Winkelmann NE, Staynov DZ. DNA methylation changes at human Th2 cytokine genes coincide with DNase I hypersensitive site formation during CD4(+) T cell differentiation. J Immunol. 2002;169:1893–1903. doi: 10.4049/jimmunol.169.4.1893. [DOI] [PubMed] [Google Scholar]
- 23.Lockstone HE, Sanderson S, Kulakova N, Baban D, Leonard A, Kok WL, et al. Gene set analysis of lung samples provides insight into pathogenesis of progressive, fibrotic pulmonary sarcoidosis. Am J Respir Crit Care Med. 2010;181:1367–1375. doi: 10.1164/rccm.200912-1855OC. [DOI] [PubMed] [Google Scholar]
- 24.Yang IV, Pedersen BS, Rabinovich E, Hennessy CE, Davidson EJ, Murphy E, et al. Relationship of DNA methylation and gene expression in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med. 2014;190:1263–1272. doi: 10.1164/rccm.201408-1452OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chen ES, Moller DR. Etiologies of sarcoidosis. Clin Rev Allergy Immunol. 2015;49:6–18. doi: 10.1007/s12016-015-8481-z. [DOI] [PubMed] [Google Scholar]
- 26.Yang IV, Pedersen BS, Liu AH, O’Connor GT, Pillai D, Kattan M, et al. The nasal methylome and childhood atopic asthma. J Allergy Clin Immunol. 2017;139:1478–1488. doi: 10.1016/j.jaci.2016.07.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Price ME, Cotton AM, Lam LL, Farré P, Emberly E, Brown CJ, et al. Additional annotation enhances potential for biologically-relevant analysis of the Illumina Infinium HumanMethylation450 BeadChip array. Epigenetics Chromatin. 2013;6:4. doi: 10.1186/1756-8935-6-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Maksimovic J, Gordon L, Oshlack A. SWAN: subset-quantile within array normalization for illumina infinium HumanMethylation450 BeadChips. Genome Biol. 2012;13:R44. doi: 10.1186/gb-2012-13-6-r44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B. 1995;57:289–300. [Google Scholar]
- 30.Xia J, Gill EE, Hancock RE. NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc. 2015;10:823–844. doi: 10.1038/nprot.2015.052. [DOI] [PubMed] [Google Scholar]
- 31.Krämer A, Green J, Pollard J, Jr, Tugendreich S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics. 2014;30:523–530. doi: 10.1093/bioinformatics/btt703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Maier LA, Sawyer RT, Tinkle SS, Kittle LA, Barker EA, Balkissoon R, et al. IL-4 fails to regulate in vitro beryllium-induced cytokines in berylliosis. Eur Respir J. 2001;17:403–415. doi: 10.1183/09031936.01.17304030. [DOI] [PubMed] [Google Scholar]
- 33.Newman LS, Bobka C, Schumacher B, Daniloff E, Zhen B, Mroz MM, et al. Compartmentalized immune response reflects clinical severity of beryllium disease. Am J Respir Crit Care Med. 1994;150:135–142. doi: 10.1164/ajrccm.150.1.8025739. [DOI] [PubMed] [Google Scholar]
- 34.Sato H, Silveira L, Spagnolo P, Gillespie M, Gottschall EB, Welsh KI, et al. CC chemokine receptor 5 gene polymorphisms in beryllium disease. Eur Respir J. 2010;36:331–338. doi: 10.1183/09031936.00107809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ramstein J, Broos CE, Simpson LJ, Ansel KM, Sun SA, Ho ME, et al. IFN-γ–producing T-helper 17.1 cells are increased in sarcoidosis and are more prevalent than T-helper type 1 cells. Am J Respir Crit Care Med. 2016;193:1281–1291. doi: 10.1164/rccm.201507-1499OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fontenot AP, Maier LA. Genetic susceptibility and immune-mediated destruction in beryllium-induced disease. Trends Immunol. 2005;26:543–549. doi: 10.1016/j.it.2005.08.004. [DOI] [PubMed] [Google Scholar]
- 37.Samuel G, Maier LA. Immunology of chronic beryllium disease. Curr Opin Allergy Clin Immunol. 2008;8:126–134. doi: 10.1097/ACI.0b013e3282f824a4. [DOI] [PubMed] [Google Scholar]
- 38.Zhou T, Casanova N, Pouladi N, Wang T, Lussier Y, Knox KS, et al. Identification of Jak-STAT signaling involvement in sarcoidosis severity via a novel microRNA-regulated peripheral blood mononuclear cell gene signature. Sci Rep. 2017;7:4237. doi: 10.1038/s41598-017-04109-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hall SC, Agrawal DK. Vitamin d and bronchial asthma: an overview of data from the past 5 years. Clin Ther. 2017;39:917–929. doi: 10.1016/j.clinthera.2017.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Burke RR, Rybicki BA, Rao DS. Calcium and vitamin D in sarcoidosis: how to assess and manage. Semin Respir Crit Care Med. 2010;31:474–484. doi: 10.1055/s-0030-1262215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Culver DA, Barna BP, Raychaudhuri B, Bonfield TL, Abraham S, Malur A, et al. Peroxisome proliferator–activated receptor γ activity is deficient in alveolar macrophages in pulmonary sarcoidosis. Am J Respir Cell Mol Biol. 2004;30:1–5. doi: 10.1165/rcmb.2003-0304RC. [DOI] [PubMed] [Google Scholar]
- 42.Zhu H, Wang G, Qian J. Transcription factors as readers and effectors of DNA methylation. Nat Rev Genet. 2016;17:551–565. doi: 10.1038/nrg.2016.83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Brocks D, Schmidt CR, Daskalakis M, Jang HS, Shah NM, Li D, et al. DNMT and HDAC inhibitors induce cryptic transcription start sites encoded in long terminal repeats. Nat Genet. 2017;49:1052–1060. doi: 10.1038/ng.3889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Shukla S, Kavak E, Gregory M, Imashimizu M, Shutinoski B, Kashlev M, et al. CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing. Nature. 2011;479:74–79. doi: 10.1038/nature10442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Yang IV, Pedersen BS, Liu A, O’Connor GT, Teach SJ, Kattan M, et al. DNA methylation and childhood asthma in the inner city. J Allergy Clin Immunol. 2015;136:69–80. doi: 10.1016/j.jaci.2015.01.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Qiu W, Baccarelli A, Carey VJ, Boutaoui N, Bacherman H, Klanderman B, et al. Variable DNA methylation is associated with chronic obstructive pulmonary disease and lung function. Am J Respir Crit Care Med. 2012;185:373–381. doi: 10.1164/rccm.201108-1382OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lappalainen T, Greally JM. Associating cellular epigenetic models with human phenotypes. Nat Rev Genet. 2017;18:441–451. doi: 10.1038/nrg.2017.32. [DOI] [PubMed] [Google Scholar]
- 48.Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, et al. Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–330. doi: 10.1038/nature14248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Relton CL, Davey Smith G. Two-step epigenetic Mendelian randomization: a strategy for establishing the causal role of epigenetic processes in pathways to disease. Int J Epidemiol. 2012;41:161–176. doi: 10.1093/ije/dyr233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Koth LL, Solberg OD, Peng JC, Bhakta NR, Nguyen CP, Woodruff PG. Sarcoidosis blood transcriptome reflects lung inflammation and overlaps with tuberculosis. Am J Respir Crit Care Med. 2011;184:1153–1163. doi: 10.1164/rccm.201106-1143OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kaminskas E, Farrell AT, Wang YC, Sridhara R, Pazdur R. FDA drug approval summary: azacitidine (5-azacytidine, Vidaza) for injectable suspension. Oncologist. 2005;10:176–182. doi: 10.1634/theoncologist.10-3-176. [DOI] [PubMed] [Google Scholar]
- 52.Saba HI. Decitabine in the treatment of myelodysplastic syndromes. Ther Clin Risk Manag. 2007;3:807–817. [PMC free article] [PubMed] [Google Scholar]
- 53.Brock MV, Hooker CM, Ota-Machida E, Han Y, Guo M, Ames S, et al. DNA methylation markers and early recurrence in stage I lung cancer. N Engl J Med. 2008;358:1118–1128. doi: 10.1056/NEJMoa0706550. [DOI] [PubMed] [Google Scholar]
- 54.Juergens RA, Wrangle J, Vendetti FP, Murphy SC, Zhao M, Coleman B, et al. Combination epigenetic therapy has efficacy in patients with refractory advanced non–small cell lung cancer. Cancer Discov. 2011;1:598–607. doi: 10.1158/2159-8290.CD-11-0214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Di Ruscio A, Ebralidze AK, Benoukraf T, Amabile G, Goff LA, Terragni J, et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature. 2013;503:371–376. doi: 10.1038/nature12598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Yu DH, Waterland RA, Zhang P, Schady D, Chen MH, Guan Y, et al. Targeted p16(Ink4a) epimutation causes tumorigenesis and reduces survival in mice. J Clin Invest. 2014;124:3708–3712. doi: 10.1172/JCI76507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Prokopenko I, Poon W, Mägi R, Prasad B R, Salehi SA, Almgren P, et al. A central role for GRB10 in regulation of islet function in man. PLoS Genet. 2014;10:e1004235. doi: 10.1371/journal.pgen.1004235. [DOI] [PMC free article] [PubMed] [Google Scholar]