Abstract
Objective
Nonalcoholic steatohepatitis (NASH) is closely associated with metabolic syndrome and increases the risk for end-stage liver disease, such as cirrhosis and hepatocellular carcinoma. Despite this, the molecular events that influence NASH pathogenesis remain poorly understood. The objectives of the current study are to delineate the transcriptomic and proteomic signatures of NASH liver, to identify potential pathogenic pathways and factors, and to critically assess their role in NASH pathogenesis.
Methods
We performed RNA sequencing and quantitative proteomic analyses on the livers from healthy and diet-induced NASH mice. We examined the association between plasma levels of TSK, a newly discovered hepatokine, and NASH pathologies and reversal in response to dietary switch in mice. Using TSK knockout mouse model, we determined how TSK deficiency modulates key aspects of NASH pathogenesis.
Results
RNA sequencing and quantitative proteomic analyses revealed that diet-induced NASH triggers concordant reprogramming of the liver transcriptome and proteome in mice. NASH pathogenesis is linked to elevated plasma levels of the hepatokine TSK, whereas dietary switch reverses NASH pathologies and reduces circulating TSK concentrations. Finally, TSK inactivation protects mice from diet-induced NASH and liver transcriptome remodeling.
Conclusions
Global transcriptomic and proteomic profiling of healthy and NASH livers revealed the molecular signatures of diet-induced NASH and dysregulation of the liver secretome. Our study illustrates a novel pathogenic mechanism through which elevated TSK in circulation promotes NASH pathologies, thereby revealing a potential target for therapeutic intervention.
Keywords: NASH, Transcriptome, Proteomics, Secreted factor, Hepatokine, Type I interferon, NAFLD, TSK
Abbreviations: ALT, Alanine aminotransferase; AST, Aspartate aminotransferase; HFD, High-fat diet; IFN, interferon; KO, Knockout; MCD, Methionine choline-deficient; NAFLD, Nonalcoholic fatty liver disease; NAS, NASH activity score; NASH, Nonalcoholic steatohepatitis; qPCR, quantitative PCR; TAG, Triglyceride; TSK, Tsukushi; WAT, White adipose tissue; WT, Wild type
Highlights
-
•
NASH triggers concordant reprogramming of the liver transcriptome and proteome.
-
•
NASH pathogenesis is linked to elevated plasma levels of the hepatokine TSK.
-
•
Dietary switch reverses NASH pathologies and reduces plasma TSK levels.
-
•
TSK inactivation protects mice from diet-induced NASH and liver transcriptome remodeling.
1. Introduction
Nonalcoholic fatty liver disease (NAFLD) is characterized by excess lipid accumulation in the liver (hepatic steatosis) and is emerging as the most common chronic liver disease [1], [2], [3], [4]. NAFLD represents the hepatic manifestation of metabolic syndrome and, not surprisingly, closely mirrors the global epidemic of obesity and type 2 diabetes. NAFLD encompasses a spectrum of liver pathologies ranging from generally benign fatty liver to steatosis that occurs in the presence of liver injury, inflammation and fibrosis. The latter was recognized as a clinically distinct disease entity termed nonalcoholic steatohepatitis (NASH) in 1980 [5]. The global burden of NAFLD is estimated to be approximately 24% among the general population with 20–30% NAFLD patients further progressing to NASH [6], [7]. NASH greatly increases the risk for end-stage liver disease, such as cirrhosis and hepatocellular carcinoma that may occur with or without cirrhosis. As a result, NASH has been projected to be the leading cause for end-stage liver disease that requires liver transplantation in near future [8]. Despite this, no effective therapeutic interventions are currently available for treating NASH, underscoring the urgent need to better understand the etiology and the progressive nature of NASH pathogenesis.
Hepatic steatosis may be caused by liver-intrinsic mechanisms, such as increased de novo lipogenesis, impaired fatty acid β-oxidation, and/or reduced lipoprotein secretion [2], [3]. In parallel, adipose tissue inflammation and dysfunction greatly increase the delivery of lipids to the liver, serving as an extrinsic factor that promotes hepatic steatosis in insulin resistant state. Beyond the protein factors mediating metabolic signaling, recent studies have implicated microRNAs and long noncoding RNAs as an integral component of the regulatory network that governs hepatic lipid homeostasis [9], [10], [11]. A lack of suitable dietary models that faithfully recapitulate key events of NASH pathogenesis has been a long-standing challenge in the field. While high-fat diet (HFD) feeding provides a robust means to induce obesity, insulin resistance and hepatic steatosis, the commonly used HFD fails to drive NAFLD progression beyond steatosis. On the contrary, methionine and choline-deficient diet (MCD) feeding leads to liver injury, inflammation, and fibrosis in mice [12]; however, these pathologies occur in the absence of obesity and insulin resistance, and as such, the MCD model is inadequate in mimicking the pathophysiological milieu of human NASH. Recent studies have demonstrated that a high-fat high-fructose diet containing 2% cholesterol is highly effective in inducing key features of NASH in mice [13], [14]. However, the molecular nature of liver transcriptome and proteome reprogramming during diet-induced NASH has not been elucidated.
Endocrine factors are important regulators of tissue metabolism and energy homeostasis. Adipose tissue hormones, such as leptin and adiponectin from white fat [15], [16], [17] and Neuregulin 4 from brown fat [18], [19], [20], gut-derived factors [21], [22], and myokines [23] participate in nutrient sensing and coordinate diverse aspects of nutrient and energy metabolism. Beyond its central role in glucose and lipid metabolism, the mammalian liver secretes an array of proteins into circulation, including coagulation factors and endocrine hormones (hepatokines) [24], [25]. FGF21 is perhaps the best studied nutrient-responsive hepatokine that has been shown to elicit beneficial effects on whole body metabolism at pharmacological doses [22], [26], [27]. By analyzing secretome gene expression, we recently identified Tsukushi (TSK) as a hepatokine that exerts powerful effects on adipose tissue thermogenesis and metabolic homeostasis [28]. In this study, we performed RNA-sequencing (RNA-seq) and quantitative proteomic analyses to elucidate the landscape of transcriptome and proteome reprogramming in NASH. We show that plasma TSK levels are tightly linked to NASH pathologies and that its inactivation powerfully attenuates diet-induced NASH pathogenesis.
2. Materials and methods
2.1. Mouse studies
All animal studies were performed following procedures approved by the Institutional Animal Care & Use Committee at the University of Michigan. Mice were maintained in a specific pathogen-free facility under 12/12-hr light/dark cycles with free access to food and water. C57/Bl6 male mice were fed standard chow (Teklad 5001 Laboratory Diet) or NASH diet containing 40% fat (of which 18% was trans-fat), 22% fructose, and 2% cholesterol (D09100301, Research Diets Inc.) starting at 3 months of age. For MCD feeding, WT C57BL/6 male and db/db mice were fed chow or MCD diet for four weeks. For NASH-chow diet switch, WT C57BL/6 male mice were fed NASH for 3 months before dividing into two groups for continued NASH diet feeding or switch to chow for two months. Tsk knockout mice were previously described [29] and maintained in C57BL/6 background.
Total liver RNA was extracted using Trizol (Alkali Scientific, TRZ-100). Liver RNA sequencing was performed using Illumina HiSeq 4000 at the University of Michigan DNA Sequencing Core. Raw sequencing read counts were normalized and processed for differential gene expression analysis using DESeq2 [30]. The significant expressed genes were determined by FDRs less than 0.05. All RNA-seq data generated in this work have been deposited into the Gene Expression Omnibus (GEO) database (GSE119340). Clustering analysis was performed using Cluster and visualized using Treeview. Gene ontology analysis was performed with Metascape (www.metascape.org).
2.2. Quantitative proteomic analysis
TMT experiments were performed as previously described [31]. Briefly, total liver proteins were extracted, reduced with 2 mM DTT (10 min) and alkylated with 50 mM iodoacetamide in dark (30 min). Proteins were digested by Lys-C (Wako, at a 1:100 enzyme/protein ratio) at room temperature for 2 h followed by overnight digestion with trypsin (Thermo Fisher Scientific, at 1:100 enzyme/protein ratio). The resulting peptides were desalted using Oasis HLB cartridges (Waters), resuspended in 200 mM HEPES (pH 8.5) to a final concentration of 1 μg/μL, and labeled with amine-based Sixplex TMT reagents (Thermo Fisher Scientific). Three replicate samples were prepared for the control samples (TMT-126, -127 and -128), and the NASH samples (TMT-129, -130 and -131), respectively.
Samples fractioned by bRPLC (basic pH reversed phase HPLC) and analyzed by LC-MS/MS. MS/MS spectra were searched against a composite database of the mouse UniProt protein database and its reversed complement using the Sequest (Rev28) algorithm. For TMT quantification, a 0.03 Th window was scanned around the theoretical m/z of each reporter ion (126:126.127725; 127:127.124760; 128:128.134433; 129:129.131468; 130:130.141141; 131:131.138176). The maximum intensity of each reporter ion was extracted, which was converted to signal-to-noise (S/N) ratios.
2.3. Plasma and liver metabolite measurements
Plasma concentrations of TAG, NEFA, cholesterol, ALT and AST were measured using the following commercial kits according to the manufacture's instruction: TAG (Sigma, TR0100); NEFA (Wako Diagnostics, 993-35191); cholesterol (Stanbio, SB-1010-430); ALT (Stanbio, 2930); AST (Stanbio, 2920). For liver hydroxyproline content, liver tissue was homogenized in water and samples were hydrolyzed by incubation with 6N hydrochloric acid at 120 °C for 3 h, followed by measurement using Hydroxyproline Colorimetric Assay Kit (BioVision, K555-100). Liver TAG was extracted as previously described [32].
2.4. Gene expression analyses
Hepatic gene expression was analyzed using qPCR, as previously described [11], [32]. Briefly, mouse livers were immediately frozen in liquid nitrogen after being removed. Total liver RNA was extracted using TRIzol method following manufacturer instructions. 2 μg of total RNA was reverse-transcribed using MMLV-RT followed by qPCR using SYBR Green (Thermo Fisher). Relative mRNA abundance was normalized to internal control genes encoding ribosomal protein 36B4. The sequences of the qPCR primers are listed in Supplementary Table S1.
2.5. Western blot analyses
Total liver protein extracts were prepared using a lysis buffer containing 50 mM Tris–HCl (pH = 7.5), 137 mM NaCl, 1 mM EDTA, 1% Triton X-100, 10% glycerol, 10 mM NaF, 10 mM Na4P2O7, 1 mM Na3VO4, and protease inhibitor cocktail. The lysates were separated by SDS-PAGE and transferred to a PVDF membrane, followed by immunoblotting with primary antibodies and secondary antibodies. The following antibodies were used: anti-TBK1 (Cell Signaling Technology, 3013); anti-phospho-TBK1S172 (Cell Signaling Technology, 5483); anti-IFIT1 (Thermo Fisher, PA3-846); anti-IFIT3 (ProteinTech, 15201-1-AP); anti-ZBP1; anti-STAT1 (Cell Signaling Technology, 9172); anti-phospho-STAT1Y701 (Cell Signaling Technology, 9171); anti-HSP90 (Santa Cruz BioTech, sc-7947). TSK antibody was generated in rabbits with a synthetic peptide corresponding to mouse TSK (amino acids 317-335; CRRLVREGAYHRQPGSSPK) and affinity purified.
2.6. Histology analyses
Liver tissues were immediately fixed in 10% formalin at 4 °C overnight after being removed and processed for paraffin embedding and H&E staining. Sirius red staining was performed as previously described [14], [20]. Percentage of Sirius red staining positive area of the total area of view was quantified using Image J. Liver histology was evaluated by a blinded hepatologist and NASH activity score (NAS) was assessed [33].
2.7. Statistical analyses
Statistical analysis was performed using GraphPad Prism 6. Statistical differences were determined using two-tailed unpaired Student's t-test. A p value of less than 0.05 (*p < 0.05, **p < 0.01, and ***p < 0.001) was considered statistically significant. Statistical methods and corresponding p values for data shown in each panel were included in figure legends.
2.8. Data availability
RNA-seq data files have been deposited into the Gene Expression Omnibus database (www.ncbi.nlm.nih.gov/geo/) with accession number GSE119340.
3. Results
3.1. Reprogramming of the liver transcriptome during diet-induced NASH
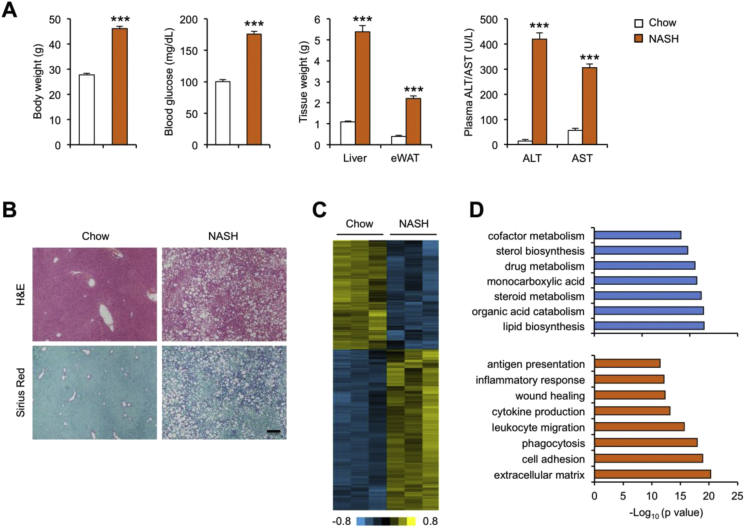
To elucidate the molecular nature of liver transcriptome and proteome reprogramming in NASH, we first performed whole liver RNA-seq on mice fed chow or NASH diet for six months. As expected, mice fed NASH diet developed obesity and insulin resistance, as indicated by significantly elevated blood glucose levels (Figure 1A). Compared to control, NASH diet-fed mice had increased adipose tissue and liver mass and exhibited severe hepatic steatosis (Figure 1B). Plasma concentrations of alanine aminotransferase (ALT) and aspartate aminotransferase (AST), markers of liver injury, were also greatly elevated following NASH diet feeding. Sirius Red staining of liver sections revealed prominent pericellular fibrosis, a hallmark of NASH pathologies (Figure 1B). These observations demonstrate that this feeding model is highly effective in inducing NASH in the context of obesity and insulin resistance.
Figure 1.
RNA-seq analysis of chow and NASH mouse livers. (A) Metabolic parameters of mice fed chow (n = 4) or NASH (n = 9) diet. (B) H&E (top) and Sirius Red (bottom) staining of paraffin-embedded liver sections. Scale bar = 100 μm. (C) Heat map of upregulated or downregulated gene clusters following NASH diet feeding. (D) Gene ontology analysis of downregulated (blue) and upregulated (orange) gene clusters in response to diet-induced NASH. Data represent mean ± sem. ***p < 0.001, NASH vs. chow; two-tailed unpaired Student's t-test.
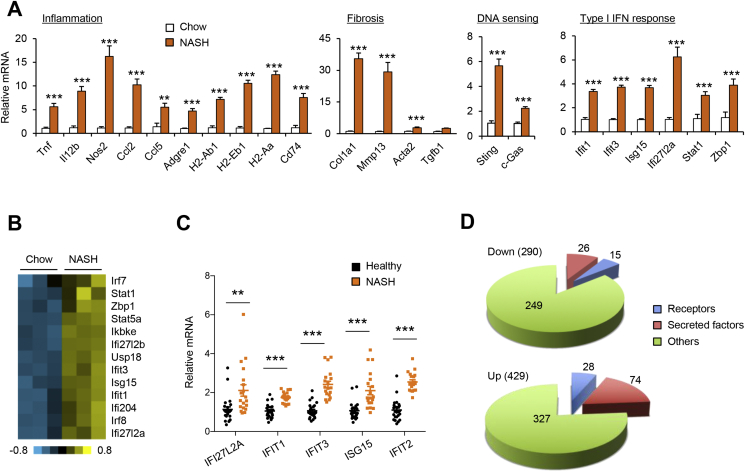
We next performed RNA-seq on whole liver RNA from three pairs of chow and NASH diet-fed mice. Differential gene expression analysis identified two clusters of genes exhibiting over two-fold decreased (290 genes) or increased (429 genes) mRNA expression in NASH livers (Figure 1C and Supplementary Table S2). Gene ontology analysis (www.metascape.org) indicated that downregulated genes were enriched for substrate metabolism, including sterol biosynthesis, cofactor metabolism and steroid metabolism, whereas the upregulated genes were enriched for pathways involved in extracellular matrix production and remodeling, wound healing and various aspects of inflammatory response (Figure 1D). Quantitative PCR (qPCR) analysis confirmed that mRNA expression of genes involved in liver fibrosis (Col1a1, Mmp13, Acta2, Tgfb1), inflammatory signaling (Tnf, Il12b, Nos2, Ccl2, Ccl5, Adgre1), MHC II antigen presentation (H2-Ab1, H2-Eb1, H2-Aa, Cd74) were strongly induced following NASH diet feeding (Figure 2A). Notably, a cluster of genes involved in cytosolic DNA sensing and type I interferon (IFN)-mediated antiviral response exhibited robust induction following diet-induced NASH, including Sting, c-Gas, Ifit1, Ifit3, Isg15, Zbp1, and Ifi27l2b (Figure 2A–B). We next analyzed a published microarray dataset containing samples from 24 healthy and 19 NASH patients (GSE89632) [34]. Similar to diet-induced NASH in mice, mRNA expression of IFIT family members, Isg15 and Ifi27l2a was significantly increased in human NASH livers (Figure 2C). These findings illustrate that hepatic type I IFN signaling is augmented in both mouse and human NASH. Interestingly, liver-specific conditional knockout of type I IFN receptor (Ifnar1) worsened hepatic steatosis and inflammation induced by MCD feeding [35], supporting a functional role of type I IFN signaling in maintaining liver homeostasis under metabolic stress conditions.
Figure 2.
Effects of NASH on hepatic gene expression. (A) qPCR analysis of gene expression in chow (n = 4) and NASH (n = 9) mouse livers. Data represent mean ± sem. **p < 0.01, ***p < 0.001, NASH vs. chow; two-tailed unpaired Student's t-test. (B) Heat map of a cluster of genes involved in antiviral and type I IFN response. (, suggesting that other pathways may be involved in the beneficial effects of TSK deficiency in the liver C) Expression of type I IFN response genes in healthy (n = 24) and NASH (n = 19) human livers. Microarray expression values were obtained from GSE89632. Data represent mean ± sem. **p < 0.01, ***p < 0.001, NASH vs. healthy; two-tailed unpaired Student's t-test. (D) Pie charts indicating the numbers of genes encoding secreted factors, membrane receptors and other proteins in the downregulated and upregulated gene clusters.
Secreted factors and membrane receptors mediate intercellular crosstalk via endocrine and/or paracrine mechanisms and serve critical functions in metabolic physiology and disease. We next examined the extent to which diet-induced NASH leads to reprogramming of the liver secretome and receptor gene expression using a mouse secretome gene list we previously described [20]. This list contains 1,275 and 757 genes that are predicted to encode secreted factors and membrane receptors, respectively. Among 290 downregulated genes in NASH livers, 26 and 15 of which were predicted to encode secreted factors and membrane receptors, respectively (Figure 2D and Supplementary Table S2). Interestingly, 74 and 28 genes encoding secreted factors and membrane receptors showed increased expression in NASH livers. Together, our RNA-seq analysis revealed global reprogramming of the liver transcriptome during NASH pathogenesis, including those encoding signaling ligands and receptors.
3.2. Quantitative proteomic analysis of healthy and diet-induced NASH livers
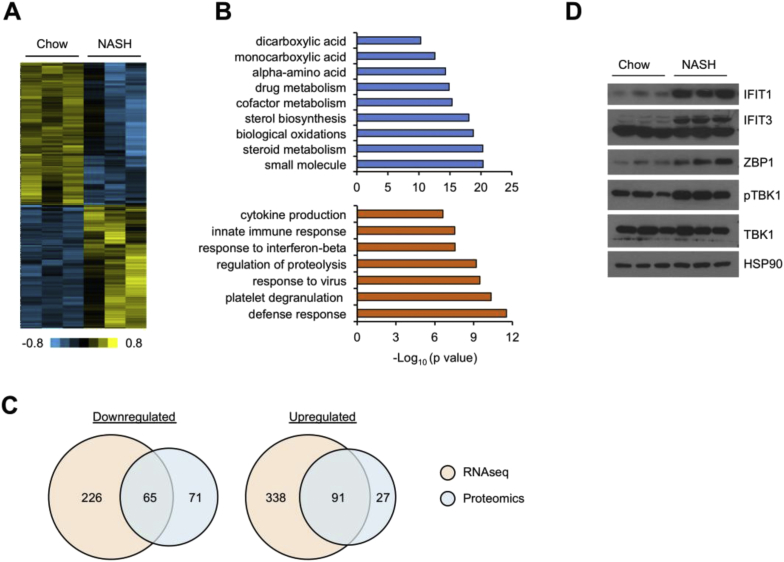
In addition to transcriptional regulation, protein translation and turnover also play important roles in the control of gene expression and function. The extent to which the liver proteome is reprogrammed during NASH pathogenesis remains largely unexplored. We next performed quantitative proteomic analysis using Sixplex Tandom Mass Tag (TMT) that allowed us to simultaneously quantify protein expression in six independent samples [31]. In this case, we analyzed the same three pairs of chow and NASH liver samples using this proteomics pipeline and obtained quantitative expression data for 7007 proteins. Using a fold change cutoff of 1.5-fold, we identified 136 and 118 proteins exhibiting decreased and increased expression in NASH livers, respectively (Figure 3A and Supplementary Table S3). Gene ontology analysis of these differentially regulated genes revealed a picture that was remarkably similar to RNA-seq results. As such, genes with increased protein expression were enriched for defense response, including antiviral and IFN response and innate immune response, whereas genes with decreased protein expression in NASH were enriched for pathways of intermediary metabolism, sterol biosynthesis and steroid metabolism (Figure 3B). Among 118 proteins induced by NASH, 91 also exhibited increased mRNA expression, based on RNA-seq data, indicating a high degree of concordance between mRNA and protein expression (Figure 3C). For the downregulated proteins, 65 of 136 of which showed reduced expression at both mRNA and protein levels. We confirmed NASH-associated induction of protein expression for several genes involved in exotic DNA sensing and type I IFN response, including IFIT1, IFIT3, phosphorylated TBK1, and ZBP1 (Figure 3D). The DNA sensing pathway is critical for stimulating type I IFN secretion in response to cytoplasmic DNA and has been implicated in metabolic inflammation [36], [37]. Whether one or more of these IFN-responsive genes play a role in modulating NASH progression remains currently unknown.
Figure 3.
Quantitative proteomic analysis of chow and NASH mouse livers. (A) Heat map representation of proteins upregulated or downregulated by over 1.5-fold following NASH diet feeding. (B) Gene ontology analysis of downregulated (blue) and upregulated (orange) protein clusters following diet-induced NASH. (C) Comparison between RNA-seq and proteomic analyses. The numbers of differentially regulated genes at both mRNA and protein levels are indicated in the intersection. (D) Immunoblotting of total liver lysates.
3.3. NASH pathogenesis is linked to marked elevation of the hepatokine TSK in plasma
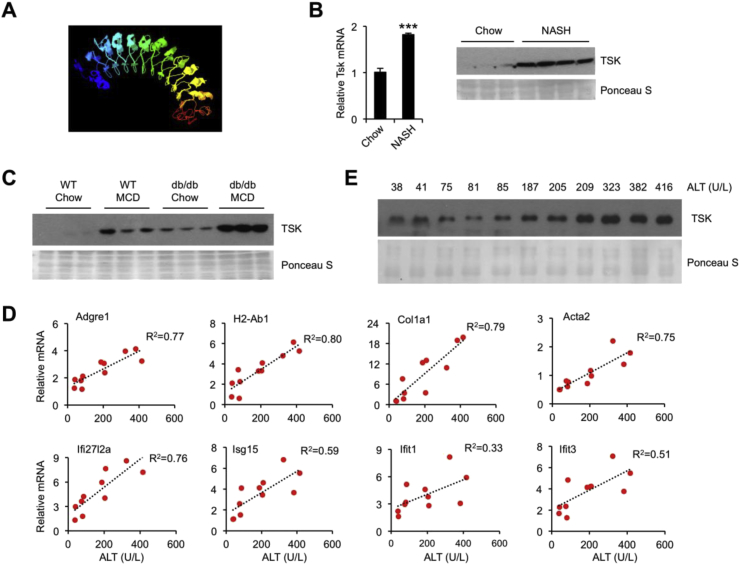
Hepatokines are a group of liver-derived secreted factors that exert pleiotropic effects on metabolic physiology and disease pathogenesis [24], [25]. For example, FGF21 [38] and Fetuins [39], [40] have been shown to regulate diverse aspects of energy and nutrient metabolism by acting on distal target tissues. The profound reprogramming of the hepatic secretome during diet-induced NASH raised the possibility that aberrant hepatokine release may contribute to the development of NASH pathologies. We recently analyzed secretome gene expression among a panel of mouse tissues and identified Tsukushi (TSK) as a novel hepatokine secreted by parenchymal hepatocytes [28]. TSK is a highly conserved proteoglycan in mammals [41] that contains ten leucine-rich repeats and is predicted to fold into a horseshoe-shaped structure (Figure 4A). The molecular identity of the TSK receptor remains currently unknown. Plasma TSK levels were abnormally elevated in obese mice and contribute to HFD-induced obesity and insulin resistance in part through attenuation of adipose tissue thermogenesis. However, whether hepatic Tsk expression and its plasma levels are dysregulated during diet-induced NASH and its role in NASH pathogenesis have not been explored.
Figure 4.
Elevated plasma TSK levels in mouse NASH models. (A) Structural modeling of mouse TSK. (B) qPCR analysis of TSK gene expression and immunoblotting of TSK in mice fed chow (n = 4) or NASH (n = 4) diet for six months. Data represent mean ± sem. ***p < 0.001, chow vs. NASH; two-tailed unpaired Student's t-test. (C) Immunoblotting of TSK in plasma from WT and db/db mice fed chow or MCD diet for four weeks. (D) Association between hepatic gene expression and plasma ALT concentrations in a cohort of mice fed NASH diet for three months. (E) Immunoblotting of TSK in mouse plasma arranged according to their plasma ALT levels, as indicated.
To address this, we first examined Tsk expression and its plasma levels in mice fed standard chow and NASH diet for six months. Compared to chow control, NASH diet feeding significantly increased TSK mRNA expression in the liver, resulting in markedly elevated levels of circulating TSK in mice following NASH diet feeding (Figure 4B). We noted Tsk mRNA induction appeared less pronounced compared to the increase of its protein levels, suggesting that additional mechanisms may modulate TSK secretion and/or turnover and influence its plasma levels. Previous studies have demonstrated that Leptin receptor-deficient (db/db) mice developed classic NASH pathologies following MCD feeding [42]. We next performed immunoblotting analysis to determine whether plasma TSK is altered in this mouse NASH model. Compared to chow-fed C57BL/6 wild type (WT) mice, elevated plasma TSK was readily observed in WT mice fed MCD and in db/db mice (Figure 4C). MCD feeding in db/db mice resulted in further increase of TSK levels in circulation, supporting the notion that plasma TSK levels are tightly linked to the severity of liver injury. To further assess this, we examined hepatic gene expression and plasma TSK levels in a cohort of mice exhibiting varying degree of liver injury following three months of NASH diet feeding. As expected, mRNA expression of Adgre1, H2-Ab1, Col1a1, Acta2, and genes involved in type I IFN response (Ifi27l2a, Isg15, Ifit1 and Ifit3) exhibited strong correlation with plasma ALT levels (Figure 4D). Importantly, plasma TSK levels also exhibited strong correlation with ALT levels in NASH mice (Figure 4E). We concluded from these studies that TSK is a hepatokine aberrantly elevated in NASH that may potentially contribute to NASH pathogenesis.
3.4. Plasma TSK is normalized following NASH reversal
In humans, weight loss is a commonly recommended approach for treating patients with NAFLD and NASH [43]; however, the extent to which NASH pathologies are reversible and the underlying mechanisms remain poorly understood. To explore reversibility of NASH pathogenesis, we divided a cohort of mice fed NASH diet for three months into two groups: one remaining on the original NASH diet and the other switched to chow diet for two more months. NASH to chow dietary switch resulted in approximately 10% lower body weight in the switch group (Figure 5A). While adipose tissue mass was slightly reduced, liver mass in the switch group was reduced by approximately 62% compared to NASH group. Additionally, plasma total cholesterol, AST and ALT concentrations were drastically lower in NASH-to-chow switch group, indicative of improved metabolic and liver injury profile (Figure 5B). In support of this, we observed improved hepatic steatosis, alleviated inflammation and reduced fibrosis in mice following the dietary switch (Figure 5C). Accordingly, NASH activity score (NAS), an indicator of NASH severity, was markedly improved (Figure 5D). Liver hydroxyproline content was lower by approximately 60% in the diet switch group.
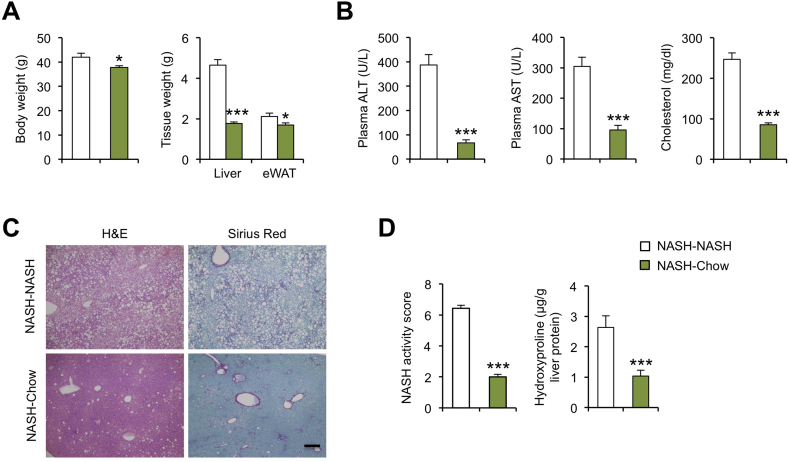
Figure 5.
A dietary model of NASH reversal in mice. (A) Body weight and tissue weight of NASH diet-fed mice kept on NASH diet (NASH–NASH; n = 7) or switched to chow diet (NASH-Chow; n = 9) for two months. (B) Plasma parameters. (C) H&E (left) and Sirius Red (right) staining of paraffin-embedded liver sections. Scale bar = 100 μm. (D) NAS score and Liver hydroxyproline content. (E) qPCR analysis of liver gene expression. Data represent mean ± sem. *p < 0.05, ***p < 0.001, NASH vs. switch; two-tailed unpaired Student's t-test.
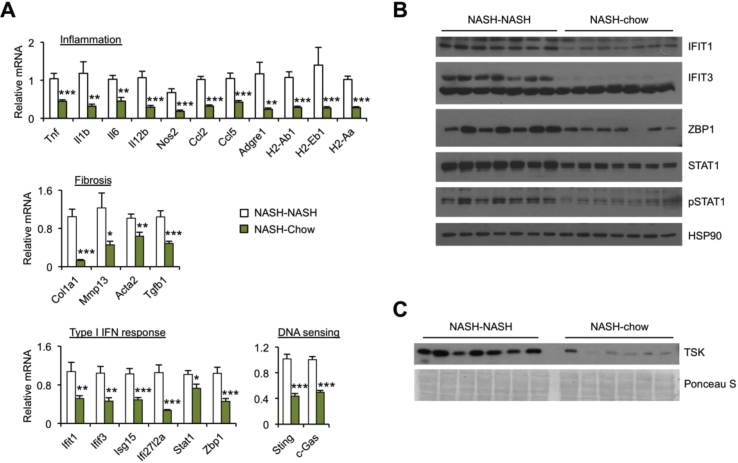
Hepatic gene expression analysis indicated that the expression of genes involved in inflammation and fibrosis was significantly attenuated following cessation of NASH diet feeding, including those involved in inflammatory response, liver fibrosis, DNA sensing and type I IFN response (Figure 6A). Consistently, the dietary switch reduced protein levels of ZBP1, STAT1 and phosphorylated STAT1, factors involved in type I IFN signaling (Figure 6B). These results illustrate that key aspects of NASH pathologies are largely reversible following weight loss caused by the NASH to chow dietary switch. Remarkably, this dietary switch resulted in a striking reduction of plasma TSK levels (Figure 6C), underscoring a tight association between circulating TSK and NASH pathologies.
Figure 6.
Hepatic gene expression analysis. (A) qPCR analysis of liver gene expression in mice kept on NASH diet (NASH–NASH; n = 7) or following switch to chow (NASH-Chow; n = 9) for two months. Data represent mean ± sem. *p < 0.05, **p < 0.01, ***p < 0.001, NASH vs. switch; two-tailed unpaired Student's t-test. (B) Immunoblots of total liver lysates. (C) Immunoblot of TSK in plasma.
3.5. TSK inactivation attenuates diet-induced NASH and restores a healthy liver transcriptome
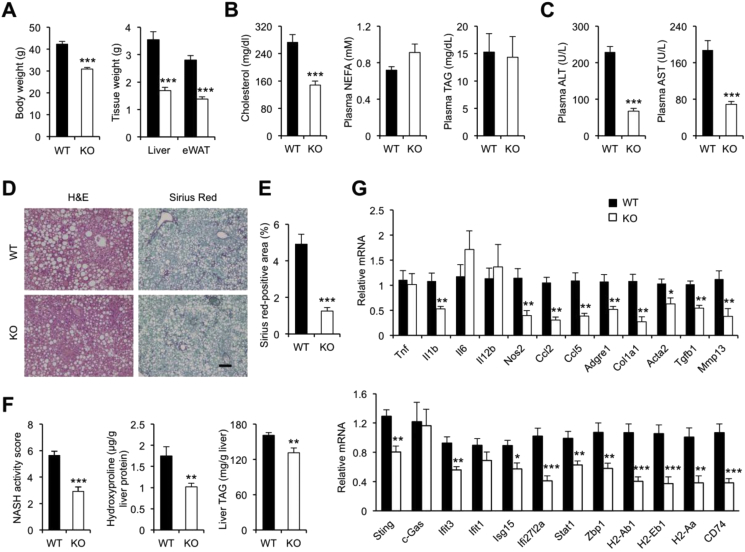
As plasma TSK levels are tightly linked to the severity of NASH pathologies and the liver NASH gene signature, we next sought to determine whether TSK is causally linked to NASH pathogenesis in mice. We recently demonstrated that mice lacking TSK appeared essentially indistinguishable from WT littermate control under chow-fed conditions and exhibited no abnormalities in glucose and lipid homeostasis. To determine whether TSK plays a role in NASH pathogenesis, we subjected WT and TSK whole-body KO mice to NASH diet feeding for five months. Similar to HFD feeding, TSK KO mice gained significantly less body weight and had smaller liver and adipose tissue mass than WT mice (Figure 7A). While plasma total cholesterol levels were significantly lower in Tsk null mice, the concentrations of non-esterified fatty acids and triglycerides remained similar between two groups (Figure 7B). As expected, WT mice exhibited hallmarks of NASH, including elevated plasma ALT and AST levels, hepatic steatosis and liver fibrosis (Figure 7C–D). Compared to control, TSK KO mice had significantly lower plasma ALT and AST levels, indicating that TSK inactivation ameliorates NASH diet-induced liver injury. Histological staining revealed that hepatic steatosis, inflammation and fibrosis were greatly diminished in TSK null mouse livers. NAS and hydroxyproline content were significantly lower in TSK null mouse livers than WT control (Figure 7E–F). TSK null mice also had lower liver TAG content. Hepatic expression of genes involved in inflammatory signaling, liver fibrosis and type I IFN signaling was significantly decreased by TSK inactivation (Figure 7G), whereas mRNA expression of FGF21 in the liver and Nrg4 and adiponectin in adipose tissue remained comparable between two groups. Together, these results demonstrate that TSK deficiency protects mice from diet-induced NASH and restores the liver transcriptome to a healthier state.
Figure 7.
TSK deficiency protects mice from diet-induced NASH. (A) Body weight and tissue weight of WT (n = 11) and Tsk KO (n = 12) mice fed NASH diet for six months. (B) Plasma metabolic parameters. (C) Plasma ALT and AST concentrations. (D) H&E (left) and Sirius Red (right) staining of liver sections (bar = 100 μm). (E) Quantitation of Sirius Red-positive area on liver sections. (F) NAS and liver hydroxyproline and TAG content. (G) qPCR analysis of hepatic gene expression. Data represent mean ± sem. *p < 0.05, **p < 0.01, ***p < 0.001, KO vs. WT; two-tailed unpaired Student's t-test.
4. Discussion
NASH pathogenesis is known to engage several non-parenchymal cell types, including hepatic stellate cells, macrophage, and other immune cells [44], [45], [46], [47], [48], which act in concert to sustain a chronic inflammatory environment and promotes pathogenic tissue remodeling and liver fibrosis. However, the molecular nature of liver transcriptome and proteome reprogramming in NASH remains incompletely understood. Our RNA-seq and quantitative proteomic profiling studies revealed remarkably concordant changes in hepatic gene expression at mRNA and protein levels during diet-induced NASH. Notably, gene ontology analyses of RNA-seq and proteomic datasets predicted the same pathways to be altered in NASH livers. We observed strong induction of NASH gene signatures enriched for immune response, inflammatory signaling, extracellular matrix production and remodeling. These results are consistent with previous transcriptomic analysis of mouse and human NASH livers [13], [49]. Unexpectedly, a cluster of genes involved in antiviral response and type I IFN signaling was also strongly induced following diet-induced NASH. Using a dietary switch model, we demonstrated that NASH pathologies are largely reversible in sync with normalization of the NASH gene signatures.
A remarkable aspect of NASH-associated changes in hepatic gene expression is the profound changes in secretome and membrane receptor gene expression. As ligand/receptor signaling plays an important role in mediating intercellular crosstalk among different liver cell types, it is conceivable that NASH pathogenesis reshapes the nature of cell–cell communications in the liver that may contribute to disease progression. In support of this, we observed a striking association between plasma levels of the hepatokine TSK and the severity of NASH pathologies. TSK is a secreted proteoglycan that has been demonstrated to physically interact with several extracellular ligands, including BMP, FGF, TGFβ1 and Wnt, and modulate their biological activities [41], [50], [51]. TGFβ1 is a growth factor that promotes HSC activation and ECM deposition [2]. As such, it is possible that TSK may directly impinge on these signaling pathways to influence NASH pathogenesis. We recently demonstrated that TSK is a negative regulator of adipose tissue sympathetic innervation, adrenergic signaling and thermogenesis [28]. Mice lacking TSK were protected from HFD-induced weight gain, brown fat whitening and insulin resistance. Therefore, we cannot rule out the possibility that TSK deficiency may protect mice from diet-induced NASH secondarily through the improvements of systemic metabolic profile. While the exact molecular and cellular elements in TSK signaling remain to be elucidated, our results strongly support elevated TSK as a novel pathogenic mechanism in driving NASH progression. These findings also raise the prospect of targeting TSK as a treatment for metabolic disease, including type 2 diabetes and NASH.
Despite the important role of TSK in NASH progression, the pathogenic stimuli that lead to its marked elevation in diet-induced NASH remain to be elucidated. FGF21 has been demonstrated to be highly responsive to various physiological signals, including starvation, certain nutrients and various physiological and environmental stresses [22], [26], [27]. Similarly, hepatic Tsk expression and its serum levels are robustly induced in response to increased energy expenditure, such as during cold exposure and following treatments with thyroid hormone and adrenergic agonist. Plasma TSK is markedly elevated in genetic and diet-induced obese mice as well as diet-induced NASH mice; however, the molecular nature of the signaling pathways underlying this aberrant induction of TSK remains currently unknown and an important unsolved question. It is possible that certain metabolic, inflammatory and/or microbiome-derived signals may promote TSK expression and secretion in metabolic disease.
Author contributions
J.D.L., X.X., and Q.W. conceived the project and designed research. X.X., Q.W., L.G., and J.Z. performed the experiments and analyzed the data. T.L. and J.D.L. performed sequencing data analyses. S.W. and Y.Y. performed quantitative proteomic analysis of chow and NASH mouse livers. X.X. and J.D.L. wrote the manuscript.
Acknowledgements
This work was supported by NIH (DK102456 and DK112800 to J.L. and GM114160 and GM122932 to Y.Y.), American Diabetes Association (1-15-BS-118, J.D.L.) and Welch Foundation (I-1800 to Y.Y.). T.L. was supported by Patten Predoctoral Fellowship provided by the University of Michigan.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.molmet.2018.12.004.
Contributor Information
Xuelian Xiong, Email: xiong.xuelian@zs-hospital.sh.cn.
Jiandie D. Lin, Email: jdlin@umich.edu.
Conflict of interest
The authors declare no conflict of interest.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Cohen J.C., Horton J.D., Hobbs H.H. Human fatty liver disease: old questions and new insights. Science. 2011;332:1519–1523. doi: 10.1126/science.1204265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Friedman S.L., Neuschwander-Tetri B.A., Rinella M., Sanyal A.J. Mechanisms of NAFLD development and therapeutic strategies. Nature Medicine. 2018;24:908–922. doi: 10.1038/s41591-018-0104-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Samuel V.T., Shulman G.I. Nonalcoholic fatty liver disease as a nexus of metabolic and hepatic diseases. Cell Metabolism. 2018;27:22–41. doi: 10.1016/j.cmet.2017.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Suzuki A., Diehl A.M. Nonalcoholic steatohepatitis. Annual Review of Medicine. 2017;68:85–98. doi: 10.1146/annurev-med-051215-031109. [DOI] [PubMed] [Google Scholar]
- 5.Ludwig J., Viggiano T.R., McGill D.B., Oh B.J. Nonalcoholic steatohepatitis: mayo Clinic experiences with a hitherto unnamed disease. Mayo Clinic Proceedings. 1980;55:434–438. [PubMed] [Google Scholar]
- 6.Younossi Z. Global burden of NAFLD and NASH: trends, predictions, risk factors and prevention. Nature Reviews Gastroenterology & Hepatology. 2018;15:11–20. doi: 10.1038/nrgastro.2017.109. [DOI] [PubMed] [Google Scholar]
- 7.Younossi Z. Global perspectives on non-alcoholic fatty liver disease and non-alcoholic steatohepatitis. Hepatology. 2018 [Google Scholar]
- 8.Pais R. NAFLD and liver transplantation: current burden and expected challenges. Journal of Hepatology. 2016;65:1245–1257. doi: 10.1016/j.jhep.2016.07.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li P. A liver-enriched long non-coding RNA, lncLSTR, regulates systemic lipid metabolism in mice. Cell Metabolism. 2015;21:455–467. doi: 10.1016/j.cmet.2015.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Szabo G., Csak T. Role of MicroRNAs in NAFLD/NASH. Digestive Diseases and Sciences. 2016;61:1314–1324. doi: 10.1007/s10620-015-4002-4. [DOI] [PubMed] [Google Scholar]
- 11.Zhao X.Y. Long noncoding RNA licensing of obesity-linked hepatic lipogenesis and NAFLD pathogenesis. Nature Communications. 2018;9:2986. doi: 10.1038/s41467-018-05383-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ramadori P., Weiskirchen R., Trebicka J., Streetz K. Mouse models of metabolic liver injury. Lab Animal. 2015;49:47–58. doi: 10.1177/0023677215570078. [DOI] [PubMed] [Google Scholar]
- 13.Clapper J.R. Diet-induced mouse model of fatty liver disease and nonalcoholic steatohepatitis reflecting clinical disease progression and methods of assessment. American Journal of Physiology - Gastrointestinal and Liver Physiology. 2013;305:G483–G495. doi: 10.1152/ajpgi.00079.2013. [DOI] [PubMed] [Google Scholar]
- 14.Guo L. Hepatic neuregulin 4 signaling defines an endocrine checkpoint for steatosis-to-NASH progression. Journal of Clinical Investigation. 2017 doi: 10.1172/JCI96324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kadowaki T. Adiponectin and adiponectin receptors in insulin resistance, diabetes, and the metabolic syndrome. Journal of Clinical Investigation. 2006;116:1784–1792. doi: 10.1172/JCI29126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Trujillo M.E., Scherer P.E. Adipose tissue-derived factors: impact on health and disease. Endocrine Reviews. 2006;27:762–778. doi: 10.1210/er.2006-0033. [DOI] [PubMed] [Google Scholar]
- 17.Waki H., Tontonoz P. Endocrine functions of adipose tissue. Annual Review of Pathology. 2007;2:31–56. doi: 10.1146/annurev.pathol.2.010506.091859. [DOI] [PubMed] [Google Scholar]
- 18.Chen Z. Nrg4 promotes fuel oxidation and a healthy adipokine profile to ameliorate diet-induced metabolic disorders. Molecular Metabolism. 2017;6:863–872. doi: 10.1016/j.molmet.2017.03.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang G.X., Zhao X.Y., Lin J.D. The brown fat secretome: metabolic functions beyond thermogenesis. Trends in Endocrinology and Metabolism. 2015;26:231–237. doi: 10.1016/j.tem.2015.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang G.X. The brown fat-enriched secreted factor Nrg4 preserves metabolic homeostasis through attenuation of hepatic lipogenesis. Nature Medicine. 2014;20:1436–1443. doi: 10.1038/nm.3713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Angelin B., Larsson T.E., Rudling M. Circulating fibroblast growth factors as metabolic regulators--a critical appraisal. Cell Metabolism. 2012;16:693–705. doi: 10.1016/j.cmet.2012.11.001. [DOI] [PubMed] [Google Scholar]
- 22.Potthoff M.J., Kliewer S.A., Mangelsdorf D.J. Endocrine fibroblast growth factors 15/19 and 21: from feast to famine. Genes & Development. 2012;26:312–324. doi: 10.1101/gad.184788.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pedersen B.K., Febbraio M.A. Muscles, exercise and obesity: skeletal muscle as a secretory organ. Nature Reviews Endocrinology. 2012;8:457–465. doi: 10.1038/nrendo.2012.49. [DOI] [PubMed] [Google Scholar]
- 24.Meex R.C.R., Watt M.J. Hepatokines: linking nonalcoholic fatty liver disease and insulin resistance. Nature Reviews Endocrinology. 2017;13:509–520. doi: 10.1038/nrendo.2017.56. [DOI] [PubMed] [Google Scholar]
- 25.Stefan N., Haring H.U. The role of hepatokines in metabolism. Nature Reviews Endocrinology. 2013;9:144–152. doi: 10.1038/nrendo.2012.258. [DOI] [PubMed] [Google Scholar]
- 26.Fisher F.M., Maratos-Flier E. Understanding the physiology of FGF21. Annual Review of Physiology. 2016;78:223–241. doi: 10.1146/annurev-physiol-021115-105339. [DOI] [PubMed] [Google Scholar]
- 27.Staiger H., Keuper M., Berti L., Hrabe de Angelis M., Haring H.U. Fibroblast growth factor 21-metabolic role in mice and men. Endocrine Reviews. 2017;38:468–488. doi: 10.1210/er.2017-00016. [DOI] [PubMed] [Google Scholar]
- 28.Wang Q. The hepatokine Tsukushi gates energy expenditure via brown fat sympathetic innervation. Nature Metabolism. 2019;1 doi: 10.1038/s42255-018-0020-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ito A. Tsukushi is required for anterior commissure formation in mouse brain. Biochemical and Biophysical Research Communications. 2010;402:813–818. doi: 10.1016/j.bbrc.2010.10.127. [DOI] [PubMed] [Google Scholar]
- 30.Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhao X. Quantitative proteomic analysis of optimal cutting temperature (OCT) embedded core-needle biopsy of lung cancer. Journal of the American Society for Mass Spectrometry. 2017;28:2078–2089. doi: 10.1007/s13361-017-1706-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Li S. Genome-wide coactivation analysis of PGC-1alpha identifies BAF60a as a regulator of hepatic lipid metabolism. Cell Metabolism. 2008;8:105–117. doi: 10.1016/j.cmet.2008.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kleiner D.E. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology. 2005;41:1313–1321. doi: 10.1002/hep.20701. [DOI] [PubMed] [Google Scholar]
- 34.Arendt B.M. Altered hepatic gene expression in nonalcoholic fatty liver disease is associated with lower hepatic n-3 and n-6 polyunsaturated fatty acids. Hepatology. 2015;61:1565–1578. doi: 10.1002/hep.27695. [DOI] [PubMed] [Google Scholar]
- 35.Wieser V. Adipose type I interferon signalling protects against metabolic dysfunction. Gut. 2018;67:157–165. doi: 10.1136/gutjnl-2016-313155. [DOI] [PubMed] [Google Scholar]
- 36.Bai J. DsbA-L prevents obesity-induced inflammation and insulin resistance by suppressing the mtDNA release-activated cGAS-cGAMP-STING pathway. Proceedings of the National Academy of Sciences of the United States of America. 2017;114:12196–12201. doi: 10.1073/pnas.1708744114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Luo X. Expression of STING is increased in liver tissues from patients with NAFLD and promotes macrophage-mediated hepatic inflammation and fibrosis in mice. Gastroenterology. 2018;155:1971–1984. doi: 10.1053/j.gastro.2018.09.010. e1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kharitonenkov A. FGF-21 as a novel metabolic regulator. Journal of Clinical Investigation. 2005;115:1627–1635. doi: 10.1172/JCI23606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Meex R.C. Fetuin B is a secreted hepatocyte factor linking steatosis to impaired glucose metabolism. Cell Metabolism. 2015;22:1078–1089. doi: 10.1016/j.cmet.2015.09.023. [DOI] [PubMed] [Google Scholar]
- 40.Zhao J. Hepatic F-box protein FBXW7 maintains glucose homeostasis through degradation of fetuin-A. Diabetes. 2018;67:818–830. doi: 10.2337/db17-1348. [DOI] [PubMed] [Google Scholar]
- 41.Ohta K. Tsukushi functions as an organizer inducer by inhibition of BMP activity in cooperation with chordin. Developmental Cell. 2004;7:347–358. doi: 10.1016/j.devcel.2004.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sahai A. Obese and diabetic db/db mice develop marked liver fibrosis in a model of nonalcoholic steatohepatitis: role of short-form leptin receptors and osteopontin. American Journal of Physiology - Gastrointestinal and Liver Physiology. 2004;287:G1035–G1043. doi: 10.1152/ajpgi.00199.2004. [DOI] [PubMed] [Google Scholar]
- 43.Diehl A.M., Day C. Cause. Pathogenesis, and treatment of nonalcoholic steatohepatitis. New England Journal of Medicine. 2017;377:2063–2072. doi: 10.1056/NEJMra1503519. [DOI] [PubMed] [Google Scholar]
- 44.Friedman S.L. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiological Reviews. 2008;88:125–172. doi: 10.1152/physrev.00013.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Krenkel O., Tacke F. Liver macrophages in tissue homeostasis and disease. Nature Reviews Immunology. 2017;17:306–321. doi: 10.1038/nri.2017.11. [DOI] [PubMed] [Google Scholar]
- 46.Poisson J. Liver sinusoidal endothelial cells: physiology and role in liver diseases. Journal of Hepatology. 2017;66:212–227. doi: 10.1016/j.jhep.2016.07.009. [DOI] [PubMed] [Google Scholar]
- 47.Robinson M.W., Harmon C., O'Farrelly C. Liver immunology and its role in inflammation and homeostasis. Cellular and Molecular Immunology. 2016;13:267–276. doi: 10.1038/cmi.2016.3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Tabibian J.H., Masyuk A.I., Masyuk T.V., O'Hara S.P., LaRusso N.F. Physiology of cholangiocytes. Comparative Physiology. 2013;3:541–565. doi: 10.1002/cphy.c120019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lefebvre P. Interspecies NASH disease activity whole-genome profiling identifies a fibrogenic role of PPARalpha-regulated dermatopontin. JCI Insight. 2017;2 doi: 10.1172/jci.insight.92264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Niimori D. Tsukushi controls the hair cycle by regulating TGF-beta1 signaling. Developmental Biology. 2012;372:81–87. doi: 10.1016/j.ydbio.2012.08.030. [DOI] [PubMed] [Google Scholar]
- 51.Ohta K. Tsukushi functions as a Wnt signaling inhibitor by competing with Wnt2b for binding to transmembrane protein Frizzled4. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:14962–14967. doi: 10.1073/pnas.1100513108. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
RNA-seq data files have been deposited into the Gene Expression Omnibus database (www.ncbi.nlm.nih.gov/geo/) with accession number GSE119340.