Abstract
Inhibitor of apoptosis (IAP) proteins are characterized by the presence of the conserved baculoviral IAP repeat (BIR) domain that is involved in protein-protein interactions. IAPs were initially thought to be mainly responsible for caspase inhibition, acting as negative regulators of apoptosis, but later works have shown that IAPs also control a plethora of other different cellular pathways. As X-linked IAP (XIAP), and other IAP, levels are often deregulated in cancer cells and have been shown to correlate with patients' prognosis, several approaches have been pursued to inhibit their activity in order to restore apoptosis. Many small molecules have been designed to target the BIR domains, the vast majority being inspired by the N-terminal tetrapeptide of Second Mitochondria-derived Activator of Caspases/Direct IAp Binding with Low pI (Smac/Diablo), which is the natural XIAP antagonist. These compounds are therefore usually referred to as Smac mimetics (SMs). Despite the fact that SMs were intended to specifically target XIAP, it has been shown that they also interact with cellular IAP-1 (cIAP1) and cIAP2, promoting their proteasome-dependent degradation. SMs have been tested in combination with several cytotoxic compounds and are now considered promising immune modulators which can be exploited in cancer therapy, especially in combination with immune checkpoint inhibitors. In this review, we give an overview of the structural hot-spots of BIRs, focusing on their fold and on the peculiar structural patches which characterize the diverse BIRs. These structures are exploited/exploitable for the development of specific and active IAP inhibitors.
Graphical ABSTRACT
1. Introduction
Resistance to cell death is considered a hallmark of cancer [1], and it represents a major issue in therapy by frustrating the efficacy of the cytotoxic compounds employed in cancer treatment. Therefore, there is a great interest in understanding the pathological determinants which protect cancer cells and prevent their capacity to undergo apoptosis or other mechanisms of cell death. To this end, many groups have focused their studies on a family of proteins referred to as inhibitor of apoptosis (IAP; [[2], [3], [4], [5]]) proteins which are often deregulated in cancer cells and whose levels have been shown to affect patients' prognosis [[6], [7], [8], [9], [10], [11], [12], [13]]. The distinctive trait which characterizes all IAPs is constituted by the presence of conserved domains called baculoviral IAP repeats (BIRs; [2,3,5]). In baculovirus, these domains are responsible for the interaction with caspases and their blockage [3,14], preventing cell suicide and allowing viral propagation. Due to the presence of BIR domains, IAPs were at first considered direct regulator of caspases [[15], [16], [17], [18], [19], [20], [21]] and they were thought to be mainly inhibitors of apoptosis (hence their name). Nevertheless, later studies have clearly shown that this view is extremely limited and that IAPs display several other functions [8,22]. Indeed, a direct interaction with executioner caspases, such as caspase-3, has been excluded, for at least 7 (neuronal apoptosis inhibitory protein, NAIP; cellular IAP-1, cIAP1; cIAP2; Survivin; Apollon; melanoma IAP, ML-IAP; and IAP-like protein-2, ILP-2) out of the 8 known IAPs. Of note, this does not imply that IAPs cannot regulate cell viability, though their pro-survival activity derives from the regulation of a plethora of different signaling pathways [8] and not from the direct inhibition of caspases.
In this regard, X-linked IAP (XIAP) represents the “black sheep” of the family [23], being the only member which interacts and inhibits both initiator and effector caspases [24]. XIAP, as NAIP, cIAP1 and cIAP2, sequence contains three BIR domains, each made of about 70 amino acids, whose fold is stabilized by a zinc atom that is coordinated by one histidine and three cysteine residues (belonging to the zinc-finger domain family). Based on the absence/presence of a deep peptide-binding, IAP binding motif (IBM), groove, the BIR domains can be grouped into type I and II. XIAP BIR1, lacking the IBM groove, belongs to type I BIR domain and it is the only XIAP BIR domain not involved in the regulation of caspases. XIAP BIR2 and BIR3 are type II BIR domains. BIR2, together with BIR1-BIR2 linker region, has been shown to interact with effector caspase-3 [21] and 7 [19]. BIR3 is known to inhibit caspase-9 activity [20] by binding its N-terminal tetrapeptide (ATPF), hence blocking the intrinsic apoptotic pathway [18].
XIAP represents a potential target in cancer treatment and several approaches have been pursued in order to inhibit its activity or to reduce its expression levels [[25], [26], [27], [28], [29], [30]]. For example, antisense oligonucleotide AEG35156 was tested both in pre-clinical settings [10,31] and in clinical trials [[32], [33], [34], [35]] to decrease the expression of XIAP and enhance the cytotoxic activity in combinatorial regiments [10]. By employing other strategies, small molecules designed to target the BIR2 domain were shown to promote the activity of caspase-3 [36], but BIR3 is indeed the most extensively studied domain for the development of BIR-targeted compounds. This is the case, for example, of Embelin, a molecule derived from the Japanese Ardisia herb, which was shown to inhibit cell growth and induce apoptosis in cancer cells expressing high levels of XIAP [37]. However, the main approach to target the BIR3 domain was stirred by XIAP natural antagonist Second Mitochondria-derived Activator of Caspases/Direct IAp Binding with Low pI (Smac/Diablo), which inspired the design and synthesis of a huge number of compounds [38], named Smac mimetics (SMs).
In this review, we summarize the structural properties of BIRs, focusing on their fold and on the crucial “hot-spots” exploited for the development of IAP-directed anti-tumor therapies, as SMs. These inhibitors, both peptidic and peptidomimetic, were designed to displace caspase-9 from BIR3 domain and promote the apoptotic machinery, but actually they resulted in unexpected outcomes and acted as pan-IAP inhibitors, eventually causing the degradation of cIAP1 and cIAP2, while inhibiting XIAP activity.
2. Mimicking the IAP Natural Antagonist Smac/Diablo
Smac/Diablo is a pro-apoptotic protein that usually localizes in the mitochondria membranes, being released in response to apoptotic stimuli [39]. Once in the cytosol, Smac/Diablo undergoes maturation with the cleavage of its N-terminal 55 amino acids. The loss of this sequence allows the exposure of the conserved IBM, present also in caspases and responsible for the interaction with the IAP BIRs [[40], [41], [42]] (Fig. 1). Through the IBM, Smac/Diablo prevents the activity of IAPs, causing the release of caspases bound with XIAP and the degradation of some members of the IAP family [43]. It has been shown that small peptides derived from the N-terminal region of mature Smac/Diablo can mimic the activity of the whole protein and display a pro-apoptotic function [44,45]. Later works identified the minimal sequence sufficient to interact with the BIR domains and demonstrated that the first four amino acids, AVPI, are sufficient to mimic the activity of the whole protein [36,46,47]. A number of modifications have been introduced on Smac/Diablo IBM sequence to generate peptidomimetics with increased affinity for the BIR domains and improved pharmacological properties [25,30,48].
Fig. 1.
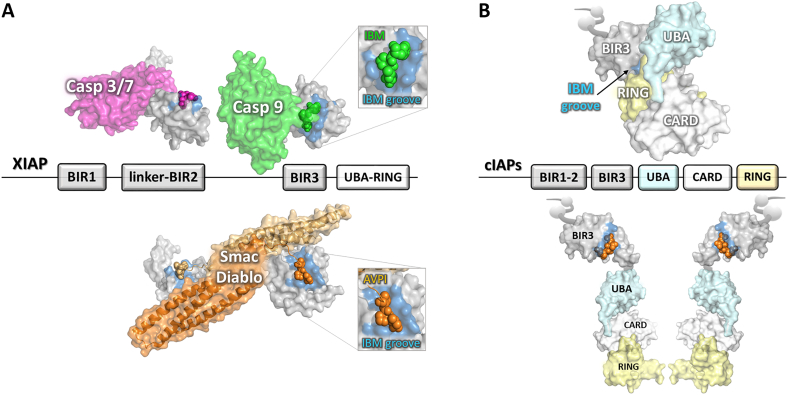
Smac/Diablo and SMs target homologous IAPs involved in different cellular pathways. IAPs are composed of three BIR domains (BIR1 to BIR3), an ubiquitin-associated (UBA) domain and a C-terminal RING domain. cIAPs additionally contain a CARD domain. (A) XIAP inhibits initiator (Caspase-9, green surface) and effector caspases (Caspase-3 and -7, magenta surface), through the interaction of its type II BIRs (BIR3 and BIR2, respectively), represented with grey boxes and surfaces. In particular, type II BIRs display a conserved IBM groove (blue region in the zoomed view), which hosts the N-terminal IAP-binding motif (IBM) reported in spheres. Smac/Diablo released from mitochondria upon stress stimuli, displaces caspases by exposing the N-terminal tetrapeptide AVPI (orange spheres), the base for SM design. (B) cIAPs are present in the cytosol as inactive monomers (panel B, on top). SM treatment induces dimerization and rapid auto-ubiquitination of cIAPs, leading to their degradation. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
3. Finding the Right Combination for IAP-Targeting Compounds
IAP-directed compounds were initially imagined as broad activators of apoptosis [36]. Accordingly, they have been tested in combination with a plethora of cytotoxic compounds [38,[49], [50], [51]]. Nonetheless, due to the crucial role of XIAP in protecting from the tumor necrosis factor (TNF) superfamily ligands, and in particular from the cytotoxic activity of TRAIL [[52], [53], [54], [55], [56]], SMs have been tested in combination with the latter molecule. The genetic and pharmacologic inhibition of XIAP has indeed been proved to sensitize cancer cells to TRAIL in preclinical models [57,58]. Later works have proved that XIAP-directed compounds have a dramatic effect also on other members of the IAP family, and in particular on cIAP1 and cIAP2 [[59], [60], [61]]. Upon stimulation with SMs, cIAPs undergo a conformational change which allows self-ubiquitination and rapid proteasome-dependent degradation [62,63]. The depletion of cIAP1 and cIAP2 causes the stabilization of NIK which is responsible for the activation of the non-canonical NF-κB pathway [62,64,65]. Of note, in about 10–15% of cancer cell lines, this event is sufficient to induce cell death due to NF-κB-dependent secretion of TNF which kills cells in an autocrine/paracrine fashion [59]. Moreover, since cIAP1 and cIAP2 have been described in a number of receptor complexes [4,66,67], it is not difficult to imagine that their depletion results in the activation and/or inhibition of different pathways. In particular, IAPs have been shown to modulate immunity-related receptors such as TLRs and NOD receptors [8,[68], [69], [70]] and protect from inflammation [71,72]. Accordingly, the treatment with SMs has been shown to display an immune modulatory activity [73,74], resulting in the massive secretion of pro-inflammatory cytokines with anti-tumor activity [75,76], but also inducing systemic toxicity [77]. Therefore, SMs have recently been proposed in combination with immune checkpoint inhibitors (ICIs; [73]), which are currently attracting a huge interest in cancer treatment [78], both in preclinical experiments [73] and in clinical trials [79]. Although the treatment with SMs in monotherapy resulted at most in partial responses in multiple myeloma [74] and glioblastoma [75] mouse models, notably, the combination with ICIs produced durable effects and was even curative in treated mice. These findings support the idea that SMs, even if not effective in cancer treatment as standalones, could be employed successfully in immune-based therapies.
4. Cancer Cell Mechanisms of Resistance to SM Treatment
As already mentioned, only a small percentage of cancer cell lines are sensitive to SMs in monotherapy [59] due to autocrine/paracrine expression of high levels of TNF [60,61] which results in apoptotic and necroptotic death [80]. A number of works focused on the mechanisms that could predict the cellular response to this class of compounds and explored strategies able to overcome cancer cell resistance. In this regard, the combination of TNF with SMs appears the most obvious way to increase their cytotoxic activity [47], but it is effective only in a small portion of cancer cell lines [81] and it is not practicable in the clinic.
Cancer cells have generally been shown to express higher levels of IAPs compared to normal cells [82], but this up-regulation is not necessarily responsible for reduced sensitivity to SMs. Accordingly, other mechanisms have been proposed. For example, CLL cells have been shown to be resistant to SMs due to their inability to form a ripoptosome [83], while other works suggest that, in the absence of cIAP1, the levels of cIAP2 massively increase [84] and this IAP is ultimately responsible for the resistance [81]. In these settings, inhibitors of phosphoinositide 3-kinase (PI3K) have been shown to prevent cIAP2 up-regulation and could therefore represent a strategy to enhance the cytotoxic activity of SMs. Biomarkers predictive of response have been proposed, especially in head and neck squamous cell carcinoma (HNSCC) [85,86], finding that the mediators of the extrinsic apoptotic pathway are essential for the efficacy of SMs, as shown also in a previous loss-of-function screening [87]. Unfortunately, the activity of SMs appears to be completely different in vivo [67], and, consequently, in vitro data may not predict their efficacy when administered systemically. Finally, mutations present in IAP genes [86] could affect the efficacy of SM treatment.
5. Structure and Function of BIRs: The BIR Fold
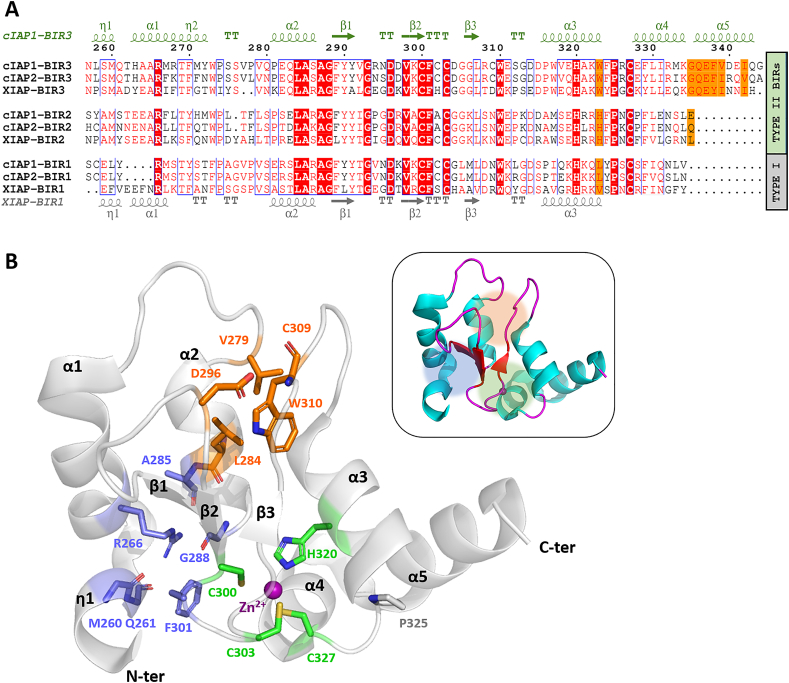
The BIR domain is composed of 70–80 amino acids (Fig. 2, A) building a central antiparallel β-sheet composed of 3 strands (β1–3) surrounded by 5 α-helices (mainly facing one side of the β-sheet). BIRs are Zn-finger domains similar to the C2H2-type [88]. The zinc binding section (Fig. 2,B, green residues and hotspot) is composed of a β-hairpin (β2-β3) followed by an α-helix (α3) forming together a left-handed bba-unit. The zinc atom is coordinated by a CCHC pattern (motif CFXC(X)16H(X)4PXC) by one histidine (in α3) and three cysteine residues (two at the end of β2 and the last between α3 and α4). Beyond the presence of the Zn binding residues, the fold is stabilized by the presence of two ‘hot-spots’ of conserved residues (residue numbered as in BIR3, Fig. 2): 1. R266-A285-G288-F301; 2. L284-D296-W310. In the first structural padlock, the N-terminal conserved residue R266 is hydrogen bonded to main chain atoms of M260, Q261 and (conserved) A285, and it is in van der Waals interaction with conserved G288 and F301. Such network of interactions stabilizes the N-terminal end of the first 2 (or 3) α-helices with the β1-turn-β2 (Fig. 2, B, light blue residues). The second ‘hot spot’ (Fig. 2, B, orange residues) is mainly due to hydrophobic interaction clustering together conserved L284, W310 and quasi-conserved V/L279 closed toward the solvent side by the hydrogen bond between conserved D296 and main chain nitrogen of amino acids 309 and 310.
Fig. 2.
The BIR domains: from the primary to the tertiary structure. (A) The sequence alignment of BIRs from cIAP1, cIAP2 and XIAP shows conserved motifs (highlighted in red when fully conserved, written in red when partially conserved), both among different BIRs of the same IAPs (Type I BIR1, compared to Type II BIR2 and BIR3) and among the same BIRs of different IAPs homologues (i.e. the BIR3 of cIAP1, cIAP2 and XIAP). The corresponding secondary structures are reported in green (cIAP1-BIR3 as a reference for Type II BIRs) and in grey (XIAP-BIR1 as reference for Type I BIRs). The alignment was performed with ESPRIPT (http://espript.ibcp.fr) [122]. (B) Structural “hot-spots” on BIRs stabilizing the BIR fold. Beyond the Zinc binding patch (residues in green, Zinc atom in purple), the second hot spot (residues in light blue) involves the fully conserved residues (highlighted in red in the sequences alignment) in the η1-α1-α2-β1-β2 elements. The third hot spot is stabilized by V279, L284, D296, C209 and W310 in orange sticks (Reference structure: cIAP1-BIR3, PDB id: 3MUP [123]). The secondary structure elements (cyan helices, red sheets and magenta loops) in the 3D structure are more clearly reported in the box, together with the location of the hot-spots. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
6. Structure and Function of BIRs: The IBM Groove
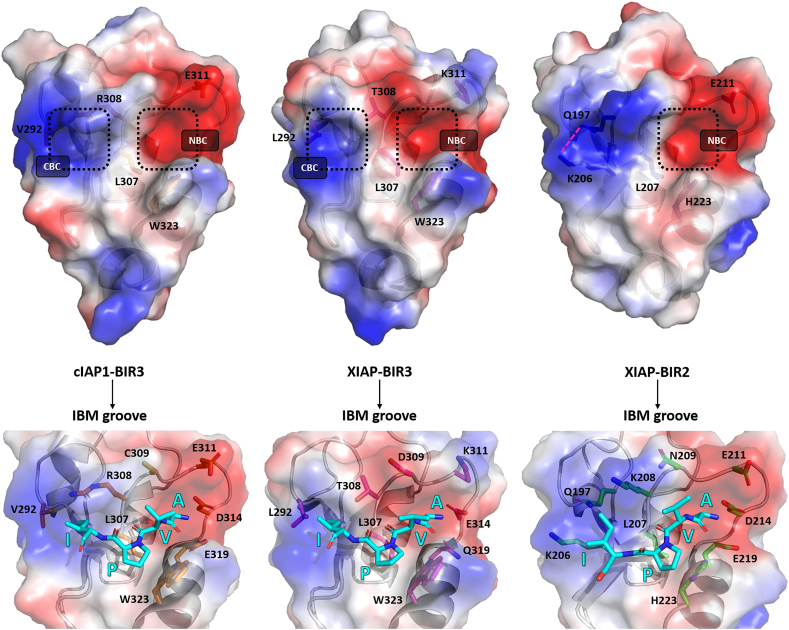
Type II BIRs are characterized by the presence of a surface cleft involved in the binding of IBM tetrapeptides: the IBM groove. It has been demonstrated that tetrapeptides and SMs bind to BIR3 in the conserved IBM cleft located between the β3 strand and the α3 helix of IAPs (i.e. cIAPs and XIAP, Fig. 3). The cleft is composed of two cavities: the N-terminal and the C-terminal binding cavities (NBC and CBC), named according to the portion of the tetrapeptide bound, and separated by a bulge (conserved L307 at the end of β3; Fig. 3). As expected, NBC is negatively charged while CBC has medium-low positive charge. The NBC is paved by the conserved Trp310 and is roughly lined by residues G306, R/T308, C/D309, E/K311, D/E314, E/Q319 and W323, in cIAP1/XIAP-BIR3, respectively (Fig. 3). The CBC corresponds to the N-terminal end of β1 closed by V/L292, in cIAP1/XIAP-BIR3, respectively.
Fig. 3.
Structural features of the IBM groove of type II BIRs hosting the tetrapeptide AVPI. Electrostatic surfaces (±70 keV) of cIAP1-BIR3 (PDB id: 3D9U), XIAP-BIR3 (PDB id: 1G73) and XIAP-BIR2 (PDB id: 4J46). The IBM groove runs from the negatively charged N-terminal binding cavity (NBC) to the positively charged C-terminal binding cavity (CBC), separated by the conserved Leucine 207/307 (in BIR2/BIR3, respectively). In XIAP-BIR2 the CBC is shallower than in BIR3 due to the interaction between Q197 and K206 (magenta hatches). On the bottom, a zoomed view of the IBM groove of each BIR domain, with the residues interacting with AVPI (in cyan sticks) highlighted in orange, magenta and green sticks, for cIAP1-BIR3, XIAP-BIR3 and XIAP-BIR2, respectively. Structures drawn with PyMOL (https://pymol.org/2/ The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC). (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
The overall features of the IBM groove are conserved in cIAP- and XIAP-BIR3, but subtle aminoacid substitutions can modulate the affinity for different SMs. In particular, in cIAP1-/XIAP-BIR3 the IBM cleft is more/less wide depending on quasi-conservative amino acids substitution: V/L292 at the end of β1 (CBC) and D/E314 (NBC). In addition, the negative charge on NBC is higher in cIAPs due to substitution E/Q319 and E/K311 in cIAPs/XIAP, respectively. The different features of the IBM cavity were exploited for the selection of SMs with different affinity for cIAP- and XIAP-BIR3, as reported in Corti et al. [89] and in Ndubaku et al. [90]. Such selective molecules can be used to study different effects in apoptosis modulations in several cell lines and develop cIAPs-targeted therapies with reduced side-effects (mainly due to XIAP inhibition). Interestingly, during optimization of IAP antagonists, Vamos and colleagues designed potent compounds which resulted selective for the unique BIR domain of ML-IAP [91]. Given the peculiar role of ML-IAP in tumorigenicity [92], such compounds could be useful to characterize novel different approaches adopted by IAPs to regulate cell death.
The IBM groove in BIR2 is different from that in BIR3 mostly for a shallow CBC due to the H-bond between Q197 and K206 that closes part of this cleft (Fig. 3). In the BIR3 domain, such residues are D/K297 and G/G306 in cIAP/XIAP, respectively. The crucial role of K206 in reducing BIR2 affinity for the Smac tetrapeptide AVPI was demonstrated by mutagenesis experiments [93]. The XIAP-BIR2 IBM groove has been shown to interact with a basic patch (R36 and R41 residues) of RIPK2 by Hrdinka and colleagues [94], thus being involved in NOD2 signaling. The role of different substituents in position 4 of AVPI tetrapeptide was extensively analyzed, in particular to enhance BIR2 vs BIR3 selectivity. As a result, bulky and flat chemical groups (like phenylhydrazine or 1-naphthyl moieties) appear to be better suited for the shallow CBC, enhancing both BIR2 affinity and selectivity [95]. In the last years, through a structure-based approach, novel XIAP-directed compounds were developed [96,97] by exploiting the increased selectivity for the BIR2 domain of XIAP compared to the BIR3 of XIAP and cIAP1. These compounds were shown to promote apoptosis without inducing cIAP1 degradation [98].
7. Type I BIR Domains: BIR1
Type I BIR domains do not inhibit caspases, but interact with adaptor proteins activating different survival pathways. In particular, XIAP-BIR1, in a dimeric form, recognizes TAB1 (TAK binding protein 1), a kinase activator, promoting the NF-κB survival pathway [99]. This likely occurs through a structural mechanism proposed by Lu and colleagues [100], where BIR1- and RING-mediated dimerization of XIAP allows the recognition and dimerization of TAB1 and subsequent dimerization and activation of TAK1 and NF-κB. cIAPs are recruited to the TNF receptor signaling complex through the interaction of cIAPs-BIR1 with elongated proteins named TRAFs (TNF receptor-associated factors). This binding event is essential for cIAPs activity, which consists of the ubiquitination of substrates involved in the canonical and non-canonical NF-κB pathways, where cIAPs positively or negatively regulate cell survival [99].
The BIR1 domain, which displays ~45% sequence similarity with BIR3, does not display the typical IBM groove, which is characteristic of the anti-apoptotic activity of type II BIRs. In fact, the residues K102/R82 in cIAP1/XIAP, respectively, display bulkier side chains than the corresponding residues in type II BIRs (E319/Q319 in cIAP1/XIAP BIR3, E219 in cIAPs/XIAP BIR2). Furthermore, the presence of L106/V86 in cIAPs/XIAP, respectively, weaken the stacking interactions that the IBM establishes with the corresponding residues in type II BIRs (W323 in cIAPs/XIAP BIR3, H223 in cIAPs/XIAP BIR2). Therefore, according to the BIR1 crystal structures [101], K102/R82 and L106/V86 would interfere with a BIR1–caspase and a BIR1–Smac putative interaction.
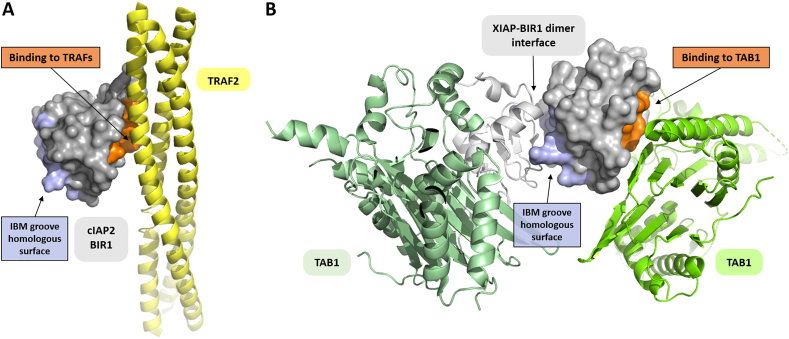
The structure of cIAP2-BIR1 in complex with TRAF2 (pdb-id 3M0A; [101]) shows two different interaction patches, but mutational studies identified the BIR1 N-terminal helix-turn-helix motif as the correct interaction surface. Such motif displays a quite shallow polar surface where the most important residues (based on mutational studies) are L30, M33, E47 and R48. Notably, these residues are located on the side of BIR1 opposite to the IBM-groove homologous surface (Fig. 4, panel A, in light blue). In the same work, Zheng and colleagues show that the surface of cIAP2-BIR1 involved in the binding to TRAF2 is the same exploited by XIAP-BIR1 for the interaction with TAB1. For this reason, such conserved surface could be an interesting target for drug design purposes [102].
Fig. 4.
The surfaces of type I BIRs mediating pro-survival macromolecular complexes. The chain D from 3M0A (cIAP2-BIR1 in complex with TRAF2) was superimposed to chain B from 2POP (XIAP-BIR1 in complex with TAB1), and therefore the two panels (A, B) display the BIR1 molecules in the same orientation. (A) The structure of cIAP2-BIR1 (grey surface) in complex with TRAF2 (yellow cartoon) reveals that the interaction of TRAF2 with the BIR1 domain occurs trough a set of aminoacids (orange area on BIR1) opposite to the surface homologous to the IBM groove (light blue area) of type II BIRs. (B) The orange patch of XIAP-BIR1 (grey surface) interacting with TAB1 (green cartoon) is the same observed for cIAP2-BIR1. The light blue area on XIAP-BIR1 is homologous to the IBM groove found on type II BIRs, but rather than binding to caspases or Smac/DIABLO, is involved in XIAP-BIR1 dimerization. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Both XIAP-BIR1 (PDB id: 2POI for XIAP-BIR1 alone and 2POP for XIAP-BIR1/TAB1 complex) and cIAP1-BIR1 (PDB id: 3M1D) crystal structures display a symmetrical dimeric assembly. The cIAP1-BIR1 dimer in the crystal counts only 4 stabilizing hydrogen bonds. The ΔiG P-value of cIAP1-BIR1 dimerization interface (0.318; http://www.ebi.ac.uk/pdbe/pisa/) indicates a stable interaction, but cIAPs-BIR1 were unable to dimerize even at high concentration (up to 60 mg/ml, [63]). In the crystal of XIAP BIR1, the dimer is stabilized by 20 hydrogen bonds and 18 salt bridges [100] and presents a ΔiG P-value of 0.905, indicating the poor hydrophobicity of the interface, often associated to crystal artifacts. Nevertheless, the XIAP-BIR1 dimer is stable also in solution and supposed to be biologically significant, since dimer-disruptive mutations resulted in a reduced ability to activate NF-κB [100]. The dimerization of XIAP-BIR1 was shown to be essential for XIAP-TAB1 interaction (Fig. 4, panel B), and is characterized by a strong network of electrostatic interactions. Such dimerization surface was targeted through in silico docking [102], identifying NF023 as an interesting moiety to be optimized for the impairment of XIAP-BIR1 dimerization and, thus, to modulate the NF-κB survival pathway.
8. Design of Compounds Specific for Different IAPs
As already mentioned, IAP-directed compounds were originally designed to specifically target XIAP [[28], [29], [30],103], but later works have shown that SMs cause the degradation of cIAP1/2 [60,104], and this is ultimately responsible for the killing of sensitive cancer cells, at least in vitro [59]. As BIRs are extremely conserved among the different IAPs, it is challenging to synthesize compounds indeed specific, but thanks to the increased knowledge of the structural properties of BIRs and the molecular basis for the interaction between SMs and the IBM groove, novel compounds have recently been described. In particular, efforts have been made to maintain the specificity for cIAP1, while reducing the inhibition of XIAP [89]. This would maintain the capacity to kill sensitive cancer cells through the activation of the non-canonical NF-κB pathway, and simultaneously would reduce the overall toxicity of SM administration [77,105]. With the latter aim, Birinapant has recently been described and is currently being tested in clinical trials [79]. This compound has been shown to display increased tolerability thanks to a decreased potency against cIAP2 and XIAP [38,105]. Of note, SM treatment rapidly deplete cIAP1, that is crucial for cIAP2 degradation [62,63], and simultaneously stimulate cIAP2 expression due to the activation of NF-κB. In this context, cIAP2 has been proposed as a possible mechanism of resistance to SM treatment [81] and therefore it could represent a potential target in combination therapies.
Finally, to increase the efficacy of SMs, hybrid compounds have been described. Indeed, thanks to the understanding of the residues that can be modified without losing affinity for the BIRs, monomeric SMs have been linked with direct activators of caspases [106], integrin inhibitors [107], nanoparticles [108] to enhance their cytotoxic activity and bioavailability. In other approaches, the ubiquitin ligase activity of IAPs has been exploited by designing proteolysis targeting chimeras (PROTAC) [109] which, by putting in direct contact IAPs with the target protein, allow the degradation of mutant huntingtin [110], BCR-ABL [111], estrogen receptor α (ERα) [112] and CRABP-II [113]. These works expand the opportunities to employ IAP-directed compounds in cancer treatment.
9. Summary and Outlook
BIR domains are conserved among IAPs and are present in multiple copies, being classified in type I and type II. Nevertheless, BIRs evolved different strategies to promote IAP pro-survival role. Although the BIR fold is a distinctive hallmark of all IAPs and it is stabilized by the presence of conserved residues, crucial amino acidic substitutions make the difference both between type I and type II BIRs, and within the same BIR type. Type I BIRs are involved in protein-protein interactions, which in turn regulate the NF-κB pathway. A deeper investigation of these mechanisms could shed new light on the relevance of type I-mediated protein-protein interaction processes in cancer, thus providing further hints in rational drug design. Type II BIRs display a functional IBM groove for the direct binding of peptides, which provided the basis for SM design. Yet, homologous IBM grooves are not identical, and selective SMs have been developed in order to study the distinct processes and to propose tailored SM-based therapies.
Despite the great expectation around IAP-directed compounds, the clinical efficacy of SMs is still limited by the lacking of predictive markers of response. Moreover, these compounds have been employed for the treatment of different types of tumors, without individuating those that could majorly benefit from this kind of treatment [79]. The insufficient efficacy of IAP-directed therapy is likely due also to the fact that IAPs are not merely inhibitor of apoptosis as initially thought [38,114] and therefore the final outcome of SM treatment is the result of a complicated network of different effects. Notably, XIAP [115], cIAP1 [116] and cIAP2 [117] knockout mice have only mild phenotypes, and this is not compatible with their expected pivotal role in regulating apoptosis, both during development and in normal cell homeostasis, though it could also be the effect of their partially redundant activities. An attempt to clarify the relevance of cIAP1, cIAP2 and XIAP was made by employing double knockout mice. It was shown that cIAP1/cIAP2 and XIAP/cIAP1 double knockouts were embryonic lethal, but XIAP/cIAP2 double knockout mice were viable and fertile [118]. This evidence allowed to hypothesize that cIAP1 could play a pivotal role in mouse development, adult physiology and NF-kB pathway regulation. In contrast with these findings, a later work showed that XIAP/cIAP1 double knockout mice are viable [84], therefore clarifying that a deeper understating of the role of these IAPs in organism development and NF-kB regulation is still necessary. Nonetheless, the development of SMs importantly contributed to a significant advance in the field, and allowed the characterization of IAP roles and to investigate several molecular mechanisms of cell death, as apoptosis itself and necroptosis [77,[119], [120], [121]]. Moreover, SMs now represent a fundamental tool to investigate several signaling pathways such as MAPKs and NF-κB [64,67].
The expectation is that a further understanding of the IAP activity will soon provide the rationale for novel combinatorial therapies. In this sense, a promising combination is constituted by immune checkpoint inhibitors which may benefit from the transient pro-inflammatory environment triggered by SMs. Therefore, the employment of SMs in cancer treatment could be successful when combined with immune-based approaches.
Acknowledgments
Acknowledgements
Daniele Lecis is supported by the Italian Ministry of Health, Italy (5 × 1000 Funds - 2013). Eloise Mastrangelo is thankful to Italian Association for the Cancer Research (AIRC) for the financial support (“My First AIRC Grant (MFAG)” project code: 17083).
Declarations of interest
None.
Contributor Information
Eloise Mastrangelo, Email: eloise.mastrangelo@unimi.it.
Daniele Lecis, Email: daniele.lecis@istitutotumori.mi.it.
References
- 1.Hanahan D., Weinberg R.A. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674. doi: 10.1016/j.cell.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 2.Crook N.E., Clem R.J., Miller L.K. An apoptosis-inhibiting baculovirus gene with a zinc finger-like motif. JVirol. 1993;67:2168–2174. doi: 10.1128/jvi.67.4.2168-2174.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Birnbaum M.J., Clem R.J., Miller L.K. An apoptosis-inhibiting gene from a nuclear polyhedrosis virus encoding a polypeptide with Cys/His sequence motifs. JVirol. 1994;68:2521–2528. doi: 10.1128/jvi.68.4.2521-2528.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rothe M., Pan M.G., Henzel W.J., Ayres T.M., Goeddel D.V. The TNFR2-TRAF signaling complex contains two novel proteins related to baculoviral inhibitor of apoptosis proteins. Cell. 1995;83:1243–1252. doi: 10.1016/0092-8674(95)90149-3. [DOI] [PubMed] [Google Scholar]
- 5.Duckett C.S., Nava V.E., Gedrich R.W., Clem R.J., Van Dongen J.L. A conserved family of cellular genes related to the baculovirus iap gene and encoding apoptosis inhibitors. EMBO J. 1996;15:2685–2694. [PMC free article] [PubMed] [Google Scholar]
- 6.Tamm I., Kornblau S.M., Segall H., Krajewski S., Welsh K. Expression and prognostic significance of IAP-family genes in human cancers and myeloid leukemias. ClinCancer Res. 2000;6:1796–1803. [PubMed] [Google Scholar]
- 7.Troeger A., Siepermann M., Escherich G., Meisel R., Willers R. Survivin and its prognostic significance in pediatric acute B-cell precursor lymphoblastic leukemia. Haematologica. 2007;92:1043–1050. doi: 10.3324/haematol.10675. [DOI] [PubMed] [Google Scholar]
- 8.Gyrd-Hansen M., Meier P. IAPs: from caspase inhibitors to modulators of NF-kappaB, inflammation and cancer. NatRevCancer. 2010;10:561–574. doi: 10.1038/nrc2889. [DOI] [PubMed] [Google Scholar]
- 9.Hundsdoerfer P., Dietrich I., Schmelz K., Eckert C., Henze G. XIAP expression is post-transcriptionally upregulated in childhood ALL and is associated with glucocorticoid response in T-cell ALL. PediatrBlood Cancer. 2010;55:260–266. doi: 10.1002/pbc.22541. [DOI] [PubMed] [Google Scholar]
- 10.Miura K., Fujibuchi W., Ishida K., Naitoh T., Ogawa H. Inhibitor of apoptosis protein family as diagnostic markers and therapeutic targets of colorectal cancer. SurgToday. 2011;41:175–182. doi: 10.1007/s00595-010-4390-1. [DOI] [PubMed] [Google Scholar]
- 11.Che X., Yang D., Zong H., Wang J., Li X. Nuclear cIAP1 overexpression is a tumor stage- and grade-independent predictor of poor prognosis in human bladder cancer patients. UrolOncol. 2012;30:450–456. doi: 10.1016/j.urolonc.2010.12.016. [DOI] [PubMed] [Google Scholar]
- 12.Yang X.H., Feng Z.E., Yan M., Hanada S., Zuo H. XIAP is a predictor of cisplatin-based chemotherapy response and prognosis for patients with advanced head and neck cancer. PLoS One. 2012;7 doi: 10.1371/journal.pone.0031601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ibrahim A.M., Mansour I.M., Wilson M.M., Mokhtar D.A., Helal A.M. Study of survivin and X-linked inhibitor of apoptosis protein (XIAP) genes in acute myeloid leukemia (AML) LabHematol. 2012;18:1–10. doi: 10.1532/LH96.11005. [DOI] [PubMed] [Google Scholar]
- 14.Zhou Q., Krebs J.F., Snipas S.J., Price A., Alnemri E.S. Interaction of the baculovirus anti-apoptotic protein p35 with caspases. Specificity, kinetics, and characterization of the caspase/p35 complex. Biochemistry. 1998;37:10757–10765. doi: 10.1021/bi980893w. [DOI] [PubMed] [Google Scholar]
- 15.You M., Ku P.T., Hrdlickova R., Bose H.R., Jr. ch-IAP1, a member of the inhibitor-of-apoptosis protein family, is a mediator of the antiapoptotic activity of the v-Rel oncoprotein. Mol.Cell. Biologicals. 1997;17:7328–7341. doi: 10.1128/mcb.17.12.7328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Takahashi R., Deveraux Q., Tamm I., Welsh K., Assa-Munt N. A Single BIR Domain of XIAP Sufficient for Inhibiting Caspases. JBiolChem. 1998;273:7787–7790. doi: 10.1074/jbc.273.14.7787. [DOI] [PubMed] [Google Scholar]
- 17.Deveraux Q.L., Roy N., Stennicke H.R., Van Arsdale T., Zhou Q. IAPs block apoptotic events induced by caspase-8 and cytochrome c by direct inhibition of distinct caspases. EMBO J. 1998;17:2215–2223. doi: 10.1093/emboj/17.8.2215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Datta R., Oki E., Endo K., Biedermann V., Ren J. XIAP regulates DNA damage-induced apoptosis downstream of caspase-9 cleavage. JBiolChem. 2000;275:31733–31738. doi: 10.1074/jbc.M910231199. [DOI] [PubMed] [Google Scholar]
- 19.Chai J., Shiozaki E., Srinivasula S.M., Wu Q., Dataa P. vol. 104. 2001. Structural Basis of Caspase-7 Inhibition by XIAP; pp. 769–780. [DOI] [PubMed] [Google Scholar]
- 20.Srinivasula S.M., Hegde R., Saleh A., Datta P., Shiozaki E. A conserved XIAP-interaction motif in caspase-9 and Smac/DIABLO regulates caspase activity and apoptosis. Nature. 2001;410:112–116. doi: 10.1038/35065125. [DOI] [PubMed] [Google Scholar]
- 21.Riedl S.J., Renatus M., Schwarzenbacher R., Zhou Q., Sun C. Structural Basis for the Inhibition of Caspase-3 by XIAP. Cell. 2001;104:791–800. doi: 10.1016/s0092-8674(01)00274-4. [DOI] [PubMed] [Google Scholar]
- 22.Damgaard R.B., Gyrd-Hansen M. Inhibitor of apoptosis (IAP) proteins in regulation of inflammation and innate immunity. DiscovMed. 2011;11:221–231. [PubMed] [Google Scholar]
- 23.Eckelman B.P., Salvesen G.S., Scott F.L. Human inhibitor of apoptosis proteins: why XIAP is the black sheep of the family. EMBO Rep. 2006;7:988–994. doi: 10.1038/sj.embor.7400795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Obexer P., Ausserlechner M.J. X-linked inhibitor of apoptosis protein - a critical death resistance regulator and therapeutic target for personalized cancer therapy. FrontOncol. 2014;4:197. doi: 10.3389/fonc.2014.00197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Seneci P., Bianchi A., Battaglia C., Belvisi L., Bolognesi M. Rational design, synthesis and characterization of potent, non-peptidic Smac mimics/XIAP inhibitors as proapoptotic agents for cancer therapy. Bioorg Med Chem. 2009;17:5834–5856. doi: 10.1016/j.bmc.2009.07.009. [DOI] [PubMed] [Google Scholar]
- 26.Cossu F., Mastrangelo E., Milani M., Sorrentino G., Lecis D. Designing Smac-mimetics as antagonists of XIAP, cIAP1, and cIAP2. Biochem Biophys Res Commun. 2009;378:162–167. doi: 10.1016/j.bbrc.2008.10.139. [DOI] [PubMed] [Google Scholar]
- 27.Nikolovska-Coleska Z., Meagher J.L., Jiang S., Kawamoto S.A., Gao W. Design and characterization of bivalent Smac-based peptides as antagonists of XIAP and development and validation of a fluorescence polarization assay for XIAP containing both BIR2 and BIR3 domains. Anal Biochem. 2008;374:87–98. doi: 10.1016/j.ab.2007.10.032. [DOI] [PubMed] [Google Scholar]
- 28.Sun H., Stuckey J.A., Nikolovska-Coleska Z., Qin D., Meagher J.L. Structure-based design, synthesis, evaluation, and crystallographic studies of conformationally constrained Smac mimetics as inhibitors of the X-linked inhibitor of apoptosis protein (XIAP) JMedChem. 2008;51:7169–7180. doi: 10.1021/jm8006849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mastrangelo E., Cossu F., Milani M., Sorrentino G., Lecis D. Targeting the X-linked inhibitor of apoptosis protein through 4-substituted azabicyclo[5.3.0]alkane smac mimetics. Structure, activity, and recognition principles. J Mol Biol. 2008;384:673–689. doi: 10.1016/j.jmb.2008.09.064. [DOI] [PubMed] [Google Scholar]
- 30.Sun H., Nikolovska-Coleska Z., Lu J., Meagher J.L., Yang C.Y. Design, synthesis, and characterization of a potent, nonpeptide, cell-permeable, bivalent Smac mimetic that concurrently targets both the BIR2 and BIR3 domains in XIAP. JAmChemSoc. 2007;129:15279–15294. doi: 10.1021/ja074725f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.LaCasse E.C., Cherton-Horvat G.G., Hewitt K.E., Jerome L.J., Morris S.J. Preclinical characterization of AEG35156/GEM 640, a second-generation antisense oligonucleotide targeting X-linked inhibitor of apoptosis. Clin Cancer Res. 2006;12:5231–5241. doi: 10.1158/1078-0432.CCR-06-0608. [DOI] [PubMed] [Google Scholar]
- 32.Lee F.A., Zee B.C., Cheung F.Y., Kwong P., Chiang C.L. Randomized phase II study of the X-linked inhibitor of apoptosis (XIAP) antisense AEG35156 in combination with sorafenib in patients with advanced hepatocellular carcinoma (HCC) AmJClinOncol. 2016;39:609–613. doi: 10.1097/COC.0000000000000099. [DOI] [PubMed] [Google Scholar]
- 33.Schimmer A.D., Estey E.H., Borthakur G., Carter B.Z., Schiller G.J. Phase I/II trial of AEG35156 X-linked inhibitor of apoptosis protein antisense oligonucleotide combined with idarubicin and cytarabine in patients with relapsed or primary refractory acute myeloid leukemia. J Clin Oncol. 2009;27:4741–4746. doi: 10.1200/JCO.2009.21.8172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mahadevan D., Chalasani P., Rensvold D., Kurtin S., Pretzinger C. Phase I trial of AEG35156 an antisense oligonucleotide to XIAP plus gemcitabine in patients with metastatic pancreatic ductal adenocarcinoma. Am J Clin Oncol. 2013;36:239–243. doi: 10.1097/COC.0b013e3182467a13. [DOI] [PubMed] [Google Scholar]
- 35.Schimmer A.D., Herr W., Hanel M., Borthakur G., Frankel A. Addition of AEG35156 XIAP antisense oligonucleotide in reinduction chemotherapy does not improve remission rates in patients with primary refractory acute myeloid leukemia in a randomized phase II study. Clin Lymphoma Myeloma Leuk. 2011;11:433–438. doi: 10.1016/j.clml.2011.03.033. [DOI] [PubMed] [Google Scholar]
- 36.Schimmer A.D., Welsh K., Pinilla C., Wang Z., Krajewska M. Small-molecule antagonists of apoptosis suppressor XIAP exhibit broad antitumor activity. Cancer Cell. 2004;5:25–35. doi: 10.1016/s1535-6108(03)00332-5. [DOI] [PubMed] [Google Scholar]
- 37.Nikolovska-Coleska Z., Xu L., Hu Z., Tomita Y., Li P. Discovery of embelin as a cell-permeable, small-molecular weight inhibitor of XIAP through structure-based computational screening of a traditional herbal medicine three-dimensional structure database. J Med Chem. 2004;47:2430–2440. doi: 10.1021/jm030420+. [DOI] [PubMed] [Google Scholar]
- 38.Fulda S. Promises and challenges of Smac mimetics as cancer therapeutics. Clin Cancer Res. 2015;21:5030–5036. doi: 10.1158/1078-0432.CCR-15-0365. [DOI] [PubMed] [Google Scholar]
- 39.Verhagen A.M., Ekert P.G., Pakusch M., Silke J., Connolly L.M. vol. 102. 2000. Identification of DIABLO, a Mammalian Protein that Promotes Apoptosis by Binding to and Antagonizing IAP Proteins; pp. 43–53. [DOI] [PubMed] [Google Scholar]
- 40.Liu Z., Sun C., Olejniczak E.T., Meadows R.P., Betz S.F. Structural basis for binding of Smac/DIABLO to the XIAP BIR3 domain. Nature. 2000;408:1004–1008. doi: 10.1038/35050006. [DOI] [PubMed] [Google Scholar]
- 41.Wu G., Chai J., Suber T.L., Wu J., Du C. Structural basis of IAP recognition by Smac/DIABLO. Nature. 2000;408:1008–1012. doi: 10.1038/35050012. [DOI] [PubMed] [Google Scholar]
- 42.Mastrangelo E., Vachette P., Cossu F., Malvezzi F., Bolognesi M. The activator of apoptosis Smac-DIABLO acts as a tetramer in solution. Biophys J. 2015;108:714–723. doi: 10.1016/j.bpj.2014.11.3471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yang Q.H., Du C. Smac/DIABLO selectively reduces the levels of c-IAP1 and c-IAP2 but not that of XIAP and livin in HeLa cells. J Biol Chem. 2004;279:16963–16970. doi: 10.1074/jbc.M401253200. [DOI] [PubMed] [Google Scholar]
- 44.Fulda S., Wick W., Weller M., Debatin K.M. Smac agonists sensitize for Apo2L/TRAIL- or anticancer drug-induced apoptosis and induce regression of malignant glioma in vivo. Nat Med. 2002;8:808–815. doi: 10.1038/nm735. [DOI] [PubMed] [Google Scholar]
- 45.Arnt C.R., Chiorean M.V., Heldebrant M.P., Gores G.J., Kaufmann S.H. Synthetic Smac/DIABLO peptides enhance the effects of chemotherapeutic agents by binding XIAP and cIAP1 in situ. J Biol Chem. 2002;277:44236–44243. doi: 10.1074/jbc.M207578200. [DOI] [PubMed] [Google Scholar]
- 46.Nikolovska-Coleska Z., Wang R., Fang X., Pan H., Tomita Y. Development and optimization of a binding assay for the XIAP BIR3 domain using fluorescence polarization. Anal Biochem. 2004;332:261–273. doi: 10.1016/j.ab.2004.05.055. [DOI] [PubMed] [Google Scholar]
- 47.Li L., Thomas R.M., Suzuki H., De Brabander J.K., Wang X. A Small Molecule Smac Mimic Potentiates TRAIL- and TNF{alpha}-Mediated Cell Death. Science. 2004;305:1471–1474. doi: 10.1126/science.1098231. [DOI] [PubMed] [Google Scholar]
- 48.Lecis D., Mastrangelo E., Belvisi L., Bolognesi M., Civera M. Dimeric Smac mimetics/IAP inhibitors as in vivo-active pro-apoptotic agents. Part II: Structural and biological characterization. Bioorg.Med. Chembiochem. 2012;20:6709–6723. doi: 10.1016/j.bmc.2012.09.041. [DOI] [PubMed] [Google Scholar]
- 49.Servida F., Lecis D., Scavullo C., Drago C., Seneci P. Novel second mitochondria-derived activator of caspases (Smac) mimetic compounds sensitize human leukemic cell lines to conventional chemotherapeutic drug-induced and death receptor-mediated apoptosis. Investig New Drugs. 2011;29:1264–1275. doi: 10.1007/s10637-010-9475-6. [DOI] [PubMed] [Google Scholar]
- 50.Wagner L., Marschall V., Karl S., Cristofanon S., Zobel K. Smac mimetic sensitizes glioblastoma cells to Temozolomide-induced apoptosis in a RIP1- and NF-kappaB-dependent manner. Oncogene. 2013;32:988–997. doi: 10.1038/onc.2012.108. [DOI] [PubMed] [Google Scholar]
- 51.Zhou B., Zhang J., Chen G., You L., Zhang T.P. Therapy of Smac mimetic SM-164 in combination with gemcitabine for pancreatic cancer. Cancer Lett. 2013;329:118–124. doi: 10.1016/j.canlet.2012.10.039. [DOI] [PubMed] [Google Scholar]
- 52.McManus D.C., Lefebvre C.A., Cherton-Horvat G., St-Jean M., Kandimalla E.R. Loss of XIAP protein expression by RNAi and antisense approaches sensitizes cancer cells to functionally diverse chemotherapeutics. Oncogene. 2004;23:8105–8117. doi: 10.1038/sj.onc.1207967. [DOI] [PubMed] [Google Scholar]
- 53.Braeuer S.J., Buneker C., Mohr A., Zwacka R.M. Constitutively activated nuclear factor-kappaB, but not induced NF-kappaB, leads to TRAIL resistance by up-regulation of X-linked inhibitor of apoptosis protein in human cancer cells. Mol Cancer Res. 2006;4:715–728. doi: 10.1158/1541-7786.MCR-05-0231. [DOI] [PubMed] [Google Scholar]
- 54.Vogler M., Walczak H., Stadel D., Haas T.L., Genze F. Targeting XIAP bypasses Bcl-2-mediated resistance to TRAIL and cooperates with TRAIL to suppress pancreatic cancer growth in vitro and in vivo. Cancer Res. 2008;68:7956–7965. doi: 10.1158/0008-5472.CAN-08-1296. [DOI] [PubMed] [Google Scholar]
- 55.Vogler M., Walczak H., Stadel D., Haas T.L., Genze F. Small molecule XIAP inhibitors enhance TRAIL-induced apoptosis and antitumor activity in preclinical models of pancreatic carcinoma. Cancer Res. 2009;69:2425–2434. doi: 10.1158/0008-5472.CAN-08-2436. [DOI] [PubMed] [Google Scholar]
- 56.Lecis D., Drago C., Manzoni L., Seneci P., Scolastico C. Novel SMAC-mimetics synergistically stimulate melanoma cell death in combination with TRAIL and Bortezomib. BrJCancer. 2010;102:1707–1716. doi: 10.1038/sj.bjc.6605687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gillissen B., Richter A., Richter A., Overkamp T., Essmann F. Targeted therapy of the XIAP/proteasome pathway overcomes TRAIL-resistance in carcinoma by switching apoptosis signaling to a Bax/Bak-independent 'type I' mode. Cell Death Dis. 2013;4 doi: 10.1038/cddis.2013.67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Allensworth J.L., Sauer S.J., Lyerly H.K., Morse M.A., Devi G.R. Smac mimetic Birinapant induces apoptosis and enhances TRAIL potency in inflammatory breast cancer cells in an IAP-dependent and TNF-alpha-independent mechanism. Breast Cancer ResTreat. 2013;137:359–371. doi: 10.1007/s10549-012-2352-6. [DOI] [PubMed] [Google Scholar]
- 59.Petersen S.L., Wang L., Yalcin-Chin A., Li L., Peyton M. Autocrine TNFα signaling renders human cancer cells susceptible to smac-mimetic-induced apoptosis. Cancer Cell. 2007;12:445–456. doi: 10.1016/j.ccr.2007.08.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Vince J.E., Wong W.W., Khan N., Feltham R., Chau D. IAP antagonists target cIAP1 to induce TNFα-dependent apoptosis. Cell. 2007;131:682–693. doi: 10.1016/j.cell.2007.10.037. [DOI] [PubMed] [Google Scholar]
- 61.Varfolomeev E., Blankenship J.W., Wayson S.M., Fedorova A.V., Kayagaki N. IAP antagonists induce autoubiquitination of c-IAPs, NF-κB activation, and TNFα-dependent apoptosis. Cell. 2007;131:669–681. doi: 10.1016/j.cell.2007.10.030. [DOI] [PubMed] [Google Scholar]
- 62.Darding M., Feltham R., Tenev T., Bianchi K., Benetatos C. Molecular determinants of Smac mimetic induced degradation of cIAP1 and cIAP2. Cell Death Differ. 2011;18:1376–1386. doi: 10.1038/cdd.2011.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Feltham R., Bettjeman B., Budhidarmo R., Mace P.D., Shirley S. SMAC-mimetics activate the E3 ligase activity of cIAP1 by promoting RING dimerisation. JBiolChem. 2011;286:17015–17028. doi: 10.1074/jbc.M111.222919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Varfolomeev E., Goncharov T., Maecker H., Zobel K., Komuves L.G. Cellular inhibitors of apoptosis are global regulators of NF-kappaB and MAPK activation by members of the TNF family of receptors. SciSignal. 2012;5:ra22. doi: 10.1126/scisignal.2001878. [DOI] [PubMed] [Google Scholar]
- 65.Tchoghandjian A., Jennewein C., Eckhardt I., Rajalingam K., Fulda S. Identification of non-canonical NF-kappaB signaling as a critical mediator of Smac mimetic-stimulated migration and invasion of glioblastoma cells. Cell Death Dis. 2013;4 doi: 10.1038/cddis.2013.70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Li X., Yang Y., Ashwell J.D. TNF-RII and c-IAP1 mediate ubiquitination and degradation of TRAF2. Nature. 2002;416:345–347. doi: 10.1038/416345a. [DOI] [PubMed] [Google Scholar]
- 67.Majorini M.T., Manenti G., Mano M., De Cecco L., Conti A. cIAP1 regulates the EGFR/Snai2 axis in triple-negative breast cancer cells. Cell Death Differ. 2018;25:2147–2164. doi: 10.1038/s41418-018-0100-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Bertrand M.J., Doiron K., Labbe K., Korneluk R.G., Barker P.A. Cellular inhibitors of apoptosis cIAP1 and cIAP2 are required for innate immunity signaling by the pattern recognition receptors NOD1 and NOD2. Immunity. 2009;30:789–801. doi: 10.1016/j.immuni.2009.04.011. [DOI] [PubMed] [Google Scholar]
- 69.Estornes Y., Bertrand M.J. IAPs, regulators of innate immunity and inflammation. Semin Cell Dev Biol. 2015;39:106–114. doi: 10.1016/j.semcdb.2014.03.035. [DOI] [PubMed] [Google Scholar]
- 70.Vandenabeele P., Bertrand M.J. The role of the IAP E3 ubiquitin ligases in regulating pattern-recognition receptor signalling. Nat Rev Immunol. 2012;12:833–844. doi: 10.1038/nri3325. [DOI] [PubMed] [Google Scholar]
- 71.Walczak H. TNF and ubiquitin at the crossroads of gene activation, cell death, inflammation, and cancer. Immunol Rev. 2011;244:9–28. doi: 10.1111/j.1600-065X.2011.01066.x. [DOI] [PubMed] [Google Scholar]
- 72.Gerlach B., Cordier S.M., Schmukle A.C., Emmerich C.H., Rieser E. Linear ubiquitination prevents inflammation and regulates immune signalling. Nature. 2011;471:591–596. doi: 10.1038/nature09816. [DOI] [PubMed] [Google Scholar]
- 73.Beug S.T., Beauregard C.E., Healy C., Sanda T., St-Jean M. Smac mimetics synergize with immune checkpoint inhibitors to promote tumour immunity against glioblastoma. NatCommun. 2017;8 doi: 10.1038/ncomms14278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Chesi M., Mirza N.N., Garbitt V.M., Sharik M.E., Dueck A.C. IAP antagonists induce anti-tumor immunity in multiple myeloma. NatMed. 2016;22:1411–1420. doi: 10.1038/nm.4229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Beug S.T., Tang V.A., Lacasse E.C., Cheung H.H., Beauregard C.E. Smac mimetics and innate immune stimuli synergize to promote tumor death. Nat Biotechnol. 2014;32:182–190. doi: 10.1038/nbt.2806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lecis D., De Cesare M., Perego P., Conti A., Corna E. Smac mimetics induce inflammation and necrotic tumour cell death by modulating macrophage activity. CellDeath Dis. 2013;4 doi: 10.1038/cddis.2013.449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Wong W.W., Vince J.E., Lalaoui N., Lawlor K.E., Chau D. cIAPs and XIAP regulate myelopoiesis through cytokine production in a RIPK1 and RIPK3 dependent manner. Blood. 2014;123:2562–2572. doi: 10.1182/blood-2013-06-510743. [DOI] [PubMed] [Google Scholar]
- 78.Imbimbo M., Lo Russo G., Blackhall F. Current status of immunotherapy for non-small-cell lung cancer. Tumori. 2016;102:337–351. doi: 10.5301/tj.5000537. [DOI] [PubMed] [Google Scholar]
- 79.Finlay D., Teriete P., Vamos M., NDP Cosford, Vuori K. Inducing death in tumor cells: roles of the inhibitor of apoptosis proteins. F1000Res. 2017;6:587. doi: 10.12688/f1000research.10625.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Muller-Sienerth N., Dietz L., Holtz P., Kapp M., Grigoleit G.U. SMAC mimetic BV6 induces cell death in monocytes and maturation of monocyte-derived dendritic cells. PLoS One. 2011;6 doi: 10.1371/journal.pone.0021556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Petersen S.L., Peyton M., Minna J.D., Wang X. Overcoming cancer cell resistance to Smac mimetic induced apoptosis by modulating cIAP-2 expression. Proc Natl Acad Sci. 2010;107:11936–11941. doi: 10.1073/pnas.1005667107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Rathore R., McCallum J.E., Varghese E., Florea A.M., Busselberg D. Overcoming chemotherapy drug resistance by targeting inhibitors of apoptosis proteins (IAPs) Apoptosis. 2017;22:898–919. doi: 10.1007/s10495-017-1375-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Maas C., Tromp J.M., van Laar J., Thijssen R., Elias J.A. CLL cells are resistant to smac mimetics because of an inability to form a ripoptosome complex. Cell Death Dis. 2013;4 doi: 10.1038/cddis.2013.305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Heard K.N., Bertrand M.J., Barker P.A. cIAP2 supports viability of mice lacking cIAP1 and XIAP. EMBO J. 2015;34:2393–2395. doi: 10.15252/embj.201592060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Raulf N., El-Attar R., Kulms D., Lecis D., Delia D. Differential response of head and neck cancer cell lines to TRAIL or Smac mimetics is associated with the cellular levels and activity of caspase-8 and caspase-10. Br J Cancer. 2014;111:1955–1964. doi: 10.1038/bjc.2014.521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Derakhshan A., Chen Z., Van Waes C. Therapeutic small molecules target inhibitor of apoptosis proteins in cancers with deregulation of extrinsic and intrinsic cell death pathways. Clin Cancer Res. 2017;23:1379–1387. doi: 10.1158/1078-0432.CCR-16-2172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Gaither A., Porter D., Yao Y., Borawski J., Yang G. A smac mimetic rescue screen reveals roles for inhibitor of apoptosis proteins in tumor necrosis factor-{alpha} signaling. Cancer Res. 2007;67:11493–11498. doi: 10.1158/0008-5472.CAN-07-5173. [DOI] [PubMed] [Google Scholar]
- 88.Krishna S.S., Majumdar I., Grishin N.V. Structural classification of zinc fingers: survey and summary. Nucleic Acids Res. 2003;31:532–550. doi: 10.1093/nar/gkg161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Corti A., Milani M., Lecis D., Seneci P., de Rosa M. Structure-based design and molecular profiling of Smac-mimetics selective for cellular IAPs. FEBS J. 2018;285:3286–3298. doi: 10.1111/febs.14616. [DOI] [PubMed] [Google Scholar]
- 90.Ndubaku C., Varfolomeev E., Wang L., Zobel K., Lau K. Antagonism of c-IAP and XIAP proteins is required for efficient induction of cell death by small-molecule IAP antagonists. ACS ChemBiol. 2009;4:557–566. doi: 10.1021/cb900083m. [DOI] [PubMed] [Google Scholar]
- 91.Vamos M., Welsh K., Finlay D., Lee P.S., Mace P.D. Expedient synthesis of highly potent antagonists of inhibitor of apoptosis proteins (IAPs) with unique selectivity for ML-IAP. ACS ChemBiol. 2013;8:725–732. doi: 10.1021/cb3005512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Abd-Elrahman I., Hershko K., Neuman T., Nachmias B., Perlman R. The inhibitor of apoptosis protein Livin (ML-IAP) plays a dual role in tumorigenicity. Cancer Res. 2009;69:5475–5480. doi: 10.1158/0008-5472.CAN-09-0424. [DOI] [PubMed] [Google Scholar]
- 93.Speer K.F., Cosimini C.L., Splan K.E. Characterization of a heterodimeric Smac-based peptide that features sequences specific to both the BIR2 and BIR3 domains of the X-linked inhibitor of apoptosis protein. Biopolymers. 2012;98:122–130. doi: 10.1002/bip.21732. [DOI] [PubMed] [Google Scholar]
- 94.Hrdinka M., Schlicher L., Dai B., Pinkas D.M., Bufton J.C. Small molecule inhibitors reveal an indispensable scaffolding role of RIPK2 in NOD2 signaling. EMBO J. 2018;37 doi: 10.15252/embj.201899372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Gonzalez-Lopez M., Welsh K., Finlay D., Ardecky R.J., Ganji S.R. Design, synthesis and evaluation of monovalent Smac mimetics that bind to the BIR2 domain of the anti-apoptotic protein XIAP. BioorgMedChemLett. 2011;21:4332–4336. doi: 10.1016/j.bmcl.2011.05.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Donnell A.F., Michoud C., Rupert K.C., Han X., Aguilar D. Benzazepinones and benzoxazepinones as antagonists of inhibitor of apoptosis proteins (IAPs) selective for the second baculovirus IAP repeat (BIR2) domain. JMedChem. 2013;56:7772–7787. doi: 10.1021/jm400731m. [DOI] [PubMed] [Google Scholar]
- 97.Kester R.F., Donnell A.F., Lou Y., Remiszewski S.W., Lombardo L.J. Optimization of benzodiazepinones as selective inhibitors of the X-linked inhibitor of apoptosis protein (XIAP) second baculovirus IAP repeat (BIR2) domain. JMedChem. 2013;56:7788–7803. doi: 10.1021/jm400732v. [DOI] [PubMed] [Google Scholar]
- 98.Goncharov T., Hedayati S., Mulvihill M.M., Izrael-Tomasevic A., Zobel K. Disruption of XIAP-RIP2 Association Blocks NOD2-Mediated Inflammatory Signaling. MolCell. 2018;69:565.e7. doi: 10.1016/j.molcel.2018.01.016. [DOI] [PubMed] [Google Scholar]
- 99.LaCasse E.C., Mahoney D.J., Cheung H.H., Plenchette S., Baird S. IAP-targeted therapies for cancer. Oncogene. 2008;27:6252–6275. doi: 10.1038/onc.2008.302. [DOI] [PubMed] [Google Scholar]
- 100.Lu M., Lin S.C., Huang Y., Kang Y.J., Rich R. XIAP induces NF-kappaB activation via the BIR1/TAB1 interaction and BIR1 dimerization. MolCell. 2007;26:689–702. doi: 10.1016/j.molcel.2007.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Zheng C., Kabaleeswaran V., Wang Y., Cheng G., Wu H. Crystal structures of the TRAF2: cIAP2 and the TRAF1: TRAF2: cIAP2 complexes: affinity, specificity, and regulation. MolCell. 2010;38:101–113. doi: 10.1016/j.molcel.2010.03.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Cossu F., Milani M., Grassi S., Malvezzi F., Corti A. NF023 binding to XIAP-BIR1: searching drugs for regulation of the NF-kappaB pathway. Proteins. 2015;83:612–620. doi: 10.1002/prot.24766. [DOI] [PubMed] [Google Scholar]
- 103.Sun H., Nikolovska-Coleska Z., Yang C.Y., Qian D., Lu J. Design of small-molecule peptidic and nonpeptidic Smac mimetics. AccChemRes. 2008;41:1264–1277. doi: 10.1021/ar8000553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Varfolomeev E.E., Ashkenazi A. Tumor necrosis factor: an apoptosis JuNKie? Cell. 2004;116:491–497. doi: 10.1016/s0092-8674(04)00166-7. [DOI] [PubMed] [Google Scholar]
- 105.Condon S.M., Mitsuuchi Y., Deng Y., LaPorte M.G., Rippin S.R. Birinapant, a smac-mimetic with improved tolerability for the treatment of solid tumors and hematological malignancies. JMedChem. 2014;57:3666–3677. doi: 10.1021/jm500176w. [DOI] [PubMed] [Google Scholar]
- 106.Manzoni L., Samela A., Barbini S., Cairati S., Penconi M. 4-Connected azabicyclo[5.3.0]decane Smac mimetics-Zn(2+) chelators as dual action antitumoral agents. BioorgMedChemLett. 2017;27:2336–2344. doi: 10.1016/j.bmcl.2017.04.032. [DOI] [PubMed] [Google Scholar]
- 107.Mingozzi M., Manzoni L., Arosio D., Dal Corso A., Manzotti M. Synthesis and biological evaluation of dual action cyclo-RGD/SMAC mimetic conjugates targeting alpha(v)beta(3)/alpha(v)beta(5) integrins and IAP proteins. OrgBiomolChem. 2014;12:3288–3302. doi: 10.1039/c4ob00207e. [DOI] [PubMed] [Google Scholar]
- 108.Seneci P., Rizzi M., Ballabio L., Lecis D., Conti A. SPION-Smac mimetic nano-conjugates: putative pro-apoptotic agents in oncology. BioorgMedChemLett. 2014;24:2374–2378. doi: 10.1016/j.bmcl.2014.03.048. [DOI] [PubMed] [Google Scholar]
- 109.Wang P., Zhou J. Proteolysis targeting chimera (PROTAC): a paradigm-shifting approach in small molecule drug discovery. CurrTopMedChem. 2018;18:1354–1356. doi: 10.2174/1568026618666181010101922. [DOI] [PubMed] [Google Scholar]
- 110.Tomoshige S., Nomura S., Ohgane K., Hashimoto Y., Ishikawa M. Discovery of small molecules that induce the degradation of huntingtin. AngewChemIntEd Engl. 2017;56:11530–11533. doi: 10.1002/anie.201706529. [DOI] [PubMed] [Google Scholar]
- 111.Shibata N., Miyamoto N., Nagai K., Shimokawa K., Sameshima T. Development of protein degradation inducers of oncogenic BCR-ABL protein by conjugation of ABL kinase inhibitors and IAP ligands. CancerSci. 2017;108:1657–1666. doi: 10.1111/cas.13284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Demizu Y., Ohoka N., Nagakubo T., Yamashita H., Misawa T. Development of a peptide-based inducer of nuclear receptors degradation. BioorgMedChemLett. 2016;26:2655–2658. doi: 10.1016/j.bmcl.2016.04.013. [DOI] [PubMed] [Google Scholar]
- 113.Itoh Y., Ishikawa M., Kitaguchi R., Okuhira K., Naito M. Double protein knockdown of cIAP1 and CRABP-II using a hybrid molecule consisting of ATRA and IAPs antagonist. BioorgMedChemLett. 2012;22:4453–4457. doi: 10.1016/j.bmcl.2012.04.134. [DOI] [PubMed] [Google Scholar]
- 114.Silke J., Meier P. Inhibitor of apoptosis (IAP) proteins-modulators of cell death and inflammation. Cold Spring Harb PerspectBiol. 2013;5 doi: 10.1101/cshperspect.a008730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Harlin H., Reffey S.B., Duckett C.S., Lindsten T., Thompson C.B. Characterization of XIAP-deficient mice. MolCellBiol. 2001;21:3604–3608. doi: 10.1128/MCB.21.10.3604-3608.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Conze D.B., Albert L., Ferrick D.A., Goeddel D.V., Yeh W.C. Posttranscriptional downregulation of c-IAP2 by the ubiquitin protein ligase c-IAP1 in vivo. MolCellBiol. 2005;25:3348–3356. doi: 10.1128/MCB.25.8.3348-3356.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Conte D., Holcik M., Lefebvre C.A., LaCasse E., Picketts D.J. Inhibitor of Apoptosis Protein cIAP2 is Essential for Lipopolysaccharide-Induced Macrophage Survival. MolCellBiol. 2006;26:699–708. doi: 10.1128/MCB.26.2.699-708.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Moulin M., Anderton H., Voss A.K., Thomas T., Wong W.W. IAPs limit activation of RIP kinases by TNF receptor 1 during development. EMBO J. 2012;31:1679–1691. doi: 10.1038/emboj.2012.18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Laukens B., Jennewein C., Schenk B., Vanlangenakker N., Schier A. Smac mimetic bypasses apoptosis resistance in FADD- or caspase-8-deficient cells by priming for tumor necrosis factor alpha-induced necroptosis. Neoplasia. 2011;13:971–979. doi: 10.1593/neo.11610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Feoktistova M., Geserick P., Kellert B., Dimitrova D.P., Langlais C. cIAPs block ripoptosome formation, a RIP1/caspase-8 containing intracellular cell death complex differentially regulated by cFLIP isoforms. MolCell. 2011;43:449–463. doi: 10.1016/j.molcel.2011.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.McComb S., Cheung H.H., Korneluk R.G., Wang S., Krishnan L. cIAP1 and cIAP2 limit macrophage necroptosis by inhibiting Rip1 and Rip3 activation. Cell Death Differ. 2012;19:1791–1801. doi: 10.1038/cdd.2012.59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Robert X., Gouet P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014;42:W320–W324. doi: 10.1093/nar/gku316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Cossu F., Malvezzi F., Canevari G., Mastrangelo E., Lecis D. Recognition of Smac-mimetic compounds by the BIR domain of cIAP1. Protein Sci. 2010;19:2418–2429. doi: 10.1002/pro.523. [DOI] [PMC free article] [PubMed] [Google Scholar]