Abstract
Mammalian microRNAs (miRNAs) play a critical role in modulating the response of immune cells to stimuli. Cannabinoids are known to exert beneficial actions such as neuroprotection and immunosuppressive activities. However, the underlying mechanisms which contribute to these effects are not fully understood. We previously reported that the psychoactive cannabinoid Δ9–tetrahydrocannabinol (THC) and the non-psychoactive cannabidiol (CBD) differ in their anti-inflammatory signaling pathways. Using lipopolysaccharide (LPS) to stimulate BV-2 microglial cells, we examined the role of cannabinoids on the expression of miRNAs. Expression was analyzed by performing deep sequencing, followed by Ingenuity Pathway Analysis to describe networks and intracellular pathways. miRNA sequencing analysis revealed that 31 miRNAs were differentially modulated by LPS and by cannabinoids treatments. In addition, we found that at the concentration tested, CBD has a greater effect than THC on the expression of most of the studied miRNAs. The results clearly link the effects of both LPS and cannabinoids to inflammatory signaling pathways. LPS upregulated the expression of pro-inflammatory miRNAs associated to Toll-like receptor (TLR) and NF-κB signaling, including miR-21, miR-146a and miR-155, whereas CBD inhibited LPS-stimulated expression of miR-146a and miR-155. In addition, CBD upregulated miR-34a, known to be involved in several pathways including Rb/E2f cell cycle and Notch-Dll1 signaling. Our results show that both CBD and THC reduced the LPS-upregulated Notch ligand Dll1 expression. MiR-155 and miR-34a are considered to be redox sensitive miRNAs, which regulate Nrf2-driven gene expression. Accordingly, we found that Nrf2-mediated expression of redox-dependent genes defines a Mox-like phenotype in CBD treated BV-2 cells. In summary, we have identified a specific repertoire of miRNAs that are regulated by cannabinoids, in resting (surveillant) and in LPS-activated microglia. The modulated miRNAs and their target genes are controlled by TLR, Nrf2 and Notch cross-talk signaling and are involved in immune response, cell cycle regulation as well as cellular stress and redox homeostasis.
Introduction
MicroRNAs (miRNAs) are evolutionary conserved, endogenous non-coding, single-stranded small RNAs (19–23 nucleotides length) that regulate gene expression through post-transcriptional repression [1–2]. It has been demonstrated that miRNAs are involved in a wide variety of physiological processes, including cell proliferation and differentiation, development, apoptosis, metabolism, angiogenesis, immunity and homeostasis [3–5]. In contrast, dysregulated miRNA expression has been linked to a variety of acute and chronic diseases, including cancer, neurodegenerative disorders, autoimmunity, heart diseases, chronic viral infections as well as acute organ injury and ischemic stroke [6–9]. A number of miRNAs were found to be significantly upregulated in response to Toll-like receptor (TLR) ligands. These include miR-155, miR-146a and miR-21, which negatively regulate TLR signaling [4, 10–11]. TLRs initiate distinct signaling pathways depending on the adaptor molecules MyD88 and TRIF, leading to activation of the transcription factors NF-κB, AP-1, IRF3 and/or IRF7, which induce the production of pro-inflammatory cytokines and type I interferon (IFN) [12].
A growing body of evidence describes the role of miRNAs in macrophages and microglial cells under defined stimuli, which induce polarized phenotypes (M1/M2) [13–16]. Microglia are the resident macrophage-like cells of the central nervous system (CNS). Microglial cells actively scan their environment and contribute to the immune surveillance of the CNS [17]. There is increasing information about diversity as well as plasticity of microglial cells, from surveillance to phagocytic and activated states, showing a more complex phenotyping than the standard M1/M2. For example, Kadl et al., [18] defined the microglial phenotype Mox, characterized by the occurrence of Nrf2-mediated redox-regulated genes. The presence of this complex set of phenotypes leads to the heterogeneity of microglial cellular responses and to the wide range of space/time activation patterns of microglia [17,19].
Preparations derived from Cannabis (marijuana and hashish) are recognized nowadays as having many physiological effects and therapeutic applications [20–21]. The beneficial effects of marijuana and its active constituents, the cannabinoids, consist of suppression of nausea and vomiting, stimulation of appetite, relief of pain as well as amelioration of the unwanted symptoms of spasticity originated by multiple sclerosis (MS) [22]. In addition, cannabinoids are known to posses pro-apoptotic, neuroprotective and anti-tumor properties [23–25]. Amongst the different cannabinoids, Δ9-tetrahydrocannabinol (THC) and cannabidiol (CBD) are recognized for their potent immunosuppressive and anti-inflammatory properties [24, 26–34]. THC mediates its psychoactive activity by binding to the cannabinoid receptor CB1 and modulates its immune response primarily through activation of the cannabinoid receptor CB2. In contrast to THC, CBD shows little agonistic activity at CB1/CB2 receptors and is therefore devoid of the unwanted psychotropic effects, characteristic of THC action on CB1 receptor. Moreover, it has been reported that CBD displays some CB1/CB2 antagonistic activities [35]. More recently, Laprairie et al., [36–37] showed that CBD can serve as a non-competitive negative allosteric modulator of the CB1 receptor with the ability to inhibit its internalization.
CBD has been shown to exhibit many physiological effects, including anti-inflammatory, anti-oxidant and neuroprotective properties [28–32, 38–44]. These effects of CBD have been related to diverse non CB1/non CB2 mechanisms [43, 45–48].
By performing microarray analysis of genome-wide mRNA in microglial BV-2 cells, our group showed that CBD is associated to genes involved in the GCN2/eIF2α/p8/ATF4/Chop-Trib3 pathway. In addition, we reported that genes stimulated by CBD (mainly related to stress response and inflammation) are regulated via the (EpRE/ARE)-Nrf2/Atf4 system and the Nrf2/Hmox axis in LPS-treated microglial cells [31, 33].
The effect of THC on miRNA expression profiles has been studied in diverse inflammatory biological models by the group of Nagarkatti [49–52] and in simian immunodeficiency virus-infected macaques by the group of Molina [53–54]. However, to our knowledge, there are no reports showing how miRNAs regulate the CBD-associated immunosuppression effects.
In this work we studied the effects of the pro-inflammatory LPS, a powerful inducer of TLR4 signaling, on the expression of miRNAs in BV-2 microglial cells as well as the effects of the cannabinoids CBD and THC on the LPS-inflammatory response. Next generation sequencing was used to identify the differentially expressed miRNAs. Highly predictive miRNA target genes and their pathways were analyzed using bioinformatics computational tools. Our results show that CBD is a potent regulator of miRNA expression, in resting (surveillant) and in LPS-activated microglia. MiRNA target genes were found to be involved in immune response, cell growth and proliferation, cell death and survival as well as stress response and redox regulation. Identification of the CBD- and THC-modulated miRNAs and their related networks and pathways provide a molecular basis for understanding the effects of these cannabinoids on activated microglia.
Materials and methods
Materials
THC was obtained from the National Institute on Drug Abuse (Baltimore, MD, USA) and CBD was provided by THC Pharm GmbH (Frankfurt, Germany). Stock solutions of cannabinoids were prepared in ethanol and diluted in culture medium before experiments. Final ethanol concentration in the medium did not exceed 0.1%. Fetal calf serum (FCS), tissue culture reagents and medium were obtained from Biological Industries (Kibbutz Beit HaEmek, Israel). LPS from Escherichia coli (serotype 055:B5) was purchased from Sigma (St. Louis, MO, USA).
Microglial cell culture
The immortalized murine BV-2 microglial cell line was kindly provided by Prof. E.J. Choi from Korea University (Seoul, Korea). BV-2 cells were grown in Dulbecco’s modified Eagle’s medium (DMEM) containing 4.5 g/l glucose, and supplemented with 5% FCS, penicillin (100 U/ml) and streptomycin (100 μg/ml) under a humidified 5% CO2 atmosphere at 37°C. Cells (1x106 cells in 100 mm plates) were pretreated for 2h with either CBD or THC (both at 10 μM), followed by the addition of LPS (100 ng/ml) for another 4h. These concentrations and time points were chosen following our previous studies [28,31,33].
Total RNA extraction
Total RNA (mRNA + miRNAs) was extracted using the NucleoSpin miRNA extraction kit (Macherey-Nagel; Düren, Germany) according to the manufacturer’s recommendation. Total RNA was quantified using the NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, DE), and the A260/230 and A260/280 ratios showing RNA purity, were examined. RNA quality was tested using the Eukaryote Total RNA Nano Assay, recorded with an Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA) and the RNA integrity number (RIN) was used as an RNA integrity parameter. Only samples with an RIN greater than 8 were used for further experiments and sequencing.
miRNA sequencing
miRNA sequencing was performed at the Informatics Center for Neurogenetics and Neurogenomics core at the University of California Los Angeles (UCLA), USA. Eighteen sequencing libraries were prepared using Illumina TruSeq Small RNA Library Preparation Kit following manufacturer's protocol. All samples were multiplexed into a single pool in order to avoid batch effects [55] and sequenced using an Illumina HiSeq 2500 sequencer (Illumina, San Diego, CA) at 69bp-paired-end sequencing, yielding between 2.8 and 5.6 million reads per sample. Quality control (QC) was performed on base qualities and nucleotide composition of sequences. Reads were aligned to the miRBase mouse reference using the BWA read aligner, run allowing for a maximum of 10 mappings per read. Between 39 and 51% (average 46%) of the reads mapped uniquely to the mouse genome. Total counts of reads aligned to known microRNAs for mouse are used as the basis for quantification of gene expression. Differentially expressed genes were identified using the Bioconductor package edgeR [56] which were then considered and ranked based on adjusted p-values of ≤ 0.1. RNA seq data has been deposited within the Gene Expression Omnibus (GEO) public repository with the accession number GSE123262 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE123262).
mRNA real time PCR (qPCR) analysis
The same RNA samples were used to test both mRNA and miRNA expressions by qPCR. For mRNA expression studies, cDNA was synthesized using the QuantiTect Reverse Transcription kit containing gDNA ‘wipe out’, (Qiagen AG, Basel, Switzerland) according to the manufacturer’s instructions. qPCR was carried out in the Rotor-Gene 3000 instrument, using Absolute Blue QPCR SYBR Green ROX mix containing the Thermo-Start DNA polymerase (Thermo Scientific, ABgene House, Epsom, Surrey, UK). Primer pairs for mRNA expression for selected transcripts were designed using the computer program Primer Quest, an online tool provided by Integrated DNA Technologies (http://test.idtdna.com/Primerquest/Home/Index). Wherever possible, designs with at least one of the primer sequences located on an intron-exon boundary were chosen, thus avoiding co-amplification of minor contaminating amounts of genomic DNA that could be present in the RNA samples. All primers were analyzed using the nucleotide program BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to ensure primer specificity for the gene of interest. Designed primers were synthesized by Metabion International AG (Planegg/Steinkirchen, Germany) with annealing temperatures of 60°C. Primers for qPCR were the followings, for B2-microglobulin: (B2m)-Forward: 5’-ATGGGAAGCCGAACATACTG-3’ and B2m-Reverse: 5’-CAGTCTCAGTGGGGGTGAAT-3’ (amplicon 176bp); for Dll1: Dll1-Forward: 5’-TGGACAGTTGCTTTGAAGAGTA-3’ and Dll1-Reverse: 5’-CAAGGAAAGACTGGCTCATAGT-3’ (amplicon 113bp), and for Notch1: Notch1-Forward: 5’- CCTGACAACTTGAGCCAGTAA-3’ and Notch1-Reverse: 5’-CGTCTGTCCTCAGTTGGATTT-3’ (amplicon 118bp). For each of the examined mRNAs, normal and mock reverse transcribed samples (in the absence of reverse transcriptase), as well as no template controls (total mix without cDNA) were run. qPCR was carried out as detailed by Juknat et al., [31]. Expression levels of genes were normalized to the reference gene B2m and expressed as fold change using the comparative cycle of threshold method as reported earlier [31]. All qPCR reactions were performed in duplicates and were repeated 3–4 times using different mRNA preparations.
miRNA qPCR analysis
For miRNA expression studies, cDNA synthesis was performed using the miRCURY LNA Universal cDNA Synthesis kit II (Exiqon, Woburn, MA), according to manufacturer’s instructions. miRNA real time qPCR was carried out using the ExiLENT SYBR Green master mix (Exiqon) and specific primers for mmu-miR-103a-3p (MIMAT0000101; Exiqon # 204063), mmu-miR-34a-5p (MIMAT0000542; Exiqon # 204486), mmu-miR-449a-5p (MIMAT0001542; Exiqon # 204481) and mmu-miR-191-5p (MIMAT0000221; Exiqon # 204306). The qPCR reactions were subjected to an initial activation step at 95°C of 10 min, followed by 40 cycles at 95°C for 10 s and at 60°C for 1 min. For quality control, synthetic RNA spike-in (UniSp6) was added to all RNA samples before reverse transcription and the UniSP6 was used to monitor the efficiency of the RT reaction. Expression of miRNAs was normalized using the means of the expression of miR-103a-3p and miR-191-5p, considered to be the most suitable and stable reference genes.
For the expression studies of miR-146a and miR-155-5p, RNA was reverse-transcribed using miScript II RT Kit (Qiagen), according to the manufacturer’s instructions. Real time qPCR was carried out using miScript SYBR Green PCR Kit (Qiagen), which contains the miScript Universal primer. Primer sequences used were the followings, for miR-146a-5p: 5’-TGAGAACTGAATTCCATGGGT-3’ (MIMAT0000158) and for miR-155-5p: 5’-TTAATGCTAATTGTGATAGGGGT-3’ (MIMAT0000165). miRNA expression was normalized to the reference gene U6 snRNA (U6 small nuclear RNA; NR_003027.2; 5’-GATGACACGCAAATTCGTGAA-3’), which was found to be expressed at high levels and showed no changes under the experimental conditions used in the present study. Real time qPCR reactions were performed at 95°C for 15 min, followed by 40 cycles at 94°C for 15 s, at 55°C for 30 s and at 70°C for 30 s. Melting curve was generated at the end of each run, in order to ensure product specificity. Fold change values in miRNA expression were calculated using the comparative cycle of threshold method as previously described [31]. All qPCR reactions were performed in duplicates in a Rotor-Gene 3000 instrument and were repeated 3–4 times using different miRNA preparations.
miRNA target predictions
Mature miRNA sequences were obtained from miRBase v21 (http://www.mirbase.org/). The prediction tools MiRTarBase (http://mirtarbase.mbc.nctu.edu.tw) and TarBase v7.0 (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=tarbase/index) were first chosen as they contain experimental evidence for the predicted miRNA targets. Other algorithms used were TargetScan v7.1 (http://www.targetscan.org/), miRanda, Release 2010 (http://www.microrna.org/; [57]) and miRDB (http://mirdb.org/miRDB/), computational programs based on seed match and conservation [58]. Sequence conservation was verified using UCSC Genome Browser (https://genome.ucsc.edu/). Sequence alignment was examined using the NCBI Basic Local Alignment Search Tool (BLAST) (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Given the high false positive rates for miRNA target prediction, a target was considered as potential if it appeared in at least three of the algorithms used.
Ingenuity pathway analysis
Pathway and global functional analysis were performed using Ingenuity Pathway Analysis (IPA), an online bioinformatics tool (Ingenuity Systems, QIAGEN Bioinformatics, Redwood City, CA, USA; www.qiagen.com/ingenuity). A data set containing the miRNA and gene identifiers as well as their corresponding expression values, were loaded to the application and each identifier was mapped using the Ingenuity Knowledge Base (IKB). The IKB analyses identify the biological functions as well as the pathways from the IPA library that are most significant to the data set. A cut-off threshold of p ≤0.005 for differential expression was used to build the networks using the IPA tools.
Statistical analysis
qPCR data is presented throughout the manuscript as fold change and is expressed as the mean ± SEM of 3–4 independent experiments. Statistical significance is analyzed using one way analysis of variance (ANOVA), followed by Tukey post-hoc test. p<0.05 was considered significant. GraphPad Prism 6.0 software (La Jolla, CA, USA) was used for statistical analysis of the data.
Results
LPS- and cannabinoid-differentially expressed miRNAs in BV-2 microglial cells
We used a high-throughput sequencing approach (miRNAseq) to analyze the expression of miRNAs in CBD-, THC-, LPS-, CBD+LPS, THC+LPS-treated microglial cells. Samples of miRNA were prepared from BV-2 microglial cells that were pre-incubated for 2 h with 10 μM CBD or 10 μM THC, followed by the addition of LPS (100 ng/ml) to the medium for another 4 h. It is important to note that, according to our previous results, CBD or THC treatment at 10 μM did not significantly affect the viability of BV-2 microglial cells during this 6 h incubation [28].
miRNA sequencing analysis revealed that 31 miRNAs were differentially regulated across the treatments (FDR 0.1; Table 1). Only the miRNAs reaching the criteria of p≤0.005 and fold change of 1.5 were considered for further analysis. They are shown as underlined (for downregulated miRNAs) or with an asterisk (for upregulated miRNAs) in Table 1. As can be seen, there are some miRNAs that although showing a fold change higher than 1.5, did not reach the cut-off threshold of p≤0.005.
Table 1. miRNA expression after treatment with LPS and cannabinoids, expressed as fold change versus control.
| MicroRNA | Accession# | LPS | CBD | CBD+LPS | THC | THC+LPS |
|---|---|---|---|---|---|---|
| mmu-miR-155-3p | MIMAT0016993 | 28.92* | 0.90 | 24.59* | 0.70 | 21.91* |
| mmu-miR-155-5p | MIMAT0000165 | 9.68* | 1.35 | 8.36* | 0.64 | 8.99* |
| mmu-mir-155 | MI0000177 | 6.51* | 1.37 | 5.26* | 0.69 | 6.43* |
| mmu-miR-451b | MI0021960 | 8.40* | 11.97* | 6.63 | 7.00 | 9.24* |
| mmu-miR-10b-5p | MIMAT0000208 | 7.40* | 1.81 | 1.38 | 0.62 | 5.09* |
| mmu-miR-146a-3p | MIMAT0016989 | 5.52* | 0.68 | 3.38* | 0.92 | 3.56* |
| mmu-miR-21a-3p | MIMAT0004628 | 3.32* | 1.04 | 2.82* | 1.01 | 2.97* |
| mmu-mir-21a | MI0000569 | 2.97* | 1.02 | 2.96* | 1.18 | 3.32* |
| mmu-miR-21a-5p | MIMAT0000530 | 2.11* | 1.15 | 1.70* | 1.09 | 1.93* |
| mmu-miR-221-5p | MIMAT0017060 | 2.44* | 0.69 | 1.95* | 0.97 | 2.17* |
| mmu-mir-29c | MI0000577 | 2.40 | 1.39 | 2.53 | 1.82 | 3.76* |
| mmu-mir-20a | MI0000568 | 2.31* | 1.08 | 1.96 | 0.84 | 2.6* |
| mmu-miR-20a-3p | MIMAT0004627 | 2.14* | 1.22 | 1.61 | 0.98 | 1.82 |
| mmu-mir-222 | MI0000710 | 1.92* | 0.83 | 1.73* | 1.02 | 1.84* |
| mmu-miR-145a-3p | MIMAT0004534 | 1.81* | 1.17 | 1.73* | 0.99 | 1.92* |
| mmu-miR-449a-5p | MIMAT0001542 | 15.62 | 36.66* | 25.15* | 7.96 | 6.27 |
| mmu-mir-34a | MI0000584 | 1.46 | 7.23* | 4.48 | 2.37 | 2.67 |
| mmu-mir-5112 | MI0018021 | 1.09 | 2.75* | 1.01 | 1.34 | 0.96 |
| mmu-mir-181b-1 | MI0000723 | 1.06 | 3.10* | 1.92 | 1.07 | 1.21 |
| mmu-miR-5129-5p | MIMAT0020640 | 1.05 | 0.03 | 0.68 | 1.20 | 0.51 |
| mmu-mir-696 | MI0004677 | 0.89 | 0.39 | 0.44 | 0.73 | 0.70 |
| mmu-mir-3096b | MI0016969 | 0.84 | 2.02* | 1.02 | 0.84 | 1.02 |
| mmu-miR-3108-3p | MIMAT0014948 | 0.78 | 0.07 | 2.05 | 1.79 | 0.85 |
| mmu-miR-3962 | MI0016967 | 0.78 | 0.03 | 0.44 | 0.75 | 0.69 |
| mmu-mir-3090 | MI0014083 | 0.71 | 1.76 | 0.19 | 0.62 | 0.61 |
| mmu-miR-6351 | MI0021879 | 0.69 | 0.01 | 0.34 | 1.08 | 0.64 |
| mmu-miR-6384 | MI0021915 | 0.43 | 5.45* | 0.82 | 0.33 | 0.59 |
| mmu-mir-666 | MI0004553 | 0.43 | 0.12 | 0.53 | 1.21 | 0.52 |
| mmu-mir-325 | MI0000597 | 0.39 | 2.72* | 0.72 | 0.97 | 0.40 |
| mmu-miR-25-5p | MIMAT0017049 | 0.36 | 1 | 0.33 | 0.61 | 0.42 |
| mmu-miR-5123 | MI0018034 | 0.07 | 0.29 | 0.63 | 0.17 | 0.19 |
#Accession number for the corresponding mmu-miRNAs, available at http://mirbase.org/; significantly upregulated miRNAs are marked with an asterisk; significantly downregulated miRNAs are underlined (p≤0.005)
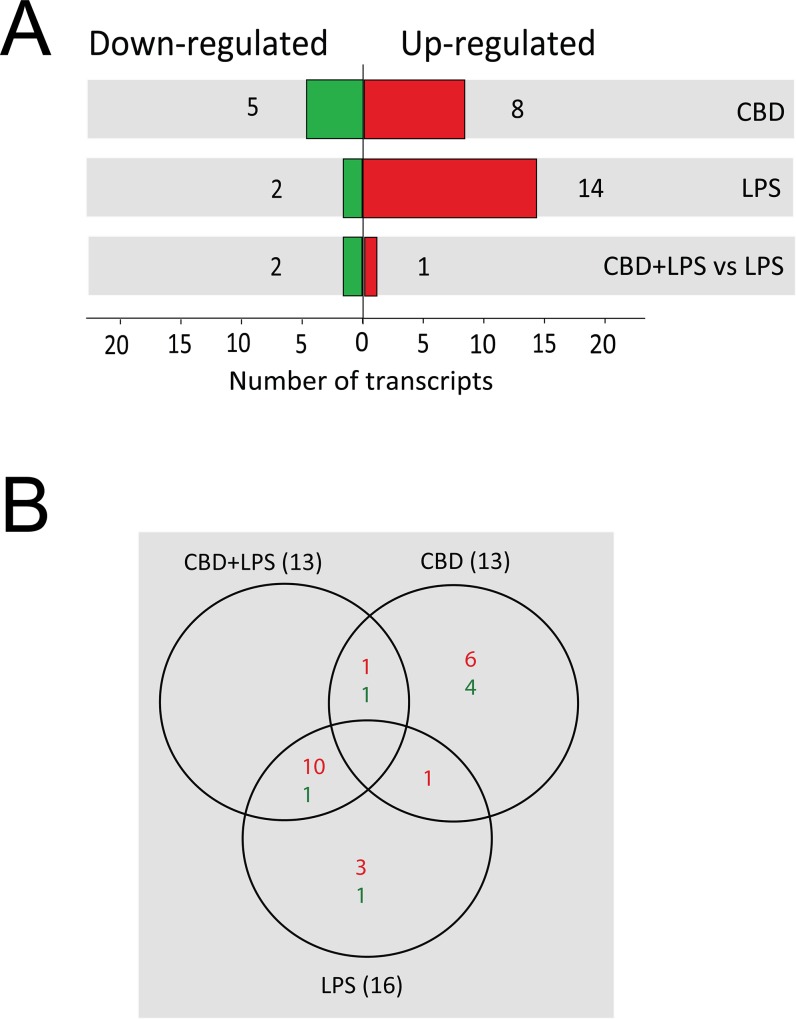
Analyzing the 16 miRNAs affected by LPS stimulation (Fig 1A and 1B; 52% of the total), we found that 14 miRNAs (87.5% of the total LPS-affected miRNAs) were significantly upregulated and that 2 miRNAs (12.5%) were significantly downregulated by the treatment (miR-5123 and miR-25-5p) (p≤0.005; Table 1). Under LPS stimulation, miR-155-3p was the most significantly upregulated miRNA (by 29-fold versus control), followed by others including miR-155-5p (by 9.7-fold), miR-451b (by 8.4-fold), miR-10b-5p (by 7.4-fold), miR-146a-3p (by 5.5-fold) and miR-21a-3p (by 3.3-fold) (p≤0.005; Table 1).
Fig 1. Deep sequencing analysis of the miRNA of BV-2 cells following treatment with LPS and cannabinoids.
miRNA was extracted from BV-2 cells treated with LPS and cannabinoids and subjected to deep sequencing analysis as described in Methods. (A) Number of differentially expressed miRNAs that were either significantly upregulated (red) or downregulated (green) across the different treatment conditions versus control untreated cells and versus LPS (p≤0.005). The differentially expressed miRNAs were identified by using a threshold of a 1.5-fold change. (B) Venn diagrams representing the numbers of BV-2 microglial miRNAs that were either upregulated or downregulated at a significance of p≤0.005 following treatments with LPS, CBD and CBD + LPS.
Most of the LPS-upregulated miRNAs were also significantly upregulated by CBD+LPS (10 miRNAs) as shown in Fig 1B, and by THC+LPS (13 miRNAs; Table 1). Moreover, in general, the addition of CBD or THC did not significantly affect the stimulatory effect of LPS. Only one miRNA, miR-10b-5p, that was upregulated by LPS stimulation (by 7.4-fold; p≤0.005; Table 1), was significantly downregulated by CBD + LPS (reduced by 81% when comparing CBD + LPS vs LPS; p≤0.005).
Following CBD treatment, the expression of 13 out of the 31 miRNAs (42% of the total) was significantly changed; 8 miRNAs (61.5% of the total CBD-affected miRNAs) were significantly upregulated by more than 2-fold and 5 miRNAs (38.5%) were downregulated by more than 50% compared to the control values (p≤0.005; Fig 1A and 1B). The miRNAs upregulated to the highest extent by CBD treatment were miR-449a-5p (by 37-fold vs control), miR-451b (by 12-fold), miR-34a (by 7.2-fold), miR-6384 (by 5.5-fold) and miR-181b-1 (by 3.1-fold) (p≤0.005; Table 1). The downregulated miRNAs included miR-6351 (reduced by 99%), miR-5129-5p (by 97%), miR-3962 (by 97%), miR-666 (by 88%) and miR-696 (by 61%) (p≤0.005; Table 1).
Interestingly, THC treatment only induced changes in one of these miRNAs (miR-5123 downregulated by 83%; p≤0.005; Table 1). From the 5 miRNAs (miR-451b, miR-29c, miR-449a-5p, miR-34a and miR-3108-3p) that show an increase (1.5-fold change) in the expression after THC treatment, only miR-451b could be considered statistically significant if we change the cut-off threshold to p≤0.05. The other miRNAs were found to be not statistically significant.
Role of CBD in the expression of miR-146a and miR-155-5p in LPS-activated microglial cells
Table 1 shows that several miRNAs are upregulated by LPS treatment. This included miR-155, miR-146a and miR-21. This result is in agreement with the finding that this trio of miRNAs is involved in many immune and inflammatory pathways, known to control TLR signaling [4, 10–11]. The effects of cannabinoids on the level of expression of miR-146a and miR-155-5p were evaluated by qPCR. The modulation of the expression of both miRNAs show a more pronounced effect of 10 μM CBD as compared to 10 μM THC (Fig 2).
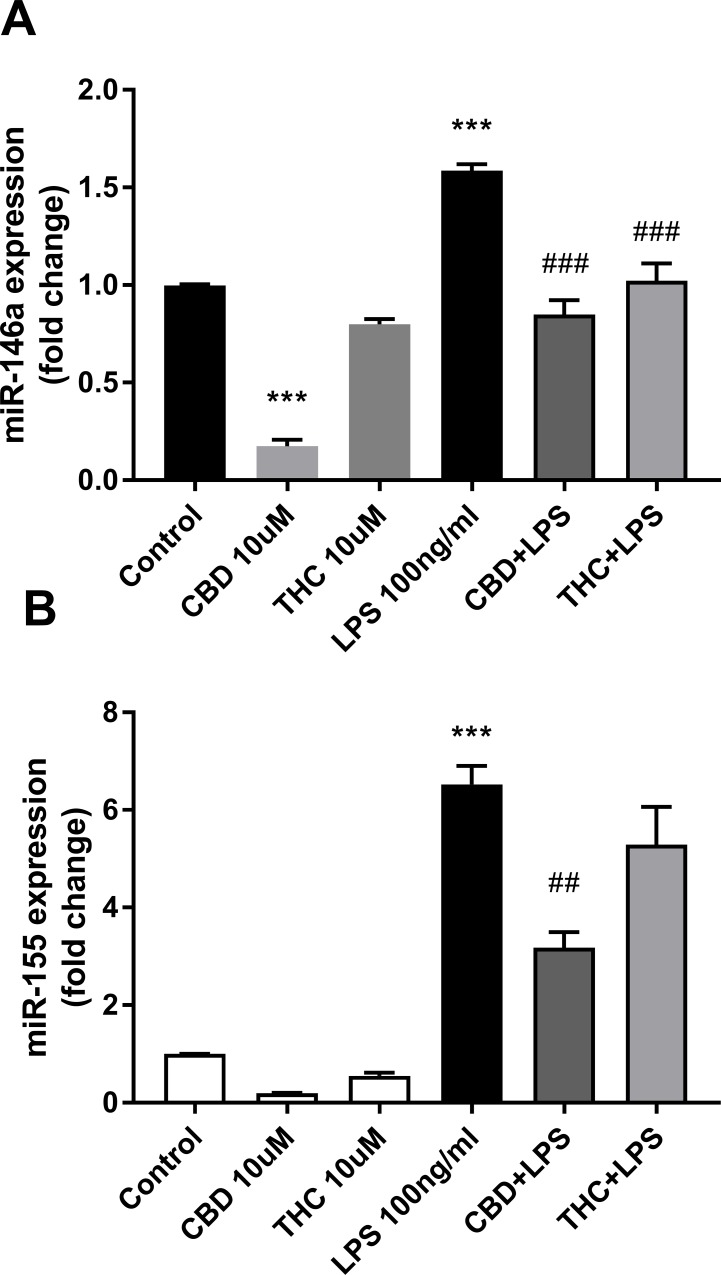
Fig 2. CBD downregulates the expression of miR-146a-5p and miR-155-5p in LPS-activated microglial cells.
Cells were treated for 2 h with the indicated concentrations of cannabinoids and then activated for 4 h with 100 ng/ml LPS. Cells were harvested and miRNA was extracted for qPCR analysis. qPCR data were plotted as the mean ± SEM of three to four independent experiments. (A) One-way ANOVA F(5,12) = 78.66 p < 0.001; (B) one-way ANOVA F(5,18) = 49.61 p < 0.001. Tukey multiple comparison post hoc test: ***p < 0.001 vs. control; ##p < 0.01 and ###p < 0.001 vs. LPS.
In resting cells, miRNA 146a expression was highly and significantly downregulated by CBD (by 83%; p<0.001; Fig 2A). In agreement with the sequencing data, both miR-146a and miR-155-5p were upregulated by treatment with LPS (by 1.6-fold; p<0.001, and 6.5-fold; p<0.001, respectively). This upregulation is reduced by pretreatment with CBD (by 47%; p<0.001, and by 51%; p<0.01, respectively) (Fig 2A and 2B). At the concentration tested, the effect of THC was lower than that of CBD; the LPS-upregulated miR-146a was reduced by 37% (THC + LPS; p<0.001; Fig 2A). THC pretreatment did not show any significant effect on LPS-upregulated miR-155-5p (THC + LPS; Fig 2B).
Target prediction was performed for both, miR-146a and miR-155. S1 and S2 Tables contain a list of genes, validated according to miRTarBase and TarBase prediction tools. These targets were chosen using bioinformatics tools as described in Methods. Indeed, we found that many of these genes are significantly affected by LPS as well as by the presence of cannabinoids during LPS activation, according to the RNAseq data (fold change set on ≥1.4-fold or ≥40% reduction; p≤0.005). Selected target genes were found to be involved in TLR signaling (Irak2, Irak3, Ikbkε/Ikkε, Cot/Tpl2/Map3k8, Inpp5d/Ship1), cytokine-mediated signaling (Il6, Il6ra), regulation of transcription (Stat1, Pparg, Tp53inp1/Trp53inp1), NF-κB signaling (Relb, Ikbkε/Ikkε), stress response (Hmox1, Tp53inp1/Trp53inp1, Nrf2/Nfe2l2), signal transduction (Cish, Socs1), Notch pathway (Notch1, Notch2, Tspan14), lipid signaling (Lipin1, S1pr1) as well as cell death and apoptosis (Fas, Cflar).
S1 Table shows that the cannabinoids reduced the expression of LPS-upregulated Il6 mRNA as previously reported by us and others [28]. Among other targets, we were interested in Pparg, even though it was identified as a prediction target for miR-146a by only one prediction tool, the miRTarBase. Pparg mRNA expression was observed to be inversely correlated with miR-146a expression in LPS-stimulated microglia, following the canonical repression pattern (S1 Table). Indeed, we previously showed that LPS downregulates the mRNA expression of both Pparg1 and Pparg2 isoforms, in BV-2 microglial cells [31], in agreement with reports showing that NF-κB drives down Pparg expression in LPS-stimulated macrophages [59]. However, CBD did not exhibit any significant effect on Pparg mRNA expression in BV-2 microglial cells eventhough Pparg has been identified as a molecular target for CBD [60].
miR-34a/449a is upregulated in microglial cells in response to CBD treatment
One of the miRNAs that attracted our attention was miR-449, that was greatly upregulated by CBD (Table 1). The miR-449 cluster, containing the highly conserved miR-449a, miR-449b and miR-449c, was found to share strong sequence homologies with the miR-34 family (miR-34a, miR-34b and miR-34c) and therefore, classified as the miR-34/449 family. MiR-34a and miR-449a share the same seed sequence (GGCAGUG), suggesting similar mRNA targets [61]. However, despite these similarities, miR-449 and miR-34 seem to have distinct functions and to be regulated differentially [62]. MiR-34a is ubiquitously expressed, except in lung tissue, showing the highest levels in brain tissues, while miR-34b/c and miR-449a/b/c are mainly expressed in lung and testis, demonstrating tissue-specific functions [61, 63].
Analysis of miR-34a and miR-449a expressions by qPCR in BV-2 microglial cells showed similar profiles (Fig 3).
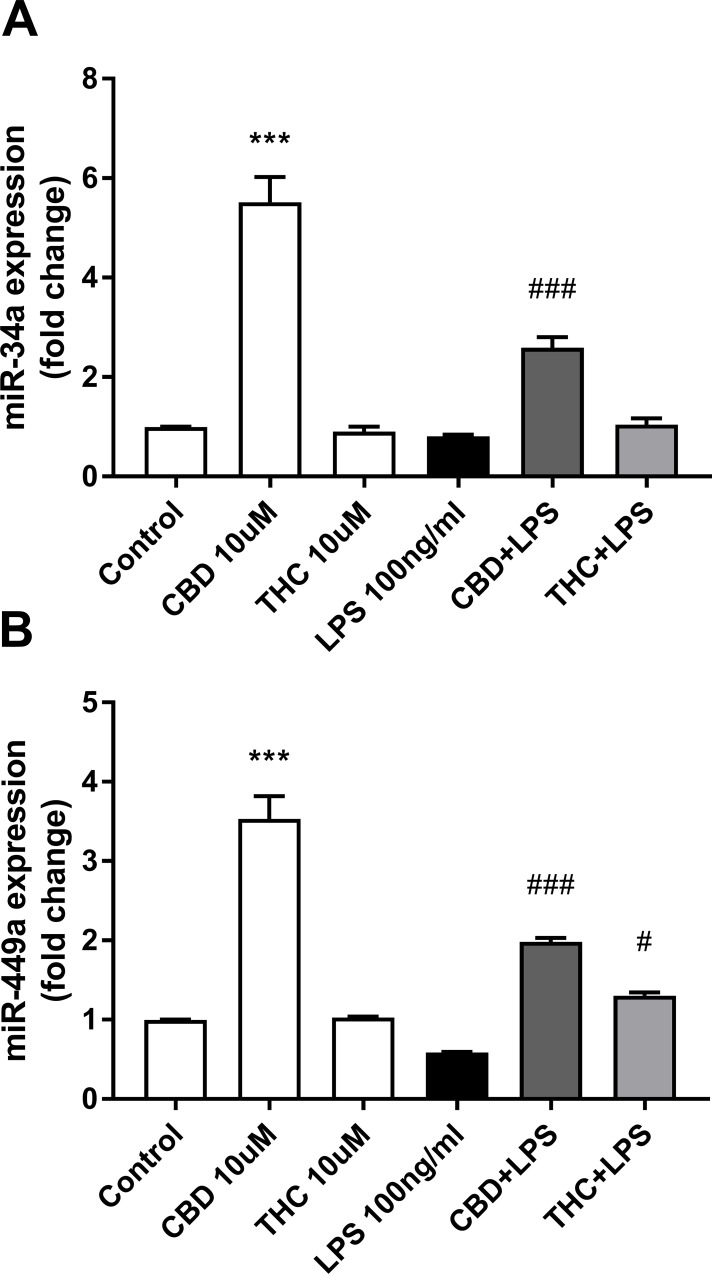
Fig 3. CBD upregulates the expression of miR-34a/449a in resting microglial cells.
Conditions are as indicated in Fig 2. qPCR data were plotted as the mean ± SEM of three to four independent experiments. (A) One-way ANOVA F(5,19) = 62.78 p < 0.001; (B) one-way ANOVA F(5,12) = 78.31 p < 0.001. Tukey multiple comparison post hoc test: ***p < 0.001 vs. control; #p < 0.05 and ###p < 0.001 vs. LPS.
Both, miR-34a and miR-449a were upregulated by CBD (by 5.5-fold; p<0.001; Fig 3A and by 3.5-fold; p<0.001; Fig 3B respectively), although miR-449a to a lower degree when compared to the sequencing data (37-fold; p≤0.005; Table 1). As pointed out for miR-146a and miR-155-5p, we observed also for miR-34a and miR-449a a more pronounced effect of CBD as compared with THC. Using qPCR, we examined miR-34a expression in BV-2 cells stimulated with different doses of LPS (0 to 5 μg/ml) and found out that it was not affected by LPS at any dose (Fig 4A); on the other hand, it was upregulated by CBD treatment in a dose dependent manner (Fig 4B). It is interesting to point out that THC does not affect the expression of miR-34a at all the concentrations tested (0 to 10 μM) under our experimental conditions (data not shown).
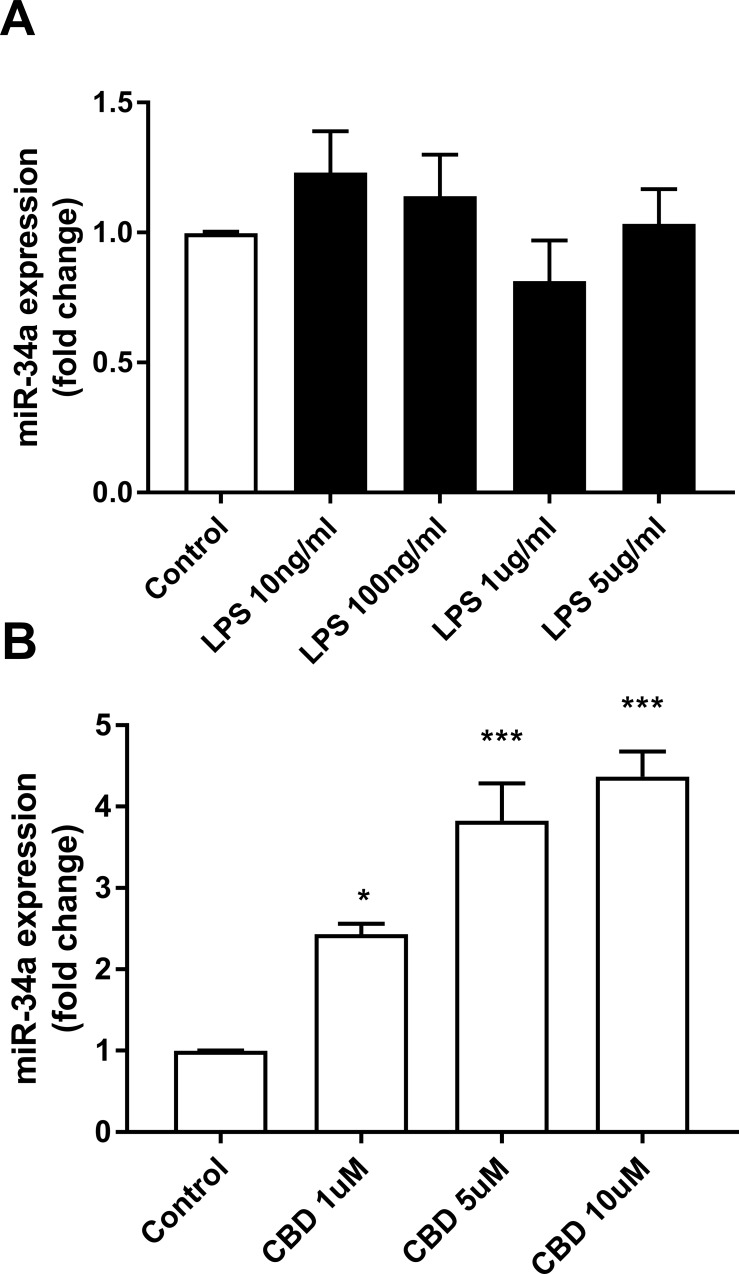
Fig 4. miR-34a expression is not affected by LPS, but upregulated by CBD.
Cells were treated for 4 h with the indicated concentrations of LPS or for 6 h with the indicated concentrations of CBD. Cells were harvested and miRNA extracted for qPCR analysis. qPCR data were plotted as the mean ± SEM of three to four independent experiments. (A) One-way ANOVA F(4,10) = 1.339 not significant; (B) one-way ANOVA F(3,8) = 28.12 p < 0.001. Tukey multiple comparison post hoc test: *p < 0.05 and ***p < 0.001 vs. control.
Potential target genes for miR-34a-5p were chosen using the target prediction tools described in Methods. S3 Table shows a selection of miR-34a target genes which were found to be affected by cannabinoids, LPS, or their combinations. Most of these target genes encode factors required for cell cycle regulation (Ccnd1, Ccne2, Cdkn1a/p21, E2f5, Cdc25a), Notch signaling (Notch1, Notch2, Dll1), inflammation (Il6, Tnf), Wnt/β-catenin signaling (Wnt1), stress response (Slc7a11/xCT, Ddit4, Ndrg1, Tp53inp1/Trp53inp1), membrane transport (Aqp9, Slc7a11/xCT), apoptosis (Gas1, Mdm2, Pdgfrb), regulation of transcription (Rora, Rela, Foxp1, Hes2, Tp53inp1/Trp53inp1), and senescence (Hbp1). Among these targets, the expression of the cell cycle-related transcripts (Ccnd1, Ccne2, E2f5, Cdc25a) was found to be inversely correlated with miR-34a expression in CBD-treated cells (S3 Table).
Among the miR-34a target genes, we validated the Notch ligand Dll1 by qPCR (Fig 5).
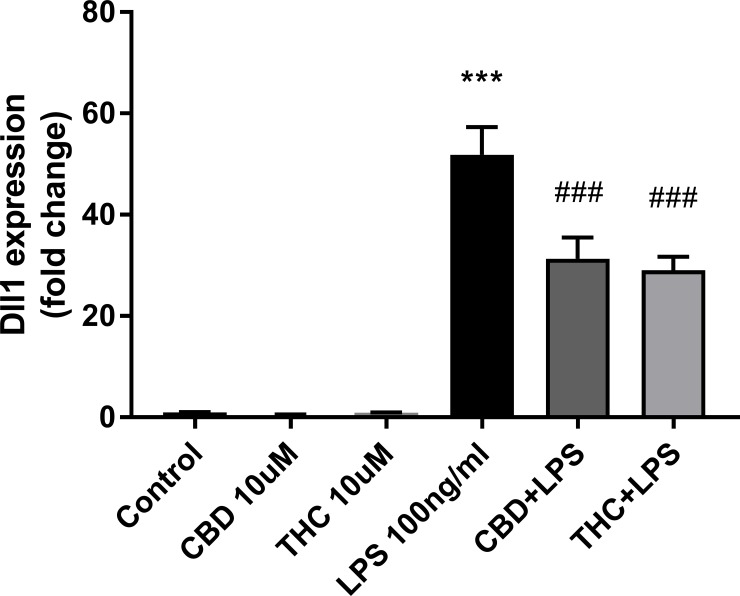
Fig 5. CBD and THC downregulate the expression of the Notch ligand Dll1 in LPS-activated microglial cells.
Conditions are as indicated in Fig 2. qPCR data were plotted as the mean ± SEM of three to four independent experiments. One-way ANOVA F(5,24) = 50.54 p < 0.001. Tukey multiple comparison post hoc test: ***p < 0.001 vs. control; ###p < 0.001 vs. LPS.
Surprisingly, in resting cells, Dll1 expression was downregulated by CBD by 56%, a result that was found to be not statistically significant, in contrast to the upregulation pattern observed in the sequencing data (upregulation by 11-fold; p≤ 0.0005; S3 Table). Analyzing the effect of LPS, we observed an upregulation of Dll1 expression by LPS (by 52-fold; p<0.001), which was decreased by pretreatment with CBD (by 40%; p<0.001) and with THC (by 44%; p<0.001) (Fig 5).
Network analysis and signaling pathways
The differential miRNA expression patterns, revealed by miRNAseq of BV-2 microglial cells treated with LPS and/or cannabinoids, were analyzed using IPA, which integrates published findings on biologically meaningful genetic or molecular gene/gene product interactions and identifies new targets as well as functionally related gene networks. Differential miRNA expression values (e.g.; LPS vs control, CBD vs control or CBD + LPS vs LPS) were entered into IPA to determine the most highly regulated networks of miRNA-mRNA interactions and to highlight the biological processes that are relevant to each of the treatments. Genes that met the p value cutoff of 0.005 for differential expression were used to build the gene networks using IPA tools. Networks describing the relationships between miRNAs and a subset of genes and their related transcripts are presented in Figs 6–8.
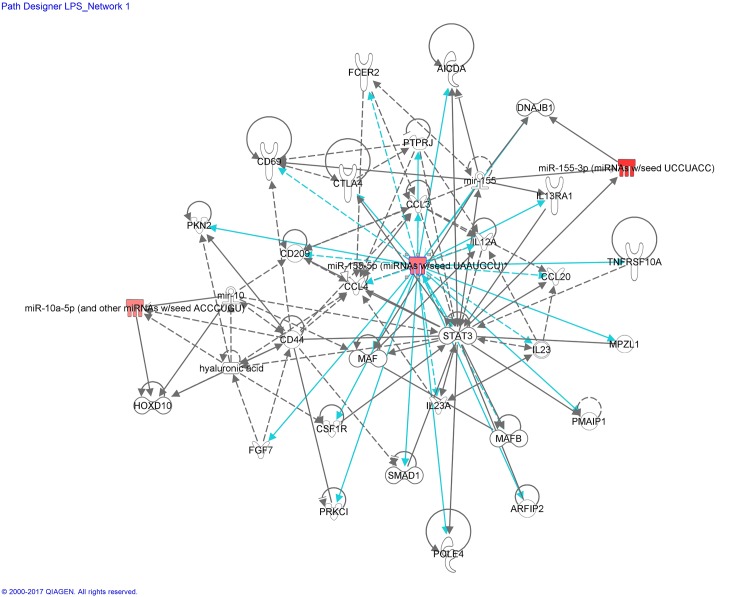
Fig 6. miRNA:mRNA gene network associated to LPS-stimulated microglial cells, identified by Ingenuity Pathway Analysis (IPA)—Network LPS_miR-155-5p.
Network analysis of the genes whose expression was affected by LPS was performed using the Ingenuity software. Each network displays the genes/gene products as nodes (different shapes representing the functional classes of gene products) and the biological relationships between the nodes as lines. Full lines represent direct interactions and broken lines, indirect interactions. Lines in gray represent interactions shown in the literature. Lines in turquoise represent the interactions provided by our mRNA/miRNA sequencing data. The color intensity of the mature miRNA node indicates the degree of upregulation (red) of the respective miRNA.
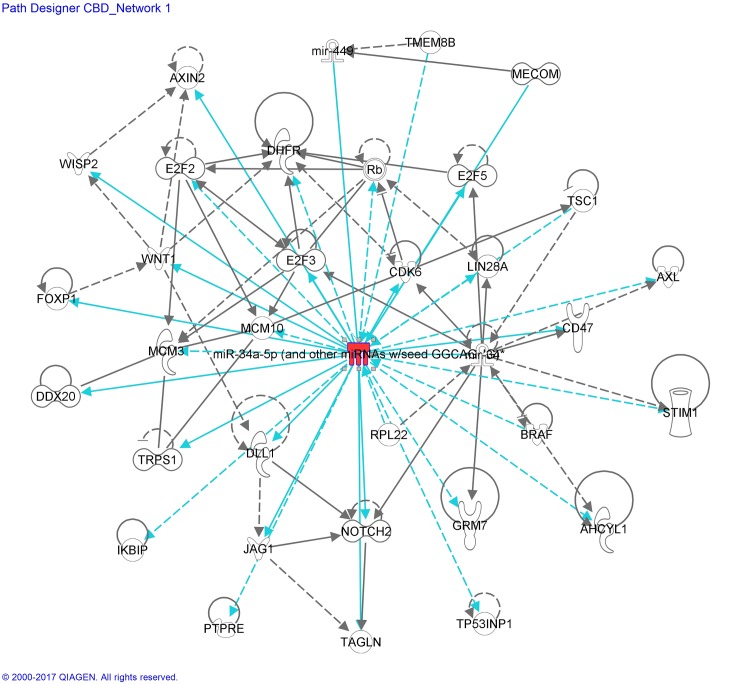
Fig 8. miRNA:mRNA gene network associated to CBD-treated microglial cells, identified by IPA—Network CBD_miR-34a-5p.
Details are as indicated in Fig 6.
Interactome analysis of LPS and associated miRNA:mRNA gene network is shown in Fig 6.
This network has in its central position the mature miRNA-155-5p (and other miRNAs with seed sequence UAAUGCU, including miR-155, considered by IPA as a duplicate). These miRNAs are upregulated by LPS (Table 1; Fig 2B). The LPS-Network interconnects miR-155-5p (possessing 15 edges, via direct connections and 9 edges, via indirect ones) with cytokines (Il12a/b, Il23a, Il13ra1), chemokines (C-C motif) (Ccl4, Ccl3, Ccl20), cell surface antigen molecules (Cd209, Cd44, Cd69, Cd23/Fcer2a, Cd152/Ctla4), transcription factors (Stat3, Smad 1, MafB), regulators of apoptotic processes (Pmaip1/Noxa), kinases (Pkn2, Prkci), phosphatases (Ptprj), other enzymes (Aicda/Arp2/Aid), regulators of signal transduction (Mpzl1) and regulators of cytoskeleton organization (Arfip2/Por1).
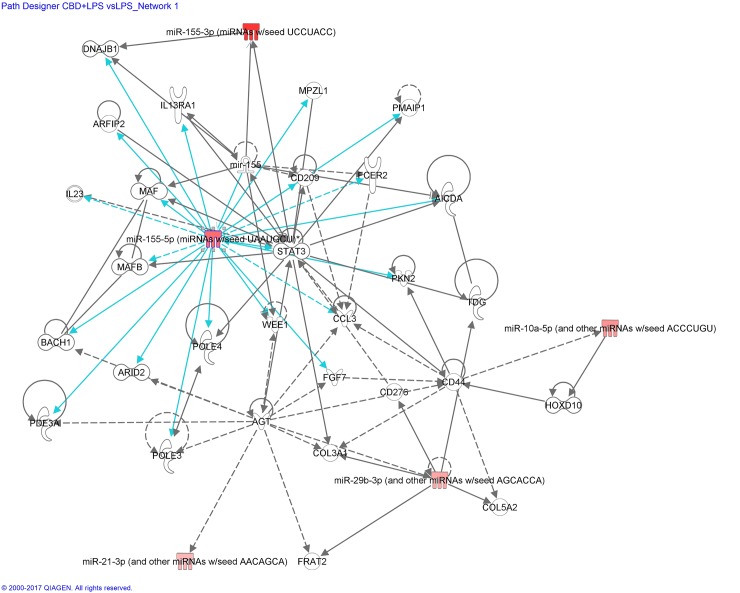
Fig 7 shows miRNA:mRNA interactions for CBD + LPS vs LPS.
Fig 7. miRNA:mRNA gene network showing the effect of CBD on LPS-stimulated microglial cells, identified by IPA—Network CBD + LPS vs LPS_miR-155-5p.
Details are as indicated in Fig 6.
Mature miR-155-5p appears to play a central role, showing 17 edges of direct connections. However, only three transcripts were exclusively found to be interconnected to miR-155-5p after CBD+LPS treatment: Bach1, a transcription factor involved in oxidative stress; Wee1, a cell cycle regulator; and Pole3/Ybl1, an enzyme with polymerase activity. The other transcripts interconnected to miR-155 after LPS treatment, were already exhibited in Fig 6.
Interactome analysis of CBD and associated miRNA:mRNA gene network is shown in Fig 8.
MiR-34a-5p (and other miRNAs with seed sequence GGCAGUG, including miR-449a-5p considered by IPA as a duplicate of miR-34a-5p) is upregulated by CBD (Table 1; Fig 3A and 3B) and plays a central role in this network, possessing 14 and 13 edges via direct and indirect connections. A series of canonical pathways, including those involved in Rb (Retinoblastoma)/E2F cell cycle (Rb, E2f2, E2f3, E2f5, Cdk6, DHFR), Notch (Dll1, Notch 2, Jag1), Wnt (Wnt, Axin2, Wisp2/Ccn5) and NF-κB (Ikbip) were found to participate in this network. Other affected genes are related to apoptosis (Ddx20, Ikbip), integrin-associated signal transduccion (Cd47), stress protein with antioxidant-associated tumor suppressive function (Tp53inp1/Trp53inp1), calcium signaling (Stim1), as well as those related to proliferation and survival (Lin28a).
Discussion
In the present study, we identified a repertoire of miRNAs that are differentially modulated by cannabinoids and/or LPS treatments in microglial BV-2 cells.
It attracted our attention the poor effect exhibited by THC in our experimental model. Only miR-5123 is significantly affected by THC. In addition, 14 miRNAs were found to be upregulated by THC+LPS treatment, from them 13 miRNAs were also affected by LPS more or less, to the same extent. THC reduces the expression of LPS-upregulated miR-146a but not of LPS-upregulated miR-155-5p. THC+LPS treatment increased the expression of miR-449a-5p but not of miR-34a-5p. Comparing the cannabinoids tested, 10 μM CBD appears to have a much greater effect than 10 μM THC on the expression of these miRNAs. These results are in line with the more robust effects of CBD on mRNA regulation in resting/surveillant and LPS-activated microglial cells, as compared with THC, shown by us previously [28,31,33].
Here we show that CBD strongly downregulates miR-146a in resting cells and that both CBD and THC inhibit the LPS-upregulated expression of this miRNA. Once LPS induces NF-κB signaling through the MyD88-dependent pathway, activated NF-κB induces the transcription of many genes, including miR-146a and the major pro-inflammatory cytokines Il-6 and Il-1β.
In addition, an excessive activation of NF-κB induces miR-146a to act as a negative regulator of inflammation, leading to a reduction of NF-κB transcriptional activity through targeting Irak1 and Traf6, and through suppression of NF-κB inducible genes such as Il-6 and Il-1β [64]. It is important to note that the key role of miR-146a as a negative feed-back regulator is fundamental to prevent a disregulated inflammation process and to control its magnitude, but miR-146a is also upregulated during immune activation through the increase in NF-κB activity [65]. We suggest that CBD is downregulating miR-146a expression by targeting NF-κB signaling, and by this way, decreasing the mRNA levels of LPS-upregulated Il-6 and Il-1β in BV-2 microglial cells [28]. In this regard, our group previously demonstrated that CBD reduces the activity of the NF-κB signaling pathway, by partially decreasing LPS-induced degradation of IκB as well as by reducing LPS-induced NF-κB p65 subunit phosphorylation [28]. In addition, CBD decreased the level of Il-6 and Il-1β proteins, providing evidence of the anti-inflammatory effects of CBD as mediated via NF-κB [28]. Although Irak1 and Traf6 were identified as direct targets for miR-146a, we did not find these adapter molecules through our target prediction tools. Moreover, studies from our group showed that 10 μM CBD did not affect the level of Irak1 and IkB proteins as well as the level of phosphorylated p65, in the absence of LPS, in 6h experiments [28]. However, we demonstrated that CBD partially decreased LPS-induced degradation of Irak1 [28], showing a possible participation of miR-146a as a negative regulator. Therefore, more studies are required in order to understand the sequence of events CBD+LPS/miR-146a/Irak1 and the final effect of CBD on NF-κB signaling.
Although miR-146a and miR-155 are co-induced by LPS, their final effects are different. MiR-146a exerts control of the inflammatory response leading to reduced inflammation by fine-tuning TLR signaling rather than abolishing the signal, whereas miR-155 is a pro-inflammatory miRNA, essential in driving the inflammatory phenotype M1 [66]. MiR-155 is the most highly upregulated miRNA in response to inflammatory stimuli, such as LPS and/or IFN-γ, in diverse immune cells [13, 67–68], including microglia [14–15, 69]. Reports show that the main target of miR-155 is the inhibitory protein of inflammation Inpp5d/Ship1 (inositol polyphosphate-5-phosphatase D/Src homology-2 domain-containing inositol 5-phosphatase 1), a negative regulator of TLR4 signaling. Our present results demonstrate that after LPS activation, miR-155-5p expression is upregulated, in a good correlation with the reduction of Inpp5d/Ship1 expression (downregulated by LPS by 60%), leading to inflammation. CBD pretreatment reverses this effect, in a good agreement with its inhibition of the LPS-upregulated miR-155-5p expression. CBD reduces the LPS-increased miR-155-5p expression by 51%, leading to a decrease in Inpp5d/Ship1 degradation (Inpp5d/Ship1 expression nearly reaches control values; p≤0.01). Although Inpp5d/Ship1 is a negative regulator of TLR4, its level of expression after CBD+LPS treatment for 6 h does not arrive to the threshold required to mounting an inflammatory response.
Another target of mir-155 is Ikbkε/Ikkε (inhibitor of kappaB kinase epsilon) which functions in the TLR4/TRIF/MyD88-independent pathway, inducing Ifnb production. Our group showed that CBD decreases the mRNA levels of LPS-upregulated Ifnb as well as the LPS-induced release of Ifnb in BV-2 cells, suggesting that CBD exerts its inhibitory activity upstream of Ifnb production [28]. As CBD reverses the LPS-upregulated Ikbkε expression reaching control values and decreases upregulated Ifnb1 mRNA expression (by 70%, p≤0.005; [33]), CBD might be acting at the level of the paralog kinases Ikbkε/Tbk1 (TANK-binding kinase 1), that are activating Irf3 and hence, Ifnb synthesis [70]. Altogether, these results provide evidence for the anti-inflammatory effects of CBD.
Among the miR-155 target transcripts, we find the stress response genes Hmox1 and Bach1, both of them upregulated by CBD (by 3.0-fold; p≤0.01 and by 40%; p≤0.005, respectively) and by CBD+LPS (by 9.4-fold; p≤0.0001 and by 80%; p≤0.0001, respectively). Interestingly, the highly pro-inflammatory miR-155 has been reported to be cytoprotective during inflammation by enhancing the induction of Hmox1 expression via inhibition of Bach1 translation in endothelial cells [71]. Moreover, miR-155 has been found to attenuate inflammation and oxidative injury via enhancement of Nrf2-mediated Hmox1 and Slc7a11/xCT gene expressions [72]. In addition, our group reported that many transcripts induced by CBD, including Hmox1 and Slc7a11/xCT are classified as Nrf2-mediated oxidative stress response genes [31,33]. This specific Nrf2-mediated redox-regulated gene expression pattern defines the microglial phenotype Mox [18], and suggests that BV-2 cells switch from the inflammatory phenotype M1 in LPS-activated microglia to the Nrf2-mediated Mox in CBD-treated cells.
CBD was found to strongly upregulate miR-34a in resting and in LPS-treated cells. MiR-34a has multiple roles including regulation of cell cycle, apoptosis, differentiation and migration, and its expression is controlled by p53 [73–74]. Accordingly, miR-34a has many potential target genes, among them, the Notch receptors and their ligands, associated with innate immunity and inflammation through Notch-TLR crosstalk [75]. LPS upregulates Notch1 and Notch2 mRNA expressions. In addition, we show that LPS upregulates Notch ligand Dll1 mRNA expression, in line with the findings that activation of microglia or macrophages with TLR4 ligands leads to induction of Notch receptors and their ligands, including Dll1, Dll4 and Jagged1 [75–78]. Our results demonstrate that LPS modulates Notch signaling by inducing the expression of Dll1. In addition, CBD and THC reduce the expression of LPS-upregulated Dll1 showing an effect of cannabinoids on Notch signaling. As CBD+LPS upregulates miR-34a expression and reduces its target gene Dll1, we suggest that miR-34a might be involved in the modulation of Notch signaling pathway, leading to reduced inflammation.
Interestingly, a cross-talk between Notch and Nrf2 signaling has been reported [79], and miR-34a has been demonstrated to be involved in the modulation of the Nrf2 anti-oxidant pathway [72]. In this regard, cellular redox homeostasis in BV-2 microglial cells, might be regulated by redox sensitive miRNAs, including miR-155 and miR-34a, through their modulation of Nrf2-driven gene expression, upregulated by CBD [31,33].
Fig 8 shows other miR-34a potential target genes, specially a group involved in the RB/E2F cell cycle. We found in our RNAseq data, several target genes related to cell cycle regulation, including Ccnd1, Ccne2, Cdk6, Cdk1a/p21, E2F5 and Cdc25a, previously shown by us, to be affected by CBD and/or LPS and their combination [31,33]. Ccnd1, Ccne2, Cdk1a/p21, E2f5 and Cdk6 are involved in the G1/S transition via the Rb-E2F signaling pathway, and Cdc25a plays a role in assisting both G1/S and G2/M progressions. We show here that CBD downregulates the mRNA expression of Ccnd1 (50%, p<0.0001), Ccne2 (50%, p<0.0001) and E2f5 (50%, p<0.001), as well as of the dual phosphatase Cdc25a (40%, p<0.0001). All together, CBD could induce a cell cycle arrest in the G1 phase leading to inhibition of the E2F pathway [80–81]. In addition, CBD upregulates Cdkn1a/p21 (3.3-fold, p<0.001), that inhibits the activity of cyclinD-Cdk4, preventing Rb phosphorylation during G1 to S phase transition. These results confirm that CBD plays a role in cell cycle arrest at the level of the G1/S phase transition. Moreover, it has been reported that regulation of the G1/S transition through modulation of Ccnd1 expression represents a central mechanism through which NF-κB/IKK signaling regulates cell growth and proliferation [82]. Our data thus suggests that CBD inhibits proliferation through G1/S phase arrest (involving Ccnd1 downregulation and Cdkn1a/p21 upregulation), and that this CBD-inhibitory effect possibly depends on the NF-κB signaling transduction pathway.
It is important to note here that many of the identified miRNA:mRNA interactions do not obey the canonical repression rule, as pointed out by others [14, 68]. For example, Irak1/2 (a target of miR-146) as well as Socs1, Cebpb and IkbkƐ (targets of miR-155) are upregulated instead of repressed, as would be expected. In this regard, it is accepted that each miRNA has many targets and that each mRNA can be regulated by more than one miRNA, suggesting a more complex regulatory mechanism than an ordinary direct repression of a target gene [83].
In conclusion, the present miRNA expression profiling highlights the role of several miRNAs in modulating inflammatory networks in LPS-activated microglia. Here we show that CBD anti-inflammatory effects are mediated via reduction in the expression of some LPS-activated miRNAs, by targeting NF-κB and IRF3 pathways, as well as by reducing LPS-upregulated Dll1 expression, involved in Notch-TLR crosstalk signaling. CBD and CBD+LPS effects are found to be linked to targets modulated by miR-146a, miR-155 and miR-34a, especially genes related to inflammation (Il6 and Ifnb1), Notch signaling (Dll1), cell cycle regulation (Ccnd1 and Cdkn1a/p21), and Nrf2-mediated oxidative stress response (Hmox and Bach1). All together, miRNAs affected by CBD and less so by THC, show to be linked to inflammatory pathways, cell cycle arrest and Nrf2-mediated cellular stress.
Supporting information
(PDF)
(PDF)
(PDF)
Data Availability
All RNAseq data are available from the Gene Expression Omnibus public repository (accession number GSE123262); https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE123262.
Funding Statement
This work was supported by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation to ZV. AJ was supported by the Israeli Ministry for Absorption in Science. We acknowledge the support of the NINDS Informatics Center for Neurogenetics and Neurogenomics (P30 NS062691) to GC. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
References
- 1.Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004. January 23;116(2):281–97. [DOI] [PubMed] [Google Scholar]
- 2.Bartel DP. MicroRNAs: Target recognition and regulatory functions. Cell. 2009. January 23;136(2):215–33. 10.1016/j.cell.2009.01.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Winter J, Jukng S, Keller S, Gregory RI, Diederich S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol. 2009. March;11(3):228–34. 10.1038/ncb0309-228 [DOI] [PubMed] [Google Scholar]
- 4.O'Neill LA, Sheedy FJ, McCoy CE. MicroRNAs: the fine-tuners of Toll-like receptor signalling. Nat Rev Immunol. 2011. March;11(3):163–75. 10.1038/nri2957 Epub 2011 Feb 18. [DOI] [PubMed] [Google Scholar]
- 5.Chen Y, Fu LL, Wen X, Liu B, Huang J, Wang JH, et al. Oncogenic and tumor suppressive roles of microRNAs in apoptosis and autophagy. Apoptosis. 2014. August;19(8):1177–89. 10.1007/s10495-014-0999-7 [DOI] [PubMed] [Google Scholar]
- 6.Zhang L, Huang J, Yang N, Greshock J, Megraw MS, Giannakakis A, et al. MicroRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci U S A. 2006. June 13;103(24):9136–41. Epub 2006 Jun 5. 10.1073/pnas.0508889103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Davidson-Moncada J, Papavasiliou FN, Tam W. MicroRNAs of the immune system. Roles in inflammation and cancer. Ann N Y Acad Sci. 2010. January;1183:183–94. 10.1111/j.1749-6632.2009.05121.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bhalala OG, Srikanth M, Kessler JA. The emerging roles of microRNAs in CNS injuries. Nat Rev Neurol. 2013. June;9(6):328–39. 10.1038/nrneurol.2013.67 Epub 2013 Apr 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ma Y, Wang J, Wang Y, Yang G-Y. The biphasic function of microglia in ischemic stroke. Prog Neurobiol. 2017. October;157:247–272. 10.1016/j.pneurobio.2016.01.005 Epub 2016 Feb 2. [DOI] [PubMed] [Google Scholar]
- 10.Quinn SR, O'Neill LA. A trio of microRNAs that control Toll-like receptor signaling. Int Immunol. 2011. July;23(7):421–5. 10.1093/intimm/dxr034 Epub 2011 Jun 6. [DOI] [PubMed] [Google Scholar]
- 11.Yang L, Seki E. Toll-like receptors in liver fibrosis: cellular crosstalk and mechanisms. Front Physiol. 2012. May 22;3:138 10.3389/fphys.2012.00138 eCollection 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.O'Neill LA, Bowie AG. The family of five: TIR-domain-containing adaptors in Toll-like receptor signalling. Nat Rev Immunol. 2007. May;7(5):353–64. 10.1038/nri2079 [DOI] [PubMed] [Google Scholar]
- 13.Graff JW, Dickson AM, Clay G, McCaffrey AP, Wilson ME. Identifying functional microRNAs in macrophages with polarized phenotypes. J Biol Chem. 2012. June 22;287(26):21816–25. 10.1074/jbc.M111.327031 Epub 2012 May 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Freilich RW, Woodbury ME, Ikezu T. Integrated expression profiles of mRNA and miRNA in polarized primary murine microglia. PLoS One. 2013. November 11;8(11):e79416 10.1371/journal.pone.0079416 eCollection 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ponomarev ED, Veremeyko T, Weiner HL. MicroRNAs are universal regulators of differentiation, activation, and polarization of microglia and macrophages in normal and diseased CNS. Glia. 2013. January;61(1):91–103. 10.1002/glia.22363 Epub 2012 May 31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cobos Jimenez V, Bradley E, Willemsen AM, van Kampen AHC, Baas F, Kootstra NA. Next-generation sequencing of microRNAs uncovers expression signatures in polarized macrophages. Physiol Genomics. 2014. February 1;46(3):91–103. 10.1152/physiolgenomics.00140.2013 Epub 2013 Dec 10. [DOI] [PubMed] [Google Scholar]
- 17.Prinz M, Priller J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat Rev Neurosci. 2014. May;15(5):300–12. 10.1038/nrn3722 Epub 2014 Apr 9. [DOI] [PubMed] [Google Scholar]
- 18.Kadl A, Meher AK, Sharma PR, Lee MY, Doran AC, Johnstone SR, et al. Identification of a novel macrophage phenotype that develops in response to atherogenic phospholipids via Nrf2. Circ Res. 2010. September 17;107(6):737–46. 10.1161/CIRCRESAHA.109.215715 Epub 2010 Jul 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Orihuela R, McPherson CA, Harry GJ. Microglial M1/M2 polarization and metabolic states. Br J Pharmacol. 2016. February;173(4):649–65. 10.1111/bph.13139 Epub 2015 May 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kogan NM, Mechoulam R. Cannabinoids in health and disease. Dialogues Clin Neurosci. 2007;9(4):413–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hill AJ,William CM, Whalley BJ, Stephens GJ. Phytocannabinoids as novel therapeutic agents in CNS disorders. Pharmacol Ther. 2012. January;133(1):79–97. 10.1016/j.pharmthera.2011.09.002 Epub 2011 Sep 6. [DOI] [PubMed] [Google Scholar]
- 22.Mechoulam R, Hanuš LO, Pertwee R, Howlett AC. Early phytocannabinoid chemistry to endocannabinoids and beyond. Nat Rev Neurosci. 2014. November;15(11):757–64. 10.1038/nrn3811 Epub 2014 Oct 15. [DOI] [PubMed] [Google Scholar]
- 23.Massi P, Valenti M, Solinas M, Parolaro D. Molecular mechanisms involved in the antitumor activity of cannabinoids on gliomas: Role for oxidative stress. Cancers (Basel). 2010. May 26;2(2):1013–26. 10.3390/cancers2021013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rieder SA, Chauhan A, Singh U, Nagarkatti M, Nagarkatti P. Cannabinoid-induced apoptosis in immune cells as a pathway to immunosuppression. Immunobiology. 2010. August;215(8):598–605. 10.1016/j.imbio.2009.04.001 Epub 2009 May 20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sánchez AJ, García-Merino A. Neuroprotective agents: Cannabinoids. Clin Immunol. 2012. January;142(1):57–67. 10.1016/j.clim.2011.02.010 Epub 2011 Mar 21 [DOI] [PubMed] [Google Scholar]
- 26.Klein TW, Cabral GA. Cannabinoid-induced immune suppression and modulation of antigen-presenting cells. J Neuroimmune Pharmacol. 2006. March;1(1):50–64. 10.1007/s11481-005-9007-x [DOI] [PubMed] [Google Scholar]
- 27.Nagarkatti P, Pandey R, Rieder SA, Hedge VL, Nagarkatti M. Cannabinoids as novel anti-inflammatory drugs. Future Med Chem. 2009. October;1(7):1333–49. 10.4155/fmc.09.93 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kozela E, Pietr M, Juknat A, Rimmerman N, Levy R, Vogel Z. Cannabinoids Δ9-tetrahydrocannabinol and cannabidiol differentially inhibit the lipopolysaccharide-activated NF-κB and interferon-β/STAT proinflammatory pathways in BV-2 microglial cells. J Biol Chem. 2010. January 15;285(3):1616–26. 10.1074/jbc.M109.069294 Epub 2009 Nov 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kozela E, Lev N, Kaushansky N, Eilam R, Rimmerman N, Levy R, et al. Cannabidiol inhibits pathogenic T cells, decreases spinal microglial activation and ameliorates multiple sclerosis-like disease in C57BL/6 mice. Br J Pharmacol. 2011. August;163(7):1507–19. 10.1111/j.1476-5381.2011.01379.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kozela E, Juknat A, Kaushansky N, Rimmerman N, Ben-Nun A, Vogel Z. Cannabinoids decrease the Th17 inflammatory autoimmune phenotype. J Neuroimmune Pharmacol. 2013. December;8(5):1265–76. 10.1007/s11481-013-9493-1 Epub 2013 Jul 28. [DOI] [PubMed] [Google Scholar]
- 31.Juknat A, Pietr M, Kozela E, Rimmerman N, Levy R, Coppola G, et al. Differential transcriptional profiles mediated by exposure to the cannabinoids cannabidiol and Δ9–tetrahydrocannabinol in BV-2 microglial cells. Br J Pharmacol. 2012a. April;165(8):2512–28. 10.1111/j.1476-5381.2011.01461.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Juknat A, Rimmerman N, Levy R, Vogel Z, Kozela E. Cannabidiol affects the expression of genes involved in zinc homeostasis in BV-2 microglial cells. Neurochem Int. 2012b. November;61(6):923–30. 10.1016/j.neuint.2011.12.002 Epub 2011 Dec 9. [DOI] [PubMed] [Google Scholar]
- 33.Juknat A, Pietr M, Kozela E, Rimmerman N, Levy R, Gao F, et al. Microarray and pathway analysis reveal distinct mechanisms underlying cannabinoid-mediated modulation of LPS-induced activation of BV-2 microglial cells. PLoS One. 2013. April 24;8(4):e61462 10.1371/journal.pone.0061462 Print 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chiurchiu V, Leuti A, Maccarrone M. Cannabinoid signaling and neuroinflammatory diseases: A melting pot for the regulation of brain immune responses. J Neuroimmune Pharmacol. 2015. June;10(2):268–80. 10.1007/s11481-015-9584-2 Epub 2015 Jan 20. [DOI] [PubMed] [Google Scholar]
- 35.Thomas A, Baillie GL, Phillips AM, Razdan RK, Ross RA, Pertwee RG. Cannabidiol displays unexpectedly high potency as an antagonist of CB1 and CB2 receptor agonists in vitro. Br J Pharmacol. 2007. March;150(5):613–23. Epub 2007 Jan 22. 10.1038/sj.bjp.0707133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Laprairie RB, Bagher AM, Kelly MEM, Dupré DJ, Denovan-Wright EM. Type I cannabinoid receptor ligands display functional selectivity in a cell culture model of striatal medium spiny projection neurons. J Biol Chem. 2014. September 5;289(36):24845–62. 10.1074/jbc.M114.557025 Epub 2014 Jul 18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Laprairie RB, Bagher AM, Kelly MEM, Denovan-Wright EM. Cannabidiol is a negative allosteric modulator of the cannabinoid CB1 receptor. Br J Pharmacol. 2015. October;172(20):4790–805. 10.1111/bph.13250 Epub 2015 Oct 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Booz GW. Cannabidiol as an emergent therapeutic strategy for lessing the impact of inflammation on oxidative stress. Free Radic Biol Med. 2011. September 1;51(5):1054–61. 10.1016/j.freeradbiomed.2011.01.007 Epub 2011 Jan 14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kozela E, Juknat A, Gao F, Kaushansky N, Coppola G, Vogel Z. Pathways and gene networks mediating the regulatory effects of cannabidiol, a nonpsychoactive cannabinoid, in autoimmune T cells. J Neuroinflammation. 2016. June 3;13(1):136 10.1186/s12974-016-0603-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kozela E, Juknat A, Vogel Z. Modulation of astrocyte activity by cannabidiol, a nonpsychoactive cannabinoid. Int J Mol Sci. 2017. July 31;18(8). pii: E1669 10.3390/ijms18081669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Massi P, Solinas M, Cinquina V, Parolaro D. Cannabidiol as potential anticancer drug. Br J Clin Pharmacol. 2013. February;75(2):303–12. 10.1111/j.1365-2125.2012.04298.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Burstein S. Cannabidiol (CBD) and its analogs: a review of their effects on inflammation. Bioorg Med Chem. 2015. April 1;23(7):1377–85. 10.1016/j.bmc.2015.01.059 Epub 2015 Feb 7. [DOI] [PubMed] [Google Scholar]
- 43.Ibeas Bih C, Chen T, Nunn AVW, Bazelot M, Dallas M, Whalley BJ. Molecular targets of cannabidiol in neurological disorders. Neurotherapeutics. 2015. October;12(4):699–730. 10.1007/s13311-015-0377-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Pisanti S, Malfitano AM, Ciaglia E, Lamberti A, Ranieri R, Cuomo G, et al. Cannabidiol: State of the art and new challenges for therapeutic applications. Pharmacol Ther. 2017. July;175:133–150. 10.1016/j.pharmthera.2017.02.041 Epub 2017 Feb 22. [DOI] [PubMed] [Google Scholar]
- 45.De Petrocellis L, Di Marzo V. Non-CB1, non-CB2 receptors for endocannabinoids, plant cannabinoids and synthetic cannabimimetics: focus on G-protein-coupled receptors and transient receptor potential channels. J Neuroimmune Pharmacol. 2010. March;5(1):103–21. 10.1007/s11481-009-9177-z Epub 2009 Oct 22. [DOI] [PubMed] [Google Scholar]
- 46.Rimmerman N, Ben-Hail D, Porat Z, Juknat A, Kozela E, Daniels MP, et al. Direct modulation of the outer mitochondrial membrane channel, voltage-dependent anion channel 1 (VDAC1) by cannabidiol: a novel mechanism for cannabinoid-induced cell death. Cell Death Dis. 2013. December 5;4:e949 10.1038/cddis.2013.471 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.McPartland JM, Duncan M, Di Marzo V, Pertwee RG. Are cannabidiol and Δ9-tetrahydrocannabivarin negative modulators of the endocannabinoid system? A systematic review. Br J Pharmacol. 2015. February;172(3):737–53. 10.1111/bph.12944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Deutsch DG. A personal retrospective: Elevating anandamide (AEA) by targeting fatty acid amide hydrolase (FAAH) and the fatty acid binding proteins (FABPs). Front Pharmacol. 2016. October 13;7:370. eCollection 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hegde VL, Tomar S, Jackson A, Rao R, Yang X, Singh UP, et al. Distinct microRNA expression profile and targeted biological pathways in functional myeloid-derived suppressor cells induced by Δ9-tetrahydrocannabinol in vivo: regulation of CCAAT/enhancer-binding protein α by microRNA-690. J Biol Chem. 2013. December 27;288(52):36810–26. 10.1074/jbc.M113.503037 Epub 2013 Nov 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Rao R, Nagarkatti PS, Nagarkatti M. Δ9 Tetrahydrocannabinol attenuates Staphylococcal enterotoxin B-induced inflammatory lung injury and prevents mortality in mice by modulation of miR-17-92 cluster and induction of T-regulatory cells. Br J Pharmacol. 2015. April;172(7):1792–806. 10.1111/bph.13026 Epub 2015 Feb 10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sido JM, Jackson AR, Nagarkatti PS, Nagarkatti M. Marijuana-derived Δ-9-tetrahydrocannabinol suppresses Th1/Th17 cell-mediated delayed-type hypersensitivity through microRNA regulation. J Mol Med (Berl). 2016. September;94(9):1039–51. 10.1007/s00109-016-1404-5 Epub 2016 Apr 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yang X, Bam M, Nagarkatti PS, Nagarkatti M. RNA-seq analysis of delta-9-tetrahydrocannabinol-treated T cells reveals altered gene expression profiles that regulate immune response and cell proliferation. J Biol Chem. 2016. July 22;291(30):15460–72. 10.1074/jbc.M116.719179 Epub 2016 Jun 6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chandra LC, Kumar V, Torben W, Vande Stouwe C, Winsauer P, Amedee A, et al. Chronic administration of Δ9-tetrahydrocannabinol induces intestinal anti-inflammatory microRNA expression during acute simian immunodeficiency virus infection of rhesus macaques. J Virol. 2015. January 15;89(2):1168–81. 10.1128/JVI.01754-14 Epub 2014 Nov 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Simon L, Song K, Vande Stouwe C, Hollenbach A, Amedee A, Mohan M, et al. Δ9-tetrahydrocannabinol (Δ9-THC) promotes neuroimmune-modulatory microRNA profile in striatum of simian immunodeficiency virus (SIV)-infected macaques. J Neuroimmune Pharmacol. 2016. March;11(1):192–213. 10.1007/s11481-015-9645-6 Epub 2015 Nov 25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Auer PL, Doerge RW. Statistical design and analysis of RNA sequencing data. Genetics. 2010. June;185(2):405–16. 10.1534/genetics.110.114983 Epub 2010 May 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010. January 1;26(1):139–40. 10.1093/bioinformatics/btp616 Epub 2009 Nov 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Betel D, Wilson M, Gabow A, Marks DS, Sander C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 2008. January;36(Database issue):D149–53. Epub 2007 Dec 23. 10.1093/nar/gkm995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Peterson SM, Thompson JA, Ufkin ML, Sathyanarayana P, Liaw L, Congdon CB. Common features of microRNA target prediction tools. Front Genet. 2014. February 18;5:23 10.3389/fgene.2014.00023 eCollection 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Necela BM, Su W, Thompson EA. Toll-like receptor 4 mediates cross-talk between peroxisome proliferator-activated receptor gamma and nuclear factor-kappaB in macrophages. Immunology. 2008. November;125(3):344–58. 10.1111/j.1365-2567.2008.02849.x Epub 2008 Apr 18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.O'Sullivan SE, Sun Y, Bennett AJ, Randall MD, Kendall DA. Time-dependent vascular actions of cannabidiol in the rat aorta. Eur J Pharmacol. 2009. June 10;612(1–3):61–8. 10.1016/j.ejphar.2009.03.010 Epub 2009 Mar 11. [DOI] [PubMed] [Google Scholar]
- 61.Lize M, Klimke A, Dobbelstein M. MicroRNA-449 in cell fate determination. Cell Cycle. 2011. September 1;10(17):2874–82. Epub 2011 Sep 1. [DOI] [PubMed] [Google Scholar]
- 62.Bao J, Li D, Wang L, Wu J, Hu Y, Wang Z, et al. MicroRNA-449 and microRNA-34b/c function redundantly in murine testes by targeting E2F transcription factor-retinoblastoma protein (E2F-pRb) pathway. J Biol Chem. 2012. June 22;287(26):21686–98. 10.1074/jbc.M111.328054 Epub 2012 May 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol. 2007. August 7;17(15):1298–307. Epub 2007 Jul 26. 10.1016/j.cub.2007.06.068 [DOI] [PubMed] [Google Scholar]
- 64.Bhaumik D, Scott GK, Schokrpur S, Patil CK, Campisi J, Benz CC. Expression of microRNA-146 suppresses NF-kappaB activity with reduction of metastatic potential in breast cancer cells. Oncogene. 2008. September 18;27(42):5643–7. 10.1038/onc.2008.171 Epub 2008 May 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Taganov KD, Boldin MP, Chang K-J, Baltimore D. NF-kB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune response. Proc Natl Acad Sci U S A. 2006. August 15;103(33):12481–6. Epub 2006 Aug 2. 10.1073/pnas.0605298103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Cardoso AL, Guedes JR, de Lima MC. Role of microRNAs in the regulation of innate immune cells under neuroinflammatory conditions. Curr Opin Pharmacol. 2016. February;26:1–9. 10.1016/j.coph.2015.09.001 Epub 2015 Sep 25. [DOI] [PubMed] [Google Scholar]
- 67.Jablonski KA, Gaudet AD, Amici SA, Popovich PG, Guerau-de-Arellano M. Control of the Inflammatory Macrophage Transcriptional Signature by miR-155. PLoS One. 2016. July 22;11(7):e0159724 10.1371/journal.pone.0159724 eCollection 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Amici SA, Dong J, Guerau-de-Arellano M. Molecular mechanisms modulating the phenotype of macrophages and microglia. Front Immunol. 2017. November 10;8:1520 10.3389/fimmu.2017.01520 eCollection 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Cardoso AL, Guedes JR, Pereira de Almeida L, Pedroso de Lima MC. miR-155 modulates microglia-mediated immune response by down-regulating SOCS-1 and promoting cytokine and nitric oxide production. Immunology. 2012. January;135(1):73–88. 10.1111/j.1365-2567.2011.03514.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Häcker H, Karin M. Regulation and function of IKK and IKK-related kinases. Sci STKE. 2006. October 17;2006(357):re13. [DOI] [PubMed] [Google Scholar]
- 71.Pulkkinen KH, Ylä-Herttuala S, Levonen AL. Heme oxygenase 1 is induced by miR-155 via reduced BACH1 translation in endothelial cells. Free Radic Biol Med. 2011. 51: 2124–2131. 10.1016/j.freeradbiomed.2011.09.014 Epub 2011 Sep 17. [DOI] [PubMed] [Google Scholar]
- 72.Cheng X, Ku CH, Siow RC. Regulation of the Nrf2 antioxidant pathway by microRNAs: New players in micromanaging redox homeostasis. Free Radic Biol Med. 2013. September;64:4–11. 10.1016/j.freeradbiomed.2013.07.025 Epub 2013 Jul 21. Review. [DOI] [PubMed] [Google Scholar]
- 73.Hermeking H. The miR-34 family in cancer and apoptosis. Cell Death Differ. 2010. February;17(2):193–9. 10.1038/cdd.2009.56 Epub 2009 May 22. [DOI] [PubMed] [Google Scholar]
- 74.Chen F, Hu S-J. Effect of microRNA-34a in cell cycle, differentiation and apoptosis: A review. J Biochem Mol Toxicol. 2012. February;26(2):79–86. 10.1002/jbt.20412 Epub 2011 Dec 12. [DOI] [PubMed] [Google Scholar]
- 75.Shang Y, Smith S, Hu X. Role of Notch signaling in regulating innate immunity and inflammation in health and disease. Protein Cell. 2016. March;7(3):159–74. 10.1007/s13238-016-0250-0 Epub 2016 Mar 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Cao Q, Lu J, Kaur C, Sivakumar V, Li F, Cheah PS, et al. Expression of Notch-1 receptor and its ligands Jagged-1 and Delta-1 in amoeboid microglia in postnatal rat brain and murine BV-2 cells. Glia. 2008. August 15;56(11):1224–37. 10.1002/glia.20692 [DOI] [PubMed] [Google Scholar]
- 77.Cao Q, Li P, Lu J, Dheen ST, Kaur C, Ling EA. Nuclear factor-κB/p65 responds to changes in the Notch signaling pathway in murine BV-2 cells and in amoeboid microglia in postnatal rats treated with the γ-secretase complex blocker DAPT. J Neurosci Res. 2010. September;88(12):2701–14. 10.1002/jnr.22429 [DOI] [PubMed] [Google Scholar]
- 78.Zhang Q, Wang C, Liu Z, Liu X, Han C, Cao X, et al. Notch signal suppresses Toll-like receptor-triggered inflammatory responses in macrophages by inhibiting extracellular signal-regulated kinase ½-mediated nuclear factor κB activation. J Biol Chem. 2012. February 24;287(9):6208–17. 10.1074/jbc.M111.310375 Epub 2011 Dec 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wakabayashi N, Chartoumpekis DV, Kensler TW. Crosstalk between Nrf2 and Notch signaling. Free Radic Biol Med. 2015. November;88(Pt B):158–167. 10.1016/j.freeradbiomed.2015.05.017 Epub 2015 May 21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Trimarchi JM, Lees JA. Sibling rivalry in the E2F family. Nat Rev Mol Cell Biol. 2002; 3(1): 11–20. 10.1038/nrm714 /DOI: [DOI] [PubMed] [Google Scholar]
- 81.Shen T, Huang S. The role of Cdc25a in the regulation of cell proliferation and apoptosis. Anticancer Agents Med Chem. 2012. July;12(6):631–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Ledoux AC, Perkins ND. NF-κB and the cell cycle. Biochem Soc Trans. 2014. February;42(1):76–81. 10.1042/BST20130156 [DOI] [PubMed] [Google Scholar]
- 83.Baumjohann D, Ansel KM. MicroRNA-mediated regulation of T helper cell differentiation and plasticity. Nat Rev Immunol. 2013. September;13(9):666–78. 10.1038/nri3494 Epub 2013 Aug 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(PDF)
(PDF)
(PDF)
Data Availability Statement
All RNAseq data are available from the Gene Expression Omnibus public repository (accession number GSE123262); https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE123262.