Abstract
The bacterial Min protein system was encapsulated in giant unilamellar vesicles (GUVs). Using confocal fluorescence microscopy, we identified several distinct modes of spatiotemporal patterns inside spherical GUVs. For osmotically deflated GUVs, the vesicle shape actively changed in concert with the Min oscillations. The periodic relocation of Min proteins from the vesicle lumen to the membrane and back is accompanied by drastic changes in the mechanical properties of the lipid bilayer. In particular, two types of oscillating membrane‐shape changes are highlighted: 1) GUVs that repeatedly undergo fission into two connected compartments and fusion of these compartments back into a dumbbell shape and 2) GUVs that show periodic budding and subsequent merging of the buds with the mother vesicle, accompanied by an overall shape change of the vesicle reminiscent of a bouncing ball. These findings demonstrate how reaction–diffusion‐based protein self‐organization can directly yield visible mechanical effects on membrane compartments, even up to autonomous division, without the need for coupling to cytoskeletal elements.
Keywords: cell division, liposomes, membranes, synthetic biology, vesicles
Creating synthetic materials that imitate properties of living organisms is a persistent challenge in science that often leads to technological advances and furthers our understanding of biological systems. One of the most prominent attributes of many living organisms is their motility. While great innovations have come from mimicking life on a macroscopic level,1 there are few artificial systems that exhibit autonomous motion on a cellular scale.2 Previously, cell‐like vesicles were created from synthetic components that exhibit oscillatory dynamic behavior driven by chemical reactions and osmotic effects.3 In recent years, first attempts were made to encapsulate biological building blocks in lipid vesicles to create structures capable of more distinct actuation.4 Herein, we present cell‐sized GUVs that autonomously and reversibly change their shape in response to the oscillatory, membrane‐interacting Min protein system.
The Min system consists of the proteins MinC, MinD, and MinE, and positions the bacterial cell division machinery in Escherichia coli.5 Based on a reaction–diffusion mechanism, the proteins MinD and MinE oscillate between the cell poles and thereby spatially regulate the assembly of the cell division complex, restricting the formation of the division ring to the cell center.6 MinD dimerizes upon adenosine triphosphate (ATP) binding, which enhances its affinity to the membrane. Membrane‐bound MinD recruits MinE, which in turn stimulates the ATPase activity of MinD, causing both proteins to detach again (see the Supporting Information, Figure S1).7
In previous work, we reconstituted MinDE protein self‐organization in vitro on supported lipid bilayers (SLBs), reproducing oscillatory patterns on a planar membrane and rendering the Min system an attractive biological model to study reaction–diffusion dynamics in 2D.8 In further research, Min protein patterns were confined within SLB‐lined microfluidic scaffolds to probe geometry sensing and recreate spatial properties of bacterial cells.9 However, these non‐deformable microstructures lack many features of living cells, in particular their plasticity and ability to ultimately divide. In a continuing attempt to confine the Min system to more cell‐like compartments, we recently fully enclosed the proteins in microdroplets and were able to observe pole‐to‐pole oscillations on the lipid monolayer at the water–oil interface, similar to the oscillations seen in E. coli.10 Unfortunately, owing to the surface tension at the water–oil interphase, microdroplets are still very rigid compartments, and the difference in the refractive indices between the two phases hampers confocal imaging of the full 3D volume of the droplets.
In spite of being conceptually straightforward, the Min oscillations had not been enclosed within a free‐standing lipid bilayer up to now, owing to several technical challenges. Herein, we report the successful encapsulation of the Min system in giant liposomes, finally leading to fully confined Min oscillations in mechanically transformable compartments.
We adapted the cDICE method,11 an emulsion transfer technique that allowed us to encapsulate purified MinD (50 % eGFP‐MinD) and MinE proteins in GUVs with negatively charged membranes (1,2‐dioleoyl‐sn‐glycero‐3‐phosphocholine (DOPC) and 1,2‐dielaidoyl‐sn‐glycero‐3‐phospho‐(1′‐rac‐glycerol) (DOPG) in a ratio of 4:1;12 Figure 1). We slightly modified the original procedure, used a customized 3D‐printed rotating chamber and larger capillaries, and dispersed the lipids in oil as aggregates,13 rather than in solution. More details about vesicle generation and protein encapsulation can be found in the Supporting Information. The generated vesicles are between 5 μm and 100 μm in diameter (Figure S2). While we detected eGFP‐labeled MinD in almost all vesicles, the fraction of GUVs exhibiting dynamic behavior varied from 50 % to 100 % between samples. After 4–5 hours, the Min oscillations began to fade owing to protein aggregation and a general decrease in protein activity.
Figure 1.

Encapsulation of MinD and MinE into giant unilamellar vesicles. A) Schematic depiction of the vesicle generation process. The aqueous protein solution is injected into a rotating chamber through a glass capillary. Droplets form at the capillary tip in the oil phase, which contains lipid aggregates. The droplets then pass through a water–oil interphase lined with lipids, forming the GUVs. B) Representative confocal image of several GUVs containing oscillating Min proteins. C) Confocal images showing the two states of a protein oscillation. Composite of fluorescence signals and the differential interference contrast (DIC) channel. eGFP‐MinD in cyan and DOPE‐ATTO655 in orange.
Sequential confocal images of the equatorial plane of the GUVs reveal several distinct modes of Min oscillations. Consistent with previous observations for Min dynamics within microdroplets,10 we observed two types of standing wave patterns and circling traveling waves (Figure 2 A–I). We further identified a second mode of traveling waves (Figure 2 J–L) and a qualitatively different type of pole‐to‐pole oscillations, which can be asymmetric and appeared to be linked to the aggregation of protein in a ring‐like structure that sets the boundary between the two poles (Figure S3 and Movie S3).
Figure 2.
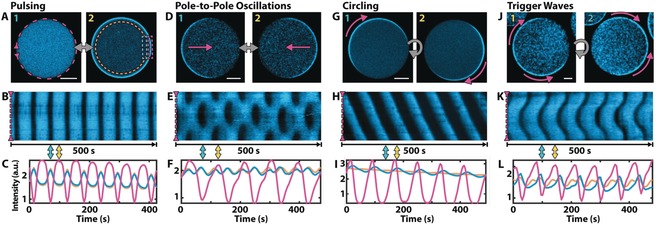
Different modes of Min oscillations in GUVs. A–C) Pulsing oscillations: MinD oscillates between the inside of the vesicles and the inner membrane surface. Dotted lines in (A) indicate ROIs for kymograph and intensity plots. D–F) Pole‐to‐pole oscillations: A protein binds alternately to the membrane of the two hemispheres of the vesicle. A second type of pole‐to‐pole oscillation is shown in Figure S3. G–I) Circling waves: Traveling waves that continuously revolve on the inside surface of the GUV. J–L) Trigger waves: Traveling waves that originate and terminate on opposing poles; here the wave origin is on the left side of the vesicle. The top panels [(A), (D), (G), and (J)] show two frames from a time series of confocal images of each oscillation mode. The fluorescence signal of eGFP‐MinD is shown in cyan. Magenta arrows indicate the directions of the waves. Scale bars: 10 μm. The center panels [(B), (E), (H), and (K)] show kymographs along the circumference of each vesicle [indicated as a magenta dotted line in (A)(1)]. The bottom panels [(C), (F), (I), and (L)] show average intensities in certain ROIs for each vesicle. ROIs are indicated by the dotted circle and boxes in (A)(2) in the respective colors of the curves. Magenta: Intensity of a membrane section (normalized); blue: intensity of the lumen close to the membrane section; orange: average intensity of entire lumen. Movie S1 shows all oscillation modes sequentially.
GUVs are ideal for confocal fluorescence microscopy and therefore allowed us to compile time series of z‐stacks of entire vesicles (Movies S2–S4), enabling the observation of reaction–diffusion dynamics on a closed 3D surface (here a sphere). Spatiotemporal reaction–diffusion patterns on closed surfaces are ubiquitous in nature and have been studied extensively in theory,14 but the only experimental record of a 3D observation thereof under controlled conditions is on catalyst‐coated millimeter‐sized beads in a Belousov–Zhabotinsky reaction solution on which spiral waves (similar to Figure 2 G–I) and trigger waves (similar to Figure 2 J–L) were observed.15
Often we see one mode transitioning into another (Movies S4 and S5 and Figure S3 D). In fact, none of the oscillation modes seemed to be temporally stable, with the exception of the pulsing mode. We observed vesicles transitioning between as many as three different modes before settling into the pulsing mode. As a general trend, the traveling wave modes (Figure 2 G–L) seemed to be the shortest lived, switching into standing wave oscillations (Figure 2 A–F) within minutes. Even though a single vesicle can exhibit different types of oscillations, our experiments indicate that vesicle size, protein concentrations, and other factors heavily affect the type of oscillations that a vesicle exhibits. Further experiments will be required to map out this large parameter space.
In addition to the spatiotemporal patterns within the spherical vesicles, we observed an unexpected effect of the protein oscillations on the morphology of the GUVs. Under hypertonic stress, vesicles that were visibly osmotically deflated (i.e., not spherical) underwent extensive shape fluctuations in concert with the MinDE oscillations.
A number of different membrane shape oscillations were observed; the two most prevalent types are shown in Figure 3. In the first case, referred to as periodic dumbbell splitting (Figure 3 A–C), the deflated vesicles assume a dumbbell‐like shape when the Min proteins are bound to the membrane. With the relocation of the Min proteins to the lumen, the “dumbbell” splits into two spherical compartments (connected by a narrow neck), before these fuse back together when the proteins bind to the membrane again. Figure 3 B shows the narrow membrane neck between the two compartments whilst split. In Figure 3, we chose a different example of a vesicle for each Figure panel, but movies for all cases are shown in Movie S6, demonstrating consistent behavior and the reproducibility of these observations. The Movie also includes a fourth example, which was recorded as z‐stacks, confirming the separation (by a neck) from a side view.
Figure 3.
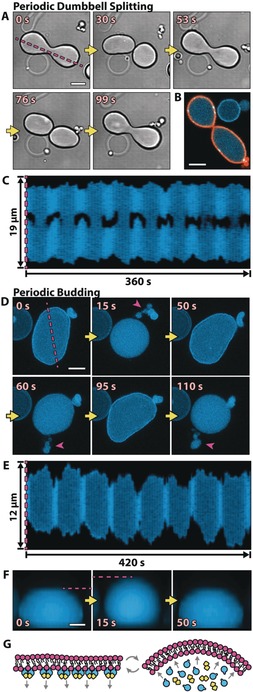
The vesicle shapes change in concert with Min oscillations. A) Time series of DIC images of a dumbbell‐shaped GUV that repetitively splits into two compartments connected by a narrow membrane neck. B) The confocal image of a shape‐oscillating GUV with fluorescently labeled lipids (orange) shows that the GUV does not undergo a full fission into two separate GUVs, but rather that the two compartments are still connected by a membrane neck. C) Kymograph along the line parallel to the axis of rotational symmetry of a dynamic GUV (see the magenta line in (A)). Whenever MinD is in the lumen, the GUV is split into two, almost separate compartments. Movie S6 features all three vesicle examples in (A) to (C). D) A GUV that undergoes periodic budding and subsequent fusion of the buds with the mother vesicle. Buds are highlighted with magenta arrows. Shown are z projections of five confocal planes. E) Kymograph (see the magenta line in (D)) of a repetitively budding vesicle, showing the reduction in size in the x and y dimensions of a GUV whenever budding occurs. F) Sequential images of a side view of a periodically budding vesicle, showing that the reduction in size in the x–y plane is accompanied by an increase in size in the z direction (see also Figure S6 C). Movie S7 shows the vesicles from (D), (E), and (F) sequentially. Scale bars: 5 μm. G) Suspected mechanism: Membrane‐bound MinD increases the surface area of the inner membrane leaflet. Upon protein detachment, the intrinsic curvature of the bilayer increases, and membrane deformations occur.
In the second case, which we refer to as periodic budding (Figure 3 D–F), the relocation of the proteins from the membrane to the lumen and back induces budding and subsequent fusion of the buds with the mother compartment (Figure 3 D). Along with the budding, the mother compartment also contracts in its x–y cross‐section (Figure 3 D, E) and expands in the z dimension (Figure 3 F), taking on an almost spherical shape, like a bouncing ball. Movie S7 includes examples of three different vesicles. Figure S4 shows sequential images of the second example, revealing further details about its shape change cycle. Most notably, the vesicle first switches from a flattened (oblate) to an elongated (prolate) shape before forming a bud and assuming a spherical shape. Similarly, when oscillations are not strong enough to trigger fission or budding, vesicles solely oscillate back and forth between a flattened and an elongated geometry, but do not become spherical (Movie S8).
We only saw extensive shape changes in vesicles that exhibited oscillations of the pulsing type; occasionally, slight deformations can be observed in other cases (e.g., in Movie S5 for the circling wave oscillation). Generally, only a small fraction of vesicles show more extensive membrane dynamics as most vesicles respond to the osmotic stress with the formation of thin membrane tubules and therefore maintain a static spherical shape (Figure S5).
For all cases in which we saw shape fluctuations, a comprehensive correlation between the changes in vesicle geometry and the location of the Min proteins can be identified. The behavior that we observed indicates that the binding of the Min proteins directly affects the membrane properties. A variety of methods have been used to study shape changes in GUVs, such as inducing these by temperature,16 changes in osmolarity,17 insertion of additional lipids into the membrane,18 and through other substances interacting with the membrane.19 In these studies, budding and fission events similar to those that we observed were reported, but they were induced externally and mostly unidirectional and were never repetitive by nature. Here, we present a system that allowed us to observe extensive shape transitions, which are not only reversible, but oscillate back and forth autonomously. Hence, we suggest that these vesicles provide a unique test bed for the study of dynamic membrane transformations in conjunction with peripheral protein association.
The specific shape oscillations that we observed seem to be a result of changes in membrane curvature caused by the periodic relocation of the proteins. The membrane‐inserted amphipathic helix of MinD7b presumably increases the surface area of the inner membrane leaflet and potentially also decreases the intrinsic spontaneous curvature of the membrane. This would result in an increase in the overall membrane curvature when the protein unbinds upon ATP hydrolysis, which would be in accord with the observed changes in vesicle morphology (Figure 3 G). The effects of inserted amphipathic helices on membrane curvature have been investigated for proteins involved in vesicular transport.20 Previous investigations on the interactions of MinD alone with free‐standing bilayers in a static system showed an increase in membrane viscosity21 and even membrane tubulation,22 but further investigations will be required to reveal the precise mechanism on a molecular level.
Dynamics and actuation of lipid vesicles as reported here have, to the best of our knowledge, not been described before. These extreme and rapid shape deformations could prove important for synthetic biology and the growing interest in creating actuation on a microscopic scale by engineering active vesicles or bioinspired “molecular robots”. Lastly, the encapsulation of the Min oscillating system in GUVs represents a crucial and necessary step for the in vitro, bottom‐up reconstitution of bacterial cell division. In this respect, it is particularly exciting that the presumed role of the Min oscillations (i.e., the positioning of the bacterial cytoskeleton to mechanically drive cell division) is, in simple vesicle compartments, accompanied by a distinctive mechanical cue itself. This may point to an even more archetypal function of this fascinating protein system, indicating how cyclic self‐organization of encapsulated biomolecules may result in autonomous division of phospholipid compartments.
Conflict of interest
The authors declare no conflict of interest.
Supporting information
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Acknowledgements
We acknowledge the MPI‐B Biochemistry Core Facility for assistance in protein purification. We thank Saša Svetina, Rumiana Dimova, and Erwin Frey for helpful discussions and Charlotte Kelley for comments on the manuscript. This work is part of the MaxSynBio consortium, which is jointly funded by the Federal Ministry of Education and Research of Germany and the Max Planck Society. We are grateful for the financial support provided by the Gottfried Wilhelm Leibniz‐Program of the DFG (SCHW716/8‐1). B.R. and P.S. acknowledge funding through the DFG Collaborative Research Centre “Spatiotemporal dynamics of bacterial cells” (TRR174/2017). B.R. is supported by a DFG fellowship through the Graduate School of Quantitative Biosciences Munich. We acknowledge support from the Center for NanoScience Munich.
T. Litschel, B. Ramm, R. Maas, M. Heymann, P. Schwille, Angew. Chem. Int. Ed. 2018, 57, 16286.
References
- 1. Taubes G., Science 2000, 288, 80. [DOI] [PubMed] [Google Scholar]
- 2. Hagiya M., Konagaya A., Kobayashi S., Saito H., Murata S., Acc. Chem. Res. 2014, 47, 1681–1690. [DOI] [PubMed] [Google Scholar]
- 3.
- 3a. Tamate R., Ueki T., Shibayama M., Yoshida R., Angew. Chem. Int. Ed. 2014, 53, 11248–11252; [DOI] [PubMed] [Google Scholar]; Angew. Chem. 2014, 126, 11430–11434; [Google Scholar]
- 3b. Oglecka K., Rangamani P., Liedberg B., Kraut R. S., Parikh A. N., eLife 2014, 3, e03695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.
- 4a. Keber F. C., Loiseau E., Sanchez T., DeCamp S. J., Giomi L., Bowick M. J., Marchetti M. C., Dogic Z., Bausch A. R., Science 2014, 345, 1135–1139; [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4b. Sato Y., Hiratsuka Y., Kawamata I., Murata S., Nomura S.-i. M., Science Robotics 2017, 2, eaal3735; [DOI] [PubMed] [Google Scholar]
- 4c. Tanaka S., Takiguchi K., Hayashi M., Commun. Phys. 2018, 1, 18. [Google Scholar]
- 5. de Boer P. A., Crossley R. E., Hand A. R., Rothfield L. I., EMBO J. 1991, 10, 4371–4380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.
- 6a. Raskin D. M., de Boer P. A., Proc. Natl. Acad. Sci. USA 1999, 96, 4971–4976; [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6b. Hu Z., Lutkenhaus J., Mol. Microbiol. 1999, 34, 82–90. [DOI] [PubMed] [Google Scholar]
- 7.
- 7a. Hu Z., Lutkenhaus J., Mol. Cell 2001, 7, 1337–1343; [DOI] [PubMed] [Google Scholar]
- 7b. Hu Z., Lutkenhaus J., Mol. Microbiol. 2003, 47, 345–355. [DOI] [PubMed] [Google Scholar]
- 8.
- 8a. Loose M., Fischer-Friedrich E., Ries J., Kruse K., Schwille P., Science 2008, 320, 789–792; [DOI] [PubMed] [Google Scholar]
- 8b. Schweizer J., Loose M., Bonny M., Kruse K., Monch I., Schwille P., Proc. Natl. Acad. Sci. USA 2012, 109, 15283–15288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.
- 9a. Zieske K., Schwille P., Angew. Chem. Int. Ed. 2013, 52, 459–462; [DOI] [PubMed] [Google Scholar]; Angew. Chem. 2013, 125, 477–481; [Google Scholar]
- 9b. Caspi Y., Dekker C., eLife 2016, 5, e19271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Zieske K., Chwastek G., Schwille P., Angew. Chem. Int. Ed. 2016, 55, 13455–13459; [DOI] [PMC free article] [PubMed] [Google Scholar]; Angew. Chem. 2016, 128, 13653–13657. [Google Scholar]
- 11. Abkarian M., Loiseau E., Massiera G., Soft Matter 2011, 7, 4610–4614. [Google Scholar]
- 12. Vecchiarelli A. G., Li M., Mizuuchi M., Mizuuchi K., Mol. Microbiol. 2014, 93, 453–463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.
- 13a. Claudet C., In M., Massiera G., Eur. Phys. J. E 2016, 39, 9; [DOI] [PubMed] [Google Scholar]
- 13b. Loiseau E., Schneider J. A., Keber F. C., Pelzl C., Massiera G., Salbreux G., Bausch A. R., Sci. Adv. 2016, 2, e1500465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.
- 14a. Grindrod P., Gomatam J., J. Math. Biol. 1987, 25, 597–610; [DOI] [PubMed] [Google Scholar]
- 14b. Zykov V. S., Müller S. C., Phys. D 1996, 97, 322–332; [Google Scholar]
- 14c. Matthews P. C., Phys. Rev. E 2003, 67, 036206; [DOI] [PubMed] [Google Scholar]
- 14d. Fuselier E. J., Wright G. B., J. Sci. Comput. 2013, 56, 535–565. [Google Scholar]
- 15. Maselko J., Showalter K., Nature 1989, 339, 609–611. [Google Scholar]
- 16.
- 16a. Berndl K., Käs J., Lipowsky R., Sackmann E., Seifert U., Europhys. Lett. 1990, 13, 659–664; [Google Scholar]
- 16b. Käs J., Sackmann E., Biophys. J. 1991, 60, 825–844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Döbereiner H. G., Käs J., Noppl D., Sprenger I., Sackmann E., Biophys. J. 1993, 65, 1396–1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.
- 18a. Farge E., Devaux P. F., Biophys. J. 1992, 61, 347–357; [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18b. Alam J. M., Yamazaki M., Chem. Phys. Lipids 2011, 164, 166–174; [DOI] [PubMed] [Google Scholar]
- 18c. Mally M., Bozic B., Hartman S. V., Klancnik U., Mur M., Svetina S., Derganc J., RSC Adv. 2017, 7, 36506–36515. [Google Scholar]
- 19.
- 19a. Tanaka T., Tamba Y., Masum S. M., Yamashita Y., Yamazaki M., Biochim. Biophys. Acta Biomembr. 2002, 1564, 173–182; [DOI] [PubMed] [Google Scholar]
- 19b. Yamashita Y., Masum S. M., Tanaka T., Yamazaki M., Langmuir 2002, 18, 9638–9641. [Google Scholar]
- 20.
- 20a. Ford M. G., Mills I. G., Peter B. J., Vallis Y., Praefcke G. J., Evans P. R., McMahon H. T., Nature 2002, 419, 361–366; [DOI] [PubMed] [Google Scholar]
- 20b. Lee M. C., Orci L., Hamamoto S., Futai E., Ravazzola M., Schekman R., Cell 2005, 122, 605–617. [DOI] [PubMed] [Google Scholar]
- 21. Mazor S., Regev T., Mileykovskaya E., Margolin W., Dowhan W., Fishov I., Biochim. Biophys. Acta Biomembr. 2008, 1778, 2496–2504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Zhou H., Lutkenhaus J., J. Bacteriol. 2003, 185, 4326–4335. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary
Supplementary


