Abstract
Background
HCV exhibits a high genetic diversity and is classified into 7 genotypes which are further divided into 86 confirmed subtypes. However, there are multiple isolates with unassigned subtypes. We aimed to amplify and characterize the full-length genome sequence of an HCV genotype 1 (HCV-1) divergent isolate (DE/17–0414) in Germany.
Methods
The HCV infection was detected in an HIV-1-positive German female within an HCV/HIV-coinfection study using a commercially available antigen-antibody HCV ELISA kit and confirmed by an in-house quantitative real-time RT-PCR assay. Preliminary genotyping was done by sequencing and phylogenetic analysis on partial NS5B region. The full-length genome sequence was determined by consensus RT-PCR assays. Resistance-associated substitutions (RASs) were analyzed using the web-based tool Geno2pheno[HCV].
Results
Partial NS5B region of the isolate DE/17–0414 showed more than 95% identity to 73–08460349-1 l and HCV_Fr_003 from France and QC316 from Canada. Full-length genome analysis of the DE/17–0414 strain showed 91.8% identity to QC316 but less than 79.6% to other HCV-1 strains. Phylogenetic analyses demonstrated that DE/17–0414, 73–08460349-1 l, HCV_Fr_003, and QC316 formed a separate subcluster within HCV-1. DE/17–0414 had a distinct 3 amino acids insertion at the N-terminal of hypervariable region 1 (HVR1) within viral envelope glycoprotein 2 (E2) and several potential antiviral RASs among the NS3 and NS5A genes.
Conclusions
We identified and analyzed an HCV-1 divergent isolate derived from an HIV-1 coinfected individual in Germany, which will be assigned to a new HCV-subtype 1o. Our understanding of the origin and transmission dynamics of this new subtype 1o requires further assessments from patients worldwide.
Keywords: Hepatitis C virus, HIV-1, HCV genotype 1 subtypes, Full-length genome, HVR1, RASs
Main text
Hepatitis C virus (HCV) causes both acute and chronic hepatitis. According to the World Health Organization (WHO), in 2015 an estimated number of 71 million people have been chronically HCV-infected globally [1]. Among these, approximately 4 to 5 million individuals are coinfected with HIV [2]. HCV/HIV-coinfections are of major public health concern, as HIV-coinfection is associated with sometimes more serious progression of HCV-infection [2]. Since 2011, direct-acting antivirals (DAAs) for various genotypes of HCV are available for standard-of-care treatment. However, there is a controversial discussion whether HIV-coinfection is associated with worse response to DAA-based therapy against chronic hepatitis C in real life than HCV-monoinfected patients [3, 4] and the occurrence of potential HCV resistance-associated substitutions (RASs) is correlated with treatment failure [5]. Therefore, detection of HIV/HCV-coinfection and monitoring of potential HCV RASs is of clinical importance [6]. HCV is a positive-strand RNA virus with a 9.7 kb single-stranded, messenger-sense RNA genome. HCV exhibits a high genetic diversity; there are 7 genotypes, further sub-divided into 86 confirmed subtypes according to the 10th International Committee on Taxonomy of Viruses (ICTV) report on the taxonomy of the family Flaviviridae [7]. Nonetheless, a number of HCV strains are phylogenetically divergent from previously described sequences, thus can only be classified into genotypes but without subtype assignment [8]. Globally, HCV genotype 1 (HCV-1) is dominant (46.2%) and different genotype/subtype prevalence evolves and correlates to epidemiological factors [9]. In Germany, a recent study reported that HCV-1a (35.9%) and HCV-1b (30.6%) are the most prevalent subtypes, followed by HCV-3 (20.6%) [10].
In this work, we aimed to amplify and characterize the full-length genome sequence of a HCV-1 divergent strain (DE/17–0414) from an HIV-1 coinfected individual from Germany. According to the “Protection against Infection Act” (IfSG; §7) diagnostic laboratories in Germany report new HIV infections anonymously to the Robert Koch Institute (RKI). Approximately 60% of the reports are submitted together with a dried serum spot (DSS) sample prepared from residual blood of the diagnosis. Antibodies and viral RNAs are isolated from these DSS and are used for sentinel studies (according IfSG §13) [11]. Within a sentinel study established at the RKI, HIV/HCV coinfections are analyzed. This includes partial sequencing for the determination of the HCV genotype. HCV-infection was serologically identified using the Monolisa HCV Ag/Ab ULTRA V2 kit (Bio-Rad, Marnes-la-Coquette, France). Viral RNA from DSS was extracted by the automated Nuclisens EasyMag platform (bioMerieux, Capronne, France) following the manufacturer’s instructions. HCV viral load was measured by an in-house quantitative RT-PCR assay targeting the 5′ noncoding region (Table 1). Preliminary HCV genotyping was done by a consensus nested RT-PCR assay targeting a 674 base pair fragment in the NS5B region corresponding to nt position 7962 to 8636 of H77 reference strain. After cDNA synthesis using the Transcriptor first-strand cDNA synthesis kit (Roche Diagnostics, Mannheim, Germany), the complete viral genome was amplified using KAPA HiFi HotStart ReadyMix PCR kit (Kapa Biosystems, Boston, USA) with HCV-1 degenerate and DE/17–0414 specific primers (Table 1). The 5′ and 3′ sequences were determined using 5′ and 3′ rapid amplification of cDNA ends (Roche Diagnostics, Mannheim, Germany). HCV amplicons were sequenced with the BigDye Terminator version 3.1 cycle sequencing kit (Applied Biosystems, California, USA) in both directions. The sequencing chromatograms were checked for overlapping multicolor peaks. Whole-genome sequence was assembled using Geneious software version 10.0.5 (Biomatters, Auckland, New Zealand) [12]. Sequence identity comparisons were performed using the BLASTn search engine (https://blast.ncbi.nlm.nih.gov). Phylogenetic analyses were completed using the Neighbor-Joining method with maximum composite likelihood nucleotide distance between coding regions and complete deletion option in MEGA software version 7 [13], Bootstrapping was performed with 1000 replicates. To identify possible recombination, identity plot and bootscan analyses of full-length sequences were performed in the SimPlot software program version 3.5.1 with a sliding window size of 300 nt and a step size of 15 nt increment [14]. Potential RASs analysis among NS3, NS5A, and NS5B regions were conducted by Geno2pheno[HCV] – a web-based interpretation system [15]. Relative numbering of nucleotide (nt), amino acid (aa), insertions and deletions used the HCV reference isolate H77 (GenBank accession number AF009606) [16].
Table 1.
Primers used for HCV quantification, genotyping and DE/17–0414 genome amplification
| Primera | Sequence (5′-3′) | Locationb | Reference |
|---|---|---|---|
| Real-time RT-PCR assay for HCV quantification | |||
| HCV-238_f | GAGGAACTACTGTCTTCACG | 49–68 | This study |
| HCV-239_r | TCGCAAGCACCCTATCAG | 310–293 | |
| HCV-240_f | TCGCAAGCACCCTATCAG | 76–94 | |
| HCV-235_r | AGTACCACAAGGCCTTTCG | 290–272 | |
| Heminested RT-PCR assay for HCV genotyping | |||
| HCV-271_f | ACCACATCMRSTCCGTGTGG | 7951–7970 | This study |
| HCV-272_f | TCCGTGTGGRARGACYTSCTRGA | 7962–7984 | |
| HCV-305_f | CTCCGTMTGGGAGGACTTGC | 7961–7980 | |
| HCV-275_r | CTSGTCATAGCYTCCGTGAA | 8635–8616 | |
| Heminested RT-PCR assays for DE/17–0414 genome amplification | |||
| HCV-235_r | AGTACCACAAGGCCTTTCG | 290–272 | This study |
| HCV-239_r | TCGCAAGCACCCTATCAG | 310–293 | |
| HCV-365_f | GGCGTTAGTATGAGTGTTGTGC | 87–108 | (Lu et al., 2014) |
| HCV-366_r | TCCCTGAAGAGTTGCGTATTCC | 939–918 | |
| HCV-367_r | AGAAAGAGCAACCGGGAAGATT | 864–843 | |
| HCV-368_f | TCTATCTTCCTTCTTGCCATCCTG | 864–887 | |
| HCV-369_f | AGGGATTTACCATGTCACCAATGA | 935–958 | |
| HCV-370_r | TCAAAGTCAGTAAGAGGTCGACAG | 1747–1724 | |
| HCV-371_f | CCCGGTGCATGGTAGACTAC | 2164–2183 | |
| HCV-372_r | CTCCACCCTCCGTTGGTTAG | 3421–3492 | |
| HCV-373_r | CCGTTGGTTAGGGAGTCAGC | 3412–3393 | |
| HCV-374_f | ACATTCTTGGCTACGTGCTGTA | 3552–3573 | |
| HCV-375_f | CCCCATTATCCAGATGTACACCAA | 3635–3658 | |
| HCV-387_r | TCTGGACTTCTCCCTCCACC | 3531–3512 | This study |
| HCV-388_f | GCCGCATCCAAACATTGAGG | 4421–4440 | |
| HCV-389_f | CGGCAAAGCTATCCCCCTAG | 4478–4497 | |
| HCV-390_r | CCCGCCTGTTTTGTCTGAGA | 5089–5070 | |
| HCV-391_f | GCATCCAAAGAGGCTGAGGT | 5565–5584 | |
| HCV-392_f | CATCCCTGCTGTCCCAACTT | 5588–5607 | |
| HCV-393_r | TTATGTCAGCTCCGCATGGG | 6456–6437 | |
| HCV-394_f | GACGCCGACCTCATAGAAGC | 7017–7036 | |
| HCV-395_r | TGGCGTAACAAGGAGTTGCT | 7708–7689 | |
| HCV-396_r | ATGGGCAGCTTGTTCTCCTC | 7678–7659 | |
| HCV-360_f | CTCACCTGCTATCTCAAGGCAA | 8487–8508 | |
| HCV-361_f | GTTATCTGTGAGAGTAGCGGGG | 8574–8595 | |
aForward primer designation end with _f; reverse primer designations end with _r
bNumbering is according to the HCV prototype strain of H77 (GenBank Acc. No. AF009606)
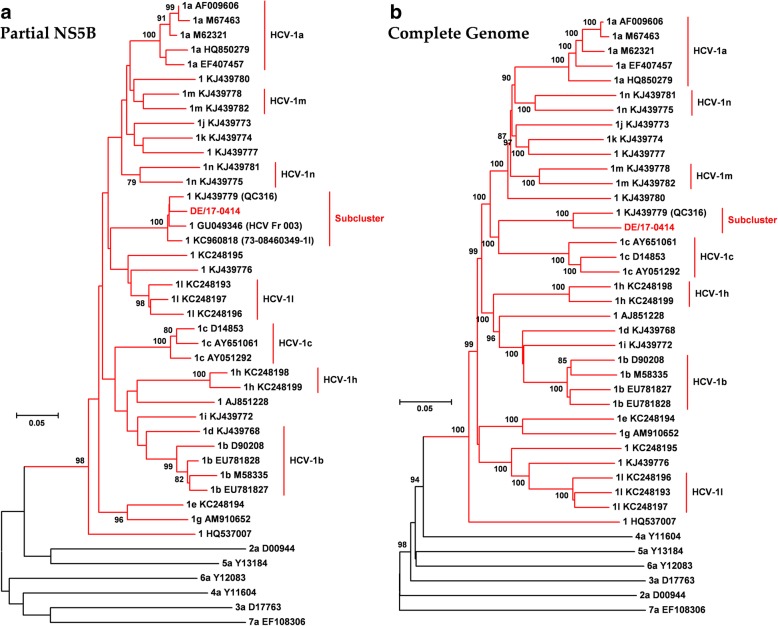
A 63-year-old German heterosexual female, diagnosed with HIV-1 in 2017, was serologically positive for antigen/antibody combination HCV test. Viral load was 1.6 × 106 IU/ml of DSS specimen. Preliminary sequence analysis based on partial NS5B sequences demonstrated that DE/17–0414 has an identity of 96.3% to the isolate QC316 (GenBank accession number KJ439779) from a Canadian immigrant with an origin in Cameroon [17]. It also shows high identities of 95.7 and 95.3% to isolates 73–08460349-1 l (GenBank accession number KC960818) and HCV_Fr_003 from France (GenBank accession number GU049346), respectively [18]. However, DE/17–0414 showed less than 83.6% identity to other HCV-1 strains. Phylogenetic analysis of representative HCV-1 to HCV-7 members of partial NS5B region suggested that DE/17–0414 belonged to HCV-1 forming an independent sub-cluster with HCV_Fr_003, 73–08460349-1 l, and QC316 (Fig. 1a). For a more comprehensive analysis of viruses belonging to the cluster, the full-length genome sequence of DE/17–0414 was amplified and sequenced. The complete genome of DE/17–0414 consisted of 9359 nt excluding the polypyrimidine tract, with a G + C content of 57.9% harboring the 10 HCV prospective genomic regions described in Table 2. The complete genome sequence of DE/17–0414 has been deposited in GenBank under the accession number MH885469. DE/17–0414 had the highest identity with the QC316 (91.8%) and less than 79.6% with any other HCV strain. Phylogenetic reconstructions based on the whole-genome sequences of HCV-1 strains showed that DE/17–0414 and QC316 formed to a separate subcluster within HCV-1 (Fig. 1b). Identity plot and bootscan analyses reflected no evidence for recombination between different HCV genotypes or HCV-1 subtypes (Fig. 2a and b). Intriguingly, a unique insertion of three aa (Q-S-R) was found at the N-terminal of hypervariable region 1 (HVR1) within viral envelope protein 2 (E2) (Fig. 3). In addition, several HCV-1 potential DAAs RASs including 36 L, 170 V (NS3 region) and 28 M, 31 M, 93H (NS5A region), were detected in DE/17–0414 (Table 3).
Fig. 1.
Phylogenetic relationships of DE/17–0414. The strain designations are indicated with geno/subtype and accession number at each branch. Clades corresponding to each genotype were supported by 100% of bootstrap replicates. Bootstrap values (> 75%) are indicated at specific nodes. Scale bars indicate the number of nt substitutions per site. HCV-1 subtypes and the new distinct sub-cluster are indicated on the right. DE/17–0414 of this study is highlighted in bold and red. (a) Phylogenetic analysis of representative HCV-1 strains based on 328 nt of partial NS5B sequences corresponding to nt positions 8283 to 8610 of H77 reference strain. (b) Phylogenetic analysis of HCV-1 complete coding region sequences
Table 2.
Genomic regions of DE/−17–0414
| Genomic region | NA numbering | AA numbering |
|---|---|---|
| 5′ UTR | 1–283 | NAa |
| Core | 284–856 | 1–191 |
| E1 | 857–1432 | 192–473 |
| E2 | 1433–2530 | 474–749 |
| p7 | 2531–2719 | 750–812 |
| NS2 | 2720–3370 | 813–1029 |
| NS3 | 3371–5263 | 1030–1660 |
| NS4A | 5264–5525 | 1661–1714 |
| NS4B | 5526–6208 | 1715–1975 |
| NS5A | 6209–7552 | 1976–2423 |
| NS5B | 7553–9328 | 2424–3014 |
| 3′ UTR | 9329–9359 | NA |
aNA for not applicable
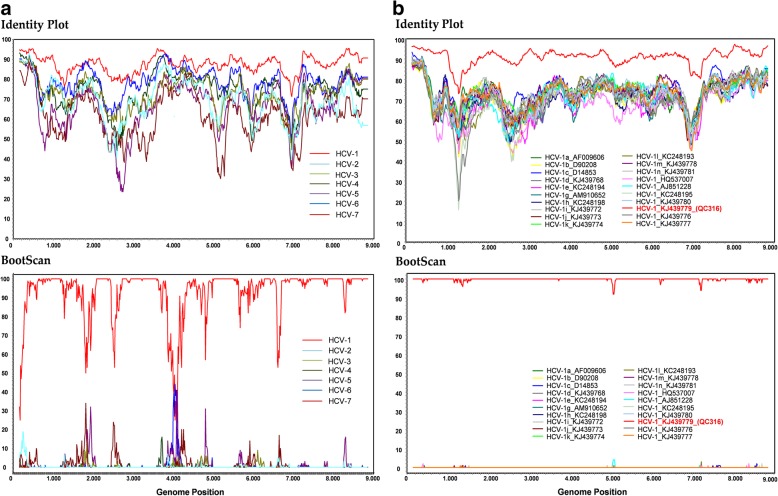
Fig. 2.
Analysis of potential recombination events of the DE/17–0414. Identity Plot and BootScan analyses of (a) HCV genotypes and (b) HCV-1 subtypes. All analyses were performed with a window of 300 nt and a step size of 15 nt under Kimura 2-parameter model. Positions containing gaps were stripped from the alignment. QC316 is highlighted in bold and red
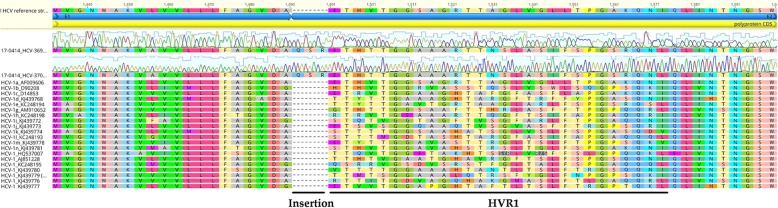
Fig. 3.
Sequence alignment of HCV-1 E1 and E2 genomic regions. The newly detected Q-S-R insertion of DE/17–0414 and HVR1 at the N-terminal of E2 is indicated at the bottom. Absolute numbering is corresponding to aa position 364 to 420 of H77 reference strain
Table 3.
Insertion and potential direct-acting antivirals resistance-associated substitutions of DE/−17–0414
| Genomic regions | Amino acid position | Reference amino acid | DE/−17–0414 | Susceptibility to DAA according to Geno2Pheno[HCV] (Kalaghatgi et al., 2016) |
|---|---|---|---|---|
| E2 | 1a-1c | – | QSR | NAa |
| NS3 | 36 | V | L | Substitution on scored position to Asunaprevir, Grazoprevir, Ledipasvir, Paritaprevir; Reduced susceptibility to Simeprevir, Telaprevir, Voxilaprevir; Resistant to Boceprevir |
| NS3 | 170 | I | V | Substitution on scored position to Voxilaprevir |
| NS5A | 28 | L | M | Substitution on scored position to Ombitasvir |
| NS5A | 31 | L | M | Substitution on scored position to Velpatasvir |
| NS5A | 93 | Y | H | Substitution on scored position to Pibrentasvir; Resistant to Daclatasvir, Elbasvir, Ledipasvir, Ombitasvir, Velpatasvir |
aNA for not available
The assignment of HCV into subtypes and genotypes is based on isolates that differ by 15–25% and by ≥30%, respectively, over their complete coding region sequence [8]. Both DE/17–0414 and QC316 exhibited close to 20% identity to other known HCV sequences. According to the ICTV criteria required for a new HCV genotype or subtype assignment which are: (1) one or more complete coding region sequence(s); (2) a distinct phylogenetic group from previously described sequences; (3) at least three epidemiologically unlinked isolates and (4) exclusion of intergenotypic or intersubtypic recombination [8]. The sequences from a total of 4 epidemiologically unlinked isolates that show more than 95% nucleotide identity have been identified for which the complete genome sequence is available for two of these and the partial NS5B sequence for the remaining two. Thus, this meets the criteria for the assignment of a new HCV subtype 1o. Subsequently, both DE/17–0414 and QC316 regarded as HCV-1o reference sequences.
The main observed genotypes/subtypes in Germany are HCV-1a, 1b and 3 [10]. In contrast, genetic diversity and distribution of other genotypes/subtypes are poorly documented. However, it is known that shifts or relative frequencies of HCV subtypes occurred in the last decades and the approval of DAAs for HCV-treatment is an additional factor, which will probably influence the subtype distribution [19]. Therefore, the knowledge on the genetic diversity of HCV is not only of epidemiological but also clinical significance. The core protein and envelope glycoproteins 1 and 2 constitute the structural elements of HCV [20]. The N-terminal of E2, called HVR1, is most divergent among HCV isolates and contributes to immune escape [21]. A distinct 3 aa (Q-S-R) insertion at the N-terminal of HVR1 was found in DE/17–0414 which exists in none of other known HCV strains. Whether the insertion is associated to HIV-coinfection and its function needs to be further analyzed. With the approval of DAA regimens testing HCV for RASs is clinically relevant. Several potential RASs were detected in the NS3 and NS5A genomic regions of DE/17–0414 on the basis of HCV subtypes 1a and 1b [6], indicating that corresponding DAAs should be avoided in this individual.
In conclusion, we identified and analyzed an HCV-1 divergent isolate from an HIV-1 coinfected individual in Germany, which will be assigned to a new subtype 1o with other three epidemiologically unrelated analogous HCV isolates. The origin and transmission dynamics of this new subtype needs further verification by more comprehensive genetic analyses of HCV strains from patients worldwide.
Acknowledgements
We are grateful for the excellent technical assistance of Marcel Schulze, Julian Heinze, Ewelina Caspers, Steffen Zander, and Hanno von Spreckelsen (RKI).
Funding
This work was supported by a grant from the German Federal Ministry of Health (BMG, grant: ZMVI1–2516-AUK-701 / BMG: 321–4471-02/157). B.W. is funded by the China Scholarship Council (CSC), Beijing, China. A.E. is funded by a scholarship from the German Academic Exchange Service (DAAD), Bonn, Germany. The content is the responsibility only of the authors and does not represent the views of the BMG, CSC or DAAD.
Availability of data and materials
All data generated or analysed during this study are included in this published article.
Abbreviations
- aa
Amino acid
- DAAs
direct-acting antivirals
- DSS
Dried serum spot
- E2
envelope glycoprotein 2
- HCV
Hepatitis C virus
- HCV-1
Hepatitis C virus genotype 1
- HIV-1
Human immunodeficiency virus type 1
- HVR1
Hypervariable region 1
- ICTV
International committee on taxonomy of viruses
- nt
Nucleotide
- RASs
Resistance-associated substitutions
- RKI
Robert koch institute
- WHO
World health organization
Authors’ contributions
NB and CTB conceptualized the study. BW, LK, PM, AE performed the experiment and data analysis. AE, AH, and PM collected specimens. BW, PM, and AH drafted the manuscript. NB, BGB, VB and CTB revised the manuscript critically. All authors read and approved the final manuscript.
Ethics approval and consent to participate
Not applicable. According to IfSG §13 (2017), the RKI is authorized to receive anonymous blood residuals from diagnostics laboratories for surveillance purposes. The KOKPIT study has been approved by the data protection officer of the RKI.
Consent for publication
Not applicable.
Competing interests
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Bo Wang, Email: WangB@rki.de.
Luise Krüger, Email: KruegerL@rki.de.
Patrycja Machnowska, Email: MachnowskaP@rki.de.
Amare Eshetu, Email: AlemayehuA@rki.de.
Barbara Gunsenheimer-Bartmeyer, Email: Gunsenheimer-BartmeyerB@rki.de.
Viviane Bremer, Email: BremerV@rki.de.
Andrea Hauser, Email: HauserA@rki.de.
Norbert Bannert, Email: BannertN@rki.de.
C.-Thomas Bock, Phone: +49-30-18754-2379, Email: bockc@rki.de.
References
- 1.WHO: Hepatitis C Fact Sheet. Available online: http://www.who.int/news-room/fact-sheets/detail/hepatitis-c (accessed 22 October 2018).
- 2.Operskalski EA, Kovacs A. HIV/HCV co-infection: pathogenesis, clinical complications, treatment, and new therapeutic technologies. Curr HIV/AIDS Rep. 2011;8(1):12–22. doi: 10.1007/s11904-010-0071-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Neukam K, Morano-Amado LE, Rivero-Juarez A, Mancebo M, Granados R, Tellez F, Collado A, Rios MJ, de Los Santos-Gil I, Reus-Banuls S, et al. HIV-coinfected patients respond worse to direct-acting antiviral-based therapy against chronic hepatitis C in real life than HCV-monoinfected individuals: a prospective cohort study. HIV Clin Trials. 2017;18(3):126–134. doi: 10.1080/15284336.2017.1330801. [DOI] [PubMed] [Google Scholar]
- 4.Schlabe S, Rockstroh JK. Advances in the treatment of HIV/HCV coinfection in adults. Expert Opin Pharmacother. 2018;19(1):49–64. doi: 10.1080/14656566.2017.1419185. [DOI] [PubMed] [Google Scholar]
- 5.Sarrazin C. The importance of resistance to direct antiviral drugs in HCV infection in clinical practice. J Hepatol. 2016;64(2):486–504. doi: 10.1016/j.jhep.2015.09.011. [DOI] [PubMed] [Google Scholar]
- 6.Pawlotsky JM. Hepatitis C virus resistance to direct-acting antiviral drugs in interferon-free regimens. Gastroenterology. 2016;151(1):70–86. doi: 10.1053/j.gastro.2016.04.003. [DOI] [PubMed] [Google Scholar]
- 7.Simmonds P, Becher P, Bukh J, Gould EA, Meyers G, Monath T, Muerhoff S, Pletnev A, Rico-Hesse R, Smith DB, et al. ICTV virus taxonomy profile: Flaviviridae. J Gen Virol. 2017;98(1):2–3. doi: 10.1099/jgv.0.000672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Smith DB, Bukh J, Kuiken C, Muerhoff AS, Rice CM, Stapleton JT, Simmonds P. Expanded classification of hepatitis C virus into 7 genotypes and 67 subtypes: updated criteria and genotype assignment web resource. Hepatology. 2014;59(1):318–327. doi: 10.1002/hep.26744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Messina JP, Humphreys I, Flaxman A, Brown A, Cooke GS, Pybus OG, Barnes E. Global distribution and prevalence of hepatitis C virus genotypes. Hepatology. 2015;61(1):77–87. doi: 10.1002/hep.27259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kartashev V, Doring M, Nieto L, Coletta E, Kaiser R, Sierra S, Group HCVES New findings in HCV genotype distribution in selected west European, Russian and Israeli regions. J Clin Virol. 2016;81:82–89. doi: 10.1016/j.jcv.2016.05.010. [DOI] [PubMed] [Google Scholar]
- 11.Hauser A, Hofmann A, Hanke K, Bremer V, Bartmeyer B, Kuecherer C, Bannert N: National molecular surveillance of recently acquired HIV infections in Germany, 2013 to 2014. Euro Surveill 2017, 22(2), pii: 30436. [DOI] [PMC free article] [PubMed]
- 12.Wang B, Yang XL, Li W, Zhu Y, Ge XY, Zhang LB, Zhang YZ, Bock CT, Shi ZL. Detection and genome characterization of four novel bat hepadnaviruses and a hepevirus in China. Virol J. 2017;14(1):40. doi: 10.1186/s12985-017-0706-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33(7):1870–1874. doi: 10.1093/molbev/msw054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lole KS, Bollinger RC, Paranjape RS, Gadkari D, Kulkarni SS, Novak NG, Ingersoll R, Sheppard HW, Ray SC. Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J Virol. 1999;73(1):152–160. doi: 10.1128/jvi.73.1.152-160.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kalaghatgi P, Sikorski AM, Knops E, Rupp D, Sierra S, Heger E, Neumann-Fraune M, Beggel B, Walker A, Timm J, et al. Geno2pheno[HCV] – a web-based interpretation system to support hepatitis C treatment decisions in the era of direct-acting antiviral agents. PLoS One. 2016;11(5):e0155869. doi: 10.1371/journal.pone.0155869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kuiken C, Combet C, Bukh J, Shin IT, Deleage G, Mizokami M, Richardson R, Sablon E, Yusim K, Pawlotsky JM, et al. A comprehensive system for consistent numbering of HCV sequences, proteins and epitopes. Hepatology. 2006;44(5):1355–1361. doi: 10.1002/hep.21377. [DOI] [PubMed] [Google Scholar]
- 17.Lu L, Li C, Xu Y, Murphy DG. Full-length genomes of 16 hepatitis C virus genotype 1 isolates representing subtypes 1c, 1d, 1e, 1g, 1h, 1i, 1j and 1k, and two new subtypes 1m and 1n, and four unclassified variants reveal ancestral relationships among subtypes. J Gen Virol. 2014;95(7):1479–1487. doi: 10.1099/vir.0.064980-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Koletzki D, Dumont S, Vermeiren H, Fevery B, De Smet P, Stuyver LJ. Development and evaluation of an automated hepatitis C virus NS5B sequence-based subtyping assay. Clin Chem Lab Med. 2010;48(8):1095–1102. doi: 10.1515/CCLM.2010.236. [DOI] [PubMed] [Google Scholar]
- 19.Schroter M, Zollner B, Schafer P, Reimer A, Muller M, Laufs R, Feucht HH. Epidemiological dynamics of hepatitis C virus among 747 German individuals: new subtypes on the advance. J Clin Microbiol. 2002;40(5):1866–1868. doi: 10.1128/JCM.40.5.1866-1868.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Penin F, Dubuisson J, Rey FA, Moradpour D, Pawlotsky JM. Structural biology of hepatitis C virus. Hepatology. 2004;39(1):5–19. doi: 10.1002/hep.20032. [DOI] [PubMed] [Google Scholar]
- 21.Bankwitz D, Vieyres G, Hueging K, Bitzegeio J, Doepke M, Chhatwal P, Haid S, Catanese MT, Zeisel MB, Nicosia A, et al. Role of hypervariable region 1 for the interplay of hepatitis C virus with entry factors and lipoproteins. J Virol. 2014;88(21):12644–12655. doi: 10.1128/JVI.01145-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data generated or analysed during this study are included in this published article.