Abstract
Lung adenocarcinoma (LAC) is a leading cause of cancer-associated mortalities, particularly in developed countries. The aberrant expression of microRNAs (miRNAs) has been proven to regulate numerous diseases in the past two decades. miRNAs have been identified in almost all human cancer types. In the present study, the role of miR-944 in LAC proliferation was examined. It was identified that miR-944 was downregulated in LAC tissues and cells, and miR-944 overexpression inhibited A549 and H1299 cell proliferation, as determined by the Cell Counting Kit-8 and colony formation assay. Signal transducer and activator of transcription 1 (STAT1) was upregulated in LAC tissues and cells. Kaplan-Meier analysis demonstrated that the 5-year overall survival in patients with high STAT1 levels was significantly reduced, compared with patients with negative and low STAT1 expression. STAT1 was the direct target of miR-944. Additionally, a miR-944 mimic inhibited A549 cell growth in vitro. Collectively, these data demonstrate that miR-944 serves a pivotal role in LAC tumor growth by targeting STAT1. The data obtained indicated that miR-944 may be a novel biomarker and could result in potential therapies for LAC.
Keywords: lung adenocarcinoma, microRNA, signal transducer and activator of transcription 1, proliferation
Introduction
Lung adenocarcinoma (LAC) is a serious disease, and accounts for ~40% of all lung cancer cases (1,2). Researches revealed that LAC is a leading cause of cancer-associated mortalities, particularly in developing countries (1,3). In China, there are >1 million patients with LAC and >200,000 mortalities associated with lung cancer annually (4). Notably, LAC is consistently identified among non-smokers (5). Despite recent advances in diagnosis, surgery, chemotherapy, targeted therapy, radiation therapy, cellular immunotherapy and radiofrequency ablation, the overall survival rate of patients with LAC remains low at an advanced stage, with a 5-year survival rate of only ~18% (6–8). Additionally, these patients also suffer complications, including postoperative metastasis and side effects from drug treatments (9). Thus, the identification of novel diagnostic methods for patients with LAC will help monitor tumor progression and guide clinical treatment, which may assist in the development of gene target-based therapy.
microRNAs (miRNAs) are small non-coding RNAs ~20–22 nt in length that serve vital roles in diseases, which threaten human health by coding specific mRNAs (10,11). In the past two decades, increasing evidence indicated that miRNAs are involved in the pathogenesis of LAC (12,13). Berrout et al (14) demonstrated that miRNA-142-3p functions as a regulatory oncogenic driver by binding transient receptor potential cation channel subfamily A member 1-fibroblast growth factor receptor 2 in LAC. Yan et al (15) reported that miR-503 modulated epithelial-mesenchymal transition in silica-induced pulmonary fibrosis by targeting phosphoinositide 3-kinase p85. Additionally, the study of Pan et al (16) indicated that miR-944 served a tumor suppressive role via the metastasis associated in colon cancer 1 (MACC1)/Met/AKT signaling pathway in gastric cancer (16). The aforementioned studies demonstrated that miRNAs serve crucial roles in LAC and other diseases. However, the role of miR-944 in LAC requires further investigation.
Signal transducer and activator of transcription (STAT)-1, a member of the STAT super family, has a number of biological functions, including acting as a tumor suppressor and preventing tumor development, and also exhibits a role in immunotherapy (17–19). A previous study reported that STAT1 served vital roles in the miR-15A and miR-16-1 signaling pathways in the regulation of colorectal tumors (20). Additionally, Zhang et al (21) reported that miR-181a/STAT1 inhibited colorectal cancer cell proliferation by regulating the phosphatase and tensin homolog/AKT signaling pathway (22). However, to the best of our knowledge, at present there is has been no report investigating whether miR-944 has a role in LAC. Collectively, the present study aimed to investigate the effects of miR-944 on cell proliferation and apoptosis in LAC.
Materials and methods
Tissue collection
A total of 25 LAC tissues from 13 males and 12 females, with a median age of 57.6 years, were obtained from patients who underwent surgery at the Third Hospital of Qiqihar Medical College (Qiqihar, China), between September 2014 and September 2016. The present study was approved by the Research Ethics Committee of Third Hospital of Qiqihar Medical College, and written informed consent was obtained from all patients. All the specimens, including cancer tissues, were diagnosed with LAC (stages I, II, and III) (23). The patients received no local or systemic treatments prior to surgery (Table I). All collected tissues were placed in liquid nitrogen immediately and stored at −80°C until required.
Table I.
Patient clinical information.
| Variables | Patients (n=25) |
|---|---|
| Age (years) | 57.6 (range 30–80 years) |
| Sex | |
| Male | 13 |
| Female | 12 |
| Tumor size | |
| ≤5 cm | 16 |
| >5 cm | 9 |
| TNM stage | |
| I/II | 15 |
| III/IV | 10 |
| Lymph node metastasis | |
| Yes | 18 |
| No | 7 |
TNM, tumor node metastasis (41).
Cell culture
LAC cells (A549, H1299, SK-Lu-1, and PC-9) were obtained from the Chinese Academy of Science Shanghai Cell Bank (Shanghai, China) and cultured in Dulbecco's modified Eagle's medium (HyClone; GE Healthcare Life Sciences, Logan, UT, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA), 100 U/ml penicillin, and 100 g/ml streptomycin (Beyotime Institute of Biotechnology, Shanghai, China). Human bronchial epithelial cells (16HBE), obtained from the Henan Xingfa Bio-Technology Co., Ltd. (Henan, China), were maintained in RPMI-1640 (HyClone; GE Healthcare Life Sciences) supplemented with 10% FBS, 100 µg/ml penicillin, and 100 µg/ml streptomycin. These cells were placed in a humidified atmosphere containing 5% CO2 at 37°C.
Gene silencing of STAT1
STAT1-small interfering (si)RNA was purchased from Shanghai GeneChem Co., Ltd. (Shanghai, China). The sequence was as follows: 5′-CCGGCTGGAAGATTTACAAGATGAACTCGAGTTCATCTTGTAAATCTTCCAGTTTTTG-3′. A control siRNA (5′-CCGGTTCTCCGAACGTGTCACGTTTCAAGAGAACGTGACACGTTCGGAGAATTTTTG-3′) was used as a negative control. Cells were transfected with 80 pmol siRNA plasmid for 48 h using Lipofectamine™ 2000 (Invitrogen; Thermo Fisher Scientific, Inc.). At 48 h following transfection A549 cells were harvested for subsequent experimentation.
Colony formation assay
A total of 8×102 A549 or H1229 cells were plated in triplicate into 60 mm dishes and cultured for 14 days in a humidified atmosphere containing 5% CO2 at 37°C. Following 14 days, colonies were stained with 0.1% crystal violet in 20% methanol for 15 min. Colonies consisting of >50 cells were counted as a single colony.
Dual luciferase reporter assay
Wild-type (WT) miR-944 (miR-944-WT), mutant miR-944 (miR-944-Mut), STAT1-WT, and STAT1-Mut were cloned into separate pMIR-REPORT Luciferase vectors (Ambion; Thermo Fisher Scientific, Inc.). A total of 8×103 A549 cells/well were seeded in 6-well plates and transfected with the specific vectors using Lipofectamine® 2000 (Invitrogen; Thermo Fisher Scientific, Inc.) for 48 h as described previously. Luciferase activity was assessed using the Dual Luciferase-reporter 1000 assay system (Promega Corporation, Madison, WI, USA). Renilla activity was used for normalization.
Kaplan-Meier method
All clinical data and the Tier 3 RNASeqV2 mRNA expression data were downloaded from https://tcga-data.nci.nih.gov/tcga/. Patients with a follow-up time or time to mortality >0 days were included in the analysis. For each gene, all samples were divided to two groups based on the median expression values, with samples with a median value placed in the high expression group. Kaplan-Meier analysis was then performed to examine the significance between the two groups. Cox proportional hazards regression was also performed with the coxph function from the R survival library (version 2.43–3; http://cran.r-project.org/web/packages/survival/index.html). Hazard ratios with 95% confidence intervals were obtained.
Cell Counting Kit (CCK)-8 assay
CCK-8 assays (Dojindo Molecular Technologies, Inc., Kumamoto, Japan) were used to determine cell viability following transfection with 50 pmol miR-944 mimic and miR-negative control (NC) in A549 and H1299 cells, according to the manufacturer's protocol and the aforementioned protocol. LAC cells were seeded into 96-well plates at a density of 2×103 cells/well and cultured for 48 h in a humidified atmosphere containing 5% CO2 at 37°C. The sequences of miR-944 mimic and NC were as follows: miR-944 NC, 5′-ACUUCAGUGGAUGUUUGCAGC-3′; and miR-944 mimic, 5′-GAGUAGGCUAAUGUUAUAAA-3′.
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was isolated from A549 cells using TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.) and cDNA was synthesized using High-Capacity cDNA Reverse Transcription kit (Applied Biosystems; Thermo Fisher Scientific, Inc.). The primers were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). qPCR was performed using Power SYBR Green PCR master mix (Thermo Fisher Scientific, Inc.) for 35 cycles at 95°C for 30 sec, 60°C for 30 sec and 72°C for 35 sec. Gene expression levels were normalized with β-actin and analyzed using the 2−ΔΔCq method (20). The primer sequences were as follows: STAT1 forward, 5′-AGCCAGTGCAAATCACGATG-3′ and reverse, 5′-CGTCAGCAAAGCCCATTTGA-3′; β-actin forward, 5′-TGTCACCAACTGGGACGATA-3′ and reverse, 5′-GGGGTGTTGAAGGTCTCAAA-3′.
Western blot analysis
All proteins were obtained from LAC cells and patients. A549 cells and patient tissues were lysed with radioimmunoprecipitation assay buffer (Beyotime Institute of Biotechnology). The proteins were then quantified using a bicinchoninic acid assay (Beyotime Institute of Biotechnology). The proteins (60–80 µg) were separated by 10% SDS-PAGE and transferred to a nitrocellulose membrane. Following blocking with 5% non-fat milk for 2 h at room temperature, the blots were probed with primary antibodies against STAT1 (catalog no. ab30645; 1:500; Abcam, Cambridge, MA, USA) and β-actin (catalog no. 4970; 1:500; Cell Signaling Technology, Inc., Danvers, MA, USA) at 4°C overnight. Following threes washes with PBS and Tween-20 for 15 min, the membranes were incubated with rabbit (catalog no. 926-32211-00; 1:10,000) or mouse (catalog no. 926-32211-01; 1:10,000) secondary antibodies (LI-COR Biosciences, Lincoln, NE, USA) at room temperature in the dark for 1 h. The blots were then visualized using an Infrared Imaging System (LI-COR Biosciences) and the band density was quantified using Odyssey 3.0 software (LI-COR Biosciences) (22). Using β-actin as an internal control, the blots were subjected to densitometry.
Tumor xenograft nude mouse model
Transfected A549 cells were subcutaneously injected into the back of total of 30 male BALB/c nude mice, aged 6 weeks old with a median weight of ~20 g (Vital River Laboratory Animal Technology, Beijing, China). The mice were kept at 21–24°C in a light/dark cycle with food and water available ad libitum. Tumor volumes were measured once each week at equal intervals. A tumor growth curve was constructed to determine the effects of the miR-944-mimic and miR-NC on tumor growth. At 21 days following injection, the mice were sacrificed with carbon dioxide at a displacement rate of 20%/min and imaged, and the tumors were dissected. The tumors in each group were harvested and weighed. Total proteins and RNAs were extracted for western blotting and RT-qPCR. All animal experiments were approved by the Third Hospital of Qiqihar Medical College's Animal Care and Use Committee and conducted according to the National Institutes of Health guidelines (23).
Statistical analysis
Statistical analysis was conducted using SPSS software (version 13.0; SPSS Inc., Chicago, IL, USA). All data were expressed as means ± standard deviation. Statistical analysis was performed using Student's non-paired t-test or one-way analysis of variance followed by Tukey's post-test. miR-944 targets were predicted using Targetscan software 7.2 (24). P<0.05 was considered to indicate a statistically significant difference.
Results
miR-944 is underexpressed in LAC tissues and cell lines
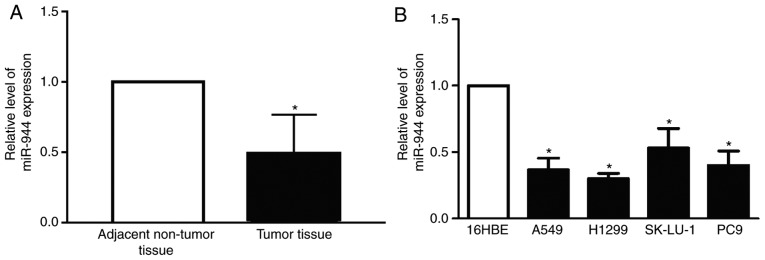
To determine the biological function of miR-944 in patients with LAC and cell lines, the present study initially evaluated the expression levels of miR-944 in 25 pairs of LAC tissues. The expression of miR-944 was significantly reduced in the LAC group compared with the normal group (Fig. 1A). Additionally, the expression of miR-944 in the four lung cancer cell lines, A549, H1299, SK-Lu-1 and PC-9, was significantly decreased, compared with normal 16HBE lung cells (Fig. 1B). However, the expression of miR-944 was reduced in A549 and H1299 cell lines, compared with SK-Lu-1 and PC9 cell lines. Therefore, for subsequent experiments the A549 and H1299 cell lines were selected. This data demonstrated that miR-944 functions as a tumor suppressor gene in LAC tissues and cell lines.
Figure 1.
miR-944 expression is reduced in LAC tissues and cell lines. (A) Relative levels of miR-944 expression were analyzed by reverse transcription-quantitative polymerase chain reaction in 25 pairs of patients with LAC. (B) The expression of miR-944 in LAC the cell lines A549, H1299, SK-Lu-1 and PC-9, and the human bronchial epithelial cell line 16HBE. Data presented as the mean ± standard deviation of three replicates. *P<0.05 vs. normal group; 16HBE group. LAC, lung adenocarcinoma; miR, microRNA.
miR-944 inhibits cell proliferation
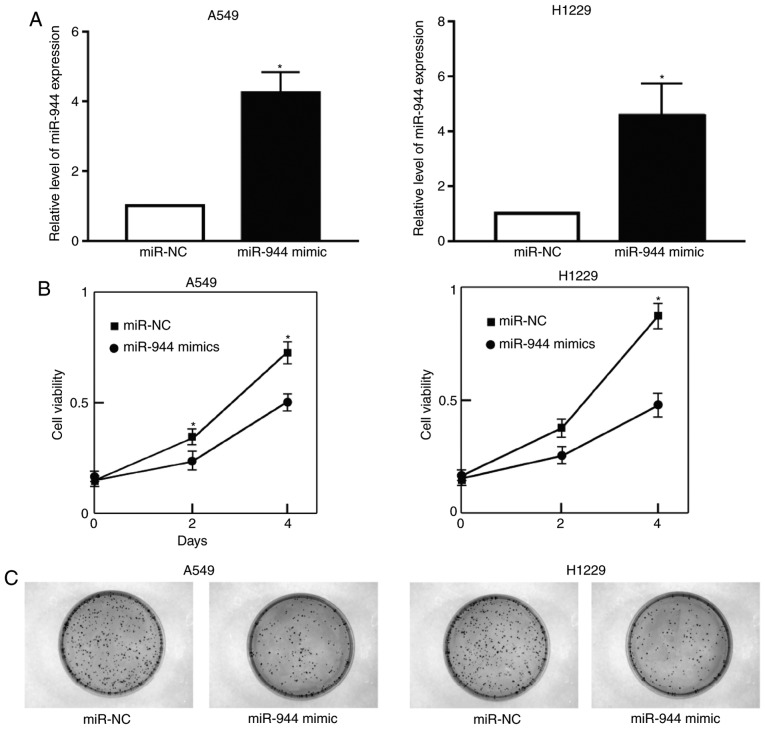
To improve the understanding of the effect of miR-944 on LAC cell proliferation, A549 and H1299 cells were treated with the miR-944 mimic or miR-NC. The expression of miR-944 was significantly increased following treatment with the miR-944 mimic in A549 and H1299 cells (Fig. 2A). Cell growth was also observed to be attenuated in miRNA-944 mimic cells, compared with miR-NC cells, as illustrated by the CCK-8 and colony formation assays (Fig. 2B and C). Collectively, these results indicate that miRNA-944 had a negative effect on the proliferation of LAC cells.
Figure 2.
miR-944 inhibits cell proliferation. (A) Relative expression levels of miR-944 following transfection with miR-944 mimic or miR-NC in A549 and H1299 cells (B) A total of 2×103 A549 and H1299 cells were plated per well in 96-well plates, and the Cell Counting Kit-8 assay was performed to detect cell proliferation absorbance at 450 nm at days 0, 2 and 4. (C) Colony formation assays were used to detect cell proliferation. *P<0.05 vs. miR-NC. NC, negative control; miR, microRNA.
STAT1 is upregulated in LAC tissue and cells
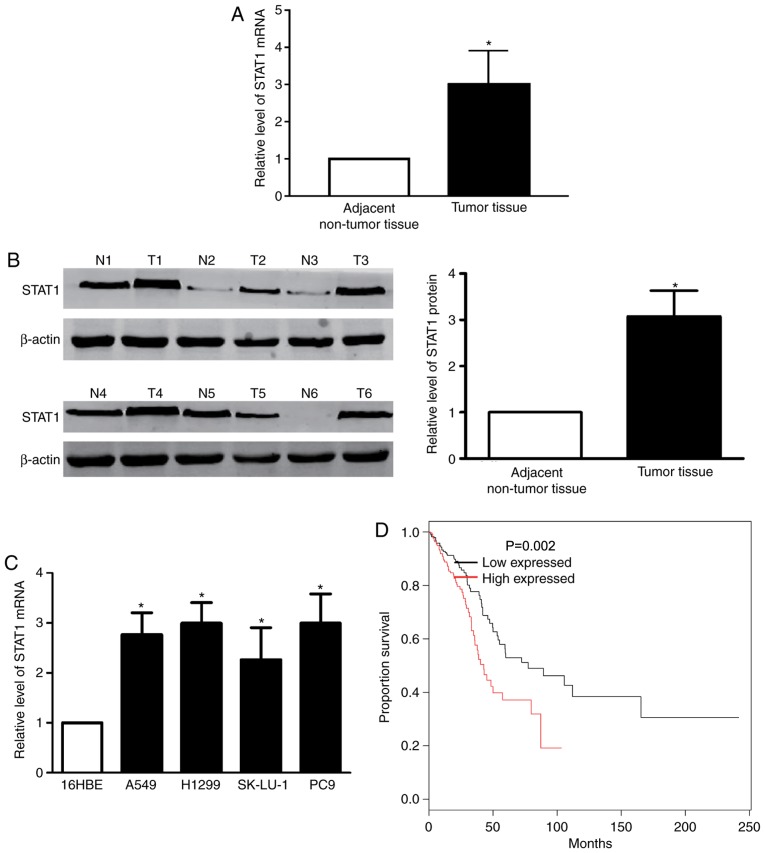
The RT-qPCR demonstrated that the STAT1 mRNA level was significantly upregulated in LAC samples, compared with paired non-malignant samples (Fig. 3A). Western blotting confirmed the results of the STAT1 mRNA level (Fig. 3B). Subsequently, the present study determined the expression of STAT1 in the LAC cell lines A549, H1299, SK-Lu-1 and PC-9. The data revealed that significantly increased levels of STAT1 mRNA expression were identified in LAC cell lines, compared with 16HBE cells (Fig. 3C). Using the Kaplan-Meier method, the overall survival times in patients with high-STAT1 expression were revealed to be significantly reduced, compared with patients with negative and low-STAT1 expression (Fig. 3D).
Figure 3.
STAT1 is upregulated in LAC tissue and cells. (A) The STAT1 mRNA level was upregulated in human LAC tissues, compared with adjacent non-tumor tissues, as determined by RT-qPCR. *P<0.05 vs. adjacent non-tumor tissues. (B) The protein level of STAT1 is upregulated in human LAC tissues, compared with adjacent non-tumor tissues. *P<0.05 vs. adjacent non-tumor tissues. (C) Relative STAT1 mRNA expression levels in 16HBE and LAC cell lines at determined by RT-qPCR. *P<0.05 vs. 16HBE. (D) The Kaplan-Meier method was used to determine survival time. STAT1, signal transducer and activator of transcription 1; RT-qPCR, reverse transcription-quantitative polymerase chain reaction; T, tumors; N, adjacent non-tumor tissues; LAC, lung adenocarcinoma.
miR-944 directly downregulates the expression of STAT1
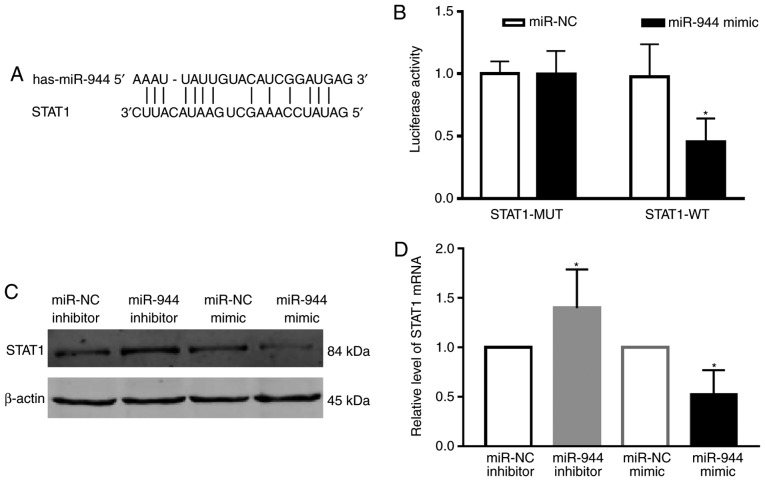
To understand how miR-944 functions in LAC, the Microrna search program (www.targetscn.org) was used to predict targets of miR-944, which revealed that STAT1 is considered to be a putative target of miR-944 (Fig. 4A). The luciferase reporter assay also demonstrated that STAT1 was a direct target gene of miR-944 (Fig. 4B). The protein level of STAT1 following transfection with the miR-944 and miR-NC mimics or miR-944 inhibitor and miR-NC inhibitor was then determined in A549 cells. The results demonstrated that the protein level of STAT1 was downregulated by treatment with the miR-944 mimic, and upregulated following treatment with the miR-944 inhibitor (Fig. 4C). These results were also confirmed at the mRNA level (Fig. 4D). Collectively, these results indicate that STAT1 was a direct target of miR-944 in LAC A549 cells.
Figure 4.
miR-944 directly downregulates the expression of STAT1. (A) The predicted targeting site of STAT1 3′-untranslated region combined with miR-944 is illustrated. (B) Luciferase assay of A549 cells. (C) The protein band mRNA levels and (D) quantification of STAT1 following transfection with miR-944 inhibitor and miR-NC, or miR-944 mimics and miR-NC. *P<0.05 vs. the miR-NC group. STAT1, signal transducer and activator of transcription 1; MUT, mutant; WT, wild-type; NC, negative control; miR, microRNA.
miR-944 inhibits tumor growth in vivo
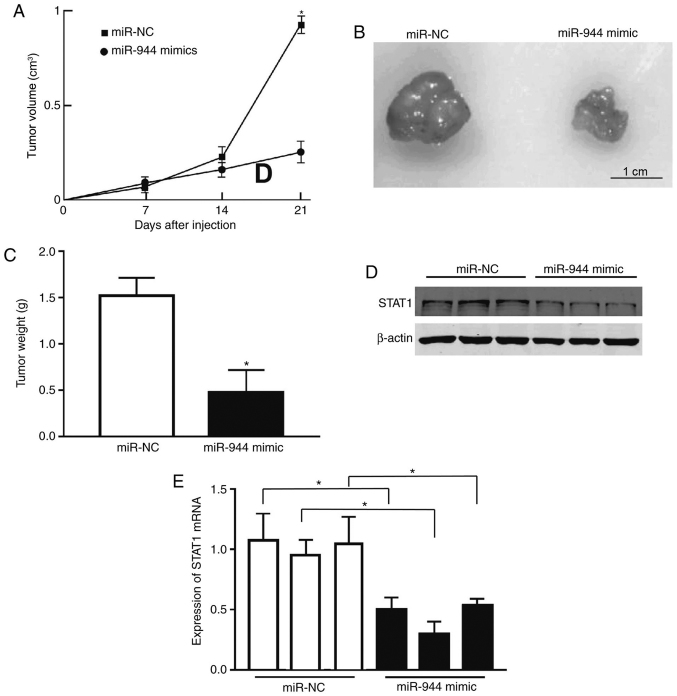
To examine the role of miR-944 in tumor proliferation in vivo, a nude mouse xenograft model was used. A549 cells were transfected with miR-NC or miR-944 mimics and injected into mice. Tumor size was measured once each week, and the growth curve as a function of the average tumor size was plotted following the injection of cells. The mice were sacrificed after 2 weeks and their bodies and xenografts were weighed. As expected, there was a significant decrease in tumor size and weight in the miR-944-overexpressing groups compared with the NC group (Fig. 5A-C). Consistent with the in vitro studies, the protein level of STAT1 in tumor tissues from the miR-944 mimic group was markedly reduced, compared with the miR-NC group, as illustrated by the immunoblotting assay (Fig. 5D). The RT-qPCR data on STAT1 mRNA measurement also exhibited the same pattern (Fig. 5E). Collectively, these observations indicate that miR-944 was a tumor suppressor in LAC.
Figure 5.
miR-944 suppresses tumor growth in vivo by targeting STAT1. (A) Tumors formed in nude mice. A total of 5×106 A549 cells were subcutaneously injected into nude mice (n=5). The mice were then sacrificed 21 day after injection. (B) Tumors were harvested, and images of representative tumors were obtained. (C) The miR-944 mimic transfection resulted in a decrease in final tumor weight. (D) The protein levels of STAT1 from tumor xenografts were measured by western blotting. (E) The mRNA levels of STAT1 in tumor xenografts were measured by reverse transcription quantitative polymerase chain reaction. A total of 30 mice were randomly divided into six groups to perform three independent experiments for each type of transfected cell. *P<0.05 vs. miR-NC. STAT1, signal transducer and activator of transcription 1; NC, negative control; miR, microRNA.
Discussion
Over the past two decades, a number of miRNAs, including LAC, have been demonstrated to exhibit functions in numerous diseases, including circulatory system diseases, cerebrovascular diseases and cancer (25–27). Recently, increasing evidence has implicated STAT1 in a number of processes associated with cancer, including cell cycle regulation, motility, differentiation and proliferation (18,20,28). The present study identified that miR-944 is downregulated in LAC tissues and cells, indicating a potential role for miR-944 in LAC. STAT1 is a direct target gene of miR-944, and overexpression of miR-944 significantly inhibited LAC cell proliferation, survival and tumor growth.
miRNAs are small non-coding RNAs that regulate 30% of gene expressions (29,30). miRNAs act as master regulators of gene expression in numerous important biological pathways, including cell cycle, apoptosis and proliferation (31), particularly in cancer. miRNAs can act as oncogenes or tumor suppressors and regulate their target genes, which are dysregulated in a numbr of cancer types, including prostate cancer, colon cancer and gastrointestinal cancer (32–34). Recent studies have demonstrated that miR-944 acted as a tumor suppressor in numerous cancer types, for example Wen et al (35) reported that miR-944 inhibits cell migration and invasion by targeting MACC1 in colorectal cancer. Additionally, He et al (36) demonstrated that miR-944 acts as a prognostic marker and promotes tumor progression in endometrial cancer. However, there is currently no data regarding the role of miR-944 in LAC proliferation. In the present study, miR-944 expression was identified to be downregulated in LAC tissue samples, compared with normal tissues. These results indicate that miR-944 may exhibit an anticancer effect of LAC. Additionally, the overexpression of miR-944 significantly inhibits LAC cell proliferation and tumor growth, which demonstrated that miR-1994 has a notable anti-proliferation effect in vivo and vitro.
STAT1 belongs to the STAT super family and has numerous functions, including reducing apoptosis, attenuating inflammation and modulating oxidative stress (37,38). Carbotti et al (39) reported that interleukin-27 triggered STAT1 phosphorylation in small cell lung cancer cells. Furthermore, Zhang et al (40) discussed the role of STAT1 in cancer. Recently, evidence indicated that STAT1 can be regulated by miRNAs. For example, Xi et al (41) reported that miR-21 depletion in macrophages promotes tumoricidal polarization and enhances PD-1 immunotherapy by STAT1 (42). Additionally, Li et al (43) indicated that miR-194 promotes osteoblast differentiation via downregulating STAT1. In the present study, STAT1 was identified as a direct target gene of miR-944. To further investigate the role of STAT1 in LAC, the expression of STAT1 in patients with LAC and LAC cell lines was examined. The results demonstrated that the expression of STAT1 is upregulated in LAC tissues. The study by Gujam et al (44) demonstrated that STAT1 and STAT3 regulate tumor microenvironment and survival in patients with invasive ductal breast cancer. In the present study, the overexpression of STAT1 was observed to decrease the 5-year survival rate of patients with LAC, these data demonstrated that STAT1 functions as an oncogene in LAC. Additionally, downregulated miR-944 can upregulate STAT1 protein and mRNA expression levels, furthermore, overexpression of miR-944 can increase the expression of STAT1. These results demonstrated that the STAT1 oncogene was validated experimentally as the novel target of miR-944. Collectively, the present study demonstrated that miR-944 significantly suppressed LAC growth through inhibition of STAT1 translation.
In summary, the present study demonstrated that miR-944/STAT1 is a novel constituent of LAC tumorigenesis and proliferation, and miR-944 regulates cell growth in LAC by targeting STAT1. This data contributes towards the improved understanding of LAC and indicates that miR-944 may be used as a clinical agent in the treatment of LAC. Although miR-944/STAT1 is not the only signaling pathway to regulate LAC cell proliferation and migration, it may provide the foundation for the development of novel methods for the diagnosis and therapy of LAC.
Acknowledgements
Not applicable.
Funding
This work was supported by the Qiqihar Science and Technology Bureau project (grant no. SFZD-2017001).
Availability of data and materials
The analyzed datasets generated during the study are available from the corresponding author on reasonable request, while preserving the necessary confidentiality and anonymity.
Authors' contributions
BM conceived and designed the study and critically revised the manuscript. JCA and HBS designed and performed the experiments, analyzed the data and wrote the manuscript. WBH and KZ were involved in drafting the manuscript and revising it critically for important intellectual content, gave advice on the experiments, designed and performed the experiments, analyzed the data, and contributed with reagents and technical assistance. All authors read and approved the final manuscript.
Ethics approval and consent to participate
The present study was approved by the Research Ethics Committee of Third Hospital of Qiqihar Medical College (Qiqihar, China), and written informed consent was obtained from all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
References
- 1.Huang TW, Lin KF, Lee CH, Chang H, Lee SC, Shieh YS. The role of thyroid transcription factor-1 and tumor differentiation in resected lung adenocarcinoma. Sci Rep. 2017;7:14222. doi: 10.1038/s41598-017-14651-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Vieira A, Ugalde Figueroa P. Anatomic bisegmentectomy for synchronous lung adenocarcinoma. J Vis Surg. 2017;3:64. doi: 10.21037/jovs.2017.03.20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zhang T, Chen L, Zhang S, Xu Y, Fan Y, Zhang L. Effects of high-intensity focused ultrasound on cisplatin-resistant human lung adenocarcinoma in vitro and in vivo. Acta Biochim Biophys Sin (Shanghai) 2017;49:1092–1098. doi: 10.1093/abbs/gmx107. [DOI] [PubMed] [Google Scholar]
- 4.Wu H, Meng S, Xu Q, Wang X, Wang J, Gong R, Song Y, Duan Y, Zhang Y. Gene expression profiling of lung adenocarcinoma in Xuanwei, China. Eur J Cancer Prev. 2016;25:508–517. doi: 10.1097/CEJ.0000000000000214. [DOI] [PubMed] [Google Scholar]
- 5.Ni J, Zhou LL, Ding L, Zhao X, Cao H, Fan F, Li H, Lou R, Du Y, Dong S, et al. PPARγ agonist efatutazone and gefitinib synergistically inhibit the proliferation of EGFR-TKI-resistant lung adenocarcinoma cells via the PPARγ/PTEN/Akt pathway. Exp Cell Res. 2017;361:246–256. doi: 10.1016/j.yexcr.2017.10.024. [DOI] [PubMed] [Google Scholar]
- 6.Bernhardt D, Adeberg S, Bozorgmehr F, Opfermann N, Hörner-Rieber J, König L, Kappes J, Thomas M, Unterberg A, Herth F, et al. Outcome and prognostic factors in single brain metastases from small-cell lung cancer. Strahlenther Onkol. 2018;194:98–106. doi: 10.1007/s00066-017-1228-4. [DOI] [PubMed] [Google Scholar]
- 7.Kim CS, Jeter MD. StatPearls. Treasure; Island (FL): 2018. Radiation therapy, early stage non-small cell lung cancer. [Google Scholar]
- 8.Li-Ming X, Zhao LJ, Simone CB, II, Cheng C, Kang M, Wang X, Gong LL, Pang QS, Wang J, Yuan ZY, Wang P. Receipt of thoracic radiation therapy and radiotherapy dose are correlated with outcomes in a retrospective study of three hundred and six patients with extensive stage small-cell lung cancer. Radiother Oncol. 2017;125:331–337. doi: 10.1016/j.radonc.2017.10.005. [DOI] [PubMed] [Google Scholar]
- 9.Peng Y, Ren W, Wang H, Li M, Feng Z, Peng Z. Surgical treatment is an effective approach for patients with synchronous multiple primary lung cancers. J Cancer Res Ther. 2017;13:702–706. doi: 10.4103/jcrt.JCRT_140_17. [DOI] [PubMed] [Google Scholar]
- 10.Nhat Tran, Alipourfard B, Abhyankar V, Nguyen K, Weidanz J, Gao J. Improved microRNA biomarkers for pathological stages in lung adenocarcinoma via clustering of dysregulated microRNA-target associations. Conf Proc IEEE Eng Med Biol Soc. 2017;2017:2708–2711. doi: 10.1109/EMBC.2017.8037416. [DOI] [PubMed] [Google Scholar]
- 11.Chang SM, Hu WW. Long non-coding RNA MALAT1 promotes oral squamous cell carcinoma development via microRNA-125b/STAT3 axis. J Cell Physiol. 2018;233:3384–3396. doi: 10.1002/jcp.26185. [DOI] [PubMed] [Google Scholar]
- 12.Bian T, Jiang D, Liu J, Yuan X, Feng J, Li Q, Zhang Q, Li X, Liu Y, Zhang J. miR-1236-3p suppresses the migration and invasion by targeting KLF8 in lung adenocarcinoma A549 cells. Biochem Biophys Res Commun. 2017;492:461–467. doi: 10.1016/j.bbrc.2017.08.074. [DOI] [PubMed] [Google Scholar]
- 13.Sui J, Yang RS, Xu SY, Zhang YQ, Li CY, Yang S, Yin LH, Pu YP, Liang GY. Comprehensive analysis of aberrantly expressed microRNA profiles reveals potential biomarkers of human lung adenocarcinoma progression. Oncol Rep. 2017;38:2453–2463. doi: 10.3892/or.2017.5880. [DOI] [PubMed] [Google Scholar]
- 14.Berrout J, Kyriakopoulou E, Moparthi L, Hogea AS, Berrout L, Ivan C, Lorger M, Boyle J, Peers C, Muench S, et al. TRPA1-FGFR2 binding event is a regulatory oncogenic driver modulated by miRNA-142-3p. Nat Commun. 2017;8:947. doi: 10.1038/s41467-017-00983-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yan W, Wu Q, Yao W, Li Y, Liu Y, Yuan J, Han R, Yang J, Ji X, Ni C. MiR-503 modulates epithelial-mesenchymal transition in silica-induced pulmonary fibrosis by targeting PI3K p85 and is sponged by lncRNA MALAT1. Sci Rep. 2017;7:11313. doi: 10.1038/s41598-017-11904-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pan T, Chen W, Yuan X, Shen J, Qin C, Wang L. miR-944 inhibits metastasis of gastric cancer by preventing the epithelial-mesenchymal transition via MACC1/Met/AKT signaling. FEBS Open Bio. 2017;7:905–914. doi: 10.1002/2211-5463.12215. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 17.Huang J, Zheng C, Shao J, Chen L, Liu X, Shao J. Overexpression of eEF1A1 regulates G1-phase progression to promote HCC proliferation through the STAT1-cyclin D1 pathway. Biochem Biophys Res Commun. 2017;494:542–549. doi: 10.1016/j.bbrc.2017.10.116. [DOI] [PubMed] [Google Scholar]
- 18.Hiller J, Hagl B, Effner R, Puel A, Schaller M, Mascher B, Eyerich S, Eyerich K, Jansson AF, Ring J, et al. STAT1 gain-of-function and dominant negative STAT3 mutations impair IL-17 and IL-22 immunity associated with CMC. J Invest Dermatol. 2018;138:711–714. doi: 10.1016/j.jid.2017.09.035. [DOI] [PubMed] [Google Scholar]
- 19.Collins-McMillen D, Stevenson EV, Kim JH, Lee BJ, Cieply SJ, Nogalski MT, Chan GC, Frost RW, III, Spohn CR, Yurochko AD. HCMV utilizes a non-traditional STAT1 activation cascade via signaling through EGFR and integrins to efficiently promote the motility, differentiation, and polarization of infected monocytes. J Virol pii. 2017 doi: 10.1128/JVI.00622-17. JVI.00622-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu R, Lu Z, Gu J, Liu J, Huang E, Liu X, Wang L, Yang J, Deng Y, Qian J, et al. MicroRNAs 15A and 16-1 activate signaling pathways that mediate chemotaxis of immune regulatory B cells to colorectal tumors. Gastroenterology. 2018;154:637–651.e7. doi: 10.1053/j.gastro.2017.09.045. [DOI] [PubMed] [Google Scholar]
- 21.Zhang X, Li X, Tan F, Yu N, Pei H. STAT1 inhibits MiR-181a expression to suppress colorectal cancer cell proliferation through PTEN/Akt. J Cell Biochem. 2017;118:3435–3443. doi: 10.1002/jcb.26000. [DOI] [PubMed] [Google Scholar]
- 22.Zhang M, Du Y, Lu R, Shu Y, Zhao W, Li Z, Zhang Y, Liu R, Yang T, Luo S, et al. Cholesterol retards senescence in bone marrow mesenchymal stem cells by modulating autophagy and ROS/p53/p21Cip1/Waf1 pathway. Oxid Med Cell Longev. 2016;2016:7524308. doi: 10.1155/2016/7524308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Daugherty A, Hegele RA, Mackman N, Rader DJ, Schmidt AM, Weber C. Complying with the national institutes of health guidelines and principles for rigor and reproducibility: Refutations. Arterioscler Thromb Vasc Biol. 2016;36:1303–1304. doi: 10.1161/ATVBAHA.116.307906. [DOI] [PubMed] [Google Scholar]
- 24.Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. doi: 10.1016/j.cell.2004.12.035. [DOI] [PubMed] [Google Scholar]
- 25.Li M, Liang Z, He S, Zeng Y, Jing Y, Fang W, Wu K, Wang G, Ning X, Wang L, et al. Genome-wide identification of leaf abscission associated microRNAs in sugarcane (Saccharum officinarum L.) BMC Genomics. 2017;18:754. doi: 10.1186/s12864-017-4053-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhao S, Gao X, Zang S, Li Y, Feng X, Yuan X. MicroRNA-383-5p acts as a prognostic marker and inhibitor of cell proliferation in lung adenocarcinoma by cancerous inhibitor of protein phosphatase 2A. Oncol Lett. 2017;14:3573–3579. doi: 10.3892/ol.2017.6603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhuang L, Shou T, Li K, Gao CL, Duan LC, Fang LZ, Zhang QY, Chen ZN, Zhang C, Yang ST, Li GF. MicroRNA-30e-5p promotes cell growth by targeting PTPN13 and indicates poor survival and recurrence in lung adenocarcinoma. J Cell Mol Med. 2017;21:2852–2862. doi: 10.1111/jcmm.13198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang Y, Chen Y, Yun H, Liu Z, Su M, Lai R. STAT1β enhances STAT1 function by protecting STAT1α from degradation in esophageal squamous cell carcinoma. Cell Death Dis. 2017;8:e3077. doi: 10.1038/cddis.2017.481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hu Y, Wang L, Gu J, Qu K, Wang Y. Identification of microRNA differentially expressed in three subtypes of non-small cell lung cancer and in silico functional analysis. Oncotarget. 2017;8:74554–74566. doi: 10.18632/oncotarget.20218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li W, Yang W, Liu Y, Chen S, Chin S, Qi X, Zhao Y, Liu H, Wang J, Mei X, et al. MicroRNA-378 enhances inhibitory effect of curcumin on glioblastoma. Oncotarget. 2017;8:73938–73946. doi: 10.18632/oncotarget.17881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhang QL, Zhu QH, Zhang F, Xu B, Wang XQ, Chen JY. Transcriptome-wide analysis of immune-responsive microRNAs against poly (I:C) challenge in Branchiostoma belcheri by deep sequencing and bioinformatics. Oncotarget. 2017;8:73590–73602. doi: 10.18632/oncotarget.20570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yang Y, Ji C, Guo S, Su X, Zhao X, Zhang S, Liu G, Qiu X, Zhang Q, Guo H, Chen H. The miR-486-5p plays a causative role in prostate cancer through negative regulation of multiple tumor suppressor pathways. Oncotarget. 2017;8:72835–72846. doi: 10.18632/oncotarget.20427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yan S, Dang G, Zhang X, Jin C, Qin L, Wang Y, Shi M, Huang H, Duan Q. Downregulation of circulating exosomal miR-638 predicts poor prognosis in colon cancer patients. Oncotarget. 2017;8:72220–72226. doi: 10.18632/oncotarget.19689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Fawzy MS, Toraih EA, Ibrahiem A, Abdeldayem H, Mohamed AO, Abdel-Daim MM. Evaluation of miRNA-196a2 and apoptosis-related target genes: ANXA1, DFFA and PDCD4 expression in gastrointestinal cancer patients: A pilot study. PLoS One. 2017;12:e0187310. doi: 10.1371/journal.pone.0187310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wen L, Li Y, Jiang Z, Zhang Y, Yang B, Han F. miR-944 inhibits cell migration and invasion by targeting MACC1 in colorectal cancer. Oncol Rep. 2017;37:3415–3422. doi: 10.3892/or.2017.5611. [DOI] [PubMed] [Google Scholar]
- 36.He Z, Xu H, Meng Y, Kuang Y. miR-944 acts as a prognostic marker and promotes the tumor progression in endometrial cancer. Biomed Pharmacother. 2017;88:902–910. doi: 10.1016/j.biopha.2017.01.117. [DOI] [PubMed] [Google Scholar]
- 37.Liu Y, Zhang J, Sun X, Su Q, You C. Down-regulation of miR-29b in carcinoma associated fibroblasts promotes cell growth and metastasis of breast cancer. Oncotarget. 2017;8:39559–39570. doi: 10.18632/oncotarget.17136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Borsini A, Cattaneo A, Malpighi C, Thuret S, Harrison NA, MRC ImmunoPsychiatry Consortium. Zunszain PA, Pariante CM. Interferon-alpha reduces human hippocampal neurogenesis and increases apoptosis via activation of distinct STAT1-dependent mechanisms. Int J Neuropsychopharmacol. 2018;21:187–200. doi: 10.1093/ijnp/pyx083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Carbotti G, Nikpoor AR, Vacca P, Gangemi R, Giordano C, Campelli F, Ferrini S, Fabbi M. IL-27 mediates HLA class I up-regulation, which can be inhibited by the IL-6 pathway, in HLA-deficient small cell lung cancer cells. J Exp Clin Cancer Res. 2017;36:140. doi: 10.1186/s13046-017-0608-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang Y, Liu Z. STAT1 in cancer: Friend or foe? Discov Med. 2017;24:19–29. [PubMed] [Google Scholar]
- 41.Xi J, Huang Q, Wang L, Ma X, Deng Q, Kumar M, Zhou Z, Li L, Zeng Z, Young KH, et al. miR-21 depletion in macrophages promotes tumoricidal polarization and enhances PD-1 immunotherapy. Oncogene. 2018;37:3151–3165. doi: 10.1038/s41388-018-0178-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li J, He X, Wei W, Zhou X. MicroRNA-194 promotes osteoblast differentiation via downregulating STAT1. Biochem Biophys Res Commun. 2015;460:482–488. doi: 10.1016/j.bbrc.2015.03.059. [DOI] [PubMed] [Google Scholar]
- 43.Li L, Li W, Chen N, Zhao H, Xu G, Zhao Y, Pan X, Zhang X, Zhou L, Yu D, et al. FLI1 exonic circular RNAs as a novel oncogenic driver to promote tumor metastasis in small cell lung cancer. Clin Cancer Res. 2018 Nov 14; doi: 10.1158/1078-0432.CCR-18-1447. (Epub ahead of print) [DOI] [PubMed] [Google Scholar]
- 44.Gujam FJ, McMillan DC, Edwards J. The relationship between total and phosphorylated STAT1 and STAT3 tumour cell expression, components of tumour microenvironment and survival in patients with invasive ductal breast cancer. Oncotarget. 2016;7:77607–77621. doi: 10.18632/oncotarget.12730. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The analyzed datasets generated during the study are available from the corresponding author on reasonable request, while preserving the necessary confidentiality and anonymity.