The role of non-coding RNAs in the development and maintenance of sex-specific characteristics in Drosophila is not well understood. Iftikhar et al. present results of a genetic screen demonstrating...
Keywords: male courtship, pheromones, miRNA, behavior, 7-T, cVA, Genetics of Sex
Abstract
Drosophila melanogaster courtship, although stereotypical, continually changes based on cues received from the courtship subject. Such adaptive responses are mediated via rapid and widespread transcriptomic reprogramming, a characteristic now widely attributed to microRNAs (miRNAs), along with other players. Here, we conducted a large-scale miRNA knockout screen to identify miRNAs that affect various parameters of male courtship behavior. Apart from identifying miRNAs that impact male–female courtship, we observed that miR-957 mutants performed significantly increased male–male courtship and “chaining” behavior, whereby groups of males court one another. We tested the effect of miR-957 reduction in specific neuronal cell clusters, identifying miR-957 activity in Doublesex (DSX)-expressing and mushroom body clusters as an important regulator of male–male courtship interactions. We further characterized the behavior of miR-957 mutants and found that these males court male subjects vigorously, but do not elicit courtship. Moreover, they fail to lower courtship efforts toward females with higher levels of antiaphrodisiac pheromones. At the level of individual pheromones, miR-957 males show a reduced inhibitory response to both 7-Tricosene (7-T) and cis-vaccenyl acetate, with the effect being more pronounced in the case of 7-T. Overall, our results indicate that a single miRNA can contribute to the regulation of complex behaviors, including detection or processing of chemicals that control important survival strategies such as chemical mate-guarding, and the maintenance of sex- and species-specific courtship barriers.
WE understand today that innate behaviors, essential for an animal’s survival and reproductive success, are often controlled by genes that operate in robust neural circuits (de Bono and Bargmann 1998; Sokolowski 2001; Lim et al. 2004; Kohl et al. 2013). It is also well understood that differences in the development of neural architecture can result in the behavioral diversification of innate behaviors, as in the case of sexually dimorphic behaviors (Kohl et al. 2013). How these robust behaviors are fine-tuned in response to rapidly changing environmental stimuli remains vaguely answered. What are the key players that regulate the balance between maintenance of the major characteristics of innate behaviors and the acquisition of appropriate stimulus-dependent adaptations? Traditionally, proteins involved in mechanisms such as transcriptional regulation and the enzymatic components of signal transduction pathways have mostly been associated with such adaptive responses (Stock et al. 1989; Abraham 2008; Zovkic et al. 2013). However, recently microRNAs (miRNAs) have emerged as important mediators of rapid and widespread changes in cells’ transcript and proteomic content in response to environmental fluctuations (Leung and Sharp 2010; Nesler et al. 2013). miRNAs are 19–20 nucleotide, noncoding RNAs that target messenger RNA (mRNA) transcripts for translation inhibition, making miRNA-mediated translational control a fast mode of protein expression regulation (Iftikhar and Carney 2016). miRNAs act by binding to the 3′-UTR of their target mRNAs and disrupting translation via the formation of RISC (RNA-Induced Silencing Complex). Complex formation can be followed by any of several proposed modes of miRNA-mediated translation inhibition, such as recruitment of deadenylation factors that destabilize the mRNA–miRNA duplex by removing the poly(A) tail or by blocking the circularization of mRNA that facilitates translation initiation (Carthew and Sontheimer 2009).
It is known that miRNAs are expressed in the central nervous system (CNS) (Morel et al. 2013; You et al. 2018), and in the past few years, studies have started to emerge that elaborate upon on this newer potential regulatory mechanism in the context of how miRNA activity can modulate behavior. Some examples of work that sheds light on roles for miRNAs in regulating behavior include studies on honeybees demonstrating that differential expression of miRNAs is linked to behavioral plasticity (Greenberg et al. 2012). Work on zebra finches shows that miR-9 and miR-140-5p are upregulated as a function of song behavior in the brain regions associated with vocal learning (Shi et al. 2013). In Drosophila melanogaster, recent reports have begun to describe the role of miRNAs in behaviors such as learning, memory, and circadian rhythms (Busto et al. 2015; You et al. 2018). An example is miR-92a, which modulates the excitability of pacemaker neurons that regulate circadian rhythms (Chen and Rosbash 2017). A further piece of evidence for the contribution of miRNAs to Drosophila behavior comes from the work of Picao-Osorio et al. (2017), which demonstrated that a single Drosophila miRNA (miR-iab4) affects larval self-righting (SR) behavior, a movement used by the larva to correct its orientation if turned upside-down. In their screen for SR behavior, a surprising 40% of all miRNAs tested impacted the behavior (Picao-Osorio et al. 2017). Interestingly, miR-iab4/iab-8 mutants have been previously reported to be sterile. The phenotype was shown to be behavioral since males were unable to achieve copulation with females as a result of being unable to fully bend their abdomens. Females were also defective for egg-laying behavior because of a muscular defect that prevented them from laying eggs or curling their abdomens (Bender 2008).
We sought to understand the role of miRNAs in the regulation of another innate but stimulus-modulated behavior displayed by Drosophila males: courtship behavior. Drosophila courtship is comprised of a series of robust, stereotypical steps, but is constantly modified based on information received from the courtship subject (Ejima et al. 2005) that includes, for example, female rejection behaviors, and aversive or attractive pheromones. The male initiates courtship by orienting himself toward the female upon receiving visual and/or olfactory cues. This step is typically followed by the male approaching the female and tapping her, allowing the male to sense the nonvolatile pheromones on the female’s body using gustatory receptors on the male’s forelegs. The male then performs what is termed “singing behavior,” which is comprised of unilateral wing extension and vibration by the male. The male then assesses the quality of his mate by licking her. The effectiveness of the male’s courtship efforts is essential in convincing the female to mate with him. The male continues to chase the female while singing and also bends his abdomen several times in attempts to mount the female. This effort continues until the receptive female bends her ovipositor such that the male mounts her and achieves copulation. Therefore, there is a constant exchange of visual, olfactory, gustatory and, auditory information between the male and female flies that allows them to contextually modify their behavior (Sokolowski 2001).
The potential for Drosophila to perform sex-specific reproductive behaviors is set by the production of the transcriptional regulator proteins Fruitless (FRU) and Doublesex (DSX). The sex-determination gene hierarchy is differentially initiated in females and males due to female-limited expression of Sex-lethal (Sxl), resulting in sex-specific splicing of transformer (tra). Expression of tra, in turn, determines the sex-specific expression of fru and dsx transcripts, and their protein products (Dauwalder et al. 2002). Expression of the male-specific form of the FRU protein (FRUM) enables male-specific courtship and mating behaviors. Similarly, sex-specific splicing of the primary transcript for dsx leads to the production of either a male- or female-specific protein isoform of DSX, DSXM and DSXF, respectively (Yamamoto and Koganezawa 2013). FRU proteins are proposed to transcriptionally regulate several target genes through chromatin modification, thereby turning on the genetic program required for masculinization in FRUM-expressing neurons (Ito et al. 2012). Neurons that express FRUM develop male typical structures and projections, whereas in the absence of FRUM, only female-type neurons develop (Kimura et al. 2005; Yamamoto and Koganezawa 2013). These sex-specific differences in the number and architecture of neurons occur in specific groups of cells, such as the PC1 cluster in the posterior region of the adult brain. Several studies collectively suggest that the PC1 neuron cluster predominantly controls the initiation of male courtship behavior (Kimura et al. 2005, 2008; Kohatsu et al. 2011). The male-specific development of this cluster requires not only the presence of FRUM but also the absence of DSXF, since the presence of DSXF results in female-specific cell death induction (Kimura et al. 2008). Reports demonstrating that artificial activation of fru-expressing or dsx-expressing neurons induces male courtship behavior in solitary males have clarified the necessity of both fru and dsx for displaying the full complement of male courtship behaviors (Pan et al. 2011).
In addition to what is known about the role of proteins of the sex-determination hierarchy, recent work highlights the possible role of miRNAs in maintaining sexual identity (Fagegaltier et al. 2014). The let-7 miRNA was shown to restrict the expression of sex-determination genes to the sex in which they are normally expressed. Fagegaltier et al. (2014) found that the levels of a downstream target of DSXF, Yp1, are reduced in female flies mutant for let-7. In male let-7 mutants, the researchers detected spurious expression of female-specific transcripts Sxl and Yp1, indicating that let-7 regulates sex-specific expression levels of these two transcripts as well. The role of let-7 in sex determination is also dependent on signaling by the insect steroid hormone ecdysone. Ecdysone signaling regulates a vast array of biological responses in Drosophila, including courtship behaviors (Dalton et al. 2009). Thus, steroid-coupled regulation of miRNAs that interact with proteins of the sex-determination hierarchy suggests a possible role for miRNAs in regulating not only physical sexual characteristics but also reproductive behaviors.
We were therefore interested in the extent to which miRNAs influence sex-specific behaviors such as male courtship. Thus far, one study has identified a role for miR-124 in suppressing male–male courtship and enhancing male attractiveness to females. miR-124 mutant males have elevated levels of traF and altered levels of male-specific pheromones (Weng et al. 2013). In our study, we conducted a genetic screen to identify miRNAs that influence D. melanogaster male–female courtship behavior. We identified numerous miRNAs that modulate behavior and confirmed two miRNAs, miR-263b and miR-317, that affect stereotypical parameters of male–female courtship, and one, miR-957, that suppressed male–male courtship. Our further characterization of the courtship behavior of miR-957 mutants indicates that the males show a reduced inhibitory response to animals with high levels of male-aversive pheromones on their bodies. Therefore, miRNAs appear to play a role in modulating sex-specific responses to pheromones.
Materials and Methods
Fly stocks
A library of 80 targeted miRNA knockout (KO) strains covering 104 miRNA genes (Chen et al. 2014) was obtained from the Bloomington Drosophila Stock Center (BDSC) (http://fly.bio.indiana.edu/). The combined set of strains deletes 130 miRNA-encoding loci either singly or in clusters. w1118 flies obtained from the BDSC were used as controls (Chen et al. 2014). Three strains in the yw background (removing miR-927, miR-210, or miR-13-b-2) were not tested in the screen since yellow mutants have been shown to have abnormal courtship behaviors (Wilson et al. 1976), and we found during our testing that mutants in the yw background had low courtship. miRNA sponge lines [UAS-miR-957.sponge/CyO; UAS-miR-957.sponge (BDSC#61443), UAS-miR263b.sponge/CyO; UAS-miR-263b.sponge (BDSC#61403), UAS-miR-317.sponge/CyO; UAS-miR-317.sponge.TM6B,Tb1 (BDSC#61434), UAS-miR-31a.sponge; UAS-miR-31a.sponge (BDSC#61383), and UAS-scramble.sponge; UAS-scramble.sponge (control)] (BDSC#61507) (Fulga et al. 2015) and miRNA expression lines [UAS-mir-957/CyO (BDSC#60609), UAS-miR-317/CyO (BDSC#59913), UAS-miR-310/TM3, Sb (BDSC#41155), UAS-miR-310,311,312,313/TM3, Sb (BDSC#41135), UAS-miR-31a/TM3, Sb, and Ser (BDSC#59869)] (Bejarano et al. 2012) were also obtained from the BDSC. The driver lines used were dsx-gal4/TM6B, Tb (BDSC#66674), fru-gal4/TM3, Sb (gift from Barry Dickson), elav-gal4/CyO (BDSC#8765), Ubi-gal4/CyO (BDSC#32551), Gr32a-gal4 (BDSC#57622) Gr39b-gal4 (BDSC#57635), and actin-gal4/SM6b. Chromosomal deficiency (Df) lines that removed specific miRNAs were also obtained from the BDSC [miRNA-310,311,312,313: Df(2R)BSC701/SM6a (BDSC#51327), miRNA-317: Df(3R)ED5454/TM6C, cu, Sb (BDSC#9080), miRNA-263b: Df(3L)BSC575/TM6C, Sb, cu (BDSC#27587), miRNA-31a: Df(2R)BSC347/CyO (BDSC#24371), and miRNA-957: Df(3L)BSC420/TM6C, Sb, cu (BDSC#24924)]. D. simulans were a gift from Mariana Mateos (Texas A&M University). Cre D[*]/TM3, Sb (BDSC#851), and mushroom body (MB) driver D52H-gal4 (on the X chromosome) were provided by Paul Hardin (Texas A&M University). Flies were maintained at 25° on a 12-hr light/dark cycle on a standard cornmeal, sugar, and agar diet.
Screen design
We screened miRNA KO strains (in the w1118 background) for defects in male courtship behavior. w1118 flies were used as controls. Out of the 80 strains available, only the 60 KO strains that produced homozygotes were selected for further evaluation (Figure 1A). During the course of the screen, stocks that showed very low viability of homozygotes were also dropped if a sufficiently large sample size was hard to attain. Therefore, 57 total strains were tested (Supplemental Material, Table S1). Single-pair mating assays (described below) were conducted in batches of 5–10 genotypes, and a control group of w1118 flies was tested in each batch. A sample size of at least 25 pairs was obtained for each genotype. All assays were conducted in dark conditions to remove the influence of visual cues on mating behavior. The parameters scored for all lines were courtship latency, mating latency, and mating success. Courtship latency is defined as the time from the introduction of the courtship subject to the beginning of courtship. Mating latency is calculated as the elapsed time from the introduction of the female subject to the start of mating. Mating success is the proportion of pairs that mate within the 1-hr timeframe for the assay.
Figure 1.
Selection of miRNA KO strains and results summary for original screen. (A) Eighty miRNA KO strains (Chen et al. 2014) in the w1118 background were selected for a screen to evaluate effects of miRNA KO alleles on male courtship behaviors. After removing those strains that did not survive as homozygotes (20 strains) or had low viability as homozygotes (3 strains), 57 strains were used to compare courtship parameters with those of a control strain. (B) Number of strains showing significant differences in the behavioral parameters tested. CI was calculated for only three strains, out of which one strain had significantly altered CI. CI, courtship index; KO, knockout; miRNA, microRNA.
From the videos of the single-pair mating assays obtained during the screen, courtship index toward the female subjects was calculated for miR-263b, miR-278, and miR-957 KO males because their courtship appeared to be very vigorous in the assay recordings. miR-957 KO males also displayed intermale courtship when kept in groups of males. Courtship index is defined as the proportion of time spent by the male courting the courtship subject as a proportion of the total time during which the behavior is evaluated. In this study, courtship index was calculated for the first 10 min from the introduction of the courtship subject unless otherwise noted. w1118 males were used for control comparisons.
For strains that showed a significantly altered value for one or more of the parameters mentioned above, validation of the phenotype was performed by placing each miRNA KO allele in trans with a deficiency chromosome that removed the miRNA. Phenotypes observed in homozygous miRNA KO animals were evaluated in these transheterozygous animals.
Behavioral testing
Single-pair mating assays:
A 5-day-old virgin male (miRNA mutant for the experimental condition and w1118 for the control) was placed in a mating chamber with a 5-day-old virgin female (Canton-S) in dark conditions at room temperature. Behavior was recorded for 1 hr using JVC-HDD Everio cameras, and we calculated the following parameters—courtship latency, mating latency, courtship index, and mating success—for each pair. The small number of pairs that did not court or mate within the hour were not included in our calculations. All experiments were carried out over several days, and control and experimental animals were always tested simultaneously.
Removal of mini-white from the miR-957 KO strain:
w[*]; TI{w[+mW.hs]=TI}mir-957[KO] males were crossed to y[1] w[67c23] P{y[+mDint2]=Crey}1b; D[*]/TM3, Sb[1] females. White-eyed progeny were collected and crossed twice with w1118 flies to confirm the removal of the mini-white gene. The flies were then backcrossed six times into the Canton-S (TM3, Sb/TM6B, and Tb) background and tested for male–male courtship. TM3, Sb/TM6B, and Tb flies were used as control animals for subsequent assays with the backcrossed miR-957 KO males.
Evaluating male courtship behavior of miR-957 KO grouped males:
Virgin miR-957 KO males were collected within 4 hr of eclosion and aged for 10 days. On the 10th day, males were aspirated into food vials in groups of 10 males 2 hr before “lights on.” The next day the males were videoed for 10 min at lights on (Villella et al. 1997). Within the group, miR-957 KO males were observed courting in pairs (several pairs courting simultaneously), as well as in groups of three or more males (known as chaining). Since males rapidly switched from courting in pairs to courting in chains, we termed the overall behavior “total courtship and chaining index” (tCCI). tCCI was calculated as the proportion of the assay time males spent courting in either chains or in multiple pairs. The same protocol was followed for observing courtship among males that were transheterozygous for the miR-957 KO mutation over a corresponding deficiency.
Single-pair male–male courtship assays:
A 5-day-old virgin male (miR-957 KO or control) was placed in a mating chamber with a 5-day-old, virgin, decapitated Canton-S male in dark conditions at room temperature. Behavior was recorded for 20 min using JVC-HDD Everio cameras. We compared the courtship of miR-957 KO or control males toward the decapitated males.
Single-pair elicitation assays:
A 5-day-old virgin Canton-S male was placed in a mating chamber with a 5-day-old, virgin, decapitated miR-957 KO male in dark conditions at room temperature. Behavior was recorded for 20 min. For control experiments, a decapitated Canton-S male was placed in the chamber as the courtship object.
Two-choice assays:
A 5-day-old virgin male (courter) was placed in a mating chamber with two 5-day-old, virgin, decapitated subjects: a Canton-S male and a similarly treated Canton-S female. The percentage of time the courter spent courting each subject in the first 10 min of the assay was recorded. The subject that was courted for the greater percentage of time was scored as the preferred subject.
Single-pair courtship assays with D. simulans females or D. melanogaster mated females:
Courtship index toward D. simulans females or D. melanogaster mated females:
A 5-day-old virgin male (miR-957 KO or control) was placed in a mating chamber with either a 5-day-old, virgin, decapitated D. simulans female or a D. melanogaster (Canton-S) female in dark conditions at room temperature. In a second experiment, individual males were presented with a 5-day-old virgin or mated D. melanogaster female. Behavior was recorded for 30 min. Courtship index was calculated for the final 10 min of the assay.
Assessment of proportion of flies displaying courtship behavior:
Pairs of flies in the above described assay were also evaluated for the presence or absence of courtship (chasing, wing extensions, and attempted copulation) for each minute of the last 5 min of the 30-min recording. This information was used to calculate the proportion of males that continued to court D. simulans females or mated D. melanogaster females after > 20 min of exposure to the female. Prior work from our laboratory demonstrated that Canton-S males significantly decrease courtship toward virgin D. simulans females, but not toward virgin D. melanogaster females, within 20 min of exposure (Ellis and Carney 2009). Males that did not court (four males paired with D. simulans females) or that mated within the timeframe of the assay (six pairs spread across all pairings) were removed from the analysis.
Sponge experiments:
UAS-miR.sponge lines for miR-263b, miR-317, or miR-957 were crossed with actin-gal4 to observe the effects of ubiquitous repression of the miRNAs. These miRNAs were specifically tested because the phenotypes observed for their KO mutants were validated with deficiency lines. Males expressing a UAS-scramble.sponge sequence were used as controls for every case (Fulga et al. 2015). For actin-gal4 > UAS-miR-263b.sponge males, single-pair mating assays were performed with females in the dark and the courtship index was calculated. For actin-gal4 > UAS-miR-317.sponge males, single-pair mating assays with females were performed in the dark and mating success was calculated. For actin-gal4 > UAS-miR-957.sponge males, tCCI was calculated for grouped males. In each case, the results were compared to actin-gal4 > UAS-scramble.sponge males.
Rescue experiments:
For miR-957, expression was restored using elav-gal4 to drive the expression of UAS-miR-957 in neurons in the miR-957 KO background. tCCI was calculated for the rescue animals and compared with miR-957 KO flies. Similarly, elav-gal4 was used to express UAS-miR-317 in the miR-317 KO/Df background. Single-pair courtship assays were performed to calculate mating success for the miR-317 rescue flies and compared with miR-317 KO/Df flies. Rescue experiments were not performed for miR-263b KO because overexpression lines were not available. We attempted to rescue expression for miR-957 and miR-317 using the ubiquitous drivers ubi-gal4 and actin-gal4, but expression with these drivers resulted in lethality in KO backgrounds for both miRNAs.
Perfuming experiments:
5-day-old virgin, Canton-S females were lightly anesthetized with CO2 and coated with 1 μl (per fly) of the pheromone solution to be tested. cis-vaccenyl acetate (cVA) was diluted at a concentration of 3 μg/ul in ethanol and 7-Tricosene (7-T) was diluted in hexane at a concentration of 3 μg/μl. Both 7-T (≥ 95% pure) and cVA (> 98% pure) were obtained from Cayman Chemical. Females to be used as controls were coated with 1 μl of the respective solvents for each pheromone. The flies were then allowed 15–20 min on the food vial to groom and recover from anesthesia, and were then decapitated. The perfumed and decapitated female subjects were then placed in mating chambers with either Canton-S or miR-957 KO 5-day-old virgin males for 30 min. Courtship index for each pair was calculated for the final 10 min of the assay.
Activity testing
For targets from the screen that were selected for further evaluation, general activity levels were compared for miRNA mutants and controls by recording the frequency at which the males crossed a line through the center of the mating chamber in the final minute before mating was achieved (McRobert et al. 2003). We did not detect activity differences for miRNA males compared to controls (Figure S1).
Statistics
Statistical analysis for behavior was done using JMP Pro statistical software (JMP Pro, Version <13.1.0>, SAS Institute, Cary, NC, 1989–2007). GraphPad Prism version 7.00 for MAC OS X (GraphPad Software, La Jolla, CA; www.graphpad.com) was used for making graphs. Values obtained for courtship latency and mating latency belonged to log normal distributions, and were therefore log transformed to obtain normally distributed values. ANOVA followed by a post hoc Dunnett’s test was performed for the transformed courtship latency and mating latency values for each batch of genotypes tested. Courtship indices were arcsine transformed to obtain normally distributed values and compared using a Student’s t-test to compare between two groups. A nonparametric Mann–Whitney U-test was used to compare the courtship index between two samples when the test for normality failed after transformation. The χ2 statistic was calculated to compare the mating success of two samples. For perfuming experiments, a two-way ANOVA was applied to test for interaction between male genotype and type of female. A one-way ANOVA followed by a post hoc Tukey–Kramer test was used for multiple comparisons (Billeter and Levine 2015).
Data availability
Table S1 contains phenotypic data and statistical analyses of miRNA KO mutants. Figure S1 contains activity measurements of miRNA KO mutant screen candidates. Figure S2 shows fru-gal4 expression in the brain. Figure S3 shows effects of overexpression of miR-957. Figure S4 is a two-choice test between perfumed and wild-type females. Supplemental material available at Figshare: https://doi.org/10.25386/genetics.7522490.
Results
miRNA KO screen
In the initial screen, we identified significant effects resulting from the removal of individual or clusters of miRNAs on several robust steps of male Drosophila courtship behavior (Table 1 and Table S1). In total, 31 of 57 KO strains tested in the original screen affected one or more of the phenotypes evaluated. For 16 strains, we detected effects on mating success (Figure 1B). miR-1017 males did not mate (N = 20) and males from a single KO strain, miR-285, had a higher mating success than the control males (N = 41, P = 0.0105). Out of the 15 KO strains for which males had reduced mating success, three strains (miR-317, miR-31a, and miR-310-311-312-313) also displayed increased mating latency, meaning that males from these strains took longer to achieve copulation (miR-317: N = 26, P = 0.0295; miR-31a: N = 24, P = 0.0327; and miR-310-311-312-313: N = 27, P = 0.0143). No other strains from the 57 strains tested had differences in mating latency. However, miR-31a mutants also took longer to initiate courtship toward females (N = 24, P = 0.0368).
Table 1. miRNA KO mutants tested for courtship parameters.
| miRNA | Phenotype observed in KO | Phenotype with transheterozygote (KO/Df) | Phenotype observed in both mutants |
|---|---|---|---|
| miR-263b | Increased courtship index (N = 27, P < 0.0001) | Increased courtship index (N = 29, P < 0.00001) | ✔ |
| miR-957 | Male–male courtship and chaining (N = 10 × 3, P = 0.0024) | Male–male courtship and chaining (N = 10 × 3, P = 0.0189) | ✔ |
| miR-317 | Increased mating latency (N = 26, P = 0.0295) | Mating latency not affected (N = 30, P = 0.6615) | X ✔ |
| Lower mating success (N = 50, P = 0.01508) | Lower mating success (N = 52, P = 0.004973) | ||
| miR-310,311,312,313 | Increased mating latency (N = 27, P = 0.0143) | Mating latency not affected (N = 29, P = 0.6362) | |
| Lower mating success (N = 70, P = 0.000001) | Mating success not affected (N = 35, P = 0.8286) | ||
| miR-31a | Increased courtship latency (N = 24, P = 0.0368) | Courtship latency not affected (N = 30, P = 0.2472) | X |
| Increased mating latency (N = 24, P = 0.0327) | Mating latency not affected (N = 30, P = 0.3103) | ||
| Lower mating success (N = 66, P = 0.000001) | Mating success not affected (N = 30, P = 0.126) | ||
| miR-285 | Higher mating success (N = 41, P = 0.0105) | Mating success not affected (N = 49, P = 0.5632) | X |
| miR-1017 | No mating (N = 20) | Mating success not affected (N = 20, P = 0.3865) | X |
miRNA KO mutants with significant variations in courtship and/or mating parameters were further tested as KO/Df transheterozygotes. miR-957 KO mutants were backcrossed into the Canton-S background after removal of the mini-white gene, and then tested for male–male courtship and chaining behavior. Df, chromosomal deficiency; KO, knockout; miRNA, microRNA.
For courtship latency, we detected differences in both directions (increased as well as decreased courtship latency) (Figure 1B). Overall, nine miRNA mutant strains displayed a significant increase in courtship latency and 10 strains had reduced courtship latency (Table S1).
We noticed that miR-263b mutants appeared to court females extremely vigorously, though we did not detect a difference in the mating latency. We then calculated the courtship index for this strain and found that it was increased (Mann–Whitney U-test, N = 27, P < 0.0001).
Validation of candidates
We selected candidates from the initial screen to determine if the observed phenotypes were maintained when we placed the KO allele in trans with a Df for the miRNA (Table 1). From the strains with reduced mating success, we selected those that also had increased mating latency and/or increased courtship latency for validation with deficiencies. Namely, these were miR-317, miR-31a, and miR-310-311-312-313. Transheterozygotes for miR-317, like the KO mutants, had lower mating success (χ2 test, N = 52, P = 0.004973). However, they did not have a significant increase in mating latency. Heteroallelic mutants for miR-31a, and miR-310-311-312-313 did not affect mating success or mating latency.
miR-263b KO mutant males had a higher courtship index than controls, as did miR-263b transheterozygous mutants (Mann–Whitney U-test, N = 29, P < 0.00001). miR-285 and miR-1017 KO/Df males did not have the originally observed phenotypes.
Finally, we further validated the phenotypes observed in miR-317 and miR-263b KO mutants by expressing miRNA sponges to reduce the expression of miR-317 and miR-263b under the control of the ubiquitous actin-gal4 driver. For actin-gal4 > UAS-miR-317.sponge males, mating success was significantly lower than for the actin-gal4 > UAS-scramble.sponge controls (N = 36, P = 0.0406). For actin-gal4 > UAS-miR-263b.sponge males, courtship index was higher than for the actin-gal4 > UAS-scramble.sponge controls, although the difference was not significant (N = 28, P = 0.0594).
Expression of miR-317 was restored via the pan-neuronal driver elav-gal4 in the miR-317 KO background, and the reduced mating success phenotype was not rescued (N = 40, P = 0.1791). We were unable to test for rescue by ubiquitous expression of the miRNA due to lethality.
Characterization of miR-957 KO grouped male behavior
As a part of the effort to identify aberrations in the courtship behavior of the strains being tested in our screen, in addition to performing male–female single-pair courtship assays, we also looked for signs of male–male courtship among grouped males of each strain. Most strikingly, we noticed that groups of miR-957 KO males displayed high levels of male–male courtship. Since the original miR-957 KO allele contains a copy of the mini-white gene, which is known to increase male–male courtship interactions in some contexts (Krstic et al. 2013), we removed the mini-white locus and evaluated the resulting miR-957 KO males for male–male courtship. After removal of mini-white, miR-957 KO males courted one another vigorously, while control males performed little male–male courtship (three groups of 10 males for each genotype; N = 30 per genotype; Mann–Whitney U-test, P = 0.0024, Figure 2 and Table 1)
Figure 2.
tCCI in grouped miR-957 KO males. (A) tCCI for miR-957 KO mutants, miR-957 KO/Df transheterozygotes, and flies with neuronal rescue of miR-957 KO compared with respective controls. Plotted tCCI values are the average of tCCI values for three groups of 10 males each for every genotype. tCCI values were compared between mutants and controls using Student’s t-tests. (B) miR-957 KO males displaying pairwise courtship or chaining. Error bars represent mean ± SE (SEM). Df, chromosomal deficiency; KO, knockout; tCCI, total courtship and chaining index.
When 10-day-old males were stored in food vials in groups of 10 males, they courted vigorously. Two patterns of behaviors were observed. First, several pairs of males were seen courting simultaneously. Pairwise courtship was intermixed with chaining behavior (groups of three or more males courting each other) (Figure 2B). Since the males frequently shifted from courting in pairs to courting in chains, we scored the behavior as a composite of pairwise courtship and chaining behavior, and termed it tCCI. miR-957 KO males showed significantly higher amounts of courtship compared to their wild-type counterparts (Figure 2A and Table 1). miR-957 transheterozygous males courted males vigorously, while control males displayed little male–male courtship (three groups of 10 males each, N = 30 for each genotype; P = 0.0189; Table 1). Pan-neural expression of miR-957 via elav-gal4 in KO males significantly reduced male–male courtship behavior (Figure 2A), suggesting a role for miR-957 neural expression in inhibiting male–male courtship interactions.
miR-957 reduction in dsx and MB neurons increases male–male courtship
To further validate the male–male courtship phenotype as well as gain insight into the location of miR-957 function, we used miRNA sponges to tissue-specifically reduce miR-957 (Figure 3). We first tested the miR-957 sponge (Fulga et al. 2015) by ubiquitously expressing it using an actin-gal4 driver. Males with ubiquitous reduction of miR-957 performed significantly more courtship toward males than actin-gal4 > UAS-scramble.sponge control males. We then checked if neuronal repression of miR-957 with the pan-neuronal driver elav-gal4 had a similar effect. The elav-gal4 > UAS-miR-957.sponge flies actively courted males, whereas very low courtship was observed for scramble controls (Figure 3).
Figure 3.
tCCI (total courtship and chaining index) for grouped males with tissue-specific reduction of miR-957. Ubiquitous (actin-gal4), pan-neuronal (elav-gal4), or neuron subpopulation (via fru-gal4, dsx-gal4, or D52H-gal4) expression of UAS-miR-957.sponge. Plotted tCCI values are the average of tCCI values for three groups of 10 males each for every genotype. tCCI values for each miR-957.sponge and gal4 combination were compared to the respective scramble control using Student’s t-tests. Error bars represent mean ± SE (SEM).
To further explore subpopulations of neurons requiring miR-957 activity to inhibit male–male interactions, we used drivers that allowed us to express the sponge in neurons expressing two well-known regulators of courtship behavior in Drosophila: dsx and fru. dsx and fru produce transcription factors that are crucial for cell sexual identity, as well as sex-specific behaviors and secondary sexual characteristics (Rideout et al. 2010; Pavlou and Goodwin 2013). To our surprise, while we observed very high levels of male–male courtship in groups of dsx-gal4 > UAS-miR-957.sponge flies, we detected little male–male courtship among fru-gal4 > UAS-miR-957.sponge males (we verified that the fru-gal4 driver was functional by testing it with UAS-gfp.nls; Figure S2). Since information from receptor neurons for sexual cues is ultimately relayed to higher-order processing centers, such as the MB via synapses formed in the glomeruli of the antennal lobe (Ziegler et al. 2013; Schultzhaus et al. 2017), we also tested for effects of reducing miR-957 in the MB using D52H-gal4 (Yi et al. 2013). Indeed, reducing miR-957 expression in the MB also resulted in high levels of male–male courtship behavior (Figure 3).
To gain additional insight into miR-957 function, we overexpressed miR-957 in the neurons where we tested the effect of its reduction to determine if increasing miR-957 levels, which should further decrease levels of its mRNA targets, also produces a male–male courtship phenotype. Overexpression of miR-957 did not result in a significant increase in male–male courtship with any of the drivers used previously to reduce miR-957 levels (Figure S3).
miR-957 mutant males court other males but do not elicit courtship
We next wanted to understand why miR-957 KO males court one another. The courtship could be a consequence of changes in the pheromonal profile of miR-957 mutant males that made them more attractive to other males, hence eliciting courtship. Another explanation could be a neurophysiological change that caused miR-957 males to direct courtship efforts toward male subjects. To distinguish between the two possibilities, we performed courtship and elicitation assays separately (Figure 4). The courtship of miR-957 KO males toward decapitated Canton-S subjects was significantly higher than that of control males (Mann–Whitney U-test, N1 = 30, N2 = 25, P = 0.005; Figure 4A). However, there was no significant difference between the courtship elicited by control or miR-957 KO males from Canton-S males (Figure 4B). These results indicate that the courtship observed among miR-957 KO grouped males was more likely a consequence of changes in their neural physiology than a change in their pheromonal profile.
Figure 4.
miR-957 KO courtship and elicitation assays. (A) Courtship of miR-957 KO (green thorax) or control males toward decapitated Canton-S male subjects. (B) Courtship elicited by control or miR-957 KO males from Canton-S males. The possibility that miR-957 mutant males have altered pheromone profiles is indicated by the purple coloration of the pheromone. CI values for experimental and control groups were compared using the Mann–Whitney U-test. N = 25–30 for each genotype. Error bars represent mean ± SE (SEM). CI, courtship index; KO, knockout.
miR-957 KO males prefer female over male subjects
We predicted that miR-957 mutant males court other males due to their inability to correctly perceive pheromones, which suggested three possibilities: (1) miR-957 mutant males have lost the ability to correctly perceive pheromones in general, causing them to lose their ability to distinguish between males and females; (2) miR-957 mutant males have altered perception of male inhibitory pheromones; or (3) they are attracted more strongly to a male stimulatory cue.
We performed competition assays to help assess the first possibility. If males generally do not perceive chemical cues correctly, their ability to distinguish males from females should be impacted. When miR-957 mutant males were presented with a decapitated wild-type male and a decapitated female simultaneously under dark conditions, control and miR-957 KO males both preferred females, indicating that miR-957 KO males have not lost the ability to differentiate females from males (Figure 5).
Figure 5.
miR-957 mutant males prefer females. The proportion of miR-957 knockout and control males that choose to court females or males for the majority of a 10-min two-choice assay. Results were compared using a χ2 test. N = 18 (control) and N = 21 (miR-957).
miR-957 KO males show a reduced inhibitory response to animals with high levels of male-aversive pheromones
The results of the competition assays indicated that miR-957 KO males respond appropriately to some chemical cues since they still prefer female courtship subjects. However, mutant males may not detect male inhibitory cues correctly. We next tested the response of miR-957 KO males toward two types of female subjects (mated D. melanogaster females or D. simulans females) known to have higher amounts of pheromones that are aversive to D. melanogaster males. Mated females have increased levels of male courtship inhibitory pheromones due to their mechanical transfer during copulation (Scott 1986; Ejima et al. 2007). These include the well-characterized cVA and 7-T pheromones (Kurtovic et al. 2007; Lacaille et al. 2007; Billeter et al. 2009; Billeter and Levine 2015; Laturney and Billeter 2016). D. simulans females have much higher concentrations of 7-T on their cuticles compared to D. melanogaster females (Jallon and David 1987), and several studies have shown that 7-T recognition is essential for preventing courtship by D. melanogaster males toward heterospecific females (Scott 1986; Lacaille et al. 2007; Fan et al. 2013).
A larger proportion of miR-957 mutant males courted mated females, although their response to virgin females was similar to that of control males (Figure 6A). miR-957 KO males also had a significantly higher courtship index than control males toward mated females (Figure 6B).
Figure 6.
miR-957 mutant males do not significantly reduce courtship efforts toward mated D. melanogaster females. (A) The proportion of miR-957 KO or control males that court decapitated D. melanogaster mated or virgin females during the last 5 min of a 30-min assay. (B) CI of miR-957 KO or control males toward decapitated D. melanogaster mated or virgin females during the last 10 min of a 30-min assay. CI values were compared using Mann–Whitney U-tests. N = 28–30 for each genotype. Error bars represent mean ± SE (SEM). CI, courtship index; CS, Canton-S; KO, knockout.
As expected, a low percentage of control males courted D. simulans females after a 25-min exposure to the heterospecific females (Figure 7A). Compared to control males, roughly twice as many miR-957 KO males continued to court D. simulans females after a 25-min exposure (Figure 7A). When we calculated the courtship index of males toward D. simulans females, we found that miR-957 KO males had a significantly higher courtship index than control males (Figure 7B).
Figure 7.
miR-957 mutant males do not significantly reduce courtship efforts toward virgin D. simulans females. (A) The proportion of miR-957 KO or control males that court decapitated virgin D. simulans or D. melanogaster females during the last 5 min of a 30-min assay. (B) CI of miR-957 KO males or control males toward decapitated virgin D. simulans or D. melanogaster females during the last 10 min of a 30-min assay. CI values were compared using Mann–Whitney U-tests. N = 28–30 for each genotype. Error bars represent mean ± SE (SEM). CI, courtship index; CS, Canton-S; KO, knockout.
These results indicated that miR-957 mutant males may not perceive male-aversive pheromones correctly, which may account for their heightened courtship toward other males.
miR-957 KO males are less responsive to 7-T and cVA
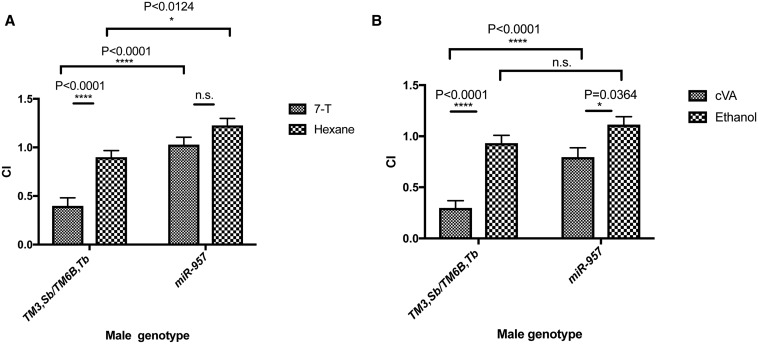
To identify the aversive pheromone(s) that miR-957 KO males are less responsive to, we exposed these males to females perfumed with 7-T or cVA, and measured the courtship index for the last 10 min of a 30-min assay (Figure 8, A and B). When males were exposed to females coated with 7-T, the male genotype as well as the substance on the female had a significant effect on courtship index, and the interaction between the male genotype and substance perfumed was also significant [two-way ANOVA: male genotype × female pheromone: F (1, 117) = 4.027; P = 0.0471; male genotype: F (1, 117) = 40.32; P < 0.0001; and female pheromone: F (1, 117) = 40.32; P < 0.0001]. These results suggested that differences in courtship index toward the different females are not consistent for either male genotype. The perfuming of 7-T on females dramatically reduces the courtship index of control males (one-way ANOVA: F (3, 118) = 22.01; P < 0.0001, Tukey’s post hoc test: control males with hexane-treated females vs. control males with 7-T treated females: P < 0.0001, N1 = 28, N2 = 29), while miR-957 KO males do not show a significant reduction in courtship index toward females perfumed with 7-T when compared to the courtship index of miR-957 KO males toward hexane-treated females (P < 0.2280, N1 = N2 = 32) (Figure 8A). There was also a significant difference between the courtship index of control males and miR-957 KO males toward 7-T-treated females (P < 0.0001). We noted that miR-957 KO males had a higher courtship index toward hexane-treated females as compared to the courtship of control males toward hexane-treated females (P = 0.0124). Regardless of the basal courtship toward hexane-treated females, it is clear that control males show a very strong behavioral response to the addition of 7-T, whereas the response is nearly absent in miR-957 KO males.
Figure 8.
miR-957 mutant males show reduced inhibitory response to 7-T and cVA. (A) CI of miR-957 KO males or control males toward 7-T- or hexane-treated females during the last 10 min of a 30-min assay. N = 28–32 for each genotype. (B) CI of miR-957 KO males or control males toward cVA- or ethanol-treated females during the last 10 min of a 30-min assay. N = 27–31 for each genotype. CI values were compared using one-way ANOVA followed by post hoc Tukey–Kramer tests. Error bars represent mean ± SE (SEM). CI, courtship index; cVA, cis-vaccenyl acetate; KO, knockout; n.s., not significant; 7-T, 7-Tricosene.
For experiments with cVA-treated females, the interaction between the male genotype and substance perfumed was significant [two-way ANOVA: male genotype × female pheromone: F (1, 111) = 3.964; P = 0.0489; male genotype: F (1, 111) = 18.23; P < 0.0001; female pheromone: F (1, 111) = 35.63; P < 0.0001]. Again, the response to the substance perfumed was genotype-dependent. Perfuming of cVA on females significantly lowered the courtship index of control males [one-way ANOVA: F (3, 111) = 20; P < 0.0001; Tukey’s post hoc test: control males with ethanol-treated females vs. control males with cVA treated females: P < 0.0001, N1 = 31, N2 = 32].
In contrast to experiments with females perfumed with 7-T, when females were perfumed with cVA, miR-957 KO males showed a reduction in courtship index toward females as compared to their courtship index toward ethanol-treated females (P = 0.0364, N1 = N2 = 27). Yet, there is a significant difference between the courtship index of control and miR-957 KO males toward cVA-treated females (P < 0.0001), compared to the difference between the courtship index of control and miR-957 KO males toward ethanol-treated females, which was not significant (P = 0.3798). Although these males are responding to cVA, the response is much lower than the response observed from control males.
These experiments do not allow us to determine whether the mutant males have a reduced inhibitory response to 7-T or if they are now attracted to 7-T. To differentiate between these two possibilities, we performed another set of two-choice experiments. miR-957 KO males were given a choice between decapitated 7-T-treated virgin females and decapitated untreated virgin females. miR-957 mutant males showed a clear preference for the untreated virgin females (Figure S4). This finding suggests that the males have a reduced inhibitory response to 7-T rather than an attraction to the pheromone.
Target mRNA prediction for miR-957
For in silico target prediction of potential mRNA targets of miR-957 we used TargetScan Fly (Ruby et al. 2007). The algorithm predicts targets by finding mRNAs that have sequences in 3′-UTRs complementary to the seed region of the miRNA. The predictions are based on a combination of factors including the local sequence environment (Bartel 2009). We found 58 conserved targets. Four out of the top six targets with the highest weightage have very high expression in the adult brain and male-biased expression in the head post mating (FlyBase) (Figure 9A). Since miR-957 KO males respond differently to pheromones, we suspected that miR-957 may be acting in the gustatory pathways. Interestingly, one of the predicted targets was Gustatory receptor 39b (Gr39b). Therefore, we asked if miR-957 is needed in Gr39b-expressing neurons to maintain correct courtship by carrying out knockdown (Figure 9) and overexpression experiments (Figure S3) for miR-957 using the Gr39b-gal4 driver. We did not detect a significant effect on male–male courtship in either case.
Figure 9.
Predicted targets of miR-957. (A) Top six weighted targets from TargetScan Fly predictions. (B) Total courtship and chaining index (tCCI) for grouped males expressing UAS-miR-957.sponge in gustatory receptor neurons. tCCI values for each miR-957.sponge and gal4 combination were compared to the respective scramble control using Student’s t-tests. Error bars represent mean ± SE (SEM).
Another gustatory receptor, GR32a, is a receptor for 7-T (Miyamoto and Amrein 2008). Since miR-957 KO males showed a reduced aversive response to 7-T, we also tested the effect of overexpressing miR-957 or expressing miR-957.sponge using Gr32a-gal4, which is expressed in neurons that detect 7-T. Again, we did not detect an effect of miR-957 overexpression on male–male courtship (Figure S3). When we used Gr32a-gal4 driver to express miR-957.sponge, we detected an increase in male–male courtship but the difference was not significant (P = 0.0557) (Figure 9).
Discussion
miRNA KO screens in Drosophila melanogaster are a powerful genetic tool for understanding behavior
miRNAs have been implicated in subtle regulation of protein expression, for example, by acting as a mechanism to counter the effects of leaky gene expression (Lai et al. 2016). Yet, several recent studies have begun to emphasize the role of miRNAs in the regulation of more complex behavioral phenotypes such as learning (Wang et al. 2012; Li et al. 2013; Busto et al. 2015), rhythmic behavior (You et al. 2018), egg laying (Fricke et al. 2014; Zhou et al. 2014; Nehammer et al. 2015), and locomotor activity (Fulga et al. 2015; Nehammer et al. 2015; Picao-Osorio et al. 2017). The impact of miRNAs on these behaviors results from the cumulative effect of altered regulation of several mRNA targets. Most studies have explored the effects of miRNA null mutations (Li et al. 2013; Chen et al. 2014; Picao-Osorio et al. 2017), and in several cases these effects were observed not just during early stages of development but also in adult organisms (Li et al. 2013; Weng et al. 2013; Chen et al. 2014). These results show that miRNAs may impact levels of several proteins within the cell without being essential for the organism’s viability. Hence, miRNA KO screens provide a powerful genetic tool to observe phenotypes resulting from complete genetic ablation of a post-transcriptional regulatory element while retaining viability of the subjects. We utilized this tool to understand the extent to which miRNAs contribute to a complex behavior in D. melanogaster by assessing effects of single KO alleles on aspects of male courtship and reproductive behavior.
Understanding the factors that dictate social behavior, such as male courtship, involves both analyzing the external stimuli and the body’s response mechanisms. To understand the genetic basis of these responses, studies have previously shown that social interactions can induce rapid changes in the transcriptomic profiles of D. melanogaster males (Ellis and Carney 2009). Moreover, evidence suggests that these gene expression patterns are influenced by the sex of the interacting partner (Ellis and Carney 2011), thereby showing that stimulus–response dynamics are more complex than previously assumed. Drosophila courtship behavior is an innate behavior that is primarily controlled by neurons that express major transcription factors of the fly sex-determination hierarchy. This circuit is responsive to sensory stimuli, and courtship behaviors can be modified based on the attractive or aversive nature of the stimuli received. One of the most potent contributors to this regulation is the complex complement of stimulatory or inhibitory pheromones (Kohl et al. 2015; Schultzhaus et al. 2017). In our current study, we identified miRNAs that impacted one or more of the stereotypical parameters of courtship. In the initial screen, ∼26% of the strains tested had reduced mating success, while ∼42% showed variations in other parameters. More specifically, compared to controls, 19 miRNA mutant strains showed a significantly different courtship latency, three strains had significantly altered mating latency, and 16 had different mating success. Overall, strains with increased mating latency always showed lower mating success, but there was no clear pattern for courtship latency. Increases or decreases in courtship latency were observed across miRNA KO strains, with no clear correlation with mating latency or mating success. miR-957, in particular, was shown to be important for the detection of male-aversive pheromones. The absence of the miRNA led to compromised sensing and/or evaluation of the courtship targets. To our knowledge, there have been no published genome-wide screens to assess the function of individual miRNAs in regulating courtship behavior.
miR-957 mutant males perceive male courtship-inhibiting pheromones differently and therefore display male–malecourtship
miR-957 is a relatively unexplored miRNA that does not have mammalian orthologs, but is conserved in four species of Drosophila and three species of mosquito (miRBase, http://www.mirbase.org) (Ruby et al. 2007; Stark et al. 2007; Kozomara and Griffiths-Jones 2014). In our study, miR-957 mutants displayed high levels of intermale courtship. These males did not elicit more courtship from wild-type males but courted males with significantly higher intensity. This result suggested that the pheromonal profile of miR-957 KO males had not changed but, instead, that they failed to perceive sexual cues from courtship subjects correctly. miR-957 mutant males courted females at levels similar to the courtship displayed by control males, and they chose female subjects when given the choice to choose between males or females. These results indicated that their perception of female attractive pheromones was intact, but miR-957 mutants failed to recognize males as futile targets for courtship. To determine if miR-957 KO males are not receiving/processing male aversive cues correctly, we tested their courtship toward two types of female subjects, D. simulans females and mated D. melanogaster females, which are known to have elevated levels of pheromones that inhibit male courtship. We found that miR-957 mutants did not reduce courtship toward mated D. melanogaster females or D. simulans females as fast as the control males, suggesting that miR-957 males are responding differently to the bouquet of pheromones that are generally sexually aversive for D. melanogaster males. A limitation of our work is that our experiments do not distinguish between developmental and adult physiological function of miR-957.
miR-957 is important for detecting pheromones, including 7-T and cVA, that contribute to chemical mate-guarding, and for maintaining the sex and species-specific courtship barriers
Male–male courtship behavior observed in miR-957 mutant males could be due to altered perception of either a specific male courtship-inhibiting pheromone or due to a change in the processing of information from multiple male sexually aversive pheromones. Potential candidate pheromones include 7-T. This pheromone has been classified by most studies as a gustatory cue associated with male–male aggression and is well described as an aversive compound for male–male courtship (Savarit et al. 1999; Billeter et al. 2009; Wang et al. 2011). 7-T is present in much higher quantities on D. melanogaster males compared to females (Laturney and Billeter 2016). Some studies have also suggested that it is only found on virgin males and is not present on virgin females (Everaerts et al. 2010). Genetic ablation of 7-T producing oenocytes results in an increase in male–male courtship, while application of 7-T onto these males rescues the phenotype (Savarit et al. 1999; Billeter et al. 2009; Wang et al. 2011), suggesting a role for 7-T in providing a sex-specific barrier to courtship. Mutating one of the receptors for 7-T, GR32a, increases male courtship toward other males as well as to mated females (Miyamoto and Amrein 2008). During copulation, Drosophila males deposit aversive pheromones onto females to prevent or delay copulations with other males, a phenomenon known as chemical mate-guarding. The fact that loss of the 7-T receptor causes males to court mated females suggests its additional role in chemical mate-guarding (Miyamoto and Amrein 2008).
Among the well-characterized volatile olfactory ligands is the male inhibitory pheromone cVA, which is produced in the male ejaculatory bulb and is transferred to females during mating (Butterworth 1969; Brieger and Butterworth 1970). cVA is also suspected to be important for reducing male–male courtship (Ejima et al. 2007; Wang and Anderson 2010). cVA appears to be another antiaphrodisiac that is not being detected by miR-957 mutant males since the males continue to court females that have been perfumed with cVA. It is also plausible that the increased male–male courtship shown by miR-957 males is due to an altered response to a combination of pheromones, including 7-T and cVA, and not to one specific pheromone. This possibility is in line with the finding that cVA and 7-T interact to reduce the attractiveness of mated females (Laturney and Billeter 2016).
Our perfuming experiments showed that miR-957 KO males do not reduce their courtship significantly toward females perfumed with 7-T, whereas the control males do so within a 30-min assay. miR-957 KO males also had higher courtship index toward hexane-treated females as compared to the courtship of control males toward hexane-treated females. Besides being defective for detecting 7-T, miR-957 mutant males may also be hypersensitive to an attractive pheromone present on the females. However, this scenario is unlikely because we did not observe a difference between the courtship of miR-957 KO males and control males toward ethanol-treated females. Additionally, the difference between the courtship index of control and miR-957 KO males toward 7-T-treated females is more pronounced compared to the difference between their courtship index toward hexane-treated females. The finding that miR-957 is important for the perception of a gustatory pheromone led us to test effects of miR-957 reduction in neurons that express Gr32a (Figure 9B), a gustatory receptor shown to be important for 7-T detection (Miyamoto and Amrein 2008). We observed an increase in male–male courtship, although it was not significantly higher (P = 0.0557). We did not detect an increase in male–male courtship upon miR-957 overexpression in Gr32a neurons (Figure S3). We also did not observe any effect of reduction (Figure 9B) or overexpression (Figure S3) of miR-957 in Gr39b-expressing neurons. Gr39b was identified as a potential target in our in silico search for miR-957 targets. Failure to detect a strong effect upon reduction of miR-957 in Gr32a or Gr39b neurons suggests that miR-957 may be needed in a broader set of neurons involved in gustatory detection, or possibly in the higher-order processing centers such as the MB.
Interestingly, miR-957 KO males reduce courtship efforts in the presence of cVA, although the response is weaker than the response shown by control males. These findings make sense in light of the emerging understanding that 7-T plays a dominant role in preventing male–male courtship as well as in interspecific mating, and in our study miR-957 KO males were less responsive to 7-T, and were observed to display both male–male courtship as well as very high heterospecific courtship (Figure 7). 7-T is one of the compounds that is transferred onto females during copulation and stays on them much longer than most seminally transferred compounds (Yew et al. 2009; Everaerts et al. 2010). cVA, on the other hand, is now viewed as a regulator of response to other pheromones by increasing or decreasing repulsion that is already there to modulate the intensity of specific social behaviors (Billeter and Levine 2015; Billeter and Wolfner 2018). cVA must be sensed in conjunction with 7-T to influence aggression (Wang et al. 2011) and courtship behavior (Laturney and Billeter 2016).
Differences in the response to 7-T and cVA could arise through a variety of mechanisms. For example, the sensory pathway for the two pheromones differs. cVA is detected primarily by the olfactory system by a specialized trichoid sensillum, while the putative receptor for 7-T is a gustatory receptor, Gr32a (Miyamoto and Amrein 2008; Wang et al. 2011; Laturney and Billeter 2016). For miR-957 to be involved in the detection of both cVA and 7-T, it would need to be expressed in both gustatory and olfactory sensing channels. Expression of miR-957 mainly in the gustatory pathways can explain the stronger impact on 7-T perception and a lesser one on the response to cVA. Apart from the requirement of miR-957 in sensing pathways, its possible expression in higher-order neuronal connections from neurons delivering aversive gustatory inputs could also explain the requirement of the miRNA in the 7-T response.
miR-957 expression is needed in dsx-expressing and MB neurons to prevent misdirected courtship
Our experiments with the miR-957 sponge showed that miR-957 expression is needed in dsx-expressing neurons for the inhibition of male–male behavior. DSX transcription factors are essential for cell sexual identity, as well as sex-specific behaviors and secondary sexual characteristics. In males, dsx is alternatively spliced to produce male-specific DSXM. Expression of DSXM is not only necessary for proper display of male courtship steps, but studies have shown it to be important for the perception of sensory cues needed for appropriate mate recognition (Rezával et al. 2016). Surprisingly, we did not observe any male–male courtship when the sponge was expressed in fru neurons. There is a partial overlap in the neurons that express fru and dsx. In particular, the majority of brain and thoracic ganglion neurons expressing dsx also express fruM. However, there are several neurons that express only dsx and are known to be sufficient to induce male courtship behavior. These include neurons in the PC1 cluster (Rezával et al. 2016). The observations from the sponge experiments thus suggested that the subfraction of dsx-expressing neurons that do not express fru is important for appropriate mate recognition. Another possibility is that miR-957 expression is needed in all dsx neurons and not only in the fraction that overlaps with fru neurons to allow correct mate recognition.
We also detected male–male courtship behavior upon suppressing miR-957 expression in the MB, which is a structure implicated in learning and olfactory processing (Kim et al. 2007), as well as mate discrimination (Ferveur et al. 1995). It has been shown that the feminization of the MB via expression of tra induces bisexual behavior in males. Since MB is also needed for olfactory processing it was speculated that MB is a structure that is important for the analysis of sex-specific pheromones (Ferveur et al. 1995). miR-957 could have a role in the MB in this context. However, we do not know if dsx is also expressed within the MB neurons. It may be that MB neurons communicate with dsx neurons in the pC1, pC2, and pC3 clusters that closely surround the MB calyces (Rideout et al. 2010), or with the higher-order dsx neurons in the ventral nerve cord. Overall the results suggest that male–male courtship in miR-957 mutant males is potentially a result of altered pheromone processing, more likely at the level of integration of sexually aversive cues in the higher-order processing centers such as the MB. This scenario may be independent of miR-957’s role in dsx neurons.
An alternative approach to miRNA reduction to assess miRNA function is tissue-specific overexpression of the miRNA. Since miRNA overexpression results in target downregulation, if we identified a phenotype from miR-957 overexpression we could follow up with a gene-specific RNA interference (RNAi) approach to validate potential targets. miRNA target downregulation can produce the same effect as target overexpression if any change in the target level from a set threshold (whether the change is positive or negative) results in the phenotype. Therefore, we used the drivers that we used to reduce miRNA levels to overexpress miR-957, but we did not observe the male–male courtship phenotype upon overexpression in any of the regions previously tested, including dsx and MB neurons (Figure S3). This result suggests either that only overexpression of miR-957’s direct targets brings about the male–male courtship phenotype or that a downstream shift in protein levels resulting from the absence of miR-957 contributes to the reduction in response to inhibitory male pheromones. The male–male courtship phenotype could be due to a large number of targets being overexpressed simultaneously, disrupting any threshold transcript or protein levels that were potentially being maintained by the miRNA.
miR-263B and miR-317 impact important parameters of male–female courtship
Our screen identified additional miRNAs that affect various aspects of male courtship and mating behaviors. miR-263b is a conserved miRNA (mammalian orthologs are in the miR-183 family) that was recently characterized in detail as part of a study to identify astroglial miRNAs that regulate circadian behavior (You et al. 2018). miR-263b is widely expressed in the CNS and has experimentally validated targets including hid and the in silico predicted target Clock (You et al. 2018). In our study, miR-263b mutants showed increased courtship directed toward wild-type females, and this phenotype was also observed with miR-263b transheterozygous mutants and sponges. The increase in courtship activity of miR-263b suggests that it is involved in the suppression of proteins that enhance courtship activity. However, no experimentally validated targets affecting courtship exist, and fru is not a predicted target for miR-263b in TargetScan, a program that calculates potential mRNA targets of miRNAs (Ruby et al. 2007).
miR-317 mutant males showed reduced mating success, and the phenotype was observed in the KO mutants as well as the KO/Df transheterozygotes. Males expressing the miR-317-sponge also showed the same phenotype. However, neuronal expression of miR-317 expression with elav-gal4 was not sufficient to rescue the mutant phenotype. Previously, miR-317 has been implicated in locomotor behavior, the startle response, morphogenesis (Yamamoto et al. 2008), and intermale aggressive behavior (Edwards et al. 2009). Additionally, miR-317 has been associated with effects on female postmating responses (Fricke et al. 2014).
Emerging roles for miRNAs in the regulation of complex behaviors
Several recent studies have documented the role of miRNAs in the regulation of transcript levels for maintenance of complex phenotypes (Varghese et al. 2010; Vodala et al. 2012; Weng et al. 2013; Busto et al. 2015; Picao-Osorio et al. 2017; You et al. 2018). A complex and robust behavior that has not received as much attention in the context of miRNA-mediated regulation is courtship behavior. By conducting a genetic screen with miRNA KO mutants, we showed that individual miRNAs can influence one or more aspects of male courtship behavior. miRNAs may be contributing to courtship behavior through involvement in the maintenance of site- and sex-specific protein expression by repressing the translation of transcripts produced from leaky expression of genes at sites where the expression of these genes needs to be limited. For example, Weng et al. (2013) demonstrated that a modest increase in the level of the female-specific tra splice form, traF, in D. melanogaster miR-124 mutant males caused reduced splicing of dsx, which led to decreased production of the male-specific isoform of dsx (dsxM).
miRNAs are recognized as being important for adaptive responses to environmental changes (He et al. 2016). Therefore, apart from restricting leaky expression, miRNAs are ideal candidates for factors that bring about fast changes in the proteomic landscape in response to rapidly changing cues from courtship subjects. Alternatively, the effects of miRNAs on courtship behaviors could be a result of their effects on the development of the male courtship circuitry, since miRNAs have been shown to have roles in neurodevelopment and dendritic remodeling (Nesler et al. 2013).
Further assessment of the effects of miRNA candidates that come from screens such as ours may involve using other experimental techniques to validate targets. Recently, You et al. (2018) attempted to validate targets of miR-263b in the context of circadian regulation by using RNAi against selective, computationally predicted targets of miR-263b (You et al. 2018). Their aim was to determine if reduction of predicted target mRNAs of miR-263b produced the same phenotype (arrhythmic behavior) seen in the mutants for the behavior. Their rationale for using RNAi against targets stemmed from the results of miR-263b overexpression experiments that showed that flies with miR-263b overexpression showed arrhythmic behavior similar to the KO mutants. However, their experiments did not identify targets for miR-263b (You et al. 2018). Other approaches that may be employed for target validation include overexpressing predicted targets and testing for phenotypes similar to those observed in the miRNA mutants.
The availability of a broad range of genetic tools for miRNA modulation in a model organism such as D. melanogaster (Iftikhar et al. 2017) has started a wave of research that explores the role of miRNAs in intricate behaviors needed for the organism’s survival. Such studies will help to add missing pieces to several neural regulatory pathways and elucidate the relevance of an additional layer of RNA-mediated gene expression control. In our study, we have shown how the role of an individual miRNA in pheromone processing shapes courtship in Drosophila, demonstrating that miRNAs, whether through widespread genomic changes or via fine-tuning of specific targets, have the potential to modify highly robust and complex phenotypes.
Acknowledgments
We thank Tai Tong and Hannah M. Lauritzen (Texas A&M University) for help evaluating behavioral data; Pedram Kajeh (Texas A&M University) for preparing fly media; Paul Hardin and Mariana Mateos (Texas A&M University) for providing us with the D52H-gal4 and D. simulans strains, respectively; and Chloe J. Bennett and Janna N. Schultzhaus for their insights into the research. This work was supported by a National Science Foundation grant (IOS 1121517) to G.E.C. All work was carried out at Texas A&M University. The authors declare that no competing interests exist.
Author contributions: H.I. and G.E.C. designed the research and wrote the manuscript; H.I. performed and analyzed data from the screen, experiments for validation of selected candidates and miR-957 mutant characterization, and perfuming experiments; N.L.J. contributed to miR-957 mutant behavioral characterization experiments the and analysis of screen experiments; M.L.M. contributed to performance of the screen and the analysis of data from the screen.
Footnotes
Supplemental material available at Figshare: https://doi.org/10.25386/genetics.7522490.
Communicating editor: M. Wolfner
Literature Cited
- Abraham W. C., 2008. Metaplasticity: tuning synapses and networks for plasticity. Nat. Rev. Neurosci. 9: 387 10.1038/nrn2356 [DOI] [PubMed] [Google Scholar]
- Bartel D. P., 2009. MicroRNAs: target recognition and regulatory functions. Cell 136: 215–233. 10.1016/j.cell.2009.01.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bejarano F., Bortolamiol-Becet D., Dai Q., Sun K., Saj A., et al. , 2012. A genome-wide transgenic resource for conditional expression of Drosophila microRNAs. Development 139: 2821–2831. 10.1242/dev.079939 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bender W., 2008. MicroRNAs in the Drosophila bithorax complex. Genes Dev. 22: 14–19. 10.1101/gad.1614208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Billeter J.-C., Levine J., 2015. The role of cVA and the Odorant binding protein Lush in social and sexual behavior in Drosophila melanogaster. Front. Ecol. Evol. 3: 75. [Google Scholar]
- Billeter J. C., Wolfner M. F., 2018. Chemical cues that guide female reproduction in Drosophila melanogaster. J. Chem. Ecol. 44: 750–769. 10.1007/s10886-018-0947-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Billeter J. C., Atallah J., Krupp J. J., Millar J. G., Levine J. D., 2009. Specialized cells tag sexual and species identity in Drosophila melanogaster. Nature 461: 987–991. 10.1038/nature08495 [DOI] [PubMed] [Google Scholar]
- Brieger G., Butterworth F. M., 1970. Drosophila melanogaster: identity of male lipid in reproductive system. Science 167: 1262 10.1126/science.167.3922.1262 [DOI] [PubMed] [Google Scholar]
- Busto G. U., Guven-Ozkan T., Fulga T. A., Van Vactor D., Davis R. L., 2015. microRNAs that promote or inhibit memory formation in Drosophila melanogaster. Genetics 200: 569–580. 10.1534/genetics.114.169623 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butterworth F. M., 1969. Lipids of Drosophila: a newly detected lipid in the male. Science 163: 1356–1357. 10.1126/science.163.3873.1356 [DOI] [PubMed] [Google Scholar]
- Carthew R. W., Sontheimer E. J., 2009. Origins and mechanisms of miRNAs and siRNAs. Cell 136: 642–655. 10.1016/j.cell.2009.01.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X., Rosbash M., 2017. MicroRNA-92a is a circadian modulator of neuronal excitability in Drosophila. Nat. Commun. 8: 14707 10.1038/ncomms14707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y. W., Song S., Weng R., Verma P., Kugler J. M., et al. , 2014. Systematic study of Drosophila microRNA functions using a collection of targeted knockout mutations. Dev. Cell 31: 784–800. 10.1016/j.devcel.2014.11.029 [DOI] [PubMed] [Google Scholar]
- Dalton J. E., Lebo M. S., Sanders L. E., Sun F., Arbeitman M. N., 2009. Ecdysone receptor acts in fruitless-expressing neurons to mediate Drosophila courtship behaviors. Curr. Biol. 19: 1447–1452. 10.1016/j.cub.2009.06.063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dauwalder B., Tsujimoto S., Moss J., Mattox W., 2002. The Drosophila takeout gene is regulated by the somatic sex-determination pathway and affects male courtship behavior. Genes Dev. 16: 2879–2892. 10.1101/gad.1010302 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Bono M., Bargmann C. I., 1998. Natural variation in a neuropeptide Y receptor homolog modifies social behavior and food response in C. elegans. Cell 94: 679–689. 10.1016/S0092-8674(00)81609-8 [DOI] [PubMed] [Google Scholar]
- Edwards A. C., Zwarts L., Yamamoto A., Callaerts P., Mackay T. F., 2009. Mutations in many genes affect aggressive behavior in Drosophila melanogaster. BMC Biol. 7: 29 10.1186/1741-7007-7-29 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ejima A., Smith B. P., Lucas C., Levine J. D., Griffith L. C., 2005. Sequential learning of pheromonal cues modulates memory consolidation in trainer-specific associative courtship conditioning. Curr. Biol. 15: 194–206. 10.1016/j.cub.2005.01.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ejima A., Smith B. P., Lucas C., van der Goes van Naters W., Miller C. J., et al. , 2007. Generalization of courtship learning in Drosophila is mediated by cis-vaccenyl acetate. Curr. Biol. 17: 599–605. 10.1016/j.cub.2007.01.053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellis L. L., Carney G. E., 2009. Drosophila melanogaster males respond differently at the behavioural and genome-wide levels to Drosophila melanogaster and Drosophila simulans females. J. Evol. Biol. 22: 2183–2191. 10.1111/j.1420-9101.2009.01834.x [DOI] [PubMed] [Google Scholar]
- Ellis L. L., Carney G. E., 2011. Socially-responsive gene expression in male Drosophila melanogaster is influenced by the sex of the interacting partner. Genetics 187: 157–169. 10.1534/genetics.110.122754 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Everaerts C., Farine J. P., Cobb M., Ferveur J. F., 2010. Drosophila cuticular hydrocarbons revisited: mating status alters cuticular profiles. PLoS One 5: e9607 10.1371/journal.pone.0009607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fagegaltier D., Konig A., Gordon A., Lai E. C., Gingeras T. R., et al. , 2014. A genome-wide survey of sexually dimorphic expression of Drosophila miRNAs identifies the steroid hormone-induced miRNA let-7 as a regulator of sexual identity. Genetics 198: 647–668. 10.1534/genetics.114.169268 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan P., Manoli D. S., Ahmed O. M., Chen Y., Agarwal N., et al. , 2013. Genetic and neural mechanisms that inhibit Drosophila from mating with other species. Cell 154: 89–102. 10.1016/j.cell.2013.06.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferveur J. F., Stortkuhl K. F., Stocker R. F., Greenspan R. J., 1995. Genetic feminization of brain structures and changed sexual orientation in male Drosophila. Science 267: 902–905. 10.1126/science.7846534 [DOI] [PubMed] [Google Scholar]
- Fricke C., Green D., Smith D., Dalmay T., Chapman T., 2014. MicroRNAs influence reproductive responses by females to male sex peptide in Drosophila melanogaster. Genetics 198: 1603–1619. 10.1534/genetics.114.167320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fulga T. A., McNeill E. M., Binari R., Yelick J., Blanche A., et al. , 2015. A transgenic resource for conditional competitive inhibition of conserved Drosophila microRNAs. Nat. Commun. 6: 7279 10.1038/ncomms8279 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenberg J. K., Xia J., Zhou X., Thatcher S. R., Gu X., et al. , 2012. Behavioral plasticity in honey bees is associated with differences in brain microRNA transcriptome. Genes Brain Behav. 11: 660–670. 10.1111/j.1601-183X.2012.00782.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- He J., Chen Q., Wei Y., Jiang F., Yang M., et al. , 2016. MicroRNA-276 promotes egg-hatching synchrony by up-regulating brm in locusts. Proc. Natl. Acad. Sci. USA 113: 584–589. 10.1073/pnas.1521098113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iftikhar H., Carney G. E., 2016. Evidence and potential in vivo functions for biofluid miRNAs: from expression profiling to functional testing: potential roles of extracellular miRNAs as indicators of physiological change and as agents of intercellular information exchange. Bioessays 38: 367–378. 10.1002/bies.201500130 [DOI] [PubMed] [Google Scholar]
- Iftikhar H., Schultzhaus J. N., Bennett C. J., Carney G. E., 2017. The in vivo genetic toolkit for studying expression and functions of Drosophila melanogaster microRNAs. RNA Biol. 14: 179–187. 10.1080/15476286.2016.1272748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Sato K., Koganezawa M., Ote M., Matsumoto K., et al. , 2012. Fruitless recruits two antagonistic chromatin factors to establish single-neuron sexual dimorphism. Cell 149: 1327–1338. 10.1016/j.cell.2012.04.025 [DOI] [PubMed] [Google Scholar]
- Jallon J. M., David J. R., 1987. Variations in cuticular hydrocarbons among the eight species of the Drosophila melanogaster subgroup. Evolution 41: 294–302. [DOI] [PubMed] [Google Scholar]
- Kim Y. C., Lee H. G., Han K. A., 2007. D1 dopamine receptor dDA1 is required in the mushroom body neurons for aversive and appetitive learning in Drosophila. J. Neurosci. 27: 7640–7647. 10.1523/JNEUROSCI.1167-07.2007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura K., Ote M., Tazawa T., Yamamoto D., 2005. Fruitless specifies sexually dimorphic neural circuitry in the Drosophila brain. Nature 438: 229–233. 10.1038/nature04229 [DOI] [PubMed] [Google Scholar]
- Kimura K., Hachiya T., Koganezawa M., Tazawa T., Yamamoto D., 2008. Fruitless and doublesex coordinate to generate male-specific neurons that can initiate courtship. Neuron 59: 759–769. 10.1016/j.neuron.2008.06.007 [DOI] [PubMed] [Google Scholar]
- Kohatsu S., Koganezawa M., Yamamoto D., 2011. Female contact activates male-specific interneurons that trigger stereotypic courtship behavior in Drosophila. Neuron 69: 498–508. 10.1016/j.neuron.2010.12.017 [DOI] [PubMed] [Google Scholar]
- Kohl J., Ostrovsky A. D., Frechter S., Jefferis G. S., 2013. A bidirectional circuit switch reroutes pheromone signals in male and female brains. Cell 155: 1610–1623. 10.1016/j.cell.2013.11.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohl J., Huoviala P., Jefferis G. S., 2015. Pheromone processing in Drosophila. Curr. Opin. Neurobiol. 34: 149–157. 10.1016/j.conb.2015.06.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozomara A., Griffiths-Jones S., 2014. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42: D68–D73. 10.1093/nar/gkt1181 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krstic D., Boll W., Noll M., 2013. Influence of the White locus on the courtship behavior of Drosophila males. PLoS One 8: e77904 10.1371/journal.pone.0077904 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurtovic A., Widmer A., Dickson B. J., 2007. A single class of olfactory neurons mediates behavioural responses to a Drosophila sex pheromone. Nature 446: 542–546. 10.1038/nature05672 [DOI] [PubMed] [Google Scholar]
- Lacaille F., Hiroi M., Twele R., Inoshita T., Umemoto D., et al. , 2007. An inhibitory sex pheromone tastes bitter for Drosophila males. PLoS One 2: e661 10.1371/journal.pone.0000661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai X., Wolkenhauer O., Vera J., 2016. Understanding microRNA-mediated gene regulatory networks through mathematical modelling. Nucleic Acids Res. 44: 6019–6035. 10.1093/nar/gkw550 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laturney M., Billeter J. C., 2016. Drosophila melanogaster females restore their attractiveness after mating by removing male anti-aphrodisiac pheromones. Nat. Commun. 7: 12322 10.1038/ncomms12322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leung A. K., Sharp P. A., 2010. MicroRNA functions in stress responses. Mol. Cell 40: 205–215. 10.1016/j.molcel.2010.09.027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li W., Cressy M., Qin H., Fulga T., Van Vactor D., et al. , 2013. MicroRNA-276a functions in ellipsoid body and mushroom body neurons for naive and conditioned olfactory avoidance in Drosophila. J. Neurosci. 33: 5821–5833. 10.1523/JNEUROSCI.4004-12.2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim M. M., Wang Z., Olazabal D. E., Ren X., Terwilliger E. F., et al. , 2004. Enhanced partner preference in a promiscuous species by manipulating the expression of a single gene. Nature 429: 754–757. 10.1038/nature02539 [DOI] [PubMed] [Google Scholar]
- McRobert S. P., Tompkins L., Barr N. B., Bradner J., Lucas D., et al. , 2003. Mutations in raised Drosophila melanogaster affect experience-dependent aspects of sexual behavior in both sexes. Behav. Genet. 33: 347–356. 10.1023/A:1023406810804 [DOI] [PubMed] [Google Scholar]
- Miyamoto T., Amrein H., 2008. Suppression of male courtship by a Drosophila pheromone receptor. Nat. Neurosci. 11: 874–876. 10.1038/nn.2161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morel L., Regan M., Higashimori H., Ng S. K., Esau C., et al. , 2013. Neuronal exosomal miRNA-dependent translational regulation of astroglial glutamate transporter GLT1. J. Biol. Chem. 288: 7105–7116. 10.1074/jbc.M112.410944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nehammer C., Podolska A., Mackowiak S. D., Kagias K., Pocock R., 2015. Specific microRNAs regulate heat stress responses in Caenorhabditis elegans. Sci. Rep. 5: 8866 10.1038/srep08866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nesler K. R., Sand R. I., Symmes B. A., Pradhan S. J., Boin N. G., et al. , 2013. The miRNA pathway controls rapid changes in activity-dependent synaptic structure at the Drosophila melanogaster neuromuscular junction. PLoS One 8: e68385 10.1371/journal.pone.0068385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan Y., Robinett C. C., Baker B. S., 2011. Turning males on: activation of male courtship behavior in Drosophila melanogaster. PLoS One 6: e21144 10.1371/journal.pone.0021144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavlou H. J., Goodwin S. F., 2013. Courtship behavior in Drosophila melanogaster: towards a ‘courtship connectome’. Curr. Opin. Neurobiol. 23: 76–83. 10.1016/j.conb.2012.09.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Picao-Osorio J., Lago-Baldaia I., Patraquim P., Alonso C. R., 2017. Pervasive behavioral effects of microRNA regulation in Drosophila. Genetics 206: 1535–1548. 10.1534/genetics.116.195776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rezával C., Pattnaik S., Pavlou H. J., Nojima T., Bruggemeier B., et al. , 2016. Activation of latent courtship circuitry in the brain of Drosophila females induces male-like behaviors. Curr. Biol. 26: 2508–2515. 10.1016/j.cub.2016.07.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rideout E. J., Dornan A. J., Neville M. C., Eadie S., Goodwin S. F., 2010. Control of sexual differentiation and behavior by the doublesex gene in Drosophila melanogaster. Nat. Neurosci. 13: 458–466. 10.1038/nn.2515 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruby J. G., Stark A., Johnston W. K., Kellis M., Bartel D. P., et al. , 2007. Evolution, biogenesis, expression, and target predictions of a substantially expanded set of Drosophila microRNAs. Genome Res. 17: 1850–1864. 10.1101/gr.6597907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Savarit F., Sureau G., Cobb M., Ferveur J. F., 1999. Genetic elimination of known pheromones reveals the fundamental chemical bases of mating and isolation in Drosophila. Proc. Natl. Acad. Sci. USA 96: 9015–9020. 10.1073/pnas.96.16.9015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultzhaus J. N., Saleem S., Iftikhar H., Carney G. E., 2017. The role of the Drosophila lateral horn in olfactory information processing and behavioral response. J. Insect Physiol. 98: 29–37. 10.1016/j.jinsphys.2016.11.007 [DOI] [PubMed] [Google Scholar]
- Scott D., 1986. Sexual mimicry regulates the attractiveness of mated Drosophila melanogaster females. Proc. Natl. Acad. Sci. USA 83: 8429–8433. 10.1073/pnas.83.21.8429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi Z., Luo G., Fu L., Fang Z., Wang X., et al. , 2013. miR-9 and miR-140–5p target FoxP2 and are regulated as a function of the social context of singing behavior in zebra finches. J. Neurosci. 33: 16510–16521. 10.1523/JNEUROSCI.0838-13.2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sokolowski M. B., 2001. Drosophila: genetics meets behaviour. Nat. Rev. Genet. 2: 879–890. 10.1038/35098592 [DOI] [PubMed] [Google Scholar]
- Stark A., Kheradpour P., Parts L., Brennecke J., Hodges E., et al. , 2007. Systematic discovery and characterization of fly microRNAs using 12 Drosophila genomes. Genome Res. 17: 1865–1879. 10.1101/gr.6593807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock J. B., Ninfa A. J., Stock A. M., 1989. Protein phosphorylation and regulation of adaptive responses in bacteria. Microbiol. Rev. 53: 450–490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varghese J., Lim S. F., Cohen S. M., 2010. Drosophila miR-14 regulates insulin production and metabolism through its target, sugarbabe. Genes Dev. 24: 2748–2753. 10.1101/gad.1995910 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villella A., Gailey D. A., Berwald B., Ohshima S., Barnes P. T., et al. , 1997. Extended reproductive roles of the fruitless gene in Drosophila melanogaster revealed by behavioral analysis of new fru mutants. Genetics 147: 1107–1130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vodala S., Pescatore S., Rodriguez J., Buescher M., Chen Y. W., et al. , 2012. The oscillating miRNA 959–964 cluster impacts Drosophila feeding time and other circadian outputs. Cell Metab. 16: 601–612. 10.1016/j.cmet.2012.10.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L., Anderson D. J., 2010. Identification of an aggression-promoting pheromone and its receptor neurons in Drosophila. Nature 463: 227–231. 10.1038/nature08678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L., Han X., Mehren J., Hiroi M., Billeter J. C., et al. , 2011. Hierarchical chemosensory regulation of male-male social interactions in Drosophila. Nat. Neurosci. 14: 757–762. 10.1038/nn.2800 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W., Kwon E. J., Tsai L. H., 2012. MicroRNAs in learning, memory, and neurological diseases. Learn. Mem. 19: 359–368. 10.1101/lm.026492.112 [DOI] [PubMed] [Google Scholar]
- Weng R., Chin J. S., Yew J. Y., Bushati N., Cohen S. M., 2013. miR-124 controls male reproductive success in Drosophila. Elife 2: e00640 10.7554/eLife.00640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson R., Burnet B., Eastwood L., Connolly K., 1976. Behavioural pleiotropy of the yellow gene in Drosophila melanogaster. Genet. Res. 28: 75–88. 10.1017/S0016672300016748 [DOI] [PubMed] [Google Scholar]
- Yamamoto A., Zwarts L., Callaerts P., Norga K., Mackay T. F., et al. , 2008. Neurogenetic networks for startle-induced locomotion in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 105: 12393–12398. 10.1073/pnas.0804889105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto D., Koganezawa M., 2013. Genes and circuits of courtship behaviour in Drosophila males. Nat. Rev. Neurosci. 14: 681–692. 10.1038/nrn3567 [DOI] [PubMed] [Google Scholar]
- Yew J. Y., Dreisewerd K., Luftmann H., Muthing J., Pohlentz G., et al. , 2009. A new male sex pheromone and novel cuticular cues for chemical communication in Drosophila. Curr. Biol. 19: 1245–1254. 10.1016/j.cub.2009.06.037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi W., Zhang Y., Tian Y., Guo J., Li Y., et al. , 2013. A subset of cholinergic mushroom body neurons requires Go signaling to regulate sleep in Drosophila. Sleep (Basel) 36: 1809–1821. 10.5665/sleep.3206 [DOI] [PMC free article] [PubMed] [Google Scholar]
- You S., Fulga T. A., Van Vactor D., Jackson F. R., 2018. Regulation of circadian behavior by astroglial microRNAs in Drosophila. Genetics 208: 1195–1207. 10.1534/genetics.117.300342 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou S., Mackay T., Anholt R. R., 2014. Transcriptional and epigenetic responses to mating and aging in Drosophila melanogaster. BMC Genomics 15: 927 10.1186/1471-2164-15-927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ziegler A. B., Berthelot-Grosjean M., Grosjean Y., 2013. The smell of love in Drosophila. Front. Physiol. 4: 72 10.3389/fphys.2013.00072 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zovkic I. B., Guzman-Karlsson M. C., Sweatt J. D., 2013. Epigenetic regulation of memory formation and maintenance. Learn. Mem. 20: 61–74. 10.1101/lm.026575.112 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Table S1 contains phenotypic data and statistical analyses of miRNA KO mutants. Figure S1 contains activity measurements of miRNA KO mutant screen candidates. Figure S2 shows fru-gal4 expression in the brain. Figure S3 shows effects of overexpression of miR-957. Figure S4 is a two-choice test between perfumed and wild-type females. Supplemental material available at Figshare: https://doi.org/10.25386/genetics.7522490.