Abstract
Antibodies protect organisms from a huge variety of foreign antigens. Antibody diversity originates from both genetic and structural levels. Antigen recognition relies on complementarity between antigen-antibody interfaces. Recent methodological advances in structural biology and the accompanying rapid increase of the number of crystal structures of proteins have enabled atomic-level manipulation of protein structures to effect alterations in function. In this study, we explored the designability of electrostatic complementarity at an antigen-antibody interface on the basis of a crystal structure of the complex. We designed several variants with altered charged residues at the interface and characterized the designed variants by surface plasmon resonance, circular dichroism, differential scanning calorimetry, and molecular dynamics simulations. Both successes and failures of the structure-based design are discussed. The variants that compensate electrostatic interactions can restore the interface complementarity, enabling the cognate antigen-antibody binding. Retrospectively, we also show that these mutational effects could be predicted by the simulations. Our study demonstrates the importance of charged residues on the physical properties of this antigen-antibody interaction and suggests that computational approaches can facilitate design of antibodies that recognize a weakly immunogenic antigen.
Introduction
Although antigens far outnumber antibody genes, sufficient antibody diversity results from genetic and structural variation1. During B cell development, V(D)J recombination, somatic mutations, and somatic insertions and deletions result in an antibody repertoire that can recognize the enormous number of antigens encountered over a lifetime with excellent specificity2. Antibody specificity is determined by the antigen binding site called the complementarity determining region (CDR) and antigen recognition is governed by several types of interactions: hydrogen bonds, electrostatic interactions, van der Waals forces, and hydrophobic interactions3–5. As the name suggests, complementarity is a key to antigen recognition. Previous works surveyed complementarity of protein-protein interactions in terms of shape, hydrophobicity, and electrostatics6–11. Among them, electrostatic interactions can be controlled by manipulating charged amino acid residues on the interaction surface. For example, generation of charge repulsion at a protein-protein interface would weaken the interaction whereas generation of charge attractions would strengthen the interaction.
Electrostatic force has been harnessed in antibody engineering12–16. Starting from crystal structures of antigen-antibody complexes, Lippow et al. exploited Poisson–Boltzmann electrostatics to improve binding affinity, showing a 100-fold improvement in affinity over the wild-type antibody12. Liu and coworkers demonstrated that substitution of multiple surface residues with charged amino acids, a technique they called supercharging, increased thermal resistance in GFP13. Miklos et al. subsequently designed thermostable antibodies based on a supercharging strategy14; they demonstrated that the designed antibodies had better thermal stability and binding affinity than the parent antibodies even though the altered positions were not in the CDRs but in the framework regions (FRs). The improved affinity was due to a faster on-rate as well as a slower off-rate as measured by surface plasmon resonance (SPR). Kiyoshi et al. also started with a crystal structure of an antibody–antigen complex and improved the affinity of the interaction using a computational saturation mutagenesis approach15. These researchers identified several affinity-enhancing mutations that were located at the periphery of the interface; interestingly, all were mutated to charged residues. Similarly, Fukunaga and Tsumoto improved the affinity of an anti-human cardiac troponin I antibody by substitution of multiple amino acids in FRs with charged residues16. These studies indicate that manipulating electrostatic interactions can improve physicochemical properties of antigen-antibody interactions. To achieve this approach, it is important to understand the physicochemical principles behind the electrostatic complementarity of antigen-antibody interactions.
Molecular dynamics (MD) simulations can reveal the effects of amino acid mutations on a protein structure17–19. Wong et al. investigated how mutations rigidified CDRs utilizing MD simulations and found that simulations could be useful in designing mutations that resulted in hydrogen bonding and tight packing of side chains17. Corrada et al. examined correlations between structures of mutated antibodies and affinities for the antigens on the basis of computational and experimental studies, respectively, and proposed a possible mechanism of rigidification18. Clark et al. attempted direct calculation of relative antigen-antibody binding affinities using a free energy perturbation protocol and replica exchange solute tempering method and was able to predict the effect of alanine mutations on the relative binding affinities19. These analyses indicate that MD simulations can reveal dynamic influences that cannot be detected by analysis of a static crystal structure and can be used to predict the effects of protein manipulations on function.
In this study, to clarify the importance of electrostatic interactions and to facilitate design of antigen-antibody interactions, we investigated the effect of surface charge manipulations at an antigen-antibody interface. As a model system, we started with a crystal structure of a complex between neutralizing antibody A6 and the interferon gamma receptor (IFNγR)20 available through the Protein Data Bank (PDB)21. SPR measurements, circular dichroism (CD) spectra, differential scanning calorimetry (DSC), and MD simulations were performed to quantify kinetics and thermodynamics of the wild-type and the designed antigen-antibody complexes. We also attempted to predict designability of the charged residues solely using a computational approach and discuss a possible strategy to efficiently elicit antibodies against weakly immunogenic antigens by surface charge manipulations.
Results
Crystal structure of an antigen-antibody complex guides surface charge manipulations at the interface
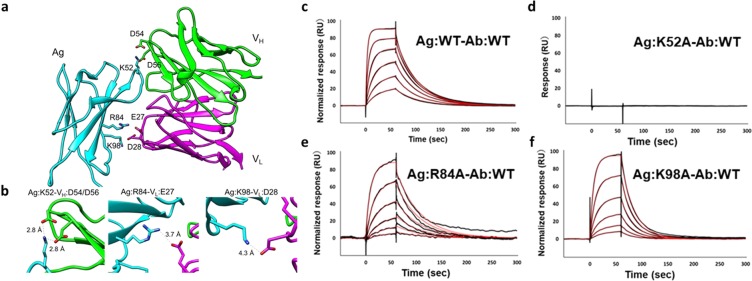
We chose as our model antigen-antibody system the complex of the extracellular domain D1 of IFNγR and the antibody A6 (Fig. 1a, PDB ID: 1JRH)20. Analysis of the crystal structure indicates that three electrostatic interactions contribute to stabilization of the interaction between antigen (Ag) and heavy and light chain variable domains of the antibody (VH, and VL, respectively) (Fig. 1b). First, K52 of the antigen interacts with the charged β-carboxyl groups of D54 and D56 of the VH domain at distances of 2.8 Å. Second, the γ-carboxyl group of E27 of the VL domain is 3.7 Å from the guanidine group of R84 of the antigen. The main chain of R84 forms part of a β-sheet, and head of exposed side chain contributes to the electrostatic interaction. Third, the β-carboxyl group of D28 of the VL is 4.3 Å from the antigen K98 ε-amino group. Both Ag:K98 and VL:D28 are exposed to solvent and in the complex interact with each other.
Figure 1.
Electrostatic interactions in the IFNγR-A6 complex and the SPR response profiles of the wild-type and the alanine mutant complexes. (a) Crystal structure of the antigen (cyan) and antibody (H chain: green, L chain: magenta) complex (from PDB ID: 1JRH, only variable regions are shown). (b) Electrostatic interactions at the interface of the wild-type complex. Each distance of electrostatic interaction is displayed as dash lines and was measured based on the crystal structure. Antibody variable regions are numbered based on the Kabat numbering scheme. SPR response profiles of (c) Ag:WT-Ab:WT, (d) Ag:K52A-Ab:WT (e) Ag:R84A-Ab:WT, and (f) Ag:K98A-Ab:WT. The concentrations of the antibody were 1000, 500, 250, 125, 62.5, and 31.25 nM. Raw data are in black; red lines are fitted curves. All fitted data were normalized and assume a 1:1 interaction. Binding kinetics were measured at 298 K.
We performed an SPR assay to experimentally quantify the interaction of the wild-type antibody (Ab:WT) with the wild-type antigen (Ag:WT); the affinity was high with a KD of 87 nM (Fig. 1c and Table 1). We first investigated the importance of each charged residue on the antigen side of the interface. To abolish electrostatic interactions, we prepared three alanine mutants of the antigen, Ag:K52A, Ag:R84A, and Ag:K98A. The SPR measurements showed that Ag:K52A did not bind detectably to Ab:WT (Fig. 1d), suggesting that K52 of the antigen is a hot spot for the interaction with the antibody. Ag:R84A and Ag:K98A bound less tightly to the Ab:WT than did Ag:WT (KDs 280 nM and 440 nM, respectively compared to 87 nM for Ag:WT; Fig. 1e–f, and Table 1). Compared to the wild-type antigen-antibody interaction, the off rates (koff) almost unchanged, whereas the on rates (kon) were slower by 3 to 4 fold for the mutant antigens, suggesting that antigen side chains R84 and K98 play important roles in the association of the antigen with the antibody, likely due to the long-range electrostatic attraction to E27 and D28 of VL, respectively. These results indicate that mutation of a single charge-charge pair is tolerated, whereas electrostatic interactions involving more than three charged residues disrupt binding completely, perhaps because the position of each residue is cooperatively controlled by multipoint binding.
Table 1.
Kinetic parameters of SPR analyses.
| Antigen-Antibody | kon (105 M−1 · s−1) | koff (10−2 · s−1) | KD (10−8 M) |
|---|---|---|---|
| Ag:WT-Ab:WT | 2.7 | 2.3 | 8.7 |
| Ag:R84A-Ab:WT | 0.74 | 2.1 | 28 |
| Ag:K98A-Ab:WT | 0.90 | 3.9 | 44 |
| Ag:K52A-Ab:WT | N/A | ||
| Ag:R84E-Ab:WT | N/A | ||
| Ag:K98D-Ab:WT | N/A | ||
| Ag:K52D-Ab:WT | N/A | ||
| Ag:R84E-V L :E27R | 2.4 | 1.7 | 7.3 |
| Ag:K98D-V L :D28K | 2.1 | 2.0 | 9.5 |
| Ag:K52D-V H :D54K/D56K | N/A |
Charge complementarity recovery restores antigen-antibody binding
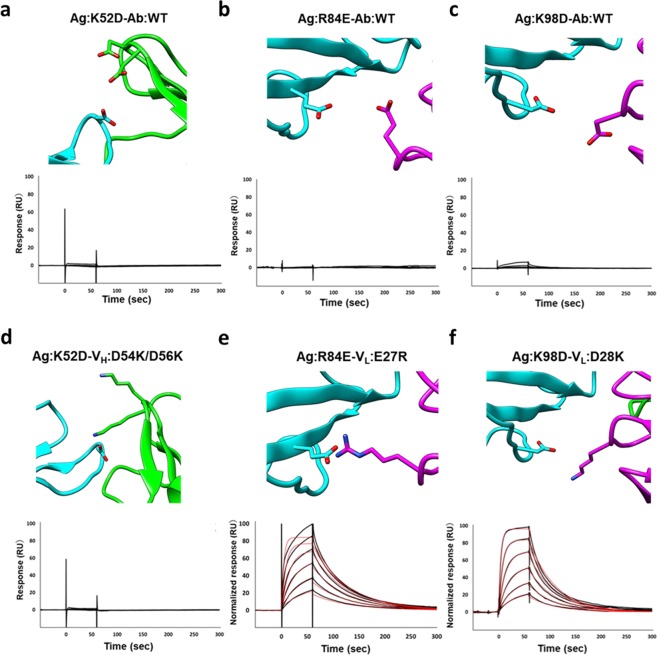
To explore the designability of electrostatic complementarity at the interface, we further designed three “charge-inverted” antigens (Ag:K52D, Ag:R84E and Ag:K98D; Fig. 2a–c). As expected, the SPR analysis of Ag:K52D and Ag:R84E showed that these charge-inverted mutants did not bind to the Ab:WT (Fig. 2a,b). The other variant, Ag:K98D, bound weakly to Ab:WT (Fig. 2c). These results suggested that a single electrostatic repulsion can disrupt the antigen-antibody interaction.
Figure 2.
Structures and SPR response profiles of the charge-inverted and charge-exchanged mutants. (a) Ag:K52D-Ab:WT, (b) Ag:R84E-Ab:WT, (c) Ag:K98D-Ab:WT, (d) Ag:K52D-VH:D54K/D56K (e) Ag:R84E-VL:E27R, and (f) Ag:K98D-VL:D28K. Structures of mutants were generated using CHARMM-GUI. The concentrations of the antibody were 1000, 500, 250, 125, 62.5, and 31.25 nM. Raw data are in black; red lines are fitted curves. All fitted data were normalized and assume a 1:1 interaction. Binding kinetics were measured at 298 K.
To determine whether compensatory changes in the antibody would restore binding, we prepared three “charge-exchanged” antigen-antibody complexes (Ag:K52D-VH:D54K/D56K, Ag:R84E-VL:E27R, and Ag:K98D-VL:D28K; Fig. 2d–f). In the antibody, the charged residues at the interface are complementary. The SPR analysis of the charge-exchanged complexes showed that Ag:R84E and Ag:K98D interacted with VL:E27R and VL:D28K, respectively, with binding affinities similar to the wild-type antigen-antibody interaction (KDs 73 nM and 95 nM, respectively, versus 87 nM for Ag:WT-Ab:WT; Fig. 2e,f, and Table 1). Moreover, the on and off rates of the Ag:R84E-VL:E27R and Ag:K98D-VL:D28K complexes were similar to those of the wild-type antigen-antibody complex.
To verify that loss of binding resulting from charge inverting mutations was caused by electrostatic repulsion, we analyzed the effects of the mutations on the structure and thermostability of antigen and antibody. The CD spectra of the engineered antigens showed that the mutation of R84E changed the secondary structure of the unbound-state of the antigen (Fig. S1a). Given that R84 is partially buried inside the antigen and is part of a β-sheet structure, it was not surprising that the mutation would alter the local structure. The SPR result indicated that the corresponding mutation in the antibody, E27R in the VL domain, restored the binding affinity to near wild-type levels. This suggests that the lack of antibody binding by Ag:R84E is mainly caused by electrostatic repulsion.
The individual CD spectra of the compensatory pair Ag:K98D and VL:D28K were similar to those of the wild-type antigen and antibody (Fig. S1a,b), suggesting that these mutants did not change the conformations, and disruption/restoration of interactions between the two mutants were derived from solely change of electrostatic effect. It is worth noting that the CD spectrum of Ag:K52D differed from that of Ag:WT; this was not expected as K52 is an exposed residue (Fig. S2). In addition, the corresponding antibody VH:D54K/D56K also differed in secondary structure from the wild-type antibody. This suggests that lack of binding of Ag:K52D to Ab:WT might be due not only to electrostatic repulsion but also structural distortion (Figs 2d and S1).
To further quantify the mutational effects on the structural stability, we carried out DSC assays. The DSC profiles showed that Ag:K52D had higher thermal stability than Ag:WT antigen; the Tm,app was 4 °C higher (Fig. 3a). However, the transition peak of the mutant was broad, and a small shoulder was observed, implying that heterogeneous conformations are present. Of antibody mutants, VH:D54K/D56K had a significantly smaller ΔH and a slightly lower Tm,app than Ab:WT (Fig. 3b). Thus, the DSC results also imply that conformations of Ag:K52D and VH:D54K/D56K differ from wild-type conformations and thereby disrupting the binding.
Figure 3.
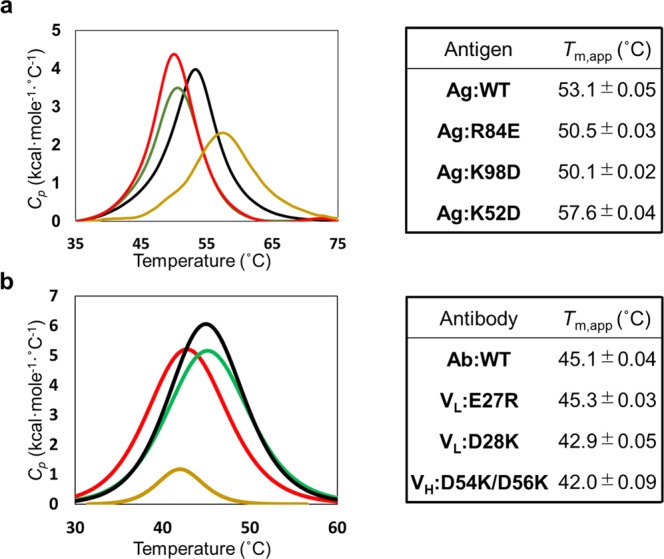
DSC analysis of thermodynamic stabilities of antigen-antibody complexes. (a) Melting curves and calculated Tm,app values of the antigens. Ag:WT (black), Ag:R84E (green), Ag:K98D (red), and Ag:K52D (orange). (b) Melting curves and calculated Tm,app values of the antibodies. Ab:WT (black), VL:E27R (green), VL:D28K (red), and VH:D54K/D56K (orange). The standard deviation of each Tm,app value was calculated from the fitting curve.
Detailed molecular characterization of mutants reveals the importance of electrostatic interactions
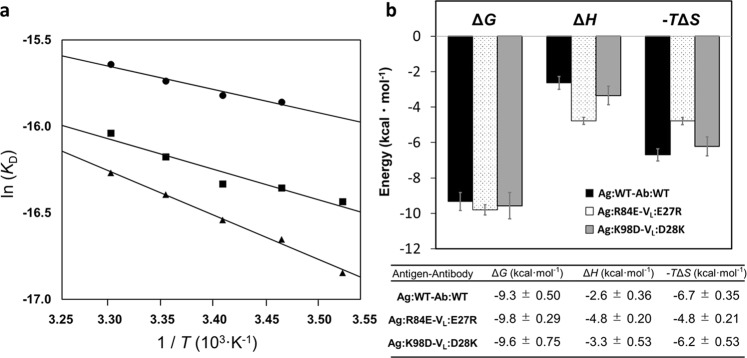
Next, we evaluated thermodynamics of the interactions for each mutant by SPR as a function of temperatures. The thermodynamic parameters were determined from van’t Hoff plots (Fig. 4a). Although the charge-exchanged mutations were kinetically similar to wild type (Fig. 2e,f, and Table 1), Ag:R84E-VL:E27R and Ag:K98D-VL:D28K had different thermodynamic profiles (Fig. 4b). Compared to the wild-type antigen-antibody interaction, the Ag:R84E-VL:E27R interaction was driven more enthalpically, whereas the balance of enthalpic and entropic contributions to the Ag:K98D-VL:D28K interaction were almost same as those of the wild-type complex.
Figure 4.
Comparison of thermodynamic profile of antigen-antibody interactions. (a) Plots of ln (KD) vs 1/T (K−1) for wild-type antigen-antibody (●), Ag:R84E-VL:E27R (▲), and Ag:K98D-VL:D28K (■). (b) Bar graphs of the equilibrium thermodynamic parameters obtained from van’t Hoff analyses. Error bars correspond to the standard deviation. The standard deviations of ΔH and −TΔS were calculated from the fitting data, respectively, and that of ΔG was calculated from ΔG = ΔH − TΔS with error propagation. Values are reported at 298 K.
To understand the molecular mechanism underlying the change in thermodynamic parameters due to charge-exchanged mutations we examined the intramolecular environment surrounding each mutated charged residue. The crystal structure of the antigen-antibody complex suggests that the guanidine group of the Ag:R84 side chain has cation-π interactions with surrounding aromatic residues such as Ag:W56 and Ag:Y96. Ag:R84 also forms an electrostatic interaction with Ag:E45 as the distance between the guanidine group of Ag:R84 and γ-carboxyl group of Ag:E45 is 2.9 Å (Fig. S2). Therefore, in addition to the salt bridge with VL:E27 of the antibody, Ag:R84 stabilizes the structure of the antigen itself.
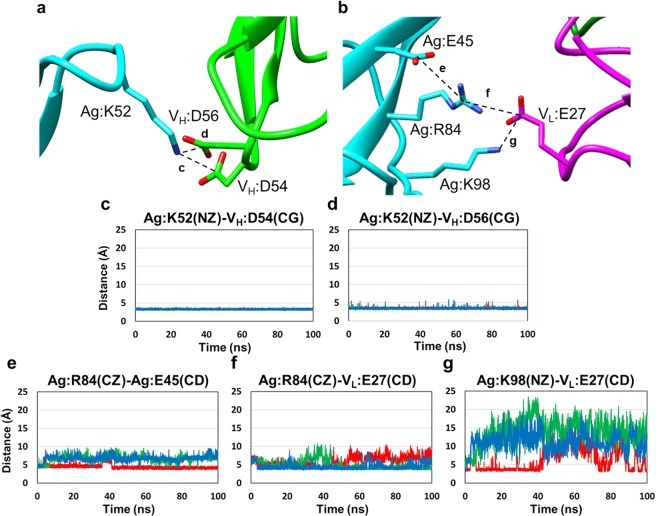
To further understand the molecular details of the reorganization of the electrostatic complementarity, we performed multiple 100 ns MD simulations of the wild-type complex as well as Ag:R84E-VL:E27R and Ag:K98D-VL:D28K complexes, which had kinetics similar to that of the wild-type complex. We first computed the root mean square deviation (RMSD) of the Cα atoms of the complexes, except for the first and last five residues of each chain, with respect to the reference crystal structure 1JRH (Fig. S3). The RMSD values were quite stable after 20 ns. Therefore, in the analyses below, we did not consider the first 20 ns of the trajectories. First, we confirmed the stable interaction at the predicted hotspot by measurement of the distances between Ag:K52 (NZ atom) and VH:D54 and VH:D56 (CG atoms). In the wild-type complex, the distance between Ag:K52 and VH:D54 was within 3.6 Å, which is the distance in the crystal structure, in ~100% of the snapshots in the last 80 ns of each simulation and the distance between Ag:K52 (NZ atom) and VH:D56 (CG atom) was within 3.5 Å in 55.3 ± 2.0% of the snapshots (Fig. 5a,c–d). In the crystal structure, Ag:R84 interacts with intramolecular aromatic residues (Ag:W56 and Ag:Y96) and a charged residue (Ag:E45) (Fig. S2). In two out of three simulations, Ag:R84 tended to interact with VL:E27 as seen in the crystal structure; this interaction was observed in 77.3% and 83.3% of the snapshots, respectively. In contrast, in the other simulation Ag:R84 tended to form a salt bridge with Ag:E45; these residues were within 4.3 Å, which is the distance between CZ and CD atoms of Arg and Glu, respectively, in the crystal structure, in 48% of the snapshots, and the interaction between Ag:R84 and VL:E27 was observed in only 26.7% (Fig. 5b,e,f). Moreover, in this simulation, VL:E27 also interacted with Ag:K98, which was spatially adjacent to the Ag:R84. The distance between VL:E27 (CD) and Ag:K98 (NZ) was 8.0 Å in the crystal structure, whereas the distance was less than or equal to 4.0 Å in 37.7% of the snapshots (Figs 1a and 5b,g). Thus, in the simulations of the wild-type complex, charge-charge interactions were well established during the simulations.
Figure 5.
Distances between charged residues that contribute to electrostatic interactions during the MD simulations. (a,b) Electrostatic interactions at the interface of the wild-type complex. Distances between each atom were displayed as dash lines. Distances between (c) Ag:K52 and VH:D54, (d) Ag:K52 and VH:D56, (e) Ag:R84 and Ag:E45, (f) Ag:R84 and VL:E27, and (g) Ag:K98 and VL:E27. Each distance was measured based on NZ, CG, CZ, and CD atoms of Lys, Asp, Arg, and Glu, respectively. Each 100 ns run was performed three times, indicated in red, green, and blue lines.
In the simulations of the Ag:K98D-VL:D28K complex, as expected, Ag:K98D interacted with VL:D28K within 4.0 Å in 62 ± 2% of the snapshots. In addition, Ag:K98D also interacted with Ag:R84 due to the charge attraction, and these residues were within 4.0 Å in 17 ± 5% of the snapshots. These salt bridges were mutually exclusive with the interaction network comprised of Ag:R84, Ag:E45, Ag:W56, and Ag:Y96, which was observed in the crystal structure. The Ag:K98D interaction with Ag:R84 placed Ag:R84 closer to VL:E27 (Fig. S4a,b). In the simulations of the Ag:R84E-VL:E27R complex, Ag:R84E interacted with VL:E27R within 4.0 Å in 88 ± 3% of the snapshots. Ag:K98 attracted Ag:R84E, making a cavity near VH:E45, VH:Y56, and VH:W96 (Fig. S4c,d). VL:E27R was located in this cavity and formed interactions electrostatic or cation-π interactions with VH:E45, VH:Y56, and VH:W96 (Fig. S4e). Therefore, as with the simulations of the wild-type complex, the charge-charge interactions were observed in the simulations of the two charge-exchanged complexes (Ag:R84E-VL:E27R and Ag:K98D-VL:D28K).
These results suggest that although the electrostatic interactions observed in the crystal structure were important in the association of mutant antigen-antibody complexes, other intermolecular and intramolecular interactions that were not obvious from the static crystal structure also play roles in the association: Surrounding residues can compensate for loss of electrostatic interactions as binding is stabilized by the surrounding residues.
Discussion
Electrostatic interactions are one of the important forces in antigen-antibody recognitions. Previous studies demonstrated that binding affinities of antibodies could be enhanced based on electrostatic interactions12,15. Another study suggested that electrostatic interactions at antigen-antibody interfaces contributed to binding specificity and affinity due to loss of conformational flexibility through geometric constraint22. In this study, we carefully characterized the electrostatic interactions of neutralizing antibody A6 with its antigen IFNγR as a model system for structure-based design of antigen-antibody complexes. Of three charged residues changed pair-wise, we succeeded in the modulation of electrostatic interactions for two combinations Ag:R84E-VL:E27R and Ag:K98D-VL:D28K. In contrast, the interactions between Ag:K52 and VH:D54/D56 were not restored by the charge-exchanged mutations (Fig. 2d–f). Our biophysical analyses using CD and DSC indicate that Ag:K52D and VH:D54K/D56K caused structural disruptions that could interfere with complex formation (Figs S1 and 3). All the three residues (Ag:K52, VH:D54/D56) are exposed to solvent (Fig. 1b) and appeared not to form intramolecular bonds in the crystal structure. However, the Ag:K52D and VH:D54K/D56K mutations might have affected the conformation of the antigen and the antibody, respectively. Our results demonstrate the importance of investigating structure and/or thermal stability coupled with interaction analysis when electrostatic mutation-based analyses are conducted.
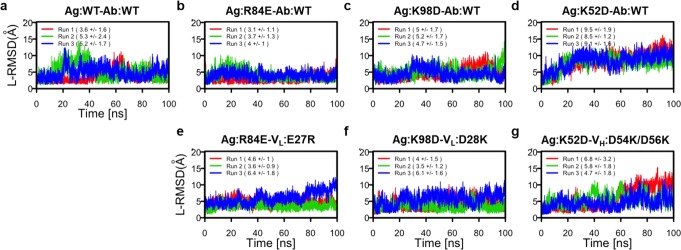
Our MD simulations were able to illustrate how the charge-exchanged residues of Ag:R84E-VL:E27R and Ag:K98D-VL:D28K complexes modulate the electrostatic complementarity. It would be difficult to observe charge association and compensation of electrostatic interactions by surrounding residues solely from the static crystal structure and/or in vitro measurements. These successes provoked us to perform additional MD simulations to evaluate the predictability of effects of mutations at the antigen-antibody interface. Therefore, in addition to the aforementioned simulation studies for two charge-exchanged variants (Ag:R84E-VL:E27R and Ag:K98D-VL:D28K), we also conducted MD simulations of three charge-inverted variants Ag:K52D-Ab:WT, Ag:R84E-Ab:WT, and Ag:K98D-Ab:WT as well as the other charge-exchanged variant Ag:K52D-VH:D54K/D56K. We analyzed the relative orientation between the antigen and antibody by computing RMSDs of the Cα atoms of the antibody after superposing the Cα atoms of the antigen in the trajectories; these values are referred to as ligand-RMSD (L-RMSD)23. The L-RMSD of the wild-type complex was stable throughout the trajectory, suggesting that the simulation resulted in the stable conformation of the antigen-antibody complex (Fig. 6a). Our SPR assays suggested that Ag:R84E-Ab:WT and Ag:K98D-Ab:WT complexes were significantly less stable than the Ag:WT-Ab:WT complex (Figs 1c and 2b,c); however, the L-RMSD plots for these two complexes were similar to that for the wild-type complex (Fig. 6b,c). For the Ag:K52D-Ab:WT complex, L-RMSD values gradually increased, and the complex appeared to dissociate (Fig. 6d), suggesting that the simulation did appropriately reflect the weakened interaction caused by charge inversion (Fig. 2a). As expected, the L-RMSD values of two charge-exchanged variants Ag:R84E-VL:E27R and Ag:K98D-VL:D28K were similar to that of the wild-type complex (Fig. 6e,f) in agreement with our SPR data. Moreover, in the MD simulations, Ag:K52D-VH:D54K/D56K did not form a stable complex, as reflected by high L-RMSD values during the trajectories (Fig. 6g), also concordant with our SPR analysis. From these observations, we suggested that L-RMSD from MD trajectories would be an appropriate metric that could be used to assess mutational effects on protein-protein interfaces.
Figure 6.
RMSDs of the antigen-antibody interactions between simulations and initial structure. L-RMSD values of (a) wild-type antigen-antibody interactions, (b–d) charge-inverted antigen-antibody interactions, and (e–g) charge-exchanged antigen-antibody interactions. Each 100 ns run was performed three times, indicated in red, green, and blue lines. Averages and standard deviations of the L-RMSDs are indicated in parentheses.
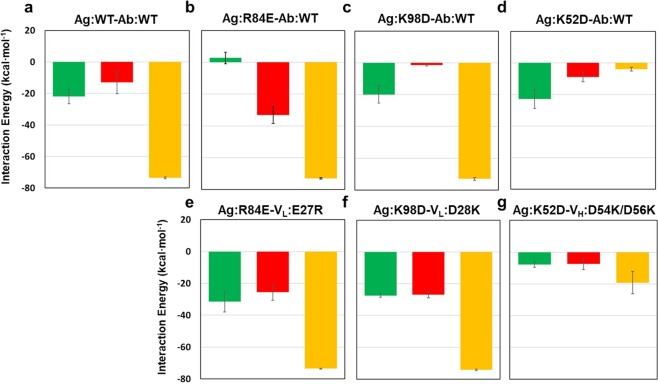
Furthermore, we also performed interaction energy calculations (van der Waals plus coulomb energy) to the mutated residues in the antigen (K52, R84, and K98), enabling to complement the prediction based on L-RMSD. In the wild-type complex and the two charge-exchanged variant Ag:R84E-VL:E27R and Ag:K98D-VL:D28K, which were stable as shown by SPR, Ag:K52 made the largest energy contributions, suggesting that Ag:K52 was a hot spot in the antigen-antibody interaction, in consistent with our SPR and L-RMSD results (Fig. 7). Importantly, the calculation showed significant loss of interaction energy due to each of the three charge-inverted mutations compared to the wild-type complex (Fig. 7a–d). Moreover, whereas the interaction energy was restored by charge-exchange mutation in Ag:R84E-VL:E27R and Ag:K98D-VL:D28K, the interaction energy was not restored in case of Ag:K52D-VH:D54K/D56K (Fig. 7e–g). Although the calculations of interaction energies isolated only the enthalpic contribution without considering entropic effects, it showed reasonable correlation with the SPR data. Thus, MD simulations followed by energy calculations could provide important guidance for antigen-antibody manipulation, enabling computer-aided rational antibody design24.
Figure 7.
Interaction energy of the antigen-antibody interactions in the simulations. Interaction energies of antigen residues with antibody in (a) the wild-type antigen-antibody complex, (b–d) the charge inverted antigen-antibody complexes, and (e–g) charge-exchanged antigen-antibody complexes. Energies of the residues at the position of 84, 98, and 52 are shown in green, red, and orange, respectively. Standard errors were indicated by black lines (n = 3).
In rational antibody engineering for therapeutic and biotechnological purposes25,26, antibody elicitation is the important first step. There are several methods for antibody generation but the most widely used method is animal immunization, where an antigen is injected into an animal. Certain antigens of interest are weakly immunogenic likely due to high homology to protein from immunized animal. Several display technologies have been developed to enable generation of antibodies, but acquisition of antibodies with high affinity, specificity, and stability is challenging.
As it has been suggested that only a single mutation can make proteins more immunogenic27, we suggest that charge conversion is a possible strategy for generation of antibodies against weakly immunogenic antigens. First, one would modulate the surface charge of the antigen to potentiate immunogenicity. After antibodies are obtained against the charge-modulated antigen, the charged residues in the antibody that interact with modified charged residue in the antigen would be identified and converted to match that of the antibody. Our study suggests that the resultant antibody should interact specifically with original antigen (Fig. S5). In addition, our result showing that charge-inverted antibody did not bind to antigen implies that antibodies generated by immunization with charge-modified antigens will not recognize homologous proteins from host animal.
Finally, although many previous studies in protein engineering have tried to improve binding affinities by introducing electrostatic pairs, binding affinity must be modulated appropriately to retain a desired function28. In this study, our designed alanine mutations resulted in the slower kon but did not abolish the binding (Fig. 1e–f). The alanine mutation to a charged residue that is involved in a single negative-positive charge pairing might be a promising approach to decrease affinity by three to five fold. Thus, design of electrostatic complementarity, guided by both experiments and molecular simulations is an effective approach for manipulation of antigen-antibody interaction.
In this study, we presented the structure-based design of electrostatic complementarity of an antigen-antibody complex, and the following biophysical analysis showed the success and failure of these designed variants in terms of the recovery of binding capability. We also suggested the availability of MD-based prediction in the design of an antigen-antibody complex. Our results supplied insights that each electrostatic interaction has different role in an antigen-antibody binding process and should be considered in view of structural and dynamic aspects for rational manipulation of electrostatic complementarity.
Methods
Expression and purification of antibodies
The single-chain variable fragment (scFv) was constructed from the DNA encoding the Fab of A6; the DNA was chemically synthesized by GeneArt. A (GGGGS)3 linker was inserted between the H chain and the L chain, and the sequence was optimized for expression in E. coli. The A6 scFv construct was expressed in vector pRA2 to result in a hexa-histidine tag at the C terminus. The DNA sequence was confirmed by the dideoxy chain-termination sequencing method.
Cells of E. coli strain BL21 (DE3) (Novagen) were transformed with the A6 scFv expression vector and grown at 28 °C in LB medium. Protein expression was induced by addition of 0.5 mM isopropyl β-D-1-thiogalactopyranoside when the optical density at 600 nm reached a value of 0.6. Cells were allowed to grow for an additional 16 h at 20 °C. The cells were harvested by centrifugation at 8,000 × g for 10 min, and the pellet thus obtained was resuspended in a solution containing 0.2 M NaCl, 1 mM EDTA, and 50 mM Tris-HCl, pH 8.0 (buffer A). Cells were subsequently lysed by sonication with an ultrasonic cell-disruptor instrument (TOMY); samples were sonicated twice for 20 min each period (Output 7, Duty 50).
The pellet containing the insoluble intracellular components was obtained by centrifugation at 8,000 × g for 30 min. The supernatant was discarded, and the insoluble fraction was resuspended in 1% Triton-X100 (Wako) in buffer A. After centrifugation at 8,000 × g for 15 min, the insoluble fraction was resuspended in water, and centrifuged at 8,000 × g for 15 min; this removed the detergent. The insoluble fraction was resuspended in acetone and centrifuged at 10,000 × g for 30 min to remove lipids. The insoluble fraction was resuspended in a water to remove remaining acetone, and the inclusion body was obtained by centrifugation at 8,000 × g for 15 min. The pellet was solubilized with 6 M guanidine-HCl, 0.4 M NaCl, 20 mM Tris-HCl, pH 8.0 (buffer B) and 5 mM imidazole and incubated overnight at 4 °C. The supernatant collected after centrifugation at 40,000 × g for 30 min was applied to an Ni-NTA agarose (QIAGEN) column equilibrated with buffer B. The column was washed sequentially with 5 and 10 mM imidazole in buffer B. The A6 scFv was eluted with 300 mM imidazole in buffer B.
The purified A6 scFv was refolded using a stepwise dialysis method. The wild-type A6 scFv was diluted to 7.5 µM, and mutants were diluted to 1 µM with buffer B, followed by stepwise dialysis to remove the denaturant. In order to increase the refolding efficiency, 0.4 M Arg-HCl was added to the dialysis solution to minimize protein aggregation when the concentration of guanidine-HCl decreased from 1 M to 0.5 M as previously described29. The refolded A6 scFv was concentrated and purified over an Ni-NTA agarose column, and subjected to size exclusion chromatography on a HiLoad 26/600 Superdex 75-pg column (GE Healthcare) equilibrated with 0.4 M NaCl, 1 mM EDTA, and 20 mM Tris-HCl, pH 8.0. The monomer peak fraction was collected.
Expression and purification of antigen
The DNA encoding the extracellular domain D1 of IFNγR was chemically synthesized by GeneArt with sequence optimized for expression in E. coli. The IFNγR construct was expressed in vector pET28b to result in a hexa-histidine tag at the C terminus. The DNA sequence was confirmed by sequencing using the dideoxy chain-termination method. Cells of E. coli strain BL21 (DE3) (Novagen) were transformed with the IFNγR expression vector and were grown at 37 °C in LB. Protein expression was induced by addition of 0.5 mM isopropyl β-D-1-galactopyranoside when the optical density at 600 nm reached a value of 0.6. Cells were allowed to grow for an additional 16 h at 20 °C. The cells were harvested by centrifugation at 8,000 × g for 10 min, and the pellet thus obtained was resuspended in a solution containing 0.5 M NaCl, 5 mM imidazole, and 20 mM Tris-HCl, pH 8.0 (buffer C). Cells were subsequently lysed two 20-min periods of sonication method with an ultrasonic cell-disruptor instrument (TOMY; Output 7, Duty 50).
The supernatant containing the soluble intracellular components was obtained by ultracentrifugation at 40,000 × g for 30 min. The soluble fraction was applied to an Ni-NTA agarose (QIAGEN) column equilibrated with buffer C. The column was washed sequentially with 5 and 10 mM imidazole in buffer C. IFNγR was eluted with 300 mM imidazole in buffer C. The purified IFNγR was subjected to size exclusion chromatography over a HiLoad 26/600 Superdex 75-pg column (GE Healthcare) equilibrated with 0.2 M NaCl, 1 mM EDTA, and 50 mM Tris-HCl, pH 8.0. The monomer peak fraction was collected.
Kinetic and thermodynamic SPR assays
The interaction between A6 and IFNγR was analyzed by SPR on a Biacore T200 instrument (GE Healthcare). A research-grade CM5 Biacore sensor chip (GE Healthcare) was activated by a short treatment with N-hydroxysuccinimide/N-ethyl-N’-(3-dimethylaminopropyl) carbodiimide hydrochloride, followed by immobilization of antigen IFNγR at a surface density of approximately 200 RU. The activated groups on the surface of the sensor were subsequently blocked by injecting 1 M ethanolamine as previously described30. The kinetic data of the binding A6 scFv to the IFNγR were obtained by injecting increasing concentrations of A6 scFv over the sensor chip at a flow rate of 30 µl/min. The measurements were carried out in PBS containing 0.005% (v/v) Tween-20. Contact time and dissociation time were 1 min and 5 min, respectively. Data analysis was performed with the BIAevaluation software (GE Healthcare). Association (kon) and dissociation (koff) rate constants were calculated by a global fitting analysis assuming a Langmuir binding model and a stoichiometry of (1:1). The dissociation constant (KD) was determined from the ratio of the rate constants as previously described31:
Enthalpy (ΔH) and entropy (ΔS) were calculated from the slope and intercept, respectively, of the plot of temperature versus dissociation constant using the van’t Hoff approximation32:
where R in the gas constant and T is the absolute temperature.
Circular dichroism spectra
CD spectra were recorded on a model J-820 CD spectrometer (JASCO). Far-UV CD measurements were performed with 0.5 mg/mL of IFNγR in PBS using a 1-mm cell and a bandwidth of 1 nm. Spectra were recorded five times for each sample.
Differential scanning calorimetry
DSC measurements of samples prepared in PBS were carried out on a MicroCal VP-Capillary DSC (Malvern Instruments) at a heating rate of 60 °C per h. To evaluate thermal stability, the temperature where heat capacity was maximum was determined as apparent Tm (Tm,app) using the software MicroCal Origin 7 (Malvern).
Accessible surface area and surface charge calculations
Accessible surface area of the antigen was calculated with the Protein Interfaces, Surfaces and Assemblies (PISA) server33. Surface charges were calculated using the protein contact potential function in PyMOL34.
Molecular dynamics simulations
MD simulations of wild-type and mutant complexes were performed using GROMACS 2016.335 with the CHARMM36m force field and the CMAP correction36,37. Using the CHARMM-GUI38, the Fv regions of the structure of the IFNγR-A6 complex (PDB ID: 1JRH) was solvated with TIP3P water in a rectangular box such that the minimum distance to the edge of the box was 15 Å under periodic boundary conditions. Na and Cl ions were added to neutralize the protein charge, then further ions were added to mimic a salt solution concentration of 0.14 M. Each system was energy-minimized for 5000 steps and equilibrated with NVT ensemble at 298 K for 1 ns. Further production run was performed for 100 ns with NPT ensemble. A cutoff distance of 12 Å for Coulomb and van der Waals interactions was used. Long-range electrostatics were evaluated through the Particle Mesh Ewald method39. LINCS algorithm40 was employed to constrain bonds involving hydrogen atoms. The time step was set to 2 fs throughout the simulations. A simulation was repeated 3 times for each system, and the snapshots were saved every 10 ps. PyMOL and UCSF Chimera41 were employed to analyze and visualize the MD trajectories and to render the molecular graphics. Interaction energy was defined as the sum of van der Waals and Coulomb interactions, which were calculated through GROMACS based on the last 80 ns of each trajectory.
Supplementary information
Acknowledgements
The super-computing resources used in this study were provided by the Human Genome Center at the Institute of Medical Science, The University of Tokyo, Japan. This work was funded in part by Japan Society for the Promotion of Science (grant no. 16H02420 to K.T., 16H06693 to M.N., and 17K18113 to D.K.) and by Japan Agency for Medical Research and Development (grant no. JP18am0101094j, JP18dm0107064h, JP18mk0101081h, JP18fm0208030h, JP18fk0108073h, JP18ak0101100h to K.T., JP18ak0101100h to M.N. and JP18fm0208022h to D.K.).
Author Contributions
This study has been conceived by S.S., H.S. and K.T. K.Y., D.K., M.K., M.N., S.N. and K.T. designed the study. K.Y., D.K. and M.K. performed the experiments. K.Y., D.K., M.N. and S.N. analyzed the data and wrote the manuscript. All the authors approved the manuscript.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Competing Interests
The authors declare no competing interests.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1038/s41598-019-40461-5.
References
- 1.de los Rios M, Criscitiello MF, Smider VV. Structural and genetic diversity in antibody repertoires from diverse species. Curr. Opin. Struct. Biol. 2015;33:27–41. doi: 10.1016/j.sbi.2015.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Janeway, C. A., Travers, P., Walport, M. & Shlomchik, M. Immunobiology: The Immune System in Health and Disease. Immuno Biol. 5th edition, New York: Garland Science (2001).
- 3.Reverberi R, Reverberi L. Factors affecting the antigen-antibody reaction. Blood Transfus. 2007;5:227–240. doi: 10.2450/2007.0047-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Akiba H, Tsumoto K. Thermodynamics of antibody-antigen interaction revealed by mutation analysis of antibody variable regions. J. Biochem. 2015;158:1–13. doi: 10.1093/jb/mvv049. [DOI] [PubMed] [Google Scholar]
- 5.Sela-Culang I, Kunik V, Ofran Y. The structural basis of antibody-antigen recognition. Front. Immunol. 2013;4:1–13. doi: 10.3389/fimmu.2013.00302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li Y, Zhang X, Cao D. The role of shape complementarity in the protein-protein interactions. Sci. Rep. 2013;3:3271. doi: 10.1038/srep03271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tsuchiya Y, Kinoshita K, Nakamura H. Analyses of homo-oligomer interfaces of proteins from the complementarity of molecular surface, electrostatic potential and hydrophobicity. Protein Eng. Des. Sel. 2006;19:421–429. doi: 10.1093/protein/gzl026. [DOI] [PubMed] [Google Scholar]
- 8.McCoy AJ, Chandana Epa V, Colman PM. Electrostatic complementarity at protein/protein interfaces. J. Mol. Biol. 1997;268:570–584. doi: 10.1006/jmbi.1997.0987. [DOI] [PubMed] [Google Scholar]
- 9.Berchanski A, Shapira B, Eisenstein M. Hydrophobic complementarity in protein-protein docking. Proteins Struct. Funct. Bioinforma. 2004;56:130–142. doi: 10.1002/prot.20145. [DOI] [PubMed] [Google Scholar]
- 10.Lawrence MC, Colman PM. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 1993;234:946–950. doi: 10.1006/jmbi.1993.1648. [DOI] [PubMed] [Google Scholar]
- 11.Kuroda D, Gray JJ. Shape complementarity and hydrogen bond preferences in protein-protein interfaces: Implications for antibody modeling and protein-protein docking. Bioinformatics. 2016;32:2451–2456. doi: 10.1093/bioinformatics/btw197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lippow SM, Wittrup KD, Tidor B. Computational design of antibody-affinity improvement beyond in vivo maturation. Nat. Biotechnol. 2007;25:1171–1176. doi: 10.1038/nbt1336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lawrence MS, Phillips KJ, Liu DR. Supercharging proteins can impart unusual resilience. J. Am. Chem. Soc. 2007;129:10110–2. doi: 10.1021/ja071641y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Miklos AE, et al. Structure-based design of supercharged, highly thermoresistant antibodies. Chem. Biol. 2012;19:449–455. doi: 10.1016/j.chembiol.2012.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kiyoshi M, et al. Affinity improvement of a therapeutic antibody by structure-based computational design: Generation of electrostatic interactions in the transition state stabilizes the antibody-antigen complex. Plos One. 2014;9:e87099. doi: 10.1371/journal.pone.0087099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fukunaga A, Tsumoto K. Improving the affinity of an antibody for its antigen via long-range electrostatic interactions. Protein Eng. Des. Sel. 2013;26:773–780. doi: 10.1093/protein/gzt053. [DOI] [PubMed] [Google Scholar]
- 17.Wong SE, Sellers BD, Jacobson MP. Effects of somatic mutations on CDR loop flexibility during affinity maturation. Proteins Struct. Funct. Bioinforma. 2011;79:821–829. doi: 10.1002/prot.22920. [DOI] [PubMed] [Google Scholar]
- 18.Corrada D, Colombo G. Energetic and dynamic aspects of the affinity maturation process: Characterizing improved variants from the bevacizumab antibody with molecular simulations. J. Chem. Inf. Model. 2013;53:2937–2950. doi: 10.1021/ci400416e. [DOI] [PubMed] [Google Scholar]
- 19.Clark AJ, et al. Free Energy Perturbation Calculation of Relative Binding Free Energy between Broadly Neutralizing Antibodies and the gp120 Glycoprotein of HIV-1. J. Mol. Biol. 2017;429:930–947. doi: 10.1016/j.jmb.2016.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sogabe S, et al. Neutralizing epitopes on the extracellular interferon gamma receptor (IFNgammaR) alpha-chain characterized by homolog scanning mutagenesis and X-ray crystal structure of the A6 fab-IFNgammaR1-108 complex. J. Mol. Biol. 1997;273:882–897. doi: 10.1006/jmbi.1997.1336. [DOI] [PubMed] [Google Scholar]
- 21.Berman H, Henrick K, Nakamura H. Announcing the worldwide Protein Data Bank. Nat. Struct. Biol. 2003;10:980. doi: 10.1038/nsb1203-980. [DOI] [PubMed] [Google Scholar]
- 22.Sinha N, Mohan S, Lipschultz CA, Smith-Gill SJ. Differences in electrostatic properties at antibodyantigen binding sites: Implications for specificity and cross-reactivity. Biophys. J. 2002;83:2946–2968. doi: 10.1016/S0006-3495(02)75302-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Janin J, et al. CAPRI: A critical assessment of PRedicted interactions. Proteins Struct. Funct. Genet. 2003;52:2–9. doi: 10.1002/prot.10381. [DOI] [PubMed] [Google Scholar]
- 24.Kuroda D, Shirai H, Jacobson MP, Nakamura H. Computer-aided antibody design. Protein Eng. Des. Sel. 2012;25:507–521. doi: 10.1093/protein/gzs024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Farkona S, Diamandis EP, Blasutig IM. Cancer immunotherapy: the beginning of the end of cancer? BMC Med. 2016;14:73. doi: 10.1186/s12916-016-0623-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sharma S, Byrne H, O’Kennedy RJ. Antibodies and antibody-derived analytical biosensors. Essays Biochem. 2016;60:9–18. doi: 10.1042/EBC20150002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ohkuri T, et al. A Protein’s Conformational Stability Is an Immunologically Dominant Factor: Evidence That Free-Energy Barriers for Protein Unfolding Limit the Immunogenicity of Foreign Proteins. J. Immunol. 2010;185:4199–4205. doi: 10.4049/jimmunol.0902249. [DOI] [PubMed] [Google Scholar]
- 28.Yu YJ, et al. Boosting brain uptake of a therapeutic antibody by reducing its affinity for a transcytosis target. Sci. Transl. Med. 2011;3:84ra44. doi: 10.1126/scitranslmed.3002230. [DOI] [PubMed] [Google Scholar]
- 29.Tsumoto K, et al. Role of arginine in protein refolding, solubilization, and purification. Biotechnol. Prog. 2004;20:1301–1308. doi: 10.1021/bp0498793. [DOI] [PubMed] [Google Scholar]
- 30.Johnsson B, Löfås S, Lindquist G. Immobilization of Proteins to a Carboxymethyldextran-Modified Gold Surface for Biospecific Interaction Analysis in Surface Plasmon Resonance Sensors. Anal. Biochem. 1991;198:268–277. doi: 10.1016/0003-2697(91)90424-R. [DOI] [PubMed] [Google Scholar]
- 31.Myszka DG. Kinetic analysis of macromolecular interactions using surface plasmon resonance biosensors. Curr. Opin. Biotechnol. 1997;8:50–57. doi: 10.1016/S0958-1669(97)80157-7. [DOI] [PubMed] [Google Scholar]
- 32.Ross PD, Subramanian S. Thermodynamics of protein association reactions: forces contributing to stability. Biochemistry. 1981;20:3096–3102. doi: 10.1021/bi00514a017. [DOI] [PubMed] [Google Scholar]
- 33.Krissinel E, Henrick K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 2007;372:774–797. doi: 10.1016/j.jmb.2007.05.022. [DOI] [PubMed] [Google Scholar]
- 34.Hagemans D, van Belzen IAEM, Morán Luengo T, Rüdiger SGD. A script to highlight hydrophobicity and charge on protein surfaces. Front. Mol. Biosci. 2015;2:56. doi: 10.3389/fmolb.2015.00056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Abraham MJ, et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 2015;1–2:19–25. doi: 10.1016/j.softx.2015.06.001. [DOI] [Google Scholar]
- 36.Huang J, et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods. 2017;14:71–73. doi: 10.1038/nmeth.4067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.MacKerell AD, Jr., Feig M, Brooks CL. Improved treatment of the protein backbone in empirical force fields. J. Am. Chem. Soc. 2004;126:698–699. doi: 10.1021/ja036959e. [DOI] [PubMed] [Google Scholar]
- 38.Jo S, Kim T, Iyer VG, Im W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 2008;29:1859–1865. doi: 10.1002/jcc.20945. [DOI] [PubMed] [Google Scholar]
- 39.Darden T, York D, Pedersen L. Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993;98:10089–10092. doi: 10.1063/1.464397. [DOI] [Google Scholar]
- 40.Hess B, Bekker H, Berendsen HJC, Fraaije JGEM. LINCS: A Linear Constraint Solver for molecular simulations. J. Comput. Chem. 1997;18:1463–1472. doi: 10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H. [DOI] [Google Scholar]
- 41.Pettersen EF, et al. UCSF Chimera - A visualization system for exploratory research and analysis. J. Comput. Chem. 2004;25:1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.