Abstract
The Activator Protein 2 (AP-2) transcription factor (TF) family is vital for the regulation of gene expression during early development as well as carcinogenesis process. The review focusses on the AP-2α and AP-2γ proteins and their dualistic regulation of gene expression in the process of carcinogenesis. Both AP-2α and AP-2γ influence a wide range of physiological or pathological processes by regulating different pathways and interacting with diverse molecules, i.e. other proteins, long non-coding RNAs (lncRNA) or miRNAs. This review summarizes the newest information about the biology of two, AP-2α and AP-2γ, TFs in the carcinogenesis process. We emphasize that these two proteins could have either oncogenic or suppressive characteristics depending on the type of cancer tissue or their interaction with specific molecules. They have also been found to contribute to resistance and sensitivity to chemotherapy in oncological patients. A better understanding of molecular network of AP-2 factors and other molecules may clarify the atypical molecular mechanisms occurring during carcinogenesis, and may assist in the recognition of new diagnostic biomarkers.
Keywords: cancer, TFAP2A, TFAP2C, transcription factors
Introduction
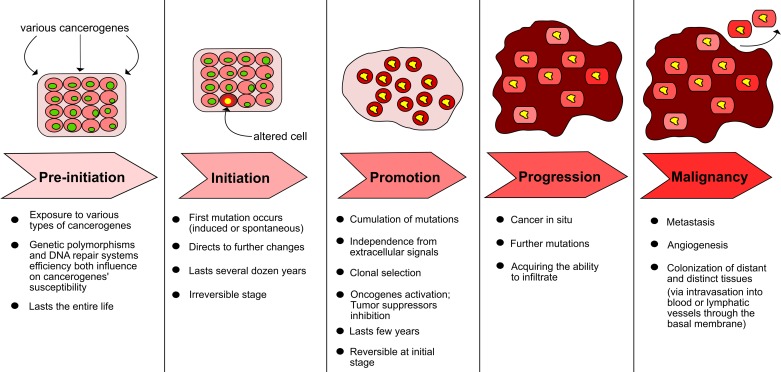
Genomic instability facilitates and speeds up tumor initiation and development, where several stepwise accumulations of dysfunction are implicated in the process. The acquisition of alterations; including but not limited to mutations at the nucleotide or chromosomal levels, where the appropriate repair checkpoints during cell division or at the epigenetic level are scarce, lead to multiple genetic changes. Initially the exposure of cells to tumor initiators (mutagens) and afterward tumor promoters (e.g. free radicals) remains the elementary event for tumor development [1,2]. Any deficiency of repair mechanisms or programmed cell death causes further errors present in metabolic processes or the control of cell division. Simultaneous activation of protooncogenes accompanied with suppressor gene inactivation or proliferation factor biosynthesis result in phenotypic changes in cells, followed by cancer cell infiltration and metastasis [3]. The steps of tumor development are presented in Figure 1.
Figure 1. Steps of cancerogenesis (based on [3]).
In addition to the number of basic factors known to play a role in cancer development [4], several theories exist regarding the process itself, three of which are the theory of Nature and Nurture [5], the Stem Cell Theory [6], and the Mutation/Epigenetic Theory, which illustrates the potential of epigenetic processes to mediate exposure–phenotype relationships [7]. Although these hypotheses are very informative, we focussed on the one described below.
Molecular paradigm concerning suppressors and oncogenes, or more precisely Somatic Mutation Theory (SMT) [8], is the hypothesis that arises in its original form in 1914 [9]. It proposes that carcinogenesis is driven by up- or down-regulation of alterations in cancer-related genes, two key examples being tumor suppressor genes (TSGs) and oncogenes (OCGs), resulting in changes in proliferation or apoptosis [10]. However, some cancer regulatory genes display both oncogenic and suppressive traits: for example, transcription factors (TFs) [11] such as the p53 protein that can function properly only if there are no mutations in any of its subunits or any Dominant-Negative Effect (DNE) [12]. Another example is Activator Protein 2γ (AP-2γ), a member of AP-2 family of TFs, that functions as an oncogene if localized in the nucleus, but this can be inhibited by interaction with WW Domain Containing Oxidoreductase (WWOX) suppressor [13]. On the other hand, the functioning of AP-2γ it has been found to act as an inducer of p21 protein expression, suggesting that it may suppress tumor development [14]. Similarly, the AP-2α protein also possesses duality of action depending on the affected signaling pathway [15,16]. The present review examines the effect of the biological functions of two members AP-2 TFs family: AP-2γ and AP-2α on cancers development and any accompanying phenomena.
General information about TFs
TFs can be considered in terms of mechanistic, functional, or structural properties. Structural classification of TFs is based on concurrent information about tertiary structure and sequence resemblance [17]. Briefly, there are ten distinguished sequence-homological superclasses of TFs, with 90% of human TFs fitting within the first three [18]. Further subdivisions, together with chosen examples, are given in Table 1.
Table 1. Superclasses of TFs (based on [19] and websites: http://gene-regulation.com/pub/databases/transfac/cl.html and http://tfclass.bioinf.med.uni-goettingen.de/).
| Superclass number | Superclass name | Quantity of classes | Quantity of families | Examples of TFs |
|---|---|---|---|---|
| 0 | Yet undefined DNA-binding domains (Superclass ‘0’) | 5 | 10 | PSPC1, RFXANK, NRF1 |
| 1 | Basic Domains | 3 | 18 | c-Jun, c-Fos, Nrf2 |
| 2 | Zinc-coordinating DNA-binding domains | 8 | 25 | ER, GATA-1, RXR-α |
| 3 | Helix–turn–helix | 7 | 22 | HOXA9, Oct-3/4, E2F-1 |
| 4 | Other all-α-helical DNA-binding domains | 2 | 8 | SOX2, TCF-7, UBF |
| 5 | α-Helices exposed by β-structures | 2 | 7 | MEF2, SRF, AIRE |
| 6 | Immunoglobulin fold | 7 | 16 | RelA, STAT1, p53 |
| 7 | β-Hairpin exposed by an α/β-scaffold | 2 | 3 | SMAD4, GCM1, NF-1A |
| 8 | β-Sheet binding to DNA | 2 | 2 | TBP, TBPL1, HMGA1 |
| 9 | β-Barrel DNA-binding domains | 1 | 1 | DbpA, YB-1, YBX2 |
Of the basic domains, Superclass 1, is termed as basic Helix–Span–Helix (bHSH) which comprises all TF AP-2 (TFAP2) representatives: α, β, γ, δ, and ε [20]. All factors in this family are crucial for gene expression in early development, regulating processes such as apoptosis or cell cycle [21]. The N-terminal region of TFAP2 factors consists of a transactivation domain, while the C-terminus contains ∼200 amino acids and is responsible for DNA binding and dimerization [22]. The dimerization domain is located inside the DNA-binding site [23], and comprise Proline/Glutamine-rich domains along with basic α-helix and bHSH. Although bHSH is unable to bind DNA when separated from the basic domain, it is still able to dimerize two family members [24]. All members of the bHSH class can recognize specific G/C-rich sequence motifs including the evolutionarily conserved binding sites GCCN3/4GGC, GCCN3/4GGG [25] or CCCCAGGC [26]. The binding region is formed of basic leucine zipper factors (bZIP) and basic helix–loop–helix factors (bHLH), which are also observed in other classes present in the basic domain superclass [24]. The members of bHSH are very similar to those of bHLH, but display a longer loop [27].
Activating enhancer-binding protein 2 – selected representatives
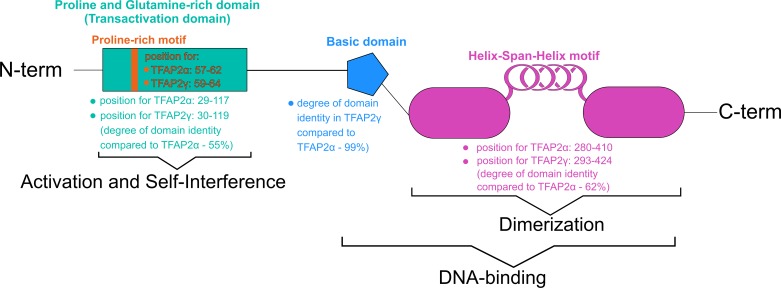
The AP-2 family, within the bHSH class, is characterized by specific regions. A schematic presentation of the domains is shown in Figure 2. TFAP2δ is a peculiar case in that it displays distinct binding affinity and a lack of critical residues with a PY (proline-rich) motif as example. PPxY can interact with WWOX proteins because it is recognized by certain WW (tryptophan) domains. Moreover, modulation capabilities as negative regulation of other AP-2 factors has been suggested in the literature [28]. In terms of gene localization, the majority of mammalian family members are encoded on chromosome 6, excluding AP-2γ and AP-2ε [22]. Selected information regarding chromosomal loci and their characteristics are summarized in Table 2.
Figure 2. Specific domains of AP-2 family with dissimilarities amongst TFAP2α and TFAP2γ (based on [22,23,29]).
Table 2. Chromosomal localization and characteristics of AP-2 family members (based on information from GeneCards and NCBI ‘Gene’ databases).
| AP-2 family member | Cytological location | Number of exons | Size/length | Orientation strand |
|---|---|---|---|---|
| TFAP2A | 6p24.3 | 11 | 26474 bases | Minus (−) |
| TFAP2B | 6p12.3 | 11 | 29910 bases | Plus (+) |
| TFAP2C | 20q13.31 | 7 | 9982 bases | Plus (+) |
| TFAP2D | 6p12.3 | 8 | 59490 bases | Plus (+) |
| TFAP2E | 1p34.3 | 7 | 21959 bases | Plus (+) |
AP-2 TFs originate in the nucleus [22]. The TFs form either homodimers or heterodimers [30]. Activity of those proteins includes modulation of transactivation potential [13], DNA-binding capability [31], degradation [32] or subcellular localization [33]; this can be achieved with the use of post-translational phosphorylation [34], sumoylation [35] or redox reactions [36]. Other proteins can either physically interact with AP-2 factors and bind to them, or only modulate their activity [22]. Furthermore, when loss of TF activity is observed due to mutation, precocious apoptosis or cell differentiation accompanied with proliferation impairment can occur [22]; however, AP-2 family members have also been found to be present at elevated levels in various types of tumors [33,37–39]. In this way, we hoped for a wider understanding of the role of TFAP2α and TFAP2γ in regulating the expression of genes that impact the clinically significant cancer phenotypes.
AP-2α
Gene
The TFAP2A gene encodes TF which can both activate [40] and inhibit [41] transcription of other genes simultaneously. It is localized on the minus strand of chromosome 6 its heterozygous mutations, mainly deletion, but also insertion or transition, can be observed in branchio-oculofacial syndrome (BOFS) [42,43]. The region contains 26474 bases at genomic location chr6:10393186-10419659 (cytogenetic band 6p24.3; Genome Reference Consortium Human Build 38) and six mRNAs are transcribed: REFSEQ NM_001032280.2, NM_001042425.1, NM_003220.2, XM_006715175.2, XM_011514833.2, and XM_017011232.1 (NCBI Reference Sequence Database). In addition, two antisense non-coding RNA molecules have been identified (Entrez gene IDs :100130275 for TFAP2A-AS1 and 109729173 for AS2).
Protein
The AP-2α protein encoded by the TFAP2A gene may act as either a homodimer or heterodimer when working with paralogs from its family. It recognizes the specific sequence 5′-GCCNNNGGC-3′ and regulates gene transcription by interacting with enhancer elements. AP-2α is also thought to be required to preserve lens integrity after vesicle formation [44]. Four transcripts, translated into distinct isoforms, have been considered in the UniProt KnowledgeBase (identifiers: P05549-1 for canonical sequence and three variants P05549-5, P05549-2, P05549-6) and these are given in Table 3. Like most of proteins, AP-2α can undergo post-translational modification (PTM), which affect protein activity or functionality (Table 4).
Table 3. Comparison of AP-2α isoforms (based on UniProt KnowledgeBase).
| AP-2α isoform | Length (amino acids) | Mass (Da) | Notes |
|---|---|---|---|
| Isoform ‘1’ (UniParc identifier: P05549-1) | 437 | 48062 | Canonical sequence; others refer to it |
| Isoform ‘2’ (UniParc identifier: P05549-5) | 431 | 47183 | Differs from canonical model in way that first 15 amino acids are substituted: MLWKLTDNIKYEDCE → MLVHSFSAM |
| Isoform ‘4′ (UniParc identifier: P05549-2) | 365 | 40557 | Altered amino acids from 296 to 437 as follows: EAVHLARDFG…SSDKEEKHRK → KRIHLLTRRN…SILLPSFPLP |
| Isoform ‘5’ (UniParc identifier: P05549-6) | 433 | 47440 | Differs from canonical model in way that first 15 amino acids are substituted: MLWKLTDNIKYEDCE → MSILAKMGDWQ |
Table 4. Localization and effect of PTMs in AP-2α (based on GeneCards database, neXtProt platform PhosphoSitePlus resource).
| Type of PTM | Position in protein | Notes (if available) |
|---|---|---|
| Sumoylation | 10 (lysine) | Leads to inhibition of transcriptional activity |
| Phosphorylation | 73 (tyrosine) | None |
| Phosphorylation | 119 (serine) | None |
| Sumoylation | 177 (lysine) | None |
| Phosphorylation | 181 (serine) | None |
| Sumoylation | 184 (lysine) | None |
| Phosphorylation | 185 (serine) | None |
| Phosphorylation | 187 (serine) | None |
| Phosphorylation | 219 (serine) | Regulate molecular association and activity; induced transcription process |
| Phosphorylation | 239 (serine) | None |
| Phosphorylation | 258 (serine) | Regulate molecular association; altered transcription process |
| Mono-methylation | 263 (arginine) | None |
| Phosphorylation | 326 (serine) | None |
| Phosphorylation | 333 (threonine) | None |
| Phosphorylation | 428 (serine) | None |
| Phosphorylation | 429 (serine) | Induced transcription process |
AP-2γ
Gene
TFAP2C is the third member of the AP-2 family and is expressed as a sequence-specific TF that activates a number of developmental genes responsible for eyes, face, and limbs formation or neural tube development. It is located on the plus strand on chromosome 20 and transcribes only one mRNA variant (NCBI Reference Sequence Database, REFSEQ accession number: NM_003222.3) and to contain 9982 bases at genomic location chr20:56629302-56639283 (cytogenetic band 20q13.31 - Genome Reference Consortium Human Build 38).
Protein
The AP-2γ TF shares the same properties with TFAP2α with regard to dimer formation, recognition of consensus sequence, and its effect on both cellular and viral enhancers. Only the canonical protein sequence and one isoform are given in the UniProt KnowledgeBase: Q92754-1 and Q92754-2, respectively (Table 5). Similar to AP-2α, it is modified after translation, however only one of them has a biological effect (Table 6).
Table 5. Comparison of AP-2γ isoforms (based on UniProt KnowledgeBase).
| AP-2γ isoform | Length (amino acids) | Mass (Da) | Notes |
|---|---|---|---|
| Isoform ‘1’ (UniParc identifier: Q92754-1) | 450 | 49,177 | Canonical sequence; others refer to it |
| Isoform ‘2’ (UniParc identifier: Q92754-2) | 281 | 31,010 | Differs from canonical model in way that first 169 amino acids are missing |
Table 6. Localization and effect of PTMs in AP-2γ (based on GeneCards database, neXtProt platform PhosphoSitePlus resource).
| Type of modification | Position in protein | Notes (if available) |
|---|---|---|
| Sumoylation | 10 (lysine) | Leads to inhibition of transcriptional activity |
| Phosphorylation | 252 (serine) | None |
| Mono-methylation | 276 (arginine) | None |
| Phosphorylation | 434 (serine) | None |
| Phosphorylation | 438 (serine) | None |
| Ubiquitylation | 444 (lysine) | None |
| Ubiquitylation | 447 (lysine) | None |
AP-2 interactions with proteins, long non-coding RNA, and miRNA molecules
Proteins
Both the AP-2α and AP-2γ proteins play an essential role in many important biological processes. Mutations in TFAP2A are known to be linked to retinal defects and a greater possibility of disturbances in eye development [45]. This gene is also important in face or limb development since its expression is observed during frontal nasal process (FNP), paired lateral nasal processes, and limb bud mesenchyme (LBM) [46]. Other processes involving both TFAP2A and TFAP2C functionality concerns generation of neural tube [47,48] or body wall [49]. However, while TFAP2C is a gene implicated in inhibition of somatic differentiation in germ cells [50] or repression of neuroectodermal differentiation and pluripotency maintenance [51], TFAP2A plays a key role in kidney development [52]. The abnormal expression of TFAP2A may lead to BOFS and anophthalmia-microphthalmia syndrome [53], while human placenta defects are associated with TFAP2C overexpression [54]. The participation of AP-2α and AP-2γ in developmental processes, along with management of other events by means of specific interactions with proteins are presented in Table 7.
Table 7. Influence of AP-2α and AP-2γ factors on selected developmental processes, diseases, and interactive molecules (based on GeneCards, Reactome databases, and Atlas of Genetics and Cytogenetics in Oncology and Haematology).
| AP-2α | AP-2γ | |
|---|---|---|
| Participation in development processess – major | Face, eye, limb, body wall, neural tube development | |
| Participation in development processess – other | Early morphogenesis of lens vesicle; kidney development | Male gonad development |
| Associated diseases | BOFS; ectopic thymus; anophthalmia-microphthalmia syndrome | Exencephaly; melanoma; pre-eclampsia |
| Mutual interactions | Interacts with WWOX, CITED2, CITED4, UBE2I, KCTD1, KCTD15, EP300; Suppresses MCAM/MUC18, C/EBPα; Stimulates transcriptional activation of PITX2 | |
| Exclusive interactions | Along with binding NPM1 – represses HSPD1 gene expression, inhibiting formation of mitochondrial chaperonin; | Along with MTA1 – mediates epigenetic regulation of ESR1 expression in breast cancer (BCa); Interacts with KDM5B |
| Stimulates APOE gene transcription in co-operation with DEK; | ||
| Interacts with RALBP1 (in complex containing NUMB and EPN1) during interphase and mitosis | ||
Both AP-2α and AP-2γ interact with WWOX protein [44,55] while the latter is described in more detail, probably due to lower affinity of the AP-2α PPxY motif (59PPPY62) than the AP-2γ PPxY motif (56PPPYFPPPY64) [13]. The binding of AP-2 factors by WWOX suppresses their transcriptional transactivation, causing sequestration of proteins in the cytoplasm [56], thus reducing their oncogenic activity by triggering their redistribution from the nucleus [13]. It was also suggested that modulation of both α and γ AP-2 factors could be important from clinical point of view [56].
In addition to the examples summarized above, three interactions considering proliferation and cell cycle regulation are worth mentioning. First, the AP-2γ homodimer is capable of binding the region of EGFR gene promoter, therefore stimulating its expression and regulating tumor growth and survival in HER2 positive breast cancer patients [57,58]. However, when it comes to AP-2α, it acts conversely to AP-2γ in terms of proliferation but with a different target. VEGFA gene promoter consists of response elements recognized by AP-2α and, as a result, its expression is inhibited [15].
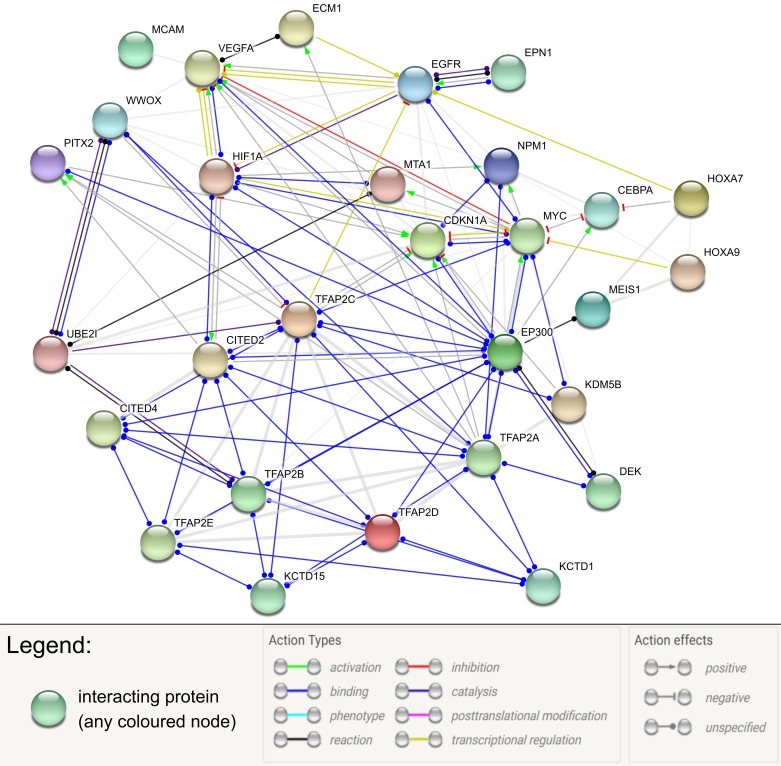
Finally, the cell cycle can be controlled by Cyclin-Dependent Kinase Inhibitor 1A (CDKN1A) regulation, but its inhibition or stimulation depends on the AP-2 member. TFAP2α is thought to stimulate CDKN1A by direct interaction with two response elements in its promoter [59,60]. In contrast, the formation of the AP2γ homodimer provides a possibility to interact with the previously mentioned KDM5B along with MYC. This complex is thought to influence the progression of the cell cycle and carcinogenesis by repressing CDKN1A expression [61]. AP-2 factors have an abundant interaction network with other molecules and are associated with several important processes (Figure 3).
Figure 3. Interaction network of AP-2 factors with selected proteins (STRING database).
Long non-coding RNA and microRNA
Just like proteins, AP-2 interacts with RNA. In many cases, this molecular feedback regulates cancer progression, tumor growth, metastasis, drug resistance, metabolism, or even the occurrence of congenital anomalies.
Long non-coding RNA (lncRNA) derived from the growth arrest-specific 5 (GAS5) gene has been found to participate in progression of glioma. In the promoter region, the functional polymorphism (rs145204276) with genetic variation in the presence or absence of specific sequence (insertion-deletion, abbreviated in/del) may influence lncRNA expression by recognition by recruited TFs. The most abundant TFs were three members of the AP-2 family – TFAP2A, B, and C, with TFAP2A gene expression correlating positively with GAS5 expression in glioma [62]. In a different case, progression of colorectal cancer (CRC) was found to be associated with down-regulated AP-2α expression through direct targetting by a CRC-associated lncRNA (CAAL) molecule [63]. The down-regulation of AP-2α had an impact on Wnt/β-catenin pathway which was activated in the presence of the key regulator CCAL; this phenomenon was also observed in hepatocellular carcinoma (HCC) [64]. In fact, the entire regulatory axis could be valuable in CRC-targetted therapy: lncRNA UCA1 is also up-regulated in CRC and involved in cellular migration, its expression correlates with that of both TFAP2A and TFAP2C, and its dependence relies on the occurrence of TF-binding sites (TFBSs) that are located upstream of the start site of UCA1 transcription [65].
An example of a molecule that maintains the balance between pluripotency and differentiation is lncPRESS1: an lncRNA that displays control over the expression of TFAP2C and other pluripotency genes in human embryonic stem cells (hESCs). lncRNA depletion leads to differentiation by altering pluripotency and down-regulating associated genes [66]. It has been proposed that a group of lncRNAs responsive to oxygen-glucose deprivation (OGD) may act as mediators of ischemic stimuli to the endothelium and that TFAP2C is one of the TFs affecting OGD-responsive lncRNAs, such as lnc-OGD4751, under specific conditions [67].
In the case of miRNA, most of the molecules described herein have been identified as regulators of tumor progression or treatment resistance. In melanoma cell lines, two miRNA molecules interacting with AP-2 family members were indicated as being significantly vital for cancer development: miR-214 and miR-638. The former is able to influence both TFAP2C [68] and TFAP2A [69], thus contributing to melanoma progression (A375P cell line); it was found that TFAP2γ is an important factor which permits miR-214 to guide tumor progression [68]. Additionally, the abolition of TFAP2A expression in melanoma was confirmed to increase malignancy [70]. The latter, miR-638, has been identified as an overexpressed molecule in metastatic melanomas in relevance to primary tumor. Promoter analysis indicated that together with TFAP2A, they develop regulatory feedback that possesses double-negative character [71]. Suppression of p53-mediated apoptosis and autophagy combined with down-regulation of AP-2α provides favorable conditions for metastasis. Lung cancer is also modulated by a couple of miRNAs interacting with the AP-2 family. The first, miR-1254, a negative regulator of heme oxygenase-1 (HO-1), is able to induce apoptosis and inhibit cell cycle progression in non-small cell lung carcinoma (NSCLC) by a two-sided approach. First, it directly targets 3′-UTR of HO-1 mRNA, eliminating its function in cancer, i.e. angiogenesis stimulation or inflammatory response inhibition [72,73]. Second, it prevents TFAP2α from functioning as a transactivator of HO-1, influencing the targetting via the non-seed sequence. When present, these conditions intensify the suppressive effect on the tumor [74].
However, overexpression of TFAP2C in NSCLC leads to excessive cell cycle activation which promotes tumor aggressiveness: the proposed model of action consists of simultaneous induction of oncogenic miR-183 and down-regulation of suppressive miR-33a. This entails, respectively, the blocking of AKAP12-mediated inhibition of cyclin D1 and the activation of cell cycle progression by cyclin-dependent kinase 6 (Cdk6) [75]. Studies of the Slug-mediated network have found that another tumor progression in NSCLC can be also be promoted by suppressing TFAP2C. Since Slug directly binds to the promoter region of miR-137, this enhances RNA expression in lung cancer cells. In turn, miR-137 suppresses AP-2γ expression directly, which increases metastasis and tumor invasion [76]. A similar pathway, but with a different effect, could be observed in neuroblastoma: miR-200a directly targets TFAP2C, thus tumor growth and cell proliferation [77]. Targetting AP-2 factors could as well lead to drug resistance, as confirmed in pancreatic cancer, where gemcitabine resistance is obtained via TFAP2C suppression by miR-10a-5p [78], or in bladder cancer where TFAP2A is suppressed by miR-193a-5p [79]. In both cases, this could indicate informative new therapeutic targets to remove resistance to treatment.
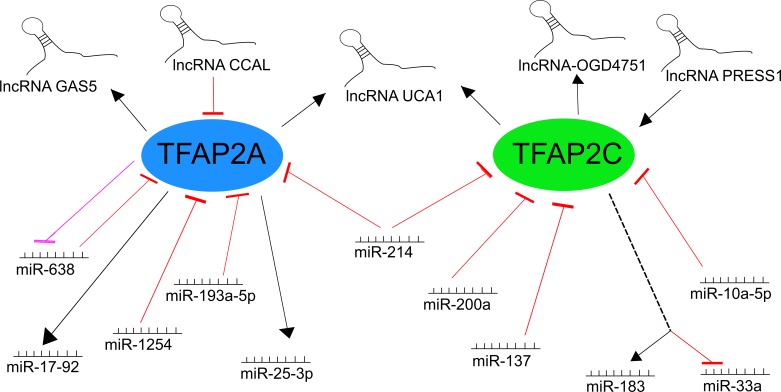
Other miRNA-TFAP2 combinations have been observed in other contexts. AP-2α was listed as a regulator of at least two miRNAs that are responsible for either metabolism regulation or proper craniofacial phenotype (or disrupted when gene is mutated). In the first case, AP-2α binds to the core promoter of miR-25-3p, thus increasing its expression and allowing it to target the Akt1 gene and enhance the metabolism in C2C12 cells [80]. In the second, it recognizes multiple binding sites in miR-17-92 chromatin that induce miRNA expression and allow proper midface development through regulation of Tbx genes. Unfortunately, occurrence of mutant miR-17-92 could result in cleft lip and palate (CL/P) which is a common congenital anomaly [81]. As clearly seen above, TFAP2A and TFAP2C are of great importance in a number of processes in which they play a controlling role or are themselves regulated by RNA molecules. A summary of the above interactions is presented in Figure 4.
Figure 4. Interaction network of AP-2α/γ factors with selected lncRNA and miRNA molecules (based on [62–81]).
Clinical data, participation in developmental processes, and cancers
Quantitative transcriptomics analyses have found TFAP2A and TFAP2C to occur predominantly in the placenta, skin, or esophagus, and for the two to be expressed at significantly different levels in the kidney, in favor of TFAP2A [82]. In terms of developmental processes, abnormal AP-2α expression may result in incorrect placenta maturation in high-risk pregnancies [83]. Likewise, high expression of AP-2γ can be preventive during pre-eclampsia, via blood pressure regulation by compensation of vasoconstructory peptides excess [84]. In case of tumorigenesis, AP-2α has demonstrated protective properties against melanoma progression, which can be disrupted following cleavage by caspase-6 [85]. However, in melanoma, AP-2γ overexpression is associated with unfavorable processes including vascular invasion or increased vascularity [86]. At last, AP-2γ is up-regulated in esophageal adenocarcinoma, suggesting it may play a progressive role [87], while high AP-2α expression correlates with longer overall survival (OS) in esophageal squamous cell carcinoma and has been proposed as a valuable prognostic biomarker in this disease [88]. Apparently, those two AP-2 family members are thought to have oncogenic or suppressive characteristics depending on the cancer tissue or interaction with other molecules. The function of TFAP2A and TFAP2C in selected cancers with implicated pathways (if any) is summarized in Tables 8 and 9.
Table 8. Regulatory role of AP-2α in various types of cancer along with involved molecular pathways (based on [15,16,89–104]).
| Protein | Type of cancer | Regulation | Function | Pathways (if available) | References |
|---|---|---|---|---|---|
| AP-2α | Hepatocellular carcinoma | Suppressor | TFAP2A overexpression decreases cell migration and invasion, and also leads to inhibition of cell growth and proliferation | HIF-1α-mediated VEGF/PEDF signaling pathway; | [15,89–92] |
| β-catenin/TCF/LEF signaling; | |||||
| Bax/cytochrome c/Apaf1/caspase9-dependent mitochondrial pathway; | |||||
| CdK-inhibitor p21WAF in p53-dependent and p53-independent pathways | |||||
| AP-2α | Breast cancer | Suppressor | Reduced TFAP2A is associated with more aggressive breast cancer | [93,94] | |
| TFAP2A high expression is related to sensitiveness to chemotherapeutic drugs (due to massive induction of apoptosis) | |||||
| AP-2α | Glioblastoma | Suppressor | TFAP2A reduces tumor cell growth, increases cell death, attenuates cell migration, and endothelial tube formation | [95] | |
| AP-2α | Melanoma | Suppressor | Comparison of stage 4 melanomas compared with non-stage 4 display, that silenced TFAP2A by aberrant CpG methylation of its promoter is most decreased in higher stages | [96] | |
| AP-2α | Gastric cancer | Suppressor | TFAP2A can reverse the multidrug resistance (MDR) | Notch signaling pathway | [97,98] |
| Lower level of TFAP2A leads to unfavorable prognosis for patients | |||||
| AP-2α | Prostate cancer | Suppressor | Loss of TFAP2A leads to decreased cellular zinc uptake which is essential for tumor development | [99] | |
| AP-2α | Colorectal cancer | Suppressor | TFAP2A negatively regulates downstream targets of β-catenin/TCF/LEF | Wnt signaling pathway | [90] |
| AP-2α | Neuroblastoma | Oncogene | TFAP2A is overexpressed in cell lines derived from high-stage tumors | [100] | |
| AP-2α | Pancreatic cancer | Oncogene | Cell lines express high nuclear levels of TFAP2A | [101] | |
| AP-2α variant 6 seems to be specific for pancreatic cancer | |||||
| AP-2α | Acute myeloid leukemia | Oncogene | TFAP2A affects Hoxa7, Hoxa9, and Meis1 which are involved in leukemogenesis | [102] | |
| AP-2α | Squamous cell carcinomas | Oncogene | AP-2α is involved in complex keratinocyte biology including proliferation, differentiation, and carcinogenesis | [103,104] | |
| AP-2α | Nasopharyngeal carcinoma | Oncogene | TFAP2A overexpression correlates with HIF-1α expression along with advanced tumor stage, local invasion, clinical progression, or poor prognosis | HIF-1α-mediated VEGF/PEDF signaling pathway | [16] |
Table 9. Regulatory role of AP-2γ in various types of cancer along with involved molecular pathways (based on [38,75–77,105–108]).
| Protein | Type of cancer | Regulation | Function | Pathways (if available) | References |
|---|---|---|---|---|---|
| AP-2γ | Breast cancer | Oncogene | TFAP2C expression is associated with proliferation, disease progression, and endocrine therapy resistance | [105] | |
| TFAP2C directly represses CDKN1A gene thereby promoting proliferation | |||||
| High TFAP2C and low CD44 expression are associated with pathologic complete response (pCR) after neoadjuvant chemotherapy | |||||
| AP-2γ | Melanoma | Oncogene | High level of TFAP2C regulates ECM1 overexpression which is associated with poor prognosis | [106] | |
| AP-2γ | Testicular cancer | Oncogene | TFAP2C is a novel marker of testicular CIS and CIS-derived tumors along with its involvement in self-renewal and survival of immature germ cells and tissue-specific stem cells | [38] | |
| AP-2γ | Neuroblastoma | Oncogene | TFAP2C knockdown results in inhibition of cell proliferation and tumor growth | [77] | |
| AP-2γ | Primary ovarian tumors | Oncogene | Overexpressed in advanced-stage cancers compared with early-stage carcinomas | [107] | |
| AP-2γ | Lung cancer | Oncogene | TFAP2C overexpression promotes cell viability, proliferation, and cell cycle progression | AK1 signaling MAPK and Snail pathways activated by TGFBR1-PAK1 signaling | [75,108] |
| AP-2γ | Lung cancer | Suppressor | TFAP2C blocks AKAP12-mediated cyclin D1 inhibition (by inducing the overexpression of oncogenic miR-183) | [76] | |
| TFAP2C activates Cdk6-mediated cell cycle progression (by down-regulating tumor-suppressive miR-33a) | |||||
| Patients with low-level expression of Slug and miR-137 but high-level expression of TFAP2C experienced better survival |
Moreover, these TFs are essential for chemotherapy sensitivity in patients. In vitro study demonstrates that AP-2α increases the sensitivity to cisplatin in endometrial cancer cells, and also proving that single nucleotide polymorphism (SNP) rs1045385 A>C variation in the 3′-UTR region of the TFAP2A gene could become a prognostic marker in the treatment of cisplatin. This is due to the fact that only the A allele is recognized by miR-200b/200c/429 family which reduces the expression of the AP-2α suppressor thereby correlating with resistance to cisplatin [109]. Furthermore, studies based on siRNA against AP-2α in MDA-MB-231 cells have shown that the use of 5-aza-2′-deoxycytidine alone for breast cancer treatment does not increase cell apoptosis or cancer sensitivity to chemotherapy; however, proper expression of TFAP2A combined with chemotherapy resulted in loss of tumorigenesis, reduction in colony formation, and heightened chemosensitivity [110]. At last, AP-2α is thought to be valuable in gastric cancer during occurrence of multidrug resistance (MDR). Using a vector overexpressing AP-2α in SGC7901/VCR cells, Lian et al. [97] demonstrated inhibition of the Notch signaling pathway which could potentially play a role in reversing drug resistance. Additionally, increased cell cycle arrest (G0/G1 phase) and apoptosis were observed during AP-2α overexpression [97]. In turn, the role of TFAP2C in terms of resistance to treatment appears to be reversed in comparison with TFAP2A. In patients with invasive breast cancer, both in vitro and clinical experiments indicated that AP-2γ overexpression determines resistance to tamoxifen or faslodex, which is possible due to inhibition of growth arrest and thus a worse outcome [111].
Conclusion
Both TFAP2A and TFAP2C hold distinct roles as oncogenes or tumor suppressors in various tumor models. However, the former tends to play a protective role, while the latter is predominantly up-regulated and participates in pathways responsible for cancer progression. Although AP-2α and AP-2γ are generally similar with regard to their activities, they do differ in a number of ways. Both act by regulating the specific expression of many genes which are frequently related to tumorigenesis, together with their products: proteins, lncRNAs, and miRNAs. Key examples of these products are p21, WWOX, EGFR, VEGF-A, lncRNA GAS5, lncRNA UCA1, miR-183 or miR-638. Further studies are needed to better understand the molecular network associated with AP-2 members, thus providing access to a wider range of elementary biomarkers which can be used to improve cancer diagnosis and treatment.
Acknowledgments
We thank Raneem Hammouz for useful suggestion(s) and helping in the preparation of this manuscript.
Abbreviations
- AP-2
activator protein 2
- bHLH
basic helix–loop–helix
- bHSH
basic helix–span–helix
- BOFS
branchio-oculofacial syndrome
- bZIP
basic leucine-zipper factor
- CCAL
colorectal cancer-associated lncRNA
- CDKN1A
cyclin-dependent kinase inhibitor 1A
- CRC
colorectal cancer
- DNE
dominant-negative effect
- FNP
frontal nasal process
- GAS5
growth arrest-specific 5
- HCC
hepatocellular carcinoma
- hESC
human embrionic stem cell
- HO-1
heme oxygenase-1
- LBM
limb bud mesenchyme
- lncRNA
long non-coding RNA
- MDR
multidrug resistance
- NSCLC
non-small cell lung carcinoma
- OCG
oncogene gene
- OGD
oxygen-glucose deprivation
- OS
overall survival
- PTM
post-translational modification
- SMT
somatic mutation theory
- TF
transcription factor
- TFAP
transcription factor activator protein
- TFBS
transcription factor-binding site
- TSG
tumor suppressor gene
- WWOX
WW domain containing oxidoreductase
Funding
This work was supported by the National Science Centre of Poland [grant number 2016/21/B/NZ2/01823].
Competing interests
The authors declare that there are no competing interests associated with the manuscript.
References
- 1.Boyland E. (1985) Tumour initiators, promoters, and complete carcinogens. Br. J. Ind. Med. 42, 716–718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Klaunig J.E., Kamendulis L.M. and Hocevar B.A. (2010) Oxidative stress and oxidative damage in carcinogenesis. Toxicol. Pathol. 38, 96–109 10.1177/0192623309356453 [DOI] [PubMed] [Google Scholar]
- 3.Cooper G.M. and Hausman R.E. (2007) The Cell: A Molecular Approach, pp. 719–728, ASM Press, Washington, D.C. [Google Scholar]
- 4.Baba A.I. and Catoi C. (2007) Comparative Oncology, The Publishing House of the Romanian Academy, Bucharest, Chapter 2, C arcinogenesis [PubMed] [Google Scholar]
- 5.Hyndman I.J. (2016) Review: the contribution of both nature and nurture to carcinogenesis and progression in solid tumours. Cancer Microenviron. 9, 63–69 10.1007/s12307-016-0183-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Jilkine A. and Gutenkunst R.N. (2014) Effect of dedifferentiation on time to mutation acquisition in stem cell-driven cancers. PLoS Comput. Biol. 10, e1003481 10.1371/journal.pcbi.1003481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cortessis V.K., Thomas D.C., Levine A.J., Breton C.V., Mack T.M., Siegmund K.D.. et al. (2012) Environmental epigenetics: prospects for studying epigenetic mediation of exposure-response relationships. Hum. Genet. 131, 1565–1589 10.1007/s00439-012-1189-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Paduch R. (2015) Theories of cancer origin. Eur. J. Cancer Prev. 24, 57–67 10.1097/CEJ.0000000000000024 [DOI] [PubMed] [Google Scholar]
- 9.Sonnenschein C. and Soto A.M. (2016) Carcinogenesis explained within the context of a theory of organisms. Prog. Biophys. Mol. Biol. 122, 70–76 10.1016/j.pbiomolbio.2016.07.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhu K., Liu Q., Zhou Y., Tao C., Zhao Z., Sun J.. et al. (2015) Oncogenes and tumor suppressor genes: comparative genomics and network perspectives. BMC Genomics 16, S8 10.1186/1471-2164-16-S7-S8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lou X., Zhang J., Liu S., Xu N. and Liao D.J. (2014) The other side of the coin: the tumor-suppressive aspect of oncogenes and the oncogenic aspect of tumor-suppressive genes, such as those along the CCND-CDK4/6-RB axis. Cell Cycle 13, 1677–1693 10.4161/cc.29082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Willis A., Jung E.J., Wakefield T. and Chen X. (2004) Mutant p53 exerts a dominant negative effect by preventing wild-type p53 from binding to the promoter of its target genes. Oncogene 23, 2330–2338 10.1038/sj.onc.1207396 [DOI] [PubMed] [Google Scholar]
- 13.Aqeilan R.I., Palamarchuk A., Weigel R.J., Herrero J.J., Pekarsky Y. and Croce C.M. (2004) Physical and functional interactions between the Wwox tumor suppressor protein and the AP-2gamma transcription factor. Cancer Res. 64, 8256–8261 10.1158/0008-5472.CAN-04-2055 [DOI] [PubMed] [Google Scholar]
- 14.Li H., Goswami P.C. and Domann F.E. (2006) AP-2gamma induces p21 expression, arrests cell cycle, and inhibits the tumor growth of human carcinoma cells. Neoplasia 8, 568–577 10.1593/neo.06367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ruiz M., Pettaway C., Song R., Stoeltzing O., Ellis L. and Bar-Eli M. (2004) Activator protein 2alpha inhibits tumorigenicity and represses vascular endothelial growth factor transcription in prostate cancer cells. Cancer Res. 64, 631–638 10.1158/0008-5472.CAN-03-2751 [DOI] [PubMed] [Google Scholar]
- 16.Shi D., Xie F., Zhang Y., Tian Y., Chen W., Fu L.. et al. (2014) TFAP2A regulates nasopharyngeal carcinoma growth and survival by targeting HIF-1alpha signaling pathway. Cancer Prev. Res. 7, 266–277 10.1158/1940-6207.CAPR-13-0271 [DOI] [PubMed] [Google Scholar]
- 17.Ehsani R., Bahrami S. and Drablos F. (2016) Feature-based classification of human transcription factors into hypothetical sub-classes related to regulatory function. BMC Bioinformatics 17, 459 10.1186/s12859-016-1349-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wingender E., Schoeps T., Haubrock M. and Donitz J. (2015) TFClass: a classification of human transcription factors and their rodent orthologs. Nucleic Acids Res. 43, D97–D102 10.1093/nar/gku1064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Matys V., Kel-Margoulis O.V., Fricke E., Liebich I., Land S., Barre-Dirrie A.. et al. (2006) TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 10.1093/nar/gkj143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Orlic-Milacic M. (2016) Transcriptional Regulation by the AP-2 (TFAP2) Family Of Transcription Factors Current Neurology and Neuroscience Reports, U.S. National Library of Medicine, https://www.ncbi.nlm.nih.gov/biosystems/1427839 [Google Scholar]
- 21.Hilger-Eversheim K., Moser M., Schorle H. and Buettner R. (2000) Regulatory roles of AP-2 transcription factors in vertebrate development, apoptosis and cell-cycle control. Gene 260, 1–12 10.1016/S0378-1119(00)00454-6 [DOI] [PubMed] [Google Scholar]
- 22.Eckert D., Buhl S., Weber S., Jager R. and Schorle H. (2005) The AP-2 family of transcription factors. Genome Biol. 6, 246 10.1186/gb-2005-6-13-246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kannan P. and Tainsky M.A. (1999) Coactivator PC4 mediates AP-2 transcriptional activity and suppresses ras-induced transformation dependent on AP-2 transcriptional interference. Mol. Cell. Biol. 19, 899–908 10.1128/MCB.19.1.899 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Williams T. and Tjian R. (1991) Characterization of a dimerization motif in AP-2 and its function in heterologous DNA-binding proteins. Science 251, 1067–1071 10.1126/science.1998122 [DOI] [PubMed] [Google Scholar]
- 25.Mohibullah N., Donner A., Ippolito J.A. and Williams T. (1999) SELEX and missing phosphate contact analyses reveal flexibility within the AP-2[alpha] protein: DNA binding complex. Nucleic Acids Res. 27, 2760–2769 10.1093/nar/27.13.2760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mitchell P.J., Wang C. and Tjian R. (1987) Positive and negative regulation of transcription in vitro: enhancer-binding protein AP-2 is inhibited by SV40 T antigen. Cell 50, 847–861 10.1016/0092-8674(87)90512-5 [DOI] [PubMed] [Google Scholar]
- 27.Bolander F.F. (2008) Molecular Endocrinology, pp. 387–443, Elsevier Academic Press, Amsterdam [Google Scholar]
- 28.Zhao F., Satoda M., Licht J.D., Hayashizaki Y. and Gelb B.D. (2001) Cloning and characterization of a novel mouse AP-2 transcription factor, AP-2delta, with unique DNA binding and transactivation properties. J. Biol. Chem. 276, 40755–40760 10.1074/jbc.M106284200 [DOI] [PubMed] [Google Scholar]
- 29.LiCalsi C., Christophe S., Steger D.J., Buescher M., Fischer W. and Mellon P.L. (2000) AP-2 family members regulate basal and cAMP-induced expression of human chorionic gonadotropin. Nucleic Acids Res. 28, 1036–1043 10.1093/nar/28.4.1036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wenke A.K. and Bosserhoff A.K. (2010) Roles of AP-2 transcription factors in the regulation of cartilage and skeletal development. FEBS J. 277, 894–902 10.1111/j.1742-4658.2009.07509.x [DOI] [PubMed] [Google Scholar]
- 31.Mazina O.M., Phillips M.A., Williams T., Vines C.A., Cherr G.N. and Rice R.H. (2001) Redistribution of transcription factor AP-2alpha in differentiating cultured human epidermal cells. J. Invest. Dermatol. 117, 864–870 10.1046/j.0022-202x.2001.01472.x [DOI] [PubMed] [Google Scholar]
- 32.Li M., Wang Y., Hung M.C. and Kannan P. (2006) Inefficient proteasomal-degradation pathway stabilizes AP-2alpha and activates HER-2/neu gene in breast cancer. Int. J. Cancer 118, 802–811 10.1002/ijc.21426 [DOI] [PubMed] [Google Scholar]
- 33.Pellikainen J., Naukkarinen A., Ropponen K., Rummukainen J., Kataja V., Kellokoski J.. et al. (2004) Expression of HER2 and its association with AP-2 in breast cancer. Eur. J. Cancer 40, 1485–1495 10.1016/j.ejca.2004.02.020 [DOI] [PubMed] [Google Scholar]
- 34.Garcia M.A., Campillos M., Marina A., Valdivieso F. and Vazquez J. (1999) Transcription factor AP-2 activity is modulated by protein kinase A-mediated phosphorylation. FEBS Lett. 444, 27–31 10.1016/S0014-5793(99)00021-6 [DOI] [PubMed] [Google Scholar]
- 35.Zhong L., Wang Y., Kannan P. and Tainsky M.A. (2003) Functional characterization of the interacting domains of the positive coactivator PC4 with the transcription factor AP-2alpha. Gene 320, 155–164 10.1016/S0378-1119(03)00823-0 [DOI] [PubMed] [Google Scholar]
- 36.Huang Y. and Domann F.E. (1998) Redox modulation of AP-2 DNA binding activity in vitro. Biochem. Biophys. Res. Commun. 249, 307–312 10.1006/bbrc.1998.9139 [DOI] [PubMed] [Google Scholar]
- 37.Jager R., Friedrichs N., Heim I., Buttner R. and Schorle H. (2005) Dual role of AP-2gamma in ErbB-2-induced mammary tumorigenesis. Breast Cancer Res. Treat. 90, 273–280 10.1007/s10549-004-4815-x [DOI] [PubMed] [Google Scholar]
- 38.Hoei-Hansen C.E., Nielsen J.E., Almstrup K., Sonne S.B., Graem N., Skakkebaek N.E.. et al. (2004) Transcription factor AP-2gamma is a developmentally regulated marker of testicular carcinoma in situ and germ cell tumors. Clin. Cancer Res. 10, 8521–8530 10.1158/1078-0432.CCR-04-1285 [DOI] [PubMed] [Google Scholar]
- 39.Beger M., Butz K., Denk C., Williams T., Hurst H.C. and Hoppe-Seyler F. (2001) Expression pattern of AP-2 transcription factors in cervical cancer cells and analysis of their influence on human papillomavirus oncogene transcription. J. Mol. Med. 79, 314–320 10.1007/s001090100211 [DOI] [PubMed] [Google Scholar]
- 40.Wang D., Shin T.H. and Kudlow J.E. (1997) Transcription factor AP-2 controls transcription of the human transforming growth factor-alpha gene. J. Biol. Chem. 272, 14244–14250 10.1074/jbc.272.22.14244 [DOI] [PubMed] [Google Scholar]
- 41.Liu H., Tan B.C., Tseng K.H., Chuang C.P., Yeh C.W., Chen K.D.. et al. (2007) Nucleophosmin acts as a novel AP2alpha-binding transcriptional corepressor during cell differentiation. EMBO Rep. 8, 394–400 10.1038/sj.embor.7400909 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Milunsky J.M., Maher T.A., Zhao G., Roberts A.E., Stalker H.J., Zori R.T.. et al. (2008) TFAP2A mutations result in branchio-oculo-facial syndrome. Am. J. Hum. Genet. 82, 1171–1177 10.1016/j.ajhg.2008.03.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Tekin M., Sirmaci A., Yuksel-Konuk B., Fitoz S. and Sennaroglu L. (2009) A complex TFAP2A allele is associated with branchio-oculo-facial syndrome and inner ear malformation in a deaf child. Am. J. Med. Genet. A 149A, 427–430 10.1002/ajmg.a.32619 [DOI] [PubMed] [Google Scholar]
- 44.Kerr C.L., Zaveri M.A., Robinson M.L., Williams T. and West-Mays J.A. (2014) AP-2alpha is required after lens vesicle formation to maintain lens integrity. Dev. Dyn. 243, 1298–1309 10.1002/dvdy.24141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Gestri G., Osborne R.J., Wyatt A.W., Gerrelli D., Gribble S., Stewart H.. et al. (2009) Reduced TFAP2A function causes variable optic fissure closure and retinal defects and sensitizes eye development to mutations in other morphogenetic regulators. Hum. Genet. 126, 791–803 10.1007/s00439-009-0730-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Donner A.L. and Williams T. (2006) Frontal nasal prominence expression driven by Tcfap2a relies on a conserved binding site for STAT proteins. Dev. Dyn. 235, 1358–1370 10.1002/dvdy.20722 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Li W. and Cornell R.A. (2007) Redundant activities of Tfap2a and Tfap2c are required for neural crest induction and development of other non-neural ectoderm derivatives in zebrafish embryos. Dev. Biol. 304, 338–354 10.1016/j.ydbio.2006.12.042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Powell D.R., Hernandez-Lagunas L., LaMonica K. and Artinger K.B. (2013) Prdm1a directly activates foxd3 and tfap2a during zebrafish neural crest specification. Development 140, 3445–3455 10.1242/dev.096164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang J., Hagopian-Donaldson S., Serbedzija G., Elsemore J., Plehn-Dujowich D., McMahon A.P.. et al. (1996) Neural tube, skeletal and body wall defects in mice lacking transcription factor AP-2. Nature 381, 238–241 10.1038/381238a0 [DOI] [PubMed] [Google Scholar]
- 50.Schafer S., Anschlag J., Nettersheim D., Haas N., Pawig L. and Schorle H. (2011) The role of BLIMP1 and its putative downstream target TFAP2C in germ cell development and germ cell tumours. Int. J. Androl. 34, e152–e158, 10.1111/j.1365-2605.2011.01167.x [DOI] [PubMed] [Google Scholar]
- 51.Pastor W.A., Liu W., Chen D., Ho J., Kim R., Hunt T.J.. et al. (2018) TFAP2C regulates transcription in human naive pluripotency by opening enhancers. Nat. Cell Biol. 20, 553–564 10.1038/s41556-018-0089-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Zainolabidin N., Kamath S.P., Thanawalla A.R. and Chen A.I. (2017) Distinct activities of Tfap2A and Tfap2B in the specification of GABAergic interneurons in the developing cerebellum. Front. Mol. Neurosci. 10, 281 10.3389/fnmol.2017.00281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Bardakjian T.W.A. and Schneider A. (2004) Microphthalmia/Anophthalmia/Coloboma Spectrum, University of Washington, Seattle, Seattle (WA) [Google Scholar]
- 54.Kuckenberg P., Kubaczka C. and Schorle H. (2012) The role of transcription factor Tcfap2c/TFAP2C in trophectoderm development. Reprod. Biomed. Online 25, 12–20 10.1016/j.rbmo.2012.02.015 [DOI] [PubMed] [Google Scholar]
- 55.Woodfield G.W., Chen Y., Bair T.B., Domann F.E. and Weigel R.J. (2010) Identification of primary gene targets of TFAP2C in hormone responsive breast carcinoma cells. Genes Chromosomes Cancer 49, 948–962 10.1002/gcc.20807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Salah Z., Aqeilan R. and Huebner K. (2010) WWOX gene and gene product: tumor suppression through specific protein interactions. Future Oncol. 6, 249–259 10.2217/fon.09.152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Park J.M., Wu T., Cyr A.R., Woodfield G.W., De Andrade J.P., Spanheimer P.M.. et al. (2015) The role of Tcfap2c in tumorigenesis and cancer growth in an activated Neu model of mammary carcinogenesis. Oncogene 34, 6105–6114 10.1038/onc.2015.59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.De Andrade J.P., Park J.M., Gu V.W., Woodfield G.W., Kulak M.V., Lorenzen A.W.. et al. (2016) EGFR is regulated by TFAP2C in luminal breast cancer and is a target for vandetanib. Mol. Cancer Ther. 15, 503–511 10.1158/1535-7163.MCT-15-0548-T [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Scibetta A.G., Wong P.P., Chan K.V., Canosa M. and Hurst H.C. (2010) Dual association by TFAP2A during activation of the p21cip/CDKN1A promoter. Cell Cycle 9, 4525–4532 10.4161/cc.9.22.13746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zeng Y.X., Somasundaram K. and el-Deiry W.S. (1997) AP2 inhibits cancer cell growth and activates p21WAF1/CIP1 expression. Nat. Genet. 15, 78–82 10.1038/ng0197-78 [DOI] [PubMed] [Google Scholar]
- 61.Wong P.P., Miranda F., Chan K.V., Berlato C., Hurst H.C. and Scibetta A.G. (2012) Histone demethylase KDM5B collaborates with TFAP2C and Myc to repress the cell cycle inhibitor p21(cip) (CDKN1A). Mol. Cell. Biol. 32, 1633–1644 10.1128/MCB.06373-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Yuan J., Zhang N., Zheng Y., Chen Y.D., Liu J. and Yang M. (2018) LncRNA GAS5 indel genetic polymorphism contributes to glioma risk through interfering binding of transcriptional factor TFAP2A. DNA Cell Biol. 37, 750–757 10.1089/dna.2018.4215 [DOI] [PubMed] [Google Scholar]
- 63.Ma Y., Yang Y., Wang F., Moyer M.P., Wei Q., Zhang P.. et al. (2016) Long non-coding RNA CCAL regulates colorectal cancer progression by activating Wnt/beta-catenin signalling pathway via suppression of activator protein 2alpha. Gut 65, 1494–1504 10.1136/gutjnl-2014-308392 [DOI] [PubMed] [Google Scholar]
- 64.Liu Y., Yang Y., Wang T., Wang L., Wang X., Li T.. et al. (2018) Long non-coding RNA CCAL promotes hepatocellular carcinoma progression by regulating AP-2alpha and Wnt/beta-catenin pathway. Int. J. Biol. Macromol. 109, 424–434 10.1016/j.ijbiomac.2017.12.110 [DOI] [PubMed] [Google Scholar]
- 65.Barbagallo C., Brex D., Caponnetto A., Cirnigliaro M., Scalia M., Magnano A.. et al. (2018) LncRNA UCA1, upregulated in CRC biopsies and downregulated in serum exosomes, controls mRNA expression by RNA-RNA interactions. Mol. Ther. Nucleic Acids 12, 229–241 10.1016/j.omtn.2018.05.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Jain A.K., Xi Y., McCarthy R., Allton K., Akdemir K.C., Patel L.R.. et al. (2016) LncPRESS1 is a p53-regulated LncRNA that safeguards pluripotency by disrupting SIRT6-mediated de-acetylation of histone H3K56. Mol. Cell 64, 967–981 10.1016/j.molcel.2016.10.039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhang J., Yuan L., Zhang X., Hamblin M.H., Zhu T., Meng F.. et al. (2016) Altered long non-coding RNA transcriptomic profiles in brain microvascular endothelium after cerebral ischemia. Exp. Neurol. 277, 162–170 10.1016/j.expneurol.2015.12.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Penna E., Orso F., Cimino D., Tenaglia E., Lembo A., Quaglino E.. et al. (2011) microRNA-214 contributes to melanoma tumour progression through suppression of TFAP2C. EMBO J. 30, 1990–2007 10.1038/emboj.2011.102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Penna E., Orso F., Cimino D., Vercellino I., Grassi E., Quaglino E.. et al. (2013) miR-214 coordinates melanoma progression by upregulating ALCAM through TFAP2 and miR-148b downmodulation. Cancer Res. 73, 4098–4111 10.1158/0008-5472.CAN-12-3686 [DOI] [PubMed] [Google Scholar]
- 70.Gershenwald J.E., Sumner W., Calderone T., Wang Z., Huang S. and Bar-Eli M. (2001) Dominant-negative transcription factor AP-2 augments SB-2 melanoma tumor growth in vivo. Oncogene 20, 3363–3375 10.1038/sj.onc.1204450 [DOI] [PubMed] [Google Scholar]
- 71.Bhattacharya A., Schmitz U., Raatz Y., Schonherr M., Kottek T., Schauer M.. et al. (2015) miR-638 promotes melanoma metastasis and protects melanoma cells from apoptosis and autophagy. Oncotarget 6, 2966–2980 10.18632/oncotarget.3070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Jozkowicz A., Was H. and Dulak J. (2007) Heme oxygenase-1 in tumors: is it a false friend? Antioxid. Redox Signal. 9, 2099–2117 10.1089/ars.2007.1659 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li C.G., Pu M.F., Li C.Z., Gao M., Liu M.X., Yu C.Z.. et al. (2017) MicroRNA-1304 suppresses human non-small cell lung cancer cell growth in vitro by targeting heme oxygenase-1. Acta Pharmacol. Sin. 38, 110–119 10.1038/aps.2016.92 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Pu M., Li C., Qi X., Chen J., Wang Y., Gao L.. et al. (2017) MiR-1254 suppresses HO-1 expression through seed region-dependent silencing and non-seed interaction with TFAP2A transcript to attenuate NSCLC growth. PLoS Genet. 13, e1006896 10.1371/journal.pgen.1006896 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Kang J., Kim W., Lee S., Kwon D., Chun J., Son B.. et al. (2017) TFAP2C promotes lung tumorigenesis and aggressiveness through miR-183- and miR-33a-mediated cell cycle regulation. Oncogene 36, 1585–1596 10.1038/onc.2016.328 [DOI] [PubMed] [Google Scholar]
- 76.Chang T.H., Tsai M.F., Gow C.H., Wu S.G., Liu Y.N., Chang Y.L.. et al. (2017) Upregulation of microRNA-137 expression by Slug promotes tumor invasion and metastasis of non-small cell lung cancer cells through suppression of TFAP2C. Cancer Lett. 402, 190–202 10.1016/j.canlet.2017.06.002 [DOI] [PubMed] [Google Scholar]
- 77.Gao S.L., Wang L.Z., Liu H.Y., Liu D.L., Xie L.M. and Zhang Z.W. (2014) miR-200a inhibits tumor proliferation by targeting AP-2gamma in neuroblastoma cells. Asian Pac. J. Cancer Prev. 15, 4671–4676 10.7314/APJCP.2014.15.11.4671 [DOI] [PubMed] [Google Scholar]
- 78.Xiong G., Huang H., Feng M., Yang G., Zheng S., You L.. et al. (2018) MiR-10a-5p targets TFAP2C to promote gemcitabine resistance in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 37, 76 10.1186/s13046-018-0739-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Zhou J., Duan H., Xie Y., Ning Y., Zhang X., Hui N.. et al. (2016) MiR-193a-5p targets the coding region of AP-2alpha mRNA and induces cisplatin resistance in bladder cancers. J. Cancer 7, 1740–1746 10.7150/jca.15620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Zhang F., Chen K., Tao H., Kang T., Xiong Q., Zeng Q.. et al. (2018) miR-25-3p, positively regulated by transcription factor AP-2alpha, regulates the metabolism of C2C12 cells by targeting Akt1. Int. J. Mol. Sci. 19, E773 10.3390/ijms19030773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Wang J., Bai Y., Li H., Greene S.B., Klysik E., Yu W.. et al. (2013) MicroRNA-17-92, a direct Ap-2alpha transcriptional target, modulates T-box factor activity in orofacial clefting. PLoS Genet. 9, e1003785 10.1371/journal.pgen.1003785 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Fagerberg L., Hallstrom B.M., Oksvold P., Kampf C., Djureinovic D., Odeberg J.. et al. (2014) Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteomics 13, 397–406 10.1074/mcp.M113.035600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Li M., Naidu P., Yu Y., Berger N.A. and Kannan P. (2004) Dual regulation of AP-2alpha transcriptional activation by poly(ADP-ribose) polymerase-1. Biochem. J. 382, 323–329 10.1042/BJ20040593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Schneider H.A., Gembruch U., Fimmers R., Schmitz J. and Muller A.M. (2015) Expression of AP-2gamma in placentas of patients with preeclampsia and of smokers. Arch. Gynecol. Obstet. 291, 1015–1021 10.1007/s00404-014-3473-4 [DOI] [PubMed] [Google Scholar]
- 85.Woenckhaus C., Giebel J., Failing K., Fenic I., Dittberner T. and Poetsch M. (2003) Expression of AP-2alpha, c-kit, and cleaved caspase-6 and -3 in naevi and malignant melanomas of the skin. A possible role for caspases in melanoma progression? J. Pathol. 201, 278–287 10.1002/path.1424 [DOI] [PubMed] [Google Scholar]
- 86.Osella-Abate S., Novelli M., Quaglino P., Orso F., Ubezio B., Tomasini C.. et al. (2012) Expression of AP-2alpha, AP-2gamma and ESDN in primary melanomas: correlation with histopathological features and potential prognostic value. J. Dermatol. Sci. 68, 202–204 10.1016/j.jdermsci.2012.09.008 [DOI] [PubMed] [Google Scholar]
- 87.Ahrens T.D., Timme S., Hoeppner J., Ostendorp J., Hembach S., Follo M.. et al. (2015) Selective inhibition of esophageal cancer cells by combination of HDAC inhibitors and Azacytidine. Epigenetics 10, 431–445 10.1080/15592294.2015.1039216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Zhang Y., Xu Y., Li Z., Zhu Y., Wen S., Wang M.. et al. (2018) Identification of the key transcription factors in esophageal squamous cell carcinoma. J. Thorac. Dis. 10, 148–161 10.21037/jtd.2017.12.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Huang W., Chen C., Liang Z., Qiu J., Li X., Hu X.. et al. (2016) AP-2alpha inhibits hepatocellular carcinoma cell growth and migration. Int. J. Oncol. 48, 1125–1134 10.3892/ijo.2016.3318 [DOI] [PubMed] [Google Scholar]
- 90.Li Q. and Dashwood R.H. (2004) Activator protein 2alpha associates with adenomatous polyposis coli/beta-catenin and Inhibits beta-catenin/T-cell factor transcriptional activity in colorectal cancer cells. J. Biol. Chem. 279, 45669–45675 10.1074/jbc.M405025200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Wajapeyee N., Britto R., Ravishankar H.M. and Somasundaram K. (2006) Apoptosis induction by activator protein 2alpha involves transcriptional repression of Bcl-2. J. Biol. Chem. 281, 16207–16219 10.1074/jbc.M600539200 [DOI] [PubMed] [Google Scholar]
- 92.Stabach P.R., Thiyagarajan M.M., Woodfield G.W. and Weigel R.J. (2006) AP2alpha alters the transcriptional activity and stability of p53. Oncogene 25, 2148–2159 10.1038/sj.onc.1209250 [DOI] [PubMed] [Google Scholar]
- 93.Pellikainen J., Kataja V., Ropponen K., Kellokoski J., Pietilainen T., Bohm J.. et al. (2002) Reduced nuclear expression of transcription factor AP-2 associates with aggressive breast cancer. Clin. Cancer Res. 8, 3487–3495 [PubMed] [Google Scholar]
- 94.Sotiriou C., Wirapati P., Loi S., Harris A., Fox S., Smeds J.. et al. (2006) Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J. Natl. Cancer Inst. 98, 262–272 10.1093/jnci/djj052 [DOI] [PubMed] [Google Scholar]
- 95.Su W., Xia J., Chen X., Xu M., Nie L., Chen N.. et al. (2014) Ectopic expression of AP-2alpha transcription factor suppresses glioma progression. Int. J. Clin. Exp. Pathol. 7, 8666–8674 [PMC free article] [PubMed] [Google Scholar]
- 96.Hallberg A.R., Vorrink S.U., Hudachek D.R., Cramer-Morales K., Milhem M.M., Cornell R.A.. et al. (2014) Aberrant CpG methylation of the TFAP2A gene constitutes a mechanism for loss of TFAP2A expression in human metastatic melanoma. Epigenetics 9, 1641–1647 10.4161/15592294.2014.988062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Lian W., Zhang L., Yang L. and Chen W. (2017) AP-2alpha reverses vincristine-induced multidrug resistance of SGC7901 gastric cancer cells by inhibiting the Notch pathway. Apoptosis 22, 933–941 10.1007/s10495-017-1379-x [DOI] [PubMed] [Google Scholar]
- 98.Wang W., Lv L., Pan K., Zhang Y., Zhao J.J., Chen J.G.. et al. (2011) Reduced expression of transcription factor AP-2alpha is associated with gastric adenocarcinoma prognosis. PLoS ONE 6, e24897 10.1371/journal.pone.0024897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Makhov P.B., Golovine K.V., Kutikov A., Canter D.J., Rybko V.A., Roshchin D.A.. et al. (2011) Reversal of epigenetic silencing of AP-2alpha results in increased zinc uptake in DU-145 and LNCaP prostate cancer cells. Carcinogenesis 32, 1773–1781 10.1093/carcin/bgr212 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Schulte J.H., Kirfel J., Lim S., Schramm A., Friedrichs N., Deubzer H.E.. et al. (2008) Transcription factor AP2alpha (TFAP2a) regulates differentiation and proliferation of neuroblastoma cells. Cancer Lett. 271, 56–63 10.1016/j.canlet.2008.05.039 [DOI] [PubMed] [Google Scholar]
- 101.Carriere C., Mirocha S., Deharvengt S., Gunn J.R. and Korc M. (2011) Aberrant expressions of AP-2alpha splice variants in pancreatic cancer. Pancreas 40, 695–700 10.1097/MPA.0b013e31821f2715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Ding X., Yang Z., Zhou F., Wang F., Li X., Chen C.. et al. (2013) Transcription factor AP-2alpha regulates acute myeloid leukemia cell proliferation by influencing Hoxa gene expression. Int. J. Biochem. Cell Biol. 45, 1647–1656 10.1016/j.biocel.2013.04.024 [DOI] [PubMed] [Google Scholar]
- 103.Oyama N., Takahashi H., Tojo M., Iwatsuki K., Iizuka H., Nakamura K.. et al. (2002) Different properties of three isoforms (alpha, beta, and gamma) of transcription factor AP-2 in the expression of human keratinocyte genes. Arch. Dermatol. Res. 294, 273–280 10.1007/s00403-002-0327-x [DOI] [PubMed] [Google Scholar]
- 104.Bennett K.L., Romigh T. and Eng C. (2009) AP-2alpha induces epigenetic silencing of tumor suppressive genes and microsatellite instability in head and neck squamous cell carcinoma. PLoS ONE 4, e6931 10.1371/journal.pone.0006931 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Williams C.M., Scibetta A.G., Friedrich J.K., Canosa M., Berlato C., Moss C.H.. et al. (2009) AP-2gamma promotes proliferation in breast tumour cells by direct repression of the CDKN1A gene. EMBO J. 28, 3591–3601 10.1038/emboj.2009.290 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Lal G., Contreras P.G., Kulak M., Woodfield G., Bair T., Domann F.E.. et al. (2013) Human Melanoma cells over-express extracellular matrix 1 (ECM1) which is regulated by TFAP2C. PLoS ONE 8, e73953 10.1371/journal.pone.0073953 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Odegaard E., Staff A.C., Kaern J., Florenes V.A., Kopolovic J., Trope C.G.. et al. (2006) The AP-2gamma transcription factor is upregulated in advanced-stage ovarian carcinoma. Gynecol. Oncol. 100, 462–468 10.1016/j.ygyno.2005.09.022 [DOI] [PubMed] [Google Scholar]
- 108.Kim W., Kim E., Lee S., Kim D., Chun J., Park K.H.. et al. (2016) TFAP2C-mediated upregulation of TGFBR1 promotes lung tumorigenesis and epithelial-mesenchymal transition. Exp. Mol. Med. 48, e273 10.1038/emm.2016.125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Wu Y., Xiao Y., Ding X., Zhuo Y., Ren P., Zhou C.. et al. (2011) A miR-200b/200c/429-binding site polymorphism in the 3′ untranslated region of the AP-2alpha gene is associated with cisplatin resistance. PLoS ONE 6, e29043 10.1371/journal.pone.0029043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Wajapeyee N., Raut C.G. and Somasundaram K. (2005) Activator protein 2alpha status determines the chemosensitivity of cancer cells: implications in cancer chemotherapy. Cancer Res. 65, 8628–8634 10.1158/0008-5472.CAN-05-1059 [DOI] [PubMed] [Google Scholar]
- 111.Gee J.M., Eloranta J.J., Ibbitt J.C., Robertson J.F., Ellis I.O., Williams T.. et al. (2009) Overexpression of TFAP2C in invasive breast cancer correlates with a poorer response to anti-hormone therapy and reduced patient survival. J. Pathol. 217, 32–41 10.1002/path.2430 [DOI] [PubMed] [Google Scholar]