Abstract
Ribonucleotide reductase (RNR) supplies the balanced pools of deoxynucleotide triphosphates (dNTPs) necessary for DNA replication and maintenance of genomic integrity. RNR is subject to allosteric regulatory mechanisms in all eukaryotes, as well as to control by small protein inhibitors Sml1p and Spd1p in budding and fission yeast, respectively. Here, we show that the metazoan protein IRBIT forms a deoxyadenosine triphosphate (dATP)–dependent complex with RNR, which stabilizes dATP in the activity site of RNR and thus inhibits the enzyme. Formation of the RNR-IRBIT complex is regulated through phosphorylation of IRBIT, and ablation of IRBIT expression in HeLa cells causes imbalanced dNTP pools and altered cell cycle progression. We demonstrate a mechanism for RNR regulation in higher eukaryotes that acts by enhancing allosteric RNR inhibition by dATP.
Ribonucleotide reductase (RNR) provides building blocks for genomic and mitochondrial DNA replication and repair, and uncontrolled RNR activity has been associated with malignant transformation and tumor cell growth. The key role of RNR in DNA synthesis has made it a target for both anticancer and antiviral therapy (1). Many chemotherapeutic RNR inhibitors are nucleoside analogs whose association with other nucleotide-binding proteins can contribute to their toxicity, and so the search continues for more specific drugs with less adverse effects (2).
IRBIT is a conserved metazoan protein that has been implicated in diverse functions (3–6). IRBIT consists of a putative enzymatic domain that has similarity to S-adenosylhomocysteine hydrolase and an essential N-terminal domain of 104 amino acids (3–5). To identify proteins that bind IRBIT, we performed immunoprecipitation from lysates of HeLa cells, followed by SDS–polyacrylamide gel electrophoresis (SDS-PAGE) and protein staining. Prominent bands were excised and analyzed by mass spectrometry. RNR’s large subunit R1 was identified as one of the most abundant proteins in the sample (Fig. 1A and table S1). We expressed RNR (R1/R2B) and IRBIT in insect cells and purified these proteins. It should be noted that only R2B (p53R2), but not R2, formed a stable complex with R1. We analyzed RNR-IRBIT interaction in the absence or presence of RNR nucleotide effectors (fig. S1, A and B). In the absence of nucleotides, IRBIT interacted with RNR directly but with low affinity, whereas the formation of the RNR-IRBIT complex was greatly stimulated in the presence of deoxyadenosine triphosphate (dATP) (fig. S1, B and C).
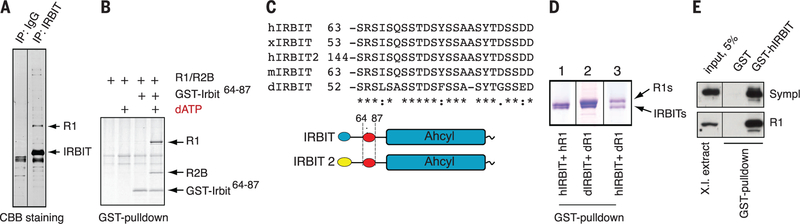
Fig. 1. IRBIT interacts with RNR in a dATP-dependent manner.
(A) IRBIT-RNR interaction. IRBIT was immunoprecipitated from asynchronous HeLa cell lysates; bound proteins were resolved on SDS-PAGE and stained with Coomassie brilliant blue (CBB). IRBIT and R1 proteins are indicated by arrows. (B) Minimal domain required for dATP-dependent IRBIT-RNR interaction. GST-IRBIT64−87 was mixed with R1/R2B and 3 mM dATP. Complexes were retrieved by glutathione-Sepharose and resolved by SDS-PAGE and stained with CBB. (C) Alignment of IRBIT domain from different metazoans. (D) R1-IRBIT interaction is evolutionarily conserved. GST-IRBIT (hIRBIT) or GST-CG9977 (dIRBIT) were mixed with either human 6His-R1 (hR1) or Drosophila 6His-R1 (dR1) along with 3 mM dATP and analyzed as in (B). (E) hIRBIT binds Xenopus R1. GST-hIRBIT was mixed with cytostatic factor–arrested Xenopus laevis egg extracts. IRBIT-bound proteins were purified and probed with antibodies against R1 and Symplekin (4).
The region of IRBIT essential for RNR interac- tion lies within amino acids 64 to 87 (fig. S1D). This fragment was sufficient for complex formation with R1/R2B in the presence of dATP (Fig. 1B). This well-conserved serine- and threonine-rich domain exists in all IRBIT-like molecules (Fig. 1C). Human IRBIT interacted strongly with Drosophila R1, whereas Drosophila IRBIT (dIRBIT) (CG9977) interacted with human RNR in a dATP-dependent manner (Fig. 1D). Human IRBIT also showed specific recruitment of Xenopus R1 from egg extracts (Fig. 1E). RNR bound IRBIT in the presence of ~1 to 3 mM dATP (fig. S1E), which is below physiological dATP levels (10 to 40 mM) (7).
dATP binds R1 at two sites: dATP bound to the low-affinity activity site (A-site) inhibits RNR, whereas dATP bound to the specificity site (S-site) promotes production of deoxyuridine diphosphate (dUDP) and deoxycytidine diphosphate (dCDP) (8). We analyzed interaction between IRBIT and a mutant that does not differentiate between dATP and ATP in the A-site [R1D57N, in which asparagine (N) replaces aspartic acid (D) at residue 57] (9). R1D57N did not stably interact with IRBIT in the presence of dATP (Fig. 2A), which indicated that the A-site is occupied by dATP when R1 is bound to IRBIT. Consistent with this idea, the RNR-dATP-IRBIT complex was not disrupted by deoxythymidine triphosphate (dTTP) or deoxyguanosine triphosphate (dGTP), which can compete only for S-site occupancy, but it was sensitive to ATP (Fig. 2B). In addition, glutathione S-transferase fused with human IRBIT (GST-hIRBIT) could bind preformed R1-dATP-dTTP complex (fig. S1F).
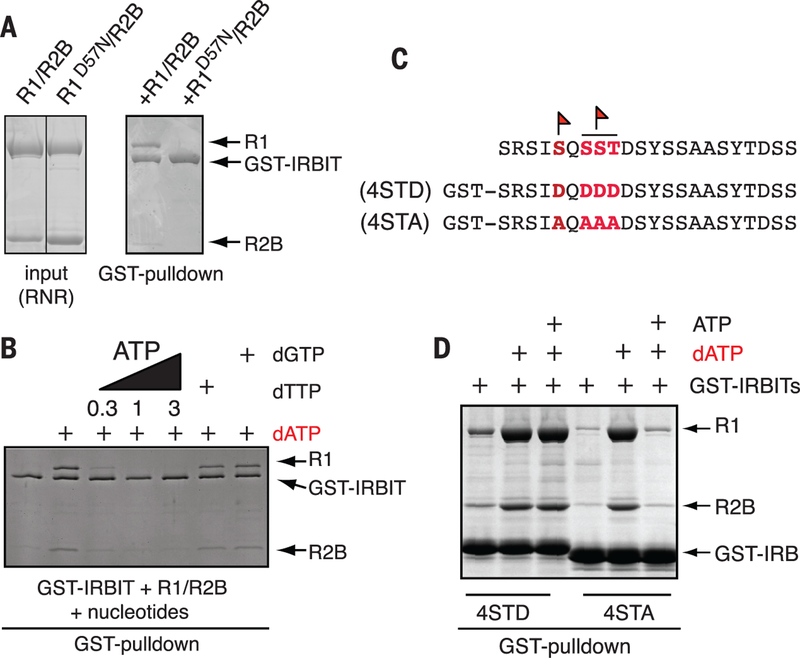
Fig. 2. IRBIT binds RNR when dATP is in the A-site and requires phosphorylation for its full binding activity.
(A) dATP occupancy of RNR’s A-site is required for IRBIT binding. GST-IRBIT was mixed with either R1/R2B or R1D57N/R2B (input, left) and 10 mM dATP and retrieved by glutathione-Sepharose (right). (B) IRBIT-RNR interaction is sensitive to ATP competition but insensitive to S-site occupancy. GST-IRBIT was mixed with R1/R2B and 3 mM dATP followed by addition of 0.3 mM ATP, 1 mM ATP, 3 mM ATP, 1 mM dTTP, or 1 mM dGTP. (C) Endogenous phosphorylation sites that were mapped on IRBIT (flags) and corresponding recombinant phosphomimetic or nonphosphorylatable mutants. (D) IRBIT phosphorylation prevents dissociation of the IRBIT-R1-dATP complex by ATP. GST-IRBIT4STD (STD) or GST-IRBIT4STA (STA) was mixed with R1/R2B and 3 mM dATP followed by addition of 1 mM ATP. In all panels, complexes were analyzed as in Fig. 1B.
Physiological levels of adenosine triphosphate (ATP) (1 to 3 mM) disrupted the IRBIT-RNR-dATP complex, which indicated that RNR bound with IRBIT was still subject to competition between ATP and dATP for A-site occupancy (Fig. 2B) (10). We suspected that some posttranslational modifications of IRBIT might increase its inhibitory potential. We analyzed the phosphorylation status of endogenous IRBIT by mass spectrometry and found that there are at least two phosphorylation sites. Both of them were mapped to the IRBIT64−87 domain: Ser68 and an undetermined residue within a SerSerThr motif (residues 70 to 72) (Fig. 2C). We compared complex formation between R1/R2B and either a phosphomimetic (4STD) or a nonphosphorylatable IRBIT domain (4STA) (Fig. 2C). Both domains bound to RNR in a dATP-dependent manner. However, high concentrations of ATP disrupted only 4STA-RNR-dATP complex but not 4STD-RNR-dATP complex, which indicated that IRBIT phosphorylation at these residues stabilizes dATP on R1 (Fig. 2D).
We hypothesized that IRBIT may recognize inactive, dATP-bound R1 and stabilize the enzyme in this state. To test this idea, we performed binding studies in the presence of IRBIT using [α−32P]dATP and R1/R2B saturated with dTTP in the S-site. The association constant of dATP binding to the A-site of R1 (kon) was virtually unchanged in the absence or in the presence of IRBIT (Fig. 3C and figs. S2D and S3A), and the A-site on R1 was saturated at ~3 mM of dATP (Fig. 3A). The dissociation rate of R1-dATP (koff) was substantially inhibited in the presence of IRBIT (Fig. 3, B and C, and fig. S3), which indicated that IRBIT stabilizes dATP in the A-site of R1 (11). IRBIT did not substantially alter dTTP binding to the S-site whether dATP was present in the reaction or not (fig. S2C). The fragment of IRBIT that binds R1 (GST-IRBIT64−87) was sufficient in itself to increase A-site affinity for dATP (fig. S2A).
Fig. 3. IRBIT stabilizes and inhibits RNR-dATP complex.
(A) IRBIT does not create additional dATP binding sites on RNR. RNR was mixed with increasing doses of [32P]dATP in the absence (circles) or presence (squares) of hIRBIT and in the presence of dTTP (1 mM) at +4°C. (B) IRBIT inhibits dATP release from RNR. Values are means T SEM. (C) Rate constants of R1-dATP interaction. Bmax (maximum number of binding sites) was derived from (A), koff was obtained from (B), and kon was calculated from fig. S2D. Kd, dissociation constant. (D) IRBIT enhances dATP inhibition of RNR. RNR was premixed with or without dATP and with or without 6His-IRBIT and tested for uridine diphosphate (UDP) reductase activity.
To measure the consequences of this binding for RNR function, we analyzed RNR activity in vitro. IRBIT did not affect RNR activity in the absence of dATP (Fig. 3D). Preformed RNR-dATP complexes exhibited 20% less activity than RNR alone. As expected, RNR in the RNR-dATP-IRBIT complex was substantially inhibited (Fig. 3D and fig. S3E).
To determine the consequences of RNR inhibition by IRBIT, we measured deoxynucleotide triphosphate (dNTP) concentration in HeLa cells that express Tet-inducible short hairpin RNA against IRBIT. IRBIT-depleted asynchronous HeLa cells showed an imbalanced pool of deoxynucleotides (Fig. 4A). These changes in dNTP levels were not due to the altered expression levels of the RNR subunits (Fig. 4B). Note that the sensitivity of dNTP levels to IRBIT depletion was more pronounced in mitosis but was minimal in G1 cells (Fig. 4A), and coprecipitation experiments showed that IRBIT interacted with R1 more strongly during mitosis than during G1 phase (Fig. 4C).
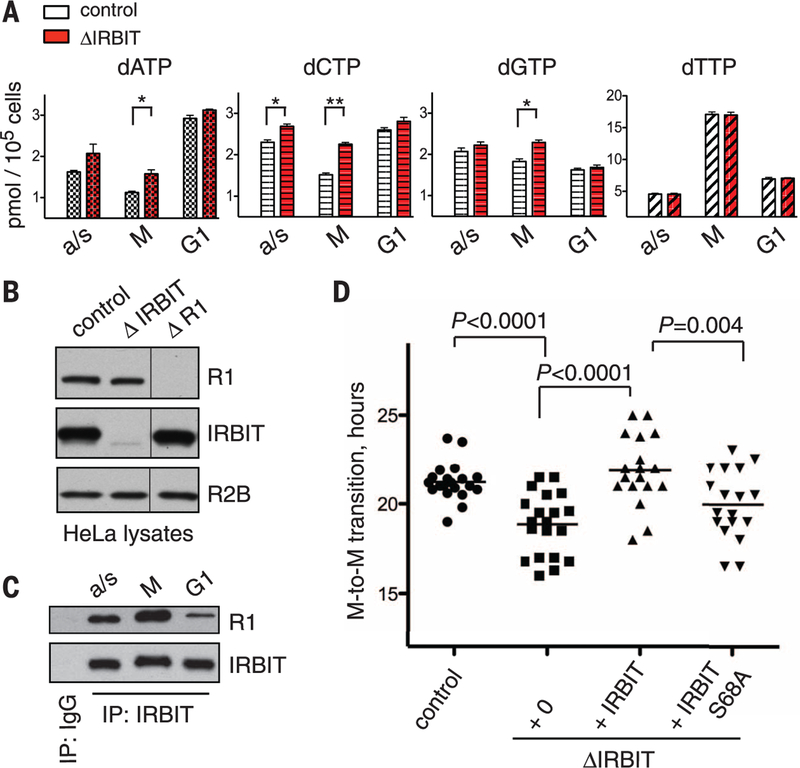
Fig. 4. IRBIT controls the endogenous dNTP pool and is required for normal cell cycle progression.
(A) dNTP levels in HeLa and in HeLaDIRBIT cells in mitotic (M) or G1 phases; a/s for asynchronous population. (B) Stability of R1 and IRBIT do not depend on each other. Total lysates of HeLa, HeLaDIRBIT, and HeLaDR1 cells were analyzed by Western blots using antibodies against IRBIT, R1, and R2B. (C) IRBIT-RNR interaction during the cell cycle. IRBIT was immunoprecipitated from corresponding lysates and analyzed for the presence of R1. (D) Cell cycle progression of HeLa cells with or without IRBIT. M-to-M transition indicates timing between anaphase and prometaphase of the next cell cycle of an individual daughter cell (n = 20). Values are means T SEM. Statistical differences were evaluated by Student’s t test (**P < 0.01).
We analyzed cell cycle progression in IRBIT-depleted cells through live imaging of HeLa cells expressing histone H2B fused with green fluorescent protein (H2B-GFP). Most of the control HeLa cells underwent mitosis every 22 hours (Fig. 4D). In contrast, depletion of IRBIT resulted in a much greater variation in the duration of interphase between individual cells. Reintroduction of wild- type IRBIT to the depleted cells rescued this phenotype, whereas IRBITS68A did not (Fig. 4D), which indicated that IRBIT function depends on phosphorylation of this residue, as suggested by our in vitro results.
The N-terminal domain of IRBIT belongs to the class of IDP (intrinsically disordered protein or peptide) (12). Budding yeast Sml1p is an IDP that binds R1 (13). We noticed that there is some similarity between the IRBIT64−87 domain and the central region of Sml1p (amino acids 46 to 72), which was not previously implicated in interaction with RNR (fig. S4) (14). Sml1p46−72, like IRBIT64−87, interacted with RNR in a dATP-dependent manner. We speculate that Sml1p may use this domain to recognize dATP-bound R1 ina manner that may be analogous to that between Sml1 and IRBIT (fig. S8).
Altogether, our results demonstrate that IRBIT interacts with RNR in a dATP-dependent manner and stabilizes dATP in the RNR A-site, potentially by stabilizing oligomeric form(s) of R1 formed in the presence of dATP (15–17) (fig. S1G). Because binding of dATP to the A-site is inhibitory to RNR activity, IRBIT inhibits RNR. Under normal physiological conditions, where ATP levels are high, such inhibition could only be achieved when IRBIT’s binding is strengthened by phosphorylation. This mechanism is likely to be critical, because cells depleted of IRBIT show substantial alteration in their cell cycle progression and because IRBIT is indispensible for embryogenesis (18). It is also possible that IRBIT-RNR acts as a part of multimeric complexes (fig. S5).
In a larger context, we note that balanced control of deoxynucleotide levels is central to maintaining the genome in an accurate fashion (figs. S6 and S7) (19). Modulation of IRBIT binding offers a fundamentally different mechanism for RNR inhibition that may circumvent toxicity issues of current RNR drugs, and it thus offers a promising target for future drug development.
Supplementary Material
ACKNOWLEDGMENTS
We are grateful to B. Cooperman and L. Thelander for reagents and for the help with RNR assay. We are indebted to M. F. Ahmad, I. Arnaoutova, P. Kalab, D. Mukhopadhyay, and C. Dealwis for their help and for the critical reading of the manuscript.
REFERENCES AND NOTES
- 1.Shao J, Zhou B, Chu B, Yen Y, Curr. Cancer Drug Targets 6, 409–431 (2006). [DOI] [PubMed] [Google Scholar]
- 2.Wijerathna SR et al. , Pharmaceuticals 4, 1328–1354 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ando H et al. , Mol. Cell 22, 795–806 (2006). [DOI] [PubMed] [Google Scholar]
- 4.Kiefer H et al. , J. Biol. Chem 284, 10694–10705 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yang D et al. , J. Clin. Invest 119, 193–202 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wash R et al. , J. Gen. Virol 93, 2118–2130 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Traut TW, Mol. Cell. Biochem 140, 1–22 (1994). [DOI] [PubMed] [Google Scholar]
- 8.Jordan A, Reichard P, Annu. Rev. Biochem 67, 71–98 (1998). [DOI] [PubMed] [Google Scholar]
- 9.Reichard P, Eliasson R, Ingemarson R, Thelander L, J. Biol. Chem 275, 33021–33026 (2000). [DOI] [PubMed] [Google Scholar]
- 10.Ormö M, Sjöberg BM, Anal. Biochem 189, 138–141 (1990). [DOI] [PubMed] [Google Scholar]
- 11.Kashlan OB, Scott CP, Lear JD, Cooperman BS, Biochemistry 41, 462–474 (2002). [DOI] [PubMed] [Google Scholar]
- 12.Tompa P, Trends Biochem. Sci 27, 527–533 (2002). [DOI] [PubMed] [Google Scholar]
- 13.Danielsson J et al. , Biochemistry 47, 13428–13437 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chabes A, Domkin V, Thelander L, J. Biol. Chem 274, 36679–36683 (1999). [DOI] [PubMed] [Google Scholar]
- 15.Ahmad MF, Dealwis CG, Prog. Mol. Biol. Transl. Sci 117, 389–410 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kashlan OB, Cooperman BS, Biochemistry 42, 1696–1706 (2003). [DOI] [PubMed] [Google Scholar]
- 17.Fairman JW et al. , Nat. Struct. Mol. Biol 18, 316–322 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cooper BJ et al. , J. Biol. Chem 281, 22471–22484 (2006). [DOI] [PubMed] [Google Scholar]
- 19.Ahluwalia D, Schaaper RM, Proc. Natl. Acad. Sci. U.S.A 110, 18596–18601 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.