Abstract
Immune checkpoint inhibitor therapy has changed clinical practice for patients with different cancers, since these agents have demonstrated a significant improvement of overall survival and are effective in many patients. However, an intrinsic or acquired resistance frequently occur and biomarkers predictive of responsiveness should help in patient selection and in defining the adequate treatment options. A deep analysis of the complexity of the tumor microenvironment is likely to further advance the field and hopefully identify more effective combined immunotherapeutic strategies. Here we review the current knowledge on tumor microenvironment, focusing on T cells, cancer associated fibroblasts and extracellular matrix. The use of 3D cell culture models to resemble tumor microenvironment landscape and to screen immunomodulatory drugs is also reviewed.
Keywords: Tumor microenvironment, Immune oncology, 3D culture models, T cells, Cancer associated fibroblasts, Extracellular matrix
Background
The use in the clinical practice of antibody-based immunotherapy, named immune checkpoint blockade (ICB), is based on the inhibition of receptors and/or ligands of Cytotoxic T-Lymphocyte Antigen Protein 4 (CTLA4) and Programmed cell death 1 (PD-1) axes. These reagents are at the forefront of immunotherapy of a wide array of cancer, previously endowed with poor prognosis [1]. However, not all patients benefit from the cure and some of them become refractory after the initial treatment response [2]. Thus, there is an urgent need to identify biomarkers of response and mechanisms of resistance to overcome the treatment failure occurring in a significant proportion of patients. The knowledge to date gathered by tumor patients treated with these drugs have indicated that a deep analysis of the tumor immune microenvironment (TME) may predict and guide response to ICB [3], again indicating that an improved understanding of the TME is crucial to improve cancer treatment. The availability of 3D experimental models able to recreate the complexity of the TME has substantially contributed to our understanding of tumor biology and has allowed more reliable studies on the effects of anti-tumor drugs. However, advancement in this field remains central for the development of new therapeutic strategies in the immune oncology era, as we have reviewed in this paper.
Tumor microenvironment (TME) and tumor immune microenvironment (TIME) in antitumor immune response and resistance to immunotherapy
Tumor development and progression relies on the dialogue among tumor cells, neighbouring stromal and immune cells, the extracellular matrix and soluble cues [4]. A deeper understanding of how cellular and molecular interactions within the TME shape tumor biology and, in turn, clinical outcome, is of tremendous importance in the new era of immune oncology.
ICB therapies targeting inhibitory receptors on T cells, such as CTLA4 and PD-1, are now approved for a broad range of tumor types, and long-term durable responses in a subset of patients represent an exceptional success in clinical oncology [5, 6]. Despite the unprecedented durable response rate observed, the majority of patients do not benefit from the treatment (primary resistance) and some others relapse after a period of response (acquired resistance) [7], indicating the urgent need to identify signatures of response to guide novel therapeutic combination overcoming ICB resistance.
Thanks to datasets and studies relative to the quantity, quality and spatial distribution of immune cells in the TME, it has been proposed that subclasses of TIME may predict and guide efficient immunotherapeutic treatments [3]. Three different immune profiles associated with responsiveness to ICB have been defined [8]. The immune-inflamed profile is characterized by the presence in the tumor core of cytotoxic T lymphocytes (CTL) which express the PD-1 molecule along with PD-L1 positive tumor cells. These inflamed ‘hot’ tumors often respond to anti-PD-1 and PD-L1 therapy. A further subclass of immune-inflamed TIME is characterized by the presence of tertiary lymphoid structures (TLSs), transient lymphoid aggregates developing at the sites of chronic inflammation, which have been correlated with clinical outcome and sensitivity to immunotherapies [9]. Notably, TLSs were found in the regression bed of neoadjuvant anti-PD-1 treated, resectable non-small cell lung cancer (NSCLC) patients [10], and their induction has been reported to enhance immunotherapy efficacy in resistant tumors [11]. Thus suggesting that the induction and manipulation of cancer associated TLSs should open new perspectives to design novel effective combination therapies [12]. The second profile is the immune-excluded profile that shows immune cells retained in the stroma surrounding tumor nests, due to their inability to penetrate the tumor bed and those tumors belong to patients with a low beneficial clinical response. The third profile, the immune-desert phenotype, is characterized by the presence of a non-inflamed TME with few or no CD8 T cells. These are the tumors more resistant to ICB [8].
Different cell populations, such as myeloid-derived suppressor cells (MDSCs), the M2 subtype of tumor-associated macrophages (TAMs), regulatory T cells (Treg cells) and cancer-associated fibroblasts (CAFs) may contribute to an immunosuppressive TME leading to ICB resistance. In accordance, different studies report that targeting and reprogramming these suppressive cells may revert this microenvironment leading to an enhanced response to immune therapy, as shown in murine and human settings. Indeed, pharmacologic targeting of the gamma isoform of phosphoinositide 3-kinase (PI3Kγ), highly expressed in myeloid cells, modulates their suppressive phenotype towards a more inflammatory phenotype and restores sensitivity to ICB. This attributed to the reshaping the TME leading to cytotoxic-T-cell-mediated tumor regression in mouse models [13]. Furthermore, the inhibition of colony-stimulating factor 1 (CSF1)/CSF1 receptor (CSF1R) signaling can functionally block tumor-infiltrating MDSCs enhancing anti-tumor T cell responses and sensitizes IDO-expressing tumors to ICB in various tumor models [14]. CSF1/CSF1R signaling also promotes a TAM immunosuppressive and pro-tumorigenic phenotype associated with a M2-like phenotype [15].
A recent paper from Peranzoni et al., reports that in human and murine tumors, CD8+ T cells poorly migrate and invade tumor nests due to their long-lasting interaction with tumor-associated macrophages in the stroma. Again, the depletion of TAMs with a CSF-1R inhibitor, restored CD8 T cell migration and infiltration into tumor islets and improved the efficacy of anti–PD-1 immunotherapies [16].
CAFs are the major component of the tumor stroma and exert profound effects on immune cells, mainly by altering the biochemical and biophysical properties of the stroma surrounding tumor cells, as detailed further in this review.
This complex landscape determines intrinsic metabolic features which, contributing to an immunosuppressive TME, may lead to resistance to immunotherapy.
Tumor hypoxia predicts poor outcome across all cancers [17], and is responsible for recruitment, polarization, and expansion of immune-suppressive stromal cell populations [18]. The cross-talk between hypoxia and immune-escape mechanisms is an emerging aspect in tumor progression and drug resistance as indicated by the enrichment of hypoxia related genes in signatures correlated with resistance to PD-1 [19]. Increased hypoxia has been associated to the release of different immunosuppressive molecules that recruit and activate multiple myeloid and lymphoid immune suppressor cells [20]. In accordance, hypoxia-targeted therapy has been reported to sensitize even the most therapeutically resistant preclinical models of prostate cancer to ICB, by reverting the highly suppressive ratio of MDSCs to CD8+ T cells present in untreated tumors and allowing T cells to infiltrate and survive in formerly hypoxic areas [21].
The mutual metabolic requirements of immune cells and tumor cells contribute to the immunosuppressive character of the TME and metabolic re-education of tumor cells could overcome metabolic immunosuppression favoring the efficacy of immunotherapy treatment [22]. An emerging pathway involved in an immunosuppressive TME is related to the production of extracellular adenosine by the ecto-enzyme CD73 [23]. CD73 elevated activity is found in many cancers and its blockade has been shown to significantly enhance the therapeutic activity of anti-PD-1 and anti-CTLA-4 monoclonal antibodies [24]. Cyclooxigenase (COX) enzymes are responsible for the synthesis of prostaglandins, with COX-2 able to induce high levels of prostaglandin E2 (PGE2), a potent immunosuppressive molecule, in a subset of cancers. Zelenay and colleagues showed that combination of cyclooxygenase-1 (COX-1) and COX-2 inhibitors with ICB can result in melanoma eradication [25].
All these results clearly demonstrate the need of a deeper knowledge of TME in terms of cellular and non cellular stromal compartments.
Cellular and non cellular stromal compartment in TME
T cells
T cells are the major players in antitumor immune response and their spatial distribution in the tumor bed and/or in the surrounding stroma strongly impact prognosis and response to therapy. In the new era of immune oncology, a great advance in the study of the immune cell subpopulations, quantification and spatial distribution has been made. The quality of immunohistochemical characterization has been greatly improved by digital pathology [26] and by the development of advanced technologies such as multiplex immunohistochemistry methods, which allow the identification of multiple biological markers in a single tissue section [27], and mass cytometry (CyTOF), an appealing platform for comprehensive phenotyping of cells in human tissues [28].
Starting from the seminal paper of Galon [29] many reports have demonstrated that solid tumors may be classified on the basis of the T cell infiltrate; intratumoral localization of T cell leads to a high “immunoscore”, which correlates with improved patient prognosis [26]. On the other hand, T cell infiltration edits the tumor during metastatic progression as previously suggested in the cancer immunoediting paradigm [30]. Angelova and Co-authors recently proposed that the tumor evolution during the metastatic process depends on the strength and quality of the local immune response at the metastatic site [31]. However, T cells may reside outside the tumor islets [32, 33], as we have observed in breast cancer where the lesions displaying undetectable HLA-A2 expression, showed peritumoral CD3+ T-cell localization compared to HLA-A2-positive tumors showing intratumoral lymphocyte localization [34]. Of relevance, tumor infiltrating lymphocytes were found in the regression bed of neo-adjuvant anti-PD-1 treated resectable NSCLC patients [10], whereas the inability of T cells to enter in the tumor bed, has been indicated as a mechanism of resistance to cancer immunotherapy [35].
T cell exclusion from the tumor site could be driven by signaling pathways related to tumor cells (intrinsic pathways) or stromal components (extrinsic pathways). The paradigm of tumor intrinsic pathways related to T cell absence into the TME is represented by the WNT/β-catenin pathway, which prevents the expression of C-C Motif Chemokine Ligand 4 (CCL4), a chemokine essential for DC and T cell recruitment [36]. Another relevant pathway related to T cell exclusion is the tyrosine kinase receptor AXL signaling pathway, strictly associated with the process of epithelial-mesenchymal transition (EMT). AXL has been identified as a mediator of immunosuppression given its role in suppressing antigen presentation and producing cytokines and chemokines supporting myeloid cell infiltrate, hampering the anti-tumor adaptive immune response [37]. In accordance, AXL levels were significantly correlated with resistance to PD-1 immunotherapy [19, 37].
A recent computational framework has been developed on the basis of Tumor Immune Dysfunction and Exclusion (TIDE), to identify factors related to the main mechanisms of tumor immune escape that could serve as a reliable surrogate biomarker to predict ICB response [38]. Moreover, by single-cell RNA sequencing (scRNAseq) of melanoma tumors, a signature associated with T cell exclusion and immune evasion has been reported as able to predict clinical responses to anti-PD-1 therapy [39].
CAF in immunoediting and ICB response
Tumor extrinsic pathways responsible of T cell exclusion from the tumor site are sustained by stromal cells that may limit T cell trafficking within the TME by different mechanisms, including the secretion of soluble factors [40].
Fibroblasts resident in tissues become activated as a consequence of various stimuli in the TME with TGFβ being the major player [41, 42] and the cancer activated fibroblasts (CAFs) are important regulators of the anti-tumor immune response [43]. Besides tissue resident fibroblasts, CAFs can also develop from mesenchymal stem cells or stellate cells, thus increasing the heterogeneity that accounts for the distinct functional subsets of these cells [44]. Of note, in breast cancer different subsets of CAFs have been associated with different immunosuppressive properties [45]. Activated CAFs produce and secrete a plethora of growth factors, chemokines and components of ECM, including collagens, fibronectin and laminins and ECM remodeling enzymes (for review see: [46]). This has a profound impact on the biochemical and biophysical properties of the stroma surrounding tumor cells, modulating the behavior of tumor cells and of the other components of TME including immune cells, with profound effects on the tumor immune contexture. Within the TME, CAFs can promote the recruitment of monocytes and their differentiation in M2 immunosuppressive macrophages via the secretion of interleukin-6 (IL-6) and Granulocyte-Macrophage Colony-Stimulating Factor (GM-CSF) [47], or in MDSC via Signal transducer and activator of transcription 3 (STAT3) activation by secreting IL-6, CCL2 (C-C Motif Chemokine Ligand 2), C-X-C Motif Chemokine Ligand 12 (CXCL12) [48]. CAFs can also promote the survival, activation, and function of neutrophils through an IL6-STAT3-PDL1 signaling cascade, impairing T-cell function through the PD1/PDL1 signaling pathway as reported in hepatocellular carcinoma (HCC) [49, 50].
CAFs are not only activated and sustained by TGFβ signalling [51], but are also the major producers of TGFβ in the TME. TGFβ has been recognized as pleiotropic regulator of immune response and a potent immunosuppressor in the TME. Inhibition of TGF-β signaling increases T cell accumulation and function in tumors [52] (For Review see [53]). Recently, stromal TGFβ has been considered as a relevant determinant of tumor responsiveness to anti-PDL1 treatment and its signaling inhibition potentiates the therapeutic effect of an anti-PDL1 blocking antibody [54]. Moreover, Mariathasan et al. in urothelial cancer have identified fibroblast-derived TGF-β signaling as a determinant of CD8+ T cell exclusion from the tumor parenchyma and localization in the fibroblast- and collagen-rich peritumoral stroma. The Authors suggest that TGFβ shapes the tumour microenvironment to restrain anti-tumour immunity by restricting T-cell infiltration. These effects have been correlated with the lack of response to ICB [55].
The recognized relevance of CAFs in the immunosuppressive TME has opened new perspectives in the identification of CAF subtypes as biomarkers of therapeutic resistance and their immunomodulatory pathways as druggable targets.
ECM in immune contexture and T cell exclusion
Cells to survive have to be anchored to extracellular matrix (ECM), a dynamic web of molecules, which provides structural support and biomechanical cues, and is fundamental in differentiation, tissue development, tissue architecture and homeostasis [56]. It has been recently recognized that the mechanical properties of the ECM are important modulators of cell behaviour, that are integrated with biochemical cues from the microenvironment to regulate tumor progression and metastatic dissemination [57, 58], also affecting the immune evasion [59]. Tumor cells reside in a stiffer environment compared to normal tissue [60] and this is mainly due to changes in ECM deposition and remodelling. Components of the ECM such as fibronectin, collagens, tenascins and laminins are secreted by both tumor and stromal cells and are organized and remodelled by a plethora of other proteins that align, cross-link, integrate or digest the deposited fibers by a complex network of signals to generate an extracellular matrix that is typical of and characterizes each tumor. Cells sense the physical properties of ECM and propagate the mechanical signals into alteration of cytoskeletal dynamics [61]. In turn, actin cytoskeleton dynamics act as platforms for gene regulation and key signaling transduction pathways involved in the cross-talk between tumor cells and TME and our group has recently demonstrated that the splicing of the actin regulator hMENA generates two alternatively expressed isoforms hMENA11a and hMENAΔv6 respectively inhibiting or inducing the secretion of several key extracellular matrix (ECM) proteins [62], modulating the ECM composition. Moreover, the actin-myosin contractility, generated by ECM stimulation, counteracts the forces transferred from ECM and further increases matrix stiffness. Yes-associated protein 1 (YAP) and WW domain containing transcription regulator 1 (TAZ) are mechanosensitive transcription factors that translocate to the nucleus in response to elevated matrix stiffness [63]. YAP function is critical for the establishment and maintenance of CAFs, which in turn, rearrange the ECM to increase tumor stiffness. YAP is activated by microenvironmental factors such as TGFβ and matrix stiffness and in turn it is required for the expression of genes regulating matrix stiffness and many pro-tumorigenic properties of fibroblasts [64]. YAP inhibition disrupts tumor-stroma interaction and suppresses pancreatic cancer progression [65] whereas YAP activation induces the expression of cytokines which recruits immunesuppressive leukocytes such as MDSCs and TAMs [66], suggesting that YAP acts as a transcriptional driver that orchestrates the immunesuppressive microenvironment within pancreatic ductal adenocarcinoma (PDAC). Tumor cell contact with rigid ECM components induces the activation of focal adhesion kinase FAK1 [67] and inhibiting FAK1 or FAK2 reduces cytokine production, the frequencies of CAFs, suppressive myeloid subsets, and CD4 + Foxp3+ Tregs, as well as ECM accumulation. Notably, FAK inhibition halts tumor growth and increases survival in a PDA mouse model, and anti-tumor activity can be further improved if combined with chemotherapy or anti-PD-1 [67].
Density and organization of ECM components also influence immune cell migration. Dynamic imaging of cell-ECM interactions showed that T-cell migration is independent by their proteolityc activity and is driven by their ability to vigorous shape change, crawling along collagen fibrils and squeezing through pre-existing matrix pores [68]. Using an ex vivo assay to track CD8 T cells in fresh human ovarian and lung cancer tissues, it has been shown that CD8 T cells accumulate and move slowly in the stroma, while the tumor islets are sites of less populated but faster T cells migration [69]. Bougherara et al., have also revealed that collagen fibers, by their orientation, spacing and density, control the distribution and migration of resident CD8 T cells within the tumor stroma [69]. Consistently, T cell motility is facilitated in loose fibronectin and collagen regions, whereas T cells poorly migrate in dense matrix areas of lung tumors. Salmon and coauthors reported that also the orientation of extracellular matrix fibers influences antitumor immunity by dictating the migratory trajectory of T cells [70]. In accordance, collagenase-mediated matrix reduction increased the ability of T cells to contact cancer cells, indicating that targeting the ECM organization may improve the immune cell access to tumor sites. This is more relevant in pancreatic cancer, where the excessive desmoplasia abrogates T-cell chemokine-guided movement toward tumor cells and where the dense collagen networks represent a physical barrier to favour intrastromal T-cell trapping [71]. To migrate into a stiffened matrix, cells need to compress their nucleus affecting the gene expression and cell migration rate (for review see [72]). Moreover, the nuclear compression induced by matrix stiffness leads to multiple damage in the nucleus and membrane at forced passage, culminating in T cell death as reported for immunosenescence and ECM aging [73].
A recent very comprehensive work of Pearce and coauthors has profiled an evolving human metastatic microenvironment of ovarian cancer, using analysis that includes gene expression, matrix proteomics, cytokine/chemokine expression, ECM organization and biomechanical properties [74]. Pearce et al., have identified a matrix response, conserved in other cancers, that predicts tissue stiffness and extent of disease. Importantly, an high matrix index correlates with Treg and Th2 signatures [74]. Since ECM is mainly produced by stromal fibroblasts, it is not surprising that the density of alpha-smooth muscle actin (α-SMA) and fibroblast activation protein alpha (α-FAP) positive cells, two markers commonly associated with CAF activation, strongly associates with a score of disease progression (high disease score) [74].
Experimental models to recapitulate TME
The extraordinary advances in immune oncology and the comprehension that the majority of the mechanisms of therapy resistance comes from the TME, impose great efforts to develop models able to resemble the complexity of the TME.
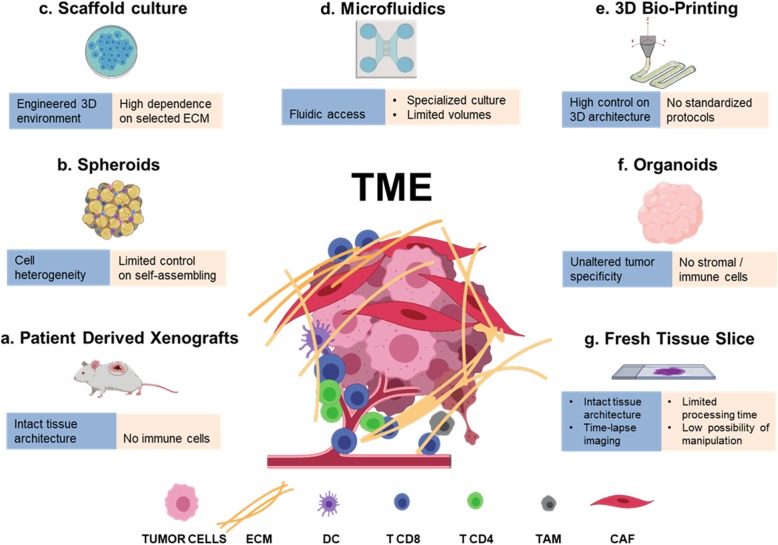
The animal models have improved our knowledge in cancer biology and have provided the scientific basis for numerous clinical trials, but they are unable to fully recapitulate the human tumor microenvironment. Recently, the development of standardized minimal information patient-derived xenograft (PDX-MI) models, with an intact ECM architecture and stromal component, represents a powerful tool to predict efficacy of cancer therapeutics [75]. These models however, lacking immune cells, are unsuitable to study the human tumor immune microenvironment, unless engrafted with functional human immune system (Fig. 1a) [76, 77]. Advantages and pitfalls of animal models developed for immune oncology research have been recently reviewed by Olson and co-authors [78].
Fig. 1.
Modelling the TME. Schematic representation of the major preclinical models and bio-fabrication techniques (a-g) employed to recapitulate TME complexity. For each model advantages (blue) and limitations (beige) are reported
The recent advances in in vitro 3D cultures are providing new models for translating basic knowledge to novel treatment in cancer [79].
Herein we report the major 3D model platforms (Fig. 1).
Bio-fabrication techniques for cancer 3D models
Tumor spheroids are 3D cellular aggregates of uniform or heterogeneous cell populations derived from tissue fragments mechanically or enzymatically partially digested (Fig. 1b). These 3D platforms are obtained in the absence of a scaffolding material, as cultured cells produce their own ECM. There are four major techniques used to induce cancer spheroids in vitro [80]: i) agitation-based techniques, in which cells are cultured in suspension using spinner flasks, and will spontaneously form multiple aggregates of diverse shape and dimension; ii) liquid overlay techniques, in which non-adhesive substrates promote cell-cell interaction and fusion, forming 3D aggregates that are cultured in static suspension condition; iii) hanging-drop techniques, where micro-reactors of static culture-medium droplets produce more consistent, isolated spheroids; iv) microfluidic reactors, in which injected cells are grouped in trapping chambers, where they can fuse in more controlled, dynamic environments. Tumor spheroids have been considered a gold-standard for cancer 3D culture, as they allow for the recapitulation of important features of TME heterogeneity [81–83], such as oxygen gradients [84, 85], and immune infiltration [86]. Nonetheless, this approach is based on the self-assembling of cells, and this limits the control over the 3D culture environment, which is certainly needed for the methodical investigation of specific TME features.
-
Scaffold-based approaches consist in the seeding or encapsulation of tumor/stromal cells in bio-materials that mimic the ECM of solid tissues (Fig. 1c) [87]. Cell seeding is done on pre-formed micro-porous or fibrous materials obtained by different techniques, such as two-phase emulsions and foams, freeze-drying or electro-spinning [88]. On the contrary, cell encapsulation is obtained by suspending cells on precursor macromolecular solutions that can undergo a biocompatible sol-gel transition, through which cells are embedded in a surrounding hydrogel, usually shaped as micro-droplet or micro-filament by means of micro-fabrication technologies, such as lithography and microfluidics [89]. Materials used as scaffolds can impair chemical and mechanical signals to cells, and can serve as tools to understand how the composition, architecture and stiffness of the ECM influence tumor proliferation [90], motility [91], matrix remodeling [92] and immune-escape [93, 94]. As an example, by employing a 3D scaffold model it has been shown that CAFs modulated the ability of specific T lymphocytes to kill breast cancer cells via TGF-β and IL-10 [95], indicating that cancer–immune-cell interaction needs a complex stroma to be evaluated. Recently, a culture platform based on alginate microencapsulation and stirred culture systems was explored to develop the 3D-3-culture, which entails the co-culture of NSCLC tumor cell spheroids, CAFs and monocytes. The Authors have demonstrated that the 3D-3-culture recreates an invasive and immunosuppressive TME, with accumulation of cytokines/chemokines, ECM elements and matrix metalloproteinases, promoting cell-cell interactions and supporting cell migration within the alginate microcapsules. Moreover, the 3D-3-culture was tested with chemo- and immunotherapeutic agents and the response to drugs was assessed in each cellular component, thus demonstrating that this 3D-3-culture constitutes a novel tool to study tumor-immune interaction in response to chemotherapeutic and immunomodulatory drugs [96].
Natural or synthetic materials can be used as scaffolds [97]; the firsts, composed of proteins and/or polysaccharides, enjoy an inherent biocompatibility and bioactivity, as they are usually native components of ECMs, but can suffer from incoherent composition, stiffness and degradability, and can potentially activate immune cells; synthetic materials, on the contrary, usually needs chemical modification with amino-acidic derivatives to increase their bio-adhesion, but can be strictly controlled in terms of bio-degradation, mechanical properties and purity. In the attempt to recapitulate the advantages of each material system, the use of hybrid composites of linked natural and synthetic macromolecules has also been tested [98]. Despite the great efforts focused on designing new reliable matrices that could mimic the in vivo complexity of TME, the most commonly used scaffold to date is the commercially available Matrigel which is an assortment of ECM proteins extracted from Englebreth-Holm-Swarm tumors in mice [99] containing also a variable amount of growth factors [100]. Even if Matrigel has been successfully employed in the 3D cultures of different tumor models [101] and in stem cell studies [102, 103] a low batch-to-batch reproducibility limits its applications. A promising trend is the use of native ECM obtained by cancer tissue decellularization, that can be employed as scaffold for cell seeding [104] or as tumor-homogenate additive component of 3D gels [105], in order to mimic in vitro the TME architectural features. This approach offers the future chance of preserving some environmental characteristics of specific, human-derived tumors that can be incorporated in engineered 3D models.
-
Microfluidics is another potent tool in cancer tissue modeling (Fig. 1d). As mentioned, microfluidic chips can be used as dynamic bioreactors for the culture of tissue spheroids [106], or for the precise shaping of micro-engineered cell-embedding hydrogels [107]; beside these applications, proper tumor-on-chip platforms have been designed to recreate controllable culture environments that integrate microfluidics, tissue engineering and biomaterials [108]. Organ-on-a-chip platforms have many biological applications that, starting from drug screening, have the potential to deeply impact the personalized medicine [109].
Recent literature presents a novel method of profiling response to PD-1 blockade using organotypic tumor spheroids cultured in collagen hydrogels suspended in a 3D microfluidic device [110]. The Authors report that the spheroids retain autologous immune cells, and that short-term culture and cytokine profiling of the organotypic tumors is feasible using this 3-D microfluidic device. This ex vivo functional immune profiling recapitulates key features of in vivo response and resistance to ICB and could represent a useful tool in the identification of biomarkers of ICB treatment response and, as the Authors reported, in the exploration of novel therapeutic combinations to enhance response to PD-1 blockade [110]. Details of the method and novel applications including RNA sequencing (RNASeq) and computational methods used to study immune cell changes in response to ex vivo ICB, have been reported in a subsequent publication where the Authors also discuss the limitations of the method [111]. A similar approach has been recently employed to demonstrate that the inhibition of cyclin-dependent kinase (CDK) 4 and 6 may activate CTL/TH1 responses to elicit antitumor immunity and that anti–PD-1 combined with CDK4/6 inhibition synergistically induced cell death ex vivo in murine-derived organotypic spheroids of colon cancer [112].
Soft-lithographic masters are used to create perfusable channels of micrometric dimension, usually molded in silicone material, that can be functionalized with adhesion proteins, filled with ECM and seeded with cells. The distinctive value offered by microfluidic culture is the presence of accessible fluidic control that is particularly effective in mimicking the vasculature component of TME, offering the possibility to induce flow-related instructions to cells [113], model invasion [114, 115], neovascularization [116, 117], metastasis formation [118–120] immune cell infiltration [121–123], and drug delivery [124, 125]. Multi-step micro-fabrication, the need of extensive user training, specific set-up equipment, the challenges associated with small-volumes protocols of culture and staining, and the difficulties in recovering seeded cells for further characterization, are among the main disadvantages of these otherwise high-performance platforms.
-
3D Bioprinting (3DBP) is an emerging technique in tissue engineering that holds great promises for tissue and cancer in vitro modeling (Fig. 1e) [126]. It consists in the application of digital fabrication technologies, specifically 3D printing, to the process of cell encapsulation. Living bio-constructs are created starting from a computer 3D model that is reproduced by robotically controlled dispensing systems that stack 2D layers of cells and biomaterials, the so-called bio-ink, in a layer-by-layer fashion to form arbitrary shapes. The bio-ink can be constituted by a dispersion of cells embedded in a pre-formed hydrogel or in a liquid solution of macromolecules that are induced to form a gel after the deposition process [127]. The deposition is achieved by using micro-metric building blocks in the form of droplets or filaments of cell-embedding ECM using either ink-jet technology [128], laser-forward transfer from donor slides [129] or by means of piston/pressure driven extrusion needles [130]. By using multiple dispensing heads or fluidic switches, it is possible to design heterogeneous culture platforms in which the spatial organization of different types of cells, tissue interface or ECM is controlled [131]. Alternatively, as we have reported, microfluidic switches can interchange the delivery of different bio-ink to a single dispensing head [132] following programmed sequences that, in harmony with the printing code, generates the desired heterogeneous structures.
This technology, thanks to the use of automated systems, enjoys great repeatability. Also, cancer and stromal cells, as well as mechanical and bio-chemical gradients, can be consistently arranged in 3D space following a pre-determined design, allowing for the systematic investigation of cellular/ECM structure-related influences on TME. Further, with 3DBP it is possible to embed cellularized and perfusable vascular structures within printed bio-constructs [133], useful for the replication of diffusive gradients, and to model cellular dynamics such as immune infiltration or cancer intra/extravasion and migration [134].
3DBP is a relatively young technique, and to date the examples of application of this bio-fabrication technique for creating cancer tissue models are limited. Nonetheless, the possibility offered in terms of precise design of TME features is great. An actual impedance that restricts the wide use of 3DBP is the absence of a consolidated technique: nowadays, many different bioprinting approaches are under development among research groups, and even if 3DBP machines start to be present in the market, most researchers build their own set-up in house. Each technique exploits specific bio-ink compositions, rheological properties and cell concentration [135], making the correlation of results difficult. Further, bioink-composition needs to be finely tuned to meet both technological and biological requisites. Material stiffness, chemistry, selected cell populations and their seeding density are all parameters that influence cell behavior in vitro [136–138] but that can also hamper the suitability of the bioink to the printing process.
Organoids are considered the more fisiological 3D culture models and various definition are available in literaure (Fig. 1f) (for an historical timeline of organoids and 3D cell cultures see Simian and Bissell [79]). Long term organoid cultures have been established from different primary and metastatic cancer tissues and have been reported able to resemble the tissue they were derived from. Their employement to predict the response to therapy is actually investigated also thanks to the effort of Human Cancer Model Initiative (HCMI), a globally accessible bank which includes information of novel cancer cell culture models including organoids [139]. Recently, they have been successfully employed to study the matched tumor specific T cell reactivity overcoming the technical limitations in obtaining primary tumor cell lines other than melanoma. In agreement, Dijkstra and co-authors have reported that the co-colture of peripheral blood lymphocytes (PBLs) with tumor organoids obtained by the autologous patient is an efficacious and unbiased strategy to generate tumor-reactive T cells from NSCLC and colorectal cancer (CRC) patients [140]. This indicates that this approach may bypass the isolation of tumor specific lymphocytes from the tumor tissue and may improve strategies for the generation of patient-specific T cells for adoptive T cell transfer.
Ex vivo tissue slices represents a promising technique which preserves tissue 3D architecture and pathway activity for short time (Fig. 1g) [141]. Recently, ex vivo assays have been developed to track T cells in fresh human tumor tissues, allowing to identify the extracellular matrix as a major stromal component in influencing T cell migration [69]. Dynamic imaging microscopy has been recently employed to study the mechanism underlying T cell exclusion by analyzing the interaction between endogenous CD8 T cells and TAMs in the tumor stroma. The translation in a murine model showed that the depletion of TAMs might improve the efficacy of anti–PD-1 immunotherapy [16]. This system may help in the screening of novel immunotherapy agents and in monitoring T cells.
Matrix biomechanics: Methods for the study
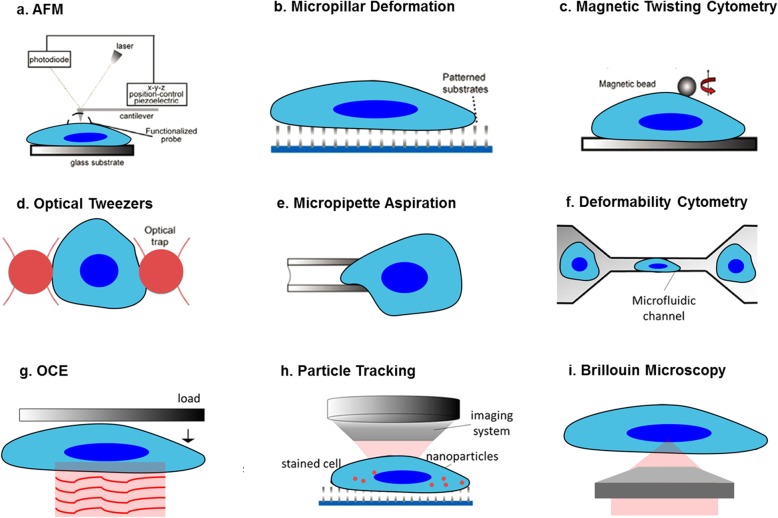
As indicated by all the data discussed in this review, ECM stiffness is a critical determinant in cancer and correlates with an immune suppressive TME. Unfortunately, our understanding on how the biomechanical properties of the extracellular matrix and the individual intracellular compartments change and contribute to the pathogenesis of cancer remains limited as a consequence of the available methods used to measure stiffness. While standard techniques require the application of invasive contact forces to the samples, others are intrinsically limited by a poor spatial resolution. The most common and widely accepted method to measure cellular elasticity, or stiffness in common language, is represented by Atomic Force Microscopy (AFM), which can reach a transverse resolution of the order of a few nanometers (Fig. 2a) [142]. AFM quantifies stiffness from the quasi-static Young’s modulus, which is measured by inducing a cellular displacement in response to the application of a sharp nanoindenter onto the superficial cellular membrane, with depths of a few nanometers [143]. In particular, the Young’s modulus is derived from the analysis performed by a variety of models of the deflection of the cantilever on which the nanoindenter is mounted. The contact process makes the AFM destructive because it can potentially invoke a cellular reaction. As a result, AFM cannot perform in-vivo measurements and the Young’s modulus can only be measured across the superficial cellular membrane in two-dimensional microenvironments where cells are tethered. Another non-negligible limitation of the AFM is given by the low axial resolution due to the unconfined contact force to the sample. As a consequence, values of the Young’s Modulus must be thought as average stiffness quantities along the strain direction. The contact mechanism together with the poor axial resolution make the AFM incapable of providing information inside the volume of neither the extracellular matrix or the intracellular compartments, where fundamental biomechanical properties of individual structures are currently unknown.
Fig. 2.
Schema of the methods to measure the cellular biomechanics properties. Standard methods, such as AFM (a), micropillar deformation (b), magnetic twisting cytometry (c), optical tweezers (d), micropipette aspiration (e), deformability cytometry (f) and OCE (g), require the application of contact forces to the extracellular matrix and measure stiffness from cellular displacement. The contact requirement makes these methods destructive and not capable to retrieve volumetric information. On the other hand, typical noncontact techniques, such as particle tracking (h), are either limited by an intrinsically low spatial resolution or require sample labelling through the use of nanoparticles. A promising method to non-invasively assess the extracellular and intracellular biomechanics in 3D is Brillouin microscopy (i), where light probes thermally activated spontaneous acoustic waves. Adapted by permission from Springer Nature: Bao G and Suresh S. Cell and molecular mechanics of biological materials. Nat Mater. 2003;2(11):715-25, © 2003 [158]
The AFM drawbacks similarly affect, to some extent, the other contact methods, where stiffness is obtained from the investigation of a sample strain in response to an applied stress. For example, elastic micropillar deformation (Fig. 2b) measures the deflection induced by the cellular focal adhesion on a patterned substrate microarray [144]. Magnetic twisting cytometry (Fig. 2c) uses magnetic beads attached to functionalized cellular surfaces [145]. The beads are controlled by external magnetic fields to induce a cellular deformation analyzed to extract the viscoelastic properties. Similarly, optical tweezers (Fig. 2d) employ a focused laser beam to control micron-size and high refractive index dielectric particles attached to the cell [146]. However, in-vivo measurements cannot be performed using optical tweezing or magnetic twisting due to the high power required and the use of particles. In micropipette aspiration (Fig. 2e), the sample is deformed by applying suction via a micropipette placed on the sample surface [147]. Recording of the cellular deformation allows to infer the mechanical properties. Similarly, deformability cytometry (Fig. 2f) measures cellular deformation by applying shear stresses or pressure gradients in suspension, which make this technique subject to significant non-linear effects [148]. Optical coherence elastography (OCE), (Fig. 2g) performs OCT measurements while inducing a certain strain to the sample using loads or ultrasound fields [149]. Although OCE provides rapid and three-dimensional biomechanical imaging, this typically requires contact with the sample and cannot perform extracellular or intracellular measurements due to the limited (> 10 μm) spatial resolution.
A noncontact method to assess stiffness at high transverse and temporal resolution is particle tracking [150]. Particle tracking (Fig. 2h) monitors and subsequently processes the Brownian motion trajectories of particles embedded in a sample to extract its viscous properties. Despite the noncontact approach, particle tracking requires a sample labelling with micro-beads. Moreover, complex models need to be applied in order to process the particle dynamics, while axial resolution is lower than tens of microns. Other noncontact techniques are those based on the application of ultrasound fields [151] or magnetic resonance [152]. However, these are intrinsically limited by a poor spatial (> 100 μm) resolution. As a result, these methods are not suitable to assess the stiffness of the extracellular matrix.
A promising, recently developed method to measure the three-dimensional biomechanical properties of both extracellular and intracellular matrixes is confocal Brillouin microscopy (Fig. 2i) [153, 154]. Brillouin light scattering is an inelastic process arising from the interaction of light with thermally activated acoustic waves that locally propagate in matter at the acoustic velocity. In Brillouin microscopy, the biomechanical properties are measured from the analysis of the Brillouin spectrum of the light scattered composed of a central elastic (Rayleigh) peak and by two inelastic (Brillouin) peaks. The frequency and the linewidth of the Brillouin peaks are related to the complex high-frequency Longitudinal elastic modulus, which bears information on both elasticity and viscosity of a sample [155]. The all-optical and label-free approach makes confocal Brillouin microscopy minimally invasive, while the optical sectioning capability enables a submicron transverse and axial resolutions [156, 157]. These key peculiarities may promote Brillouin microscopy as a novel tool of choice to perform measurements of the three-dimensional biomechanics of extracellular and intracellular compartments in physiological and in-vivo environments. In turn, Brillouin microscopy may elicit fundamental insights on the biomechanical role of the extracellular matrix and its variations during the different stages in cancer progression.
Conclusions
Immune oncology has revolutionized the therapeutic landscape for at least a portion of cancer patients. However, many critical questions remain opened and need urgent answers to identify patient responsive to ICB therapy and define novel combined therapies. It is largely demonstrated that the study of TIME and the identification of TIME subclasses is crucial for improving immunotherapy strategies [3].
For a progress to occur in the field, a close cooperation among biologists, bioengineers, biophysics, bioinformatics and clinicians has to be encouraged to allow the standardization of exciting new 3D platforms based on advances in biotechnologies and with the potential to impact the clinical practice.
Acknowledgements
The authors apologize to those authors whose work could not be cited due to space limitations.
Funding
PN is supported by the Italian Association for Cancer Research AIRC: 5 × 1000, 12182, and IG 15224.
Availability of data and materials
Not applicable.
Abbreviations
- AFM
Atomic force microscopy
- CAF
Cancer-associated fibroblast
- CCL4
C-C motif chemokine ligand 4
- CDK
Cyclin-dependent kinase
- COX
Cyclooxygenase
- CRC
Colorectal cancer
- CSF1
Colony-stimulating factor 1
- CSF1R
Colony-stimulating factor 1 receptor
- CTL
Cytotoxic T lymphocyte
- CTLA4
Cytotoxic T-lymphocyte antigen protein 4
- CXCL12
C-X-C motif chemokine ligand 12
- EMT
Epithelial-mesenchymal transition
- FAK
Focal adhesion kinase
- GM-CSF
Granulocyte-macrophage colony-stimulating factor
- HCC
Hepatocellular carcinoma
- HLA
Human leukocyte antigen
- HNSCC
Head and neck squamous cell carcinoma
- ICB
Immune checkpoint blockade
- IFNγ
Interferon-γ
- IL-2
Interleukin-2
- IL-6
Interleukin-6
- MDSC
Myeloid-derived suppressor cell
- NSCLC
Non-small cell lung cancer
- OCE
Optical coherence elastography
- PBL
Peripheral blood lymphocytes
- PD-1
Programmed cell death 1
- PDAC
Pancreatic ductal adenocarcinoma
- PD-L1
Programmed cell death Ligand 1
- PDPN
Podoplanin
- PDX
Patient derived xenograft
- PGE2
Prostaglandin E2
- PI3K
Phosphoinositide 3-kinase
- RNASeq
RNA sequencing
- STAT3
Signal transducer and activator of transcription 3
- TAM
Tumor-associated macrophage
- TAZ
WW domain containing transcription regulator 1
- TGFβ
Transforming growth factor β
- TIDE
Tumor immune dysfunction and exclusion
- TIL
Tumor-infiltrating lymphocytes
- TIM3
T-cell immunoglobulin and mucin-domain containing-3
- TIME
Tumor immune environment
- TLS
Tertiary lymphoid structure
- TME
Tumor microenvironment
- Treg
Regulatory T
- YAP
Yes-associated protein 1
- α-FAP
Fibroblast activation protein alpha
- α-SMA
Alpha-smooth muscle actin
Authors’ contributions
PN developed the idea and revised the manuscript. PN and FDM contributed to study design. FDM, CC, PT and GA drafted the manuscript and prepared the figures; GR critically revised the manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Francesca Di Modugno, Email: francesca.dimodugno@ifo.gov.it.
Cristina Colosi, Email: cristinacolosi@gmail.com.
Paola Trono, Email: paola.trono@ifo.gov.it.
Giuseppe Antonacci, Email: giuseppe.antonacci.iit@gmail.com.
Giancarlo Ruocco, Email: giancarlo.ruocco@iit.it.
Paola Nisticò, Email: paola.nistico@ifo.gov.it.
References
- 1.Gong J, Chehrazi-Raffle A, Reddi S, Salgia R. Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: a comprehensive review of registration trials and future considerations. J Immunother Cancer. 2018;6(1):8. doi: 10.1186/s40425-018-0316-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pitt JM, Vetizou M, Daillere R, Roberti MP, Yamazaki T, Routy B, et al. Resistance mechanisms to immune-checkpoint blockade in Cancer: tumor-intrinsic and -extrinsic factors. Immunity. 2016;44(6):1255–1269. doi: 10.1016/j.immuni.2016.06.001. [DOI] [PubMed] [Google Scholar]
- 3.Binnewies M, Roberts EW, Kersten K, Chan V, Fearon DF, Merad M, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med. 2018;24(5):541–550. doi: 10.1038/s41591-018-0014-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bissell MJ, Radisky D. Putting tumours in context. Nat Rev Cancer. 2001;1(1):46–54. doi: 10.1038/35094059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wei SC, Duffy CR, Allison JP. Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 2018;8(9):1069–1086. doi: 10.1158/2159-8290.CD-18-0367. [DOI] [PubMed] [Google Scholar]
- 6.Aldarouish M, Wang C. Trends and advances in tumor immunology and lung cancer immunotherapy. J Exp Clin Cancer Res. 2016;35(1):157. doi: 10.1186/s13046-016-0439-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A. Primary, adaptive, and acquired resistance to Cancer immunotherapy. Cell. 2017;168(4):707–723. doi: 10.1016/j.cell.2017.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen DS, Mellman I. Elements of cancer immunity and the cancer-immune set point. Nature. 2017;541(7637):321–330. doi: 10.1038/nature21349. [DOI] [PubMed] [Google Scholar]
- 9.Dieu-Nosjean MC, Goc J, Giraldo NA, Sautes-Fridman C, Fridman WH. Tertiary lymphoid structures in cancer and beyond. Trends Immunol. 2014;35(11):571–580. doi: 10.1016/j.it.2014.09.006. [DOI] [PubMed] [Google Scholar]
- 10.Cottrell TR, Thompson ED, Forde PM, Stein JE, Duffield AS, Anagnostou V, et al. Pathologic features of response to neoadjuvant anti-PD-1 in resected non-small-cell lung carcinoma: a proposal for quantitative immune-related pathologic response criteria (irPRC) Ann Oncol. 2018;29(8):1853–1860. doi: 10.1093/annonc/mdy218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Johansson-Percival A, He B, Li ZJ, Kjellen A, Russell K, Li J, et al. De novo induction of intratumoral lymphoid structures and vessel normalization enhances immunotherapy in resistant tumors. Nat Immunol. 2017;18(11):1207–1217. doi: 10.1038/ni.3836. [DOI] [PubMed] [Google Scholar]
- 12.Zhu G, Falahat R, Wang K, Mailloux A, Artzi N, Mule JJ. Tumor-associated tertiary lymphoid structures: gene-expression profiling and their bioengineering. Front Immunol. 2017;8:767. doi: 10.3389/fimmu.2017.00767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.De Henau O, Rausch M, Winkler D, Campesato LF, Liu C, Cymerman DH, et al. Overcoming resistance to checkpoint blockade therapy by targeting PI3Kgamma in myeloid cells. Nature. 2016;539(7629):443–447. doi: 10.1038/nature20554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Holmgaard RB, Brachfeld A, Gasmi B, Jones DR, Mattar M, Doman T, et al. Timing of CSF-1/CSF-1R signaling blockade is critical to improving responses to CTLA-4 based immunotherapy. Oncoimmunology. 2016;5(7):e1151595. doi: 10.1080/2162402X.2016.1151595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pyonteck SM, Akkari L, Schuhmacher AJ, Bowman RL, Sevenich L, Quail DF, et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat Med. 2013;19(10):1264–1272. doi: 10.1038/nm.3337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Peranzoni E, Lemoine J, Vimeux L, Feuillet V, Barrin S, Kantari-Mimoun C, et al. Macrophages impede CD8 T cells from reaching tumor cells and limit the efficacy of anti-PD-1 treatment. Proc Natl Acad Sci U S A. 2018;115(17):E4041–E4E50. doi: 10.1073/pnas.1720948115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Saggar JK, Yu M, Tan Q, Tannock IF. The tumor microenvironment and strategies to improve drug distribution. Front Oncol. 2013;3:154. doi: 10.3389/fonc.2013.00154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Becker JC, Andersen MH, Schrama D, Thor SP. Immune-suppressive properties of the tumor microenvironment. Cancer Immunol Immunother. 2013;62(7):1137–1148. doi: 10.1007/s00262-013-1434-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell. 2016;165(1):35–44. doi: 10.1016/j.cell.2016.02.065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li Y, Patel SP, Roszik J, Qin Y. Hypoxia-driven immunosuppressive metabolites in the tumor microenvironment: new approaches for combinational immunotherapy. Front Immunol. 2018;9:1591. doi: 10.3389/fimmu.2018.01591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jayaprakash P, Ai M, Liu A, Budhani P, Bartkowiak T, Sheng J, et al. Targeted hypoxia reduction restores T cell infiltration and sensitizes prostate cancer to immunotherapy. J Clin Invest. 2018;128(11):5137–5149. doi: 10.1172/JCI96268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chang CH, Qiu J, O'Sullivan D, Buck MD, Noguchi T, Curtis JD, et al. Metabolic competition in the tumor microenvironment is a driver of Cancer progression. Cell. 2015;162(6):1229–1241. doi: 10.1016/j.cell.2015.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Beavis PA, Stagg J, Darcy PK, Smyth MJ. CD73: a potent suppressor of antitumor immune responses. Trends Immunol. 2012;33(5):231–237. doi: 10.1016/j.it.2012.02.009. [DOI] [PubMed] [Google Scholar]
- 24.Allard B, Pommey S, Smyth MJ, Stagg J. Targeting CD73 enhances the antitumor activity of anti-PD-1 and anti-CTLA-4 mAbs. Clin Cancer Res. 2013;19(20):5626–5635. doi: 10.1158/1078-0432.CCR-13-0545. [DOI] [PubMed] [Google Scholar]
- 25.Zelenay S, van der Veen AG, Bottcher JP, Snelgrove KJ, Rogers N, Acton SE, et al. Cyclooxygenase-dependent tumor growth through evasion of immunity. Cell. 2015;162(6):1257–1270. doi: 10.1016/j.cell.2015.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Pages F, Mlecnik B, Marliot F, Bindea G, Ou FS, Bifulco C, et al. International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet. 2018;391(10135):2128–2139. doi: 10.1016/S0140-6736(18)30789-X. [DOI] [PubMed] [Google Scholar]
- 27.Dixon AR, Bathany C, Tsuei M, White J, Barald KF, Takayama S. Recent developments in multiplexing techniques for immunohistochemistry. Expert Rev Mol Diagn. 2015;15(9):1171–1186. doi: 10.1586/14737159.2015.1069182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Leelatian N, Doxie DB, Greenplate AR, Mobley BC, Lehman JM, Sinnaeve J, et al. Single cell analysis of human tissues and solid tumors with mass cytometry. Cytometry B Clin Cytom. 2017;92(1):68–78. doi: 10.1002/cyto.b.21481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pages C, et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science. 2006;313(5795):1960–1964. doi: 10.1126/science.1129139. [DOI] [PubMed] [Google Scholar]
- 30.Dunn GP, Bruce AT, Ikeda H, Old LJ, Schreiber RD. Cancer immunoediting: from immunosurveillance to tumor escape. Nat Immunol. 2002;3(11):991–998. doi: 10.1038/ni1102-991. [DOI] [PubMed] [Google Scholar]
- 31.Angelova M, Mlecnik B, Vasaturo A, Bindea G, Fredriksen T, Lafontaine L, et al. Evolution of metastases in space and time under immune selection. Cell. 2018;175(3):751–765. doi: 10.1016/j.cell.2018.09.018. [DOI] [PubMed] [Google Scholar]
- 32.Zhang L, Conejo-Garcia JR, Katsaros D, Gimotty PA, Massobrio M, Regnani G, et al. Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. N Engl J Med. 2003;348(3):203–213. doi: 10.1056/NEJMoa020177. [DOI] [PubMed] [Google Scholar]
- 33.Vose BM, Moore M. Human tumor-infiltrating lymphocytes: a marker of host response. Semin Hematol. 1985;22(1):27–40. [PubMed] [Google Scholar]
- 34.Nistico P, Mottolese M, Cascioli S, Benevolo M, Del Bello D, Di Modugno F, et al. Host immunosurveillance contributes to the control of erbB-2 overexpression in HLA-A2-breast-cancer patients. Int J Cancer. 1999;84(6):598–603. doi: 10.1002/(SICI)1097-0215(19991222)84:6<598::AID-IJC10>3.0.CO;2-7. [DOI] [PubMed] [Google Scholar]
- 35.Joyce JA, Fearon DT. T cell exclusion, immune privilege, and the tumor microenvironment. Science. 2015;348(6230):74–80. doi: 10.1126/science.aaa6204. [DOI] [PubMed] [Google Scholar]
- 36.Spranger S, Bao R, Gajewski TF. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature. 2015;523(7559):231–235. doi: 10.1038/nature14404. [DOI] [PubMed] [Google Scholar]
- 37.Aguilera TA, Giaccia AJ. Molecular pathways: oncologic pathways and their role in T-cell exclusion and immune evasion-a new role for the AXL receptor tyrosine kinase. Clin Cancer Res. 2017;23(12):2928–2933. doi: 10.1158/1078-0432.CCR-17-0189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–1558. doi: 10.1038/s41591-018-0136-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Jerby-Arnon L, Shah P, Cuoco MS, Rodman C, Su MJ, Melms JC, et al. A Cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell. 2018;175(4):984–997. doi: 10.1016/j.cell.2018.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Haddad R, Saldanha-Araujo F. Mechanisms of T-cell immunosuppression by mesenchymal stromal cells: what do we know so far? Biomed Res Int. 2014;2014:216806. doi: 10.1155/2014/216806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kuzet SE, Gaggioli C. Fibroblast activation in cancer: when seed fertilizes soil. Cell Tissue Res. 2016;365(3):607–619. doi: 10.1007/s00441-016-2467-x. [DOI] [PubMed] [Google Scholar]
- 42.Lohr M, Schmidt C, Ringel J, Kluth M, Muller P, Nizze H, et al. Transforming growth factor-beta1 induces desmoplasia in an experimental model of human pancreatic carcinoma. Cancer Res. 2001;61(2):550–555. [PubMed] [Google Scholar]
- 43.Fearon DT. The carcinoma-associated fibroblast expressing fibroblast activation protein and escape from immune surveillance. Cancer Immunol Res. 2014;2(3):187–193. doi: 10.1158/2326-6066.CIR-14-0002. [DOI] [PubMed] [Google Scholar]
- 44.Kalluri R. The biology and function of fibroblasts in cancer. Nat Rev Cancer. 2016;16(9):582–598. doi: 10.1038/nrc.2016.73. [DOI] [PubMed] [Google Scholar]
- 45.Costa A, Kieffer Y, Scholer-Dahirel A, Pelon F, Bourachot B, Cardon M, et al. Fibroblast heterogeneity and immunosuppressive environment in human breast Cancer. Cancer Cell. 2018;33(3):463–479. doi: 10.1016/j.ccell.2018.01.011. [DOI] [PubMed] [Google Scholar]
- 46.Ohlund D, Elyada E, Tuveson D. Fibroblast heterogeneity in the cancer wound. J Exp Med. 2014;211(8):1503–1523. doi: 10.1084/jem.20140692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Cho H, Seo Y, Loke KM, Kim SW, Oh SM, Kim JH, et al. Cancer-stimulated CAFs enhance monocyte differentiation and Protumoral TAM activation via IL6 and GM-CSF secretion. Clin Cancer Res. 2018;24(21):5407–5421. doi: 10.1158/1078-0432.CCR-18-0125. [DOI] [PubMed] [Google Scholar]
- 48.Mace TA, Ameen Z, Collins A, Wojcik S, Mair M, Young GS, et al. Pancreatic cancer-associated stellate cells promote differentiation of myeloid-derived suppressor cells in a STAT3-dependent manner. Cancer Res. 2013;73(10):3007–3018. doi: 10.1158/0008-5472.CAN-12-4601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Cheng Y, Li H, Deng Y, Tai Y, Zeng K, Zhang Y, et al. Cancer-associated fibroblasts induce PDL1+ neutrophils through the IL6-STAT3 pathway that foster immune suppression in hepatocellular carcinoma. Cell Death Dis. 2018;9(4):422. doi: 10.1038/s41419-018-0458-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.He G, Zhang H, Zhou J, Wang B, Chen Y, Kong Y, et al. Peritumoural neutrophils negatively regulate adaptive immunity via the PD-L1/PD-1 signalling pathway in hepatocellular carcinoma. J Exp Clin Cancer Res. 2015;34:141. doi: 10.1186/s13046-015-0256-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kojima Y, Acar A, Eaton EN, Mellody KT, Scheel C, Ben-Porath I, et al. Autocrine TGF-beta and stromal cell-derived factor-1 (SDF-1) signaling drives the evolution of tumor-promoting mammary stromal myofibroblasts. Proc Natl Acad Sci U S A. 2010;107(46):20009–20014. doi: 10.1073/pnas.1013805107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Chou CK, Schietinger A, Liggitt HD, Tan X, Funk S, Freeman GJ, et al. Cell-intrinsic abrogation of TGF-beta signaling delays but does not prevent dysfunction of self/tumor-specific CD8 T cells in a murine model of autochthonous prostate cancer. J Immunol. 2012;189(8):3936–3946. doi: 10.4049/jimmunol.1201415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ziani L, Chouaib S, Thiery J. Alteration of the antitumor immune response by Cancer-associated fibroblasts. Front Immunol. 2018;9:414. doi: 10.3389/fimmu.2018.00414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Tauriello DVF, Palomo-Ponce S, Stork D, Berenguer-Llergo A, Badia-Ramentol J, Iglesias M, et al. TGFbeta drives immune evasion in genetically reconstituted colon cancer metastasis. Nature. 2018;554(7693):538–543. doi: 10.1038/nature25492. [DOI] [PubMed] [Google Scholar]
- 55.Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554(7693):544–548. doi: 10.1038/nature25501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Lu P, Takai K, Weaver VM, Werb Z. Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb Perspect Biol. 2011;3(12). [DOI] [PMC free article] [PubMed]
- 57.Wei SC, Yang J. Forcing through tumor metastasis: the interplay between tissue rigidity and epithelial-mesenchymal transition. Trends Cell Biol. 2016;26(2):111–120. doi: 10.1016/j.tcb.2015.09.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Bissell MJ, Hines WC. Why don't we get more cancer? A proposed role of the microenvironment in restraining cancer progression. Nat Med. 2011;17(3):320–329. doi: 10.1038/nm.2328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Pickup MW, Mouw JK, Weaver VM. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 2014;15(12):1243–1253. doi: 10.15252/embr.201439246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Paszek MJ, Zahir N, Johnson KR, Lakins JN, Rozenberg GI, Gefen A, et al. Tensional homeostasis and the malignant phenotype. Cancer Cell. 2005;8(3):241–254. doi: 10.1016/j.ccr.2005.08.010. [DOI] [PubMed] [Google Scholar]
- 61.Gilbert PM, Weaver VM. Cellular adaptation to biomechanical stress across length scales in tissue homeostasis and disease. Semin Cell Dev Biol. 2017;67:141–152. doi: 10.1016/j.semcdb.2016.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Di Modugno F, Spada S, Palermo B, Visca P, Iapicca P, Di Carlo A, et al. hMENA isoforms impact NSCLC patient outcome through fibronectin/beta1 integrin axis. Oncogene. 2018;37(42):5605–5617. doi: 10.1038/s41388-018-0364-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Dupont S, Morsut L, Aragona M, Enzo E, Giulitti S, Cordenonsi M, et al. Role of YAP/TAZ in mechanotransduction. Nature. 2011;474(7350):179–183. doi: 10.1038/nature10137. [DOI] [PubMed] [Google Scholar]
- 64.Calvo F, Ege N, Grande-Garcia A, Hooper S, Jenkins RP, Chaudhry SI, et al. Mechanotransduction and YAP-dependent matrix remodelling is required for the generation and maintenance of cancer-associated fibroblasts. Nat Cell Biol. 2013;15(6):637–646. doi: 10.1038/ncb2756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Jiang Z, Zhou C, Cheng L, Yan B, Chen K, Chen X, et al. Inhibiting YAP expression suppresses pancreatic cancer progression by disrupting tumor-stromal interactions. J Exp Clin Cancer Res. 2018;37(1):69. doi: 10.1186/s13046-018-0740-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Murakami S, Shahbazian D, Surana R, Zhang W, Chen H, Graham GT, et al. Yes-associated protein mediates immune reprogramming in pancreatic ductal adenocarcinoma. Oncogene. 2017;36(9):1232–1244. doi: 10.1038/onc.2016.288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Jiang H, Hegde S, Knolhoff BL, Zhu Y, Herndon JM, Meyer MA, et al. Targeting focal adhesion kinase renders pancreatic cancers responsive to checkpoint immunotherapy. Nat Med. 2016;22(8):851–860. doi: 10.1038/nm.4123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wolf K, Te Lindert M, Krause M, Alexander S, Te Riet J, Willis AL, et al. Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force. J Cell Biol. 2013;201(7):1069–1084. doi: 10.1083/jcb.201210152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Bougherara H, Mansuet-Lupo A, Alifano M, Ngo C, Damotte D, Le Frere-Belda MA, et al. Real-time imaging of resident T cells in human lung and ovarian carcinomas reveals how different tumor microenvironments control T lymphocyte migration. Front Immunol. 2015;6:500. doi: 10.3389/fimmu.2015.00500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Salmon H, Franciszkiewicz K, Damotte D, Dieu-Nosjean MC, Validire P, Trautmann A, et al. Matrix architecture defines the preferential localization and migration of T cells into the stroma of human lung tumors. J Clin Invest. 2012;122(3):899–910. doi: 10.1172/JCI45817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hartmann N, Giese NA, Giese T, Poschke I, Offringa R, Werner J, et al. Prevailing role of contact guidance in intrastromal T-cell trapping in human pancreatic cancer. Clin Cancer Res. 2014;20(13):3422–3433. doi: 10.1158/1078-0432.CCR-13-2972. [DOI] [PubMed] [Google Scholar]
- 72.Hallmann R, Zhang X, Di Russo J, Li L, Song J, Hannocks MJ, et al. The regulation of immune cell trafficking by the extracellular matrix. Curr Opin Cell Biol. 2015;36:54–61. doi: 10.1016/j.ceb.2015.06.006. [DOI] [PubMed] [Google Scholar]
- 73.Moreau JF, Pradeu T, Grignolio A, Nardini C, Castiglione F, Tieri P, et al. The emerging role of ECM crosslinking in T cell mobility as a hallmark of immunosenescence in humans. Ageing Res Rev. 2017;35:322–335. doi: 10.1016/j.arr.2016.11.005. [DOI] [PubMed] [Google Scholar]
- 74.Pearce OMT, Delaine-Smith RM, Maniati E, Nichols S, Wang J, Bohm S, et al. Deconstruction of a metastatic tumor microenvironment reveals a common matrix response in human cancers. Cancer Discov. 2018;8(3):304–319. doi: 10.1158/2159-8290.CD-17-0284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Meehan TF, Conte N, Goldstein T, Inghirami G, Murakami MA, Brabetz S, et al. PDX-MI: minimal information for patient-derived tumor xenograft models. Cancer Res. 2017;77(21):e62–ee6. doi: 10.1158/0008-5472.CAN-17-0582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Wang M, Yao LC, Cheng M, Cai D, Martinek J, Pan CX, et al. Humanized mice in studying efficacy and mechanisms of PD-1-targeted cancer immunotherapy. FASEB J. 2018;32(3):1537–1549. doi: 10.1096/fj.201700740R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Shultz LD, Goodwin N, Ishikawa F, Hosur V, Lyons BL, Greiner DL. Human cancer growth and therapy in immunodeficient mouse models. Cold Spring Harb Protoc. 2014;2014(7):694–708. doi: 10.1101/pdb.top073585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Olson B, Li Y, Lin Y, Liu ET, Patnaik A. Mouse models for Cancer immunotherapy research. Cancer Discov. 2018;8(11):1358–1365. doi: 10.1158/2159-8290.CD-18-0044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Simian M, Bissell MJ. Organoids: a historical perspective of thinking in three dimensions. J Cell Biol. 2017;216(1):31–40. doi: 10.1083/jcb.201610056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Nunes AS, Barros AS, Costa EC, Moreira AF, Correia IJ. 3D tumor spheroids as in vitro models to mimic in vivo human solid tumors resistance to therapeutic drugs. Biotechnol Bioeng. 2019;116(1):206–226. doi: 10.1002/bit.26845. [DOI] [PubMed] [Google Scholar]
- 81.Khawar IA, Park JK, Jung ES, Lee MA, Chang S, Kuh HJ. Three dimensional mixed-cell spheroids mimic stroma-mediated Chemoresistance and invasive migration in hepatocellular carcinoma. Neoplasia. 2018;20(8):800–812. doi: 10.1016/j.neo.2018.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Sutherland RM. Cell and environment interactions in tumor microregions: the multicell spheroid model. Science. 1988;240(4849):177–184. doi: 10.1126/science.2451290. [DOI] [PubMed] [Google Scholar]
- 83.Song Y, Kim SH, Kim KM, Choi EK, Kim J, Seo HR. Activated hepatic stellate cells play pivotal roles in hepatocellular carcinoma cell chemoresistance and migration in multicellular tumor spheroids. Sci Rep. 2016;6:36750. doi: 10.1038/srep36750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Daster S, Amatruda N, Calabrese D, Ivanek R, Turrini E, Droeser RA, et al. Induction of hypoxia and necrosis in multicellular tumor spheroids is associated with resistance to chemotherapy treatment. Oncotarget. 2017;8(1):1725–1736. doi: 10.18632/oncotarget.13857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Riffle S, Hegde RS. Modeling tumor cell adaptations to hypoxia in multicellular tumor spheroids. J Exp Clin Cancer Res. 2017;36(1):102. doi: 10.1186/s13046-017-0570-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Herter S, Morra L, Schlenker R, Sulcova J, Fahrni L, Waldhauer I, et al. A novel three-dimensional heterotypic spheroid model for the assessment of the activity of cancer immunotherapy agents. Cancer Immunol Immunother. 2017;66(1):129–140. doi: 10.1007/s00262-016-1927-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Ferreira LP, Gaspar VM, Mano JF. Design of spherically structured 3D in vitro tumor models -advances and prospects. Acta Biomater. 2018;75:11–34. doi: 10.1016/j.actbio.2018.05.034. [DOI] [PubMed] [Google Scholar]
- 88.Ma PX. Scaffolds for tissue fabrication. Mater Today. 2004;7(5):30–40. doi: 10.1016/S1369-7021(04)00233-0. [DOI] [Google Scholar]
- 89.Kang A, Park J, Ju J, Jeong GS, Lee SH. Cell encapsulation via microtechnologies. Biomaterials. 2014;35(9):2651–2663. doi: 10.1016/j.biomaterials.2013.12.073. [DOI] [PubMed] [Google Scholar]
- 90.Loessner D, Stok KS, Lutolf MP, Hutmacher DW, Clements JA, Rizzi SC. Bioengineered 3D platform to explore cell-ECM interactions and drug resistance of epithelial ovarian cancer cells. Biomaterials. 2010;31(32):8494–8506. doi: 10.1016/j.biomaterials.2010.07.064. [DOI] [PubMed] [Google Scholar]
- 91.Ulrich TA, de Juan Pardo EM, Kumar S. The mechanical rigidity of the extracellular matrix regulates the structure, motility, and proliferation of glioma cells. Cancer Res. 2009;69(10):4167–4174. doi: 10.1158/0008-5472.CAN-08-4859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Wolf K, Wu YI, Liu Y, Geiger J, Tam E, Overall C, et al. Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat Cell Biol. 2007;9(8):893–904. doi: 10.1038/ncb1616. [DOI] [PubMed] [Google Scholar]
- 93.Florczyk SJ, Liu G, Kievit FM, Lewis AM, Wu JD, Zhang M. 3D porous chitosan-alginate scaffolds: a new matrix for studying prostate cancer cell-lymphocyte interactions in vitro. Adv Healthc Mater. 2012;1(5):590–599. doi: 10.1002/adhm.201100054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Dustin ML, de Fougerolles AR. Reprogramming T cells: the role of extracellular matrix in coordination of T cell activation and migration. Curr Opin Immunol. 2001;13(3):286–290. doi: 10.1016/S0952-7915(00)00217-X. [DOI] [PubMed] [Google Scholar]
- 95.Phan-Lai V, Florczyk SJ, Kievit FM, Wang K, Gad E, Disis ML, et al. Three-dimensional scaffolds to evaluate tumor associated fibroblast-mediated suppression of breast tumor specific T cells. Biomacromolecules. 2013;14(5):1330–1337. doi: 10.1021/bm301928u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Rebelo SP, Pinto C, Martins TR, Harrer N, Estrada MF, Loza-Alvarez P, et al. 3D-3-culture: a tool to unveil macrophage plasticity in the tumour microenvironment. Biomaterials. 2018;163:185–197. doi: 10.1016/j.biomaterials.2018.02.030. [DOI] [PubMed] [Google Scholar]
- 97.Nyga A, Cheema U, Loizidou M. 3D tumour models: novel in vitro approaches to cancer studies. J Cell Commun Signal. 2011;5(3):239–248. doi: 10.1007/s12079-011-0132-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Pradhan S, Hassani I, Seeto WJ, Lipke EA. PEG-fibrinogen hydrogels for three-dimensional breast cancer cell culture. J Biomed Mater Res A. 2017;105(1):236–252. doi: 10.1002/jbm.a.35899. [DOI] [PubMed] [Google Scholar]
- 99.Orkin RW, Gehron P, McGoodwin EB, Martin GR, Valentine T, Swarm R. A murine tumor producing a matrix of basement membrane. J Exp Med. 1977;145(1):204–220. doi: 10.1084/jem.145.1.204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hughes CS, Postovit LM, Lajoie GA. Matrigel: a complex protein mixture required for optimal growth of cell culture. Proteomics. 2010;10(9):1886–1890. doi: 10.1002/pmic.200900758. [DOI] [PubMed] [Google Scholar]
- 101.Benton G, Kleinman HK, George J, Arnaoutova I. Multiple uses of basement membrane-like matrix (BME/Matrigel) in vitro and in vivo with cancer cells. Int J Cancer. 2011;128(8):1751–1757. doi: 10.1002/ijc.25781. [DOI] [PubMed] [Google Scholar]
- 102.Ramos-Hryb AB, Da-Costa MC, Trentin AG, Calloni GW. Matrigel supports neural, melanocytic and chondrogenic differentiation of trunk neural crest cells. Int J Dev Biol. 2013;57(11–12):885–890. doi: 10.1387/ijdb.130206gw. [DOI] [PubMed] [Google Scholar]
- 103.Choi NY, Park YS, Ryu JS, Lee HJ, Arauzo-Bravo MJ, Ko K, et al. A novel feeder-free culture system for expansion of mouse spermatogonial stem cells. Mol Cells. 2014;37(6):473–479. doi: 10.14348/molcells.2014.0080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Genovese L, Zawada L, Tosoni A, Ferri A, Zerbi P, Allevi R, et al. Cellular localization, invasion, and turnover are differently influenced by healthy and tumor-derived extracellular matrix. Tissue Eng Part A. 2014;20(13–14):2005–2018. doi: 10.1089/ten.tea.2013.0588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Wang Z, Wang C, Abudukeremu A, Rui X, Liu S, Zhang X, et al. Engineering a tumor microenvironment-mimetic niche for tissue regeneration with xenogeneic Cancer cells. Adv Sci (Weinh) 2018;5(3):1700666. doi: 10.1002/advs.201700666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Hsiao AY, Torisawa YS, Tung YC, Sud S, Taichman RS, Pienta KJ, et al. Microfluidic system for formation of PC-3 prostate cancer co-culture spheroids. Biomaterials. 2009;30(16):3020–3027. doi: 10.1016/j.biomaterials.2009.02.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Sugimoto M, Kitagawa Y, Yamada M, Yajima Y, Utoh R, Seki M. Micropassage-embedding composite hydrogel fibers enable quantitative evaluation of cancer cell invasion under 3D coculture conditions. Lab Chip. 2018;18(9):1378–1387. doi: 10.1039/C7LC01280B. [DOI] [PubMed] [Google Scholar]
- 108.Tsai HF, Trubelja A, Shen AQ, Bao G. Tumour-on-a-chip: microfluidic models of tumour morphology, growth and microenvironment. J R Soc Interface. 2017;14(131). [DOI] [PMC free article] [PubMed]
- 109.Skardal A, Shupe T, Atala A. Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discov Today. 2016;21(9):1399–1411. doi: 10.1016/j.drudis.2016.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Jenkins RW, Aref AR, Lizotte PH, Ivanova E, Stinson S, Zhou CW, et al. Ex vivo profiling of PD-1 blockade using Organotypic tumor spheroids. Cancer Discov. 2018;8(2):196–215. doi: 10.1158/2159-8290.CD-17-0833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Aref AR, Campisi M, Ivanova E, Portell A, Larios D, Piel BP, et al. 3D microfluidic ex vivo culture of organotypic tumor spheroids to model immune checkpoint blockade. Lab Chip. 2018;18(20):3129–3143. doi: 10.1039/C8LC00322J. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Deng J, Wang ES, Jenkins RW, Li S, Dries R, Yates K, et al. CDK4/6 inhibition augments antitumor immunity by enhancing T-cell activation. Cancer Discov. 2018;8(2):216–233. doi: 10.1158/2159-8290.CD-17-0915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Rizvi I, Gurkan UA, Tasoglu S, Alagic N, Celli JP, Mensah LB, et al. Flow induces epithelial-mesenchymal transition, cellular heterogeneity and biomarker modulation in 3D ovarian cancer nodules. Proc Natl Acad Sci U S A. 2013;110(22):E1974–E1983. doi: 10.1073/pnas.1216989110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Nagaraju S, Truong D, Mouneimne G, Nikkhah M. Microfluidic tumor-vascular model to study breast Cancer cell invasion and Intravasation. Adv Healthc Mater. 2018;7(9):e1701257. doi: 10.1002/adhm.201701257. [DOI] [PubMed] [Google Scholar]
- 115.Liu T, Lin B, Qin J. Carcinoma-associated fibroblasts promoted tumor spheroid invasion on a microfluidic 3D co-culture device. Lab Chip. 2010;10(13):1671–1677. doi: 10.1039/c000022a. [DOI] [PubMed] [Google Scholar]
- 116.Cross VL, Zheng Y, Won Choi N, Verbridge SS, Sutermaster BA, Bonassar LJ, et al. Dense type I collagen matrices that support cellular remodeling and microfabrication for studies of tumor angiogenesis and vasculogenesis in vitro. Biomaterials. 2010;31(33):8596–8607. doi: 10.1016/j.biomaterials.2010.07.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Aung A, Theprungsirikul J, Lim HL, Varghese S. Chemotaxis-driven assembly of endothelial barrier in a tumor-on-a-chip platform. Lab Chip. 2016;16(10):1886–1898. doi: 10.1039/C6LC00184J. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Zhang Q, Liu T, Qin J. A microfluidic-based device for study of transendothelial invasion of tumor aggregates in realtime. Lab Chip. 2012;12(16):2837–2842. doi: 10.1039/c2lc00030j. [DOI] [PubMed] [Google Scholar]
- 119.Jeon JS, Bersini S, Gilardi M, Dubini G, Charest JL, Moretti M, et al. Human 3D vascularized organotypic microfluidic assays to study breast cancer cell extravasation. Proc Natl Acad Sci U S A. 2015;112(1):214–219. doi: 10.1073/pnas.1417115112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Jeon JS, Zervantonakis IK, Chung S, Kamm RD, Charest JL. In vitro model of tumor cell extravasation. PLoS One. 2013;8(2):e56910. doi: 10.1371/journal.pone.0056910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Agliari E, Altavilla M, Barra A, Dello Schiavo L, Katz E. Notes on stochastic (bio)-logic gates: computing with allosteric cooperativity. Sci Rep. 2015;5:9415. doi: 10.1038/srep09415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Businaro L, De Ninno A, Schiavoni G, Lucarini V, Ciasca G, Gerardino A, et al. Cross talk between cancer and immune cells: exploring complex dynamics in a microfluidic environment. Lab Chip. 2013;13(2):229–239. doi: 10.1039/C2LC40887B. [DOI] [PubMed] [Google Scholar]
- 123.Parlato S, De Ninno A, Molfetta R, Toschi E, Salerno D, Mencattini A, et al. 3D microfluidic model for evaluating immunotherapy efficacy by tracking dendritic cell behaviour toward tumor cells. Sci Rep. 2017;7(1):1093. doi: 10.1038/s41598-017-01013-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Fan ZH, Tan W. DNA nanospheres with microfluidics: a promising platform for cancer diagnosis? Nanomedicine (Lond) 2013;8(11):1731–1733. doi: 10.2217/nnm.13.163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Tang Y, Soroush F, Sheffield JB, Wang B, Prabhakarpandian B, Kiani MF. A biomimetic microfluidic tumor microenvironment platform mimicking the EPR effect for rapid screening of drug delivery systems. Sci Rep. 2017;7(1):9359. doi: 10.1038/s41598-017-09815-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Knowlton S, Onal S, Yu CH, Zhao JJ, Tasoglu S. Bioprinting for cancer research. Trends Biotechnol. 2015;33(9):504–513. doi: 10.1016/j.tibtech.2015.06.007. [DOI] [PubMed] [Google Scholar]
- 127.Lee JM, Yeong WY. Design and printing strategies in 3D bioprinting of cell-hydrogels: a review. Adv Healthc Mater. 2016;5(22):2856–2865. doi: 10.1002/adhm.201600435. [DOI] [PubMed] [Google Scholar]
- 128.Gudapati H, Dey M, Ozbolat I. A comprehensive review on droplet-based bioprinting: past, present and future. Biomaterials. 2016;102:20–42. doi: 10.1016/j.biomaterials.2016.06.012. [DOI] [PubMed] [Google Scholar]
- 129.Schiele NR, Corr DT, Huang Y, Raof NA, Xie Y, Chrisey DB. Laser-based direct-write techniques for cell printing. Biofabrication. 2010;2(3):032001. doi: 10.1088/1758-5082/2/3/032001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Ozbolat IT, Hospodiuk M. Current advances and future perspectives in extrusion-based bioprinting. Biomaterials. 2016;76:321–343. doi: 10.1016/j.biomaterials.2015.10.076. [DOI] [PubMed] [Google Scholar]
- 131.Jung JW, Lee JS, Cho DW. Computer-aided multiple-head 3D printing system for printing of heterogeneous organ/tissue constructs. Sci Rep. 2016;6:21685. doi: 10.1038/srep21685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Colosi C, Shin SR, Manoharan V, Massa S, Costantini M, Barbetta A, et al. Microfluidic bioprinting of heterogeneous 3D tissue constructs using low-viscosity bioink. Adv Mater. 2016;28(4):677–684. doi: 10.1002/adma.201503310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Datta P, Ayan B, Ozbolat IT. Bioprinting for vascular and vascularized tissue biofabrication. Acta Biomater. 2017;51:1–20. doi: 10.1016/j.actbio.2017.01.035. [DOI] [PubMed] [Google Scholar]
- 134.Huang TQ, Qu X, Liu J, Chen S. 3D printing of biomimetic microstructures for cancer cell migration. Biomed Microdevices. 2014;16(1):127–132. doi: 10.1007/s10544-013-9812-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Hospodiuk M, Dey M, Sosnoski D, Ozbolat IT. The bioink: a comprehensive review on bioprintable materials. Biotechnol Adv. 2017;35(2):217–239. doi: 10.1016/j.biotechadv.2016.12.006. [DOI] [PubMed] [Google Scholar]
- 136.Zhao Y, Yao R, Ouyang L, Ding H, Zhang T, Zhang K, et al. Three-dimensional printing of Hela cells for cervical tumor model in vitro. Biofabrication. 2014;6(3):035001. doi: 10.1088/1758-5082/6/3/035001. [DOI] [PubMed] [Google Scholar]
- 137.Zhou X, Zhu W, Nowicki M, Miao S, Cui H, Holmes B, et al. 3D bioprinting a cell-laden bone matrix for breast Cancer metastasis study. ACS Appl Mater Interfaces. 2016;8(44):30017–30026. doi: 10.1021/acsami.6b10673. [DOI] [PubMed] [Google Scholar]
- 138.Wang X, Zhang X, Dai X, Li X, Diao J, Xu T. Tumor-like lung cancer model based on 3D bioprinting. 3 Biotech. 2018;8(12):501. doi: 10.1007/s13205-018-1519-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Drost J, Clevers H. Organoids in cancer research. Nat Rev Cancer. 2018;18(7):407–418. doi: 10.1038/s41568-018-0007-6. [DOI] [PubMed] [Google Scholar]
- 140.Dijkstra KK, Cattaneo CM, Weeber F, Chalabi M, van de Haar J, Fanchi LF, et al. Generation of tumor-reactive T cells by co-culture of peripheral blood lymphocytes and tumor organoids. Cell. 2018;174(6):1586–1598. doi: 10.1016/j.cell.2018.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Vaira V, Fedele G, Pyne S, Fasoli E, Zadra G, Bailey D, et al. Preclinical model of organotypic culture for pharmacodynamic profiling of human tumors. Proc Natl Acad Sci U S A. 2010;107(18):8352–8356. doi: 10.1073/pnas.0907676107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Kuznetsova TG, Starodubtseva MN, Yegorenkov NI, Chizhik SA, Zhdanov RI. Atomic force microscopy probing of cell elasticity. Micron. 2007;38(8):824–833. doi: 10.1016/j.micron.2007.06.011. [DOI] [PubMed] [Google Scholar]
- 143.Chaudhuri O, Koshy ST, Branco da Cunha C, Shin JW, Verbeke CS, Allison KH, et al. Extracellular matrix stiffness and composition jointly regulate the induction of malignant phenotypes in mammary epithelium. Nat Mater. 2014;13(10):970–978. doi: 10.1038/nmat4009. [DOI] [PubMed] [Google Scholar]
- 144.Tan JL, Tien J, Pirone DM, Gray DS, Bhadriraju K, Chen CS. Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc Natl Acad Sci U S A. 2003;100(4):1484–1489. doi: 10.1073/pnas.0235407100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Gardel ML, Shin JH, MacKintosh FC, Mahadevan L, Matsudaira P, Weitz DA. Elastic behavior of cross-linked and bundled actin networks. Science. 2004;304(5675):1301–1305. doi: 10.1126/science.1095087. [DOI] [PubMed] [Google Scholar]
- 146.Zhang H, Liu KK. Optical tweezers for single cells. J R Soc Interface. 2008;5(24):671–690. doi: 10.1098/rsif.2008.0052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Hochmuth RM. Micropipette aspiration of living cells. J Biomech. 2000;33(1):15–22. doi: 10.1016/S0021-9290(99)00175-X. [DOI] [PubMed] [Google Scholar]
- 148.Otto O, Rosendahl P, Golfier S, Mietke A, Herbig M, Jacobi A, et al. Real-time deformability cytometry as a label-free indicator of cell function. Conf Proc IEEE Eng Med Biol Soc. 2015;2015:1861–1864. doi: 10.1109/EMBC.2015.7318744. [DOI] [PubMed] [Google Scholar]
- 149.Kennedy BF, Wijesinghe P, Sampson DD. The emergence of optical elastography in biomedicine. Nat Photonics. 2017;11(4):215–221. doi: 10.1038/nphoton.2017.6. [DOI] [Google Scholar]
- 150.Wirtz D. Particle-tracking microrheology of living cells: principles and applications. Annu Rev Biophys. 2009;38:301–326. doi: 10.1146/annurev.biophys.050708.133724. [DOI] [PubMed] [Google Scholar]
- 151.Ophir J, Cespedes I, Ponnekanti H, Yazdi Y, Li X. Elastography: a quantitative method for imaging the elasticity of biological tissues. Ultrason Imaging. 1991;13(2):111–134. doi: 10.1177/016173469101300201. [DOI] [PubMed] [Google Scholar]
- 152.Manduca A, Oliphant TE, Dresner MA, Mahowald JL, Kruse SA, Amromin E, et al. Magnetic resonance elastography: non-invasive mapping of tissue elasticity. Med Image Anal. 2001;5(4):237–254. doi: 10.1016/S1361-8415(00)00039-6. [DOI] [PubMed] [Google Scholar]
- 153.Antonacci G, Braakman S. Biomechanics of subcellular structures by non-invasive Brillouin microscopy. Sci Rep. 2016;6:37217. doi: 10.1038/srep37217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Scarcelli G, Polacheck WJ, Nia HT, Patel K, Grodzinsky AJ, Kamm RD, et al. Noncontact three-dimensional mapping of intracellular hydromechanical properties by Brillouin microscopy. Nat Methods. 2015;12(12):1132–1134. doi: 10.1038/nmeth.3616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Antonacci G, de Turris V, Rosa A, Ruocco G. Background-deflection Brillouin microscopy reveals altered biomechanics of intracellular stress granules by ALS protein FUS. Commun Biol. 2018;1:139. doi: 10.1038/s42003-018-0148-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Antonacci G, Foreman MR, Paterson C, Torok P. Spectral broadening in Brillouin imaging. Appl Phys Lett. 2013;103(22):221105. doi: 10.1063/1.4836477. [DOI] [Google Scholar]
- 157.Antonacci G, Pedrigi RM, Kondiboyina A, Mehta VV, de Silva R, Paterson C, et al. Quantification of plaque stiffness by Brillouin microscopy in experimental thin cap fibroatheroma. J R Soc Interface. 2015;12(112). [DOI] [PMC free article] [PubMed]
- 158.Bao G and Suresh S. Cell and molecular mechanics of biological materials. Nat Mater. 2003;2(11):715–25. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.