Abstract
In the present study, we aimed at determining the potential role of rs12917 polymorphism of the O-6-methylguanine-DNA methyltransferase (MGMT) gene in the occurrence of cancer. Based on the available data from the online database, we performed an updated meta-analysis. We retrieved 537 articles from our database research and finally selected a total of 54 case–control studies (21010 cases and 34018 controls) for a series of pooling analyses. We observed an enhanced risk in cancer cases compared with controls, using the genetic models T/T compared with C/C (P-value of association test <0.001; odds ratio (OR) = 1.29) and T/T compared with C/C+C/T (P<0.001; OR = 1.32). We detected similar positive results in the subgroups ‘Caucasian’, and ‘glioma’ (all P<0.05; OR > 1). However, we detected negative results in our analyses of most of the other subgroups (P>0.05). Begg’s and Egger’s tests indicated that the results were free of potential publication bias, and sensitivity analysis suggested the stability of the pooling results. In summary, the T/T genotype of MGMT rs12917 is likely to be linked to an enhanced susceptibility to cancer overall, especially glioma, in the Caucasian population.
Keywords: MGMT, polymorphism, Cancer, meta-analysis
Introduction
In humans, the O-6-methylguanine-DNA methyltransferase (MGMT) protein, encoded by the MGMT gene located on chromosome 10 (10q26) [1], is involved in the DNA repair process [2,3]. By means of methyl transfer, MGMT removes alkylating agents from the DNA direct reversal repair pathway and thus repairs the DNA [2,3]. Two potential functional polymorphisms have been identified in the MGMT gene, namely rs12917 (Leu84Phe) and rs2308321 (Ile143Val) [4,5]. In addition, the promoter methylation status of the gene is reportedly correlated with several clinical diseases, such as glioblastoma [6,7], gastric cancer [8], and oral carcinoma [9].
Both genetic and environmental factors contribute to the occurrence and progression of clinical cancers [10,11]. A number of studies have been conducted on the potential genetic effect of MGMT rs12917 polymorphism on its susceptibility to cancer, but the results were inconclusive. Before 2013, only three relative meta-analyses investigated the potential role of this polymorphism in the overall risk for cancer [12–14]. Based on the currently available data, we performed an updated meta-analysis to reassess the genetic relationship between MGMT rs12917 polymorphism and cancer risk. We enrolled a total of 54 case–control studies for the study.
Materials and methods
Database searching strategy
To identify potential publications, we searched four online electronic databases (PubMed, Embase, Cochrane Library, and WANFANG) up through August 2018. We used the terms ‘MeSH (Medical Subject Headings)’ and ‘Entry Terms’ to search PubMed and Cochrane Library, and ‘Emtree’ and ‘Synonyms’ for Embase. The search string we used for PubMed was as follows: (((((((((((((((O(6)-Methylguanine-DNA Methyltransferase [MeSH Terms]) OR Methylated-DNA-Protein-Cysteine S-Methyltransferase) OR Methylated DNA Protein Cysteine S Methyltransferase) OR S-Methyltransferase, Methylated-DNA-Protein- Cysteine) OR O(6)-Methylguanine Methyltransferase) OR O(6)-Alkylguanine-DNA Alkyltransferase) OR O(6)-MeG-DNA Methyltransferase) OR O(6)-Methylguanine DNA Transmethylase) OR Guanine-O(6)-Alkyltransferase) OR O(6)-AGT) OR DNA Repair Methyltransferase II) OR DNA Repair Methyltransferase I) OR MGMT)) AND ((((((((Polymorphism, Genetic [MeSH Terms]) OR Polymorphisms, Genetic) OR Genetic Polymorphisms) OR Genetic Polymorphism) OR Polymorphism (Genetics)) OR Polymorphisms (Genetics)) OR Polymorphism) OR Polymorphisms)) AND ((((((((((((((((((Neoplasms [MeSH Terms]) OR Neoplasia) OR Neoplasias) OR Neoplasm) OR Tumors) OR Tumor) OR Cancer) OR Cancers) OR Malignant Neoplasms) OR Malignant Neoplasm) OR Neoplasm, Malignant) OR Neoplasms, Malignant) OR Malignancy) OR Malignancies) OR Benign Neoplasms) OR Neoplasms, Benign) OR Benign Neoplasm) OR Neoplasm, Benign).
Article screening strategy
We designed our inclusion and exclusion criteria according to Patient, Intervention, Comparison and Outcome and Study design (PICOS) principles. We ruled out duplicates and screened improper articles. Exclusion criteria were as follows: (P), non-cancer patients; (I), other variants, gene expression or methylation; (C), lack of study controls or P-value of Hardy–Weinberg equilibrium (HWE) <0.05; (O), lack of full genotype frequency data; (S), review, meta, poster, or meeting abstract. Eligible articles had to be designed as case–control studies, targetting the genetic relationship between MGMT rs12917 and cancer risk and containing the full genotype (C/C, C/T, T/T) frequencies in both cancer cases and negative controls.
Data extraction and quality assessment
After extracting usable data, we listed the basic information in tables. We assessed methodological quality via the Newcastle–Ottawa Scale (NOS) [15]. High-quality articles with NOS score > 5 were regarded as eligible and included in our statistical analysis.
Statistical analysis
We used STATA software version 12.0-SE (StataCorp, College Station, TX) to perform our analyses. We first assessed the inter-study heterogeneity using Cochran’s Q statistic and the I2 test. A P-value of Cochran’s Q statistic < 0.1 or I2 value > 50% was considered to show a high level of heterogeneity. We thus used the DerSimonian–Laird association test with a random-effects model. Otherwise, we used the Mantel–Haenszel association test with a fixed-effects model. The P-value of association test, summary odds ratio (OR), along with the corresponding 95% confidence interval (CI) could be obtained for the allele (T compared with C), homozygous (T/T compared with C/C), recessive (T/T compared with C/C+C/T), heterozygous (C/T compared with C/C), dominant (C/T+T/T compared with C/C), and carrier (T compared with C) models.
We performed subgroup analyses by race, cancer type, and control source. Additionally, we assessed possible publication bias by means of Begg’s and Egger’s tests and evaluated the robustness of the results through sensitivity analysis.
Results
Eligible case–control studies
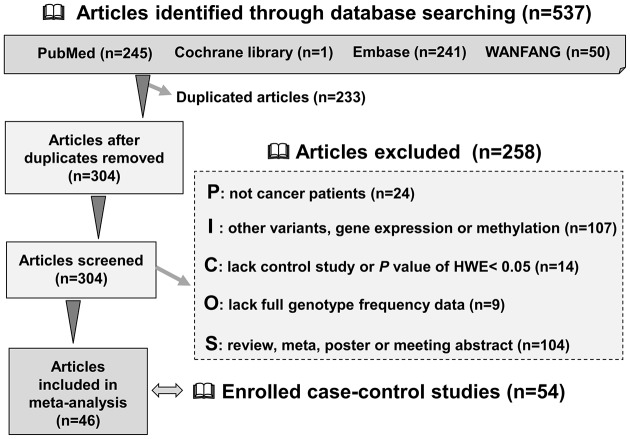
Figure 1 depicts the flowchart for the identification of eligible case–control studies. We initially obtained a total of 537 articles by searching four databases, including PubMed (245 articles), Cochrane Library (1 article), Embase (241 articles), and WANFANG (50 articles). We then excluded 233 duplicates plus another 258 articles based strictly on our screening strategy. Finally, we identified 46 full-text articles for inclusion [4,5,16–59]. After data extraction and quality evaluation, we enrolled a total of 54 case–control studies free of poor quality (all NOS score > 5) in our pooling analyses. The basic information and genotype frequency distribution are presented in Supplementary Table S1 and Table 1, respectively.
Figure 1. Flowchart for the identification of eligible case–control studies.
Table 1. Genotype and allele frequency of MGMT rs12917 in the enrolled case–control studies.
| Authors | Year | Genotype (case) | Allele (case) | Cancer type (case) | Genotype (control) | Allele (control) | HWE (control) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| C/C | C/T | T/T | C | T | C/C | C/T | T/T | C | T | χ2 | P | |||
| Agalliu et al. [16] | 2010 | 949 | 269 | 32 | 2167 | 333 | Prostate cancer1 | 916 | 298 | 23 | 2130 | 344 | 0.05 | 0.83 |
| 106 | 35 | 6 | 247 | 47 | Prostate cancer2 | 60 | 20 | 1 | 140 | 22 | 0.22 | 0.64 | ||
| Akbari et al. [17] | 2009 | 142 | 53 | 1 | 337 | 55 | Esophageal cancer | 185 | 63 | 2 | 433 | 67 | 1.84 | 0.17 |
| Betti et al. [18] | 2011 | 95 | 36 | 2 | 226 | 40 | MPM3 | 179 | 64 | 8 | 422 | 80 | 0.59 | 0.44 |
| 50 | 17 | 1 | 117 | 19 | MPM4 | 32 | 12 | 0 | 76 | 12 | 1.10 | 0.29 | ||
| Bye et al. [19] | 2011 | 225 | 111 | 10 | 561 | 131 | Esophageal cancer1 | 300 | 155 | 14 | 755 | 183 | 1.28 | 0.26 |
| 120 | 65 | 11 | 305 | 87 | Esophageal cancer5 | 294 | 116 | 13 | 704 | 142 | 1.28 | 0.26 | ||
| Chae et al. [20] | 2006 | 344 | 84 | 4 | 772 | 92 | Lung cancer | 341 | 81 | 10 | 763 | 101 | 3.65 | 0.06 |
| Chuang et al. [21] | 2011 | 1105 | 307 | 43 | 2517 | 393 | Head and neck cancer | 2256 | 823 | 81 | 5335 | 985 | 0.33 | 0.57 |
| Doecke et al. [22] | 2008 | 416 | 136 | 14 | 968 | 164 | Esophageal cancer | 1029 | 281 | 27 | 2339 | 335 | 2.25 | 0.13 |
| Felini et al. [23] | 2007 | 289 | 84 | 6 | 662 | 96 | Glioma | 369 | 84 | 6 | 822 | 96 | 0.24 | 0.63 |
| Feng et al. [24] | 2008 | 96 | 58 | 47 | 250 | 152 | Esophageal cancer | 87 | 85 | 29 | 259 | 143 | 1.20 | 0.27 |
| Gu et al. [25] | 2009 | 152 | 60 | 2 | 364 | 64 | Melanoma | 168 | 43 | 1 | 379 | 45 | 1.01 | 0.31 |
| Hall et al. [26] | 2007 | 548 | 193 | 38 | 1289 | 269 | UADT | 730 | 281 | 23 | 1741 | 327 | 0.44 | 0.51 |
| Han et al. [27] | 20061 | 344 | 82 | 8 | 770 | 98 | Endometrial cancer | 822 | 242 | 21 | 1886 | 284 | 0.42 | 0.52 |
| Han et al. [28] | 20062 | 964 | 279 | 33 | 2207 | 345 | Breast cancer | 1,306 | 382 | 26 | 2994 | 434 | 0.10 | 0.75 |
| Hu et al. [29] | 2013 | 389 | 130 | 24 | 908 | 178 | Glioma | 405 | 84 | 6 | 894 | 96 | 0.48 | 0.49 |
| Hu et al. [4] | 2007 | 418 | 77 | 5 | 913 | 87 | Lung cancer | 421 | 93 | 3 | 935 | 99 | 0.78 | 0.38 |
| Huang et al. [30] | 2017 | 76 | 12 | 2 | 164 | 16 | Glioma | 75 | 14 | 1 | 164 | 16 | 0.14 | 0.71 |
| Huang et al. [31] | 2007 | 372 | 156 | 11 | 900 | 178 | Cervical cancer | 592 | 198 | 10 | 1382 | 218 | 2.12 | 0.15 |
| Huang et al. [32] | 2010 | 151 | 25 | 0 | 327 | 25 | Oral cancer | 89 | 21 | 0 | 199 | 21 | 1.22 | 0.27 |
| Huang et al. [33] | 20051 | 190 | 82 | 8 | 462 | 98 | Gastric cancer | 279 | 99 | 9 | 657 | 117 | 0.00 | 0.95 |
| Huang et al. [34] | 20052 | 386 | 117 | 11 | 889 | 139 | Head and neck cancer | 529 | 204 | 21 | 1262 | 246 | 0.06 | 0.80 |
| Inoue et al. [35] | 2003 | 55 | 18 | 0 | 128 | 18 | Primary brain cancer | 160 | 55 | 9 | 375 | 73 | 2.24 | 0.13 |
| Kiczmer [36] | 2018 | 49 | 11 | 9 | 109 | 29 | Head and neck cancer | 168 | 66 | 5 | 402 | 76 | 0.25 | 0.61 |
| Kietthubthew et al. [37] | 2006 | 84 | 21 | 1 | 189 | 23 | Oral cancer | 130 | 33 | 1 | 293 | 35 | 0.50 | 0.48 |
| Li et al. [38] | 2005 | 132 | 34 | 1 | 298 | 36 | Bladder cancer | 173 | 28 | 3 | 374 | 34 | 2.11 | 0.15 |
| Liu et al. [39] | 20021 | 53 | 7 | 0 | 113 | 7 | Lung cancer | 89 | 11 | 0 | 189 | 11 | 0.34 | 0.56 |
| Liu et al. [40] | 20022 | 21 | 3 | 0 | 45 | 3 | Gynecologic tumor | 89 | 11 | 0 | 189 | 11 | 0.34 | 0.56 |
| 26 | 8 | 0 | 60 | 8 | Digestive system cancer | 89 | 11 | 0 | 189 | 11 | 0.34 | 0.56 | ||
| Liu et al. [41] | 2006 | 82 | 16 | 2 | 180 | 20 | Esophageal cancer | 57 | 8 | 0 | 122 | 8 | 0.28 | 0.60 |
| Liu et al. [42] | 2009 | 299 | 62 | 8 | 660 | 78 | Glioma | 267 | 89 | 7 | 623 | 103 | 0.02 | 0.89 |
| Loh et al. [43] | 2011 | 146 | 37 | 5 | 329 | 47 | Cancer | 894 | 212 | 14 | 2000 | 240 | 0.13 | 0.72 |
| Lu et al. [44] | 2006 | 142 | 45 | 4 | 329 | 53 | Gastric cancer | 186 | 59 | 6 | 431 | 71 | 0.26 | 0.61 |
| McKean-Cowdin et al. [45] | 2009 | 774 | 204 | 20 | 1752 | 244 | Glioblastoma | 1,480 | 453 | 35 | 3413 | 523 | 0.00 | 0.96 |
| O’Mara et al. [46] | 2011 | 889 | 261 | 23 | 2039 | 307 | Endometrial cancer6 | 810 | 270 | 19 | 1890 | 308 | 0.42 | 0.52 |
| 278 | 108 | 11 | 664 | 130 | Endometrial cancer7 | 296 | 103 | 7 | 695 | 117 | 0.33 | 0.57 | ||
| Palli et al. [47] | 2010 | 210 | 77 | 4 | 497 | 85 | Gastric cancer | 395 | 131 | 11 | 921 | 153 | 0.00 | 0.97 |
| Rajaraman et al. [48] | 2010 | 265 | 77 | 9 | 607 | 95 | Glioma | 348 | 117 | 12 | 813 | 141 | 0.33 | 0.57 |
| 102 | 23 | 4 | 227 | 31 | Meningioma | 348 | 117 | 12 | 813 | 141 | 0.33 | 0.57 | ||
| 52 | 12 | 2 | 116 | 16 | Acoustic neuroma | 348 | 117 | 12 | 813 | 141 | 0.33 | 0.57 | ||
| Ritchey et al. [49] | 2005 | 123 | 36 | 2 | 282 | 40 | Prostate cancer | 213 | 32 | 1 | 458 | 34 | 0.03 | 0.86 |
| Shah et al. [50] | 2012 | 64 | 26 | 2 | 154 | 30 | Esophageal cancer | 57 | 20 | 0 | 134 | 20 | 1.72 | 0.19 |
| Shen et al. [51] | 2005 | 778 | 265 | 21 | 1821 | 307 | Breast cancer | 824 | 263 | 20 | 1911 | 303 | 0.03 | 0.85 |
| Shen et al. [52] | 2007 | 432 | 112 | 11 | 976 | 134 | NHL | 373 | 110 | 12 | 856 | 134 | 1.27 | 0.26 |
| Shi et al. [53] | 2011 | 253 | 47 | 3 | 553 | 53 | AML | 459 | 91 | 4 | 1009 | 99 | 0.05 | 0.83 |
| Stern et al. [54] | 2007 | 251 | 40 | 1 | 542 | 42 | Colorectal cancer | 959 | 194 | 13 | 2112 | 220 | 0.81 | 0.37 |
| Tranah et al. [55] | 2006 | 147 | 33 | 6 | 327 | 45 | Colorectal cancer8 | 1,634 | 471 | 32 | 3739 | 535 | 0.09 | 0.77 |
| 204 | 47 | 6 | 455 | 59 | Colorectal cancer9 | 330 | 93 | 6 | 753 | 105 | 0.04 | 0.85 | ||
| Wang et al. [5] | 2006 | 832 | 259 | 30 | 1923 | 319 | Lung cancer | 872 | 272 | 19 | 2016 | 310 | 0.18 | 0.67 |
| Yang et al. [56] | 2009 | 33 | 14 | 1 | 80 | 16 | NHL | 289 | 58 | 5 | 636 | 68 | 1.10 | 0.29 |
| Zhang et al. [57] | 2008 | 352 | 53 | 1 | 757 | 55 | Biliary track cancer | 631 | 144 | 7 | 1406 | 158 | 0.15 | 0.70 |
| Zhang et al. [58] | 2010 | 563 | 151 | 7 | 1277 | 165 | Head and neck cancer | 933 | 284 | 17 | 2150 | 318 | 0.78 | 0.38 |
| Zienolddiny et al. [59] | 2006 | 189 | 102 | 13 | 480 | 128 | Lung cancer | 247 | 106 | 10 | 600 | 126 | 0.12 | 0.73 |
Abbreviations: AML, acute myeloid leukemia; MPM, malignant mesothelioma; NHL, non-Hodgkin’s lymphoma; UADT, upper aerodigestive tract.
1Data from Caucasian population. 2Data from African population. 3With population-based control. 4With hospital-based control. 5Data from mixed population. 6Data from Australia. 7Data from Poland. 8With controls from Nurses’ Health Study (NHS). 9With controls from Physicians’ Health Study (PHS) cohorts
Meta-analysis data
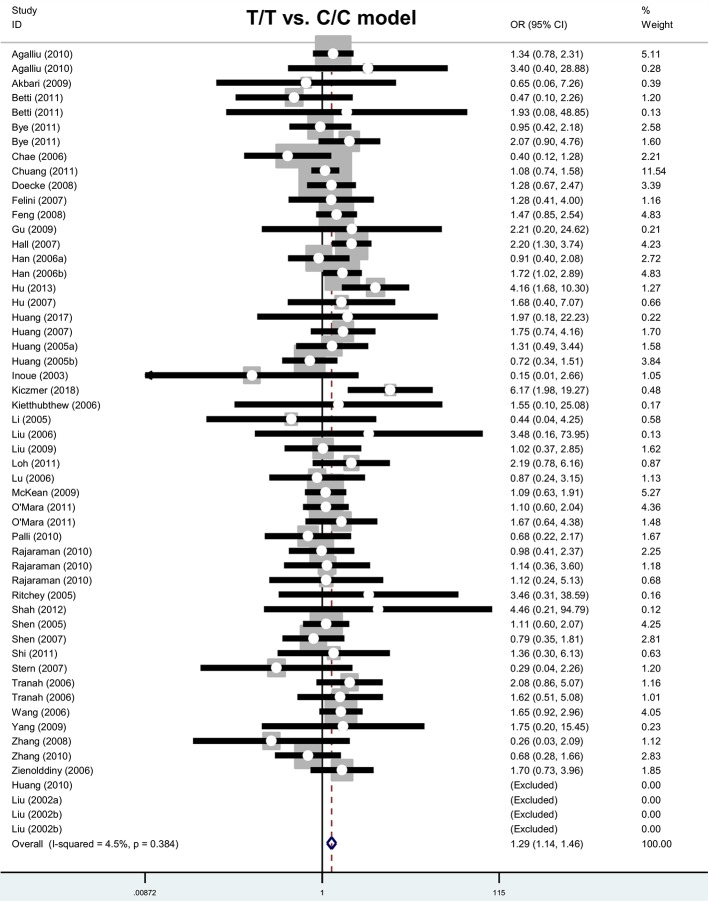
First, we studied the association between the MGMT rs12917 polymorphism and cancer risk via an overall meta-analysis. As shown in Table 2, we included a total of 54 case–control studies with 21010 cases and 34018 controls under the genetic models of allele T compared with C, C/T compared with C/C, C/T+T/T compared with C/C, and carrier T compared with C; meanwhile, we included 50 studies with 20716 cases and 33608 controls under the models of T/T compared with C/C and T/T compared with C/C+C/T. For the homozygous, recessive and carrier genetic models, we performed a Mantel–Haenszel association test with a fixed-effects model, and we observed no high degree of heterogeneity (Table 2; all P-values of heterogeneity > 0.1; I2 < 50%). For other models (all P-values of heterogeneity <0.001), we performed a DerSimonian–Laird association test with a random-effects model. Pooling data (Table 2) indicated an increased risk of cancer in cases compared with controls for the T/T compared with C/C (P-value of association test <0.001; OR = 1.29) and T/T compared with C/C+C/T (P<0.001; OR = 1.32) genetic models. Nevertheless, we failed to detect any statistical difference between cancer cases and negative controls under other genetic models (Table 2; all P>0.05). Forest plot data are shown in Figure 2 and Supplementary Figures S1–S5; they revealed that the T/T genotype of the MGMT rs12917 polymorphism was likely to be associated with an increased susceptibility to cancer.
Table 2. Meta-analysis of the association between MGMT rs12917 and cancer susceptibility.
| Models | Sample size | Heterogeneity | Association | |||||
|---|---|---|---|---|---|---|---|---|
| Study | Case | Control | I2 | P | Fixed/random | P | OR (95% CI) | |
| Allele T compared with C | 54 | 21010 | 34018 | 50.1% | <0.001 | Random | 0.354 | - |
| T/T compared with C/C | 50 | 20716 | 33608 | 4.5% | 0.384 | Fixed | <0.001 | 1.29 (1.14–1.46) |
| T/T compared with C/C+C/T | 50 | 20716 | 33608 | 3.2% | 0.410 | Fixed | <0.001 | 1.32 (1.17–1.49) |
| C/T compared with C/C | 54 | 21010 | 34018 | 46.1% | <0.001 | Random | 0.442 | - |
| C/T+T/T compared with C/C | 54 | 21010 | 34018 | 47.7% | <0.001 | Random | 0.976 | - |
| Carrier T compared with C | 54 | 21010 | 34018 | 20.0% | 0.104 | Fixed | 0.642 | - |
-, OR (95% CI) data were not provided, when P-value of association >0.05.
Figure 2. Forest plot of meta-analysis (T/T compared with C/C model).
Subgroup analysis data
Next, we carried out four subgroup analyses by race, cancer type, and control source. For the T/T compared with C/C model (Table 3), the association test data showed an increased cancer risk in the subgroups ‘Caucasian’ (P<0.001; OR = 1.35), ‘glioma’ (P=0.022; OR = 1.70), ‘population-based control (PB)’ (P<0.001; OR = 1.32) and ‘hospital-based control (HB)’ (P<0.030; OR = 1.39). Figure 3 and Supplementary Figures S6–S7 present the forest plot data.
Table 3. Data of subgroup analysis under T/T compared with C/C model.
| Factor | Subgroup | Sample size | Heterogeneity | Association | ||||
|---|---|---|---|---|---|---|---|---|
| Study | Case | Control | I2 | P | P | OR (95% CI) | ||
| Race | Caucasian | 27 | 13158 | 20678 | 0.0% | 0.573 | <0.001 | 1.35 (1.15, 1.58) |
| African | 3 | 796 | 1104 | 0.0% | 0.538 | 0.560 | - | |
| Asian | 16 | 4031 | 6152 | 28.6% | 0.136 | 0.088 | - | |
| Cancer type | Urinary system cancer | 4 | 1725 | 1768 | 0.0% | 0.526 | 0.174 | - |
| Esophageal cancer | 8 | 2131 | 3907 | 0.0% | 0.781 | 0.069 | - | |
| Lung cancer | 4 | 2357 | 2475 | 40.7% | 0.167 | 0.155 | - | |
| Head and neck cancer | 14 | 5863 | 10581 | 39.5% | 0.064 | 0.138 | - | |
| Gastric cancer | 3 | 762 | 1175 | 0.0% | 0.692 | 0.891 | - | |
| Blood cancer | 3 | 906 | 1401 | 0.0% | 0.702 | 0.882 | - | |
| Colorectal cancer | 3 | 735 | 3732 | 38.5% | 0.197 | 0.416 | - | |
| Brain cancer | 9 | 2998 | 5030 | 17.4% | 0.288 | 0.106 | - | |
| Glioma | 5 | 1735 | 1884 | 37.9% | 0.168 | 0.022 | 1.70 (1.08, 2.68) | |
| Control source | PB | 39 | 16526 | 26488 | 6.3% | 0.358 | <0.001 | 1.32 (1.14, 1.52) |
| HB | 8 | 2482 | 4148 | 3.2% | 0.405 | 0.030 | 1.39 (1.03, 1.86) | |
-, OR (95% CI) data were not provided, when P-value of association > 0.05.
Figure 3. Forest plot of subgroup analysis by race (T/T compared with C/C model).
For the T/T compared with C/C+C/T model (Table 4), we also observed positive correlations in the subgroups ‘Caucasian’ (P<0.001; OR = 1.37), ‘Asian’ (P=0.036; OR = 1.37), ‘glioma’ (P=0.026; OR = 1.68), ‘PB’ (P<0.001; OR = 1.32), and ‘HB’ (P=0.004; OR = 1.52). Supplementary Figures S8–S10 present the forest plot data.
Table 4. Data of subgroup analysis under T/T compared with C/C+C/T model.
| Factor | Subgroup | Sample size | Heterogeneity | Association | ||||
|---|---|---|---|---|---|---|---|---|
| Study | Case | Control | I2 | P | P | OR (95% CI) | ||
| Race | Caucasian | 27 | 13158 | 20678 | 0.0% | 0.528 | <0.001 | 1.37 (1.17, 1.60) |
| African | 3 | 796 | 1104 | 0.0% | 0.542 | 0.535 | - | |
| Asian | 16 | 4031 | 6152 | 27.2% | 0.150 | 0.036 | 1.37 (1.02, 1.83) | |
| Cancer type | Urinary system cancer | 4 | 1725 | 1768 | 0.0% | 0.527 | 0.152 | - |
| Esophageal cancer | 8 | 2131 | 3907 | 0.0% | 0.725 | 0.021 | - | |
| Lung cancer | 4 | 2357 | 2475 | 40.0% | 0.467 | 0.174 | - | |
| Head and neck cancer | 14 | 5863 | 10581 | 37.5% | 0.077 | 0.064 | - | |
| Gastric cancer | 3 | 762 | 1175 | 0.0% | 0.718 | 0.815 | - | |
| Blood cancer | 3 | 906 | 1401 | 0.0% | 0.769 | 0.901 | - | |
| Colorectal cancer | 3 | 735 | 3732 | 39.6% | 0.191 | 0.344 | - | |
| Brain cancer | 9 | 2998 | 5030 | 3.0% | 0.410 | 0.088 | - | |
| Glioma | 5 | 1735 | 1884 | 23.7% | 0.263 | 0.026 | 1.68 (1.07, 2.65) | |
| Control source | PB | 39 | 16526 | 26488 | 2.5% | 0.426 | <0.001 | 1.32 (1.15, 1.52) |
| HB | 8 | 2482 | 4148 | 11.0% | 0.344 | 0.004 | 1.52 (1.14, 2.03) | |
-, OR (95% CI) data was not provided, when P-value of association > 0.05.
We did not detect positive results for the other genetic models (Supplementary Tables S2–S5; P<0.05) except for the subgroups ‘colorectal cancer’ (Supplementary Table S3; P=0.041; OR = 0.79), ‘HB’ (Supplementary Table S3; P=0.027; OR = 0.86) under the C/T compared with C/C model; and the subgroup ‘head and neck cancer’ (Supplementary Table S5; P=0.020; OR = 0.92) under the carrier T compared with C model. Thus, the T/T genotype of MGMT rs12917 may have been associated with an increased risk of cancer in cases, especially the glioma cases, in the Caucasian population.
Publication bias and sensitivity analysis
Begg’s and Egger’s tests indicated that results were free of possible publication bias (Supplementary Table S6; P>0.05 for Begg’s test, >0.05 for Egger’s test). A Begg’s funnel plot with pseudo–95% confidence limits under the T/T compared with C/C model is shown in Figure 4. In addition, we observed the same stable results in our subsequent sensitivity analysis; data from this analysis under the homozygous model (Figure 5) are presented as an example.
Figure 4. Begg’s funnel plot with pseudo-95% confidence limits (T/T compared with C/C model).
Figure 5. Sensitivity analysis result (T/T compared with C/C model).
Discussion
We observed conflicting conclusions about the genetic role of MGMT rs12917 polymorphism in its susceptibility to different cancers. For instance, the polymorphism seems to be associated with the risk of esophageal cancer in the Chinese population [41], but not in the Kashmiri population [50]. This merits a quantitative synthesis via the meta-analytic approach. Although there were already three meta-analyses of the MGMT rs12917 polymorphism and its role in the overall risk for cancer [12–14], expanding the sample size and employing a distinct analysis strategy led to better results in our updated pooling analysis.
We did our best to gather candidate articles from four online databases. After screening them based on strict inclusion and exclusion criteria, we enrolled only the case–control studies that were of high quality and those that followed HWE. We ultimately included a total of 46 articles in our updated meta-analysis. After data extraction, we enrolled 54 case–control studies with 21010 cases and 34018 controls in the meta-analysis. We used the carrier, allele, homozygous, recessive, heterozygous, and dominant genetic models, and also confirmed the stability of the statistical results via sensitivity analysis.
In 2010, Zhong et al. [12] performed the first meta-analysis on this topic, reviewing 28 case–control studies from 26 articles [4,5,20,22,23,26–28,31,33–35,37,38,42,45,49,51,52,54,55,59–63]. Another 24 case–control studies [16–19,21,24,25,29,30,32,36,39–41,43,44,46–48,50,53,56–58] were included in our study. We excluded three studies not in-line with the HWE principle [61–63] and one that focussed only on colorectal adenomatous or hyperplastic polyps but not on colorectal cancer [60]. In 2013, Du et al. [14] enrolled 41 case–control studies with 16643 cancer cases and 26720 negative controls from 37 articles [5,16–20,22–24,26–28,31–34,37–41,43,44,46,47,49–59,64] in a meta-analysis. We excluded one of these studies [64] from our meta-analysis because it did not meet the requirement of full genotype frequency in both case and control groups. Finally, we enrolled another ten case–control studies [4,21,25,29,30,35,36,42,45,48]. In addition, when compared with another meta-analysis of Liu et al. (2013) [13], which consisted of 44 case–control studies from 37 articles [4,5,16,17,19,20,22,23,25–27,31–33,35,37,38,42,43,45–47,49,51,52,54–63,65,66], we excluded four studies that were not in HWE [61–63,66], one that did not analyze colorectal cancer [60], and one that included other genetic variants [65]. We also added another 15 new case–control studies [18,21,24,28–30,34,36,39–41,44,48,50,53] for the analysis.
Our updated pooling analysis data demonstrated that cases had an overall enhanced risk for cancer when compared with negative controls under the T/T compared with C/C and T/T compared with C/C+C/T genetic models, especially in the European-descended population, which is partly consistent with the data of previous analyses [12–14]. Moreover, we observed that the MGMT rs12917 polymorphism is likely to be associated with the susceptibility to glioma, which is partly in-line with the two studies on the association between DNA repair gene polymorphisms and glioma risk [67,68]. Nevertheless, owing to the limitation of sample size, the previous three meta-analyses of the overall risk for cancer did not conduct subgroup analyses of ‘glioma’ [12–14].
Some of the limitations to our meta-analysis are as follows:
Although the sample sizes enrolled were quite large (21010 cases and 34018 controls), genotype data were very limited in many subgroup analyses. For instance, we used only three case–control studies in our analyses of the subgroups for gastric [33,44,47], blood [52,53,56], and colorectal [54,55] cancers. Even for the subgroup analysis of ‘glioma’, with positive correlations under the T/T compared with C/C and T/T compared with C/C+C/T models, only five case–control studies [23,29,30,42,48] were included.
We did not investigate the genetic effects of the MGMT rs12917 polymorphism in combination with other variants, such as rs2308321 of MGMT, rs25487 of X-ray cross-complementing group 1 (XRCC1), and rs13181 of xeroderma pigmentosum complementation group D (XPD), in certain specific cancers.
We extracted certain demographic information such as the mean age at diagnosis and the sex of subject, but not other confounding factors such as lifestyle and clinical features. Moreover, we did not perform the relevant stratified meta-analyses due to lack of sufficient usable data.
We detected significant heterogeneity amongst studies under the allele T compared with C, C/T compared with C/C, C/T+T/T compared with C/C, and carrier T compared with C genetic models. Complicating factors such as race and cancer type may be sources of inter-study heterogeneity. For instance, we detected decreased levels of heterogeneity in the ‘Caucasian’ and ‘esophageal cancer’ subgroups. Although we observed a positive conclusion in the ‘glioma’ subgroup, we failed to detect reduced inter-study heterogeneity. Only five case–control studies [23,29,30,42,48] were enrolled.
There may be other undetected or unpublished articles containing potential eligible case–controls in other geographical locations or languages; in other words, our study may suffer from selection bias.
Last but most important, our meta-analysis found a positive conclusion between MGMT rs12917 and the risk of cancer in general for the T/T compared with C/C and T/T compared with C/C+C/T models. Considering the distinct etiopathogenesis or pathogenesis of different kinds of cancers, more studies of large-scale populations of different ethnicities are required for a more scientific elucidation of MGMT rs12917’s functional role in each particular cancer type.
To sum up, our updated pooling analysis offered additional evidence that MGMT rs12917 polymorphism is likely to be associated with an enhanced susceptibility to cancer overall, especially glioma, in the Caucasian population.
Supporting information
Figure S1.
Figure S2.
Figure S3.
Figure S4.
Figure S5.
Figure S6.
Figure S4.
Figure S8.
Figure S9.
Figure S10.
Table S1. Basic information of included studies.
Table S2. Data of subgroup analysis under allele T vs. C model.
Table S3. Data of subgroup analysis under C/T vs. C/C model.
Table S4. Data of subgroup analysis under C/T+T/T vs. C/C model.
Table S5. Data of subgroup analysis under carrier T vs. C model.
Table S6. Publication bias result.
Abbreviations
- HB
hospital-based control
- HWE
Hardy–Weinberg equilibrium
- MeSH
Medical Subject Heading
- MGMT
O-6-methylguanine-DNA methyltransferase
- NOS
Newcastle–Ottawa scale
- OR
odds ratio
- PB
population-based control
Author contribution
Z.S. and H.W. conceived and designed the study. Z.S. and M.K. were responsible for the data extraction and statistical analysis. Z.S. wrote the manuscript and H.W. revised the manuscript.
Competing interests
The authors declare that there are no competing interests associated with the manuscript.
Funding
The authors declare that there are no sources of funding to be acknowledged.
References
- 1.Natarajan A.T., Vermeulen S., Darroudi F., Valentine M.B., Brent T.P., Mitra S. et al. (1992) Chromosomal localization of human O6-methylguanine-DNA methyltransferase (MGMT) gene by in situ hybridization. Mutagenesis 7, 83–85 10.1093/mutage/7.1.83 [DOI] [PubMed] [Google Scholar]
- 2.Christmann M., Verbeek B., Roos W.P. and Kaina B. (2011) O(6)-Methylguanine-DNA methyltransferase (MGMT) in normal tissues and tumors: enzyme activity, promoter methylation and immunohistochemistry. Biochim. Biophys. Acta 1816, 179–190 [DOI] [PubMed] [Google Scholar]
- 3.Nikolova T., Roos W.P., Kramer O.H., Strik H.M. and Kaina B. (2017) Chloroethylating nitrosoureas in cancer therapy: DNA damage, repair and cell death signaling. Biochim. Biophys. Acta 1868, 29–39 [DOI] [PubMed] [Google Scholar]
- 4.Hu Z., Wang H., Shao M., Jin G., Sun W., Wang Y. et al. (2007) Genetic variants in MGMT and risk of lung cancer in Southeastern Chinese: a haplotype-based analysis. Hum. Mutat. 28, 431–440 10.1002/humu.20462 [DOI] [PubMed] [Google Scholar]
- 5.Wang L., Liu H., Zhang Z., Spitz M.R. and Wei Q. (2006) Association of genetic variants of O6-methylguanine-DNA methyltransferase with risk of lung cancer in non-Hispanic Whites. Cancer Epidemiol. Biomarkers Prev. 15, 2364–2369 10.1158/1055-9965.EPI-06-0437 [DOI] [PubMed] [Google Scholar]
- 6.Staedtke V., a Dzaye O.D. and Holdhoff M. (2016) Actionable molecular biomarkers in primary brain tumors. Trends Cancer 2, 338–349 10.1016/j.trecan.2016.06.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Binabaj M.M., Bahrami A., ShahidSales S., Joodi M., Joudi Mashhad M. and Hassanian S.M. (2018) The prognostic value of MGMT promoter methylation in glioblastoma: a meta-analysis of clinical trials, J. Cell. Physiol. 233, 378–386, 10.1002/jcp.25896 [DOI] [PubMed] [Google Scholar]
- 8.Zhang Z., Xin S., Gao M. and Cai Y. (2017) Promoter hypermethylation of MGMT gene may contribute to the pathogenesis of gastric cancer: A PRISMA-compliant meta-analysis. Medicine (Baltimore) 96, e6708 10.1097/MD.0000000000006708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jayaprakash C., Radhakrishnan R., Ray S. and Satyamoorthy K. (2017) Promoter methylation of MGMT in oral carcinoma: A population-based study and meta-analysis. Arch. Oral Biol. 80, 197–208 10.1016/j.archoralbio.2017.04.006 [DOI] [PubMed] [Google Scholar]
- 10.Theodoratou E., Timofeeva M., Li X., Meng X. and Ioannidis J.P.A. (2017) Nature, nurture, and cancer risks: genetic and nutritional contributions to cancer. Annu. Rev. Nutr. 37, 293–320 10.1146/annurev-nutr-071715-051004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sugimura H. (2016) Susceptibility to human cancer: from the perspective of a pathologist. Pathol. Int. 66, 359–368 10.1111/pin.12418 [DOI] [PubMed] [Google Scholar]
- 12.Zhong Y., Huang Y., Huang Y., Zhang T., Ma C., Zhang S. et al. (2010) Effects of O6-methylguanine-DNA methyltransferase (MGMT) polymorphisms on cancer: a meta-analysis. Mutagenesis 25, 83–95 10.1093/mutage/gep050 [DOI] [PubMed] [Google Scholar]
- 13.Liu J., Zhang R., Chen F., Yu C., Sun Y., Jia C. et al. (2013) MGMT Leu84Phe polymorphism contributes to cancer susceptibility: evidence from 44 case-control studies. PLoS ONE 8, e75367 10.1371/journal.pone.0075367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Du L., Wang H., Xiong T., Ma Y., Yang J., Huang J. et al. (2013) The polymorphisms in the MGMT gene and the risk of cancer: a meta-analysis. Tumour Biol. 34, 3227–3237 10.1007/s13277-013-0893-x [DOI] [PubMed] [Google Scholar]
- 15.Stang A. (2010) Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur. J. Epidemiol. 25, 603–605 10.1007/s10654-010-9491-z [DOI] [PubMed] [Google Scholar]
- 16.Agalliu I., Kwon E.M., Salinas C.A., Koopmeiners J.S., Ostrander E.A. and Stanford J.L. (2010) Genetic variation in DNA repair genes and prostate cancer risk: results from a population-based study. Cancer Causes Control 21, 289–300 10.1007/s10552-009-9461-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Akbari M.R., Malekzadeh R., Shakeri R., Nasrollahzadeh D., Foumani M., Sun Y. et al. (2009) Candidate gene association study of esophageal squamous cell carcinoma in a high-risk region in Iran. Cancer Res. 69, 7994–8000 10.1158/0008-5472.CAN-09-1149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Betti M., Ferrante D., Padoan M., Guarrera S., Giordano M., Aspesi A. et al. (2011) XRCC1 and ERCC1 variants modify malignant mesothelioma risk: a case-control study. Mutat. Res. 708, 11–20 10.1016/j.mrfmmm.2011.01.001 [DOI] [PubMed] [Google Scholar]
- 19.Bye H., Prescott N.J., Matejcic M., Rose E., Lewis C.M., Parker M.I. et al. (2011) Population-specific genetic associations with oesophageal squamous cell carcinoma in South Africa. Carcinogenesis 32, 1855–1861 10.1093/carcin/bgr211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chae M.H., Jang J.S., Kang H.G., Park J.H., Park J.M., Lee W.K. et al. (2006) O6-alkylguanine-DNA alkyltransferase gene polymorphisms and the risk of primary lung cancer. Mol. Carcinog. 45, 239–249 10.1002/mc.20171 [DOI] [PubMed] [Google Scholar]
- 21.Chuang S.C., Agudo A., Ahrens W., Anantharaman D., Benhamou S., Boccia S. et al. (2011) Sequence variants and the risk of head and neck cancer: pooled analysis in the INHANCE consortium. Front. Oncol. 1, 13 10.3389/fonc.2011.00013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Doecke J., Zhao Z.Z., Pandeya N., Sadeghi S., Stark M., Green A.C. et al. (2008) Polymorphisms in MGMT and DNA repair genes and the risk of esophageal adenocarcinoma. Int. J. Cancer 123, 174–180 10.1002/ijc.23410 [DOI] [PubMed] [Google Scholar]
- 23.Felini M.J., Olshan A.F., Schroeder J.C., North K.E., Carozza S.E., Kelsey K.T. et al. (2007) DNA repair polymorphisms XRCC1 and MGMT and risk of adult gliomas. Neuroepidemiology 29, 55–58 10.1159/000108919 [DOI] [PubMed] [Google Scholar]
- 24.Feng X.X., Li Z.F., Wang L.B., Zhang J.B. and Lu Z.X. (2008) Relationship between MGMT gene polymorphism and susceptibility of esophageal cancer. Zhong Guo Gong Gong Wei Sheng 24, 697–699 [Google Scholar]
- 25.Gu F., Qureshi A.A., Kraft P., Guo Q., Hunter D.J. and Han J. (2009) Polymorphisms in genes involved in DNA repair, cell growth, oxidative stress and inflammatory response, and melanoma risk. Br. J. Dermatol. 161, 209–212 10.1111/j.1365-2133.2009.09219.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hall J., Hashibe M., Boffetta P., Gaborieau V., Moullan N., Chabrier A. et al. (2007) The association of sequence variants in DNA repair and cell cycle genes with cancers of the upper aerodigestive tract. Carcinogenesis 28, 665–671 10.1093/carcin/bgl160 [DOI] [PubMed] [Google Scholar]
- 27.Han J., Hankinson S.E. and De Vivo I. (2006) Polymorphisms in O6-methylguanine DNA methyltransferase and endometrial cancer risk. Carcinogenesis 27, 2281–2285 10.1093/carcin/bgl099 [DOI] [PubMed] [Google Scholar]
- 28.Han J., Tranah G.J., Hankinson S.E., Samson L.D. and Hunter D.J. (2006) Polymorphisms in O6-methylguanine DNA methyltransferase and breast cancer risk. Pharmacogenet. Genomics 16, 469–474 10.1097/01.fpc.0000215065.21718.4c [DOI] [PubMed] [Google Scholar]
- 29.Hu Y.Z., Wang D.H., Gong H.D., Li G.Z., Li J.W., Wang L.J. et al. (2013) Correlation of polymorphism of DNA repair gene XRCC1 and MGMT with susceptibility to glioma in a Han population in northeastern China. Chin. J. Cancer Prev. Treat. 20, 1629–1633 [Google Scholar]
- 30.Huang G.Y., Zhang X., Wu Y.J., Cao C.N., Pu J., Ming Y. et al. (2017) Association between genetic polymorphisms of MGMT and CASP3 and the susceptibility to glioma. Guizhou Med. J. 41, 118–140 [Google Scholar]
- 31.Huang J., Ye F., Chen H., Lu W. and Xie X. (2007) Amino acid substitution polymorphisms of the DNA repair gene MGMT and the susceptibility to cervical carcinoma. Carcinogenesis 28, 1314–1322 10.1093/carcin/bgm003 [DOI] [PubMed] [Google Scholar]
- 32.Huang S.H., Chang P.Y., Liu C.J., Lin M.W. and Hsia K.T. (2010) O6-methylguanine-DNA methyltransferase gene coding region polymorphisms and oral cancer risk. J. Oral Pathol. Med. 39, 645–650 10.1111/j.1600-0714.2009.00880.x [DOI] [PubMed] [Google Scholar]
- 33.Huang W.Y., Chow W.H., Rothman N., Lissowska J., Llaca V., Yeager M. et al. (2005) Selected DNA repair polymorphisms and gastric cancer in Poland. Carcinogenesis 26, 1354–1359 10.1093/carcin/bgi084 [DOI] [PubMed] [Google Scholar]
- 34.Huang W.Y., Olshan A.F., Schwartz S.M., Berndt S.I., Chen C., Llaca V. et al. (2005) Selected genetic polymorphisms in MGMT, XRCC1, XPD, and XRCC3 and risk of head and neck cancer: a pooled analysis. Cancer Epidemiol Biomarkers Prev. 14, 1747–1753 10.1158/1055-9965.EPI-05-0162 [DOI] [PubMed] [Google Scholar]
- 35.Inoue R., Isono M., Abe M., Abe T. and Kobayashi H. (2003) A genotype of the polymorphic DNA repair gene MGMT is associated with de novo glioblastoma. Neurol. Res 25, 875–879 10.1179/016164103771954005 [DOI] [PubMed] [Google Scholar]
- 36.Kiczmer P., Prawdzic Senkowska A., Strzelczyk J.K., Szydlo B., Biernacki K., Osadnik T. et al. (2018) The role of MGMT polymorphisms rs12917 and rs11016879 in head and neck cancer risk and prognosis. Acta Biochim. Pol. 65, 87–92 10.18388/abp.2017_1613 [DOI] [PubMed] [Google Scholar]
- 37.Kietthubthew S., Sriplung H., Au W.W. and Ishida T. (2006) Polymorphism in DNA repair genes and oral squamous cell carcinoma in Thailand. Int. J. Hyg. Environ. Health 209, 21–29 10.1016/j.ijheh.2005.06.002 [DOI] [PubMed] [Google Scholar]
- 38.Li C., Liu J., Li A., Qian L., Wang X., Wei Q. et al. (2005) Exon 3 polymorphisms and haplotypes of O6-methylguanine-DNA methyltransferase and risk of bladder cancer in southern China: a case-control analysis. Cancer Lett. 227, 49–57 10.1016/j.canlet.2005.03.043 [DOI] [PubMed] [Google Scholar]
- 39.Liu R.Q. and Zhuang Z.X. (2002) Single-nucleotide polymorphisms of human O6-methylguanine-DNA methyltransferase (MGMT) gene in lung cancer patients from south china. Wei Sheng Du Li Xue Za Zhi 16, 1–5 [Google Scholar]
- 40.Liu R.Q., Zhuang Z.X., He C.H. and He Y. (2002) Relationship between genetic polymorphisms of human O6-methylguanine-DNA methyltransferase (MGMT) gene. Ai Zheng Ji Bian Tu Bian 14, 101–106 [Google Scholar]
- 41.Liu S.H., Su M., Cheng L., Sun B.L. and Lu Z.H. (2006) Polymorphisms of O6-methylguanine-DNA methyltransferase gene in Chinese Chaoshan esophageal cancer patients. Ai Zheng Ji Bian Tu Bian 18, 100–104 [Google Scholar]
- 42.Liu Y., Scheurer M.E., El-Zein R., Cao Y., Do K.A., Gilbert M. et al. (2009) Association and interactions between DNA repair gene polymorphisms and adult glioma. Cancer Epidemiol. Biomarkers Prev. 18, 204–214 10.1158/1055-9965.EPI-08-0632 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Loh Y.H., Mitrou P.N., Wood A., Luben R.N., McTaggart A., Khaw K.T. et al. (2011) SMAD7 and MGMT genotype variants and cancer incidence in the European Prospective Investigation into Cancer and Nutrition (EPIC)-Norfolk Study. Cancer Epidemiol. 35, 369–374 10.1016/j.canep.2010.09.011 [DOI] [PubMed] [Google Scholar]
- 44.Lu Y., Xu Y.C., Shen J., Yu R.B., Niu J.Y. and Guo J.T. (2006) Study on the association between the role of polymorphisms of the O6-methylguanine-DNA methyltransferase gene and gastric cancer hereditary susceptibility. Ji Bing Kong Zhi Za Zhi 10, 222–225 [Google Scholar]
- 45.McKean-Cowdin R., Barnholtz-Sloan J., Inskip P.D., Ruder A.M., Butler M., Rajaraman P. et al. (2009) Associations between polymorphisms in DNA repair genes and glioblastoma. Cancer Epidemiol. Biomarkers Prev. 18, 1118–1126 10.1158/1055-9965.EPI-08-1078 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.O’Mara T.A., Ferguson K., Fahey P., Marquart L., Yang H.P., Lissowska J. et al. (2011) CHEK2, MGMT, SULT1E1 and SULT1A1 polymorphisms and endometrial cancer risk. Twin Res. Hum. Genet. 14, 328–332 10.1375/twin.14.4.328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Palli D., Polidoro S., D’Errico M., Saieva C., Guarrera S., Calcagnile A.S. et al. (2010) Polymorphic DNA repair and metabolic genes: a multigenic study on gastric cancer. Mutagenesis 25, 569–575 10.1093/mutage/geq042 [DOI] [PubMed] [Google Scholar]
- 48.Rajaraman P., Hutchinson A., Wichner S., Black P.M., Fine H.A., Loeffler J.S. et al. (2010) DNA repair gene polymorphisms and risk of adult meningioma, glioma, and acoustic neuroma. Neuro Oncol. 12, 37–48 10.1093/neuonc/nop012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ritchey J.D., Huang W.Y., Chokkalingam A.P., Gao Y.T., Deng J., Levine P. et al. (2005) Genetic variants of DNA repair genes and prostate cancer: a population-based study. Cancer Epidemiol. Biomarkers Prev. 14, 1703–1709 10.1158/1055-9965.EPI-04-0809 [DOI] [PubMed] [Google Scholar]
- 50.Shah M.A., Shaff S.M., Lone G.N. and Jan S.M. (2012) Lack of influence of MGMT codon Leu84Phe and codon Ileu143Val polymorphisms on esophageal cancer risk in the Kashmir valley. Asian Pac. J. Cancer Prev. 13, 3047–3052 10.7314/APJCP.2012.13.7.3047 [DOI] [PubMed] [Google Scholar]
- 51.Shen J., Terry M.B., Gammon M.D., Gaudet M.M., Teitelbaum S.L., Eng S.M. et al. (2005) MGMT genotype modulates the associations between cigarette smoking, dietary antioxidants and breast cancer risk. Carcinogenesis 26, 2131–2137 10.1093/carcin/bgi179 [DOI] [PubMed] [Google Scholar]
- 52.Shen M., Purdue M.P., Kricker A., Lan Q., Grulich A.E., Vajdic C.M. et al. (2007) Polymorphisms in DNA repair genes and risk of non-Hodgkin’s lymphoma in New South Wales, Australia. Haematologica 92, 1180–1185 10.3324/haematol.11324 [DOI] [PubMed] [Google Scholar]
- 53.Shi J.Y., Ren Z.H., Jiao B., Xiao R., Yun H.Y., Chen B. et al. (2011) Genetic variations of DNA repair genes and their prognostic significance in patients with acute myeloid leukemia. Int. J. Cancer 128, 233–238 10.1002/ijc.25318 [DOI] [PubMed] [Google Scholar]
- 54.Stern M.C., Conti D.V., Siegmund K.D., Corral R., Yuan J.M., Koh W.P. et al. (2007) DNA repair single-nucleotide polymorphisms in colorectal cancer and their role as modifiers of the effect of cigarette smoking and alcohol in the Singapore Chinese Health Study. Cancer Epidemiol. Biomarkers Prev. 16, 2363–2372 10.1158/1055-9965.EPI-07-0268 [DOI] [PubMed] [Google Scholar]
- 55.Tranah G.J., Bugni J., Giovannucci E., Ma J., Fuchs C., Hines L. et al. (2006) O6-methylguanine-DNA methyltransferase Leu84Phe and Ile143Val polymorphisms and risk of colorectal cancer in the Nurses’ Health Study and Physicians’ Health Study (United States). Cancer Causes Control 17, 721–731 10.1007/s10552-006-0005-y [DOI] [PubMed] [Google Scholar]
- 56.Yang F., Shi J.Y., Xu L., Ren L.J., Zhang Q.H., Zhao W.L. et al. (2009) Genetic susceptibility of single nucleotide polymorphism in MGMT to non-Hodgkin lymphoma. Zhonghua Xue Ye Xue Za Zhi 30, 622–625 [PubMed] [Google Scholar]
- 57.Zhang M., Huang W.Y., Andreotti G., Gao Y.T., Rashid A., Chen J. et al. (2008) Variants of DNA repair genes and the risk of biliary tract cancers and stones: a population-based study in China. Cancer Epidemiol. Biomarkers Prev. 17, 2123–2127 10.1158/1055-9965.EPI-07-2735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zhang Z., Wang L., Wei S., Liu Z., Wang L.E., Sturgis E.M. et al. (2010) Polymorphisms of the DNA repair gene MGMT and risk and progression of head and neck cancer. DNA Repair (Amst.) 9, 558–566 10.1016/j.dnarep.2010.02.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zienolddiny S., Campa D., Lind H., Ryberg D., Skaug V., Stangeland L. et al. (2006) Polymorphisms of DNA repair genes and risk of non-small cell lung cancer. Carcinogenesis 27, 560–567 10.1093/carcin/bgi232 [DOI] [PubMed] [Google Scholar]
- 60.Bigler J., Ulrich C.M., Kawashima T., Whitton J. and Potter J.D. (2005) DNA repair polymorphisms and risk of colorectal adenomatous or hyperplastic polyps. Cancer Epidemiol. Biomarkers Prev. 14, 2501–2508 10.1158/1055-9965.EPI-05-0270 [DOI] [PubMed] [Google Scholar]
- 61.Jiao L., Bondy M.L., Hassan M.M., Wolff R.A., Evans D.B., Abbruzzese J.L. et al. (2006) Selected polymorphisms of DNA repair genes and risk of pancreatic cancer. Cancer Detect. Prev. 30, 284–291 10.1016/j.cdp.2006.05.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Krzesniak M., Butkiewicz D., Samojedny A., Chorazy M. and Rusin M. (2004) Polymorphisms in TDG and MGMT genes - epidemiological and functional study in lung cancer patients from Poland. Ann. Hum. Genet. 68, 300–312 10.1046/j.1529-8817.2004.00079.x [DOI] [PubMed] [Google Scholar]
- 63.Moreno V., Gemignani F., Landi S., Gioia-Patricola L., Chabrier A., Blanco I. et al. (2006) Polymorphisms in genes of nucleotide and base excision repair: risk and prognosis of colorectal cancer. Clin. Cancer Res. 12, 2101–2108 10.1158/1078-0432.CCR-05-1363 [DOI] [PubMed] [Google Scholar]
- 64.Hung R.J., Baragatti M., Thomas D., McKay J., Szeszenia-Dabrowska N., Zaridze D. et al. (2007) Inherited predisposition of lung cancer: a hierarchical modeling approach to DNA repair and cell cycle control pathways. Cancer Epidemiol. Biomarkers Prev. 16, 2736–2744 10.1158/1055-9965.EPI-07-0494 [DOI] [PubMed] [Google Scholar]
- 65.Hazra A., Chanock S., Giovannucci E., Cox D.G., Niu T., Fuchs C. et al. (2008) Large-scale evaluation of genetic variants in candidate genes for colorectal cancer risk in the nurses’ health study and the health professionals’ follow-up study. Cancer Epidemiol. Biomarkers Prev. 17, 311–319 10.1158/1055-9965.EPI-07-0195 [DOI] [PubMed] [Google Scholar]
- 66.Khatami F., Noorinayer B., Mohebi S.R., Ghiasi S., Mohebi R., Hashemi M. et al. (2009) Effects of amino acid substitution polymorphisms of two DNA methyltransferases on susceptibility to sporadic colorectal cancer. Asian Pac. J. Cancer Prev. 10, 1183–1188 [PubMed] [Google Scholar]
- 67.Adel Fahmideh M., Schwartzbaum J., Frumento P. and Feychting M. (2014) Association between DNA repair gene polymorphisms and risk of glioma: a systematic review and meta-analysis. Neuro Oncol. 16, 807–814 10.1093/neuonc/nou003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Liu K. and Jiang Y. (2017) Polymorphisms in DNA repair gene and susceptibility to glioma: a systematic review and meta-analysis based on 33 studies with 15 SNPs in 9 genes. Cell Mol. Neurobiol. 37, 263–274 10.1007/s10571-016-0367-y [DOI] [PMC free article] [PubMed] [Google Scholar]