Abstract
Background
Increased activity of the chaperones Hsp70 and Hsp90 is a common feature of solid tumours. Translocase of the outer mitochondrial membrane 34 (Tomm34) is a cochaperone of both Hsp70 and Hsp90 that was found to be overexpressed in colorectal, hepatocellular, lung and breast carcinomas. The expression profile of Tomm34 in ovarian cancer has not been investigated. Therefore, the aim of the current study was to investigate the expression pattern of Tomm34 in ovarian carcinomas and analyse its correlation with clinico-pathological parameters.
Results
Epithelial ovarian cancers (140) were histologically classified based on their morphology and graded into two types comprising 5 histologic subgroups. Type I carcinomas comprise low grade serous (LGSC), clear cell (CCOC) and endometrioid (ENOC), type II comprises high grade serous carcinomas (HGSC) and solid, pseudoendometrioid, transitional carcinomas (SET). Tomm34 was more highly expressed in type II than type I carcinomas (p < 0.0001). Comparing tumours based on the mutation in the TP53 gene revealed similar results, where mutant tumours exhibited significantly higher levels of Tomm34 (p < 0.0001). The decreased levels of Tomm34 in type I carcinomas were particularly evident in clear cell and mucinous carcinomas. The expression of Tomm34 was also positively correlated with FIGO stage (r = 0.23; p = 0.007). Tomm34 levels also indicated poor prognosis for patients with mutant p53.
Conclusions
Our data indicate that Tomm34 is commonly expressed at high levels in epithelial ovarian cancers, except for the clear cell and mucinous subtypes. The expression of Tomm34 corresponds with the dualistic model of ovarian cancer pathogenesis where high grade, type II tumours exhibit higher expression of Tomm34 in contrast to type I tumours. These data are also comparable to the previous findings that Tomm34 is a marker of progression and poor prognosis in human cancer.
Electronic supplementary material
The online version of this article (10.1186/s13048-019-0498-0) contains supplementary material, which is available to authorized users.
Keywords: Tomm34, Chaperone, Epithelial ovarian cancer, Tumour, Immunohistochemistry, Ovary, Heat shock protein
Background
Epithelial ovarian cancer accounts for approximately 90% of ovarian tumours and is usually diagnosed only at advanced stages of the International Federation of Gynecology and Obstetrics (FIGO) system, accounting for its high mortality rate [1, 2]. Epithelial ovarian cancer is not a single disease and a dualistic model has been proposed to describe morphological subtypes and cell of origin [2, 3]. Type I ovarian cancers are low grade and develop slowly (including endometrioid, clear cell, mucinous and low grade serous adenocarcinomas) whereas type II cancers are the most common form, representing high-grade serous adenocarcinomas. It is also recognized that high and low-grade serous adenocarcinomas originate from precursor lesions in the fallopian tube, whilst the other histological type I tumours arise from endometriosis, germ cells or transitional cells, with an important role for distinct genetic alterations influencing the tumour characteristics [4–6]. Most notably, mutations in the p53 tumour suppressor are an overwhelming characteristic of Type II high grade serous tumours [7, 8]. Whilst many low grade lesions can be treated with surgery alone, high grade epithelial ovarian cancer is a difficult-to-treat disease that requires surgery plus combination chemotherapy, and even then recurrence is common (70–80% within two years), although this is improving with current targeted therapies for subsets of patients [9].
Increased chaperone activities are a universal feature of cancer and anti-Hsp90 or anti-Hsp70 therapeutics are under investigation for the treatment of various cancer types [10], including ovarian cancer [11, 12]. Chaperone activities are dependent on their interactions with co-chaperones that provide either protein folding or protein degradation functions. The protein folding activity of chaperones is hyperactive in cancers due to enhanced interactions of phosphorylated Hsp90 with Hsp70/Hsp90-organising protein (HOP, also known as STIP1) and reduced interaction with C-terminal Hsp70 interacting protein (CHIP, also known as STUB1) [13].
Translocase of the outer mitochondrial membrane 34 (Tomm34) is an additional component of the cellular chaperone system involved in protein folding. As the name suggests, Tomm34 was initially identified as being involved in mitochondrial protein processing [14, 15]. Subsequent studies have shown that Tomm34 interacts with both Hsp70 and Hsp90 and modifies their protein folding activities [16–20]. In cancer, high levels of Tomm34 have been reported in bladder, colorectal and breast cancers compared to their normal tissue counterparts [21–26]. In these cancers, Tomm34 promotes colorectal cancer cell growth [21] and is a biomarker of poor outcome in early invasive breast cancer [22] and bladder cancer [26]. As a tumour-associated protein, Tomm34 peptide vaccination is under investigation as a therapeutic option for colorectal cancer, with significant Tomm34 cytotoxic T-lymphocyte (CTL) response observed [27–29].
Previous data have indicated that the ovary also expresses TOMM34 mRNA [21] although its expression in ovarian cancer has not been reported. Here, we investigated the levels of Tomm34 in ovarian cancers of mixed subtypes using immunohistochemistry. The data were correlated with tumour type and clinicopathological variables and demonstrate that Tomm34 is expressed at high levels in type II carcinomas and correlates with high FIGO stage.
Results
Patient details
The patients (136) ranged in age from 29 to 86 years old (mean 59, median 59). Histological diagnosis classified tumours based on their morphology and grade into two types comprising 6 histologic subgroups. Type I carcinomas comprise low grade serous (LGSC), clear cell ovarian carcinoma (CCOC), endometrioid ovarian carcinoma (ENOC) and mucinous ovarian carcinoma (MOC). Type II include high grade serous carcinomas (HGSC) and solid, pseudoendometrioid, transitional carcinomas (SET). Since the carcinomas with serous and endometrioid morphology exhibited higher heterogeneity, the assignment of individual samples to histological subtypes was further verified by analysis of p53 status using sequencing and immunohistochemistry (Fig. 1). Serum CA125 levels varied from 6.6–42,415 U/ml, with 6 patients showing levels of less than 35. Thirty four (25%) cancers were FIGO stage 1; 15 (11%) FIGO 2; 65 (48%) FIGO 3 and 20 (15%) FIGO 4 (1 missing, n = 135). Fifty-one patients (38%) had residual disease (3 missing, n = 133) and 46 patients were alive and 90 were dead at last follow up. Twenty patients (14%) had a secondary tumour. Detailed information on individual cases is given in additional file 1.
Fig. 1.
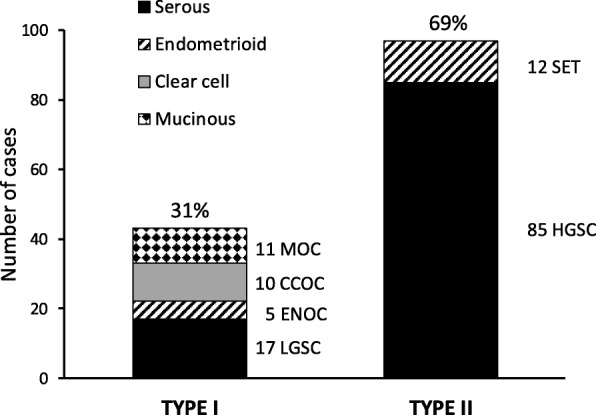
Distribution of histologic groups of 140 epithelial ovarian cancers. High grade tumours with endometrioid morphology and TP53 mutation were classified as type II SET subgroup tumours
Tomm34 staining
Tomm34 staining was seen in the cytoplasm of tumour cells. In the cohort of 136 ovarian cancers, Tomm34 was absent (score of 0) in 14 tumours, 35 cancers were scored as class 1; 44 as class 2; and 43 as class 3 (Fig. 2).
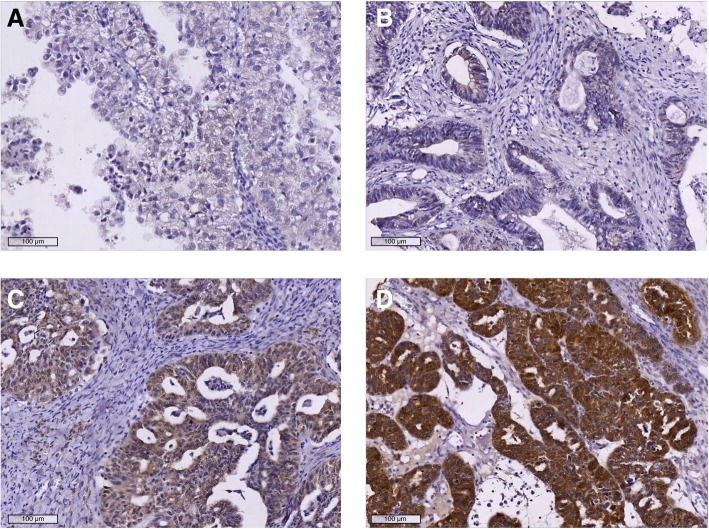
Fig. 2.
Representative staining patterns of Tomm34 in ovarian cancer. a histoscore 0, CCOC, grade 3. b histoscore 1, ENOC, grade 1. c histoscore 2, HGSC, grade 3. d histoscore 3, HGSC, grade 3. Magnification 100x
We tested whether the level of Tomm34 corresponds with the histological type of tumour and whether it correlates with the dualistic model dividing the tumours into 2 types according to their pathogenesis. Figure 3a shows lower expression levels of Tomm34 in type I tumours (MOC, CCOC, LGSC and ENOC) compared to type II tumours (SET and HGSC). The lowest levels of Tomm34 were detected in MOC and CCOC (10 and 11 cases) when compared to the other samples t(138) = 5.6; p < 0.0001. The most statistically significant differences in Tomm34 were found when the cohort was classified according to tumours of types I and II t(138) = 6.6; p < 0.0001 (Fig. 3b). Comparing tumours bearing mutation in the TP53 gene (94 cases) with TP53 wild-type tumours (39 cases; 7 missing) revealed that TP53 mutant tumours exhibited significantly higher levels of Tomm34 t(130) = 4.7; p < 0.0001.
Fig. 3.
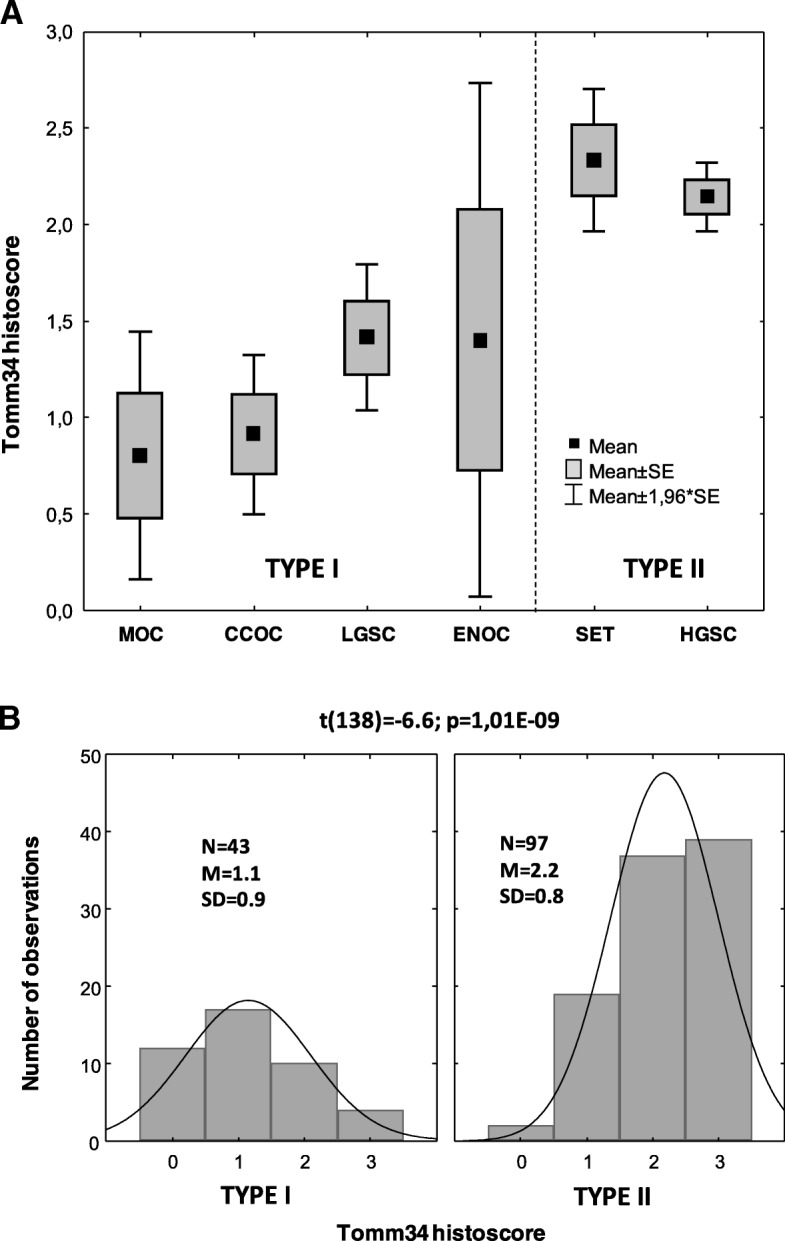
Expression of Tomm34 according to histological classification. a Distribution according to histological group. b The independent samples t-test revealed the most significant difference in Tomm34 between type I and type II tumours
Tomm34 staining also correlated with tumour progression represented by FIGO scale (Fig. 4). In accordance with this finding, higher values of Tomm34 were also found in patients who had residual disease after surgery (n = 51) versus the rest (n = 82) using independent samples t-test (p = 0.036). We also found that high Tomm34 levels correlate with pT status but not with presence of secondary tumour or response to therapy (Table 1). Analysis of patients with serous cancers as a separate group showed that Tomm34 staining correlated with pT status and high FIGO stage, but not with tumour grade or any of the other variables analysed (Table 2).
Fig. 4.
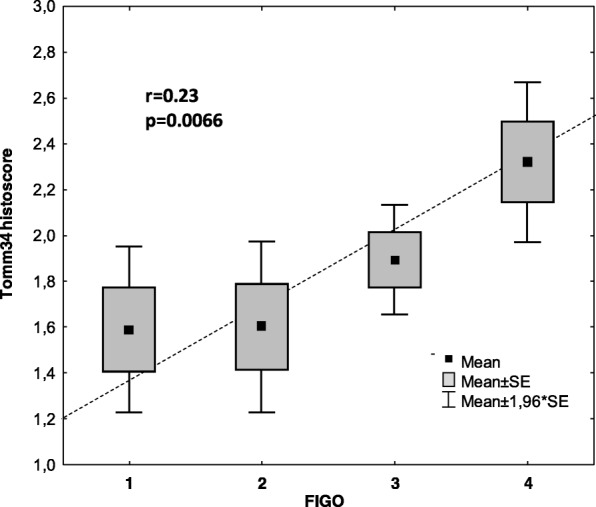
Relationship between FIGO stage and Tomm34 histoscore (0–3). Pearson correlation r = 0.23; p = 0.0066
Table 1.
Tomm34 correlates with tumour grade, FIGO stage and tumour type in ovarian cancer (n = 140)
| Variable | Tomm34 0–1 | Tomm34 2–3 | P value |
|---|---|---|---|
| Tomm 34 score | 50 | 90 | |
| pT 1–2 | 29 | 22 | |
| pT 3–4 | 19 | 57 | 0.0004 |
| pN 0 | 19 | 18 | |
| pN 1 | 11 | 16 | 0.454 |
| M0 | 44 | 59 | |
| M1 | 4 | 15 | 0.123 |
| Secondary No | 30 | 76 | |
| Secondary Yes | 8 | 11 | 0.456 |
| Refractory | 2 | 4 | |
| Resistant | 7 | 15 | |
| Sensitive | 29 | 63 | 0.966 |
pT size of primary tumour, pN degree of spread to regional lymph nodes given by histopathologic examination and M presence of distant metastasis. Tomm34 0–1 = IHC score of 0 or 1; Tomm34 2–3 = IHC score of 2 or 3; P value = Fisher exact 2-tailed test
Table 2.
Tomm34 staining in serous ovarian cancers (n = 102)
| Variable | Tomm 0–1 | Tomm 2–3 | P value |
|---|---|---|---|
| Tomm 34 | 29 | 73 | |
| pT 1–2 | 15 | 16 | |
| pT 3–4 | 12 | 49 | 0.0071 |
| pN 0 | 8 | 13 | |
| pN 1 | 7 | 14 | 1 |
| M0 | 26 | 48 | |
| M1 | 2 | 14 | 0.134 |
| Secondary No | 24 | 59 | |
| Secondary Yes | 4 | 11 | 1 |
| Refractory | 0 | 4 | |
| Resistant | 5 | 14 | |
| Sensitive | 19 | 48 | 0.707 |
pT size of primary tumour, pN degree of spread to regional lymph nodes given by histopathologic examination and M presence of distant metastasis. Tomm34 0–1 = IHC score of 0 or 1; Tomm34 2–3 = IHC score of 2 or 3; P value = Fisher exact 2-tailed test
To investigate the effects of Tomm34 on patient survival in more detail, we also analysed publicly available data using Kaplan-Meier Plotter for Ovarian Cancer (http://kmplot.com/analysis/index.php?p=service&cancer=ovar). These analyses indicated that Tomm34 mRNA levels associate with poor overall survival (p = 0.036; HR = 1.29 (1.02–1.64; n = 506), poor progression-free survival (p = 0.0026; HR = 1.42 (1.13–1.8); n = 483) and poor post-progression free survival (p = 0.012; HR = 1.4 (1.07–1.82); n = 325) in patients with mutant p53 cancers (Additional file 2).
Discussion
Cancer cells require a high level of protein synthesis due to their rapid proliferation and exhibit a high degree of stress due to genomic instability and lack of both oxygen and essential nutrients. To overcome such difficulties, cancer cells show up-regulation of many stress-induced proteins, including components of the protein chaperone system, particularly Hsp70 and Hsp90 [30–32]. Chaperone inhibition is therefore a clinically promising area [10], including for the treatment of ovarian cancer [11, 12]. Metabolic abnormalities are also a common finding in human cancers and the high rate of glucose uptake by tumour cells is utilised clinically for cancer imaging by 18F-2-deoxyglucose uptake and accumulation. Although it was originally thought that aerobic glycolysis in cancer cells (the Warburg effect) represented a lack of mitochondrial function, it is now clear that altered mitochondrial function represents a redeployment of glycolytic nutrients from catabolism to anabolism, required to meet the biosynthetic requirements of rapidly proliferating cancer cells, and NADPH production to help maintain redox balance [33–36].
These alterations to mitochondrial function relate to altered chaperoning of mitochondrial proteins [37]. Thus, targeting abnormal mitochondrial function and chaperoning of mitochondrial proteins is also a promising strategy for cancer therapeutics [38]. Hsp90 and Hsp70 play critical roles in mitochondrial protein stabilisation and folding [39] and various co-chaperones such as p23, Hsp40 and HOP are also involved [40–42]. In this respect, Tomm34 was originally identified as a potential mitochondrial import protein [15] and Tomm34 antibodies inhibit transport of preproteins into the mitochondria, whilst expression stimulates mitochondrial preprotein maturation and TOMM34 siRNA inhibits this process [14]. Tomm34 is also reported to be increased as a component of compensatory adaptations to maintain normal rates of protein import in response to mitochondrial abnormalities [43]. On the other hand, and in agreement with our immunostaining data, Tomm34 exists predominantly in the cytoplasm rather than in mitochondria, suggesting it is involved in the transport of mitochondrial preproteins in an unfolded state prior to import [14, 17, 44].
Our data indicate that Tomm34 is commonly expressed at high levels in human ovarian cancers, except for the MOC and CCOC subtype, where high level Tomm34 is rarely seen. Within the different sub-types of ovarian cancer, high levels of Tomm34 associate with higher stage and higher grade cancers and similar findings are seen within the sub-type of serous cancers. Most notably, we have found that Tomm34 associates with poor survival of patients with p53-mutant ovarian cancers. These data are also comparable to the findings that Tomm34 is a marker of poor outcome and a predictor of distant metastasis in breast cancer [22, 23]. Thus, as with breast cancer patients, Tomm34 may serve as part of a panel of markers for prognostic determination in ovarian cancers. In this respect, proteomic analysis of an animal model of ovarian cancer revealed up-regulation of numerous proteins involved in metabolic processes, including endoplasmic stress responses, mitochondrial systems and chaperones such as Hsp70 and Grp78 [45].
The mechanism for high level expression of Tomm34 is unclear from our studies. Data from the COSMIC database (http://cancer.sanger.ac.uk/cosmic) indicate that only 1.5% of 729 ovarian cancers tested show copy number variation gain of TOMM34, indicating that gene amplification is a rare event. However, Tomm34 is transcriptionally regulated by NRF-1 and NRF-2 [46, 47] and these latter are implicated in directing metabolic reprogramming during stress [48] and are often hyperactive in cancers. Although further research will be required to elucidate the mechanisms that regulate Tomm34 in ovarian and other cancers, our data imply that Tomm34 has pro-tumourigenic actions and may serve as a useful prognostic indicator and potential therapeutic target in ovarian cancers.
Conclusions
The co-chaperone Tomm34 is frequently expressed in epithelial ovarian cancers. Immunohistochemical identification of Tomm34 may be a useful adjunct to provide prognostic information and may serve as an immunogenic target for therapy.
Methods
Ovarian cancer samples and TMA construction
A total of 140 samples of ovarian cancer with anonymised clinicopathological and survival data were available for immunohistochemical staining of Tomm34. All tissues had been removed during surgery, fixed in formalin and processed into paraffin wax for histopathological diagnosis. Tissue microarrays were constructed using five separate cores (1.5 mm diameter each) selected from different regions of each tumour. Clinicopathological information includes age, histological subtype, p53 status, TNM status, grade, FIGO stage, CA125 levels, tumour response to therapy, progression free survival and overall survival. Histological classification was performed as follows: type I carcinomas comprise low grade serous carcinoma (LGSC), clear cell ovarian carcinoma (CCOC), endometrioid ovarian carcinoma (ENOC) and mucinous ovarian carcinoma (MOC). Type II include high grade serous carcinomas (HGSC) and solid, pseudoendometrioid, transitional carcinomas (SET). The study was approved by the MMCI biobank and all patients gave informed consent for the use of their tissues for research.
Tomm34 antibody generation and characterization
Full-length recombinant Tomm34 purified from E. coli was used for immunization of mice. The hybridomas producing monoclonal antibodies were generated by Moravian Biotechnology. The clones were characterized by immunoblotting and immunohistochemistry. Hybridoma clone Tomm34–4.1 was used due to its highest sensitivity and specificity in immunohistochemistry. Specificity of the antibody was tested on MCF7 Tomm34 −/− cell line, where the Tomm34 gene was removed by CRISPR knock-out (Additional file 3).
Tissue microarrays, immunohistochemistry and scoring
Tissue blocks of primary tumours were fixed in 4% neutral formaldehyde for approximately 24 h before processing into paraffin wax. Representative cores of tumour were selected from the diagnostic blocks by an experienced histopathologist. The tissue microarrays comprised 45 tissue cores consisting of nine tumour samples each represented by five cores from the original paraffin block. Endogenous peroxidase was blocked with 3% hydrogen peroxide in phosphate buffered saline (PBS), pH 7.5, for 15 min. Antigen retrieval was performed in 1 mM EDTA–NaOH (pH 8.0) for 40 min at 93 °C. Primary antibodies were applied overnight at 4 °C at 2 μg/ml in antibody diluent (DakoCytomation, Denmark). The antibody was visualized using peroxidase labelled polymer conjugated to goat anti-mouse immunoglobulins and EnVision+ System containing DAB chromogen (DakoCytomation, Denmark). The nuclei were counterstained with Gill’s haematoxylin before permanent mounting. Neoplastic cells were classified into four staining intensities: 0 no staining, 1 weak staining, 2 moderate staining, 3 strong staining.
Statistical analyses
The independent samples t-test was used to compare two sets of independent of and identically distributed samples; P values < 0.05 were considered significant. Relationships between two continuous variables were tested by Pearson correlation where correlation coefficient r was considered significant if P values < 0.05.
Additional files
Clinicopathological data of individual cases (PDF 276 kb)
Survival analysis based on Tomm34 expression in epithelial ovarian cancers with p53 mutation. (A) Overall survival, (B) progression free survival and (C) post progression survival. (PDF 204 kb)
Detection of Tomm34 by mouse monoclonal antibody Tomm34.4.1. MCF7 cells with knockout expression of Tomm34 gene were transfected with constructs encoding HA-tag labelled full-length Tomm34 protein and its TPR1 (aa 1–188) and TPR2 (188–309) domains. Cell lysates were separated by SDS-PAGE, blotted and the membranes were probed with either anti-HA tag antibody or Tomm34.4.1 monoclonal antibody. Protein loading was tested by probing the membrane with anti-Actin antibody. (PDF 158 kb)
Acknowledgments
Funding
The work was supported by the Czech Science Foundation (GACR 16-07321S) and partially supported by the Ministry of Education, Youth and Sports of the Czech Republic; National Programme of Sustainability I (MEYS – NPS I – LO1413) and by the Ministry of Health Development of Research Organization, MH CZ - DRO (MMCI, 00209805). The MMCI biobank is supported by grant MEYS-LM2015089 from the Ministry of Education, Youth and Sports, Czech Republic and co-funded by ADOPT BBMRI-ERIC supported by EU Horizon 2020, grant agreement No. 676550. There was no additional external funding received for this study.
Availability of data and materials
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Authors’ contributions
PM designed and performed experiments, analysed data and edited the manuscript; PJC analysed data and wrote the manuscript; RN performed histopathological analysis, preparation of TMAs and immunohistochemical scoring; FT performed experiments and analysed data; RH performed experiments and analysed data; JC analysed clinical data; VB performed experiments, analysed data and edited the manuscript; BV supervised the project, analysed data and edited the manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to participate
Ethical approval for the use of anonymised excess tumour tissues for this research was granted by the Biobank of clinical samples at the Masaryk Memorial Cancer Institute, a member of the pan-European BBMRI (membership number AO192).
Competing interests
RN, PM and BV are associated with Moravian Biotechnology, the company that produced and supplied the Tomm34 monoclonal antibody. The company did not provide financial support for the studies and had no influence on the design, execution or analysis of the experiments. The other authors have declared that no competing interests exist.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Petr Muller, Email: muller@mou.cz.
Philip J. Coates, Email: philip.coates@mou.cz
Rudolf Nenutil, Email: nenutil@mou.cz.
Filip Trcka, Email: pilif.trcka@gmail.com.
Roman Hrstka, Email: hrstka@mou.cz.
Josef Chovanec, Email: josef.chovanec@mou.cz.
Veronika Brychtova, Email: vebrychtova@mou.cz.
Borivoj Vojtesek, Phone: (+420) 5413 3300, Email: vojtesek@mou.cz.
References
- 1.Cannistra SA. Cancer of the ovary. N Engl J Med. 2004;351(24):2519–2529. doi: 10.1056/NEJMra041842. [DOI] [PubMed] [Google Scholar]
- 2.Kurman RJ, Shih Ie M. The dualistic model of ovarian carcinogenesis. Revisited, Revised, and Expanded Am J Pathol. 2016;186(4):733–747. doi: 10.1016/j.ajpath.2015.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Levanon K, Crum C, Drapkin R. New insights into the pathogenesis of serous ovarian cancer and its clinical impact. J Clin Oncol. 2008;26(32):5284–5293. doi: 10.1200/JCO.2008.18.1107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Stuckelberger S, Drapkin R. Precious GEMMs: emergence of faithful models for ovarian cancer research. J Pathol. 2018;245(2):129–131. doi: 10.1002/path.5065. [DOI] [PubMed] [Google Scholar]
- 5.Cochrane DR, Tessier-Cloutier B, Lawrence KM, Nazeran T, Karnezis AN, Salamanca C, et al. Clear cell and endometrioid carcinomas: are their differences attributable to distinct cells of origin? J Pathol. 2017;243(1):26–36. doi: 10.1002/path.4934. [DOI] [PubMed] [Google Scholar]
- 6.Kolin DL, Dinulescu DM, Crum CP. Origin of clear cell carcinoma: nature or nurture? J Pathol. 2018;244(2):131–134. doi: 10.1002/path.5009. [DOI] [PubMed] [Google Scholar]
- 7.Cancer Genome Atlas Research N Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474(7353):609–615. doi: 10.1038/nature10166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Vang R, Levine DA, Soslow RA, Zaloudek C, Shih Ie M, Kurman RJ. Molecular alterations of TP53 are a defining feature of ovarian high-grade serous carcinoma: a Rereview of cases lacking TP53 mutations in the Cancer genome atlas ovarian study. Int J Gynecol Pathol. 2016;35(1):48–55. doi: 10.1097/PGP.0000000000000207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gadducci A, Guarneri V, Peccatori FA, Ronzino G, Scandurra G, Zamagni C, et al. Current strategies for the targeted treatment of high-grade serous epithelial ovarian cancer and relevance of BRCA mutational status. J Ovarian Res. 2019;12(1):9. doi: 10.1186/s13048-019-0484-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Garcia-Carbonero R, Carnero A, Paz-Ares L. Inhibition of HSP90 molecular chaperones: moving into the clinic. Lancet Oncol. 2013;14(9):e358–e369. doi: 10.1016/S1470-2045(13)70169-4. [DOI] [PubMed] [Google Scholar]
- 11.Elstrand MB, Stavnes HT, Trope CG, Davidson B. Heat shock protein 90 is a putative therapeutic target in patients with recurrent advanced-stage ovarian carcinoma with serous effusions. Hum Pathol. 2012;43(4):529–535. doi: 10.1016/j.humpath.2011.05.022. [DOI] [PubMed] [Google Scholar]
- 12.Liu H, Xiao F, Serebriiskii IG, O'Brien SW, Maglaty MA, Astsaturov I, et al. Network analysis identifies an HSP90-central hub susceptible in ovarian cancer. Clin Cancer Res. 2013;19(18):5053–5067. doi: 10.1158/1078-0432.CCR-13-1115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Muller P, Ruckova E, Halada P, Coates PJ, Hrstka R, Lane DP, et al. C-terminal phosphorylation of Hsp70 and Hsp90 regulates alternate binding to co-chaperones CHIP and HOP to determine cellular protein folding/degradation balances. Oncogene. 2013;32(25):3101–3110. doi: 10.1038/onc.2012.314. [DOI] [PubMed] [Google Scholar]
- 14.Chewawiwat N, Yano M, Terada K, Hoogenraad NJ, Mori M. Characterization of the novel mitochondrial protein import component, Tom34, in mammalian cells. J Biochem. 1999;125(4):721–727. doi: 10.1093/oxfordjournals.jbchem.a022342. [DOI] [PubMed] [Google Scholar]
- 15.Nuttall SD, Hanson BJ, Mori M, Hoogenraad NJ. hTom34: a novel translocase for the import of proteins into human mitochondria. DNA Cell Biol. 1997;16(9):1067–1074. doi: 10.1089/dna.1997.16.1067. [DOI] [PubMed] [Google Scholar]
- 16.Tsaytler PA, Krijgsveld J, Goerdayal SS, Rudiger S, Egmond MR. Novel Hsp90 partners discovered using complementary proteomic approaches. Cell Stress Chaperones. 2009;14(6):629–638. doi: 10.1007/s12192-009-0115-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Faou P, Hoogenraad NJ. Tom34: a cytosolic cochaperone of the Hsp90/Hsp70 protein complex involved in mitochondrial protein import. Biochim Biophys Acta. 2012;1823(2):348–357. doi: 10.1016/j.bbamcr.2011.12.001. [DOI] [PubMed] [Google Scholar]
- 18.Trcka F, Durech M, Man P, Hernychova L, Muller P, Vojtesek B. The assembly and intermolecular properties of the Hsp70-Tomm34-Hsp90 molecular chaperone complex. J Biol Chem. 2014;289(14):9887–9901. doi: 10.1074/jbc.M113.526046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Durech M, Trcka F, Man P, Blackburn EA, Hernychova L, Dvorakova P, et al. Novel entropically driven conformation-specific interactions with Tomm34 protein modulate Hsp70 protein folding and ATPase activities. Mol Cell Proteomics. 2016;15(5):1710–1727. doi: 10.1074/mcp.M116.058131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Trcka F, Durech M, Vankova P, Chmelik J, Martinkova V, Hausner J, et al. Human stress-inducible Hsp70 has a high propensity to form ATP-dependent antiparallel dimers that are differentially regulated by Cochaperone binding. Mol Cell Proteomics. 2019;18(2):320–337. doi: 10.1074/mcp.RA118.001044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shimokawa T, Matsushima S, Tsunoda T, Tahara H, Nakamura Y, Furukawa Y. Identification of TOMM34, which shows elevated expression in the majority of human colon cancers, as a novel drug target. Int J Oncol. 2006;29(2):381–386. [PubMed] [Google Scholar]
- 22.Aleskandarany MA, Negm OH, Rakha EA, Ahmed MA, Nolan CC, Ball GR, et al. TOMM34 expression in early invasive breast cancer: a biomarker associated with poor outcome. Breast Cancer Res Treat. 2012;136(2):419–427. doi: 10.1007/s10549-012-2249-4. [DOI] [PubMed] [Google Scholar]
- 23.Aleskandarany MA, Soria D, Green AR, Nolan C, Diez-Rodriguez M, Ellis IO, et al. Markers of progression in early-stage invasive breast cancer: a predictive immunohistochemical panel algorithm for distant recurrence risk stratification. Breast Cancer Res Treat. 2015;151(2):325–333. doi: 10.1007/s10549-015-3406-3. [DOI] [PubMed] [Google Scholar]
- 24.Miyata Y, Kumagai K, Nagaoka T, Kitaura K, Kaneda G, Kanazawa H, et al. Clinicopathological significance and prognostic value of Wilms' tumor gene expression in colorectal cancer. Cancer Biomark. 2015;15(6):789–797. doi: 10.3233/CBM-150521. [DOI] [PubMed] [Google Scholar]
- 25.Zhang B, Wang J, Wang X, Zhu J, Liu Q, Shi Z, et al. Proteogenomic characterization of human colon and rectal cancer. Nature. 2014;513(7518):382–387. doi: 10.1038/nature13438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ahmed MAH, Ali MH, Abbas HH, Elatrash GA, Foda AAM. Expression of TOMM34 and its Clinicopathological correlations in urothelial carcinoma of the bladder. Pathol Oncol Res. 2018. [DOI] [PubMed]
- 27.Hazama S, Nakamura Y, Takenouchi H, Suzuki N, Tsunedomi R, Inoue Y, et al. A phase I study of combination vaccine treatment of five therapeutic epitope-peptides for metastatic colorectal cancer; safety, immunological response, and clinical outcome. J Transl Med. 2014;12:63. doi: 10.1186/1479-5876-12-63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Matsushita N, Aruga A, Inoue Y, Kotera Y, Takeda K, Yamamoto M. Phase I clinical trial of a peptide vaccine combined with tegafur-uracil plus leucovorin for treatment of advanced or recurrent colorectal cancer. Oncol Rep. 2013;29(3):951–959. doi: 10.3892/or.2013.2231. [DOI] [PubMed] [Google Scholar]
- 29.Kawamura J, Sugiura F, Sukegawa Y, Yoshioka Y, Hida JI, Hazama S, et al. Cytotoxic T lymphocyte response to peptide vaccination predicts survival in stage III colorectal cancer. Cancer Sci. 2018;109(5):1545–1551. doi: 10.1111/cas.13547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Calderwood SK, Gong J. Heat shock proteins promote cancer: It's a protection racket. Trends Biochem Sci. 2016;41(4):311–323. doi: 10.1016/j.tibs.2016.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Murphy ME. The HSP70 family and cancer. Carcinogenesis. 2013;34(6):1181–1188. doi: 10.1093/carcin/bgt111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Miyata Y, Nakamoto H, Neckers L. The therapeutic target Hsp90 and cancer hallmarks. Curr Pharm Des. 2013;19(3):347–365. doi: 10.2174/138161213804143725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chandel NS, Jasper H, Ho TT, Passegue E. Metabolic regulation of stem cell function in tissue homeostasis and organismal ageing. Nat Cell Biol. 2016;18(8):823–832. doi: 10.1038/ncb3385. [DOI] [PubMed] [Google Scholar]
- 34.Pavlova NN, Thompson CB. The emerging hallmarks of Cancer metabolism. Cell Metab. 2016;23(1):27–47. doi: 10.1016/j.cmet.2015.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wallace DC. Mitochondria and cancer. Nat Rev Cancer. 2012;12(10):685–698. doi: 10.1038/nrc3365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Weinberg SE, Chandel NS. Targeting mitochondria metabolism for cancer therapy. Nat Chem Biol. 2015;11(1):9–15. doi: 10.1038/nchembio.1712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Rasola A, Neckers L, Picard D. Mitochondrial oxidative phosphorylation TRAP(1)ped in tumor cells. Trends Cell Biol. 2014;24(8):455–463. doi: 10.1016/j.tcb.2014.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Siegelin MD. Inhibition of the mitochondrial Hsp90 chaperone network: a novel, efficient treatment strategy for cancer? Cancer Lett. 2013;333(2):133–146. doi: 10.1016/j.canlet.2013.01.045. [DOI] [PubMed] [Google Scholar]
- 39.Young JC, Hoogenraad NJ, Hartl FU. Molecular chaperones Hsp90 and Hsp70 deliver preproteins to the mitochondrial import receptor Tom70. Cell. 2003;112(1):41–50. doi: 10.1016/s0092-8674(02)01250-3. [DOI] [PubMed] [Google Scholar]
- 40.Baker MJ, Palmer CS, Stojanovski D. Mitochondrial protein quality control in health and disease. Br J Pharmacol. 2014;171(8):1870–1889. doi: 10.1111/bph.12430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hansen KG, Schlagowski A, Herrmann JM. Escorted by chaperones: Sti1 helps to usher precursor proteins from the ribosome to mitochondria. FEBS J. 2016;283(18):3335–3337. doi: 10.1111/febs.13821. [DOI] [PubMed] [Google Scholar]
- 42.Stojanovski D, Bohnert M, Pfanner N, van der Laan M. Mechanisms of protein sorting in mitochondria. Cold Spring Harb Perspect Biol. 2012;4(10). [DOI] [PMC free article] [PubMed]
- 43.Joseph AM, Rungi AA, Robinson BH, Hood DA. Compensatory responses of protein import and transcription factor expression in mitochondrial DNA defects. Am J Physiol Cell Physiol. 2004;286(4):C867–C875. doi: 10.1152/ajpcell.00191.2003. [DOI] [PubMed] [Google Scholar]
- 44.Mukhopadhyay A, Avramova LV, Weiner H. Tom34 unlike Tom20 does not interact with the leader sequences of mitochondrial precursor proteins. Arch Biochem Biophys. 2002;400(1):97–104. doi: 10.1006/abbi.2002.2777. [DOI] [PubMed] [Google Scholar]
- 45.Chuffa LG, Lupi Junior LA, Seiva FR, Martinez M, Domeniconi RF, Pinheiro PF, et al. Quantitative proteomic profiling reveals that diverse metabolic pathways are influenced by melatonin in an in vivo model of ovarian carcinoma. J Proteome Res. 2016;15(10):3872–3882. doi: 10.1021/acs.jproteome.6b00713. [DOI] [PubMed] [Google Scholar]
- 46.Blesa JR, Prieto-Ruiz JA, Abraham BA, Harrison BL, Hegde AA, Hernandez-Yago J. NRF-1 is the major transcription factor regulating the expression of the human TOMM34 gene. Biochem Cell Biol. 2008;86(1):46–56. doi: 10.1139/o07-151. [DOI] [PubMed] [Google Scholar]
- 47.Satoh J, Kawana N, Yamamoto Y. Pathway analysis of ChIP-Seq-based NRF1 target genes suggests a logical hypothesis of their involvement in the pathogenesis of neurodegenerative diseases. Gene Regul Syst Bio. 2013;7:139–152. doi: 10.4137/GRSB.S13204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hayes JD, Dinkova-Kostova AT. The Nrf2 regulatory network provides an interface between redox and intermediary metabolism. Trends Biochem Sci. 2014;39(4):199–218. doi: 10.1016/j.tibs.2014.02.002. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Clinicopathological data of individual cases (PDF 276 kb)
Survival analysis based on Tomm34 expression in epithelial ovarian cancers with p53 mutation. (A) Overall survival, (B) progression free survival and (C) post progression survival. (PDF 204 kb)
Detection of Tomm34 by mouse monoclonal antibody Tomm34.4.1. MCF7 cells with knockout expression of Tomm34 gene were transfected with constructs encoding HA-tag labelled full-length Tomm34 protein and its TPR1 (aa 1–188) and TPR2 (188–309) domains. Cell lysates were separated by SDS-PAGE, blotted and the membranes were probed with either anti-HA tag antibody or Tomm34.4.1 monoclonal antibody. Protein loading was tested by probing the membrane with anti-Actin antibody. (PDF 158 kb)
Data Availability Statement
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.