Abstract
Genetic regulators and environmental stimuli modulate T-cell activation in autoimmunity and cancer. The enzyme co-factor tetrahydrobiopterin (BH4) is involved in the production of monoamine neurotransmitters, the generation of nitric oxide, and pain1,2. Here we uncover a link between these processes, identifying a fundamental role for BH4 in T-cell biology. We find that genetic inactivation of GTP cyclohydrolase 1 (GCH1, the rate-limiting enzyme in the synthesis of BH4) and inhibition of sepiapterin reductase (SPR, the terminal enzyme in its synthetic pathway) severely impair the proliferation of mature mouse and human T cells. BH4 production in activated T cells is linked to alterations in iron metabolism and mitochondrial bioenergetics. In vivo blockade of BH4 synthesis abrogates T-cell-mediated autoimmunity and allergic inflammation, while enhancing BH4 levels through GCH1 overexpression augments responses by CD4- and CD8-expressing T cells, increasing their antitumour activity in vivo. Administration of BH4 to mice markedly reduces tumour growth and expands the population of intratumoral effector T cells. Kynurenine—a tryptophan metabolite that blocks antitumour immunity—inhibits T-cell proliferation in a manner that can be rescued by BH4. Finally, we report the development of a potent SPR antagonist for possible clinical use. Our data uncover GCH1, SPR and their downstream metabolite BH4 as critical regulators of T-cell biology that can be readily manipulated to either block autoimmunity or enhance anticancer immunity.
GCH1—the first enzyme in the de novo BH4-synthesis pathway— is known to be expressed in activated T cells3,4. Using isolated CD4+ and CD8+ T cells from a Gch1-Gfp reporter mouse line1 (where Gfp encodes green fluorescent protein), we confirmed that GCH1 is induced in activated T cells in response both to phorbol myristate acetate (PMA)/ionomycin and to stimulation of T-cell receptors (TCRs) by anti-CD3/CD28 antibodies (Extended Data Fig. 1a-c). To explore the function of the GCH1/BH4 pathway in these cells, we generated mice in which Gch1 is knocked out specifically in T cells by crossing Lck–Cre driver mice with Gch1 (fl/fl)5 mice (producing Gch1;Lck animals). These Gch1;Lck mice showed normal numbers of thymic and peripheral T cells compared with Cre-only controls (Extended Data Fig. 1d); that is, lack of GCH1 does not influence T-cell development or peripheral T cell homeostasis. Stimulation of mature peripheral CD4+ T cells from Gch1;Lck mice revealed, as expected, severely reduced GCH1 protein and BH4 production relative to controls (Fig. 1a, b). Shortly after TCR engagement (at 16 hours), we observed no differences between Gch1;Lck and control T cells in either the expression of surface activation markers or the secretion of interleukin (IL)-2 (Fig. 1c, d). Similar results were obtained with CD8+ T cells (data not shown). However, TCR-stimulated Gch1-deficient CD4+ and CD8+ T cells did display markedly reduced proliferation (Fig. 1e, f and Extended Data Fig. 1e, f). In contrast to this antiproliferative effect on peripheral T cells, Gch1 ablation did not affect the proliferation of DN3a thymocytes co-cultured with OP9–DL1 stromal cells (Extended Data Fig. 1 g–i). Moreover, there were no obvious differences in the survival of thymocytes or of mature naive peripheral T cells (Extended Data Fig. 2a, b).
Fig. 1|. The BH4 pathway is indispensable for effective T-cell proliferation in vitro and in vivo.

a, Immunoblot of GCH1 after 24 hours of TCR stimulation with anti-CD3/CD28 antibodies in CD4+ T cells. The experiment was repeated three times with similar results. Staining for actin acts as a control. b, BH4 production upon 24 hours of anti-CD3/CD28 stimulation in purified CD4+ control and Gch1-null T cells. Individual data (dots and squares; n = 5 mice in each case) are shown as means ± s.e.m. c, d, Representative fluorescence-activated cell sorting (FACS) blot depicting early activation markers (CD62L and CD25; c) and IL-2 secretion (d) before and after T-cell stimulation (16 hours). Data (n = 5 independent samples) are shown as means ± s.e.m. The experiment was repeated two independent times with similar results. FITC, fluorescein isothiocyanate. Naive T cells, CD25low, CD62Lhi; activated T cells, CD25hi, CD62Llow. e, Proliferation of CD4+ T cells after three days of stimulation of control and Gch1;Lck mice. Cell Trace Violet gets diluted in proliferating cells (see Methods). Representative data are shown from more than 15 experiments with similar results. f, Quantification of CD4+ T cell proliferation from individual Gch1;Lck (left; n = 10) and Gch1;RORc (right; n = 7) mice. Data are shown as means ± s.e.m. g, h, Transfer colitis model of intestinal autoimmunity. g, Schematic outline (top) and colitis scores of transferred control and Gch1- ablated CD4+ T cells into Rag1−/−hosts (bottom). D, day. Data (n = 10 mice) are shown as means ± s.e.m. h, Representative immunofluorescence depicting intestinal infiltration of various immune cells (CD3ε+, CD4+, CD11c+ and myeloperoxidase (MPO)+ cells). Scale bar represents 200 μm. i, Allergic airway inflammatory disease model (top) and quantification of inflammatory cells in bronchoalveolar lavage fluids (BALFs; bottom). Data are shown as means ± s.e.m.; n = 35 for control mice; n = 31 for Gch1;Lck mice. OVA, ovalbumin. j, Percentage increase in ear swelling after re-challenge using the 2,4,6-trinitrochlorobenzene (TNCB)-dependent skin hypersensitivity model. Data are shown as means ± s.e.m.; n = 8 for control mice; n = 9 for Gch1;Lck mice. NS, not significant; *P< 0.05; **P< 0.01; ***P < 0.001 (two-tailed Student’s t-test for panels b, d, f, i, j; two-way analysis of variance (ANOVA) with Dunnett’s comparison for panel g).
To validate these findings, we crossed a different T-cell-specific Cre mouse line, RORgammact Cre6, with Gch1 (fl/fl)5 mice. Again, specific loss of GCH1 in T cells did not affect thymocyte development or peripheral T cell homeostasis (data not shown). However, we again found that GCH1 is a key regulator of mature T cell proliferation (Fig. 1e and Extended Data Fig. 2c, d). B-cell-specific deletion of Gch1 using MB1-Cre7 did not affect B-cell development or function (Extended Data Fig. 2e–h). Moreover, loss of GCH1 had no effect on the development, numbers or suppressive capacity of peripheral regulatory T cells (Extended Data Fig. 3a–f). We conclude that GCH1 induction and BH4 synthesis are required for effective proliferation of CD4+ and CD8+ T cells.
To investigate whether Gch1-ablated, BH4-deficent T cells are defective in vivo, we studied several models of T-cell-dependent inflammation. In a colitis model in which naïve, CD4+ T cells are transferred into hosts lacking recombination-activating gene 1 (Rag1)8, transfer of Gch1-deficient CD4+ T cells resulted in a substantially lower influx of immune cells into the gut, with less colonic inflammation and colitis development (Fig. 1g, h). Although numbers of colonic and mesenteric lymph node CD4+ T cells were greatly reduced, the production of the inflammatory T cell cytokines IL-17A and interferon (IFN)-γ were apparently not affected by selective Gch1 deletion (Extended Data Fig. 3g–i). Next, we used a model of type-2 allergic airway inflammation in which immune cells—particularly CD4+ T-helper-2 cells and eosinophils—are central to disease pathology9,10. Compared with controls, Gch1;Lck mice showed substantially fewer CD45+ cells, eosinophils and T cells in bronchoalveolar lavage (Fig. 1i). Moreover, T-cell-dependent ovalbumin-induced immune responses were reduced during primary immunization and re-challenge (Extended Data Fig. 4a). Gch1;Lck mice also showed greatly reduced inflammatory responses in a T-cell-mediated skin dermatitis model11 (Fig. 1j) and in the experimental autoimmune encephalomyelitis (EAE) model of multiple sclerosis12,13 (Extended Data Fig. 4b, c). Therefore, genetic ablation of Gch1 in T cells alleviates T-cell- mediated inflammatory intestinal, airway, skin and brain diseases.
Inhibiting GCH1 pharmacologically is challenging because of its inaccessible active sites14,15. Therefore, we used an inhibitor, SPRi3, that targets sepiapterin reductase (SPR)—the terminal enzyme in the de novo BH4-synthesis pathway (Fig. 2a and Extended Data Fig. 4d). Purified naive CD4+ T cells treated with SPRi3 showed lower BH4 levels than did vehicle-treated cells following TCR stimulation (Fig. 2a). SPRi3-treated, TCR-stimulated CD4+ and CD8+ cells displayed a defect in proliferation that was similar to that of T cells in which Gch1 was genetically ablated (Fig. 2b), without affecting the survival of non-stimulated T cells or the induction of early activation markers (Extended Data Fig. 4e, f). Pulse labelling with 5-ethynyl-2ʹ-deoxyuridine (EdU) revealed that SPRi3 treatment and Gch1 deficiency resulted in substantially fewer S-phase cells after TCR stimulation than did vehicle treatment, culminating in increased cell death (Extended Data Fig. 4g). In vivo, SPRi3 administration significantly ameliorated colitis, greatly diminishing the intestinal infiltration of T cells and other immune cells after CD4+ T cell transfer (Fig. 2c). SPRi3 treatment also reduced immune-cell infiltration into the lungs after challenge involving inhaled ovalbumin in sensitized mice (Fig. 2d). To determine whether these findings translate to human T cells, we isolated human peripheral blood mononuclear cells from different healthy donors (n = 4). Following anti-CD3/CD28 stimulation, SPRi3-treated, freshly isolated human T cells also exhibited greatly reduced proliferation compared with vehicle-treated cells (Fig. 2e). Moreover, we observed a substantial decrease in proliferative capacity in SPRi3-treated purified human effector CD4+ T cells after anti-CD3/CD8 re-stimulation (Fig. 2f).
Fig. 2|. Pharmacological inhibition of the BH4 pathway ameliorates T-cell-mediated inflammation.

a, BH4 production in 24-hour-activated control (n = 5) and Gch1-ablated (n = 5) CD4+ T cells and in wild-type cells treated with SPRi3 (50 μM; n = 4). The experiment was repeated two independent times with similar results. Data are shown as means ± s.e.m. At the top is shown the mechanism of action of SPRi3. b, Representative three-day T-cell-proliferation histogram (right) and quantification (left) of stimulated wild-type T cells (CD4+ and CD8+) treated with vehicle (n = 10) or SPRi3 (50 μM; n = 8). Data are shown as means ± s.e.m. c, Colitis model, involving transfer of wild-type CD4+ T cells into Rag1−/− hosts treated with vehicle or SPRi3 (300 mg kg–1; n = 8 each). Data (bottom left) are shown as means ± s.e.m. Right panels show representative images of intestinal immune infiltration. Scale bar represents 200 μm. d, Allergic airway inflammatory disease model in control mice treated with SPRi3 (300 mg kg–1; n = 14) or vehicle (n = 15). Data are shown as means ± s.e.m. e, f, Proliferation of vehicle- and SPRi3-treated (50 μM) naïve human (n = 4 donors) peripheral blood mononuclear cells (PBMCs) (e) and purified effector human CD4+ T cells (f) re-stimulated for 3 days. Data are shown as means ± s.e.m. g, Western immunoblot of iron regulators in 24-hour-activated peripheral CD4+ T cells from control and Gch1;Lck mice. The experiment was repeated three independent times with similar results. HO-1, haem oxygenase-1. h, Total iron content from unstimulated and 24-hour-stimulated CD4+ T cells from control (n = 17) and Gch1;Lck (n = 22) mice. Data are shown as means ± s.e.m. i, Oxygen-consumption rate (OCR) in unstimulated and 16-hour-stimulated CD4+ T cells from control and Gch1;Lck mice (n = 6 each). Data are shown as means ± s.e.m. j, Top, dose-dependent reduction of ferri-cytochrome c (cytoc-Fe3+) to ferro-cytochrome c (cytoc-Fe2+) by BH4; and bottom, diagram of the reduction pathway. DHPR, dihydropyridine receptor; qBH2, quinonoid dihydrobiopterin. k, l, Representative oxygen uptake rate in permeabilized, 16-hour-stimulated CD4+ T cells from vehicle-treated and SPRi3-treated (50 μM) wild-type cells before and after the addition of cytoc-Fe2+ (k), and quantification of oxygen rate upon supplementation of cytoc-Fe2+ (n = 4 independent experiments) (l). Data are shown as means ± s.e.m. NS, not significant; *P < 0.05; **P < 0.01; ***P < 0.001 (one-way ANOVA with Dunnett’s comparison for panel a; two-tailed Student’s t-test for panels b, d–f, h, i, l; two-way ANOVA with Sidak’s comparison for panel c).
To explore the molecular mechanisms responsible for the proliferation deficit, we carried out gene-expression profiling in TCR-stimulated CD4+ T cells from control and Gch1;Lck mice. Analysis of the greatly altered genes confirmed that loss of GCH1 did not affect early T-cell activation (data accessible through Gene Expression Omnibus (GEO), accession number GSE108101 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE108101)). Biogenic amines or their amino-acid precursors—production of which involves BH4 as a co-factor2—were also unaffected (Extended Data Fig. 4j, k and Supplementary Table 1). Intriguingly, several genes involved in iron homeostasis or its availability were upregulated in the absence of GCH1—a finding confirmed by western blot of activated Gch1-ablated T cells (Fig. 2g). Total iron levels were greatly reduced in TCR-activated Gch1-ablated CD4+ T cells compared with control cells (Fig. 2h). As one of the most upregulated genes in Gch1-ablated cells was mitoferrin, a mitochondrial iron transporter, and because iron is critical for mitochondrial respiration16, we analysed the energy needs of Gch1-deficient activated T cells. Both Gch1-deficient and SPRi3-treated CD4+ T cells synthesized less ATP than control cells after anti-CD3/CD28 stimulation (Extended Data Fig. 5a, b). Furthermore, levels of both lactate and pyruvate were enhanced in activated Gch1-deficient T cells, indicating augmented glycolysis (Extended Data Fig. 5c), and suggesting that the loss of GCH1 expression affects mitochondrial respiration.
Following anti-CD3/CD28 stimulation, mitochondrial respiration and oxygen consumption were much lower in BH4-deficient T cells than in control cells (Fig. 2i and Extended Data Fig. 5d–g). Cytochrome c—a redox-active protein containing haem groups that reversibly alternate between their Fe2+ and Fe3+ oxidation states—is important for the mitochondrial electron-transport chain (ETC), and we confirmed earlier reports17–19 that BH4 efficiently reduces ferri (Fe3+)-cytochrome c to ferro (Fe2+)-cytochrome c at doses that are physiological in activated T cells (Fig. 2j). Critically, we could rescue ETC function by providing reduced cytochrome c to BH4-deficient cells (Fig. 2k, l). Moreover, impaired ETC in activated Gch1-ablated and SPRi3-treated CD4+ T cells was associated with elevation of superoxide reactive oxygen species (ROS; Extended Data Fig. 6a, b). The superoxide scavenger N-acetylcysteine (NAC) only partially rescued the proliferation defect of Gch1-ablated T cells, and NAC addition did not rescue the iron deficiency observed in activated Gchl-ablated CD4+ T cells, nor did it enhance ATP production (Extended Data Fig. 6c–f), suggesting that the enhanced ROS are the result of mitochondrial dysregulation. BH4 is a co-factor for nitric oxide synthase (NOS) enzymes and is required for nitric oxide (NO) production2. However, under our experimental conditions we did not observe detectable expression of inducible NOS or NO production until several days after T-cell activation (Extended Data Fig. 6g–j). Our data indicate that antigen-receptor-stimulated, BH4- depleted T cells display a defective iron-redox cycling of cytochrome c, which leads to mitochondrial dysfunction.
Given that SPRi3 has relatively low potency and a short half-life, we developed a novel SPR inhibitor, QM385, which is structurally distinct from SPRi3 (Fig. 3a). QM385 binds with high affinity to human SPR in a cell-free assay, and efficiently reduced BH4 levels in anti-CD3/CD28 activated mouse splenocytes and in anti-CD3/CD28 activated human peripheral blood mononuclear cells (Extended Data Fig. 7a–c). QM385 is orally bioavailable, has a long half-life (Supplementary Table 2), and dose-dependently reduces plasma levels of BH4 while concurrently increasing levels of sepiapterin (Fig. 3b)—a sensitive biomarker of SPR inhibition1. QM385 did not inhibit a panel of physiologically important targets or closely related reductases (Supplementary Table 3), and in vivo administration did not result in detectable adverse effects. QM385 treatment resulted in markedly less proliferation of CD4+ T cells in vitro (Fig. 3c). Moreover, in activated CD4+ T cells, QM385 substantially reduced ETC function and ATP levels, and led to an elevation in superoxide ROS (Extended Data Fig. 7d–f). Altogether, QM385 phenocopies the effects of SPRi3 in vitro, albeit at much lower concentrations. Importantly, oral administration of QM385 to mice for three days greatly reduced the number of inflammatory T cells and eosinophils in the ovalbumin-induced and house dust mite (HDM) airway allergic inflammation models (Fig. 3d and Extended Data Fig. 7g), which are T cell dependent20. QM385 effectively inhibited the proliferation of human CD4+ T cells at low doses (Extended Data Fig. 7h). We have, therefore, developed a novel SPR inhibitor that blocks T-cell proliferation and autoimmunity at nanomolar potency and with good oral bioavailability, and this or similar compounds could potentially be used to treat T-cell-mediated autoimmune and allergic diseases.
Fig. 3|. Development of an orally available, potent, small-molecule SPR inhibitor to limit BH4 production.

a, Chemical structure of SPRi3 and QM385. b, Dose-dependent reduction in plasma BH4 levels by QM385, and respective dose-dependent increase in plasma sepiapterin levels. BQL, below quantifiable limits (less than 0.3 ng ml–1 for sepiapterin). Data are shown as means ± s.e.m.; n = 6 mice for each condition. c, Left, representative histograms depicting three-day proliferation of stimulated CD4+ T cells with vehicle (n = 4 mice) and QM385 treatment (2.5 μM or 5 μM; n = 3 mice each); and right, quantitative analysis of total cell numbers. Data are shown as means ± s.e.m. The experiment was repeated two independent times with similar results. d, Top, diagram showing the HDM allergy model with dose-response administration of QM385 peritoneally twice a day for three consecutive days as indicated. Bottom, quantification of T cells and eosinophils in BALF. Data are shown as box-and-whisker plots (running from minimal to maximal values), for which individual data points are given. Vehicle, n = 10; 0.3 mg kg–1, n = 5; 1 mg kg–1, n = 4; 3 mg kg–1, n = 6/7. Absolute numbers in the BALF are shown. NS, not significant; *P < 0.05; **P < 0.01; ***P < 0.001 (one-way ANOVA with Dunnett’s multiple comparisons for panels d, e).
To investigate whether an increase in GCH1 and BH4 enhances T-cell function in vivo, we crossed the Lck-Cre driver line to Cre-recombinase- inducible, haemagglutinin (HA)-tagged, human-GCHl-overex-pressing (GOE) mice1, generating GOE;Lck animals (Extended Data Fig. 8a). T-cell development and homeostasis of peripheral CD4+ and CD8+ T cells were unaffected in these mice, although in the periphery there was a marked increase in the proportion of effector T cells (Extended Data Fig. 8b–d). Anti-CD3/CD28-stimulated CD4+ T cells from the GOE;Lck mice had elevated BH4 levels compared with controls (Fig. 4a), and displayed enhanced T-cell proliferation upon activation (Fig. 4b). GCH1 overexpression in unstimulated naïve T cells did not result in proliferation or any overt spontaneous autoimmunity. To confirm that elevated GCH1 increases the proliferation of activated T cells, we used additional T-cell-specific Cre lines to drive GCH1–HA expression, namely the CD4-Cre and tamoxifen-inducible ERT-Cre lines, both of which show enhanced T-cell proliferation and cytokine production (Extended Data Fig. 8e–j). In GOE;Lck mice we also observed more inflammatory cells, including T cells, in the ovalbumin-induced allergic inflammation asthma model (Extended Data Fig. 9a), and greater severity in the in vivo T-cell-transfer colitis model (Extended Data Fig. 9b, c). Overproduction of BH4 in regulatory T cells did not affect their suppressive function in transfer colitis (Extended Data Fig. 9d). Administration of sepiapterin to anti-CD3/ CD28-stimulated CD4+ T cells also increased BH4 levels and enhanced the proliferation of stimulated CD4+ and CD8+ T cells (Fig. 4c and Extended Data Fig. 9e–g). Furthermore, treatment of stimulated CD4+ T cells with BH4 itself increased both proliferation and IL-2 secretion (Extended Data Fig. 9h, i), and the proliferative and S-phase-entry defects observed in Gch1 -ablated T cells were rescued with either sepiapterin (Extended Data Fig. 9e, f) or BH4 (Extended Data Fig. 9j). In activated Gch1 -ablated T cells, sepiapterin supplementation also restored iron levels, reduced superoxide, and increased ATP production (Extended Data Fig. 10a–c), reinforcing that these deficits are due to reduced BH4 levels.
Fig. 4|. Enhanced BH4 production results in enhanced T-cell proliferation and anticancer immunity.
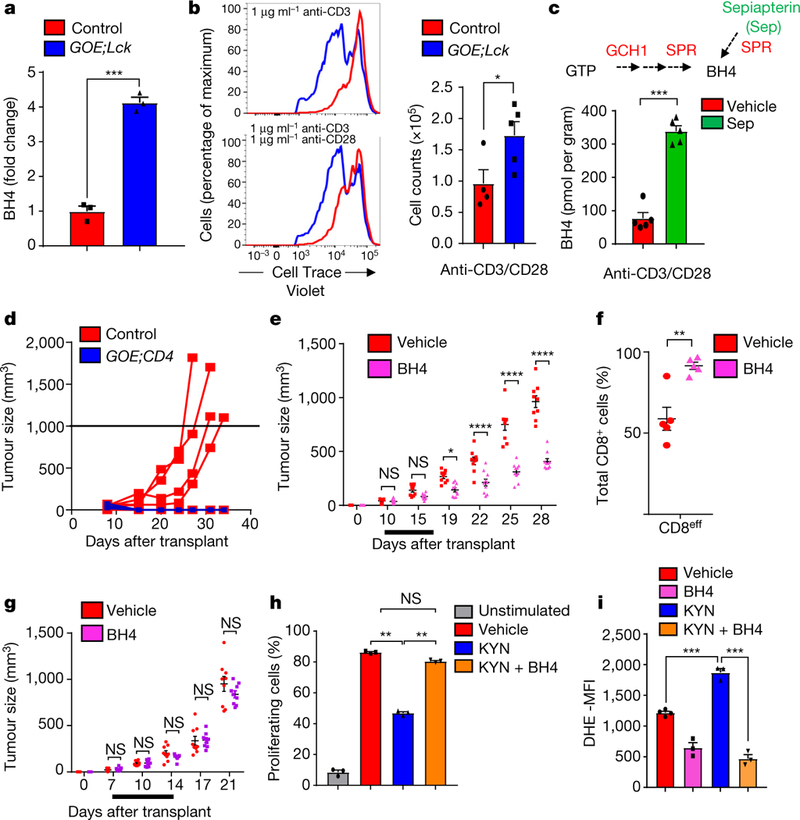
a, Fold change in BH4 levels after 24-hour-stimulation of CD4+ T cells. Data for individual mice (n = 3) are given as means ± s.e.m. b, Representative histograms after three days of CD4+ T cell proliferation, from control (n = 4) and GOE;Lck (n = 5) mice. Data are means ± s.e.m. c, BH4 measurements after 24 hours in activated wild-type CD4+ T cells treated with vehicle or sepiapterin (Sep; 5 μM). Data shown for individual mice (n = 5) are given as means ± s.e.m. d, Breast-cancer model, involving orthotopic injection of E0771 breast- cancer cells into syngeneic control (n = 6) and GOE;Lck (n = 7) mice. e, Effect of BH4 supplementation on cancer growth in the E0771 model. Supplementation with BH4 (n = 10 mice) or vehicle (n = 9 mice) was carried out for seven days as indicated (black line). f, Quantification of intratumoral effector CD8+ T cells (CD44+CD62Llo) assayed from E0071 tumours on day 28 of vehicle treatment (n = 5 mice) or BH4 treatment (n = 5 mice). Data are means ± s.e.m. g, Effect of BH4 supplementation on cancer growth in Rag2−/− female hosts. BH4 and vehicle supplementation (n = 9 mice each) was carried out for seven days as indicated (black line). Data are means ± s.e.m. h, Quantification of proliferation of stimulated CD4+ T cells treated with kynurenine (KYN; 50 μM) or BH4 (10 μM). Data are means ± s.e.m.; n = 3 samples for each condition. Experiment was repeated two independent times with similar results. i, Quantification of the mean fluorescent intensity (MFI) of dihydroethidium (DHE, a superoxide ROS indicator) in stimulated wild-type CD4+ T cells treated with vehicle, KYN (50 μM), BH4 (10 μM) or KYN (50 μM) plus BH4 (10 μM) for 20 hours. Data are means ± s.e.m.; n = 3 samples for each condition. The experiment was repeated two independent times with similar results. NS, not significant; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001 (two-tailed Student’s t-test for panels a, b, c, f; two-way ANOVA with Sidak’s comparison for panels d, g; one-way ANOVA with Tukey’s comparison for panels h,1).
To address whether hyperactivation of the BH4 pathway in T cells promotes anticancer immunity, we orthotopically injected E0771 breast-cancer cells into syngeneic mice to generate mammary tumours21. GOE;CD4 mice, unlike controls, completely rejected tumour growth (Fig. 4d). Moreover, treatment of mice carrying established E0771-derived mammary tumours with BH4 slowed the growth of the tumours (Fig. 4e). Tumours in BH4-treated mice displayed increased frequencies of activated effector CD4+ and CD8+ cells among the infiltrating T cells, compared with vehicle-treated mice (Fig. 4f and Extended Data Fig. 10d). BH4 treatment in Rag2−/− hosts had no effect on breast-cancer growth, confirming that the effect of BH4 is via effects on the adaptive immune system (Fig. 4g). We validated these results with a second orthotopic model, the TC-1 cancer line (Extended Data Fig. 10e–g). Kynurenine—a tryptophan metabolite—inhibits T-cell proliferation22. Interestingly, xanthurenic acid, a kynurenine metabolite, blocks SPR activity23. We found that kynurenine treatment inhibits SPR in activated T cells, as shown by increased sepiapterin levels (Extended Data Fig. 10h). Adding kynurenine to T-cell cultures also reduced T-cell proliferation and increased ROS in activated T cells, both of which were fully restored by addition of BH4 (Fig. 4h, i and Extended Data Fig. 10i, j).
In conclusion, we have revealed that BH4 is required for the effective proliferation of mature T cells in vitro and in vivo, and that this is mechanistically linked to iron metabolism and mitochondrial respiration. Of relevance in this context, nutritional iron deficiency is associated with impaired T-cell proliferation and delayed-type hypersensitivity responses, while humoral immunity is largely preserved24,25. Iron-deficiency anaemia is also associated with an increased incidence of cancer26,27. Notably, we have further found that BH4 is required for T-cell-driven autoimmunity and allergic inflammation and that its inhibition by kynurenine links the immunosuppressive tumour environment to impaired T-cell function. Moreover, increasing BH4 levels can overcome this inhibition to enhance immunity and inhibit tumour growth. Therefore, blocking the synthesis of BH4 could be a viable way to abrogate proinflammatory auto-aggressive T cells in T-cell-driven pathological diseases, whereas its elevation could be a novel way to enhance antitumour immunity.
METHODS
Mice.
Mice expressing enhanced GFP (eGFP) under the Gch1 promoter were used to label cells that upregulate Gch1 after T-cell activation. Mice with a Cre-dependent GCH1-HA overexpression cassette to induce BH4 overproduction, and Gch1 floxed mice in which BH4 production is prevented, have been previously reported1,5. For both gain- and loss-of-function experiments, we bred GCH1-HA and Gch1 floxed mice to the T-cell-specific lines LCK-Cre28, CD4-Cre29 or RORgammact-Cre, the ubiquitous tamoxifen-inducible Rosa26-CreERT2 line30, and the B-cell-specific line MB1-Cre. All animal experiments were approved by the Austrian Animal Care and Use Committee.
Compounds.
Sepiapterin (Sep; 11.225), tetrahydrobiopterin (BH4, 11.212) were purchased from Schircks Labs, Switzerland. For in vitro use, both sepiapterin and BH4 were dissolved in dimethylsulfoxide (DMSO) to a stock concentration of 10 mM. SPRi3 has been developed previously and was used as instructed1. For T-cell assays, sepiapterin was used at a concentration of 5 μM, BH4 at a concentration of 10μM and SPRi3 at a concentration of 50μM unless otherwise stated in the figure legends. For in vivo use, BH4 was reconstituted in sterile saline under argon gas. Kynurenine (K8625) and NAC (A9165) were purchased from Sigma.
Determination of BH4 levels.
Levels of BH4 (tetrahydrobiopterin) and oxidized biopterins (BH2 and biopterin) were determined by high-performance liquid chromatography (HPLC) followed by electrochemical and fluorescent detection, respectively, following an established protocol31. Cell pellets were freeze–thawed in ice-cold resuspension buffer (50 mM phosphate-buffered saline (PBS), 1 mM dithioerythriol, 1 mM EDTA, pH 7.4). After centrifugation at 13,200 r.p.m. for 10 min at 4 °C, supernatant was removed and ice-cold acid precipitation buffer (1 M phosphoric acid, 2 M trichloroacetic acid, 1 mM dithioerythritol) was added. Following centrifugation at 13,200 r.p.m. for 10 min at 4 °C, the supernatant was removed and injected onto the HPLC system. Quantification of BH4 and oxidized biopterins was obtained by comparison with external standards and normalized to protein concentration, determined by the bicinchoninic acid (BCA) protein assay (Pierce).
Determination of sepiapterin levels by HPLC.
Supernatant samples were precipitated by the addition of one volume (1/1, v/v) of 5% trichloroacetic acid (TCA) plus 6.5 mM dithiothreitol (DTE). Afterwards, samples were centrifuged (10,000g for 10 min at 4 °C) and 20 μl was analysed. HPLC analysis of sepiapterin was done using a Beckman System Gold (Beckman Instruments, Fullerton, CA, USA) by using a Waters Atlantis dC-18, 5-μm RP column (4.6 mm × 250 mm; temperature 35 °C), with a flow rate set at 0.5 ml min–1 and isocratic elution of mobile phase (92% phosphate buffer (15 mM); 8% acetonitrile (90%), pH 6.4). Identification and quantification of sepiapterin was done using a multiwavelength fluorescence detector (excitation wavelength 425 nm, emission wavelength 530 nm, module 2,475; Waters, Milford, USA) and expressed as nM of sepiapterin.
Lymphocyte proliferation.
T cells were purified from spleens and lymph nodes of mice using microbeads (CD4+; CD8+, naïve CD4+, Miltenyi Biotec). We coated 96 U-shaped plates with anti-CD3 (4 μg ml–1; Biolegend), with or without anti-CD28 (2 μg ml–1; Biolegend) at the indicated concentrations (unless otherwise stated in the figure legends) in PBS for 3 h at 37 °C. T cells were then plated at at 105 cells per well in Iscove’s modified Dulbecco medium (IMDM) plus penicillin streptomycin plus L-glycine plus 10% fetal calf serum (FCS). Beta-mercaptoethanol was omitted. Phorbol myristate (PMA; 50 ng ml–1) and ionomycin (50 ng ml–1) were also used to stimulate purified T cells for 24 h. Purified and activated T cells were cultured for 24 h; the expression of activation markers (CD62L, CD25, CD44 and CD69) was analysed using flow cytometry; and the supernatant was collected for measurement of IL-2 and IFNγ concentrations using ELISA kits (Biolegend). Purified T cells were also stained with the Cell Violet Trace Proliferation Kit (Invitrogen) and cultured for three days; proliferation was assayed by flow cytometry using viable cells (4,6-diamidino-2-phenylindole (DAPI)-negative). In addition, purified T cells were cultured with purified splenic dendritic cells and soluble anti-CD3 antibody (1 μg ml–1) for three days. To analyse the expression of inducible (i)NOS, we stimulated purified CD4+ T cells, assayed fixed and permeabilized cells at various time points, and stained them for iNOS levels. B cells were purified using microbeads (CD19+; Miltenyi Biotec) from the spleen, loaded with cell tracer, stimulated with lipopolysaccharide (LPS; 1 μg ml–1) and analysed for proliferation as described above. For the class- switch recombination experiment, CD43− B cells were isolated from spleens by magnetic-activated cell sorting (MACS; Miltenyi Biotec) and stimulated for five days with LPS (20 μg ml–1) to induce switching to IgG3 expression. Percentages of switched B lymphocytes were assessed by flow cytometry.
EdU staining.
The cell-cycle status of T cells was assessed using the Click-iT EdU flow cytometry cell proliferation assay (Invitrogen). Briefly, purified CD4+ T cells were activated with anti-CD3 (4 μg ml–1) and anti-CD28 (2 μg ml–1) as described above. EdU was pulsed into the wells for 4 h after 16 h of stimulation. The cells were prepared and stained with EdU as per the manufacturer’s instructions.
Mitochondrial respiration and metabolomics.
Mitochondrial respiratory parameters were measured with high-resolution respirometry (Oxygraph-2k, Oroboros Instruments, Innsbruck, Austria)32. Routine respiration was measured by incubating cells in a buffer containing 110 mM sucrose, 60 mM K-lactobionate, 20 mM K-HEPES, 10 mM KH2PO4, 3 mM MgCl2, 0.5 mM egtazic acid (EGTA) and 1 gl–1 fatty-acid-free bovine serum albumin at 37°C (pH 7.2). Total capacity was induced by titration of carbonyl cyanide-4-(trifluoromethoxy)phenylhydrazone (Sigma Aldrich) in steps of 0.5 μM. To assess complex -I- and complex-II-linked respiration, cells were permeabilized with digitonin (8 μM). Complex-I-linked state 3 respiration was induced by adding 5 mM glutamate/5 mM malate and 1 mM adenosine diphosphate (ADP). Complex-II-linked state 3 respiration was induced with 10 mM succinate after adding the complex I inhibitor rotenone (1 ng ml–1). To restore respiratory function in activated CD4+ T cells, cells were permeabilized with digitonin (12μM) and exogenous reduced cytochrome c (2.5 μM; Abcam, b140219) was added. Respiration rates were obtained by calculating the negative time derivative of the measured oxygen concentration. Oxygen-consumption rates were measured using Seahorse technology. To measure ATP, purified T cells were either left unstimulated or stimulated with plate-bound anti-CD3 (4 μg ml–1) and anti-CD28 (2 μg ml–1) for the times indicted in the Figures. ATP was measured using the CellTiter-Glo Luminescent cell viability assay (Promega). To determine ROS levels, purified T cells were activated with anti-CD3 plate-bound anti-CD3 (4 μg ml–1) and anti-CD28 (2 μg ml–1) for 10 h. Cells were washed once with Hank’s balanced salt solution (HBSS) and stained in 10 μM DHE (Invitrogen) for 30 min at 37 °C. Cells were washed twice with HBSS and assayed by flow cytometry. Profiling of biogenic amines by hydrophilic interaction liquid chromatography/quadrupole time-of-flight (HILIC-QTOF) mass spectrometry was performed on cell pellets and supernatants from unstimulated and TCR-stimulated purified T cells by the West Coast Metabolomics Center (UC Davis). For NO2 measurements, we used the Griess reagent system (TB229, Promega). Purified T cells were stimulated with anti-CD3 and anti-CD28 antibodies as described above, and the supernatant was collected at various time points for nitrite measurements. Peritoneal macrophages stimulated with LPS (100 ng ml–1) for 24 h were used as a positive control.
Flow cytometry.
Antibody labelling of cells was carried out in FACS staining buffer (PBS supplemented with 2% FCS and 2 mM EDTA) on ice for 30 min after blocking Fc receptors. See Supplementary Table 4 for a list of antibodies used in this study. Cells were recorded on an LSR II flow cytometer (BD Biosciences), and data were analysed using FlowJo v10.0.6 software (Tree Star). Absolute splenocyte and thymus numbers were determined by counting total cells with a CASY1 counter, and subsequent calculation of T-cell and B-cell numbers was based on ratios from FACS experiments.
Protein blotting.
Protein blotting was carried out using standard protocols. Blots were blocked for 1 h with 5% bovine serum albumin (BSA) in TBST (1 × Tris- buffered saline (TBS) and 0.1% Tween-20) and were then incubated overnight at 4°C with primary antibodies (see Supplementary Table 4), diluted in 5% BSA in TBST (1/1,000 dilution). Blots were washed three times in TBST for 15 min and were then incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (1/2,500 dilution; GE Healthcare, NA9340V) for 45 min at room temperature, washed three times in TBST for 15 min and visualized using enhanced chemiluminescence (ECL Plus, Pierce, 1896327).
OP9-DL1 co-cultures.
OP9 bone marrow stromal cells expressing the Notch ligand DL-1 (OP9-DL1; provided by J. C. Zúñiga-Pflücker, University of Toronto) were maintained as described33. We plated 104 OP9-DL1 per well in 48-well plates 4–12 h before the start of thymocyte cultures. DN3a thymocytes were sorted as TCRβ– TCRγδ– CD4– CD8a– CD28– CD25hi CD44–/lo cells using a BD FACS Aria sorter. Cell Trace Violet labelling of the sorted cells was performed in 1 μM Cell Trace Violet solution in PBS containing 0.1% BSA for 7 min at 37 °C. Cells were washed with medium containing 20% FCS. Thymocytes were then plated on the OP9-DL1 monolayers in the presence of 5 ng ml–1 Flt3L (a tyrosine-kinase-3 inhibitor). Co-cultures were performed in α-minimal essential medium (αMEM) supplemented with 10 mM HEPES (pH 7.5), 1 mM sodium pyruvate, 100 units per ml penicillin, 0.1 mg ml–1 streptomycin, and 20% heat-inactivated FBS.
Adoptive transfer model of colitis.
We injected 5 × 105 MACS-purified naive CD4+CD62L+ T cells from control and GCH1;Lck mice intraperitoneally into 6- to 8-week-old Rag1−/− mice. After the cell transfer, Rag1−/− recipients were weighed weekly and monitored by mini-endoscopy. For monitoring of colitis activity, we used a high-resolution video endoscopic system (Karl Storz). To determine colitis activity, we anaesthetized mice by injecting intraperitoneally a mixture of ketamine (Ketavest 100 mg ml–1; Pfizer) and xylazine (Rompun 2%; Bayer Healthcare) and monitored them by mini-endoscopy at the indicated time points. Endoscopic scoring of five parameters (translucency, granularity, fibrin, vascularity and stool) was performed (Supplementary Table 5)34. For histological analysis, colonic cross-sections were stained with haematoxylin and eosin (H&E). Immunofluorescence of cryo-sections was performed using the tyramide signal amplification (TSA) Cy3 system (PerkinElmer) and a fluorescence microscope (IX70; Olympus) using primary antibodies against F4/80, MPO, CD3, CD4 and CD11c. In brief, cryo-sections were fixed in ice-cold acetone for 10 min, and then incubated sequentially with methanol, avidin/biotin (Vector Laboratories) and protein-blocking reagent (DAKO) to eliminate unspecific background staining. Slides were then incubated overnight with primary antibodies specific for the respective antigen. Subsequently, the slides were incubated for 30 min at room temperature with biotinylated secondary antibodies (Dianova). All samples were finally treated with streptavidin-HRP and stained with tyramide (Cy3) according to the manufacturer’s instructions (Perkin Elmer). Before examination, nuclei were counterstained with Hoechst 3342 (Invitrogen). For experiments involving transfer of regulatory T cells35, 500,000 conventional T cells (CD4+ CD25- CD45RBhi) and 500,000 regulatory T cells (CD4+ CD25+ CD45RBlo) were transferred intraperitoneally into Rag2−/− hosts. For GOE;CD4 CD4+ T cells, 150,000 cells were transferred. Body weights were monitored over the course of the experiment35.
OVA immunization and airway hyperresponsiveness.
For the OVA immunization study, immunization was performed using 100 μg OVA per mouse in 200 μl Alum intraperitoneally. Blood was collected from the tail vein 14 days after injection to check IgG and IgM titres. Three weeks later, a further intraperitoneal injection was carried out and again blood collected two weeks later to measure the rechallenge responses. For measurements of lung function, deeply anaesthetized mice (pentobarbital (60 mg kg–1) underwent a tracheotomy with a 20G sterile catheter. A computer-based analysis of airway hyperresponsiveness was then performed using a Flexivent (SCIREQ) apparatus10. Mice were ventilated at a tidal volume of 9 ml kg–1 with a frequency of 150 b.p.m.; positive end-expiratory pressure was set at 2 cm H2O. Lung resistance and elastance of the respiratory system was determined in response to in-line aerosolized methacholine challenges (0, 1, 3, 10, 30 and 100 mg ml–1). Methacholine was dissolved in sterile PBS. The mean elastance and resistance of ten measurements by doses was calculated. For bronchoalveolar lavage (BAL) on day 21, mice were anaesthetized following an intraperitoneal injection of urethane (200 μl; 35%) and a 20G sterile catheter inserted longitudinally into the trachea. We injected 2 ml of ice-cold PBS containing protease inhibitors (Roche) into the lungs, harvested and stored on ice. BAL fluid underwent a 400g centrifugation (15 min; 4 °C), the supernatant was discarded and cells were resuspended in 200 μl BAL fluid10. Bronchoalveolar lavage fluid (BALF) cells were resuspended in FACS buffer (PBS, 2% FCS, EDTA), and incubated with Fc block (0.5 mg ml–1; 10 min; BD Biosciences). Cells were then stained with monoclonal antibodies (FITC-conjugated anti-mouse CD45, BD Biosciences, catalogue number 553,079; phycoerythrin (PE)-conjugated anti-mouse Syglec-F, BD Biosciences, catalogue number 552,126; allophycocyanin (APC)-conjugated anti-mouse GR-1, eBiosciences, catalogue number 17–5931-81; PE-Cy7-conjugated anti-mouse CD3e, catalogue number 25–0031-81; peridinin chlorophyll protein complex (PerCP)-conjugated anti-mouse F4/80, BioLegend, catalogue number 123,125; 45 min, 4 °C on ice) before data acquisition on a FACS Canto II (BD Biosciences). A leukocyte differential count was performed during flow-cytometry analysis of cells expressing the common leukocyte antigen CD45 (BD Pharmigen; catalogue number 553,079). Specific cell populations were identified as follows: macrophages as F4/80Hi Ly6gNeg, eosinophils as F4/80Int Ly6gLo SiglecFHi, neutrophils as F4/80Lo Ly6gHi SiglecFNeg, and T lymphocytes as F4/80Neg Ly6gNeg CD3Pos. Total BAL cell counts were performed using a standard haemocytometer, with absolute cell numbers calculated as total BAL cell number multiplied by the percentage of cell subpopulation as determined by FACS10.
HDM allergy model.
For HDM-induced lung inflammation, C57Bl/6 animals (female, 6–12 weeks old) were sensitized for two consecutive days with 25 μg HDM extract (Dermatophagoides pteronyssinus, Greer Laboratories, XPB82D382.5) intra-nasally Six days after the last sensitizing dose, mice were challenged with 12.5 μg of HDM for five consecutive days, with 3 mg kg–1 QM-760 (in 1% Tween-80 and 0.5% sodium carboxymethyl cellulose; Sigma-Aldrich, St Louis, MO, USA) administered by oral gavage, twice daily on days 3–5 during the challenge phase. BALF was removed and analysed for the following immune-cell subsets three days after the last challenge—T cells: CD45+, Thy1+, CD3+, CD11b–, Siglec-F–, Ly6C/G–; eosinophils: CD45+, Thy1+, CD11b+, Siglec-F+, CD3–, Ly6C/G–.
Skin hypersensitivity.
The skin-contact hypersensitivity model was performed as described11. Briefly, to induce contact hypersensitivity, mice were sensitized on day zero by applying 100 μl of 7% 2,4,6-trinitrochlorobenzene (TNCB; Sigma)/acetone or acetone alone as a vehicle control on the shaved abdomen. On day five, mice were challenged on the dorsum of both ears with 20 μl of 1% TNCB/acetone. Ear thickness was measured immediately before and 24 h after the challenge.
Experimental autoimmune encephalitis.
Experimental autoimmune encephalitis (EAE) was induced in control and Gch1;Lck mice by immunization with an emulsion of 100 μg myelin oligodendrocyte glycoprotein (MOG)35–55 in complete Freund’s adjuvant (CFA), supplemented with 5 mg ml–1 Mycobacterium tuberculosis (Difco). We injected 100 μl MOG/CFA subcutaneously above the inguinal lymph node on both sides of the mouse. We then injected 200 μl pertussis toxin/ PBS (50 ng μl–1; List Biological Labs) intraperitoneally per mouse on days zero and one. Scoring for EAE was performed as described over the course of 45 days36.
Orthotopic cancer models.
E0771 cells were orthotopically injected into syngeneic control and GOE;Lck mice as described21. In brief, cells were harvested for injection into mice by trypsin digestion for 5 min, washed in HBSS, counted, diluted in this salt solution and orthotopically injected into the fat pad of the fourth mammary gland (2.5 × 105 cells per 200 μl per mouse). BH4 administration was delivered intraperitoneally (100 mg kg–1) after tumours were palpable (day 10) and treatment was continued for seven days. Tumours were measured using digital calipers; the size of the tumour was expressed as length (mm) by width (mm) by height (mm) equals tumour size (mm3). The tumour cell line TC-1 was derived from primary lung epithelial cells of C57BL/6 mice. The cells were immortalized with the amphotropic retrovirus vector LXSN16E6E7 and subsequently transformed with the pVEJB plasmid expressing the activated human c-H-RAS oncogene37. This cell line was treated and injected into wild-type and Rag2−/− mice as described above. After collagenase/dispase digestion of the tumours, intratunoral effector CD4+ and CD8+ T cells were characterized by flow cytometry (CD62Llo, CD44hi).
Microarray analysis.
Purified CD4+ T cells from control and Gch1;Lck mice were stimulated with plate-bound anti-CD3 (4 μg ml–1) and anti-CD28 (2 μg ml–1) antibodies for 16 h, and total RNA was extracted by sequential Qiazol extraction and purification through the RNeasy micro kit with on-column genomic DNA digestion (Qiagen). RNA quality was determined by an Agilent 2100 Bioanalyzer using the RNA Pico Chip (Agilent). RNA was amplified into complementary DNA using the Ambion wild-type expression kit for whole-transcript expression arrays, with poly-A controls from the Affymetrix Genechip Eukaryotic Poly-A RNA control kit. Images from Agilent arrays were processed using Agilent Feature Extraction Software 10.7.3.1. Raw intensity data were processed in R v3.4.0 using limma v3.34.3, applying normexp background calculation, lowess within-array and Aquantile between-array normalization methods. The normalized values were used to calculate log2-transformed Cy5/Cy3 ratios. Differential-expression analysis was performed by fitting a linear model to the normalized data and computing empirical Bayes test statistics in limma, accommodating a mean-variance trend38. The false-discovery rate was controlled by Benjamini-Hochberg adjustment. The data discussed herein have been deposited in the National Center for Biotechnology Information (NCBI)’s Gene Expression Omnibus39 and are accessible through GEO accession number GSE108101 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE108101).
Ferric/ferrous reduction.
The enzymatic activity of BH4 was assayed as described40. The enzymatic conversion of qBH2 to BH4 was followed by the reduction of ferricytochrome c to ferrocytochrome c by BH4. Ferricytochrome c reduction was determined by reading the increasing ferrocytochrome c absorbance signal at 550 nm. The experiment was run for 40 min at pH 7.4 and recorded at 10-s intervals in 200 μl buffer containing 50 μM ferricytochrome c, 1 μM 6-methyltetrahydropterin (6MPH4), 20 nM DHPR, 50 μM NADH and selected inhibitors. A control lacking DHPR was run in parallel to assess the rate of non-enzymatic reduction of qBH2 by NADH. The extinction coefficients used for ferrocytochrome c and NADH are respectively 29,500 (reduced, 550 nm, H2O) and 6,220 (340 nm, H2O) (in l mol–1 cm–1). Generally, 50 μM of ferrocytochrome c and ferricytochrome cin buffer were measured in isolated wells to assess the completion of the reaction.
Iron measurements.
Total iron content was measured as described41. Briefly, intracellular iron measurements were carried out by using a PerkinElmer Analyst 800 equipped with a transversely heated graphite atomizer (THGA). A Zeeman-effect background correction was realized by a 0.8-T magnetic field, oriented longitudinally with respect to the optical path. A PerkinElmer Lumina single-element iron hollow cathode lamp was driven at a constant current of 30 mA after proper equilibration (that is, for 20 min or more). For the absorption measurements, the 248.3-nm line (spectral bandwidth 0.7 nm) was chosen. FACS-purified naïve CD4+ T cells from control and Gch1;Lck mice were left untreated or stimulated (anti-CD3 and anti-CD28) for 12 h. The cells were then pelleted and frozen at –80 °C. Next, the samples were suspended in 200 μl of a 0.1% (v/v) solution of nitric acid (Rotipuran Supra, 69%, Carl Roth GmbH, Karlsruhe, Germany) in high-purity water (Milli-Q, Merck-Millipore, Darmstadt, Germany) by extended periods (that is, for 30 min or more) of vortexing and ultrasonication at 30–40 kHz. After an initial estimation of the sample’s iron quantity, a five-point linear calibration was established in the range between 0 (that is, less than 0.004 μM) and 0.106 μM. The calibration standards were prepared by diluting a 0.1 M standard stock solution of (NH4)2Fe(SO4)2 (Merck-Millipore, Darmstadt, Germany) with a 0.1% (v/v) aqueous solution of nitric acid (vide supra). The absence of detectable iron (that is, less than 0.004 μM) in the dilution agent, as well as in the sample cups, and the glassware was verified throughout the analyses. A linear fit of the 15 data points (k = 0.978, d = 0.006 μM) yielded a coefficient of determination of 0.992 where k is the slope of the linear fit and d is the axis intercept on the y-axis. Samples with iron concentrations exceeding the calibration range (that is, 0.106 μM or more) were diluted appropriately. The blank solution, the calibration standards and the samples were supplied to the atomizer in randomized fashion as triplicates, using a PerkinElmer AS-800 autosampler with an injection volume of 20 μl. The solvent was evaporated by a slow temperature gradient to 130 °C; ashing took place at a maximum temperature of 1,000 °C; and the atomization profile was read at 2,000 °C. The graphite tube, which was protected against oxidation by high-purity argon (99.999%; Messer Austria GmbH, Gumpoldskirchen, Austria), was cleaned out after each analysis at 2,450 °C. The integrity of each analysis was verified by a visual inspection of the respective time-dependent atomization profile.
Human T-cell-proliferation assays.
Proliferation of PBMCs, obtained from healthy blood donors, was assessed following cell exposure to Dynabeads human T-activator CD3/CD28 (bead/cell ratio 1/2) and IL-2 (30 international units per ml), in the absence or presence of vehicle (DMSO) or SPRi3 (50 μM). PBMCs were resuspended in RPMI 1640 medium supplemented with 2 mM L-glutamine, 100 units per ml penicillin, 100 mg ml–1 streptomycin, 1% non-essential amino acids and 10% FBS, seeded at 2.5 × 105 cells per well and cultured for five days. For the last 18 h of culture, cells were pulsed with 0.25 mCi per well 3H-thymidine. Incorporated thymidine was measured by liquid scintillation spectroscopy. In addition, we also determined the proliferation of alloreactive human T cells. PBMCs from a healthy donor were stimulated with M21 tumour cells. Alloreactive T cells (based on MHC mismatch) were cultured for two weeks. Afterwards, effector CD4+ T cells were sorted (with regulatory T cells excluded), labelled with carboxyfluorescein succinimidyl ester (CFSE), and stimulated with anti-CD3 and anti-CD28 antibodies for five days, supplemented with either DMSO or SPRi3 (50 μM). For QM385 studies, PBMCs from two donors were stimulated with plate-bound anti-CD3 and anti-CD28 antibodies (1 μg ml–1 each). On day three of stimulation, the number of CD4+ T cells was counted by FACS. PBMCs were isolated from healthy subjects (from the Blood Donor Center at the Children’s Hospital Boston, MA, USA). Human studies received Institutional Review Board (IRB) approval (number 2011P000202) from the Beth Israel Deaconess Medical Center Ethics Committee, and written consent was obtained from all study participants before inclusion in the study.
QM385 compound analysis.
For the SNAP (soluble NSF attachment proteins)-based competition time-resolved FRET (TR-FRET) assay, purified SNAP-SPR and sulfasalazine–SNAP–meGFP were labelled with a twofold excess of benzylguanine/terbium cryptate conjugate (K2-benzylamide-BG; Cisbio) or benzylguanine/sulfasalazine (BG-SSZ), respectively, and purified with NAP5 columns (GE Healthcare) to remove excess labelling reagents. The final reaction mixture contained 2.0 nM terbium–SNAP–SPR, 70 nM SSZ–SNAP–meGFP, 10 μM NADPH and 10 μM NADP+ in buffer A (50 mM HEPES-NaOH pH 7.4, 0.15 M NaCl, 0.5 μg μl−1 BSA, 0.05% Triton X-100, 1 mM DTT). Signal was measured after 3 h of incubation with varying concentrations of QM385, using Infinite F500 (TECAN). The excitation wavelength was 320 nm and emission wavelengths were 485 nm (K2-benzylamide-BG; Cisbio) and 520 nm (BG-SSZ) respectively. For BH4 measurements, 50,000 human PBMCs were plated in 96-well plates coated with 1 μg ml–1 human anti-CD3 antibody. Cells were incubated with 1 μg ml–1 soluble human anti-CD28 antibody for 48 h with varying doses of QM385 as indicated in Extended Data Fig. 7c. Cells were then harvested for liquid chromatography mass spectrometry (LC-MS) BH4 measurements. Similar experiments were performed on anti-CD3/28-activated mouse splenocytes.
Plasma levels of QM385.
To formulate 10 mg kg–1 of QM385 for oral administration, we added 6.312 ml of 1% Tween-80 plus 0.5% hydroxypropyl methylcellulose (HPMC) in 50 mM carbonate buffer (pH 9.0) into a tube containing 7.98 mg QM385, then vortexed the mixture for 1–2 min and sonicated it for 30–35 min. Solutions were prepared just before use. The intravenous dosing solution was prepared in 10% dimethylacetamide (DMAC), 10% solutol HS15, 80% (10% (2-hydroxypropyl)-(3-cyclodextrin) in saline (w/v). Approximately 100 μl of blood sample was collected via the tail vein into EDTA-2K tubes. The blood samples were maintained in wet ice first, and centrifuged to obtain plasma (2,000g, 4°C, 5 min) within 15 min of sampling. An aliquot of 30 μl serum sample was added with 100 μl acetonitrile containing 10 ng ml–1 dexamethasone. The mixture was vortexed for 2 min and centrifuged at 14,000 r.p.m. for 5 min. An aliquot of 10 μl supernatant was injected for LC-MS/MS analysis. We also prepared a calibration curve of 0.100–1,000 ng ml–1 for QM385 in diluted blood from C57BL/6 mice.
Statistical analyses.
All values are expressed as means ± s.e.m. Details of the statistical tests used are stated in the figure legends. Briefly, Student’s t-test was used to compare between two groups. One-way ANOVA followed by Dunnett’s post-hoc test for multiple comparisons was used for analysis between multiple groups. Two-way ANOVA was used to compare two groups over time. In all tests, P ≤ 0.05 was considered significant.
Extended Data
Extended Data Fig. 1|. Upregulation of Gchl and BH4 in activated T cells.

a, Percentage of CD62L10 GFP+ cells from purified Gch1-Gfp CD4+ T cells stimulated for 24 h with PMA/ionomycin (50 ng ml–1 each). Data are shown as means ± s.e.m., from n = 3 samples. The experiment was repeated two independent times. ***P < 0.001 (two-tailed Student’s t-test). b,c, Representative Gch1-Gfp expression in 16-h-activated (CD62Llo) CD4+ T cells after anti-CD3/CD28 stimulation (b) and representative dose-response of anti-(α)CD3/CD28 stimulation of purified CD4+ Gch1- Gfp T cells for 24 h (c). The experiment was repeated two independent times with similar results. d, Cell numbers of various immune populations in the thymus (left) and spleen (right) from control (n = 3) and Gch1;Lck (n = 3) 8-week-old mice. Data from individual mice are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). e, f, CD4+ (e) and CD8+ (f) T-cell proliferation after three days of anti-CD3/28 stimulation, from control and Gch1;Lck mice. g, Representative histogram depicting the proliferation of DN3a thymocytes from control and Gch1;Lck mice cultured on OP9-Dl1 stromal cells for five days. The experiment was repeated two independent times with similar results. h, i, Representative FACS blot depicting the differentiation into CD4+ and CD8+ T cells of DN3a thymocytes from control and Gch1;Lck mice cultured on OP9-Dl1 stromal cells for five days (h), and quantification of the differentiated cell types from n = 3 animals (i). Data from individual mice are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 2|. Normal T-cell development and B-cell biology in the absence of Gch1.

a, Thymocyte cell death induced over 24 h by various stimuli: anti-CD3 (0.5 μg ml–1 and 5 μg ml–1), Fas ligand (0.2 μg ml–1 and 2 μg ml–1), dexamethasone (Dex, 0.1 μg ml–1 and 0.5 μg ml–1) and γ-irradiation (1 Gray (Gy)). Data are shown as means ± s.e.m. n = 3 for each genotype. NS, not significant (two-tailed Student’s t-test). b, Death by neglect of purified CD4+ T cells cultured without stimulation for up to 56 h. Data are shown as means ± s.e.m. n = 3 for each genotype. NS, not significant (two-tailed Student’s t-test). c, d, Proliferation of CD4+ T cells from control and Gch1;RORc mice after three days of anti-CD3/28 stimulation. Panels show representative FACS proliferation traces (c) and representative dose response (d). Experiments were repeated independently more than six times with similar results. e, Representative FACS plots from spleens of control and Gch1;MB1 mice. MB1-Cre is an early B-cell deleter Cre line using endogenous CD79a B-cell specific expression. The experiment was repeated two independent times with similar results. f, g, Representative FACS histogram depicting the proliferation of wild-type B cells treated with vehicle (DMSO) or SPRi3 (50 μM) (f), and of B cells from control and Gch1;MB1 mice in response to LPS (1 μg ml–1) after three days (f). Shaded grey peaks represent unstimulated cells. FACS plots are representative of two independent experiments showing similar results. n = 3 mice per group. h, Class-switch recombination. FACS analysis of splenic CD43- B cells from control and Gch1;MB1 mice stimulated with LPS (20 μg ml–1) for five days to induce class-switch recombination to IgG3. FACS plots are representative of two independent experiments showing similar results.
Extended Data Fig. 3|. Development of regulatory T cells and their function in Gch1-ablated mice.

a, b, Representative FACS plot depicting CD4+ FoxP3+ regulatory T cells (T regs; a) and quantification of T-reg proportions as well as absolute numbers in the spleen (b) of control and Gch1;RORc mice (n = 6 each). Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). c,d, In vitro T-reg suppression assay, in which naïve, wild-type CD4+ T cells were activated in the presence of varying ratios of T-reg cells from control and Gch1;RORc mice for four days. Representative histogram showing the suppressive capacity of control and Gch1;RORc T-reg cells (c) and quantification of proliferation with various ratios of T-reg cells (d). n = 4 samples. Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; NS, not significant (two-tailed Student’s t-test with multiple comparisons). Tconv, conventional CD4+ T cells (CD4+, CD25neg, CD45RBhi). e, Naive CD4+ transfer colitis model, with co-transfer of FACS-purified T-reg cells from control (n = 4) and Gch1;RORc (n = 4) mice. As a control, Tconv cells (from n = 16 mice) with no co-transfer of T-reg cells were used. Changes to initial body weight (BW) were scored over five weeks. Data are shown as means ± s.e.m. *P < 0.05; ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). f, Total numbers of CD4+ splenic T cells at two weeks post-transfer in mice (n = 3) transferred with naïve CD4+ cells only (‘no T regs’) and mice transferred with T regs from control or Gch1-ablated (Gch1;RORc) mice. Data are shown as means ± s.e.m. ***P < 0.001; ****P < 0.0001; NS, not significant (one-way ANOVA with Dunnett’s multiple comparison test). g, Transfer colitis model of intestinal autoimmunity. Body-weight changes are plotted relative to initial weight in mice transferred with naïve CD4+ T cells from control or Gch1;Lck mice (n = 10 each). Data are shown as means ± s.e.m. NS, not significant (two-way ANOVA with Sidak’s multiple comparisons). h, i, Proportion of CD4+ T cells in the draining mesenteric lymph nodes in week 4 (h), and profiles of ntracellular cytokines (IFN-γ and IL-17) from transferred control and Gch1;Lck cells (i). Data are shown as means ± s.e.m. n = 10 for each genotype for panel h and n = 5 for each genotype for panel i. ***P < 0.001; NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 4|. Blockage of GCH1/BH4 abrogates T-cell- mediated autoimmunity.

a, OVA immunization of control and Gch1;Lck mice. T-cell-dependent IgG responses and T-cell-independent IgM responses are shown two weeks after OVA immunization (left panels, 100 (μg OVA in 200 μg alum) as well as two weeks after re-challenge (right panels). n = 5 for control mice; n = 6 for Gch1;Lck mice. Data are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (tw-tailed Student’s t-test with multiple comparisons). b,c, EAE model of autoimmunity towards the central nervous system. Data are shown as means ± s.e.m. b, EAE scores of control and Gch1;Lck mice. n = 6 for each genotype. ****P < 0.0001 (linear regression analysis was performed on the slope of each curve). c, Mean maximal EAE severity in control and littermate Gch1;Lck mice. *P < 0.05 (Mann-Whitney test). d, Schematic representation of the de novo, salvage and recycling arms of the BH4 pathway. The dotted arrow indicates non-enzymatic reactions; solid arrows indicate enzymatic reactions. DHFR, dihydrofolate reductase; GTP, guanosine triphosphate; PCDB, pterin-4α-carbinolamine dehydratase; PTPS, 6-pyruvoyl tetrahydropterin synthase; QDPR, quinoid dihydropteridine reductase; SPR, sepiapterin reductase. e, Representative FACS plots depicting activation marker profiles of purified wild-type control CD4+ T cells left unstimulated or stimulated with anti-CD3/28 antibodies for 16 h and then treated with vehicle (DMSO), SPRi3 (50 μM) or sepiapterin (5 μM). The experiment was repeated two independent times with similar results. f, Cell survival as defined by the percentage of DAPI– annexinV– cells from purified CD4+ T cells stimulated for 24 h or 48 h with anti-CD3/28 antibodies and then treated with vehicle (DMSO), SPRi3 (50 μM) or sepiapterin (5 μM). The experiment was repeated two independent times with similar results. g, h, Representative FACS blots depicting EdU cell-cycle analysis after 28hours anti-CD3/CD28 stimulation of control, Gch1;RORc, and SPRi3-treated control CD4+ T cells. EdU was pulsed for the last 4 hours (g) and quantification of S-phase entry (h). Data from individual mice are shown ± s.e.m.. ***P < 0.001 (One-way ANOVA with Dunnett’s multiple comparisons test). i, Quantification of subG1 (dead cells) populations after 24- and 48-hour stimulation. EdU was pulsed for the last 4 hours of each time point. Data from individual mice are shown ± s.e.m.). **P < 0.01; NS, not significant (Multiple t-test comparisons). j, k, Amino-acid profiles in the supernatants (j) and cell pellets (k) from 24-h anti-CD3/CD28-stimulated CD4+ T cells from control and Gch1;Lck mice. n = 3 for each genotype. Data are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test).
Extended Data Fig. 5|. Mitochondrial dysfunction in BH4-depleted T cells after activation.

a, b, ATP measurements in control (n = 3) and Gch1;Lck (n = 3) CD4+ T cells (a) and in wild-type CD4+ T cells treated with DMSO vehicle (n = 3) or SPRi3 (50 μM; n = 3) (b), either left unstimulated or assayed at the indicated time points after T-cell activation with anti-CD3/28 antibodies. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). c, Metabolomic measurements of lactate and pyruvate levels in cell pellets of 16-h anti-CD3/28-activated CD4+ T cells from control and Gch1;Lck mice. Data are shown as means ± s.e.m. n = 4 for each genotype. *P < 0.05 (two-tailed Student’s t-test). d, Routine and total capacitance oxygen respiration in intact, 16-h anti-CD3/CD28- stimulated CD4+ T cells from control and Gch1;Lck mice. Data from individual mice are indicated ± s.e.m. n = 4 for each genotype. **P < 0.01 (two-tailed Student’s t-test). e, f, Oxygen uptake rate in permeabilized, 16-h anti-CD3/CD28-stimulated CD4+ T cells from control (n = 4) and Gch1;ROR (n = 4) mice (e) and wild-type CD4+ T cells treated with DMSO or SPRi3 (50 μM (n = 5 each) (f). Data from individual mice are indicated ± s.e.m. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test).g, Left, representative oxygen consumption traces of complex-I-linked and complex-II-linked ETC activity from 16-h-activated wild-type CD4+ T cells treated with vehicle or SPRi3 (50 μM). Right, relative complex-I- and complex-II-linked activities in activated control cells treated with vehicle (n = 4) or SPRi3 (50 μM; n = 4). Data are shown as means ± s.e.m. NS, not significant; *P < 0.05 (two-tailed Student’s t-test).
Extended Data Fig. 6|. Enhanced superoxide levels independent of iNOS coupling observed in BH4-deficient activated T cells.

a, b, Representative FACS histogram (a) and quantification of the mean fluorescent intensity (MFI; b) showing levels of DHE (dihydroethidium, a superoxide ROS indicator) in unstimulated and 20-h anti-CD3/28- activated CD4+ T cells from control and GCH1;RORc mice as well as control cells treated with SPRi3 (50 μM). n = 3 samples per group. The experiment was repeated three independent times with similar results. c, d, Proliferation of control (n = 6) and Gch1;Lck (n = 9) CD4+ T cells and treatment with the superoxide scavenger NAC (500 μM; n = 4 each). Representative three-day proliferation histograms are shown in panel c; quantification is shown in panel d. Data are given as means ± s.e.m. Individual mice for each genotype are shown. ****P < 0.0001 (one-way ANOVA with Tukey’s multiple comparison test). e, Total iron content from unstimulated or 24-h anti-CD3/28-stimulated CD4+ T cells (untreated or treated with 500 μM NAC) from control (n = 17, 4, respectively and Gch1;RORc (n = 22, 6, respectively) mice. Data are shown as means ± s.e.m. Individual mice for each genotype are shown. **P < 0.01 (two-tailed Student’s t-test with Tukey’s multiple comparisons). f, ATP measurements from stimulated wild-type CD4+ T cells treated with DMSO, sepiapterin and NAC for 24 h. Data are shown as means ± s.e.m. n = 5 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). g, Intracellular iNOS expression in purified CD4+ control T cells left untreated or anti-CD3/CD28-stimulated for 12 h, 24 h or 72 h. The experiment was repeated two independent times with similar results. h, i, Representative histogram showing iNOS expression in control and Gch1 -ablated CD4+ T cells stimulated with anti-CD3/CD28 antibodies for 72 h (h) and the percentage of iNOS+ cells was quantified over time (i). n = 4 for each genotype. Data are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). j, Nitrite measurements in the supernatant of stimulated cells from panel i. Peritoneal, thioglycollate- elicited macrophages stimulated with LPS (100 ng ml–1) for 24 h were used as a positive control. Data are shown as means ± s.e.m. N = 4 for each genotype. NS, not significant (2-tailed, Student’s t-test).
Extended Data Fig. 7|. Functional evaluation of the SPR blocker QM385.

a, The BH4 pathway, indicating how QM385 acts on SPR, limiting BH4 production and correspondingly increasing sepiapterin levels, which can be used as a biomarker for QM385-mediated SPR inhibition. b, c, A representative concentration-response curve showing the binding affinity of QM385 to human SPR, tested in vitro by TR-FRET (b); and reduction of BH4 levels upon QM385 treatment in anti-CD3/28-stimulated mouse splenocytes (left panel, two independent experiments) and human PBMCs (right panel, two independent experiments) (c). Indicated in red are the calculated IC50 values for each assay. The binding-effect assay was repeated 162 independent times with similar results. d, The oxygen-uptake rate in permeabilized, 16-h anti-CD3/CD28-stimulated wild-type CD4+ T cells treated with DMSO or QM385 (2.5 μM). Data from individual mice (n = 3) are indicated ± s.e.m. ***P < 0.001 (two-tailed Student’s t-test). e, ATP measurements of unstimulated (n = 8) and 24-h-activated wild-type CD4+ T cells treated with DMSO vehicle (n = 4) or varying doses of QM385 (n = 4 for each dose). Data are shown as means ± s.e.m. NS, not significant; **P < 0.01 (one-way ANOVA with Dunnett’s multiple comparisons). f, Fold changes in DHE levels between CD4+ T cells treated with DMSO or QM385 (2.5 μM) and activated for 20 h. Data from individual mice (n = 4) are indicated ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). g, Allergic airway inflammatory disease model and quantification of inflammatory cells in bronchoalveolar lavage fluids (BALFs). Data are shown as box-and-whisker plots (running from minimal to maximal values); individual data points are shown. n = 15 for vehicle-treated mice; n = 17 for QM385-treated mice. QM385 (1 mg kg–1) was administered orally (peritoneally) twice a day for three consecutive days as depicted in the diagram. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). h, Proliferation of human CD4+ T cells from two donors performed in triplicate samples. Anti-CD3/28 T cells were stimulated with varying doses of QM385 and total counts were measured. Data are shown as means ± s.e.m. **P < 0.01; P < 0.05 (one-way ANOVA with Dunnett’s multiple comparisons).
Extended Data Fig. 8|. Increased numbers of effector T cells in naïve mice overexpressing Gch1, and enhanced T-cell proliferation after stimulation.

a, Representative immunoblot to detect GCH1 and the HA tag in naïve CD4+ T cells from control and GOE;Lck-overexpressing mice. The experiment was repeated three times with similar findings. b, c, The proportion of splenic T and B cells (b) and the proportion of CD4+ and CD8+ T cells among the splenic T cell (TCRβ+) population (c), from control (n = 4) and GOE;Lck (n = 4) mice. Data for individual mice aged eight weeks are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). d, Quantification of naïve (CD44lo CD62Lhi), memory (CD44hi CD62Lhi) and effector (CD44hi CD62Llo) T-cell subtypes from the spleen of control (n = 6) and GOE;Lck (n = 10) mice. Data for individual mice are shown as means ± s.e.m. **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). e, Cell numbers for B cells, T cells, and CD4+ and CD8+ T cells in the spleens of control and GOE;CD4 mice. Data from individual mice (n = 4 for each genotype) are shown as means ± s.e.m. NS, not significant (two-tailed Student’s t-test). f, Proportion of CD4+ and CD8+ naïve, memory and effector T cells in the spleens of naïve control and GOE;CD4 mice. Data for individual mice (n = 4 for each genotype) are shown as means ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; NS, not significant (two-tailed Student’s t-test). g, Representative histograms depicting dose-dependent proliferation of control and GOE;CD4 CD4+ T cells stimulated for three days with anti-CD3/28 antibodies. Experiments were repeated more than three times with comparable results. h, IL-2 and IFN-γ secretion after three days of stimulation (with anti-CD3/28 antibodies) of control and GOE;CD4 CD4+ T cells. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; ***P < 0.0001 (two-tailed Student’s t-test). i, j, Cells from control (n = 3) and GOE;ERT (n = 3) mice were stimulated with anti-CD3/28 antibodies for three days and treated with 4-hydroxytamoxifen (4-OHT; 0.5 μM) to induce Gch1 overexpression in vitro. i, Quantification of proliferation of CD4+ T cells; j, cytokine secretion. Data from individual mice are shown as means ± s.e.m. **P < 0.01; ***P < 0.001 (two-tailed Student’s t-test).
Extended Data Fig. 9|. T cells overproducing BH4 display enhanced ATP production, proliferation and autoimmunity.
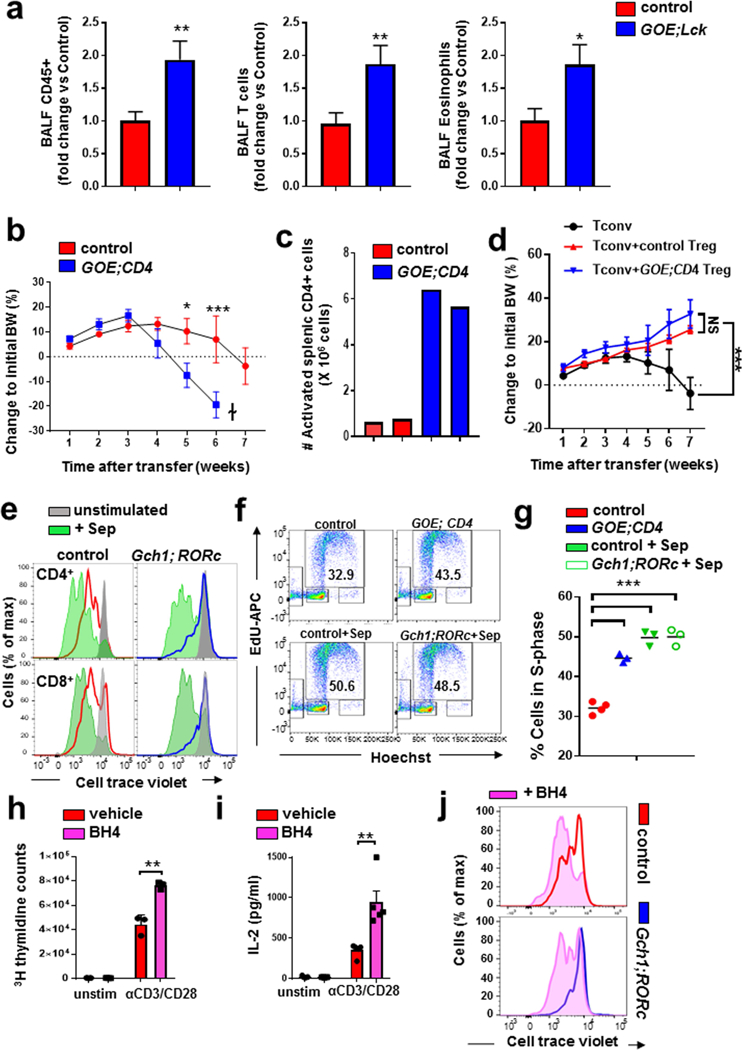
a, Allergic airway inflammatory disease model and fold change of inflammatory cells in BALFs, comparing control and GOE;Lck mice. Data are shown as means ± s.e.m. n = 18 for control mice; n = 17 for GOE;Lck mice. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). b, Transfer colitis model. Changes in body weight of Rag2−/− mice transferred with control (n = 6 animals) or GOE;CD4 (n = 5) naïve CD4+ T cells. Data are shown as means ± s.e.m. *P < 0.05; ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). c, Total numbers of activated (CD62Llo CD44hi) CD4+ splenic T cells at three weeks post-transfer in mice transferred with control or GOE;CD4 naïve CD4+ T cells. Data for two mice from each group are shown. d, Transfer colitis model, involving transfer of naive CD4+ T cells (150,000 cells) and co-transfer of FACS-purified T-reg cells from control (n = 5) and GOE;CD4 (n = 6) mice. Changes to initial body weights were scored over seven weeks. Data are shown as means ± s.e.m. ***P < 0.001; NS, not significant (two-way ANOVA with Tukey’s multiple comparison test). e, Representative histograms depicting the proliferation of purified unstimulated and anti-CD3/28-stimulated CD4+ and CD8+ wild-type and Gch1;RORc T cells treated for three days with sepiapterin (5 μM). The profile for the unstimulated T cells of each genotype is shown in grey. Experiments were repeated three independent times with comparable results. f, g, Representative FACS plots showing EdU-based cell-cycle analysis following 28-h anti-CD3/28 stimulation of control CD4+ T cells, GOE;CD4 CD4+ T cells, control CD4+ T cells treated with sepiapterin (5 μM), and GCH1;RORc CD4+ T cells treated with sepiapterin (5 μM) (f); and quantification of the S-phase-entry population (g). EdU was pulsed for the last 4 h of stimulation. n = 4 mice for control; n = 3 mice for all other genotypes. ***P < 0.001 (one-way ANOVA with Dunnett’s multiple comparisons test). h, i, Effect of BH4 on the proliferati-n (3H-thymidine incorporation; h) and IL-2 secretion (i) of CD4+ wild-type T cells activated with anti-CD3/28 antibodies for 24 h and treated with vehicle (n = 3/4) or BH4 (10 μM; n = 3/4). Data are shown for individual mice as means ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). j, Representative histograms depicting the proliferation of control and Gch1;RORc CD4+ T cells after three days of anti-CD3/28 stimulation supplemented with BH4 (10 μM). FACS blots are representative of two independent experiments with comparable results.
Extended Data Fig. 10|. Overactivation of the GCH1/BH4 pathway leads to enhanced anti-tumour immunity.

a, Total iron content from 24-h anti-CD3/28 stimulated CD4+ T cells (untreated or treated with 5μM sepiapterin) from control (n = 5/4) and Gch1;RORc (n = 5) mice. Data are shown as means ± s.e.m. individual mice for each genotype are shown. *P < 0.05 (one-way ANOVA with Tukey’s multiple comparisons. b, Representative FACS histogram depicting DHE levels (left) and quantification of the mean fluorescent intensity (right) in unstimulated and 20-h anti-CD3/28-activated CD4+ T cells from control and GOE;CD4 littermates as well as wild-type cells treated with sepiapterin (5 μM). n = 3 for each condition. Data are shown as means ± s.e.m. **P < 0.01; ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). c, ATP measurements for stimulated wild-type CD4+ T cells treated with DMSO, sepiapterin (5 μM) or SPRi3 (50 μM) for 24 h. Data are shown as means ± s.e.m. n = 3 for each genotype. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test with multiple comparisons). d, Quantification of intratumoral effector CD4+ T cells (CD44+ CD62Llo) assayed from E0071 tumours on day 28 for vehicle- and BH4-treated mice. Data are shown as means ± s.e.m. n = 5 mice for each condition. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). e, Effect of BH4 supplementation on H-Ras- transformed TC-1 tumour growth. TC-1 tumour cells were orthotopically injected; once the tumours were palpable (day 7), BH4 (35 mg kg–1; n = 15) or vehicle (saline; n = 10) was therapeutically administered for seven days as indicated. Data are shown for individual mice as means ± s.e.m. ***P < 0.001; ****P < 0.0001 (two-way ANOVA with Sidak’s multiple comparisons). f, Quantification of intratumoral effector CD8+ T cells (CD44+ CD62Llo) assayed from TC-1 tumours on day 21 in vehicle- or BH4-treated mice (n = 9 mice for each genotype). Data are shown as means ± s.e.m. **P < 0.01 (two-tailed Student’s t-test). g, Effect of BH4 supplementation on TC-1 tumour growth in Rag2−/− hosts. TC-1 tumour cells were orthotopically injected into Rag2−/− female mice; once the tumours were palpable (day 7), BH4 (35 mg kg–1; n = 5) or vehicle (saline; n = 5) was administered. BH4 and vehicle supplementation was carried out for seven days as indicated on the graph. Data are shown for individual mice as means ± s.e.m. NS, not significant (two-way ANOVA with Sidak’s multiple comparisons). h, Sepiapterin levels in the supernatant of wild-type CD4+ T cells stimulated with anti-CD3/28 antibodies for 20 h and treated with vehicle or kynurenine (KYN; 150 μM). Culture medium was also included for comparison. BQL, below quantifiable levels. Data are shown as means ± s.e.m. n = 4 independent samples for each condition. ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). i, Representative histogram depicting proliferation of three-day anti-CD3/28-activated wild-type CD4+ T cells treated with vehicle or kynurenine (50 μM). j, Representative FACS histograms depicting DHE levels in anti-CD3/28-stimulated wild-type CD4+ T cells treated with vehicle (DMSO), kynurenine alone (50 μM) or kynurenine (50 μM) plus BH4 (10 μM) for 20 h. The experiment was repeated three independent times with comparable results.
Supplementary Material
Acknowledgements
We thank all members of our laboratories for helpful discussions and Life Science Editors for editorial support. We thank Shanghai ChemPartners for running the drug metabolism and pharmacokinetic assays associated with QM385. J.M.P. is supported by grants from IMBA, the Austrian Ministry of Sciences and the Austrian Academy of Sciences, and the T. Von Zastrow Foundation as well as a European Research Council (ERC) Advanced Grant and an Era of Hope Innovator award. C.J.W. is supported by a National Institutes of Health (NIH) R35 grant (NS105076). We also acknowledge the Christian Doppler Laboratory for Iron Metabolism and Anemia Research as a funding body for our research (M.S. and G.W.). M.R. is supported by EMD Serono, Canada, and a MS Network Transitional Career Development Award.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Reviewer information Nature thanks R.S. Johnson, L. O’Neill and N. Restifo for their contribution to the peer review of this work.
Competing interests The authors declare no competing interests.
Extended data is available for this paper at https://doi.org/10.1038/s41586-018-0701-2.
Supplementary information is available for this paper at https://doi.org/10.1038/s41586-018-0701-2.
Reprints and permissions information is available at http://www.nature.com/reprints.
Reporting summary. Further information on experimental design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The microarray data set is accessible through GEO accession number GSE108101. All other data sets generated and/or analysed during this study are available from the corresponding authors upon reasonable request.
Online content
Any methods, additional references, Nature Research reporting summaries, source data, statements of data availability and associated accession codes are available at https://doi.org/10.1038/s41586-018-0701-2.
References
- 1.Hennet T, Hagen FK, Tabak LA & Marth JD T-cell-specific deletion of a polypeptide N-acetylga lactosaminyl-tra nsferase gene by site-di rected recombination. Proc. Natl Acad. Sci. USA 92, 12070–12074 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Sawada S, Scarborough JD, Killeen N & Liftman DR A lineage-specific transcriptional silencer regulates CD4 gene expression during T lymphocyte development. Cell 77, 917–929 (1994). [DOI] [PubMed] [Google Scholar]
- 3.Ventura A et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007). [DOI] [PubMed] [Google Scholar]
- 4.Crabtree MJ et al. Quantitative regulation of intracellular endothelial nitric-oxide synthase (eNOS) coupling by both tetrahydrobiopterin-eNOS stoichiometry and biopterin redox status: insights from cells with tet-regulated GTP cyclohydrolase I expression. J. Biol. Chem. 284, 1136–1144 (2009). [DOI] [PubMed] [Google Scholar]
- 5.Banerjee A et al. Cellular and site-specific mitochondrial characterization of vital human amniotic membrane. Cell Transplant. 27, 3–11 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schmitt TM & Zúñiga-Pflücker JC Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 17, 749–756 (2002). [DOI] [PubMed] [Google Scholar]
- 7.Becker C et al. In vivo imaging of colitis and colon cancer development in mice using high resolution chromoendoscopy. Gut 54, 950–954 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Collison LW & Vignali DAA In vitro Treg suppression assays. Methods Mol. Biol. 707, 21–37 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Boivin N, Baillargeon J, Doss PMIA, Roy AP & Rangachari M Interferon-β suppresses murine Th 1 cell function in the absence of antigen-presenting cells. PLoS One 10, e0124802 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lin KY et al. Treatment of established tumors with a novel vaccine that enhances major histocompatibility class II presentation of tumor antigen. Cancer Res. 56, 21–26 (1996). [PubMed] [Google Scholar]
- 11.Smyth GK Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 10.2202/1544-6115.1027 (2004). [DOI] [PubMed] [Google Scholar]
- 12.Edgar R, Domrachev M & Lash AE Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Arai N, Narisawa K, Hayakawa H & Tada K Hyperphenylalaninemia due to dihydropteridine reductase deficiency: diagnosis by enzyme assays on dried blood spots. Pediatrics 70, 426–430 (1982). [PubMed] [Google Scholar]
- 14.Theurl I et al. On-demand erythrocyte disposal and iron recycling requires transient macrophages in the liver. Nat. Med. 22, 945–951 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Latremoliere A et al. Reduction of neuropathic and inflammatory pain through inhibition of the tetrahydrobiopterin pathway. Neuron 86, 1393–1406 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Werner ER, Blau N & Thöny B Tetrahydrobiopterin: biochemistry and pathophysiology. Biochem. J. 438, 397–414 (2011). [DOI] [PubMed] [Google Scholar]
- 17.Chen W et al. Role of increased guanosine triphosphate cyclohydrolase-1 expression and tetrahydrobiopterin levels upon T cell activation. J. Biol. Chem. 286, 13846–13851 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ziegler I et al. Control of tetrahydrobiopterin synthesis in T lymphocytes by synergistic action of interferon-Y and interleukin-2. J. Biol. Chem. 265, 17026–17030 (1990). [PubMed] [Google Scholar]
- 19.Chuaiphichai S et al. Cell-autonomous role of endothelial GTP cyclohydrolase 1 and tetrahydrobiopterin in blood pressure regulation. Hypertension 64, 530–540 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Eberl G & Littman DR Thymic origin of intestinal αβ T cells revealed by fate mapping of RORγt+ cells. Science 305, 248–251 (2004). [DOI] [PubMed] [Google Scholar]
- 21.Hobeika E et al. Testing gene function early in the B cell lineage in mb1-cre mice. Proc. Natl Acad. Sci. USA 103, 13789–13794 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ṥledzińska A et al. TGF-β signalling is required for CD4+ T cell homeostasis but dispensable for regulatory T cell function. PLoS Biol. 11, e1001674 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Talbot S et al. Silencing nociceptor neurons reduces allergic airway inflammation. Neuron 87, 341–354 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Haworth O, Cernadas M, Yang R, Serhan CN & Levy BD Resolvin E1 regulates interleukin 23, interferon-γ and lipoxin A4 to promote the resolution of allergic airway inflammation. Nat. Immunol. 9, 873–879 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Martin SF et al. Toll-like receptor and IL-12 signaling control susceptibility to contact hypersensitivity. J. Exp. Med. 205, 2151–2162 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rangachari M & Kuchroo VK Using EAE to better understand principles of immune function and autoimmune pathology. J. Autoimmun. 45, 31–39 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Rangachari M et al. Bat3 promotes T cell responses and autoimmunity by repressing Tim-3-mediated cell death and exhaustion. Nat. Med. 18, 1394–1400 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nar H et al. Active site topology and reaction mechanism of GTP cyclohydrolase I. Proc. Natl Acad. Sci. USA 92, 12120–12125 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nar H et al. Atomic structure of GTP cyclohydrolase I. Structure 3, 459–466 (1995). [DOI] [PubMed] [Google Scholar]
- 30.Volani C et al. Dietary iron loading negatively affects liver mitochondrial function. Metallomics 9, 1634–1644 (2017). [DOI] [PubMed] [Google Scholar]
- 31.Archer MC, Vonderschmitt DJ & Scrimgeour KG Mechanism of oxidation of tetrahydropterins. Can. J. Biochem. 50, 1174–1182 (1972). [DOI] [PubMed] [Google Scholar]
- 32.Eberlein G, Bruice TC, Lazarus RA, Henrie R & Benkovic SJ The interconversion of the 5,6,7,8-tetrahydro-, 7,8-dihydro-, and radical forms of 6,6,7,7-tetramethyldihydropterin. A model for the biopterin center of aromatic amino acid Mixed function oxidases. J. Am. Chem. Soc. 106, 7916–7924 (1984). [Google Scholar]
- 33.Capeillere-Blandin C, Mathieu D & Mansuy D Reduction of ferric haemoproteins by tetrahydropterins: a kinetic study. Biochem. J. 392, 583–587 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hondowicz BD et al. Interleukin-2-dependent allergen-specific tissue-resident memory cells drive asthma. Immunity 44, 155–166 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ewens A, Mihich E & Ehrke MJ Distant metastasis from subcutaneously grown E0771 medullary breast adenocarcinoma. Anticancer Res. 25, 3905–3915 (2005). [PubMed] [Google Scholar]
- 36.Curti A et al. Indoleamine 2,3-dioxygenase-expressing leukemic dendritic cells impair a leukemia-specific immune response by inducing potent T regulatory cells. Haematologica 95, 2022–2030 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Haruki H, Hovius R, Pedersen MG & Johnsson K Tetrahydrobiopterin biosynthesis as a potential target of the kynurenine pathway metabolite xanthurenic acid. J. Biol. Chem. 291, 652–657 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Oppenheimer SJ Iron and its relation to immunity and infectious disease. J. Nutr. 131, 616S–633S (2001). [DOI] [PubMed] [Google Scholar]
- 39.Cassat JE & Skaar EP Iron in infection and immunity. Cell Host Microbe 13, 509–519 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Liu C-J, Chen K-W Hu Y-W. Hong Y-C., Huang Y-C., Chiou T-J & Tzeng C-H. Chronic iron deficiency anemia and cancer risk. Blood 120, 5172 (2012). [Google Scholar]
- 41.Hung N et al. Risk of cancer in patients with iron deficiency anemia: a nationwide population-based study. PLoS One 10, e0119647 (2015); correction 10.1371/journal.pone.0125951 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


