Abstract
BACKGROUND:
Diagnosing giant cell-rich bone tumors can be challenging on limited biopsies. H3 histone family member 3A (H3F3A) (G34W/V/R/L) mutations are present in the majority of giant cell tumors (GCTs) of bone and H3 histone family member 3B (H3F3B) (K36M) mutations are present in nearly all chondroblastomas, but are absent in histologic mimics. Mutation-specific immunohistochemistry (IHC) is highly specific for GCT and chondroblastoma in surgical excisions. The objective of the current study was to validate H3G34W and H3K36M IHC in the diagnosis of giant cell-rich bone tumors on fine-needle aspiration and core needle biopsy specimens.
METHODS:
IHC was performed using monoclonal antibodies against histone H3.3 G34W and K36M in GCTs of bone (26 cases, including 2 malignant cases), GCT of Paget disease (1 case), chondroblastoma (8 cases), aneurysmal bone cyst (7 cases), and osteosarcoma (13 cases) with available fineneedle aspiration and/or core needle biopsy specimens from 2 institutions. H3F3A and H3F3B Sanger sequencing was performed on all 4 H3G34W IHC-negative GCTs.
RESULTS:
IHC for H3G34W was positive in 22 of 26 GCTs (85%) and negative in all histologic mimics. IHC for H3K36M was positive in all 8 chondroblastomas and negative in all histologic mimics. IHC results were concordant between biopsy and surgical specimens in 152 of 158 samples (96%). Sequencing identified alternate H3F3A G34L and G34V mutations in 1 IHC-negative GCT each, but no mutation was found in the remaining 2 cases.
CONCLUSIONS:
H3G34W and H3K36M IHC is highly specific for GCT and chondroblastoma, respectively, among giant cell-rich bone tumors, and is useful for confirming the diagnosis in limited biopsies. The presence of alternate H3F3A mutations accounts for the H3G34W IHC negativity in a subset of GCT of bone cases.
Keywords: Chondroblastoma, core needle biopsy, fine-needle aspiration, giant cell-rich, giant cell tumor of bone, H3 histone family member 3A (H3F3A), H3 histone family member 3B (H3F3B), H3G34W, H3K36M, immunohistochemistry
INTRODUCTION
Giant cell-rich primary bone tumors are defined by the presence of prominent osteoclasts or osteoclast-like giant cells and include giant cell tumor (GCT) of bone, GCT of Paget disease, chondroblastoma, aneurysmal bone cyst, and a subset of osteosarcomas.1 Correlation with clinical and radiologic characteristics, such as tumor site and patient age (indicative of an immature vs mature skeleton), is critical to establish a correct diagnosis. A clear distinction of benign or locally aggressive tumors (ie, conventional GCT, chondroblastoma, or aneurysmal bone cyst) from malignant GCT or osteosarcoma is of paramount importance, with a substantial impact on prognosis and clinical management.2,3
GCT of bone has a predilection for young adults with a mature skeleton and usually arises in the epiphysis of long bones.4 Fewer than 10% of GCTs of bone demonstrate malignant transformation and/or distant metastatic spread. In contrast, chondroblastoma occurs in children and adolescents with an immature skeleton and involves the epiphysis of long bones, with or without extension into the articular cartilage.4 Aneurysmal bone cysts usually affect patients in their first 2 decades of life with a slight female predominance, and is most common in the metaphysis of long bones.4 Osteosarcoma usually occurs in the metadiaphysis of the long bones of adolescents and young adults, with a male predominance.4
Although the diagnosis of giant cell-rich bone tumors on surgical curetting or excision specimens usually is straightforward, the evaluation of these tumors on small biopsies, such as fine-needle aspiration (FNA) or core needle biopsy (CNB), can be challenging,5–10 especially in tumors that arise in unusual anatomic sites and/or show atypical morphologic features. With the increasing use of minimally invasive procedures to obtain samples for the initial diagnosis of bone tumors in recent years, recognition of a spectrum of clinical, radiologic, and morphologic features, with the judicious application of immunohistochemical and/or genetic studies, becomes essential in the evaluation of many primary bone tumors.
Recently, highly recurrent somatic driver mutations in H3 histone family member 3A (H3F3A) (located at 1q42.12) and H3 histone family member 3B (H3F3B) (located at 17q25.1) encoding replication-independent histone proteins (H3.3) of identical amino acid sequence have been identified in 92% of GCTs of bone and 95% of chondroblastomas, respectively.11 In GCT of bone, H3F3A G34W mutations are most common (85%−95%), but alternate G34V, G34R, and G34L mutations have been reported in a subset of cases.11–15 Chondroblastomas harbor K36M mutations in >90% of cases, resulting from H3F3B mutations and rarely from H3F3A mutations.11,15 Other giant cell-rich bone tumors, such as aneurysmal bone cyst and giant cell-rich osteosarcoma, lack these mutations with very rare exceptions.11,13,14,16
Immunohistochemistry (IHC) using monoclonal antibodies directed against the mutant H3.3G34W and H3.3K36M proteins has been shown to be highly specific for the diagnosis of GCT of bone17 and chondroblastoma,18 respectively, on surgical excision specimens. In GCT of bone and chondroblastoma, H3F3A and H3F3B mutations as well as resulting mutant H3.3G34W and H3.3K36M proteins are restricted to the mononuclear stromal cell population and are absent in nonneoplastic multinucleated giant cells and their precursors.11
Although H3F3A sequencing previously has been shown to detect mutations in approximately 90% of GCTs of bone in CNB samples,14 to the best of our knowledge, IHC for H3G34W and H3K36M has not been systematically studied in FNA and/or CNB specimens. The objective of the current study was to validate the H3G34W and H3K36M mutation-specific antibodies in the diagnosis of giant cell-rich bone tumors on FNA and CNB specimens, with correlation of H3F3A and H3F3B Sanger sequencing in a subset of cases.
MATERIALS AND METHODS
Tumor Samples
Cases were retrieved from the cytopathology and surgical pathology files of Brigham and Women’s Hospital and Massachusetts General Hospital. A total of 55 giant cell-rich tumors including 26 GCTs of bone (including 2 malignant cases), 1 GCT of Paget disease, 8 chondroblastomas, 7 aneurysmal bone cysts, and 13 osteosarcomas were selected. The diagnosis of each case was confirmed by subsequent curetting and/or surgical resection specimens. Clinical information was retrieved from the electronic medical record and the pathology reports. A total of 158 specimens from 55 tumors in 55 patients were included, which comprised 42 FNA samples, 53 CNB specimens, and 63 curettings or surgical excisions. All needle biopsies in both institutions were performed by interventional radiologists via computed tomography guidance using a percutaneous approach. At Brigham and Women’s Hospital, concurrent FNA and CNB samples were obtained with a 22-gauge, 15-cm Chiba needle (Cook Medical LLC, Bloomington, Indiana) for FNA and a 16-gauge, 15-cm Temno biopsy needle (Merit Medical Systems, South Jordan, Utah) for CNB, both through a 13.5-gauge, 10-cm introducer needle. Only CNB specimens were obtained at Massachusetts General Hospital. Both alcohol-fixed Papanicolaou-stained and air-dried modified Romanowsky-stained smears were prepared from FNA. Formalin-fixed, paraffin-embedded (FFPE) cell blocks were prepared from needle rinses and/or dedicated FNA passes using human plasma and thrombin for clot formation. The hematoxylin and eosin-stained cell block and histologic sections were rereviewed by at least 2 pathologists who subspecialized in soft tissue and bone pathology (I.M.S., G.P.N., J.L.H., and X.Q.) to confirm the diagnosis in selected cases. The majority of surgical specimens were treated with decalcification before processing using either Rapid Cal Immuno solution (BBC Biochemical, Mount Vernon, Washington) for bone CNB samples or Decal Stat solution (Thermo Fisher Scientific, Waltham, Massachusetts) for excisions. No ancillary IHC stains were performed in the initial diagnostic workup.
Immunohistochemistry
IHC was performed on FFPE tissue or cell block sections measuring 4-lm thick after pressure cooker antigen retrieval (Target Retrieval Solution [pH 6.1]; Dako, Carpinteria, California) using rabbit monoclonal antibodies directed against histone H3.3 G34W (1:2000 dilution, clone RM263; RevMAb, South San Francisco, California) and histone H3.3 K36M (1:4000 dilution, clone RM193; RevMAb) with the EnVision Plus detection system (Dako). Appropriate positive (confirmed GCT of bone and chondroblastoma) and negative (normal skeletal muscle, colon, and skin) controls were used throughout. Staining was scored as positive if >5 % of the tumor cells were positive.
DNA Isolation and Polymerase Chain Reaction
Sanger sequencing of H3F3A and H3F3B (including codons 34 and 36) was performed on all 4 H3G34W-negative GCTs of bone (cases 9, 13, 23, and 34), 1 H3G34W-positive malignant GCT of bone (case 33), 1 H3G34W-negative GCT of Paget disease (case 35), and 1 H3K36M-positive chondroblastoma (case 3). DNA was isolated from FFPE tissue blocks using standard protocols (QIAamp DNA FFPE Tissue Kit; Qiagen, Valencia, California). Genomic DNA primers for H3F3A (amplicon sizes of 128 base pair [bp] and 170 bp) and H3F3B (amplicon size of 148 bp) were designed using Primer3 Input software (version 0.4.0) (http://bioinfo.ut.ee/primer3–0.4.0/). Polymerase chain reaction (PCR) products were purified using ExoSAP-IT reagent (ExoSAP-IT PCR Product Cleanup Reagent; Applied Biosystems, Foster City, California). Primer sequences and sequencing conditions, which have been described previously,19 are available upon request. PCR reaction and sequencing were performed as per standard protocols.19
RESULTS
Clinicopathologic, IHC, and genomic data for all 55 patients with giant cell-rich bone tumors are summarized in Table 1.
TABLE 1.
Summary of Clinicopathologic, Immunohistochemical, and Genomic Findings in 55 Giant Cell-Rich Bone Tumors
| Case No. | Diagnosis | Sex | Age, Years | Tumor Site | Biopsy Type | IHC Biopsy (No. of Samples) | IHC Excision (No. of Samples) | Genomic |
|---|---|---|---|---|---|---|---|---|
| 1 | Chondroblastoma | Male | 19 | Distal femur | FNA and CNB | H3K36M (2) | H3K36M (1) | |
| 2 | Chondroblastoma | Female | 15 | Navicular bone | FNA and CNB | H3K36M (2) | Negative (1) | |
| 3 | Chondroblastoma | Male | 28 | Calcaneus | FNA and CNB | H3K36M (1) | H3K36M (1) | H3F3B c.110A>T;p.K36Ma |
| 4 | Chondroblastoma | Female | 17 | Proximal humerus | CNB | H3K36M (1) | Negative (1) | |
| 5 | Chondroblastoma | Female | 13 | Proximal humerus | CNB | H3K36M (1) | Negative (1) | |
| 6 | Chondroblastoma | Female | 29 | Distal femur | CNB | H3K36M (1) | Negative (1) | |
| 7 | Chondroblastoma | Male | 28 | Ilium | CNB | H3K36M (1) | Negative (1) | |
| 8 | Chondroblastoma | Male | 28 | Distal tibia | CNB | Negative (1) | H3K36M (1) | |
| 9 | GCT of bone | Female | 22 | Distal femur | FNA and CNB | Negative (2) | Negative (1) | WTa |
| 10 | GCT of bone | Female | 75 | Distal femur | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 11 | GCT of bone | Female | 24 | Proximal femur | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 12 | GCT of bone | Male | 84 | Distal femur | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 13 | GCT of bone | Female | 20 | Sacrum | FNA and CNB | Negative (2) | Negative (1)b | WTa,c |
| 14 | GCT of bone | Male | 27 | Sacrum | FNA | H3G34W (1) | H3G34W (1) | |
| 15 | GCT of bone | Male | 38 | Distal radius | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 16 | GCT of bone | Male | 48 | Ischium | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 17 | GCT of bone | Male | 27 | Distal humerus | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 18 | GCT of bone | Female | 30 | Metatarsal bone | FNA and CNB | H3G34W (2) | H3G34W (5) | |
| 19 | GCT of bone | Female | 34 | Proximal tibia | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 20 | GCT of bone | Female | 44 | Ilium | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 21 | GCT of bone | Male | 53 | Distal tibia | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 22 | GCT of bone | Male | 41 | Ilium | FNA and CNB | H3G34W (2) | H3G34W (3) | |
| 23 | GCT of bone | Male | 25 | Proximal femur | FNA and CNB | Negative (2) | Negative (1) | H3F3A c.104G>T;p.G34Va |
| 24 | GCT of bone | Female | 83 | Distal femur | FNA and CNB | H3G34W (2) | H3G34W (1) | |
| 25 | GCT of bone | Male | 29 | 10th rib | CNB | H3G34W (1) | H3G34W (1) | |
| 26 | GCT of bone | Male | 52 | Distal ulna | CNB | H3G34W (1) | H3G34W (1) | |
| 27 | GCT of bone | Male | 55 | Distal femur | CNB | H3G34W (1) | H3G34W (1) | |
| 28 | GCT of bone | Male | 22 | Distal radius | CNB | H3G34W (1) | H3G34W (1)b | |
| 29 | GCT of bone | Male | 17 | Distal femur | CNB | H3G34W (1) | H3G34W (1) | |
| 30 | GCT of bone | Male | 54 | Distal femur | CNB | H3G34W (1) | H3G34W (1) | |
| 31 | GCT of bone | Female | 31 | Proximal humerus | CNB | H3G34W (1) | H3G34W (1) | |
| 32 | GCT of bone | Female | 26 | Distal femur | CNB | H3G34W (1) | H3G34W (1) | |
| 33 | Malignant GCT | Female | 18 | T7 vertebra | FNA and CNBd | H3G34W (3)e | H3G34W (1)f | H3F3A c.103G>T;p.G34Wa |
| 34 | Malignant GCT | Female | 62 | Proximal tibia | FNA and CNB | Negative (4) | Negative (1)b | H3F3A c.103_104GG>TT;p.G34La |
| 35 | GCT of Paget disease | Male | 88 | Distal humerus | FNA and CNB | Negative (2) | Negative (2) | WTa |
| 36 | ABC | Female | 23 | Ulnar shaft | CNB | Negative (1) | Negative (1) | |
| 37 | ABC, solid variant | Female | 21 | Rib | FNA and CNB | Negative (2) | Negative (1) | |
| 38 | ABC, solid variant | Female | 16 | Proximal femur | FNA | Negative (1) | Negative (1) | |
| 39 | ABC | Female | 33 | Sacrum | FNA and CNB | Negative (2) | Negative (1) | |
| 40 | ABC | Female | 17 | Proximal tibia | FNA and CNB | Negative (2) | Negative (1) | |
| 41 | ABC, solid variant | Female | 33 | Ulnar shaft | FNA and CNB | Negative (2) | Negative (1) | |
| 42 | ABC | Female | 10 | Distal femur | CNB | Negative (1) | Negative (1) | |
| 43 | Osteosarcoma | Female | 40 | Distal femur | FNA and CNB | Negative (2) | Negative (1) | |
| 44 | Osteosarcoma | Female | 33 | Proximal femur | FNA and CNB | Negative (2) | Negative (1) | |
| 45 | Osteosarcoma | Female | 41 | Distal tibia | FNA and CNB | Negative (2) | Negative (1) | |
| 46 | Osteosarcoma | Female | 25 | Ilium | FNA and CNB | Negative (2) | Negative (1) | |
| 47 | Osteosarcoma | Male | 72 | Acetabulum | FNA and CNB | Negative (3) | Negative (1) | |
| 48 | Osteosarcoma | Female | 19 | Distal femur | FNA and CNB | Negative (2) | Negative (1) | |
| 49 | Osteosarcoma | Male | 18 | Proximal fibula | FNA | Negative (2) | Negative (1) | |
| 50 | Osteosarcoma | Female | 55 | Femur shaft | FNA and CNB | Negative (2) | Negative (1) | |
| 51 | Osteosarcoma | Female | 61 | Proximal femur | FNA | No tumor (1) | Negative (2) | |
| 52 | Osteosarcoma | Male | 24 | Proximal femur | FNA and CNB | Negative (2) | Negative (1) | |
| 53 | Osteosarcoma | Female | 40 | Ilium | FNA and CNB | Negative (2) | Negative (1) | |
| 54 | Osteosarcoma | Male | 25 | Distal femur | FNA and CNB | Negative (2) | Negative (1) | |
| 55 | Osteosarcoma | Male | 48 | Proximal tibia | FNA and CNB | Negative (2) | Negative (1) |
Abbreviations: ABC, aneurysmal bone cyst; CNB, core needle biopsy; FNA, fine-needle aspiration; GCT, giant cell tumor; H3F3A, H3 histone family member 3A; H3F3B, H3 histone family member 3B; IHC, immunohistochemistry; WT: wild-type.
Sequencing was performed on curetting and/or excision.
Preceded by denosumab and/or chemotherapy.
Sequencing was performed on fine-needle aspiration.
Preceded by radiotherapy.
Negative staining in 1 biopsy specimen (decalcified).
Preceded by chemotherapy and/or radiotherapy.
Clinicopathologic Findings
Among the 26 patients with GCTs of bone, 14 were male and 12 female with a median age of 32.5 years (range, 17–84 years). Conventional GCT of bone was found for the most part in the lower extremity (13 cases), followed by the limb girdle (5 cases), upper extremity (5 cases), and trunk (1 case). The 2 malignant GCTs of bone arose in the lower extremity and trunk, respectively. The single case of GCT of Paget disease arose in the humerus of an 88-year-old man with Paget disease who presented clinically with pathologic fracture. Of the 8 patients with chondroblastoma, 4 were male and 4 were female with a median age of 23.5 years (range, 13–28 years). Chondroblastomas were located in the lower extremity (5 cases), upper extremity (2 cases), and limb girdle (1 case). All 7 patients with aneurysmal bone cyst were female with a median age of 21 years (range, 10–33 years). Aneurysmal bone cysts were located in the lower extremity (3 cases), upper extremity (2 cases), limb girdle (1 case), and trunk (1 case). Thirteen patients with osteosarcoma comprised 5 males and 8 females with a median age of 40 years (range, 18–72 years); the tumors for the most part were located in the lower extremity (10 cases) or limb girdle (3 cases).
The original diagnoses of patients with GCT of bone or chondroblastoma on FNA, CNB, and curettage/excision specimens are summarized in Table 2.
TABLE 2.
FNA, CNB, and Excision Diagnosis of Chondroblastoma and GCT of Bone
| Diagnosis | Case No. | FNA | CNB | Curettage/Excision | Change in Diagnosis |
|---|---|---|---|---|---|
| Chondroblastoma | 1 | Chondroblastoma | Chondroblastoma | Chondroblastoma | No |
| 2 | Chondroblastoma | Chondroblastoma | Chondroblastoma | No | |
| 3 | ABCa | No tumor present | Chondroblastoma | IHC/sequencing (FNA)a | |
| 4 | Not performed | Chondroblastoma | Chondroblastoma | No | |
| 5 | Not performed | Chondroblastoma | Chondroblastoma | No | |
| 6 | Not performed | Chondroblastoma | Chondroblastoma | No | |
| 7 | Not performed | Chondroblastoma | Chondroblastoma | No | |
| 8 | Not performed | Chondroblastoma | Chondroblastoma | No | |
| GCT of bone | 9 | Giant cell-rich spindle cell neoplasm | Giant cell-rich spindle cell neoplasm | GCT of bone | No |
| 10 | Rare atypical spindle/epithelioid cells, multinucleated giant cells | GCT of bone | GCT of bone | No | |
| 11 | GCT of bone | GCT of bone | GCT of bone | No | |
| 12 | GCT of bone | GCT of bone | GCT of bone | No | |
| 13 | GCT of bone | GCT of bone | GCT of bone | No | |
| 14 | GCT of bone | Not performed | GCT of bone | No | |
| 15 | GCT of bone | GCT of bone | GCT of bone | No | |
| 16 | GCT of bone | Benign histiocytic proliferationb | GCT of bone | No | |
| 17 | GCT of bone | GCT of bone | GCT of bone | No | |
| 18 | GCT of bone | GCT of bone | GCT of bone | No | |
| 19 | GCT of bone | GCT of bone | GCT of bone | No | |
| 20 | GCT of bone | Nondiagnosticb | GCT of bone | No | |
| 21 | GCT of bone | GCT of bone | GCT of bone | No | |
| 22 | GCT of bone | GCT of bone | GCT of bone | No | |
| 23 | Giant cell-rich neoplasm | GCT of bone | GCT of bone | No | |
| 24 | Giant cell-rich neoplasm | Giant cell-rich neoplasm | Giant cell-rich neoplasm: favor GCT of bone | IHC (biopsy and excision)c | |
| 25 | Not performed | GCT of bone | GCT of bone | No | |
| 26 | Not performed | GCT of bone | GCT of bone | No | |
| 27 | Not performed | GCT of bone | GCT of bone | No | |
| 28 | Not performed | GCT of bone | GCT of bone | No | |
| 29 | Not performed | GCT of bone | GCT of bone | No | |
| 30 | Not performed | GCT of bone | GCT of bone | No | |
| 31 | Not performed | GCT of bone | GCT of bone | No | |
| 32 | Not performed | GCT of bone | GCT of bone | No | |
| 33 | GCT of bone | GCT of bone | Chondroblastoma/ osteosarcoma | IHC/sequencing (biopsy and excision)d | |
| 34 | GCT of bone/ osteosarcoma | GCT of bone/ osteosarcoma | Osteosarcoma | Sequencing (excision)d |
Abbreviations: ABC, aneurysmal bone cyst; CNB, core needle biopsy; FNA, fine-needle aspiration; GCT, giant cell tumor; IHC, immunohistochemistry.
Reclassified as chondroblastoma.
IHC highlighted tumor cells.
Reclassified as GCT of bone.
Reclassified as malignant GCT of bone.
In 26 patients, the diagnosis of GCT of bone was established on FNA alone (3 cases, including 2 cases with nondiagnostic concurrent CNB specimens), CNB alone (10 cases, including 2 cases with inconclusive concurrent FNA specimens), FNA with concurrent CNB specimens (11 cases), and on curetting/excision (2 cases). In the last 2 cases, indefinite diagnoses such as “giant cell-rich neoplasm” was rendered on both FNA and CNB samples. Before excision, 3 patients with GCT of bone (cases 13, 28, and 34) received nonsurgical treatment with denosumab, a monoclonal antibody to receptor activator of nuclear factor jB (RANK) ligand.20 Histologically, these treated tumors demonstrated a marked decrease in giant cells accompanied by hyalinization and osteoid formation. One patient with malignant GCT of bone underwent radiotherapy (case 33) and the other patient (case 34) was treated with chemotherapy after their malignant diagnoses; no significant treatment effect was observed histologically.
FNA specimens from cases of GCT of bone were cellular in 17 of 19 samples. Smear preparations demonstrated oval to spindled cells in cohesive clusters bordered by multiple multinucleated giant cells. Cell blocks showed bland mononuclear spindle cells with admixed, evenly distributed, multinucleated giant cells and focal hemosiderin deposition (Figs. 1A and 1B). The GCT of Paget disease was cytomorphologically indistinguishable from GCT of bone (data not shown).
Figure 1.
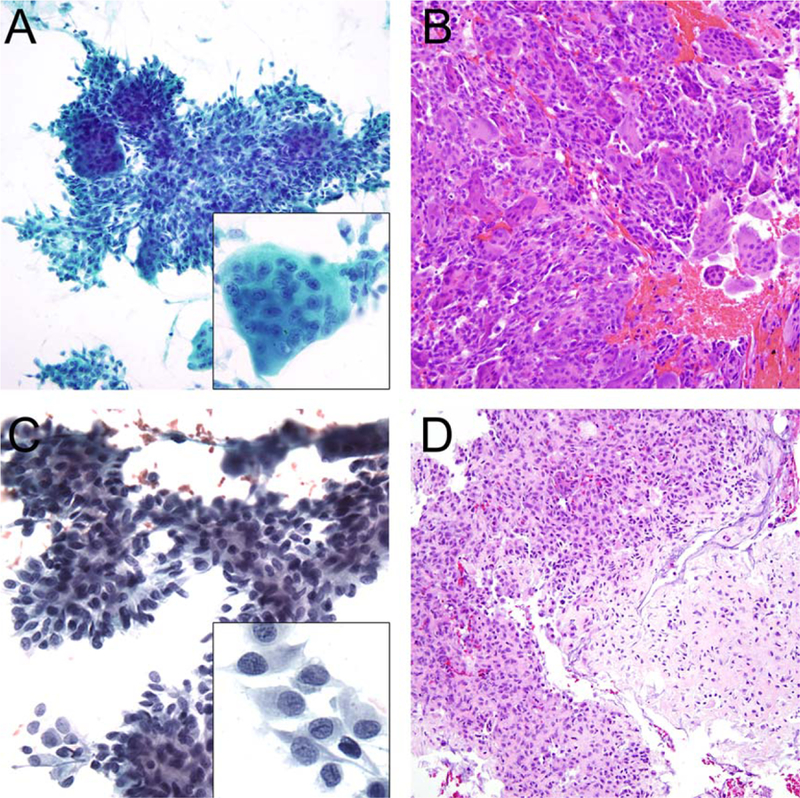
Giant cell tumor of bone shown on (A) a fine-needle aspiration (FNA) specimen (Papanicolaou stain) demonstrating oval to spindled cells in cohesive clusters bordered by multiple multinucleated giant cells (Inset). (B) The cell block (H & E) showed bland mononuclear spindle cells with admixed, evenly distributed multinucleated giant cells. Chondroblastoma is shown on (C) an FNA specimen (Papanicolaou stain) demonstrating cellular clusters of uniformly round to oval cells with loose matrix material and scattered multinucleated giant cells. At high power, the tumor cells showed well-defined cell borders, clear to vacuolated cytoplasm, and oval nuclei with longitudinal grooves (Inset). (D) The cell block (H & E) showed multinucleated giant cells and mononuclear cells embedded in a chondroid matrix.
Of the 8 cases of chondroblastoma, 2 were diagnosed on FNA with concurrent CNB specimens, 5 cases were diagnosed on CNB alone, and 1 case was diagnosed on surgical curetting (Table 2). Two FNA cases of chondroblastoma demonstrated characteristic cellular smears with clusters of uniformly round to oval cells with loose matrix material and scattered multinucleated giant cells. The tumor cells showed well-delineated cell borders, clear to vacuolated cytoplasm, and oval nuclei with longitudinal grooves (Figs. 1C and 1D). Case 3 was a chondroblastoma arising in the calcaneus of a 28-year-old man. The initial FNA was diagnosed as aneurysmal bone cyst and the concurrent CNB specimen was nondiagnostic. This tumor was reclassified as chondroblastoma after re-review of the curetting. The FNA specimens of this case were very sparsely cellular with occasional oval epithelioid cells and osteoclast-like giant cells in a bloody background, and the cell block sections demonstrated scant mononuclear cells with admixed multinucleated giant cells and dense eosinophilic matrix.
In 7 patients, aneurysmal bone cyst was diagnosed on FNA with a concurrent CNB specimen in 4 cases, on CNB alone in 2 cases, and on FNA alone in 1 case. FNA specimens demonstrated small fragments of spindle cells with admixed multinucleated giant cells in a prominent hemorrhagic background (Figs. 2A and 2B). Eleven cases of osteosarcoma were diagnosed on FNA with concurrent CNB specimens whereas 2 cases were diagnosed on FNA alone. The FNA specimens exhibited loosely cohesive pleomorphic tumor cells with admixed multinucleated giant cells (Figs. 2C and 2D).
Figure 2.

Aneurysmal bone cyst shown on (A) a Papanicolaou-stained fine-needle aspiration (FNA) smear demonstrating small fragments of spindle cells with admixed multinucleated giant cells (Inset) in a hemorrhagic background. (B) The cell block (H & E) revealed multinucleated giant cells located at the periphery of fragments of spindle cell-rich stroma. Osteosarcoma is shown on (C) a Papanicolaou-stained FNA specimen demonstrating loosely cohesive pleomorphic tumor cells with admixed multinucleated giant cells. However, at high power, numerous atypical osteoblasts with atypical mitoses were evident (Inset). (D) The cell block (H & E) showed highly atypical tumor cells with pleomorphism and mitoses surrounded by lacey osteoid.
Immunohistochemistry
For GCTs of bone, IHC for H3G34W was positive in 22 of 26 cases (85%) with at least 2 positive specimens. Of a total of 78 specimens (Table 1), IHC for H3G34W was positive in 64 samples (82%), including 14 of 19 FNA samples (74%), 22 of 27 CNB samples (81%), and 28 of 32 excisions (88%). These specimens included 2 GCTs of bone (Table 1, case 23 and case 34) with alternate G34V and G34L mutations (see Sanger sequencing below), which are not detected by the H3G34W antibody. In 2 CNB cases (Table 2, case 16 and case 20), “benign histiocytic proliferation” and “nondiagnostic” were reported, but IHC for H3G34W highlighted the few neoplastic cells in these CNB samples. IHC for H3G34W demonstrated positive staining in all 3 samples (FNA, CNB, and excision) of a “giant cell-rich tumor” in the distal femur of an 83-year-old woman with unusual histology showing extensive hyalinization (case 24), and therefore the diagnosis of GCT of bone finally was confirmed. In all positive cases, only the mononuclear stromal cells were positive for H3G34W and the nonneoplastic multinucleated giant cells and histiocytes were negative (Fig. 3A).
Figure 3.

Immunohistochemistry for H3G34W in (A) a giant cell tumor of bone was positive in a subset of mononuclear tumor cells (approximately 40% of cells) and negative in multinucleated giant cells as well as the remaining mononuclear cells and inflammatory cells. H3G34W was negative in (B) chondroblastoma, (C) aneurysmal bone cyst, and (D) osteosarcoma.
In 1 malignant GCT of bone (case 33) that originally was classified as GCT of bone on FNA and CNB, a diagnosis of osteosarcoma (variably chondroblastoma-like or giant cell-rich) was made on the surgical resection specimen based on the presence of areas with significant cytologic atypia and the consultation opinions from multiple bone pathology experts worldwide. The tumor cells expressed H3G34W on all the biopsy and resection specimens (Fig. 4A-D), thereby supporting the diagnosis of malignant GCT of bone. The second malignant GCT of bone (case 34) initially was diagnosed as GCT of bone on both FNA and CNB specimens, but recurred 32 months later as “high-grade spindle cell sarcoma, suggestive of osteosarcoma” on the repeat CNB and subsequent surgical resection specimen (Fig. 5A and 5C). This case was negative for H3G34W in both biopsy and resection specimens but was shown to harbor an alternate H3F3A mutation (see below and Fig. 5B and 5D). Neoadjuvant and/or denosumab therapy appeared to have no effect on H3G34W expression in any of these cases.
Figure 4.

Malignant giant cell tumor (GCT) of bone (case 33). (A) The core needle biopsy (H & E) showed nonneoplastic multinucleated giant cells (Inset) in a background of mononuclear cells, consistent with typical GCT of bone at the time of initial biopsy. (B) Immunohistochemistry (IHC) for H3G34W was positive in tumor cells (approximately 60%) and negative in admixed nonneoplastic cells. (C) A subsequent surgical resection performed 4 months later revealed pleomorphic tumor cells with scattered multinucleated giant cells, frequent mitoses, and bone formation, mimicking osteosarcoma. (D) IHC for H3G34W highlighted the neoplastic nuclei, including atypical forms, confirming the diagnosis of GCT of bone with malignant transformation. (E) Sanger sequencing confirmed the presence of an H3 histone family member 3A (H3F3A) mutation (c.103G>T;p.G34W) with a mutant allele fraction of approximately 40%. (F) Wild-type sequence from normal tissue is shown as the reference.
Figure 5.

Giant cell tumor (GCT) of bone with malignant transformation (case 34). (A) The core needle biopsy (H & E) showed nonneoplastic multinucleated giant cells in a background of mononuclear round and spindle cells, consistent with GCTof bone, but (B) immunohistochemistry for H3G34W was negative. (C) On surgical resection performed 3 years later, the recurrent tumor showed morphologic features of an unclassified spindle cell sarcoma with nuclear atypia and frequent mitoses and only scattered giant cells. (D) Again, H3G34W staining was negative. (E) Sanger sequencing detected an alternate H3 histone family member 3A (H3F3A) mutation (c.103_104GG>TT;p.G34L) with a mutant allele fraction of approximately 40% that was not detected by the H3G34W antibody, confirming the diagnosis of GCT of bone with malignant transformation. (F) Wild-type sequence from normal tissue is shown as the reference.
The 3 conventional H3G34W-negative GCTs of bone demonstrated classic clinical presentations and arose in the distal femur of a 22-year-old woman (case 9), in the sacrum of a 20-year-old woman (case 13), and in the proximal femur of a 25-year-old man (case 23) (Fig. 6).
Figure 6.

Conventional giant cell tumor (GCT) of bone (case 23). (A) The core needle biopsy (H & E) showed fragments of ovoid to spindly mononuclear cells with admixed multinucleated giant cells (Inset) in a hemorrhagic background. (B) Immunohistochemistry for H3G34W was negative. (C) The resection specimen showed morphologic features of conventional GCT of bone, but (D) tumor cells lacked H3G34W expression. (E) Sanger sequencing confirmed the presence of an alternate H3 histone family member 3A (H3F3A) mutation (c.104G>T;p.G34V) with a mutant allele fraction of approximately 15%. (F) Wild-type sequence from normal tissue is shown as the reference.
In cases of chondroblastoma, IHC for H3K36M was found to be positive in all 8 cases (100%). The positive staining was detected in a total of 13 of 18 biopsy and excision specimens (72%), including 3 of 3 FNA specimens (100%), 6 of 7 CNB specimens (86%), and 4 of 8 curettings (50%). Positive H3K36W IHC nuclear staining highlighted mononuclear tumor cells only (Fig. 7B).
Figure 7.

Immunohistochemistry for H3K36M was negative in (A) giant cell tumor of bone, but positive in (B) the neoplastic cells of chondroblastoma. (C) Aneurysmal bone cyst and (D) osteosarcoma were negative for H3K36M.
All GCTs of bone were negative for H3K36M (Table 1) (Fig. 7A), and all chondroblastoma cases were negative for H3G34W (Fig. 3B).
All biopsy and excision specimens for the cases of GCT of Paget disease (1 case), aneurysmal bone cysts (7 cases), and osteosarcoma (13 cases) were negative for both H3G34W and H3K36M (Table 1) (Figs. 3C to 3D and Figs. 7C to 7D, respectively).
Taken together, the specificity and sensitivity of H3G34W IHC for the diagnosis of GCTs of bone among giant cell-rich bone tumors were 100% and 85%, respectively. Both the specificity and sensitivity of H3K36M IHC for the diagnosis of chondroblastoma were 100%. IHC was concordant between the biopsy and surgical specimens in 152 of 158 samples (96%). All 42 FNA specimens demonstrated concordant IHC results with concurrent CNB and/or subsequent excision specimens. Six of 158 specimens for which previous or subsequent samples demonstrated positive staining for H3G34W or H3K36M were negative, including 2 CNB samples and 4 curetting/excision specimens.
Sanger Sequencing
Two GCTs of bone that were negative for H3G34W on IHC were found to have alternate H3F3A codon 34 mutations: case 23 harbored an alternate H3F3A G34V mutation (c.104G>T) (mutant allele fraction of approximately 15%) (Fig. 6) and case 34 harbored an alternate H3F3A G34L mutation (c.103_104GG>TT) (mutant allele fraction of approximately 40%) (Fig. 5). As expected, one H3G34W-positive malignant GCT of bone (case 33) harbored an H3F3A G34W mutation (c.103G>T;p.G34W) (mutant allele fraction of approximately 30%) (Fig. 4). All 3 cases were H3F3B wild-type (data not shown). No H3F3A or H3F3B mutations could be detected in the remaining 2 H3G34W IHC-negative GCTs of bone and the GCT of Paget disease (data not shown). As expected, sequence analyses in an H3K36M-positive chondroblastoma (case 3) demonstrated an H3F3B K36M (c.110A>T) mutation (Fig. 8) and did not demonstrate an H3F3A mutation.
Figure 8.

Chondroblastoma (case 3). (A) The cell block (H & E) showed a small cluster of mononuclear cells with admixed multinucleated giant cells (Inset). (B) Immunohistochemistry for H3K36M was positive in tumor cells (approximately 60%). The curetting (H & E stain, C) showed tumor cells with round to ovoid nuclei and eosinophilic cytoplasm, embedded in abundant eosinophilic chondroid matrix with (D) expression of H3K36M. (E) Sanger sequencing confirmed the presence of an H3 histone family member 3B (H3F3B) mutation (c.110A>T;p.K36M). (F) Wild-type sequence from normal tissue is shown as the reference.
DISCUSSION
The utility of FNA and/or CNB in the evaluation of primary bone tumors currently is limited to a few large tertiary centers10,21 due to a combination of the uneasiness of general cytopathologists with regard to primary bone tumors, a high insufficient rate in certain tumors,6 mounting challenges in distinguishing highly malignant giant cell-rich osteosarcoma from locally aggressive giant cell-rich neoplasms such as GCT of bone and chondroblastoma,7–10 and the lack of helpful diagnostic markers for these difficult differential diagnoses. Recent advances in understanding the underlying genetics in bone tumors, notably H3.3 mutations in GCT of bone and chondroblastoma,11 have not only broadened our understanding of the pathogenesis of these bone tumors but also have provided us with novel diagnostic tools, thereby opening a new era of using FNA and/or CNB in the evaluation of primary bone tumors.
Amary et al17 recently demonstrated a high sensitivity for H3G34W IHC in detecting GCT of bone in full tissue and tissue microarray sections: 90.6% of all cases and 97.8% of cases with alternate H3.3G34V, G34R, or G34L mutations were excluded. In the current study, using the same rabbit monoclonal antibody, IHC for H3G34W demonstrated a slightly lower but similar sensitivity of 85% of all cases including 2 cases with alternate mutations. Therefore, in tumors with an otherwise classic morphology and clinical presentation, negative H3G34W staining should not exclude a diagnosis of GCT of bone. Sequencing analysis and/or additional IHC with variant mutant-specific antibodies may aid in confirming the diagnosis, if necessary.22
The reported frequency of H3F3A mutations in GCT of bone appears to depend on the sequencing approaches, and ranges from 69% for Sanger sequencing13 to 96% for targeted massively parallel sequencing.16 With a detection rate of approximately 15% to 20% mutant alleles,23 Sanger sequencing may not always be sensitive enough and previously was shown to miss an H3F3A mutation at a mutant allele fraction of 12%, which was detected by pyrosequencing.14 Because nonneoplastic cells constitute up to 50% of cells in GCT of bone and chondroblastoma, Sanger sequencing may lead to false-negative results. Case 23 in the current study demonstrated low mutant allele fractions of only approximately 15% (Fig. 6E), reaching just above the threshold of detection for Sanger sequencing. Therefore, it is possible that the 2 remaining cases of H3G34W-negative GCT of bone in the current study, which appeared to be wild-type, harbored H3.3 G34 mutations at allele fractions that were beyond the sensitivity of detection.
Cases 33 and 34 illustrate the challenges of distinguishing malignant GCT of bone from osteosarcoma (even in excision specimens) based on morphology alone. In case 33 with a confirmed H3F3A G34W mutation, IHC for H3G34W was positive in biopsy samples demonstrating conventional GCT of bone as well as on the final surgical resection specimen showing morphologic features of chondroblastoma-like osteosarcoma, thus confirming the diagnosis of GCT of bone with malignant transformation (Fig. 4). In case 34, IHC for H3G34W was negative in the initial FNA/CNB, which demonstrated morphology of classic GCT of bone, in the FNA/CNB obtained at the time of disease recurrence, as well as in the surgical resection specimen showing a malignant spindle cell neoplasm. In conjunction with aggressive imaging features, the morphologic findings of a malignant spindle cell neoplasm were interpreted as osteosarcoma. Sanger sequencing revealed an alternate H3F3A G34L mutation, which is not detected by the mutation-specific H3G34W antibody, thereby supporting the belief that this “osteosarcoma” in fact represents malignant transformation of GCT of bone (Fig. 5).
H3.3 G34W/G34R mutations have been reported in rare cases of osteosarcoma and chondrosarcoma.11,14,17 In agreement with Amary et al,17 we believe that such cases represent malignancy in GCT of bone rather than true osteosarcoma. In a recent study, 6 of 106 osteosarcomas were found to harbor H3F3A hotspot mutations, including H3.3 G34W, G34R, and K27M.24 However, the authors demonstrated that the DNA methylation profiles of these H3.3 G34-mutant osteosarcomas differed from H3F3 wild-type osteosarcomas and were more closely related to GCT of bone,24 further supporting the view that these tumors could represent malignant GCT of bone rather than osteosarcoma.
In the current study, the treatment of patients with GCT of bone with denosumab, chemotherapy, or radiotherapy did not appear to alter the intensity or extent of H3G34W staining, in keeping with the findings reported in a very recent study by Yamamoto et al.22 In contrast, Luke et al reported a reduction in the H3G34W-positive cells in GCT of bone after treatment with denosumab.25 At the molecular level, conflicting results also have been reported. Although some groups demonstrated preservation of the H3F3A mutation in 2 cases of GCT of bone after denosumab therapy,20 others found an apparent loss of mutant alleles after treatment in 2 of 3 cases.14 In the current study, the H3F3A G34L mutation was detected in surgical resection specimens after treatment with denosumab (case 34). Additional study is needed to clarify these conflicting results.
Similar to the results reported by Amary et al,18 the results of the current study indicate that H3K36M IHC was highly sensitive and specific for the diagnosis of chondroblastoma, and was found to be positive in all 8 cases (100%) and negative in all histologic mimics. The ability of H3K36M IHC to detect a few clusters of chondroblastoma cells on FNA cell block sections highlights the diagnostic value of the mutation-specific IHC in very scant FNA specimens. The calcaneus tumor from a 28-year-old man (case 3) initially was misdiagnosed as an aneurysmal bone cyst on FNA. The histology on surgical curetting demonstrated features typical of chondroblastoma and the diagnosis was confirmed further by H3K36M positivity and by the presence of an H3F3B mutation by Sanger sequencing (Fig. 8).
In the current series, the original diagnosis was changed in 3 of 34 tumors (8.8%) (Table 2) based on the results of IHC (2 cases; cases 3 and 33) and Sanger sequencing (1 case; case 34). IHC for H3G34W would have helped to establish the definitive diagnosis of GCT of bone in an additional 4 FNA cases and 4 CNB cases, in which only nondiagnostic, benign histiocytic proliferation, or equivocal “giant cell-rich neoplasm” diagnoses were rendered. These results highlight the diagnostic value of mutation-specific antibodies in difficult cases, especially in FNA/CNB specimens.
Although giant cell reparative granuloma (mostly located in the jaw) and brown tumor of hyperparathyroidism enter the differential diagnosis of giant cell-rich bone tumors on small biopsies, these lesions are rare and infrequently sampled by FNA. Therefore, we were not able to include these entities in the current study. However, previous studies performed on surgical resections by Amary et al18 and Yamamoto et al22 included large numbers of cases of giant cell granuloma of the jaw, fibrous dysplasia, tenosynovial GCT, and brown tumors, none of which were found to express H3G34W or H3K36M by IHC.
The findings of the current study confirm the high specificity of mutation-specific H3G34W and H3K36M antibodies for the diagnosis of GCT of bone and chondroblastoma respectively, in FNA and CNB specimens, which often are the first samples to be evaluated in the diagnostic workup of giant cell-rich bone tumors. These markers are of high diagnostic usefulness, especially in certain challenging settings when clinicoradiologic presentations and/or cytomorphologic features are not definitive. The presence of alternate H3F3A mutations accounts for H3G34W IHC negativity in a subset of GCT of bone cases. The detection of H3F3A mutations by H3G34W IHC and/or by sequencing in malignant tumors arising from GCT of bone aids in the distinction between malignant GCT of bone and osteosarcoma.
FUNDING SUPPORT
No specific funding was disclosed.
We thank Dr. Jeffrey Mito for critical review of the manuscript and Ms. Mei Zheng for excellent technical support.
Footnotes
Presented at the 107th Annual Meeting of the United States and Canadian Academy of Pathology (USCAP 2018); March 17–23, 2018; Vancouver, British Columbia, Canada.
CONFLICT OF INTEREST DISCLOSURES
The authors made no disclosures.
REFERENCES
- 1.Orosz Z, Athanasou NA. Giant cell-containing tumors of bone. Surg Pathol Clin 2017;10:553–573. [DOI] [PubMed] [Google Scholar]
- 2.van der Heijden L, Dijkstra PDS, Blay JY, Gelderblom H. Giant cell tumour of bone in the denosumab era. Eur J Cancer 2017;77: 75–83. [DOI] [PubMed] [Google Scholar]
- 3.Biermann JS, Chow W, Reed DR, et al. NCCN Guidelines Insights: Bone Cancer, Version 2.2017. J Natl Compr Canc Netw 2017;15:155–167. [DOI] [PubMed] [Google Scholar]
- 4.Fletcher C, Bridge JA, Hogendoorn PCW, Mertens F. WHO Classification of Tumours of Soft Tissue and Bone Lyon, France: IARC Press; 2013. [Google Scholar]
- 5.VandenBussche CJ, Sathiyamoorthy S, Wakely PE Jr, Ali SZ. Chondroblastic osteosarcoma: cytomorphologic characteristics and differential diagnosis on FNA. Cancer Cytopathol 2016;124:493–500. [DOI] [PubMed] [Google Scholar]
- 6.Creager AJ, Madden CR, Bergman S, Geisinger KR. Aneurysmal bone cyst: fine-needle aspiration findings in 23 patients with clinical and radiologic correlation. Am J Clin Pathol 2007;128:740–745. [DOI] [PubMed] [Google Scholar]
- 7.Fanning CV, Sneige NS, Carrasco CH, Ayala AG, Murray JA, Raymond AK. Fine needle aspiration cytology of chondroblastoma of bone. Cancer 1990;65:1847–1863. [DOI] [PubMed] [Google Scholar]
- 8.Agarwal S, Agarwal T, Agarwal R, Agarwal PK, Jain UK. Fine needle aspiration of bone tumors. Cancer Detect Prev 2000;24:602–609. [PubMed] [Google Scholar]
- 9.Kilpatrick SE, Pike EJ, Geisinger KR, Ward WG. Chondroblastoma of bone: use of fine-needle aspiration biopsy and potential diagnostic pitfalls. Diagn Cytopathol 1997;16:65–71. [DOI] [PubMed] [Google Scholar]
- 10.Cardona DM, Dodd LG. Bone cytology: a realistic approach for clinical use. Surg Pathol Clin 2012;5:79–100. [DOI] [PubMed] [Google Scholar]
- 11.Behjati S, Tarpey PS, Presneau N, et al. Distinct H3F3A and H3F3B driver mutations define chondroblastoma and giant cell tumor of bone. Nat Genet 2013;45:1479–1482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Al-Ibraheemi A, Inwards CY, Zreik RT, et al. Histologic spectrum of giant cell tumor (GCT) of bone in patients 18 years of age and below: a study of 63 patients. Am J Surg Pathol 2016;40:1702–1712. [DOI] [PubMed] [Google Scholar]
- 13.Cleven AH, Hocker S, Briaire-de Bruijn I, Szuhai K, Cleton-Jansen AM, Bovee JV. Mutation analysis of H3F3A and H3F3B as a diagnostic tool for giant cell tumor of bone and chondroblastoma. Am J Surg Pathol 2015;39:1576–1583. [DOI] [PubMed] [Google Scholar]
- 14.Kervarrec T, Collin C, Larousserie F, et al. H3F3 mutation status of giant cell tumors of the bone, chondroblastomas and their mimics: a combined high resolution melting and pyrosequencing approach. Mod Pathol 2017;30:393–406. [DOI] [PubMed] [Google Scholar]
- 15.Nohr E, Lee LH, Cates JM, Perizzolo M, Itani D. Diagnostic value of histone 3 mutations in osteoclast-rich bone tumors. Hum Pathol 2017;68:119–127. [DOI] [PubMed] [Google Scholar]
- 16.Presneau N, Baumhoer D, Behjati S, et al. Diagnostic value of H3F3A mutations in giant cell tumour of bone compared to osteoclast-rich mimics. J Pathol Clin Res 2015;1:113–123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Amary F, Berisha F, Ye H, et al. H3F3A (histone 3.3) G34W immunohistochemistry: a reliable marker defining benign and malignant giant cell tumor of bone. Am J Surg Pathol 2017;41:1059–1068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Amary MF, Berisha F, Mozela R, et al. The H3F3 K36M mutant antibody is a sensitive and specific marker for the diagnosis of chondroblastoma. Histopathology 2016;69:121–127. [DOI] [PubMed] [Google Scholar]
- 19.Ravegnini G, Marino-Enriquez A, Slater J, et al. MED12 mutations in leiomyosarcoma and extrauterine leiomyoma. Mod Pathol 2013;26:743–749. [DOI] [PubMed] [Google Scholar]
- 20.Girolami I, Mancini I, Simoni A, et al. Denosumab treated giant cell tumour of bone: a morphological, immunohistochemical and molecular analysis of a series. J Clin Pathol 2016;69:240–247. [DOI] [PubMed] [Google Scholar]
- 21.Domanski HA, Qian X, Stanley DE. Bone. In: Domanski HA, ed. Atlas of Fine Needle Aspiration Cytology 2nd ed. London: Springer; 2018. [Google Scholar]
- 22.Yamamoto H, Iwasaki T, Yamada Y, et al. Diagnostic utility of histone H3.3 G34W, G34R, and G34V mutant-specific antibodies for giant cell tumors of bone. Hum Pathol 2018;73:41–50. [DOI] [PubMed] [Google Scholar]
- 23.Tsiatis AC, Norris-Kirby A, Rich RG, et al. Comparison of Sanger sequencing, pyrosequencing, and melting curve analysis for the detection of KRAS mutations: diagnostic and clinical implications. J Mol Diagn 2010;12:425–432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Koelsche C, Schrimpf D, Tharun L, et al. Histone 3.3 hotspot mutations in conventional osteosarcomas: a comprehensive clinical and molecular characterization of six H3F3A mutated cases. Clin Sarcoma Res 2017;7:9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Luke J, von Baer A, Schreiber J, et al. H3F3A mutation in giant cell tumour of the bone is detected by immunohistochemistry using a monoclonal antibody against the G34W mutated site of the histone H3.3 variant. Histopathology 2017;71:125–133. [DOI] [PubMed] [Google Scholar]


