Abstract
With the growing requirement for novel prognostic biomarkers for pancreatic cancer, many studies have focused on clinical and/or genomic variables. Although many studies have been performed, carbohydrate antigen 19-9 is the only biomarker in clinical use. Therefore, the present study examined whether γ-secretase genes, including presenilin (PSEN), nicastrin (NCSTN), presenilin enhancer protein 2 (PSENEN), and anterior pharynx-defective 1 (APH1-), could serve as prognostic factors for pancreatic cancer. The cohorts selected included >100 pancreatic cancer patients. Patient data were downloaded from The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GSE21501). The prognostic roles of the γ-secretase genes were analyzed by several survival analysis methods. Among the γ-secretase genes, the prognosis tended to be worse in the 2 cohorts with increasing expression of PSEN1, APH1A, and PSENEN, while the remaining genes were the opposite in the 2 cohorts. Notably, although the patient characteristics were quite different, APH1A was statistically significantly associated with prognosis in the 2 cohorts. The hazard ratio of APH1A for overall survival was 1.598 (TCGA) and 2.724 (GSE21501). These results contribute to the study of γ-secretase in pancreatic cancer. We believe that γ-secretase, particularly APH1A, will be a new prognostic biomarker for pancreatic cancer.
Keywords: γ-secretase, Notch signaling, pancreatic cancer, prognosis, The Cancer Genome Atlas, Gene Expression Omnibus
Introduction
Pancreatic cancer is the fourth-leading cause of cancer deaths in the US and has a very poor prognosis with a mortality approaching 100% (1–4). Only 15–20% of pancreatic cancer cases are diagnosed early due to nonspecific symptoms and approximately 50% of patients have metastasis at diagnosis (5,6). As precision and personalized medicine emerged, research on the genomics of patients with various cancers to identify novel biomarkers is actively being pursued (7,8).
γ-Secretase is an intramembrane-cleaving protease composed of an essential protein complex including γ-secretase complex-presenilin (PSEN), nicastrin (NCSTN), presenilin enhancer protein 2 (PSENEN), and anterior pharynx-defective 1 (APH1-) (9,10). Either PSEN1 or PSEN2 contains NCSTN, the catalytic site, and the PSENEN and APH1 (APH1A or APH1B) substrate binding site (10,11). Notch signaling is one of the most evolutionarily conserved pathways involved in cell fate control during development and postnatal tissue differentiation (10,12). Aberrations in this signaling pathway have been identified in several diseases, including Alzheimer's disease, and malignancies (10,12–15). γ-Secretase is not only able to cleave type I transmembrane proteins, but also the amyloid precursor protein and NOTCH receptors (12). Because abnormal activation of the Notch pathway plays a role in tumorigenesis, γ-secretase, which mediates Notch signals, is a promising therapeutic target (12,16).
Several Notch signal inhibitors have been developed, including γ-secretase inhibitors (GSIs), siRNA, and monoclonal antibodies (17,18). In in vitro studies, the administration of the GSI MRK-003 with gemcitabine decreased pancreatic adenocarcinoma cancer cell growth and suppressed cancer stem cells (18–20). In this context, γ-secretase is thought to play a significant role in tumorigenesis and survival in pancreatic cancer.
The present study retrospectively examined the prognostic significance of γ-secretase genes in patients with pancreatic cancer in independent cohorts. Among these genes, high expression of APH1A was significantly associated with a worse prognosis in patients with pancreatic cancer.
Materials and methods
Patients
The patients' data were downloaded from The Cancer Genome Atlas (TCGA) (21,22) and the Gene Expression Omnibus (GEO, GSE21501, GSE28735, GSE15471, GSE16515) in October 2017 (23–26). We included only cohorts (TCGA, GSE21501) containing more than 100 patients with pancreatic cancer in which survival information was available. In TCGA and GSE21501, the cancer staging system follows The American Joint Committee on Cancer (AJCC) (27). These processes were performed using R software version 3.5.0 (The R Foundation for Statistical Computing, 2018), using the ‘cgdsr’ and ‘GEOquery’ packages.
Survival and statistical analysis
Survival analyses were performed to predict the overall survival (OS). We used following three methods: i) Kaplan-Meier survival curves with log-rank test to evaluate the accuracy of the discrimination, ii) Uno's C-index in the time-dependent area under the curve (AUC) analysis, and iii) AUC values in receiver operating characteristics (ROC) at three years as we used previously (28–31). These values were calculated using the R packages ‘survival’ and ‘survAUC’. C-indices and AUC values of 0.6 or greater were considered acceptable for survival predictions. In the Kaplan-Meier analyses, we determined the optimal cutoff value with the maximal Uno's C-index by five-fold cross-validation as we performed before (30,32). We used univariate Cox regression to identify the effect of the genes on prognosis. To compare the γ-secretases expression value between tumor and non-tumor tissues, we used Wilcoxon signed rank test or t test according to their expression distribution in GSE28735, GSE15471 and GSE16515. All statistical analyses were performed using R.
Results
Patient distribution
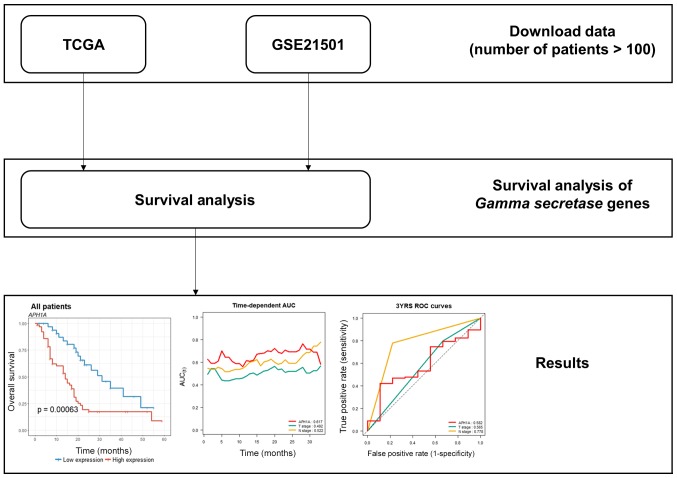
In total, 172 patients (TCGA) and 102 patients (GSE21501) were included in this study. Their information is described in Table I and the characteristics of each cohort were quite different. The patients in TCGA were almost all at low stages (AJCC staging system, Stage I, 21; Stage II, 145; Stage III 3; Stage IV, 5), while the patients in GSE21501 were almost all at high stages (T1, 2; T2, 16; T3, 79; T4, 1; N0, 28; N1, 73). A flowchart of the present study is described in Fig. 1. There are 45, 36 pairs of pancreatic tumor and adjacent non-tumor tissues in GSE28735, GSE15471. GSE16515 dataset consists of 36 tumors and 16 normal samples.
Table I.
Patients' information used in current research in the TCGA and GSE21501 cohorts.
| Group | Description | TCGA | GSE21501 |
|---|---|---|---|
| Male | – | 94 | − |
| Female | – | 78 | − |
| Stage I | – | 21 | − |
| Stage II | – | 145 | − |
| Stage III | – | 3 | − |
| Stage IV | – | 5 | − |
| T1 | Confined to the pancreas, ≤2 cm in diameter | − | 2 |
| T2 | Confined to the pancreas, 2 cm < diameter ≤4 cm | − | 16 |
| T3 | Confined to the pancreas, >4 cm in diameter | − | 79 |
| T4 | Extends outside the pancreas and into nearby major blood vessels | − | 1 |
| N0 | No spread to nearby lymph nodes | − | 28 |
| N1 | Spreads to no more than 3 nearby lymph nodes | − | 73 |
Figure 1.
Study flowchart. TCGA, The Cancer Genome Atlas.
Survival curve of γ-secretases
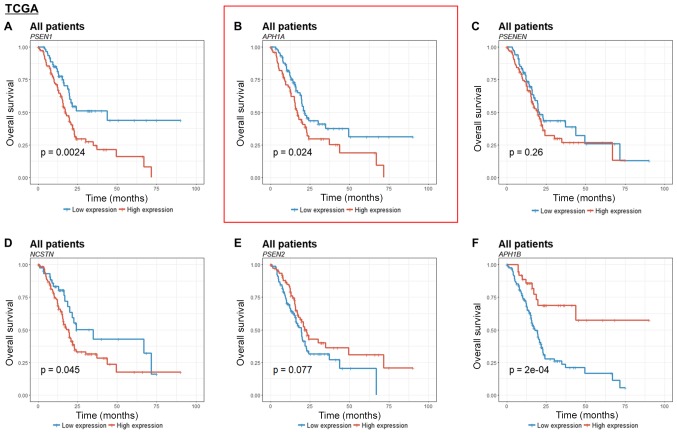
To identify the discriminatory power of γ-secretase genes as a categorical variable, we analyzed Kaplan-Meier curves with log-rank test for gene expression and survival. Among the γ-secretase genes, only APH1A was significantly associated with survival in both cohorts (Figs. 2B and 3B). High expression of PSEN1 and PSENEN showed a poor prognosis as the expression levels increased in both cohorts and were statistically significant in GSE21501 and TCGA respectively (Figs. 2A and C, and 3A and C). The other genes were not statistically significant or showed opposite trends in each cohort (Figs. 2 and 3). The number of patients and deaths in the high- and low-risk groups divided by each gene are listed in Table II. The prognostic values were further confirmed using univariate Cox regression analysis (Table III).
Figure 2.
Kaplan-Meier estimates of the overall survival of pancreatic cancer patients according to γ-secretase gene expression in the TCGA cohort. (A) PSEN1, (B) APH1A, (C) PSENEN, (D) NSCTN, (E) PSEN2 and (F) APH1B. TCGA, The Cancer Genome Atlas; PSEN, presenilin; APH1, anterior pharynx-defective 1; PSENEN, presenilin enhancer protein 2; NCSTN, nicastrin.
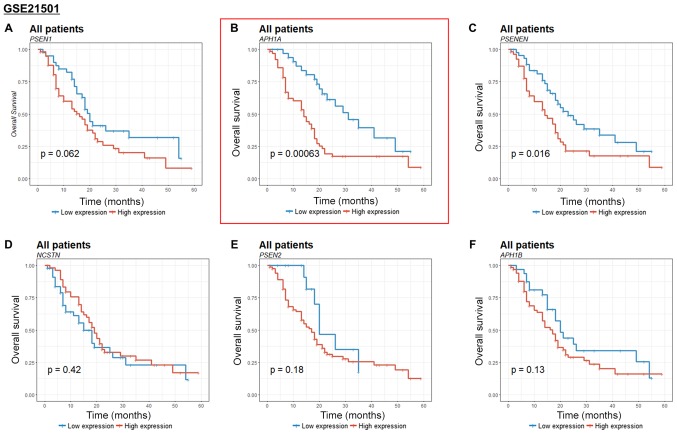
Figure 3.
Kaplan-Meier estimates of the overall survival of pancreatic cancer patients according to γ-secretase gene expression in the GSE21501 cohort. (A) PSEN1, (B) APH1A, (C) PSENEN, (D) NSCTN, (E) PSEN2, (F) APH1B. PSEN, presenilin; APH1, anterior pharynx-defective 1; PSENEN, presenilin enhancer protein 2; NCSTN, nicastrin.
Table II.
Number of patients (event) divided by each gene cutoff value.
| Gene | Group | TCGA | GSE21501 |
|---|---|---|---|
| Total values (n) | All patients | 172 | 102 |
| PSEN1 | High expression (event) | 107 (68) | 61 (42) |
| Low expression (event) | 65 (24) | 41 (24) | |
| APH1A | High expression (event) | 75 (49) | 68 (49) |
| Low expression (event) | 97 (43) | 34 (17) | |
| PSENEN | High expression (event) | 101 (57) | 56 (40) |
| Low expression (event) | 71 (35) | 46 (26) | |
| NCSTN | High expression (event) | 127 (74) | 57 (38) |
| Low expression (event) | 45 (18) | 45 (28) | |
| PSEN2 | High expression (event) | 79 (39) | 86 (59) |
| Low expression (event) | 93 (53) | 16 (7) | |
| APH1B | High expression (event) | 37 (10) | 69 (47) |
| Low expression (event) | 135 (82) | 33 (19) |
PSEN, presenilin; APH1, anterior pharynx-defective 1; PSENEN, presenilin enhancer protein 2; NCSTN, nicastrin.
Table III.
Univariate cox regression results of each gene in TCGA and GSE21501 cohorts.
| A, TCGA | ||||
|---|---|---|---|---|
| Genes | P-value | Hazard ratio | 95% confidence interval | |
| PSEN1 | 0.00291b | 2.0355 | 1.275 | 3.25 |
| APH1A | 0.025a | 1.598 | 1.061 | 2.408 |
| PSENEN | 0.26 | 1.275 | 0.8353 | 1.946 |
| NCSTN | 0.0469a | 1.7011 | 1.007 | 2.873 |
| PSEN2 | 0.0782 | 1.4574 | 0.9583 | 2.216 |
| APH1B | <0.0001c | 3.2734 | 1.691 | 6.335 |
| B, GSE21501 | ||||
| Genes | P-value | Hazard ratio | 95% confidence interval | |
| PSEN1 | 0.0835 | 1.5772 | 0.9414 | 2.642 |
| APH1A | <0.0001c | 2.724 | 1.517 | 4.892 |
| PSENEN | 0.0248a | 1.7894 | 1.076 | 2.975 |
| NCSTN | 0.371 | 1.257 | 0.7623 | 2.071 |
| PSEN2 | 0.195 | 1.747 | 0.7517 | 4.06 |
| APH1B | 0.153 | 1.4939 | 0.861 | 2.592 |
P<0.05
P<0.01
P<0.0001. PSEN, presenilin; APH1, anterior pharynx-defective 1; PSENEN, presenilin enhancer protein 2; NCSTN, nicastrin; TCGA, The Cancer Genome Atlas.
C-index and AUC values of γ-secretase genes
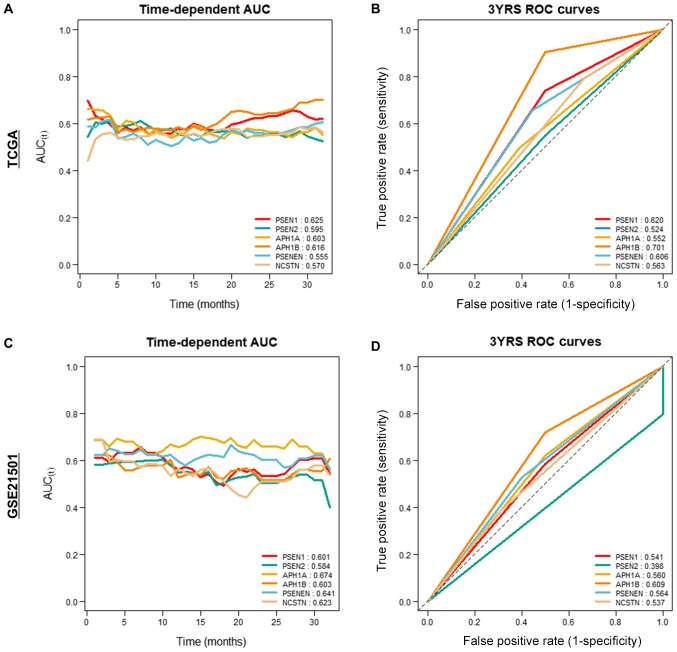
We compared γ-secretase genes in both cohorts to evaluate their prognostic value. We analyzed the gene expression values as continuous variables using Uno's C-indices and AUC values at three years. APH1A had high C-index values in the two independent cohorts compared to those of the other genes (Fig. 4). The three-year AUC value was also significantly higher than that of the other factors across the two cohorts (Fig. 4).
Figure 4.
Time-dependent AUC and ROC curve at 3 years in the (A and B) TCGA and (C and D) ICGC cohorts. AUC, area under the curve; ROC, receiver operator characteristic; TCGA, The Cancer Genome Atlas; ICGC, International Cancer Genome Consortium; YRS, years.
The differences of expression values between normal and tumor tissues
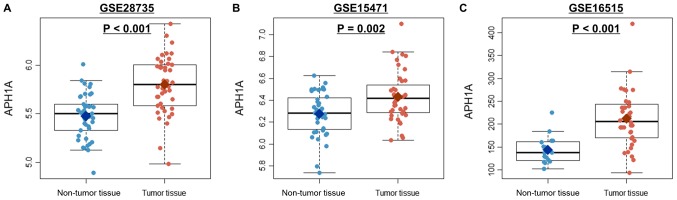
By using Wilcoxon rank sum test, we compared the expression of γ-secretase genes in three independent cohorts. As shown in Table IV, the expression patterns of the γ-secretase genes except APH1B and NCSTN were statistically significant in agreement with the three independent cohorts. APH1A, the only statistically significant gene in survival curves, was found to be more expressed in cancer tissues compared to normal tissues in the three cohorts (Fig. 5 and Table IV).
Table IV.
Differences in gene expression between cancerous and normal tissues.
| GSE28735 | GSE15471 | GSE16515 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Genes | P-value | Cancer (mean) | Normal (mean) | P-value | Cancer (mean) | Normal (mean) | P-value | Cancer (mean) | Normal (mean) |
| PSEN1 | <0.001 | 6.462 | 5.932 | <0.001 | 5.677 | 5.503 | <0.001 | 85.603 | 50.961 |
| APH1A | <0.001 | 5.799 | 5.476 | 0.002 | 6.416 | 6.282 | <0.001 | 211.812 | 144.006 |
| PSENEN | <0.001 | 6.347 | 6.038 | 0.006 | 7.726 | 7.456 | <0.001 | 168.60 | 107.950 |
| NCSTN | 0.016 | 6.162 | 6.024 | <0.001 | 6.635 | 6.732 | 0.0501 | 154.240 | 134.207 |
| PSEN2 | <0.001 | 5.251 | 5.689 | <0.001 | 6.045 | 6.625 | 0.032 | 38.227 | 58.335 |
| APH1B | 0.41 | 5.123 | 5.035 | <0.001 | 6.952 | 6.688 | 0.504 | 57.535 | 60.265 |
PSEN, presenilin; APH1, anterior pharynx-defective 1; PSENEN, presenilin enhancer protein 2; NCSTN, nicastrin.
Figure 5.
APH1A expression differences between cancer and normal tissues. (A) GSE28735, (B) GSE15471 and (C) GSE16515. APH1A, anterior pharynx-defective 1A.
Discussion
The present study investigated the prognostic significance of γ-secretase genes in pancreatic cancer using two independent cohorts each containing more than 100 patients. Although the characteristics of the two cohorts were quite different, high expression levels of PSEN1, APH1A, and PSENEN were associated with a poor prognosis in both cohorts. Especially, APH1A is the only statistically significant gene in all analyses.
Owing to the poor survival rate of pancreatic cancer, it is necessary to identify prognostic markers for patients to determine the precise treatment strategy. Although certain studies have attempted to predict the survival of patients with pancreatic cancer based on clinical variables and/or expression profiles, carbohydrate antigen 19-9 (CA-19-9) is the only biomarker approved by the US Food and Drug Administration (FDA) (33,34). As the development of genetics and the importance of data sharing have been emphasized, relatively rare pancreatic cancer data have been collected and released. For the implementation of precision medicine, a number of cohorts and biomarkers verified in data from many patients are needed. Hence, we included and analyzed cohorts with more than 100 patients (21–23).
Notch signaling promotes the tumorigenesis of lung cancer in the hypoxic state (12,35). In breast cancer, this signaling has been implicated in tumorigenesis (36–38). In this context, GSIs have been tested for their therapeutic activity in cancer cell lines (breast and lung) and several clinical trials (12,37,39). Moreover, the activation of Notch signals has been implicated in the progression of pancreatic cancer and MRK-003, a potent pan-NOTCH inhibitor has been shown to be an effective treatment for pancreatic adenocarcinoma (19). In our study, as the expression of some Notch substrates (PSEN1, APH1A, and PSENEN) increases, the prognosis of the patient worsens. Also, their expression levels of cancer are significantly higher than normal tissues. Thus, higher expression of them may play a significant role in tumor progression in pancreatic cancer. Our findings suggest that MRK-003 is in line with findings that is effective in the suppression of pancreatic cancer.
Some studies have reported that the different composition of six γ-secretase subunits have various enzymatic functions and cause varying physiological outcome. For instance, some researchers suggested that complexes in which the PSEN1 and/or APH1A subunits were the major constituents could cause Alzheimer's disease if PSEN2 and/or APH1B were changed to a major member (40,41). In the current study, APH1A, PSEN1, and PSENEN increased in cancer tissues compared to normal tissues, while PSEN2 decreased and APH1B and NCSTN did not change. These results suggested that they may play a role in the progression of pancreatic cancer. Especially, APH1A is closely related to survival, so APH1A is considered to play the most important role in changed γ-secretase subunits. Therefore, in future studies of pancreatic cancer, it will be necessary to study the role and function of APH1A as a major component of γ-secretase.
In conclusion, we found that the survival rate of patients with pancreatic cancer with high PSEN1, APH1A, and PSENEN expression was low. Although further studies are needed to assess the role of γ-secretase in tumorigenesis, our findings suggest that some γ-secretase substrates, especially APH1A, may have a critical role on the development and progression of pancreatic cancer. We believe that these findings will contribute to the prognostic prediction of pancreatic cancer.
Acknowledgements
Not applicable.
Funding
This work was supported by Pusan National University Research Grant, 2018 and the Basic Science Research Program through a National Research Foundation of Korea (NRF) grant funded by the Korean government (MOE; grant no. NRF-2016R1A2B4014593). It was also supported by the Medical Research Center Program through the National Research Foundation of Korea grant funded by the Korea government (MSIP; grant no. NRF-2015R1A5A2009656).
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Authors' contributions
YHJ and MH analyzed data and wrote the manuscript. SWK, MJK and MEH collected the data. MH performed the statistical analysis. MEH contributed to the conception of the study. CSL and CKO performed the statistical analysis. SOO and YHK conceived and designed the study, supervised the project and gave final approval of the version to be published. All authors read and approved the final manuscript.
Ethics approval and consent to participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interest
The authors declare that they have no conflict of interest.
References
- 1.Kleeff J, Michalski C, Friess H, Büchler MW. Pancreatic cancer: From bench to 5-year survival. Pancreas. 2006;33:111–118. doi: 10.1097/01.mpa.0000229010.62538.f2. [DOI] [PubMed] [Google Scholar]
- 2.Kwon HJ, Kim SG. Surgical treatment for advanced pancreatic cancer. Korean J Hepatobiliary Pancreat Surg. 2012;16:89–92. doi: 10.14701/kjhbps.2012.16.3.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lowenfels AB, Maisonneuve P. Epidemiology and risk factors for pancreatic cancer. Best Pract Res Clin Gastroenterol. 2006;20:197–209. doi: 10.1016/j.bpg.2005.10.001. [DOI] [PubMed] [Google Scholar]
- 4.Yeo CJ, Cameron JL, Lillemoe KD, Sitzmann JV, Hruban RH, Goodman SN, Dooley WC, Coleman J, Pitt HA. Pancreaticoduodenectomy for cancer of the head of the pancreas. 201 patients. Ann Surg. 1995;221:721–733. doi: 10.1097/00000658-199506000-00011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Eto S, Ishikawa M, Asanoma M, Tashiro Y, Matsuyama K, Oshio T. A long-term survival case of advanced biliary cancer with repeated resection due to recurrence in the pancreaticogastrostomy site after pancreaticoduodenectomy. Ann Hepatobiliary Pancreat Surg. 2018;22:173–177. doi: 10.14701/ahbps.2018.22.2.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kim HS, Jang JY, Han Y, Lee KB, Joo I, Lee DH, Kim JR, Kim H, Kwon W, Kim SW. Survival outcome and prognostic factors of neoadjuvant treatment followed by resection for borderline resectable pancreatic cancer. Ann Surg Treat Res. 2017;93:186–194. doi: 10.4174/astr.2017.93.4.186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Arnedos M, Vicier C, Loi S, Lefebvre C, Michiels S, Bonnefoi H, Andre F. Precision medicine for metastatic breast cancer-limitations and solutions. Nat Rev Clin Oncol. 2015;12:693–704. doi: 10.1038/nrclinonc.2015.123. [DOI] [PubMed] [Google Scholar]
- 8.Friedman AA, Letai A, Fisher DE, Flaherty KT. Precision medicine for cancer with next-generation functional diagnostics. Nat Rev Cancer. 2015;15:747–756. doi: 10.1038/nrc4015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Selkoe D, Kopan R. Notch and presenilin: Regulated intramembrane proteolysis links development and degeneration. Annu Rev Neurosci. 2003;26:565–597. doi: 10.1146/annurev.neuro.26.041002.131334. [DOI] [PubMed] [Google Scholar]
- 10.Zhao G, Liu Z, Ilagan MX, Kopan R. Gamma-secretase composed of PS1/Pen2/Aph1a can cleave notch and amyloid precursor protein in the absence of nicastrin. J Neurosci. 2010;30:1648–1656. doi: 10.1523/JNEUROSCI.3826-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De Strooper B. Aph-1, Pen-2, and Nicastrin with Presenilin generate an active gamma-Secretase complex. Neuron. 2003;38:9–12. doi: 10.1016/S0896-6273(03)00205-8. [DOI] [PubMed] [Google Scholar]
- 12.Ran Y, Hossain F, Pannuti A, Lessard CB, Ladd GZ, Jung JI, Minter LM, Osborne BA, Miele L, Golde TE. γ-Secretase inhibitors in cancer clinical trials are pharmacologically and functionally distinct. EMBO Mol Med. 2017;9:950–966. doi: 10.15252/emmm.201607265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Halawany AME, Sayed NSE, Abdallah HM, Dine RSE. Protective effects of gingerol on streptozotocin-induced sporadic Alzheimer's disease: Emphasis on inhibition of β-amyloid, COX-2, alpha-, beta-secretases and APH1a. Sci Rep. 2017;7:2902. doi: 10.1038/s41598-017-02961-0. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 14.Penton AL, Leonard LD, Spinner NB. Notch signaling in human development and disease. Semin Cell Dev Biol. 2012;23:450–457. doi: 10.1016/j.semcdb.2012.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.International Cancer Genome Consortium. Hudson TJ, Anderson W, Artez A, Barker AD, Bell C, Bernabé RR, Bhan MK, Calvo F, Eerola I, et al. International network of cancer genome projects. Nature. 2010;464:993–998. doi: 10.1038/nature08987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shih IeM, Wang TL. Notch signaling, gamma-secretase inhibitors, and cancer therapy. Cancer Res. 2007;67:1879–1882. doi: 10.1158/0008-5472.CAN-06-3958. [DOI] [PubMed] [Google Scholar]
- 17.Takebe N, Nguyen D, Yang SX. Targeting notch signaling pathway in cancer: Clinical development advances and challenges. Pharmacol Ther. 2014;141:140–149. doi: 10.1016/j.pharmthera.2013.09.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yuan X, Wu H, Xu H, Xiong H, Chu Q, Yu S, Wu GS, Wu K. Notch signaling: An emerging therapeutic target for cancer treatment. Cancer Lett. 2015;369:20–27. doi: 10.1016/j.canlet.2015.07.048. [DOI] [PubMed] [Google Scholar]
- 19.Mizuma M, Rasheed ZA, Yabuuchi S, Omura N, Campbell NR, de Wilde RF, De Oliveira E, Zhang Q, Puig O, Matsui W, et al. The gamma secretase inhibitor MRK-003 attenuates pancreatic cancer growth in preclinical models. Mol Cancer Ther. 2012;11:1999–2009. doi: 10.1158/1535-7163.MCT-12-0017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.So JY, Wahler J, Das Gupta S, Salerno DM, Maehr H, Uskokovic M, Suh N. HES1-mediated inhibition of Notch1 signaling by a Gemini vitamin D analog leads to decreased CD44(+)/CD24(−/low) tumor-initiating subpopulation in basal-like breast cancer. J Steroid Biochem Mol Biol. 2015;148:111–121. doi: 10.1016/j.jsbmb.2014.12.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cancer Genome Atlas Research Network. Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, Shmulevich I, Sander C, Stuart JM. The cancer genome atlas pan-cancer analysis project. Nat Genet. 2013;45:1113–1120. doi: 10.1038/ng.2764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–404. doi: 10.1158/2159-8290.CD-12-0095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Stratford JK, Bentrem DJ, Anderson JM, Fan C, Volmar KA, Marron JS, Routh ED, Caskey LS, Samuel JC, Der CJ, et al. A six-gene signature predicts survival of patients with localized pancreatic ductal adenocarcinoma. PLoS Med. 2010;7:e1000307. doi: 10.1371/journal.pmed.1000307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Badea L, Herlea V, Dima SO, Dumitrascu T, Popescu I. Combined gene expression analysis of whole-tissue and microdissected pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in tumor epithelia. Hepatogastroenterology. 2008;55:2016–2027. [PubMed] [Google Scholar]
- 25.Pei H, Li L, Fridley BL, Jenkins GD, Kalari KR, Lingle W, Petersen G, Lou Z, Wang L. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell. 2009;16:259–266. doi: 10.1016/j.ccr.2009.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhang G, He P, Tan H, Budhu A, Gaedcke J, Ghadimi BM, Ried T, Yfantis HG, Lee DH, Maitra A, et al. Integration of metabolomics and transcriptomics revealed a fatty acid network exerting growth inhibitory effects in human pancreatic cancer. Clin Cancer Res. 2013;19:4983–4993. doi: 10.1158/1078-0432.CCR-13-0209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK, Meyer L, Gress DM, Byrd DR, Winchester DP. The Eighth Edition AJCC Cancer Staging Manual: Continuing to build a bridge from a population-based to a more ‘personalized’ approach to cancer staging. CA Cancer J Clin. 2017;67:93–99. doi: 10.3322/caac.21388. [DOI] [PubMed] [Google Scholar]
- 28.Cho SH, Pak K, Jeong DC, Han ME, Oh SO, Kim YH. The AP2M1 gene expression is a promising biomarker for predicting survival of patients with hepatocellular carcinoma. J Cell Biochem. 2019;120:4140–4146. doi: 10.1002/jcb.27699. [DOI] [PubMed] [Google Scholar]
- 29.Ha M, Kim JY, Han ME, Kim GH, Park SY, Jeong DC, Oh SO, Kim YH. TMEM18: A novel prognostic marker in acute myeloid leukemia. Acta Haematol. 2018;140:71–76. doi: 10.1159/000492742. [DOI] [PubMed] [Google Scholar]
- 30.Han ME, Kim JY, Kim GH, Park SY, Kim YH, Oh SO. SAC3D1: A novel prognostic marker in hepatocellular carcinoma. Sci Rep. 2018;8:15608. doi: 10.1038/s41598-018-34129-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Goh TS, Lee JS, Il Kim J, Park YG, Pak K, Jeong DC, Oh SO, Kim YH. Prognostic scoring system for osteosarcoma using network-regularized high-dimensional Cox-regression analysis and potential therapeutic targets. J Cell Physiol. 2019 Jan 3; doi: 10.1002/jcp.28065. (Epub ahead of print) [DOI] [PubMed] [Google Scholar]
- 32.Ha M, Han ME, Kim JY, Jeong DC, Oh SO, Kim YH. Prognostic role of TPD52 in acute myeloid leukemia: A retrospective multicohort analysis. J Cell Biochem. 2019;120:3672–3678. doi: 10.1002/jcb.27645. [DOI] [PubMed] [Google Scholar]
- 33.Fong ZV, Winter JM. Biomarkers in pancreatic cancer: Diagnostic, prognostic, and predictive. Cancer J. 2012;18:530–538. doi: 10.1097/PPO.0b013e31827654ea. [DOI] [PubMed] [Google Scholar]
- 34.Winter JM, Yeo CJ, Brody JR. Diagnostic, prognostic, and predictive biomarkers in pancreatic cancer. J Surg Oncol. 2013;107:15–22. doi: 10.1002/jso.23192. [DOI] [PubMed] [Google Scholar]
- 35.Chen Y, De Marco MA, Graziani I, Gazdar AF, Strack PR, Miele L, Bocchetta M. Oxygen concentration determines the biological effects of NOTCH-1 signaling in adenocarcinoma of the lung. Cancer Res. 2007;67:7954–7959. doi: 10.1158/0008-5472.CAN-07-1229. [DOI] [PubMed] [Google Scholar]
- 36.Al-Hussaini H, Subramanyam D, Reedijk M, Sridhar SS. Notch signaling pathway as a therapeutic target in breast cancer. Mol Cancer Ther. 2011;10:9–15. doi: 10.1158/1535-7163.MCT-10-0677. [DOI] [PubMed] [Google Scholar]
- 37.Han J, Shen Q. Targeting γ-secretase in breast cancer. Breast Cancer (Dove Med Press) 2012;4:83–90. doi: 10.2147/BCTT.S26437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Harrison H, Farnie G, Brennan KR, Clarke RB. Breast cancer stem cells: Something out of notching? Cancer Res. 2010;70:8973–8976. doi: 10.1158/0008-5472.CAN-10-1559. [DOI] [PubMed] [Google Scholar]
- 39.Kondratyev M, Kreso A, Hallett RM, Girgis-Gabardo A, Barcelon ME, Ilieva D, Ware C, Majumder PK, Hassell JA. Gamma-secretase inhibitors target tumor-initiating cells in a mouse model of ERBB2 breast cancer. Oncogene. 2012;31:93–103. doi: 10.1038/onc.2011.212. [DOI] [PubMed] [Google Scholar]
- 40.Peltonen HM, Haapasalo A, Hiltunen M, Kataja V, Kosma VM, Mannermaa A. Γ-secretase components as predictors of breast cancer outcome. PLoS One. 2013;8:e79249. doi: 10.1371/journal.pone.0079249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Placanica L, Tarassishin L, Yang G, Peethumnongsin E, Kim SH, Zheng H, Sisodia SS, Li YM. Pen2 and presenilin-1 modulate the dynamic equilibrium of presenilin-1 and presenilin-2 gamma-secretase complexes. J Biol Chem. 2009;284:2967–2977. doi: 10.1074/jbc.M807269200. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.