Abstract
Although invasion reproductive numbers (IRNs) are utilized frequently in continuous-time models with multiple interacting pathogens, they are yet to be explored in discrete-time systems. Here, we extend the concept of IRNs to discrete-time models by showing how to calculate them for a set of two-pathogen SIS models with coinfection. In our exploration, we address how sequencing events impacts the basic reproductive number (BRN) and IRN. As an illustrative example, our models are applied to rhinovirus and respiratory syncytial virus co-circulation. Results show that while the BRN is unaffected by variations in the order of events, the IRN differs. Furthermore, our models predict copersistence of multiple pathogen strains under cross-immunity, which is atypical of analogous continuous-time models. This investigation shows that sequencing events has important consequences for the IRN and can drastically alter competition dynamics.
Keywords: Discrete-time model, Coinfection, Invasion reproductive number, Competitive exclusion
1. Introduction
The basic reproductive number, , is a critical threshold in mathematical epidemiology. This quantity, defined as the “expected number of secondary cases produced, in a completely susceptible population, by a typical infected individual during its entire period of infectiousness” [(Diekmann, Heesterbeek, & Metz, 1990), p. 365], is used to determine whether a disease will persist, with indicating disease persistence. has been studied in numerous continuous and discrete-time epidemic models (e.g. (Allen, 1994; Allen & van den Driessche, 2008; Crawford & Kribs-Zaleta, 2009; Hernandez-Ceron, Feng, & van den Driessche, 2013; Lewis, Rencławowicz, van Den Driessche, & Wonham, 2006; Qiu, Kong, Li, & Martcheva, 2013)) and has proved useful for informing disease control strategies. Although the basic reproductive number (BRN) is invaluable, it has its restrictions. One main restriction is that it can only be used to describe disease spread in a naive population.
As many communities around the globe are endemic for at least one pathogen (e.g. Chagas disease in Latin America, malaria in parts of Africa, and dengue in regions of South America), it becomes necessary to study invasion reproductive numbers (IRNs). An IRN describes the number of secondary infections produced by an infected individual in a population where one (or more) other pathogen is endemic (Porco & Blower, 1998). This quantity displays the same threshold behavior as , namely if the IRN of a pathogen is greater than 1, the pathogen can spread in a population endemic with the other disease(s).
IRNs have been utilized frequently in continuous-time models (e.g. (Crawford & Kribs-Zaleta, 2009; Martcheva & Pilyugin, 2006; Nuno, Chowell, Wang, & Castillo-Chavez, 2007; Qiu et al., 2013; Zhang, Sandland, Feng, Xu, & Minchella, 2007)); however, they are yet to be explored in discrete-time systems. This may be due to the lack of multiple-pathogen discrete-time models in literature. While discrete-time competition models have been studied for over forty years and shown to exhibit both competitive exclusion and coexistence under different conditions (Cushing, Levarge, Chitnis, & Henson, 2004; Edmunds et al., 2003; Franke & Yakubu, 1991; Hassell & Comins, 1976; Smith & Zhao, 2001), the only discrete-time multiple-pathogen models, to the best of our knowledge, are discussed in (Perez-Velazquez, 1999) and (Allen, Kirupaharan, & Wilson, 2004). In the former article, a discrete-time SIS model of two competing pathogen strains with demography and cross-immunity is analyzed. The basic reproductive number of each strain is calculated and stability analyses are conducted for the disease-free and boundary equilibria. The study also asserts that a coexistence equilibrium exists if and only if the basic reproductive numbers of both strains are identical. Since most parameter values used in this model did not satisfy the coexistence condition, the principle of competitive exclusion is supported. Additionally, through numerical simulations, the author in (Perez-Velazquez, 1999) illustrates how demography can result in complex population dynamics such as period doubling and chaos. It is important to highlight that nowhere in the analysis of (Perez-Velazquez, 1999) is the IRN of the two strains mentioned or alluded to.
In (Allen et al., 2004), Allen et al. present single-patch and two-patch discrete-time SIS models for n cocirculating pathogens with complete cross-immunity and no demography. Here too, the authors compute the BRNs and address stability of the disease-free and single-strain endemic equilibria. Their analysis reveals that the principle of competitive exclusion holds in a single patch, where the pathogen with the greatest BRN dominates (given that the BRN is greater than 1). However, coexistence of multiple pathogen strains is possible in two patches and is impacted by both dispersal probabilities and BRNs. Although (Allen et al., 2004) examines disease persistence in a population with multiple pathogen strains, we note that at no point in the study are IRNs mentioned.
The primary purpose of this current work is to extend the derivation of IRNs to discrete-time models, using as illustrative examples a set of two-pathogen SIS models. Unlike (Perez-Velazquez, 1999) and (Allen et al., 2004), the models that we propose assume that coinfection by both pathogen strains is possible. Like (Allen et al., 2004), however, we leave demography implicit (as explained in the next section) because our interest is in developing the concept of IRNs in discrete-time models using the simplest possible multiple-pathogen coinfection scenario. By extending IRNs in this fashion, our research adds to the analytical tools by which discrete-time models can be examined and enriches literature on multiple-pathogen discrete-time systems.
The secondary purpose of this article is to investigate how different assumptions on the ordering of events in discrete-time models impact the BRN and IRN, and ultimately the types of biological conclusions that can be drawn from them. When creating discrete-time models, it is common practice to specify the order of events (e.g. (Bravo de la Parra, Marvá, Sánchez, & Sanz, 2017; Hilker & Liz, 2013; Klepac & Caswell, 2011; Lewis et al., 2006; Perez-Velazquez, 1999; Weide & Hilker, 2019)). This is because different orderings can lead to differing conclusions as described in the West Nile virus study of (Lewis et al., 2006) where changing the order of disease-induced mortality, natural mortality, birth, infection, and transfer resulted in distinct forms of the BRN. Although various sequential formulations of our coinfection model can be derived using different ordering of events, we consider only three sequential models where either (a) all infection events occur before recovery, (b) infection and recovery are intertwined, or (c) recovery precedes infection. These three formulations encompass a wide range of infection/recovery patterns that may arise in nature and thus any conclusions obtained from them can be extended to a plethora of other models that follow a similar pattern.
In addition, we consider a formulation of the coinfection model that assumes that events occur simultaneously, as in continuous-time systems. The model of (Allen et al., 2004) assumes a small enough time step that all event types can occur simultaneously without emptying any of the compartments. Although the structure of the model in (Perez-Velazquez, 1999) is partially ordered, with both infection types occurring simultaneously, a fully simultaneous “ordering” of events, as presented in this work, has not previously been explored in a discrete-time model using arbitrary time step size. While a specific order of events may be necessary when describing certain biological processes (e.g. animal life cycles), there are instances where events in discrete-time models can occur simultaneously (e.g. when describing disease transmission). In such scenarios, a different formulation of discrete-time models is required, namely the simultaneous formulation that we propose.
The present study extends the single-patch model of (Allen et al., 2004) in two ways: by formulating it in terms of an arbitrary time step, following notation such as in (Castillo-Chavez & Yakubu, 2001; Lewis et al., 2006; Smith & Zhao, 2001), and by allowing coinfection. Relaxing the assumption of a small time step requires addressing the order of events, while relaxing the assumption of complete cross-immunity to consider a broad spectrum of coinfection interactions vastly increases the size of the model. Because of the complexity involved in articulating and analyzing the general n-pathogen model, we begin by developing the models and analysis using two pathogens only. The following section develops the different formulations of the two-pathogen discrete-time SIS coinfection model. We derive and compare their respective BRNs and IRNs to develop generalizations about the impact of ordering on these key epidemiological quantities. Next, the models are applied to a specific example of rhinovirus (RV) and respiratory syncytial virus (RSV) co-circulation to provide insight on the relationship between these two pathogens at the population level. Finally, we extend the models and results to the more general case with n pathogens cocirculating.
2. Two-pathogen models
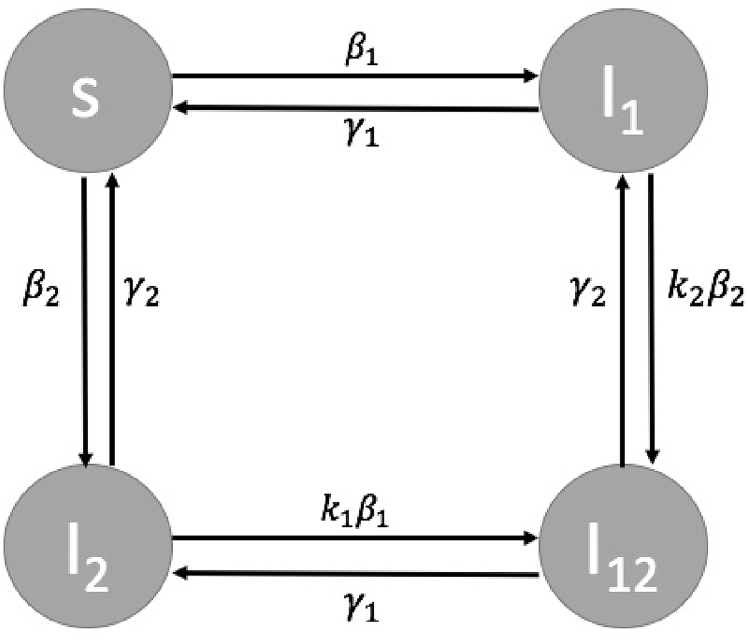
The discrete-time models that we first consider describe the dynamics of two pathogen strains competing for susceptible humans within a population, as shown in Fig. 1. In the models, humans are compartmentalized into four classes: susceptible (S), infected with pathogen 1 (), infected with pathogen 2 (), and infected with both pathogens (). Throughout the following sections, we take the proportion of people infected with pathogen 1 to be , and the proportion of people infected with pathogen 2 to be , where the constant N represents the total human population. The infection and recovery rates of each pathogen are given by and respectively, while the relative likelihood of coinfection with pathogen i given infection with the other pathogen is represented by . We note that, ecologically, inversely measures the level of interspecific competition, with high values indicating low interspecific competition. The parameter represents the proportion of people who do not get infected with pathogen i by a single infective within a certain time interval and , is the proportion that do not recover from pathogen i in that time interval. In our study, we take the time step, , to be 1. Hence, and . The proportion of susceptibles who do not get infected with pathogen i in unit time when proportion of the population is infectious with pathogen i is therefore represented by .
Fig. 1.
SIS coinfection model for two pathogens.
Some authors have studied the interactions of demographic changes and infection dynamics, but our interest is focused on the infection dynamics alone in a population at equilibrium, similarly to Allen et al. (Allen et al., 2004). In a simple SIS monoinfection cycle, recovery and demographic renewal are mathematically equivalent, since (assuming no vertical transmission) both have the effect of replacing infectives with an equivalent number of susceptibles, so that a single event (and associated parameter) can account for both. Here our interest is instead on the interplay among infections, and the impact of order of events on discrete-time reproduction numbers.
In the subsections below, we develop the simultaneous formulation of the discrete-time SIS coinfection model, referred to as SIM, along with the three sequential formulations which we call SEQ1, SEQ2, and SEQ3.
2.1. SIM model formulation ()
The complication in extending the model of (Allen et al., 2004) (which already assumes that infection and recovery occur simultaneously) to allow both arbitrary time steps and coinfection involves the result of coexposure to multiple pathogens during a single time step. Such coexposure should qualitatively reflect the altered susceptibilities caused by a primary exposure, specifically the special cases (no interaction) and (full cross-immunity) of which this model is a generalization. To address this issue, we assume that multiple exposures during a single time step occur in random order with equal frequency, and define some notation. We let and then define
gives the proportion of coexposed individuals first exposed to pathogen who become coinfected with both infections i and j in a single time step. For , we have (unaltered susceptibility means coexposure implies coinfection), while for , (complete cross-immunity means monoinfection only). We also note that as , remains defined. More broadly, when , (reduced susceptibility means not all coexposed become coinfected), while for , (under unreduced susceptibility, all coexposed become coinfected).
For the two-pathogen model, by assumption half of the susceptibles coexposed to both pathogens 1 and 2 in a single time step are first exposed to pathogen 1, and the other half to pathogen 2 first. All coexposed individuals are assumed to become infected by their first exposure, j. Whether they also become infected by their second exposure, i, depends on the altered susceptibility to that second pathogen, as well as on the prevalence of that pathogen, . of those coexposed at time step t first exposed to pathogen 1 will become coinfected; the remaining will become infected only with pathogen 1. Similarly of those coexposed first exposed to pathogen 2 will also develop pathogen 1. Thus overall, of the coexposed (in S), the proportions moving to classes , and , respectively, are , , and . This leads to the following model.
| (1) |
Reading the equations, at time t, a susceptible individual can become infected with either pathogen 1 or pathogen 2. The proportion of people that do not get infected with either pathogen is represented by . These individuals remain in the susceptible class during the next time step and are joined by people in the and classes who recover from infection at time t and do not develop the other infection, and by members of who recover from both infections in the same time step.
The individuals in at time consist of those susceptibles S who developed infection 1 but not infection 2, those in who neither recovered from their infection nor were coinfected at the previous time step (), those in who developed infection 1 and recovered from infection 2, and those coinfected who recovered from infection with pathogen 2 but not from infection with pathogen 1. The population in at each time step is composed in a similar fashion.
Humans in the coinfected class, , who remain in that class at time are those who neither recovered from infection with pathogen 1 nor pathogen 2 during the previous time step (). Adding to this class are those individuals from who developed infection 2 but did not recover from infection 1, and a similar proportion from who developed infection 1 but did not recover from infection 2. In addition, some of the susceptibles who were dually exposed develop both infections.
We note that for the special case , model (1) simplifies to
| (2) |
where represents the class of individuals susceptible to pathogen 1 and are those infected with pathogen 1. A similar reduction of the SIM model can be obtained when . For this case, model (1) simplifies to
| (3) |
where denotes those within the population that are susceptible to pathogen 2 and represents those infected with pathogen 2. Since the total population, N, is constant, (2) and (3) can further be written respectively as
| (4) |
and
| (5) |
2.2. SEQ1 model formulation ()
In the first sequential model formulation of the discrete-time SIS model, we assume that infection with pathogen 1 occurs first and is followed by infection with pathogen 2, then recovery from pathogen 1, and finally recovery from pathogen 2. From this ordering of events, we obtain the following system of difference equations for the SEQ1 model (a step-by-step formulation of this system is provided in Appendix A):
| (6) |
We note that for the special case , model (6) simplifies to
| (7) |
while for it simplifies to
| (8) |
2.3. SEQ2 model formulation ()
The second sequential formulation of the SIS model assumes the following order of events: infection with pathogen 1, recovery from pathogen 1, infection with pathogen 2, and finally recovery from pathogen 2. From this, we obtain the system of difference equations for the SEQ2 model:
| (9) |
For or 2, this model has the same simplifications as SEQ1.
2.4. SEQ3 model formulation ()
In the third sequential model, we assume that recovery from pathogen 1 occurs first and is followed by recovery from pathogen 2, then infection with pathogen 1, and finally infection with pathogen 2. The system of difference equations for SEQ3 is
| (10) |
The SEQ3 model can be simplified for . However, this simplification is different from that of SEQ1 and SEQ2 due to the fact that recovery from each pathogen occurs before infection in SEQ3, while recovery occurs after infection in SEQ1 and SEQ2. For or respectively, the SEQ3 model becomes
| (11) |
or
| (12) |
3. Basic reproductive numbers ()
The BRN of each model is calculated using the next generation matrix approach outlined in (Allen & van den Driessche, 2008). When modeling scenarios with n pathogens, it is common for the BRN to have the form , . While measures the ability of any combination of pathogens to invade a susceptible population, measures the ability of pathogen i to invade it. In the following subsections, we denote the overall BRN of each model by , , where k describes the model at hand (s indicating the simultaneous model and 1, 2, and 3 indicating each of the sequential models). Additionally, we represent each model's pathogen i-only BRN by .
3.1. SIM BRN
Our analysis begins with the calculation of the basic reproductive number of the simultaneous model. Using the next generation matrix approach of (Allen & van den Driessche, 2008), we obtain the vector of new infections that survive the time interval
and the vector of all other transitions
Differentiating these with respect to the infected states (, , ) and evaluating at the unique disease-free equilibrium (N,0,0,0), we have
The basic reproductive number of the SIM model, , is calculated as max max, where is the 3 × 3 identity matrix. In this expression, can be interpreted as the product of the number of infections in one time step (β) by the average duration of infection in time steps ().
3.2. SEQ1 BRN ()
Using the next generation operator method to calculate the BRN of the SEQ1 model (6), our and matrices are
and
Differentiating with respect to the infected classes (, , ) and evaluating at the unique disease-free equilibrium (N,0,0,0), we have
The BRN of the SEQ1 model, , is calculated as
It is clear that . The reason for this is embedded in the order of events in SEQ1. Since recovery follows infection in SEQ1, an infected individual must avoid recovery in order to be counted as infected after each time step. A proportion of infected individuals do not recover, hence the term in . This term is not present in due to the concurrent nature of infection and recovery events in the simultaneous model.
3.3. SEQ2 BRN ()
To calculate the BRN of the SEQ2 model, we use the following and matrices:
and
Differentiating with respect to the infected classes (, , ) and evaluating at the unique disease-free equilibrium (N,0,0,0), we have
The BRN for the SEQ2 model, , is calculated as
Here, the expression matches that of . This is because just as in SEQ1, recovery from a given pathogen follows infection in the SEQ2 model.
3.4. SEQ3 BRN ()
Finally, we compute the BRN for the SEQ3 model. In this case, we use
and
Differentiating with respect to the infected classes (, , ) and evaluating at the unique disease-free equilibrium (N,0,0,0), we have
The BRN of the SEQ3 model, , is calculated as
Note that the BRNs are identical for all three sequential models. This occurs because the SEQ1 and SEQ2 models differ only in coinfection terms (altered susceptibility to a second infection), but BRNs are defined in the context of a single pathogen invading an uninfected population. Meanwhile, the SEQ1 and SEQ3 models describe the same infection cycle (infection, recovery, infection, recovery, etc.). The only difference between them is the point at which the population is observed within that cycle; in SEQ1 it is observed after recovery and in SEQ3 after infection.
3.5. Global stability of disease-free equilibrium
To conduct global stability analysis of the disease-free equilibrium, we illustrate our approach with the SEQ1 model; the same approach can be used for the other models. The proportion of the population involving pathogen i is denoted by , where and . In Theorem 1, we show that the DFE is GAS when for all .
Theorem 1
For the SEQ1 model, if for some i, then . Furthermore, if for all , then the DFE is globally asymptotically stable (GAS).
Proof. From the SEQ1 equations,
If , , so that
Thus, when (which is equivalent to ), . Since the sequence is monotone decreasing and bounded below by 0, .
If , and we have
In this case, when (which is equivalent to ) then and so . Therefore, if , .
For , we have
If , , so that
meaning that when (equivalently, ), . Since the sequence is monotone decreasing and bounded below by 0, .
If , then , , and we have
In this case, when (which is equivalent to ) then and so . Therefore, if , .
Clearly, if for all , then all of the infectious classes go to 0 and the DFE is GAS. ∎
4. Single-strain endemic equilibria
As in most epidemiological models, when , a pathogen can invade a susceptible population. More specifically here, when for pathogen i, an equilibrium exists in which only pathogen i is endemic. This result holds for all four sequences of events explored here (SEQ1, SEQ2, SEQ3, and SIM) as well as for models considering any number of pathogens. For any given pathogen i, if all classes involving infection with a pathogen other than i (including coinfection with pathogen i and some other pathogen) are set to 0, then there remain only two compartments to identify, namely S and , which sum to N. The resulting equilibrium condition is equivalent to that of the corresponding one-pathogen model (equation (13) for the SIM model, (14) for SEQ1 and 2, and (15) for SEQ3).
| (13) |
| (14) |
| (15) |
The existence of a unique single-pathogen endemic equilibrium (EE) for the SIM and SEQ models (which is independent of the number of pathogens involved in the formulations) is described in Theorem 2, and is followed by local stability analysis in Theorem 3.
Theorem 2
For equations (13), (14), (15), if , a single-strain EE exists and is unique.
Proof. We begin by providing a detailed proof for the existence and uniqueness of the single-strain EE for (14) and note that the existence and uniqueness of the single-strain EE for (13) and (15) can be proved in an analogous fashion.
The fixed point condition for (14) is given by
and can be rewritten as
(or else ). Adding to both sides and dividing by N yields
where .
Differentiating with respect to x, we have
The last term, where , is positive since on (0,1) and (note )). Thus, on , meaning that f is monotone increasing within that interval.
Since f is positive and increasing on (0,1) and , has a unique solution in (0,1) if and only if . Thus, a unique pathogen i EE (and hence a unique root of f) exists if and only if , which is equivalent to . ∎
Stability of these EEs involves invasion reproductive numbers, which are developed in section 5. The following theorem makes use of the IRN expressions derived in section 5.2.
Theorem 3
(i) When , the unique EE is LAS in the 1-pathogen system (13), (14), or (15). (ii) When and the IRN of all other pathogens with regard to pathogen i is less than 1, say for , the unique single-strain EE for pathogen i is LAS in the n-pathogen system.
Proof. We begin by providing a detailed proof of part (i) for (14), and note that the proof for (13) and (15) can be obtained in a similar manner.
For (14), let . The EE is LAS iff , where x is the pathogen i EE.
Since we cannot solve explicitly for x in the EE expression (as is transcendental for x), we consider the inverse and view as a function of x. Given , we have
Notice that in the second line of equalities, is less than 1. This forces which implies that since the function is monotone increasing on (0,1). Substituting into we obtain
To prove , we wish to show, using techniques from multivariable calculus, that max on . Notice that the domain does not include the boundary (i.e. or ). This is because implies (as iff ), a contradiction to our assumption that . In addition, implies that (from ), a contradiction to the requirement that .
Note that F has no critical points in . We will prove this by showing that on its domain (since a critical point in the domain must satisfy ). For , we require
If we let , we obtain on and . Hence, on . Likewise, letting , we notice on and , meaning that on the domain. This proves that . We note that, similarly, on the interior of .
Since
and
is a critical point of F. Also, , for , , for , and , for . Thus, max and is attained on the boundary lines and . Since the boundary is not included in our domain, we have . This proves that the unique EE is LAS.
For (ii), we again prove LAS for the EE of (14) in the SEQ1 model with . The proof proceeds similarly for the other models and for . We calculate the Jacobian matrix for the SEQ1 model and substitute the single-strain EE for pathogen 1. The eigenvalues of the resulting block-triangular matrix are 1, , , and . The first eigenvalue, 1, comes from the redundancy of the 4 × 4 Jacobian matrix since N is constant. The second eigenvalue, , is clearly in , and the third eigenvalue is which is proven to be in (0,1) by part (i) of this theorem. We complete the proof of (ii) by showing that the fourth eigenvalue, denoted , is also in .
Since for any value of , it follows that . Likewise, we have through the following sequence of inequalities which hold iff the previous inequality is satisfied:
The eigenvalues obtained after substituting the single-strain EE for pathogen 2 into the Jacobian matrix for SEQ1 are 1, , , and . With these values, the LAS of the pathogen 2 EE follows in an analogous manner as that of the pathogen 1 EE (in this case, proof that the fourth eigenvalue is in (0,1) follows directly from ). ∎
In order to examine the co-persistence of two or more pathogens in an endemic state, we first consider IRNs, which provide threshold conditions for a disease to invade a population endemic with another disease(s).
5. Invasion reproductive numbers ()
To obtain the invasion reproductive number (IRN) in a two-pathogen model when pathogen i is endemic (i.e. ), we reclassify the infectious classes and apply the next generation operator method to the compartments representing invading pathogens. With pathogen 1 resident, for instance, the invading pathogen is taken to be pathogen 2, making the infectious classes and . The IRN for pathogen 2 invading a pathogen-1-endemic population is denoted since the resident pathogen can then be left implicit.
5.1. SIM IRN
As established in the previous section, unique single-strain endemic equilibria exist when the corresponding BRN exceeds 1. Here, to derive we consider the pathogen 1 endemic equilibrium that exists when , so that and are the invading classes. Letting
and
we evaluate the resulting F and T matrices at the unique pathogen 1 endemic equilibrium . These matrices are
where the pathogen 1 prevalence is the unique root of ,
The pathogen 2 IRN for the SIM model is then
In this expression, the BRN, , is multiplied by the weighted-average susceptibility of 1 and , where 1 is weighted by the proportion of the population not infected with pathogen 1 that either remain uninfected in the next time step or, if they become infected with pathogen 1 and are coexposed, develop both infections, and is weighted by those already infected with pathogen 1. We note that when , .
In a similar manner, the pathogen 1 IRN for the SIM model is
where is the unique root of . This expression has an interpretation analogous to that of Furthermore, when , .
5.2. SEQ1 IRN ()
To obtain the IRN of the SEQ1 model when pathogen 1 is endemic (i.e. ), we reclassify the infectious classes and use the next generation operator method. The infectious classes are now taken to be and . Letting
and
we evaluate the resulting F and T matrices at the unique pathogen 1 endemic equilibrium .
where
and
Here, and . Since , the Jury criterion requirement is satisfied. Through the set of inequalities
we see that and thus .
Using the F and T matrices, . In this model, the pathogen 1 prevalence is the unique root of . The IRN when pathogen 2 is endemic is computed in a similar manner as and calculated to be . Here, the pathogen 2 prevalence is the unique solution to . We note that when , and when , .
is a multiple of that is the weighted average of 1 and . The value 1 is weighted by the proportion of the population that started out not infected by pathogen 1 and remained uninfected by pathogen 1 during one time step, while is weighed by the rest of the population. Likewise, is a multiple of that consists of a weighted average of 1 and , with 1 weighted by the proportion of the population that is not infected with pathogen 2 and weighted by the proportion that is infected with pathogen 2.
5.3. SEQ2 IRN ()
We obtain the IRN when pathogen 1 is endemic in a similar manner as above. Here, we let
and
and evaluating the resulting F and T matrices at the unique pathogen 1 endemic equilibrium , we have
where
Since the T matrix presented here is identical to that of Section 5.2, we know that .
With these F and T matrices, we obtain
where is the same as that in SEQ1. The factor which is multiplied by in the expression for is a simplification of
In this expression, the value 1 is weighted by the sum of the proportion of individuals who were initially infected with pathogen 1 but recovered (), the proportion of people initially uninfected with pathogen 1 who became infected with pathogen 1 and recovered from said infection (), and the proportion of people initially uninfected by pathogen 1 who remained uninfected during one time step ). Moreover, is weighted by the sum of the proportion of individuals who were infected with pathogen one at the beginning of the time step and did not recover (), and the proportion of individuals initially uninfected with pathogen 1 who became infected and did not recover within the time period ().
The IRN when pathogen 2 is endemic is calculated to be
where is the same as that in SEQ1. This is identical to and results from the fact that infection with pathogen 1 occurs first in both SEQ1 and SEQ2. Therefore, the point at which we count the number of secondary pathogen 1 infections is identical for both models and occurs when the population is divided into only two categories: those infected with pathogen 2 and those not infected with pathogen 2. In addition, when , and when , .
5.4. SEQ3 IRN ()
To calculate the IRN of the SEQ3 model when pathogen 1 is endemic, we use
and
Evaluating the resulting F and T matrices at the unique pathogen 1 endemic equilibrium we have
where
Here, and . To prove that , we use the approach described in Section 5.2, but replace x (in the determinant and trace expressions of Section 5.2) with .
With these F and T matrices,
where is the unique solution to . The factor by which is multiplied in the expression for is a simplification of
In this expression, 1 is weighted by the sum of the proportion of people originally infected with pathogen 1 who recovered and were not reinfected with the pathogen and the proportion of people who were originally uninfected with pathogen 1 and remained uninfected during one time step. The parameter is weighted by the sum of the proportion of people originally infected with pathogen 1 who did not recover, the proportion of people originally infected with pathogen 1 who recovered and were re-infected, and the proportion of people originally uninfected with pathogen 1 but who got infected with the pathogen during one time step. Infections are transmitted by the proportion of people who do not recover from infection with pathogen 1.
The IRN when pathogen 2 is endemic is , where is the unique root of . is multiplied by a weighted average of 1 and , with weighted by the proportion of people originally infected with pathogen 2 who did not recover in one time step and 1 is weighted by the rest of the population. This expression is similar to that of and except for the fact that is multiplied by in ( represents the proportion of people who do not recover from infection with pathogen 2). The inclusion of the factor is due to recovery occurring first in the SEQ3 model, hence an individual must fail to recover from infection with pathogen 2 in order to be counted in the IRN. Although the IRNs for the SEQ1 and SEQ3 models appear to be different, they are in fact identical as detailed in Appendix C. Lastly, notice that when , and when , .
5.5. Dual-endemic equilibrium
It is straightforward to verify numerically that when both IRNs for a given model exceed 1, a unique dual-endemic equilibrium exists and is globally asymptotically stable.
6. BRN/IRN threshold comparisons ()
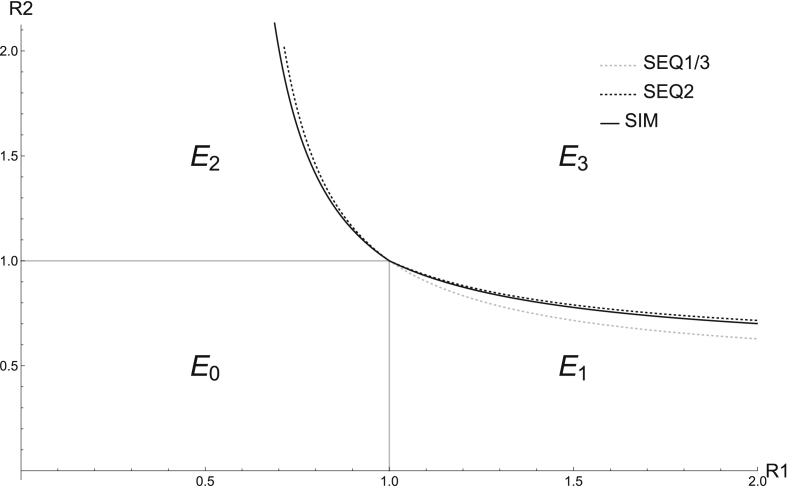
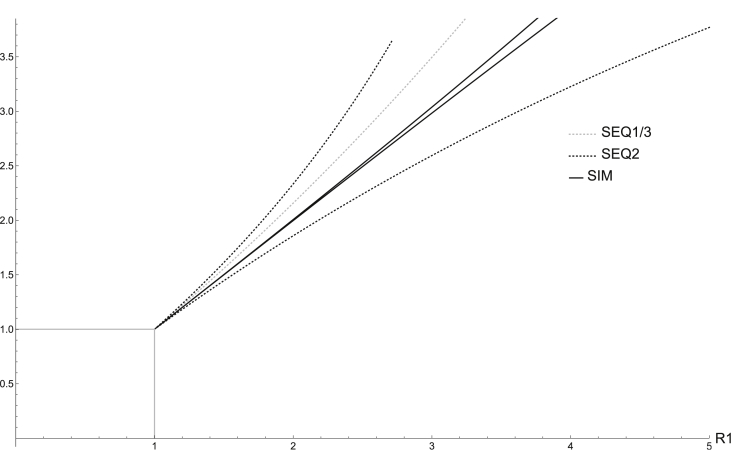
Through the above computations, it is evident that there is great variation in the IRN expressions for the SIM, SEQ1, SEQ2, and SEQ3 models. However, what remains unclear is how the different models' IRN thresholds compare (i.e. whether one model's threshold is more or less restrictive than another). To better analyze the IRNs, we create IRN threshold curves for when and for the various models equal 1 and begin by plotting these curves on the vs. axes in Mathematica. The vs. plane has been utilized in previous studies on co-circulating pathogens (e.g. (Crawford & Kribs-Zaleta, 2009; Kribs & Mitchell, 2015; Martcheva & Pilyugin, 2006)) and allows for a primarily epidemiological comparison of our different models. As seen in Fig. 2, Fig. 3, the threshold curves divide the plane into four distinct regions of possible model outcomes (extinction of both pathogens, , persistence of only pathogen 1, , persistence of only pathogen 2, , and co-persistence of both pathogens, ).
Fig. 2.
BRN/IRN Threshold Curves. In this graph, and . In region , we see extinction of both pathogens, in , the persistence of only pathogen 1, in , the persistence of only pathogen 2, and in , co-persistence of both pathogens.
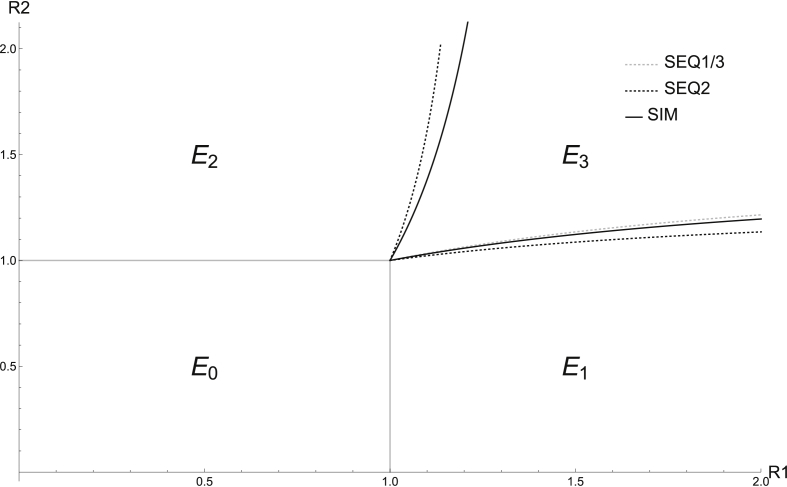
Fig. 3.
BRN/IRN Threshold Curves for. In this graph, and . In region , we see extinction of both pathogens, in , the persistence of only pathogen 1, in , the persistence of only pathogen 2, and in , co-persistence of both pathogens.
As an example of how these curves were generated, consider the curve. We fixed the and parameters while varying . For each value of , we solved for the pathogen 2 endemic equilibrium, and used this equilibrium value to numerically solve for in the equation . With this pair of and values, we calculated and for the respective models. Generating the curves was done in a similar manner while varying and solving for the pathogen 1 endemic equilibrium. To minimize the effect of parameter values on the threshold graphs, we keep and in Fig. 2, Fig. 3. We find that while the SEQ2 model predicts a larger region of co-persistence of both pathogens than the other models for , it predicts a smaller region of co-persistence for .
While the SEQ1 and SEQ3 IRN curves and the SEQ1 and SEQ2 curves will always overlap due to the models' identical and/or expressions, in general, the ordering of the curves in Fig. 2, Fig. 3 is not always consistent. For example, since if and only if (meaning that it is easier for strain two to survive in SEQ2 than SEQ1 for ), and vice versa for , we find that the SEQ1 threshold curve will be above that of SEQ2 for , with the ordering switched for (Appendix B). In addition, the position of the SIM model curves relative to other curves varies with parameter values. For example, if , we have , but for a different set of values, namely, , we have .
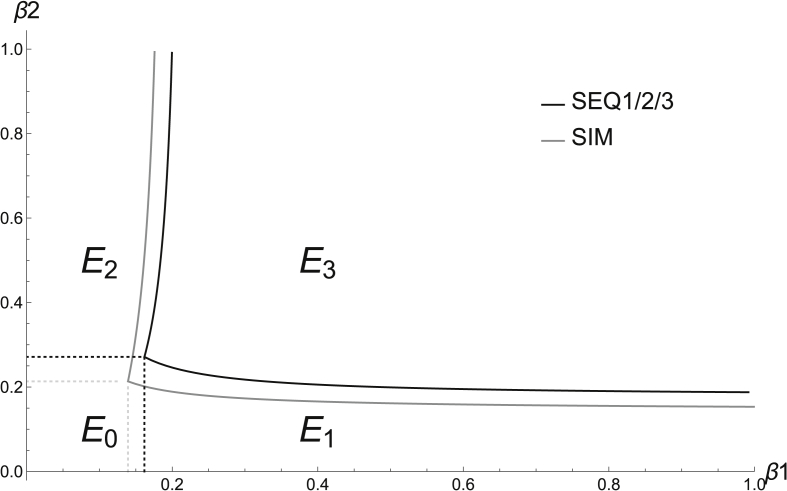
In addition to focusing on the epidemiological variation between the models in the vs. plane, we can observe how the behavior of each model changes with respect to raw parameter values such as and . Since the various models have different BRN's, viewing the IRN threshold curves on a vs. axis allows for the visualization of an additional region where certain models predict disease extinction while others do not. This is the case depicted in Fig. 4. The region between the gray and black boxes illustrates and values for which the sequential models predict disease extinction but the SIM model does not. In fact, parts of that space represent the , , and regions predicted by the SIM models. In addition, the L shaped region created by the two sets of and curves illustrates a space where the SIM model predicts co-persistence of both pathogens but the sequential models do not. The region of co-persistence predicted by the SIM model in this case is larger than that of the sequential models. This results from the fact that recovery occurs after infection in the sequential models. Infected individuals must avoid recovery in order to be counted in the BRNs of the sequential models, which reduces the BRNs and thus makes it less likely to predict persistence of the pathogens.
Fig. 4.
BRN/IRN Threshold Curvesvs.. In this graph, , , and . In region , we see extinction of both pathogens, in , the persistence of only pathogen 1, in , the persistence of only pathogen 2, and in , co-persistence of both pathogens. The dotted gray box delineates the region for the SIM model while the dotted black box delineates the region for the SEQ1, SEQ2, and SEQ3 models.
Through numerical simulations, we are able to investigate how the different model BRN/IRN threshold curves behave as the likelihood of coinfection approaches zero or infinity. For all of the discrete-time models, we find that the region of co-persistence of the two pathogens widens as increases, but becomes smaller as decreases (as illustrated in Fig. 2, Fig. 3). As the likelihood of coinfection increases, the interspecific competition between the two pathogens decreases. This causes the relationship between the pathogens to become more and more mutualistic, resulting in both strains aiding each other to become established within the community. Unlike autonomous continuous-time models, when (i.e. when the two strains exhibit cross-immunity and interspecific competition is at its maximum), we see that for a wide range of parameter values (one of which is shown in Fig. 5), all of our discrete-time models predict a region of co-persistence of both pathogens.
Fig. 5.
Coexistence with Complete Cross-Immunity. The parameter values used to generate this figure are . With these parameter values, we witness the possibility of coexistence of the two pathogen strains in the SEQ and SIM models.
To further explore the competition dynamics between two pathogen strains in our SEQ and SIM models under the assumption of complete cross-immunity, we observe what happens in the limit as approaches zero (recall that in formulating the discrete-time models, we assume ). In this scenario, we find that all of the models have the same BRN and IRNs. The BRN in this case is max max , the pathogen 1 IRN () is , and pathogen 2 IRN () is . Notice that the BRN and IRN expressions are similar to what one expects to find in a continuous-time formulation of the coinfection models. Co-persistence of the two pathogen strains is possible if , , , and are greater than 1. As shown in Theorem 4, it is impossible for both IRNs to be greater than 1. This means that in the limit as T approaches 0, competitive exclusion occurs if and are greater than 1.
Theorem 4
and cannot both be greater than 1 in the limit as T approaches 0.
Proof. Notice that , and . Consequently . Since they are reciprocals of each other, they cannot both exceed 1. ∎
7. RV and RSV coinfection
We now apply the above models to describe RV and RSV circulation within a human population. RV and RSV are two of the main causes of common respiratory tract infections such as pneumonia, bronchiolitis, and the common cold. Since there are currently no vaccines for these viruses, reinfection by either pathogen, and even co-infection by both pathogens have been reported (Greer et al., 2009; Hogan, Glass, Moore, & Anderssen, 2016; Zlateva et al., 2014). During co-infections, it is believed that RSV and RV behave in an antagonistic manner. This is supported by numerous studies which show that infection by one virus is associated with a reduced likelihood of infection by the other virus (Greer et al., 2009; Karppinen, Toivonen, Schuez-Havupalo, Waris, & Peltola, 2016; Martin, Fairchok, Stednick, Kuypers, & Englund, 2013).
Most previous RV/RSV mathematical models have examined these viruses individually (e.g. (Adler & Kim, 2013; Hogan et al., 2016; Pinky & Dobrovolny, 2016)). The only model to consider RV and RSV coinfection (Pinky & Dobrovolny, 2016) focuses on the within-host competition dynamics of the two viruses. There is currently no model that considers RV and RSV co-circulation at the population level. In addition, although RV and RSV hospital case reports are given in daily, weekly, or monthly intervals as mentioned in (Eggo, Scott, Galvani, & Meyers, 2016; Leecaster et al., 2011; Weber, Weber, & Milligan, 2001), none of the existing models are situated in discrete time. Given the available viral infection data sets, it is natural (and one may even argue more appropriate) to consider a discrete-time model of RV and RSV as we do here.
In all of our models, pathogen 1 is taken to be RSV and pathogen 2 is RV. Most of the parameter values used in the models were obtained from previously published studies and are shown in Table 3. The transmission rate for RV is the average of the estimated child to child, adult to child, and adult to adult transmission rates found in (Eggo et al., 2016). Due to the inhibitory relationship between the two viruses, we take .
Table 2.
Pathogen 1 & 2 Endemic Equilibria. In this table, is the proportion of people infected with pathogen 1 at the pathogen 1 endemic equilibrium and is the proportion of people infected with pathogen 2 at the pathogen 2 endemic equilibrium.
| Endemic Equilibria | SEQ1, SEQ2 | SEQ3 | SIM |
|---|---|---|---|
| is unique root of | |||
| is unique root of |
Table 3.
RV & RSV Parameter Values. RSV is taken to be pathogen 1 in the discrete-time models and RV is pathogen 2.
| Symbol | Value/Range | Source |
|---|---|---|
| 0.28 days−1 | Moore, Jacoby, Hogan, Blyth, and Mercer (2014) | |
| 0.36 days−1 | Eggo et al. (2016) | |
| 0.1 days−1 | Weber et al. (2001) | |
| 0.14 days−1 | Gwaltney, Hendley, Simon, and Jordan (1967) | |
| , | () | Assumed |
To illustrate the kind of variation in results that may occur when using the discrete-time models to describe RV and RSV spread, we take . The BRNs and IRNs obtained for this particular choice of are in Table 4. These values place us within the E1 region of SEQ1 and SEQ3 and the E3 region of the SEQ2 and SIM models. While all of the models predict that RV can spread in a completely susceptible population, only the SEQ2 and SIM models predict that the virus will spread in a population endemic with RSV. On the other hand, the SEQ1 and SEQ3 models illustrate the unusual phenomenon that the presence of one virus (RSV) actually protects against invasion by another (RV), a phenomenon also depicted in (Crawford & Kribs-Zaleta, 2009) for two co-circulating HPV strains. Switching the order of RSV and RV, namely letting RSV be pathogen 2 and RV be pathogen 1 while keeping , shows that our discrete-time systems are sensitive to the order of each pathogen. In this scenario (not depicted here), the IRN for both pathogens is greater than 1 in all of the models. Conflicting results such as these show how model assumptions about the order of disease infection and recovery events impact epidemiological conclusions.
Table 4.
BRNs and IRNs for RV (pathogen 1) and RSV (pathogen 2). Calculations use values from Table 4 and .
| Model | ||||
|---|---|---|---|---|
| SEQ1 | 2.66 | 2.40 | 1.29 | 0.91 |
| SEQ2 | 2.66 | 2.40 | 1.29 | 1.05 |
| SEQ3 | 2.66 | 2.40 | 1.29 | 0.91 |
| SIM | 2.94 | 2.76 | 1.16 | 1.03 |
8. Multiple pathogens ()
Writing and analyzing models for more than two (say n) cocirculating pathogens becomes complex rapidly because of the number of classes () and reproduction numbers () such models have. As discussed in (Mitchell & Kribs, 2019), even the notation for such models becomes complicated. In general, coinfection classes exist for any subset A of pathogens. Assuming that any such subset A of pathogens can be considered resident, the system then has an overall invasion reproductive number measuring the ability of any pathogen not in A to invade, as well as more specific IRNs measuring the ability of a specific pathogen i not in A to invade. We therefore first consider a three-pathogen model, in order to illustrate the changes from two-pathogen models. Following the approach of earlier sections, we use the SEQ1 formulation (all infection events, in numerical order, followed by all recovery events, in numerical order) to demonstrate results, but similar results can be shown for the other orders of events.
8.1. Three-pathogen SEQ1 model
We can write a discrete-time three-pathogen model using the SEQ1 event sequence (infection with pathogens 1, 2, and 3, followed by recovery from pathogens 1, 2, and 3) in matrix form as if we define the following notation:
-
•
Let the vector of state variables be
-
•
Let the prevalences for each pathogen be , , .
-
•
Let be the factor of increased vulnerability to infection with pathogen i of individuals already infected with at least one other pathogen.
-
•
Denote the proportion of individuals already infected with some pathogen (not i) who become, and remain, infected with pathogen i (for at least 1 time step) as Denote the remainder of such individuals, who do not acquire and retain infection i from time t to time , as .
Finally, then, the matrix M is given by
where , , , , , , and .
In the special case , this system reduces to the single-pathogen model (14) for transmission of pathogen i where and (e.g., equation (7) for ), or to the two-pathogen SEQ1 model which excludes pathogen i.
The basic reproductive number can be calculated using next-generation matrix methods. The key matrices here are
where is the identity matrix, (), and thus .
The proof that (Theorem 1) for , as is identical to that for the two-pathogen model. Therefore the disease-free equilibrium is globally asymptotically stable when . Similarly, the existence of three single-pathogen endemic equilibria (each unique for the respective pathogen) when the respective (Theorem 2) follows the proof for one or two pathogens. Also as before, the conditions for local stability of these equilibria (Theorem 3) involve the system's overall IRN with respect to pathogen i being less than 1, but here that IRN takes a more complicated form.
To study coinfection one computes the IRNs using appropriately adjusted next-generation methods. For instance, to compute , the overall IRN when [only] pathogen 1 is endemic, we consider the reduced state vector representing the invading pathogens, equal to zero at the pathogen-1 equilibrium. Now the matrices are
where , , . The overall IRN with pathogen 1 resident is thus , where
That is, the IRN for pathogen 2 or 3 invading when pathogen 1 is resident is the given pathogen's BRN multiplied by the average susceptibility of that population to the invading infection. The term in brackets is a weighted average of a factor of 1 (for those of the population who avoid infection with pathogen 1) and a factor of (for the rest of the population, who do become infected with pathogen 1).
The overall IRN of the system when pathogen 2 or 3 is resident has the same structure but not an identical formulation, given the way that order of infection affects susceptibility.
In the case where two of the pathogens are already resident in the population—say, pathogens 1 and 2, with , , and (so that such a dual-endemic equilibrium exists)—the system's overall IRN is the same as the IRN for the remaining pathogen, e.g., . In this case, only the four compartments involving pathogen 3 count as infected, leading to the next-generation matrices
where
and thus we have, finally,
Again the last factor is a weighted average susceptibility at the equilibrium where pathogens 1 and 2 are endemic.
The other two IRNs involving two resident pathogens have the same structure but, again, not identical forms due to the effects of the ordering of events. Numerical analysis appears to indicate that when all three such dual-resident IRNs exceed 1, a triple-endemic equilibrium exists and is globally stable.
8.2. n-pathogen model
To extend the three-pathogen model to n pathogens, one must define state variables (compartments), of them involving i-fold coinfection (). Following the pattern for and , one can see that the derivation of remains straightforward, with all relevant matrices upper triangular, and the next-generation matrix's eigenvalues along the diagonals, with once again, . Again the disease-free equilibrium is globally asymptotically stable for , while n single-pathogen endemic equilibria (each unique for the given pathogen) exist iff the corresponding , and are LAS iff and . Multi-endemic equilibria involving a subset A of m of the n pathogens exist when the m different IRNs, each involving one pathogen in A invading the equilibrium at which the other pathogens in A are endemic, all exceed 1. These A-endemic equilibria are stable when other pathogens not in A cannot invade them.
9. Discussion & concluding remarks
Discrete-time systems are sensitive to ordering and are more prone to chaotic and other complex behavior than continuous-time models, but may nevertheless be more appropriate to describe scenarios where the biology of a system demands discrete representation. However, it is important to note that the conclusions obtained from these models depend on underlying assumptions about the sequencing of events. In this paper, we develop multiple formulations of a novel two-pathogen discrete-time coinfection model and provide a detailed analysis of how differences in the ordering of infection and recovery events impact two critical quantities, the BRN and IRN, for each pathogen. Our work is the first to extend the derivation of IRNs to discrete-time systems and to introduce discrete-time coinfection models with arbitrary time steps. To the authors' knowledge, in extending the model to pathogens, this is also the first paper to derive IRNs where multiple pathogens are assumed resident.
Our results show that whether events in a discrete-time model are sequenced or not impacts the BRN. Due to the order of events in the sequential models, a term appears in their BRNs. This occurs because an individual must fail to recover during a time step in order to be counted in the BRN. Since events occur simultaneously in the SIM model, an infected individual does not need to evade recovery in order to be counted in the BRN, hence the lack of a term in the SIM model's BRN expression. This difference in BRNs makes it possible to find parameter values such that the sequential models predict disease extinction () while the SIM model predicts disease invasion (), a contradiction that can be especially alarming from a public health standpoint.
One peculiar finding of this research is the potential for the SEQ and SIM models to predict a region of coexistence for the two pathogen strains under the assumption of complete cross-immunity. This result differs from the competitive exclusion principle that is typical of autonomous continuous-time models (e.g. (Pelosse & Kribs-Zaleta, 2012)) and that is observed in the partially ordered model of (Perez-Velazquez, 1999), where only infection events occur simultaneously. The fact that competitive exclusion is seen in the Perez-Velazquez model but not in any of our models (when cross-immunity is considered) highlights the significance of ordering assumptions in discrete-time models. It is essential for researchers to be mindful of such assumptions when formulating their models, as differences in formulation can drastically impact model predictions. For example, assuming cross-immunity, the partially ordered model in (Perez-Velazquez, 1999) suggests that control strategies should target the pathogen that is more likely to spread, while the SEQ and SIM models presented here suggest that control aimed at both pathogens may be needed.
As shown in Theorem 4, the apparent contradiction in competition dynamics between our discrete-time coinfection models and an analogous continuous-time version (both with complete cross-immunity) is resolved in the limit as approaches 0. Under this scenario, all of the discrete-time models predict competitive exclusion and have BRN and IRN expressions similar to those of a continuous-time formulation of the coinfection model. This result shows that it becomes more difficult for both IRNs to exceed 1 for small time steps, and that the region of co-persistence in the vs. graphs (e.g. Fig. 2, Fig. 3) constricts for small T. Furthermore, the result illustrates that time steps are crucial in the study of discrete-time systems since different time steps can produce fundamentally different model conclusions.
Given that events in the coinfection model are ordered, our work illustrates that ordering affects the IRN but not the BRN of the two pathogens. As seen in Table 1, the IRN (especially the pathogen 2 IRN) is more sensitive to assumptions on the sequence of events than the BRN. This is because many factors are involved in calculating the IRN that are not needed for the BRN, factors such as whether an individual was infected with the endemic strain at the beginning of a time step, whether he or she recovered from the endemic strain, became infected with the invading strain, etc. It is common knowledge in discrete-time mathematical modeling that ordering of events can affect important epidemiological quantities such as the BRN (Lewis et al., 2006). However, since IRNs have not been studied in discrete time until now, this research shows that ordering has implications of which mathematical biologists working with discrete-time models may not previously have been aware.
Table 1.
Summary of BRNs and IRNs for two-pathogen models.
| Model | BRN | Pathogen 1 IRN | Pathogen 2 IRN |
|---|---|---|---|
| SEQ1 | max | ||
| SEQ2 | same as SEQ1 | same as SEQ1 | |
| SEQ3 | same as SEQ1 | same as SEQ1 | same as SEQ1 |
| SIM | max |
Results obtained from our various model formulations are analogous to the findings of (Hilker & Liz, 2013) and (Weide & Hilker, 2019) which are situated in the context of population ecology. In (Hilker & Liz, 2013), Hilker and Liz investigate whether hydra effect, the unusual increase in a species' population size in response to an increase in its mortality rate, is impacted by the timing of harvesting and reproduction in discrete-time models of standard harvesting strategies. Through rigorous mathematical proofs, they find no qualitative distinctions between two models that differ in the ordering of harvesting and reproduction because the models essentially describe the same process (harvesting, reproduction, harvesting, reproduction, etc.). This result is further echoed in the predator-prey models of (Weide & Hilker, 2019), where switching the order of density-dependent prey regulation and predation results in identical qualitative conclusions. In our study, we also find that when the general order of events is preserved between models, as seen in SEQ1 and SEQ3, which simply take census at different points in the process, there is no difference in the BRN or IRN of the models. However, differences become apparent when we consider the SEQ2 model which describes a completely different order than SEQ1 and SEQ3. This illustrates that the order of events in discrete-time models matters when more than two interacting events are involved, as altering the order of a subset of events can result in distinct disease cycles.
Our application of the different formulations of the discrete-time coinfection model to RSV and RV co-circulation provides a concrete illustration of the wide array of results that can be obtained from these models. Under certain assumptions, only the SEQ1 and SEQ3 models show the potential for the presence of RSV to protect a population from invasion of RV. This conclusion, however, changes with the order of each pathogen, indicating that the protective ability of RSV is only observed when it is given the advantage of infecting the population first since infection with pathogen 2 is then applied to a less receptive population. This conclusion stresses the importance of the biological assumptions embedded in mathematical models.
While this current work provides valuable insight into the dynamics of various formulations of discrete-time coinfection models, we acknowledge that the model that we propose is simplistic in nature. In reality, the dynamics of co-circulating pathogen strains can lead to complex mathematical systems, some examples of which are described in (Crawford & Kribs-Zaleta, 2009; Kribs-Zaleta & Mubayi, 2012; Qiu et al., 2013). In our future work, we will extend this discrete-time coinfection model to incorporate various transmission routes, such as vector-borne transmission, so that it can be applicable to a wider range of diseases. We hope that this initial exploration of IRNs in discrete-time systems, and the simultaneous formulation of such systems, will serve as a catalyst for other researchers to contribute to the study of multiple-pathogen discrete-time models, paying careful attention to how model assumptions on the order of events (or the lack thereof) affect key epidemiological conclusions.
Acknowledgements
The authors thank Dr. Pauline van den Driessche for valuable feedback on initial work developing the models and calculating reproductive numbers, Dr. Frank Hilker for suggesting relevant articles and providing a preprint of (Weide & Hilker, 2019), and an anonymous reviewer for suggestions which improved the paper. Furthermore, the authors acknowledge the National Science Foundation as this material is based upon work supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. 1746052. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation.
Handling Editor: J. Wu
Footnotes
Peer review under responsibility of KeAi Communications Co., Ltd.
Appendix A. Formulation of SEQ1 Model
For the SEQ1 model, let , , , and represent the time when infection with pathogen 1, infection with pathogen 2, recovery from pathogen 1, and recovery from pathogen 2 occur respectively, with . In addition, let . The system of difference equations obtained after infection with pathogen 1 is
After infection with pathogen 2 we have,
After recovery from pathogen 1, we obtain
After recovery from pathogen 2, we have
Since , the system of equations for the SEQ1 model can be written as
| (16) |
The systems of difference equations for the SEQ2 and SEQ3 models can be obtained step-by-step in a similar fashion.
Appendix B. SEQ1 & SEQ2 Threshold Curve
To prove that iff (if and only if) , notice the following sequence of inequalities which follow iff the previous inequality is satisfied:
A similar proof can be given to show iff .
Appendix C. SEQ1 & SEQ3 IRN Expressions
Since , we will prove that the SEQ1 and SEQ3 equations are identical by showing that , where is the unique root of and is the unique root of (uniqueness of these roots is discussed in Appendix B).
To see this, note that
This shows that is a root of . Since is the unique root of , we have .
An analogous proof showing that , where is the unique root of and is the unique root of , can be used to show that the two models' equations are identical.
References
- Adler F.R., Kim P.S. Models of contrasting strategies of rhinovirus immune manipulation. Journal of Theoretical Biology. 2013;327:1–10. doi: 10.1016/j.jtbi.2013.02.010. [DOI] [PubMed] [Google Scholar]
- Allen L.J. Some discrete-time SI, SIR, and SIS epidemic models. Mathematical Biosciences. 1994;124(1):83–105. doi: 10.1016/0025-5564(94)90025-6. [DOI] [PubMed] [Google Scholar]
- Allen L.J., Kirupaharan N., Wilson S.M. SIS epidemic models with multiple pathogen strains. Journal of Difference Equations and Applications. 2004;10(1):53–75. [Google Scholar]
- Allen L.J., van den Driessche P. The basic reproduction number in some discrete-time epidemic models. Journal of Difference Equations and Applications. 2008;14(10–11):1127–1147. [Google Scholar]
- Bravo de la Parra R., Marvá M., Sánchez E., Sanz L. Discrete models of disease and competition. Discrete Dynamics in Nature and Society. 2017;2017 [Google Scholar]
- Castillo-Chavez C., Yakubu A.-A. Dispersal, disease and life-history evolution. Mathematical Biosciences. 2001;173(1):35–53. doi: 10.1016/s0025-5564(01)00065-7. [DOI] [PubMed] [Google Scholar]
- Crawford B., Kribs-Zaleta C.M. The impact of vaccination and coinfection on HPV and cervical cancer. Discrete and Continuous Dynamical Systems - Series B. 2009;12(2):279–304. [Google Scholar]
- Cushing J.M., Levarge S., Chitnis N., Henson S.M. Some discrete competition models and the competitive exclusion principle. Journal of Difference Equations and Applications. 2004;10(13–15):1139–1151. [Google Scholar]
- Diekmann O., Heesterbeek J.A.P., Metz J.A. On the definition and the computation of the basic reproduction ratio R0 in models for infectious diseases in heterogeneous populations. Journal of Mathematical Biology. 1990;28(4):365–382. doi: 10.1007/BF00178324. [DOI] [PubMed] [Google Scholar]
- Edmunds J., Cushing J.M., Costantino R.F., Henson S.M., Dennis B., Desharnais R.A. Park's Tribolium competition experiments: A non-equilibrium species coexistence hypothesis. Journal of Animal Ecology. 2003;72(5):703–712. [Google Scholar]
- Eggo R.M., Scott J.G., Galvani A.P., Meyers L.A. Respiratory virus transmission dynamics determine timing of asthma exacerbation peaks: Evidence from a population-level model. Proceedings of the National Academy of Sciences. 2016:201518677. doi: 10.1073/pnas.1518677113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franke J.E., Yakubu A.-A. Mutual exclusion versus coexistence for discrete competitive systems. Journal of Mathematical Biology. 1991;30(2):161–168. [Google Scholar]
- Greer R.M., McErlean P., Arden K.E., Faux C.E., Nitsche A., Lambert S.B. Do rhinoviruses reduce the probability of viral co-detection during acute respiratory tract infections? Journal of Clinical Virology. 2009;45(1):10–15. doi: 10.1016/j.jcv.2009.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gwaltney J.M., Hendley J.O., Simon G., Jordan W.S. Rhinovirus infections in an industrial population: Ii. characteristics of illness and antibody response. Journal of the American Medical Association. 1967;202(6):494–500. [PubMed] [Google Scholar]
- Hassell M.P., Comins H.N. Discrete time models for two-species competition. Theoretical Population Biology. 1976;9(2):202–221. doi: 10.1016/0040-5809(76)90045-9. [DOI] [PubMed] [Google Scholar]
- Hernandez-Ceron N., Feng Z., van den Driessche P. Reproduction numbers for discrete-time epidemic models with arbitrary stage distributions. Journal of Difference Equations and Applications. 2013;19(10):1671–1693. [Google Scholar]
- Hilker F.M., Liz E. Harvesting, census timing and “hidden” hydra effects. Ecological Complexity. 2013;14:95–107. [Google Scholar]
- Hogan A.B., Glass K., Moore H.C., Anderssen R.S. Exploring the dynamics of respiratory syncytial virus (RSV) transmission in children. Theoretical Population Biology. 2016;110:78–85. doi: 10.1016/j.tpb.2016.04.003. [DOI] [PubMed] [Google Scholar]
- Karppinen S., Toivonen L., Schuez-Havupalo L., Waris M., Peltola V. Interference between respiratory syncytial virus and rhinovirus in respiratory tract infections in children. Clinical Microbiology and Infections. 2016;22(2) doi: 10.1016/j.cmi.2015.10.002. 208–e1. [DOI] [PubMed] [Google Scholar]
- Klepac P., Caswell H. The stage-structured epidemic: Linking disease and demography with a multi-state matrix approach model. Theoretical Ecology. 2011;4(3):301–319. [Google Scholar]
- Kribs-Zaleta C.M., Mubayi A. The role of adaptations in two-strain competition for sylvatic Trypanosoma cruzi transmission. Journal of Biological Dynamics. 2012;6(2):813–835. doi: 10.1080/17513758.2012.710339. [DOI] [PubMed] [Google Scholar]
- Kribs C.M., Mitchell C. Host switching vs. host sharing in overlapping sylvatic Trypanosoma cruzi transmission cycles. Journal of Biological Dynamics. 2015;9(1):247–277. doi: 10.1080/17513758.2015.1075611. [DOI] [PubMed] [Google Scholar]
- Leecaster M., Gesteland P., Greene T., Walton N., Gundlapalli A., Rolfs R. Modeling the variations in pediatric respiratory syncytial virus seasonal epidemics. BMC Infectious Diseases. 2011;11(1):105. doi: 10.1186/1471-2334-11-105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis M.A., Rencławowicz J., van Den Driessche P., Wonham M. A comparison of continuous and discrete-time West Nile virus models. Bulletin of Mathematical Biology. 2006;68(3):491–509. doi: 10.1007/s11538-005-9039-7. [DOI] [PubMed] [Google Scholar]
- Martcheva M., Pilyugin S.S. The role of coinfection in multi disease dynamics. SIAM Journal on Applied Mathematics. 2006;66(3):843–872. [Google Scholar]
- Martin E.T., Fairchok M.P., Stednick Z.J., Kuypers J., Englund J.A. Epidemiology of multiple respiratory viruses in childcare attendees. The Journal of Infectious Diseases. 2013;207(6):982–989. doi: 10.1093/infdis/jis934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchell C., Kribs C. Invasion reproductive numbers for periodic epidemic models. Infectious Disease Modelling. 2019 doi: 10.1016/j.idm.2019.04.002. (in press) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore H.C., Jacoby P., Hogan A.B., Blyth C.C., Mercer G.N. Modelling the seasonal epidemics of respiratory syncytial virus in young children. PLoS One. 2014;9(6) doi: 10.1371/journal.pone.0100422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nuno M., Chowell G., Wang X., Castillo-Chavez C. On the role of cross-immunity and vaccines on the survival of less fit flu-strains. Theoretical Population Biology. 2007;71(1):20–29. doi: 10.1016/j.tpb.2006.07.002. [DOI] [PubMed] [Google Scholar]
- Pelosse P., Kribs-Zaleta C.M. The role of the ratio of vector and host densities in the evolution of transmission modes in vector-borne diseases. the example of sylvatic Trypanosoma cruzi. Journal of Theoretical Biology. 2012;312:133–142. doi: 10.1016/j.jtbi.2012.07.028. [DOI] [PubMed] [Google Scholar]
- Perez-Velazquez J. Biometric Department, MTBI Cornell University; 1999. Competitive exclusion in a discrete-time epidemic model with two competing strains. Technical Report BU-1514-M. [Google Scholar]
- Pinky L., Dobrovolny H.M. Coinfections of the respiratory tract: Viral competition for resources. PLoS One. 2016;11(5) doi: 10.1371/journal.pone.0155589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porco T.C., Blower S.M. Designing HIV vaccination policies: Subtypes and cross-immunity. Interfaces. 1998;28(3):167–190. [Google Scholar]
- Qiu Z., Kong Q., Li X., Martcheva M. The vector–host epidemic model with multiple strains in a patchy environment. Journal of Mathematical Analysis and Applications. 2013;405(1):12–36. [Google Scholar]
- Smith H.L., Zhao X.-Q. Competitive exclusion in a discrete-time, size-structured chemostat model. Discrete and Continuous Dynamical Systems - Series B. 2001;1(2):183–191. [Google Scholar]
- Weber A., Weber M., Milligan P. Modeling epidemics caused by respiratory syncytial virus (RSV) Mathematical Biosciences. 2001;172(2):95–113. doi: 10.1016/s0025-5564(01)00066-9. [DOI] [PubMed] [Google Scholar]
- Weide V., Varriale M.C., Hilker F.M. Hydra effect and paradox of enrichment in discrete-time predator-prey models. Mathematical Biosciences. 2019;310:120–127. doi: 10.1016/j.mbs.2018.12.010. [DOI] [PubMed] [Google Scholar]
- Zhang P., Sandland G.J., Feng Z., Xu D., Minchella D.J. Evolutionary implications for interactions between multiple strains of host and parasite. Journal of Theoretical Biology. 2007;248(2):225–240. doi: 10.1016/j.jtbi.2007.05.011. [DOI] [PubMed] [Google Scholar]
- Zlateva K.T., de Vries J.J., Coenjaerts F.E., van Loon A.M., Verheij T., Little P. Prolonged shedding of rhinovirus and re-infection in adults with respiratory tract illness. European Respiratory Journal. 2014:erj01721–erj02013. doi: 10.1183/09031936.00172113. [DOI] [PubMed] [Google Scholar]