Abstract
The past decade has witnessed an explosion in the use of super-resolution fluorescence microscopy methods in biology and other fields. Single-molecule localization microscopy (SMLM) is one of the most widespread of these methods and owes its success in large part to the ability to control the on-off state of fluorophores through various chemical, photochemical, or binding-unbinding mechanisms. We provide here a comprehensive overview of switchable fluorophores in SMLM including a detailed review of all major classes of SMLM fluorophores, and we also address strategies for labeling specimens, considerations for multichannel and live-cell imaging, potential pitfalls, and areas for future development.
Graphical Abstract

1. Introduction
Fluorescence microscopy is a workhorse technique in biological research due in large part to its ability to probe the distributions of specific molecules or structures in fixed and living specimens. Despite its widespread use in biology, however, fluorescence microscopy has long suffered from poor spatial resolution that obscures key details of interest. The limited spatial resolution results from the diffraction of light into the far field (i.e., at a distance of many wavelengths from the emitter) and is referred to as the diffraction limit of light microscopy or Abbe’s diffraction limit, in honor of Ernst Abbe. Nearly 150 years ago, Abbe articulated the inverse relationship between spatial resolution and numerical aperture1, as summarized by the equation d ≈ λ/2NA, where d is spatial resolution, λ is the wavelength of light, and NA is numerical aperture of the microscope objective lens. In practical terms, this means that objects closer than 200-250 nm are unresolved by traditional approaches to light microscopy. Although electron microscopy and atomic force microscopy can achieve much better spatial resolution, these techniques generally have a poor ability to report on distributions of specific molecules, lack compatibility with living specimens, and/or are limited to measurement of surface features.
The 2014 Nobel Prize in Chemistry recognized the development of optical microscopy techniques that circumvent the 200-250 nm diffraction limit of light and that now allow researchers to image distributions of molecules with a resolution of 100 nm or better. The prize was awarded to Stefan Hell for the development of stimulated emission depletion microscopy (STED), to W.E. Moerner for the optical detection of single molecules, and to Eric Betzig for the development of single-molecule localization microscopy (SMLM) which will be the main focus of this review.2–4 These techniques and related subsequent developments are now widely utilized in studying diverse problems in biology and other disciplines at most major research institutions worldwide, a testament to the ability of transformative methodologies to stimulate discovery and innovation.
Previous reviews have discussed super-resolution methods and/or SMLM in general,5–13 including some that have provided guides/protocols for SMLM14–18 and others that have focused on fluorescent probes for SMLM.19–25 Here, we provide a comprehensive and up to date review of fluorophores and probes that enable SMLM, including a history of the field, a review of the major classes of SMLM fluorophores, strategies and considerations for sequential localization, a discussion of key methods for specimen labeling, a discussion of multichannel and live SMLM, cautionary notes about some potential artifacts or pitfalls, and we also discuss future directions. We use the acronym SMLM, throughout, to acknowledge the work of many groups who have contributed to this field and in consideration that the initial four publications of the field from 2006 used different names/acronyms (i.e., STORM26, PALM27, fPALM28, PAINT29), with subsequent variations introducing many more names to be described in a later section.
2. Basics of SMLM
2.1. SMLM Principle
In SMLM, individual fluorophores on a specimen are detected sequentially and localized with low position uncertainty in order to build up a high-resolution image of the labeled specimen (Figure 1). The fluorophores may be introduced through the use of antibodies, fluorescent proteins, small molecules, or other labels, and fixed or living specimens may be imaged using the procedure. The raw data typically consist of movies containing thousands of frames, where each frame contains a sufficiently sparse subset of molecules to allow detection of individual molecules. Automated analysis software is then used to determine the positions of the individual molecules in all frames of the movie. Because each molecule emits a large number of photons, the position of each molecule may be determined with a low uncertainty (typically in the range 20-50 nm, FWHM (full width at half maximum)). From a list of the known molecule positions, software is used to render a high resolution SMLM image.
Figure 1.
Basic principle of SMLM. Scale bar indicates the ~250 nm diffraction limit of visible light.
2.2. History of SMLM
Multiple lines of work converged in the first few years of the new millennium, resulting in several independent publications of the basic SMLM idea within the span of ~four months in 2006.26–29 Below is a brief account of some of the key developments leading up to the breakthrough year for SMLM. Rather than a strictly chronological account, this story has been organized according to five themes that include: the optical detection of single molecules and photoswitches; the localization of multiple overlapping emitters; single-molecule FRET (Förster resonance energy transfer) and single-molecule tracking; the pursuit of improved spatial resolution; the SMLM breakthrough year; and analogs of SMLM used in diverse imaging modalities
2.2.1. Optical detection of single molecules and photoswitches
Single molecules were first optically detected by frequency-modulation absorption spectroscopy in 1989 by Moerner et al.30 This breakthrough was followed by two key papers in 1990 where single molecules were detected by fluorescence in crystals at low temperature by Orrit et al.31 and at room temperature in solution by Shera et al.32 It was evident from the first fluorescence-based studies and from others published shortly thereafter that fluorophores exhibit frequent transitions between distinct states which in some cases could be driven by light as a photoswitch.31–37 In 1997, a noteworthy paper from Dickson et al. reported the observation of reversible photoswitching of individual fluorescent protein molecules (Figure 2).38 Around this same time, it was also shown that other fluorescent emitters such as quantum dots, conjugated fluorescent polymers, and individual photosynthetic reaction centers each exhibited switching or blinking behavior.39–41 Although reversible photochromic effects42,43 have been known for many decades prior to the optical detection of single molecules, it was nevertheless surprising to observe fluctuations and/or switching behavior among such broad classes of chromophores.
Figure 2.
On-off photoswitching of YFP. A single YFP molecule was illuminated with 488 nm light prior to odd-numbered frames, creating a nonfluorescent state. Illumination of nonfluorescent YFP with 405 nm light prior to even-numbered frames returned the molecule to the fluorescent state. The process could be repeated many times. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 38, Copyright 1997.
In addition to the 1997 single-molecule paper by Dickson et al.38, researchers began to discover that fluorescent proteins exhibited a range of intriguing photoswitching properties, including ensemble studies which reported light-dependent fluorescence enhancement as well as green to red or red to green switching behavior.44–48 Robust photoactivatable or photoswitchable fluorescent proteins were engineered and/or selected for beginning with photoactivatable GFP49 and Kaede50 in 2002; these were soon followed by kindling FP51, Dronpa52, and many others.53–57 In the intervening years, the palette of photoactivatable or photoswitchable fluorescent proteins has grown rapidly and is discussed further, below, as well as in several reviews on the subject.25,58–66
2.2.2. Localization of several overlapping emitters
In 1995, Betzig proposed a general strategy to circumvent the diffraction limit of light microscopy by sequentially identifying distinct features labeling a structure of interest that are packed more densely than the diffraction limit.67 While the proposal, which focused on spectrally distinguishable emitters, resembled super-resolution methodology previously proposed by Burns et al. in 1985 for localizing spatially overlapping point emitters with distinct spectral properties68, Betzig clearly articulated how detection of individual fluorophores could achieve spatial resolution that surpasses Abbe’s diffraction limit. Experiments by van Oijen et al. in 1998 were able to detect up to seven spectrally distinct emitters within a diffraction limited region by using these schemes.69 Several years later, in 2004, two reports of super-resolution methods based on localization by sequential photobleaching were able to identify up to ~5 emitters per diffraction-limited region with identical spectra;70,71 this was followed in 2005 by work from Lidke et al. proposing and demonstrating the identification of two blinking quantum dots with identical spectra within a diffraction-limited region.72
2.2.3. Single-molecule FRET and single-molecule tracking
By the late 1990s, single-molecule FRET had been developed and began to be applied to the study of molecular interactions and dynamics of proteins and nucleic acids, etc.73–75 In short order, the fluorophores Cy3 and Cy5 became popular choices for single-molecule FRET, and in 2005 two groups independently published observations of reversible photoswitching exhibited by the fluorophore Cy5 in deoxygenated solutions containing a thiol, a solution commonly used for single-molecule FRET studies.76,77 While this photoswitching or blinking behavior had been previously observed by other researchers as a nuisance in single-molecule FRET studies, the two reports in 2005 recognized that the remarkable photoswitching properties of Cy5 could be potentially useful and suggested applications as memory devices or short-range probe of intermolecular distance.
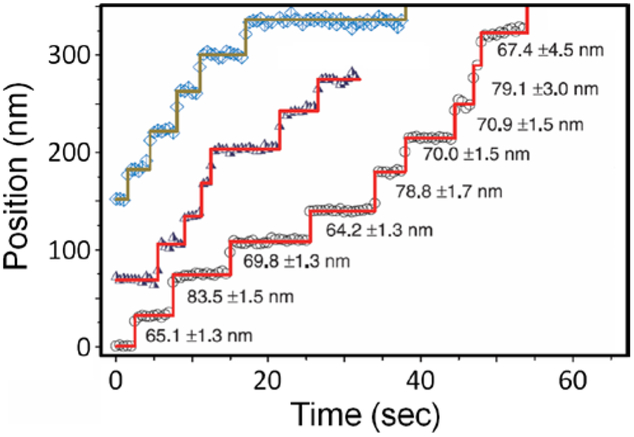
In parallel with the above, the use of fluorescence microscopy had proliferated rapidly among biologists due to technical advances in instrumentation, fluorescent probes, imaging methodology, and data analysis.78 Biophysical experiments studying motor proteins and other dynamic processes had long before developed strategies for detecting few nanometer displacements of relatively large objects such as metal nanoparticles or beads.79 By 2003, Yildiz et al. extended the concept to few nanometer localization of single fluorescent molecules using FIONA (fluorescence imaging with one nanometer accuracy, Figure 3), work that was particularly influential in the development of SMLM methods.80–82
Figure 3.
Trajectories of the motor protein Myosin V translocating along immobilized actin filaments, where the abrupt vertical jumps indicate individual stepping events. This was achieved by labeling Myosin V dimers on one head with a Cy3 fluorophore and then localizing the fluorophores to within a few nm uncertainty. From Ref. 80, adapted with permission from AAAS.
2.2.4. The pursuit of improved resolution
In the decades preceding the development of SMLM, many innovative methods were developed for improving the spatial resolution of fluorescence microscopy. Confocal microscopy progressed quickly in the late 1980s and has become the gold standard against which subsequent methods are compared, generally achieving a spatial resolution of ≥200 nm along the transverse dimension and ≥500 nm along the axial dimension.78 One of the earliest super-resolution methods was near-field scanning optical microscopy (NSOM), developed in the mid 1980s83–86, which scanned a small aperture or tip over a specimen and enabled measurement of surface features at <100 nm lateral spatial resolution.87,88 Dual objective lens confocal microscopy, also known as 4pi microscopy, was first reported by Hell et al. in 199489 and was further developed by Gustafsson et al. with <100 nm axial resolution.90 Stimulated emission depletion (STED) microscopy was developed by Hell and colleagues in the 1990s, published first as a concept in 199491 and then realized experimentally by Klar et al. in 1999 and in 2000 with better than 100 nm lateral spatial resolution.92,93 Structured illumination microscopy was developed by Lanni et al.94, Heintzmann et al.95, and Gustafsson et al.96 during the 1990s and enabled up to a twofold improvement of spatial resolution over the diffraction limit along the lateral and axial dimensions. All of these methods were further improved in subsequent years, and some are commonly used by biologists to this day. In any event, by the early 2000s it was clear that improving the spatial resolution of fluorescence microscopy was a critically important task for biology and the problem had captured the attention of a large community of researchers.
2.2.5. The SMLM breakthrough year
With the advent of robust photoactivatable or photoswitchable fluorophores, relatively mature hardware and software for localization of single fluorophores down to a few nm, published concept or demonstrations for super-resolution microscopy including some which used point-emitters, and a growing interest in developing tools to improve the spatial resolution of fluorescence microscopy, the stage was set for rapid progress. This progress became evident within the span of a few months in late 2006, when four independent groups published experimental demonstrations of SMLM including STochastic Optical Reconstruction Microscopy (STORM) from the group of Xiaowei Zhuang, PhotoActivated Localization Microscopy (PALM) from Eric Betzig and Harald Hess and collaborators, Point Accumulation for Imaging in Nanoscale Topography (PAINT) from the group of Robin Hochstrasser, and fluorescence Photoactivation Localization Microscopy (fPALM) from the group of Samuel Hess.26–29 All four of these papers utilized serial detection of multiple emitters with identical spectral properties within a diffraction-limited area, in contrast with earlier proposals or demonstrations. STORM, PALM, and fPALM achieved sequential detection by sequentially activating single photoswitchable fluorescent proteins or organic fluorophores bound to the specimen26–28 whereas PAINT utilized transient binding of hydrophobic fluorophores that are activated upon binding to a lipid membrane.29 Notably, PAINT built on earlier work from the same group using diffusing probes that activate upon binding.97 Together, these methods formed the key foundations of SMLM that would be further developed and elaborated by growing community of researchers.
2.2.6. Diverse analogs of SMLM
It is interesting to observe with the advantage of hindsight that point-by-point acquisition and reconstruction methods resembling SMLM are used in diverse imaging modalities. In a first example, low electron dose electron microscopy methods from as far back as 1978 utilize low illumination current such that individual electrons are detected sequentially, individually localized, and accumulated to reconstruct an image.98 A similar procedure has recently become popular in cryo-electron microscopy methods which use ‘super-resolution mode’ for sub-pixel localization of individual electrons incident upon a direct detection camera.99,100 In a second example, medical radioisotope imaging (i.e., positron emission tomography and single photon emission computed tomography) also uses sequential detection of individual decay events with sub-pixel localization, although these data require considerable additional processing to reconstruct the original distribution of a radioactive contrast agent.101 In a final example, ultrasound imaging has now adopted the SMLM approach for sequential localization of microbubbles in order to achieve super-resolution ultrasound images of tissue vasculature.102–105
2.3. Alphabet Soup
Single-molecule localization microscopy in many ways built off of earlier single-molecule techniques bearing colorful acronyms including Fluorescence Imaging One Nanometer Accuracy (FIONA)80, Defocus Orientation and Position Imaging (DOPI)106, Single-molecule High Resolution Colocalization (SHREC)107, Single-molecule High Resolution Imaging with Photobleaching (SHRImP)70, Nanometer-Localized Multiple Single-molecule microscopy (NALMS)71.
The proliferation of new names or acronyms continued unabated with the advent of SMLM in 2006, with four papers published under four different names: Stochastic Optical Reconstruction Microscopy (STORM)26; PhotoActivated Light Microscopy (PALM)27; Point Accumulation for Imaging in Nanoscale Topography (PAINT)29; and fluorescence PALM (FPALM)28. These were shortly followed by many others, including: PALM with Independently Running Acquisition (PALMIRA)108; Ground Stated Depletion with Individual Molecule Return (GSDIM)109; direct STORM (dSTORM)110; Single Particle Tracking PALM (sptPALM)111; Photoactuated Unimolecular Logical Switching Attained Reconstruction (PULSAR)112; Spectral Precision Distance Microscopy (SPDM)113; interferometric PALM (iPALM)114; Reversible Photobleaching Microscopy (RPM)115; Nanometer Accuracy by Stochastic Catalytic reactions (NASCA),116 Single-molecule Active Control Microscopy (SMACM)117; Chemically Improved Resolution for Optical Nanoscopy (CHIRON)118; super-resolution power-dependent active intermittency PAINT (SPRAI-PAINT)119; Binding-Activated Light Microscopy (BALM)120; Bleaching/blinking assisted Localization Microscopy (BaLM)121; Complementation Assisted Light Microscopy (CALM)122; Enzymatic Turnover Activated Localization Microscopy (ETALM)123; Tracking and Localization Microscopy (TALM)124). While these various reports have contributed a range of strategies for detection, switching mechanisms, or other innovations, the barrage of acronyms can be bewildering to nonspecialists.
2.4. Requirements for SMLM fluorophores
In SMLM, many individual points (referred to as localizations) must be collected sequentially in order to outline a structure of interest with sufficient detail (Figure 4), a conceptually similar requirement to fulfilling the Nyquist sampling criterion. In that case, the best achievable resolution cannot be less than twice the average distance between neighboring localizations, or in other words, for D dimensions (where D is typically 2 or 3), the maximum spatial resolution is 2/(localization density)1/D.125 Due to the stochastic nature of the samples in SMLM, which are not, in general, evenly spaced, Legant et al.126 have suggested that the localization density should be exceeded by a factor of ~5 fold in SMLM in order to reliably achieve the nominal Nyquist resolution, with the caveat that smaller structures or less dense proteins/molecules would not require such high densities (see Section 9.4). Additionally, the position of each localization must be known with low uncertainty.
Figure 4.
Impact of density of localizations on resolution achievable by SMLM. A high density of localizations (top row) resolves even the finest details on all of the test grids, while lower densities of localizations (middle and bottom rows) only resolves details about the larger grids. Reproduced by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 125, Copyright 2008.
The requirements for a high density of localizations and a low position uncertainty in turn place very specific requirements on fluorophores for SMLM. First, the fluorophores must be able to be switched between a bright and a dark state, with most molecules in the dark state, and the contrast between these states should be very high. For instance, in order to detect one bright fluorophore in a diffraction-limited volume out of a background of 100 dark fluorophores, the ratio of brightness of the on and off states, referred to as the on-off contrast ratio, should be greater than ~100:1. Although the on-off contrast ratios are seldom reported, many of the popular SMLM fluorophores have on-off contrast ratios much greater than ~1000:1. Second, the fluorophores must have a low duty cycle, or fraction of time spent in the bright state prior to bleaching, regardless of whether the fluorophore is reversibly photoswitchable or irreversibly photoactivatable. In the case where there are 100 fluorophores in a diffraction-limited volume, the fluorophores must spend much less than 1% of their time in the bright state in order to detected individually (i.e., must have a duty cycle of <0.01) in order to avoid a very high incidence of double/multiple localizations (see Section 9.3). The relationship of duty cycle to on/off rates is shown in Figure 5 and, notably, is applicable whether using reversible switching or irreversible activation followed by bleaching. It should be noted that the duty cycle of a fluorophore is in general dependent upon illumination conditions and chemical environment; researchers commonly illuminate a specimen with violet light (~405 nm) during the acquisition of a SMLM movie in order to acquire localizations more quickly (i.e., increasing the duty cycle above tabulated value) while taking care to avoid excessive double/multiple localization artifacts. Third, the fluorophores must be bright enough to allow single-molecule detection and they must emit a large number of photons during an on burst, where position uncertainty scales roughly as the inverse square root of the number of detected photons.80,81,127 The ideal SMLM fluorophore would therefore have a low duty cycle, a high on-off contrast ratio, a large number of emitted photons per burst, as well as other important but less fundamental properties that include the number of switching cycles, the utility of the fluorophore in labeling structures of interest, and low phototoxicity when imaging living specimens.
Figure 5.
A) Simplified analysis of reversible switching, B) irreversible activation, and C) estimate of duty cycle, where koff is the rate of switching off, kon is the rate of switching on, koff,2 is the rate of bleaching for a reversible switch, DC is the duty cycle, ton is the average time spent in the on state prior to bleaching, and toff is the average time spent in the off state. Notably, DC is approximately the ratio of the on and off rates for both reversible switching and irreversible activation.
2.5. Fluorophore switching based on blinking/bleaching/binding of probes
In SMLM, individual fluorophores are imaged on a dark background and positions are determined through analysis of the distribution of fluorescence in the on state. A set of related methods uses analysis of two or more subsequent frames to determine distributions of fluorophores either by observing the abrupt appearance/disappearance of a signal on top of a nonzero background or through statistical analysis of fractional fluctuations of the signal. In some cases, photoswitching properties that may not be suitable for observing individual molecules by SMLM may still be useful for these related methods. Similarly, hardware requirements for these methods may in some cases be relaxed.
The activation or bleaching of a single fluorophore in the presence of other emitting fluorophores which do not change may be detected by subtracting consecutive frames (Figure 7). A fluorophore which appears abruptly due to photoactivation or binding will produce a bright peak in a difference image, while a fluorophore which disappears abruptly due to bleaching or unbinding will produce a valley. Although the presence of a high background increases localization uncertainty,81 individual fluorophores may nonetheless be detected with sub-diffraction resolution provided that one or fewer transitions occur per diffraction-limited area between consecutive images.
Figure 7.
Schematic illustration of photobleaching based SMLM wherein consecutive frames (Fn) are subtracted to localize individual bleached molecules. A similar strategy can be used to identify individual activated molecules in consecutive frames.
Two reports published in 2004 introduced the acronyms SHRImP (single-molecule high-resolution imaging with photobleaching) and NALMS (nanometer-localized multiple single-molecule microscopy) and used sequential photobleaching to localize a handful of fluorophores per diffraction limited area.70,71 On the same theme, generalized SHRImP (gSHRImP), QDB3, and bleaching/blinking assisted localization microscopy (BaLM) were published in 2011 for the detection of both bleaching and blinking events through difference analysis of consecutive frames.121,129,130 A more computationally intensive approach termed 3B Analysis (Bayesian analysis of blinking and bleaching) concurrently determines the positions of relatively large ensembles of molecules from image stacks consisting a few hundred frames based on blinking and bleaching signals observed within the stack.131 SOFI (super-resolution optical fluctuation imaging) and photobleaching microscopy with nonlinear processing (PiMP) instead use statistical methods to analyze spatially and temporally varying fluorescence fluctuations within the specimen in order to extract sub-diffraction images.132–134 Given the large variety of ways to extract sub-diffraction-limit information from the fluctuating fluorescence emission of a specimen, it is likely that fluorophores which are unsuitable for one method may find good application in another.
3. Switchable fluorophores for SMLM
A wide range of fluorophores has been utilized for SMLM, including synthetic organic fluorophores, fluorescent proteins, and quantum dots. These fluorophores rely upon a diverse set of switching mechanisms to convert between fluorescent and nonfluorescent forms so that many individual fluorophores may be sequentially detected in an image series within a diffraction limited volume for SMLM. We review the main fluorophore categories, below, with an emphasis on organic fluorophores spanning a range of chemical classes.
3.1. Major classes of organic fluorophores
To accompany the discussion, below, on organic fluorophores for SMLM, we have prepared a supporting document containing the structures of most of the organic fluorophores we discuss in this paper together with a summary of the key switching reactions (see Supporting Information).
3.1.1. Cyanines
Cyanine dyes are perhaps the most popular fluorophore class for SMLM, particularly the red-absorbing fluorophore Cy5 and its close structural relatives such as Alexa Fluor 647. The photoswitching behavior of Cy5 was first reported by Bates et al. and Heilemann et al. in 2005, where both groups detected the switching behavior through observation of single molecules transitioning reversibly between fluorescent and nonfluorescent states (Figure 8A).76,77 The reports showed that red light could be used to switch Cy5 to a long-lived nonfluorescent state and that shorter wavelength light could be used to activate nonfluorescent Cy5 back to a fluorescent state; Heilemann et al. utilized strong visible light for activation, while Bates et al. utilized much weaker visible light to excite Cy3 “activator” dyes that were positioned within 1-2 nm of Cy5 molecules and could activate dark-state Cy5 molecules.76,77 Photoswitching in both cases was achieved in a solution commonly used for single-molecule FRET microscopy which contains a primary thiol such as β-mercaptoethylamine (MEA) or β-mercaptoethanol (BME) as well as an enzymatic oxygen scavenging system (see also Section 3.4).76,77
Figure 8.
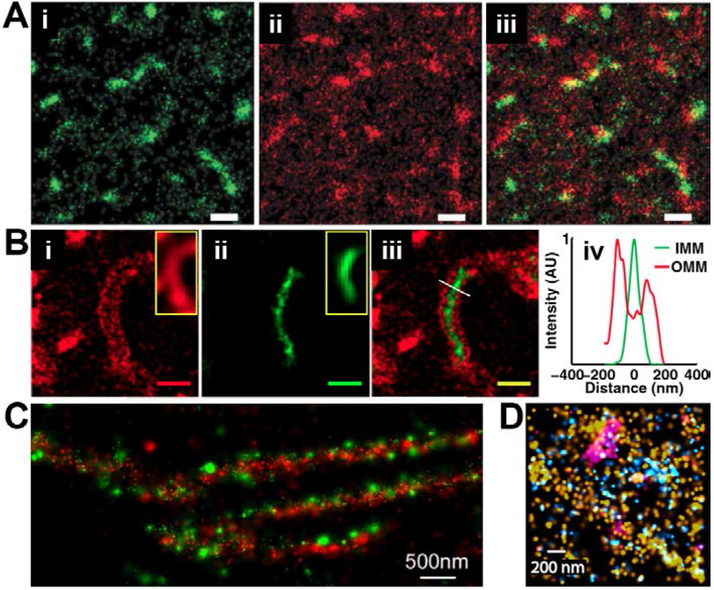
Photochemical and chemical reactions switch cyanine dyes between fluorescent and nonfluorescent states. A) Photoswitching of a single Cy5 molecule in a deoxygenated solution containing a thiol. The Cy5 molecule was continuously illuminated with 633 nm light, switching off at ~500 ms. When intense 488 nm light also illuminated the sample at time ~5 sec, the Cy5 fluorescence was recovered and the molecule repeatedly switched between bight and dark states. B) Chemical quenching of an ensemble of Cy5 molecules with the phosphine TCEP and photoactivation with brief pulses ultraviolet (UV) light (gray arrows). C) UV light-induced activation of reduction-quenched Cy3B. Reproduced with permission from Ref. 77, Copyright 2005 American Chemical Society. Reproduced with permission from Ref. 135, Copyright 2013 American Chemical Society. Reproduced by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 136, Copyright 2012.
The first application of Cy5 photoswitching to SMLM was reported in the original STORM paper in 2006 by Rust et al., where the Cy3-Cy5 dye pair was utilized to image circular RecA-coated plasmid DNA.26 The same group later showed that Cy5.5 and Cy7 also exhibit good switching properties and that each of the reporter molecules (Cy5, Cy5.5, and Cy7) may be paired with a number of activator molecules (e.g., Alexa Fluor 405, Cy2, Cy3) in order to create a palette of activator-reporter pairs for multicolor super-resolution microscopy (Figure 9A-B).137,138 Bock et al. showed in 2007 that Cy5 could be used for SMLM without an activator dye;139 this “activator-free” approach was later substantially expanded upon by Heilemann et al. and other researchers using a number of cyanine or non-cyanine fluorophores and is often referred to as dSTORM (or direct STORM).109,110,140 Importantly, Cy5 (and its close structural analog Alexa Fluor 647) is still considered one of the best overall fluorophores of any class for SMLM due to its particularly suitable photoswitching properties including a high photons per localization, a low duty cycle, and reasonably large number of switching cycles.
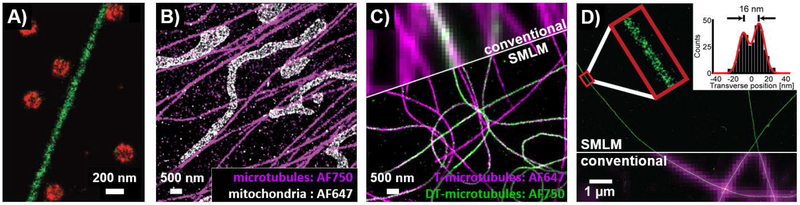
Figure 9.
SMLM imaging using cyanine dyes. A) SMLM zoom-in view of a cell immunostained for microtubules (green) and clathrin-coated pits (red) with activator-reporter fluorophore pairs. Alexa Fluor 647 was the reporter fluorophore used on both structures but Cy2 was used as an activator dye on microtubules and a Cy3 was used as an activator dye on clathrin-coated pits. SMLM images of cells immunostained for B) microtubules and mitochondria or C) tyrosinated (T) and detyrosinated (DT) microtubules using multi-reporter imaging with Alexa Fluor 647 and Alexa Fluor 750. D) SMLM image of microtubules grown in vitro, and directly labeled with Cy3B (green, with partial overlay of SMLM image on magenta conventional image); inset shows cross-sectional profile of zoomed in region (red box) with the successful resolution of the inner microtubule walls (~16 nm). In A-C, images were acquired under conditions where fluorophores can reversibly switch on and off, whereas D was acquired using reductive caging as described in the text. From Ref. 137, adapted with permission from AAAS. Adapted with permission from Ref. 138, Copyright 2012 John Wiley and Sons. Adapted with permission from Ref. 135, Copyright 2013 American Chemical Society. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 136, Copyright 2012.
Dempsey et al. used single-molecule imaging and mass spectrometry to study the structure and mechanism of formation of the dark state of Cy5.141 They found that upon illumination with red light, thiolate anions added covalently to the bridge of Cy5, disrupting the conjugation and shifting the absorption to the ultraviolet (Figure 10A).141 The site of addition to the Cy5 polymethine bridge was identified as most likely the γ-carbon of the bridge (Figure 10A).141 It was later shown by Vaughan et al. through mass spectrometry and NMR spectroscopy that the phosphine TCEP (tris 2-carboxyethyl phosphine) adds reversibly to the γ-carbon of the bridge of Cy5 even in the absence of illumination with red light (Figure 8B, Figure 10A), further supporting the idea that the γ-carbon of the bridge of Cy5 is also the site of addition of the thiolate anion.135 This TCEP-based switching approach also provided an improved two-color imaging approach using Cy5 and Cy7 (or close structural relatives of Cy5 and Cy7 such as Alexa Fluor 647 and Alexa Fluor 750) by boosting the relatively low photon output of Cy7 approximately 3-fold compared to thiol-based imaging.135
Figure 10.
Schematic illustration of switching reactions of cyanine fluorophores. A) Cy5 is switched off by BME in the presence of intense red light or by TCEP (without illumination) to form a non-fluorescent adduct in which a thiolate anion or the phosphine TCEP is bound to the γ position of the polymethine bridge, switching it to a nonfluorescent state. Illumination with ultraviolet light dissociates the adduct, returning Cy5 to its bright form. B) Cy3B and other cyanines may be chemically reduced by NaBH4 to a non-fluorescent form which is photoactivatable.
Unlike the fluorophores Cy5, Cy5.5, and Cy7 which have polymethine bridges that are either 5 or 7 carbons long, the fluorophore Cy3 does not photoswitch well in deoxygenated, thiol-containing solutions.128,141 This may reflect a lack of access to the three-carbon bridge of Cy3 or differences in the rates of other photochemical steps. Instead, the rigidified fluorophore Cy3B (Figure 10B) was found to photoswitch reasonably well in deoxygenated solutions, whether or not a thiol was included in the solution,128 implying that Cy3B may use a different mechanism of photoswitching such as a long-lived radical as has been shown to be the case for some rhodamine fluorophores.142 Glembockyte and Cosa recently showed that Cy3B exhibits remarkably efficient geminate recombination of photo-generated radical ion pairs in the presence of selected reducing agents (particularly for BME) which may help in the development of optimized imaging cocktails for SMLM (see also Section 3.4).143 Regardless, the fluorophores Cy3B, Cy5, and Cy7 and their close structural relatives exhibit good photoswitching properties that make the cyanine class of fluorophores some of the most heavily used in SMLM.128
Hydrocyanines are cyanines that have been rendered nonfluorescent through chemical reduction and that, under the right circumstances, can be activated back to a fluorescent state.144–146 This class of molecules, whose first example dates back at least to Richard Kuhn in 1932,144 has been used in various applications including as sensors for electron beams145 or reactive oxygen species146 as well as in SMLM.136 Hydrocyanines are typically formed through net addition of a hydride anion to Cα of the cyanine polymethine bridge (Figure 10A). Vaughan et al. developed an in situ method for “reductive caging” in which the cyanines Cy3, Cy3B, Cy5, or Cy5.5 could be reduced to hydrocyanines in situ through brief treatment of a fixed and labeled specimen with dilute aqueous sodium borohydride.136 The hydrocyanines were then photoactivated in an imaging cocktail (see Section 3.4) that suppresses bleaching and blinking, yielding 4-20 times more detected photons per localization (Figure 8C) and a very low resulting position uncertainty (Figure 9D). Although the improved photon yield resulted in greatly improved localization, the method was only able to activate ~12-40% of fluorophores and did not exhibit an improvement in duty cycle over thiol-based switching.136 Lehmann et al. have extended this approach to two or three color SMLM with a range of other cyanine and non-cyanine fluorophores.147 Michie et al. have developed a new rigidized Cy5 variant called Cy5B which has a recovery yield of ~38%, which is an approximately six-fold improvement over its non-rigidized analog.148
3.1.2. Rhodamines
Rhodamines are a popular class of fluorophores for many applications in fluorescence microscopy due to their high quantum yield, large extinction coefficient, and high photostability, and several rhodamines are commonly used in SMLM. The relatively easy synthesis of rhodamines also makes them good candidates for further innovation by synthetic chemists.
Several popular, commercially available rhodamine fluorophores exhibit good photoswitching in deoxygenated solutions containing a thiol, including ATTO 488, Alexa Fluor 488, and Alexa Fluor 568.109,128,140 Thus, a number cyanines and rhodamines (as well as other fluorophores) can be switched well using a single “imaging cocktail” (Section 3.4). Unlike the red and far-red cyanines such as Cy5 and Cy7, however, many rhodamines are photoswitched to a dark state that has been shown by electron paramagnetic resonance spectroscopy to consist of a long-lived radical (Figure 11, upper left).142 The radical state is believed to be generated by the photo-reduction of its triplet state by thiol in solution.142
Figure 11.
Reversible switching reactions of rhodamine fluorophores. Illumination of a fluorescent rhodamine (green, lower left) in a deoxygenated solution containing a thiol leads to the generation of a nonfluorescent, long-lived radical (upper left) while chemical reduction leads to the generation of a non-fluorescent, reduced rhodamine (lower right). Oxidation or illumination with ultraviolet light can return the radical or reduced rhodamine back to the fluorescent state. The inset diagram illustrates the thiol-induced photo-reduction of the rhodamine triplet state to a radical species (F•) which is followed by oxidation back to the ground state. Adapted from Ref. 142 with permission of The Royal Society of Chemistry (RSC) on behalf of the Centre National de la Recherche Scientifique (CNRS) and the RSC.
Rhodamines are also good candidates for reductive caging and show very good photon-recovery yields by UV illumination (e.g., 66% for ATTO 488) compared to cyanines (12–40% recovery).136,147 The net addition of a hydride anion to the central carbon bridge of the xanthene group breaks its conjugation (Figure 11, lower right),147 thereby quenching fluorescence emission. UV illumination removes the hydride and restores the caged rhodamine back to its fluorescent form prior to imaging and bleaching for SMLM. Lehmann et al. screened a variety of rhodamine dyes for reductive caging and found that ATTO 488, ATTO 514, and ATTO 532 also have good recovery yields of ~30-60%.147
One of the most well-known reactions of rhodamines is that their absorption/emission can be reversibly switched off through formation of a spiro-ring on the central carbon of the xanthene (Figure 12, top).149 An intramolecular nucleophile such as a carboxylate, hydroxy, or amide, is generally required to form the spiro-ring and break the conjugation of the xanthene chromophore. Changes of pH or the concentration of certain metal ions or small molecules can shift the equilibrium in either direction by coordination effects or by specific reactions,150,151 while tuning electron density on the xanthene ring can also shift the equilibrium.152 This spiro-ring equilibrium reaction has been widely exploited in the development of various rhodamine-based sensors or switches150 and has been adapted for use in SMLM.153
Figure 12.
Schematic illustration of switching reaction of rhodamine fluorophore via ring opening and intramolecular spirocyclization reactions (top). Examples of spiro-rhodamine dyes used for SMLM are shown (bottom).
Different rhodamine skeletons show different sensitivities to environmental variations which can greatly influence the open/close equilibrium. For example, Lukinavičius et al. showed that the red-shifted dye Si-rhodamine (SiR, absorption/emission = 645 nm/661 nm) with a carboxylic acid group at the 2-position of the phenyl ring (SiTMR in Figure 12, shown in its nonfluorescent spiro-ring form) is quenched while free in solution but becomes ~5x brighter when a ligand conjugated to the dye binds to a reactive protein tag such as SNAP.154 SiTMR thus exhibits decreased background signal while improving cell permeability because the favored spiro-ring form of the dye masks charges on the zwitterionic, open form of the dye.154 At the same time, the bound, open form of the dye exhibits suitable photoswitching properties for SMLM, allowing, for instance, high quality imaging of histone proteins in the nuclei of living specimens without interference from unbound probes.154
Recently, Lavis’s group has made substantial improvements to the fundamental spectroscopic and photoswitching properties of rhodamine fluorophores.155 For instance, they discovered that the replacement of the classical N,N-dialkyl groups with four-membered azetidine rings substantially improved the fluorescence quantum yield of several rhodamine dyes (e.g., Figure 13A, R=H, JF646 (Janelia Fluor)).156 By further adjusting the substituents on the azetidine ring, they were able to fine-tune the ring-closed/open equilibrium,152 resulting in a series of novel fluorogenic/chromogenic ligands such as JF646-Halo and JF635-Halo, which provide a 21-fold and 113-fold absorbance enhancement, respectively, upon binding a HaloTagged protein (Figure 13B). Binding of these probes to HaloTagged proteins also stabilizes their ring-open (bright) form and with much higher on-off contrast than that of the N,N-dialkyl equivalent ligand while retaining good switching properties (Figure 13C).
Figure 13.
A) Structure of two example fluorogenic/chromogenic HaloTag ligands JF646-Halo and JF635-Halo. B) Absorbance enhancement of JF646-Halo and JF635-Halo after binding with HaloTag protein. C) No wash fluorescence imaging of COS7 cells expressing HaloTag-histone H2B fusion and then labeled with JF646-Halo (i), JF635-Halo (ii) and SiTMR-Halo (iii). Scale bar, 15 μm. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 152, Copyright 2017.
Beyond providing a means for reducing background signal, the open-closed equilibrium of spiro forms of rhodamines can, themselves, form the basis of the on-off switching chemistry for SMLM. For example, spiro-lactams are an easily formed nonfluorescent form of rhodamine fluorophores (RSA-1 and RSA-2 in Figure 12) which in general exhibit higher stability than spiro-lactones, although acid or certain other analytes can shift the equilibrium toward the ring-open state.153 The increased stability of spiro-lactams makes them suitable to caging a wider range of rhodamine dyes compared to spiro-lactones, and interestingly, by proper tuning of substituents on the amide moiety, the spiro-lactam ring can be designed to be opened by UV light irradiation before spontaneously converting back to the ring-closed form (Figure 12, top). For example, Fölling et al. incorporated a 4-aminophthalimide substituent (RSA-1) such that electron-withdrawing groups on the phthalimide, together with its original phenylacetyl, stabilized the charge on the amide nitrogen atom by delocalization and improving the photoactivation process.157–159 The resulting dye RSA-1 can be activated with ultraviolet light (~310-380 nm) in order to open the spiro-lactam ring such that the dye becomes fluorescent, emitting up to 900 detected photons per molecule, before returning to the spiro-lactam form. Fölling et al. also demonstrated that two-photon activation can extend this type of rhodamine spiro-lactam for deep 3D imaging with axial optical sections of 0.5-1 μm (Figure 14A-C).159 Further optimizations were performed, either by fluorination, sulfonation or other functionalizations160 on xanthene or phenyl-amide moieties,158,161 or by introducing other new rhodamine frameworks,158 to improve photoactivation and tune the optical properties of dyes for multicolor imaging (Figure 14D-H).158,162 Lee and co-workers hypothesized that extension of the conjugation area on the phenyl-lactam moiety will further lower the energy required to photoactivate the ring-closed isomer163 which is favorable for retaining the fluorescence from already activated isomers. Inspired by this, RSA-2 was developed (Figure 12), showing more efficient photoactivation in response to 405 nm illumination. The pyridinium-derived activated ester linker furnished a convenient handle for labeling of the specimen of interest. Using RSA-2 as photoswitchable fluorescent marker, they quantified the distribution of cell stalk lengths at a mixed population of C. crescentus cells (Figure 14I).163
Figure 14.
SMLM images obtained using various photoswitchable spiro-lactam rhodamines. A) A 3D SMLM image of a 5 μm silica bead whose surface has been stained with PC-Rh590 and imaged in 16 layers. B) The middle focal plane from A, and C) an associated line profile showing a lateral FHWM of ~30 nm. D-H) Multicolor images of mixtures of silica beads labeled with photoswitchable spiro-lactam rhodamine fluorophores SRA545 (blue), SRA577 (green), and SRA617 (red). I) Montage of SMLM images of C. crescentus cells labeled with RSA-2. Scale bars, 1 μm for B, D and I, 200 nm for E-H. Adapted with permission from Ref. 159, Copyright 2008 John Wiley and Sons. Adapted with permission from Ref. 162, Copyright 2008 American Chemical Society. Adapted with permission from Ref. 163, Copyright 2014 American Chemical Society.
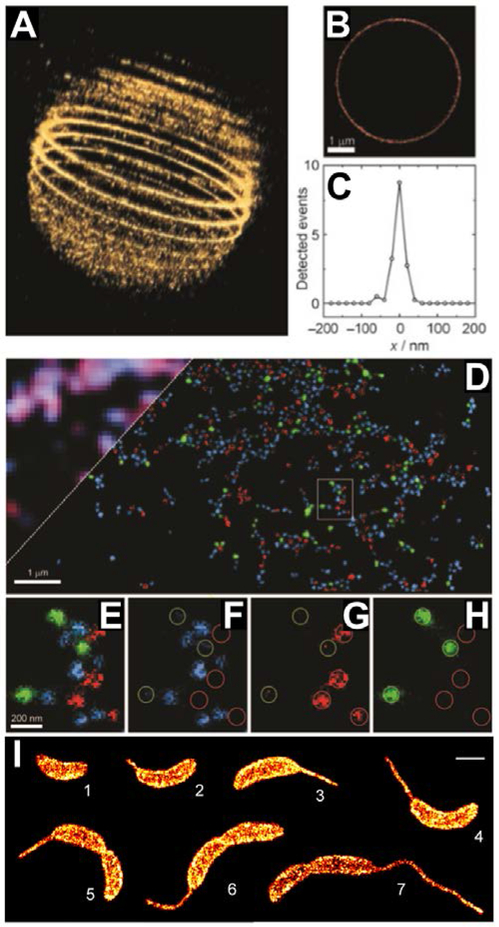
Rather than use light to switch caged fluorophores, the equilibrium between ring-closed (nonfluorescent) and ring-open (fluorescent) forms may be tuned in order to allow passive on-off switching that is suitable for SMLM. Uno et al. have achieved this by preparing and optimizing a series of rhodamine derivatives bearing various intramolecular nucleophiles at the 2-position of the phenyl ring.164 Among these, HMSiR (Figure 12) exhibited good switching properties in fixed or living specimens with ~2,600 detected photons per switching event and a duty cycle of ~0.01 (Figure 15A).164 Although passive switching lacks the convenient adjustability of on/off rates achievable by photoswitching-based approaches, passive switching avoids the use of possibly toxic reagents or near-ultraviolet illumination for live-cell imaging. SR imaging of HMSiR labeled RecA filaments by SMLM revealed a circular structure (Figure 15B-C) with a feature width (51.4 ± 3.2 nm, FWHM) that was comparable to that achieved with thiol-based Cy5 photoswitching in deoxygenated solutions (47.2 ± 2.0 nm, FWHM).
Figure 15.
Use of the spontaneously-blinking fluorophore HMSiR for SMLM. A) Single-molecule fluorescence time traces of antibody-bound HMSiR in the absence of thiol and an oxygen-scavenging system. B) Conventional and C) SMLM image of circular, plasmid DNA decorated with HMSiR-labeled RecA filaments. D) Cross-sectional profile of yellow boxed-region in C, showing an apparent ~52 nm width. Scale bars, 500 nm. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 164, Copyright 2014.
Belov et al.165 reported a novel caging strategy for rhodamines by using a diazoketone group to form nonfluorescent spirocyclic rhodamines, termed Rhodamine NN dyes (Figure 16). Unlike spiro-lactams, which after activation spontaneously return to their ring-closed nonfluorescent form, the irradiation of non-fluorescent Rhodamine NN with UV light converts the rhodamine to the fluorescent spiro-open form while also undergoing subsequent chemical steps that degrade the caging group (fluorescent products are shown in Figure 16).165 Thus, the uncaged rhodamine fluorophores are locked in the ring-open until they photobleach. The flexible diazoketone caging group can be incorporated into a variety of rhodamine and carbopyronine fluorophores in order to span a wide spectral range.166,167
Figure 16.
Photo-activation reactions of rhodamine derivatives by photo-cleavage of diazoketone on spiro-ring (top). Examples of diazoketone caged rhodamine dyes, PA-JF549, PA-JF646 and their corresponding Halo- and SNAP tag ligands (bottom).
Grimm et al. recently reported improved photoactivatable rhodamine NN dyes168 based on combination of diazoketone caging165 with the azetidine-rhodamine dyes JF549 and JF646.156 The resulting dyes, such as PA-JF549 and PA-JF646, exhibit good caging/uncaging properties and good cell permeability that are well suited for SMLM and single-molecule tracking experiments (Figure 16 bottom).168 Upon photoactivation, PA-JF549 and PA-JF646 each generate fluorescent products with spectroscopic properties consistent with uncaged rhodamine dyes (Figure 17A). The photoconversion showed improved yields when attached to Halo- or SNAP-tag proteins. Compared with the genetically encoded fluorescent protein mEos3.2 and the commercially available TMR-HaloTag probe, PA-JF549-Halo also shows higher detected photons per particle per frame, longer tracks in single-molecule tracking experiments, and higher calculated localization precision in SMLM (Figure 17B-C). Grimm et al. used PA-JF549-Halo, together with the spectrally independent far-red Si-rhodamine fluorophore PA-JF646, to perform single-molecule tracking and SMLM in live embryonic stem cells expressing histone H2B–SNAP-tag and HaloTag–Sox2 (Figure 17D).
Figure 17.
A) PA-JF646-Halo can be photoactivated, and when bound to a Halo-tagged protein, the photoactivation yield is improved. Comparison of B) photons per localization per frame and C) track length (number of frames per localization) for PA-JF549-Halo and mEos3.2 showing that PA-JF549-Halo is brighter and produces longer tracks. D) Single-molecule tracking and SMLM in a live embryonic stem cell expressing histone H2B–SNAP-tag and HaloTag-Sox2 labeled with PA-JF646-SNAP and PA-JF549-Halo, respectively; the apparent diffusion rate of Sox2 is lower for the fraction of Sox2 that colocalizes with H2B than for the fraction that does not colocalize (H2B data not shown here). Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 168, Copyright 2016.
Another well-known rhodamine caging strategy in combination with formation of a spiro-ring is the introduction of 2-nitrobenzyl derivatives as photo-cleavable caging groups (Figure 18),169–172 a caging strategy originally developed by Mitchison with caged fluorescein in 1989.173 Among these, Wysocki et al. recently established an improved strategy to functionalize rhodamine (and fluorescein) dyes with 4,5-dimethoxy-2-nitrobenzyl (DMNB) groups on the aniline moiety of the xanthene ring (RhQ8, Figure 18).174 A key advance in their work was the use of reduced fluorophore leuco-dye intermediates in order to improve the efficiency of rhodamine acylation with the DMNB caging groups.174 UV irradiation can cleave the DMNB group in order to restore the dye to its fluorescent form and this caging strategy can also be applied to other structurally similar fluorophores, such as silicon-rhodamine,175 fluorescein,174 carborhodamine, and carbofluorecein.176 However, the use of two caging groups (e.g., RhQ8 in Figure 18) potentially increases the irradiation time required to achieve full uncaging and also leads to the possibility of incomplete uncaging and an associated variation in the emission signal.170,177 Banala et al.178 caged rhodamine 110 with one DMNB group (BG-cRhod1 in Figure 18) while the other aniline group was modified with a functional linker for conjugation with benzylguanine (BG) to react with SNAP-tag fusion proteins. Photo-cleavage of the DMNB group by illumination with ultraviolet light recovers the fluorescence emission,177 leading to a large fluorescence enhancement.
Figure 18.
Photo-switching of 4,5-dimethoxy-2-nitrobenzyl (DMNB) group caged rhodamine derivatives by photo-cleavage reaction. Example dyes RhQ8 and BG-cRhod1 used for SR-imaging by SMLM are shown at bottom.
3.1.3. Spiropyrans and oxazine-auxochromes
Spiropyrans (SPs) are classic photochromic molecules which have attracted interest for their use in SMLM.179,180 Similar to the on-off switching behavior of spirocyclic rhodamines, SPs can exist in a spiro-closed form (SP) or a spiro-open form called the merocyanine (MC) form. Additionally, illumination with suitable wavelengths of light can shift the equilibrium between the SP and MC forms (Figure 19 top).179,181 The spiro-cyclization is formed from an internal nucleophile under visible light irradiation, which converts the Cα carbon on the bridge from sp2 to sp3 hybridization and breaks the original π-conjugation to either generate a nonfluorescent product182 or to generate a spectrally blue-shifted emissive product.183,184 UV or blue light will cause photo-elimination of the nucleophile and recovers the π-conjugation to generate MC, while visible light illumination accelerates the spiro-ring-close conversion to dark SP state again. The SP-MC conversion shows good reversibility by UV/visible light irradiation, and similar to the rhodamine spiro-lactam system, there are no requirements for special fluorescence quenching agents or a strict oxygen-free environment. Hu et al.112 have utilized SP-MC photoswitching for SMLM, in order to demonstrate the imaging of several nanoparticles within a diffraction-limited volume (Figure 20). In their work, illumination with 532 nm light shifted the equilibrium to a nonfluorescent SP state while at the same time exciting fluorescence from only those molecules remaining in the MC form. As with some other photoswitches, illumination with UV light provides a convenient option for activating fluorophores. Multiple modification options enable functionalization of various kinds of particles with SPs allowing investigation of their structure or organization at sub-diffraction-limit resolution.185–188
Figure 19.
Photoswitching reactions of spiropyran (SP, top) and oxazine-auxochromes (OA, bottom).
Figure 20.
70-nm spiropyran nanoparticles as imaged by conventional fluorescence microscopy (top) and SMLM (bottom). Reproduced with permission from Ref. 112, Copyright 2008 American Chemical Society.
Oxazine-auxochromes (OAs) are fluorophore dyads that, like SPs, exhibit intramolecular spirocyclic ring-open/close reactions that induce a large spectroscopic shift or fluorescence on/off switching suitable for SMLM.189–191 The oxazine moiety of the dyad acts as a molecular switch that can modulate the fluorescence of a range of appended fluorophores. Deniz et al. and Petriella et al. reported OAs in which the appended fluorophores include coumarin, pyrene, fluorene, BODIPY, or cyanine fluorophores and all but the oxazine-cyanine dyads existed primarily in the ring-closed form. 189–193 Among these, the oxazine-coumarin OA-1 (Figure 19 bottom) exhibited a good fluorescence quantum yield and reversible switching sufficient for SMLM on nanoparticles189 and immunolabeled cells.191 Based on their modular and easily tunable design, OAs are important photoswitchable compounds that with further development may emerge as powerful tools for SMLM.
3.1.4. 1,2-dithienylethenes
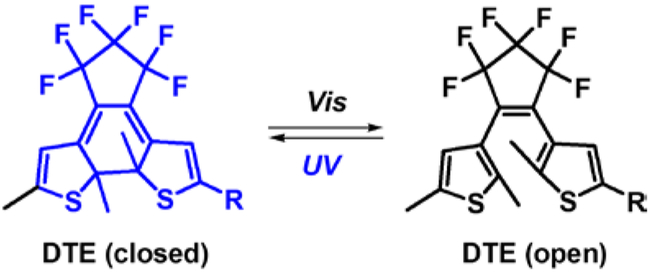
1,2-dithienylethene (DTE) fluorophores are another classic class of photochromic compounds with excellent fatigue resistance (many switching cycles before degradation), fast response, good thermal stability, and large spectral shift between the two isomers, which are in general desirable characteristics for SMLM.194–196 DTEs undergo photo-isomerization reactions between ring-open and ring-closed states accompanied with de-coloration and coloration (Figure 21) that occurs differently from that of rhodamine-spiro dyes or spiropyran cyclization. For the two latter dyes, UV irradiation cleaves one covalent bond on the spiro-ring, inducing a ring-opening reaction, and restoring the π-conjugation system to the colored state. For DTE dyes, the ring-open form shows no absorption in the visual region. UV irradiation enables the formation of a C-C bond between two thiophene moieties to produce the ring-closed form of DTE which typically absorbs in the visible range and for a subset of DTE variants is fluorescent.197, 198
Figure 21.
Photoswitching of DTE based on light-induced ring-opening/closing.
DTEs are useful as reversible photoswitchable fluorescence quenchers in quencher-reporter dyads. This is achieved by linking a DTE molecule to a fluorescent reporter molecule, such as naphthalimide,199 perylenemonoimide,200 perylenebisimide201 or tetraphenylethene,202 whose fluorescence emission spectral range of ~500 nm to ~700 nm overlaps well with the absorption spectrum of the closed-ring form of DTE. Photo-isomerization of DTE switches the linked fluorophore or aggregation-induced emission luminogen202,203 between on/off states based on intramolecular energy transfer, and this on/off switching facilitates SMLM.199,200,202 Strategic modification can provide DTEs with good emission properties,197,198 enabling their use in SMLM.
More recently, Roubinet et al. reported the synthesis and use of new hydrophilic DTEs for SMLM of biological specimens using antibody-conjugated DTEs. In this approach, DTEs are directly used as a photoswitch where they switch transiently between fluorescent and non-fluorescent states.204
3.1.5. Dicyanodihydrofurans
Dicyanodihydrofuran (DCDHF)-based fluorophores have large extinction coefficients, high fluorescence quantum yields, and good photo stability;205–207 through combination with caging and activation strategies, DCDHFs have been used in SMLM. DCDHFs are strong electron acceptors when combined with a conjugated unit (e.g., benzene, thiophene, alkene, styrene) and when linked to an electron donor (e.g., amine, hydroxyl) a typical D-π-A (donor—acceptor) system will be constructed. This electron pull-push character offers DCDHFs a bright fluorescence emission with adjustable spectral properties through the use of various substitutions, although these and other properties are relatively sensitive to the local environment (e.g., viscosity, polarity, pH).208,209
Several methods have been developed to switch DCDHFs off and on for use in SMLM. The DCDHF D-π-A system can be blocked by replacing the amine with azido group, a mildly electron-withdrawing group, to produce a non-fluorescent, caged DCDHF molecule. Generally, the aryl azide groups are able to be photoactivated to release one nitrogen molecule and give an aryl nitrene, which is prone to undergo rearrangement to seven-membered azepine heterocycles.210 However, in DCDHFs, a high electron-deficiency will stabilize the aryl nitrene and preferentially convert the nitrene to an amine or amino derivative upon reaction with the surrounding molecules.210,211 The photoconversion process of azide to amine restores the D-π-A system as well as its fluorescence emission (Figure 22).212,213 This can be a general strategy to construct photoswitching in other D-π-A fluorophores.211 The photoactivated process in DCDHFs system is irreversible, like the above-mentioned DMNB or diazo caged rhodamines; once activated, the fluorophore cannot revert to its dark state. This irreversible uncaging property is useful in SMLM since it can result in more photons being collected in a single burst rather than several separate small bursts (that in general cannot be linked to the same molecule) so that a superior localization precision can be obtained for each detected molecule. Lee et al. reported an azido-chloroalkane-DCDHF probe and used it to label Halo-tagged proteins (see Section 6.2) and to image protein distributions by SMLM.213
Figure 22.
Example DCDHF fluorophores (top) and fluorogenic reaction of azido-DCDHFs based on photoactivated conversion of caging groups.
3.1.6. Summary of commonly used organic fluorophores
A wide range of organic fluorophores for SMLM are commercially available or have been reported in the literature, and we believe it is critical for the field that SMLM fluorophore developers report key SMLM fluorophore properties including photons/localization, duty cycle, and number of cycles. Along these lines, Dempsey et al. compared 26 commercially available organic fluorophores for SMLM using single-molecule measurements of photoswitching properties with dyes on antibodies as well as SMLM imaging of standard specimens.128 They identified superior fluorophores in each of four spectral bands for low-crosstalk multichannel imaging (ATTO 488, Cy3B, Cy5 or Alexa Fluor 647, DyLight 750 or Alexa Fluor 750) and showed that all exhibited good performance in a deoxygenated solution containing a thiol (see Section 3.4 on imaging cocktail).128 Bittel et al. showed that unconjugated fluorophores embedded in polyvinyl alcohol film behave similarly to antibody-conjugated fluorophores in an aqueous solution, thus paving the way for a linker-free fluorophore evaluation that could save time-consuming functionalization steps when evaluating newly synthesized fluorophores.214 That study evaluated several of the same fluorophores evaluated in the Dempsey et al. paper as well as some that were not, including BODIPY FL.
We have included here a summary of switching properties of selected organic fluorophores (Table 1) which shows duty cycle, number of detected photons per localization, and number of switching cycles before bleaching, by thiol-,128 TCEP-based switching,135 or reductive caging136,147 methods. For example, using thiol-based switching, many fluorophores have a good duty cycle, but Alexa Fluor 647 also emits a large number of photons per localization and exhibits a high number of cycles, and Alexa Fluor 647 therefore remains the most popular choice to date. On the other hand, the fluorophore Cy3B works quite well reductive caging for approximately one photoswitching event and a very high number of detected photons per localization.
Table 1.
Switching properties of selected organic fluorophores for SR-imaging by SMLM.
| Organic dye | λex/ (nm) | λem (nm) | λon/off (nm) | ε (M−1cm−1) | QY | Duty cycle | # Photons | # Cycles | Refs |
|---|---|---|---|---|---|---|---|---|---|
| Alexa Fluor 647a,f | 650 | 665 | 405/647 | 239,000 | 0.33 | 1.2×10−3 | 5,202 | 26 | 128 |
| Alexa Fluor 647c,f | 650 | 665 | 405/647 | 239,000 | 0.33 | 1.3×10−4 | ~2400 | NR | 135 |
| Alexa Fluor 750a,f | 749 | 775 | 405/750 | 240,000 | 0.12 | 1×10−4 | 703 | 6 | 128 |
| Alexa Fluor 750c,f | 749 | 775 | 405/750 | 240,000 | 0.12 | 4×10−4 | 2800 | NR | 135 |
| ATTO 488a,f | 501 | 523 | 405/488 | 90,000 | 0.8 | 2.2×10−3 | 1,110 | 49 | 128 |
| ATTO 488b,g | 501 | 523 | 405/488 | 90,000 | 0.8 | 2×10−3 | ~104 | ~1 | 128,136 |
| BODIPY FLa,g | 503 | 512 | 405/488 | 80,000 | 0.97 | 1.8×10−2 | 756 | 3.4 | 214,215 |
| Cy3Ba,f | 559 | 570 | 405/561 | 130,000 | 0.67 | 4×10−4 | 2,057 | 5 | 128 |
| Cy3Bb,g | 559 | 570 | 405/561 | 130,000 | 0.67 | 3×10−3 | ~105 | ~1 | 128,136 |
| Amino-DCDHFe,i | 594 | 613 | 407 | 54,100 | 0.39 | NR | 2.3×106 | 1 | 212 |
| DTE (4-Et)e,f | 488 | 582 | 488 | 37,300 | 0.45 | 4×10−4 | 200-300 | 14 | 204 |
| Dyomics 654a,f | 654 | 675 | 405/647 | 220,000 | NR | 1.8×10−3 | 3,014 | 19 | 128 |
| DyLight 750a,f | 752 | 778 | 405/750 | 220,000 | NR | 2×10−4 | 749 | 6 | 128 |
| ER-Tracker Reda,h | 587 | 615 | 405/561 | 64, 300 | NR | 3×10−4 | 820 | 28 | 216 |
| HMSiRe,g | 650 | 672 | NR | 100,000 | 0.39 | NR | 2,600 | ≥20 | 164 |
| PA-JF549d,h | 561 | 571 | 405 | 101,000 | 0.88 | 1×10−5 | 1.9×104 | 1 | 156,168 |
| PA-JF646d,h | 637 | 664 | 405 | 152,000 | 0.54 | 1.6×10−6 | NR | 1 | 156,168 |
| OA-1 (open)e,j | 532 | 650 | 355/532 | 83,000 | 0.02 | NR | 222 | NR | 189 |
| SiRa,h | 645 | 662 | NR/640 | 100,000 | 0.39 | NR | 630 | NR | 154 |
| SP (open)e,j | 532 | 665 | UV/532 | 52,000 | 0.17 | NR | 1.8×105 | NR | 112,217 |
Note: λex (excitation wavelength); λem (emission wavelength); λon/off (wavelengths for switching the fluorescence on and off, respectively); ε (extinction coefficient); QY (fluorescence quantum yield); duty cycle (fraction of time spent in the on state prior to photobleaching); # Photons (number of detected photons per localization event); # cycles (number of switching cycles); Refs (journal references for published data).
indicate the composition of the imaging cocktail: a) deoxygenated + thiol; b) deoxygenated + MVAA (methyl viologen and ascorbic acid) where fluorophores had been previously reduced with NaBH4; c) deoxygenated + TCEP + MVAA; d) deoxygenated; and e) no additives to the aqueous buffer.
indicate the specimen used for fluorophore characterizations: f) fixed cells; g) antibody adsorbed to coverglass; h) live cells; i) fluorophores immobilized in PMMA (for quantum yield measurements) or gelatin (for other measurements); j) fluorophores on nanoparticles. ‘N.R.’ indicates the parameter was not reported.
3.2. Fluorescent proteins
FPs are a powerful class of fluorophores which have been widely utilized in SMLM since its inception. Recent articles have recently reviewed FPs for SMLM19,59,65,218,219 and we provide here a relatively brief summary with a few key details. A major benefit of FPs is that they are genetically encoded and do not require addition of exogenous ligands, thus enabling labeling of target proteins with high specificity and high efficiency (potentially stoichiometric). FPs are also compact, with molecular weights near ~30 kDa and a size of ~3-4 nm across, so there is minimal linkage error between the chromophore and the target. In terms of switching properties, FPs for SMLM tend to be substantially dimmer than the preferred organic fluorophores, although many of the preferred FPs have duty cycles in the range of 10−4-10−6 which are over 10-100 times lower (i.e., better) than that of the preferred organic fluorophores (Table 1). Finally, many FPs tend to dimerize or aggregate, leading to artifacts in protein localization, and researchers have invested considerable time to design probes that are sufficiently monomeric.
Based on their switching properties, FPs for SMLM may be divided into three classes: irreversible photoactivatable FPs (PA-FPs), irreversible photoconvertible FPs (PC-FPs), and reversible photoswitchable FPs (PS-FPs). The representative photoswitching mechanisms are illustrated in Figure 23 and a summary of photoswitching properties of selected FPs are shown in Table 2.
Figure 23.
Representative photo-modulation mechanisms of FPs (only the FP chromophores are shown). A) Photoactivatable FPs (PA-FPs) such as PA-GFP convert irreversibly from a native dark from to a green fluorescent form by photo-induced decarboxylation of a glutamate residue (pink arrow) near to the chromophore. B) Photoconvertible FPs (PC-FPs) such as EosFP convert irreversibly from a green fluorescent form to a red fluorescent form upon illumination with violet or ultraviolet light. The photochromic shift occurs due to cleavage of the peptide backbone which extends the conjugation of the chromophore and red-shifts the absorption and emission spectra by 50–60 nm. C) Photoswitchable FPs (PS-FPs) such as Dronpa undergo reversible cis–trans isomerization, where the cis form is fluorescent and the trans form is dark. Note that certain FPs such as PAmCherry 220 and IrisFP 221 may use a combination of these switching mechanisms.
Table 2.
Photo-switching properties of selected FPs for SR-imaging by SMLM.
| FPs | λex (nm) | λem (nm) | λon/off (nm) | ε (M−1cm−1) | QY | on/off contrast |
Duty Cycle |
Photons/ localization |
# Switching Cycles |
clustering | Ref. |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Dendra2 | 490/553 | 507/573 | 405/488 | 45,000/35,000 | 0.5/0.55 | 300 | 4×10−6 | 686 | 2.7 | - | 56,62,63,240 |
| Dreiklang | 511 | 529 | 340/412 | 83,000 | 0.41 | 75 | N.R. | 400-700 | >160 | N.R. | 61,237,241 |
| Dronpa | 503 | 518 | 405/488 | 95,000 | 0.85 | 20 | 8×10−4 | 262 | 60-70 | - | 25,63,236,242 |
| mEos2 | 506/573 | 519/584 | 405/573 | 56,000/46,000 | 0.84/0.66 | 200 | 3×10−6 | 1000 | 2.8 | + | 62,63,227,228 |
| mEos3.2 | 507/572 | 516/580 | 405/572 | 63,400/32,200 | 0.84/0.55 | N.R. | 3×10−6 | 1000 | 2.4 | - | 62,63,229 |
| mMaple | 489/566 | 505/583 | 380/566 | 15,000/30,000 | 0.74/0.56 | 400 | 2×10−6 | 1000 | 3.4 | + | 62,63,228 |
| mMaple3 | 489/566 | 506/583 | 405/561 | N.R. | N.R. | N.R. | 6×10−7 | 675 | 2.8 | - | 63 |
| PAmCherry1 | 564 | 595 | ~370 | 18,000 | 0.46 | 4000 | 8×10−6 | 725 | 1 | + | 62,63,223 |
| PAGFP | 504 | 517 | 405 | 17,400 | 0.79 | 200 | 1.3×10−3 | 313 | N.R. | - | 49,63 |
Note: λex (excitation wavelength); λem (emission wavelength); λon/off (wavelengths for switching the fluorescence on and off, respectively); ε (extinction coefficient); QY (fluorescence quantum yield); on/off contrast (ratio of intensity of the emissive form to the dark form); duty cycle (fraction of time spent in the on state); photons/localization (number of detected photons per localization event); # switching cycles (number of switching cycles); ‘N.R.’ indicates the parameter was not reported;‘+’ indicates clustering or aggregation was observed in the ClpP assay by Wang et al.63 (‘-’ indicates no clustering).
The first PA-FP, PA-GFP, was reported by Patterson et al.49 Illumination at 405 nm shifts its absorption peak from 400 nm to 504 nm, with a 200-fold fluorescence enhancement when excited at 488 nm (Figure 23A, Table 2). van Thor et al. used X-ray crystallography to determine that the photoactivation process results from the light-induced decarboxylation of a glutamate residue near to the chromophore which irreversibly converts PA-GFP from a dark, neutral cis form to a brightly fluorescent, anionic cis form (Figure 23A).222 The photoactivated form of PA-GFP shows good stability at physiological temperature and pH environment,49 accompanied by good overall brightness and fusion protein compatibility. PA-GFP, as well as other FPs, were utilized by Betzig et al. in their original 2006 paper.27
PA-FPs can implement fluorogenic emission in the green (PA-GFP), red (PA-TagRFP,223 PA-mCherry1220 and PA-mRFP-154) and far-red (PA-mKate224) channels, allowing multi-color SR imaging by SMLM. Subach et al. implemented two-color SMLM imaging using PA-GFP tagged clathrin light chain (CLC) and PAmCherry1 tagged transferrin receptor (TfR) in COS-7 cells to show the nanoscale distributions of CLC and TfR (Figure 24A).220 PA-mKate shows far-red shifted fluorescence emission and may be used in three-color SR imaging when paired with other green and red emitting PA-FPs. Gunewardene et al. performed three-color SMLM in cells co-expressing PAmKate-TfR, PAmCherry1b-actin, and Dendra2-hemagglutinin (HA, Figure 24D).224 Their results revealed correlations between these membrane proteins and membrane-associated actin structures: TfR and HA are both correlated with actin but are either uncorrelated or anticorrelated with each other. Like photoactivatable organic dyes, the best PA-FPs have several advantages including a low fluorescence background, a low duty cycle, and in some cases a single burst of photons. However, PA-FPs are invisible prior to photoactivation so that it may be difficult for researchers at the microscope to select transformed cells exhibiting the phenotype of interest. This issue is addressed by using PC-FPs.
Figure 24.
Multicolor SMLM using fluorescent proteins. A) Two-color SMLM in COS-7 cells showing (i) PA-GFP-tagged clathrin light chain (green), (ii) PAmCherry1-tagged transferrin receptor (red), and (iii) overlay. B) Two-color SMLM of mitochondria in EpH4 cells showing outer membrane labeled with (i) PAmCherry1-Lk-BclXl (red), (ii) inner membrane labeled with BCS1L-Lk-rsKame (green), (iii) two-channel overlay, and (iv) intensity profile along white line in iii. C) Two-color SMLM of tdEos-tagged paxillin (green) and PS-CFP2-tagged zyxin (red) in an HFF-1 cell reveals little overlap despite appearing to co-localize using conventional microscopy. D) Three-color SMLM image of Dendra2-hemagglutinin (cyan), PAmCherry1b-actin (yellow), and PAmKate-transferrin receptor (magenta) in fixed HAb2 mouse fibroblast cells. Scale bars are all 500 nm except as indicated otherwise. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 220, Copyright 2009. Adapted with permission from Ref. 231, Copyright 2014 National Academy of Sciences, U. S. A. Adapted with permission from Ref 232, Copyright 2007 National Academy of Sciences, U. S. A. Adapted from Ref. 224, Copyright 2011, with permission from Elsevier.
PC-FPs are generally photoconverted from one fluorescent state to another by violet or blue light irradiation. The photoconversion process is attributed to the photo-cleavage of the peptide backbone on histidine, which creates a new double bond formation between its Cα and Cβ atoms and thereby extends the π-electron conjugation from the chromophore to the histidine imidazole ring (Figure 23B).225,226 For example, EosFP57 exists initially in a green fluorescent state and upon illumination with UV or near-UV light will convert to a red fluorescent form (Figure 23B), permitting on/off and off/on switching in green and red channels, respectively. The activation light (~400 nm) is well separated in wavelength from the excitation wavelength (~510 nm for the green channel and ~570 nm for the red channel), which reduces the risk of unintended photoconversion.
Many PC-FPs are popular in SMLM including EosFP, mMaple, Dendra and subsequent variations, due their high on-off contrast, low duty cycle, and relatively large number of detected photons per localization (Table 2).56,62,63,227–230 mEos2 and mEos3.2 were engineered to have a reduced tendency to dimerize or aggregate while still retaining the spectroscopic properties of EosFP.57,227,229 mMaple2 also shows minimal aggregation with good switching properties while exhibiting a superior efficiency of expression/folding/maturation.63,228 mEos4 was developed to be able to survive fixation conditions used for electron microscopy (EM) and thus nicely enables correlative SMLM and EM.230 A major limitation of PC-FPs, however, is their dual-channel emission property; by converting from a green fluorescent form to a red one, two fluorescent channels are occupied, creating serious challenges for multi-channel imaging with PC-FPs.
PS-FPs show reversible on/off photoswitching between fluorescent and nonfluorescent states. This process is reminiscent of reversible thiol-based photoswitching of organic dyes, however, special reagents (e.g., a deoxygenated solution containing a thiol, Section 3.4) are not generally required to switch PS-FPs. Dronpa50 and its variants 233,234 are among the best known examples of PS-FPs. Intense irradiation with ~488 nm light converts Dronpa from an anionic cis form that is fluorescent to a neutral trans form that is nonfluorescent, while illumination of the nonfluorescent form with ~405 nm light converts it back to the fluorescent form (Figure 23C). This deactivation/activation cycle can be repeated many times and allows Dronpa’s application to single-molecule localization techniques and SMLM.232,235,236 For instance, Rosenbloom and co-workers paired the Dronpa variant rsKame with PAmCherry1 to capture two-color SMLM images of the inner and outer mitochondrial membranes (Figure 24).231 Other Dronpa variants, like bsDronpa,234 Padron234 and rsFastlime233 have been developed which show markedly improved photostability (8.5 to 34-fold) compared with Dronpa.
The PS-FPs, Dreiklang237 and reversibly switchable enhanced GFP (rsEGFP)238 reported by Brakemann, Grotjohann and co-workers also have good fatigue resistance, allowing photoswitching for more than 160 and 1000 cycles, respectively. Among them, Dreiklang shows a larger number of detected photons per localization than Dronpa (Table 2). It is worth mentioning that the fluorescence excitation of Dreiklang is decoupled from its photoswitching. Light irradiation around its excitation maximum (511 nm) does not activate dark Dreiklang, thereby avoiding unintended deactivation or activation and enabling control over the length and probability of a photoswitching cycle. However, Dreiklang is sensitive to temperature, exhibiting poorer fatigue resistance and duty cycle at higher temperature.
There are yet other classes of fluorescent proteins, including some which exhibit a combination of photoconversion and photoswitching behaviors. For instance, mIrisFP is photoconvertible between green and red fluorescent states while both these two states exhibit reversible photoswitching between on and off forms.221 This dual PC/PS mode of mIrisFP enables the combination of pulse-chase experiments with SMLM to implement dynamic studies of protein complexes.239
Although FPs are generally dimmer compared with the preferred organic dyes,128 their high compatibility and specificity of labeling makes them particularly useful for living specimens.59 Finally, it should be mentioned that expression of exogenous protein tags raises several issues. For instance, overexpression or oligomerization/aggregation may perturb the target protein or specimen, and their use may be impacted by species-specific codon usage, maturation time of the fluorescent tag, and changes in the expression levels of the fusion protein (see Section 9.2).63
3.3. Fluorescent quantum dots
Quantum dots (QDs) are nanoscale semiconductors whose fluorescence emission spectra are tunable by varying particle size and composition (Figure 25).243 Compared with most organic fluorophores or fluorescent proteins, QDs generally exhibit intense brightness244 and resistance to photobleaching,245 both of which are valuable general properties for fluorescence imaging, but face challenges due to their relatively large size and limited cell permeability for live-cell applications. While spontaneous QD blinking is a well-known phenomenon that had been reported approximately a decade before the first SMLM publications39, the duty cycle of spontaneously blinking QDs tends to be quite high and the switching rates rather heterogeneous, making the fluorophores unfavorable for SMLM. Nonetheless, Lidke et al.72 in 2005 and Lagerholm et al.246 in 2006 both demonstrated the identification of two QDs within a diffraction-limited region.
Figure 25.
Quantum dot emission spectra are tunable through control of A) particle size and B) particle composition. Reproduced with permission from Ref. 243, Copyright 2015 American Chemical Society.
Spontaneously blinking QDs have been used with good results for super-resolution optical fluctuation imaging (SOFI, Section 2.5),133 however, SOFI attains improved spatial resolution through statistical analysis of signal fluctuations (an ensemble measurement) rather than requiring observation of individual fluorophores. In the original SOFI publication,133 Dertinger and co-workers reported ~55 nm resolution (FWHM) using commercially available quantum dots consisting of CdSe cores surrounded by ZnS shells. Subsequent researchers were able to tune the blinking properties either by synthesis of QDs with a thinner ZnS shell247 or by addition of ascorbic acid in solution in order to increase the blinking rate.248 Through a variation of SMLM called QDB3 (Section 2.5), Wang et al. were able to use the spontaneous blinking of quantum dots to study cell surface receptors on breast cancer cells.130
Hoyer and co-workers exploited stochastic ‘blueing’ (or blue-shifting) of QDs to enable use of QDs in a low duty cycle mode that is suitable for SMLM.249 In QD blueing, the CdSe cores gradually oxidized from the outside inwards such that the emission wavelength shift to shorter and shorter wavelengths over time until the CdSe core is oxidized and becomes nonluminescent. By this means, a detection band between the excitation wavelength and the unoxidized QD emission allows observation of individual QDs as they are stochastically blue-shifted but before irreversible bleaching. Tehrani et al.250 and Xu et al.251 extended the application of QD blueing to multi-color SMLM and 3D-SMLM. As an illustration in Figure 26, QD565 and QD705 were used for studying the morphology and localization of microtubules and mitochondria. Their work also demonstrated that the rate of QD blueing can be tuned and optimized by modulation of laser intensity and the mounting medium (oxygen concentration), resulting in better localization data for SR image reconstruction. Alternative strategies for producing photoswitchable QDs with low duty cycle that are potentially well-suited to SMLM include using chemical caging,252 light-driven modulation,253 or FRET when decorated with a photoswitchable organic fluorophore.254
Figure 26.
Two-color SMLM using QD blueing (stochastic photoconversion). A-B) Widefield and corresponding SMLM fluorescence image of a fixed HepG2 cell immunostained for microtubules with QD565 (cyan) and for mitochondria with QD705 (red). C-F) Zoom-in views of boxed regions from A and B. Scale bars are 2 μm (A-B) and 500 nm (C-F). Reproduced with permission from Ref. 251, Copyright 2015 American Chemical Society.
3.4. Imaging cocktails
Fluorophores are sensitive to their chemical environment, and it is common in SMLM to use various “imaging cocktails” in order to facilitate or enhance photoswitching properties of a given fluorophore. While a range of imaging cocktails have been reported in the literature, including several fluorophore-specific examples described above, we will confine our comments to a few limited examples below that are of particular importance or special interest.
Dempsey et al. showed that a wide range of commercial organic fluorophores photoswitch well in deoxygenated solutions containing a thiol and a buffer (e.g., 100 mM Tris pH 7.4).128 Deoxygenation is often achieved enzymatically by addition of a mixture of glucose oxidase and catalase255, along with glucose (the substrate of glucose oxidase), to convert glucose and dissolved oxygen into gluconic acid while lowering the concentration of dissolved oxygen. Several alternative oxygen scavenging systems have been reported in the SMLM and single-molecule fluorescence literature, including the use of other enzymes such as protocatechuate deoxygenase, galactose oxidase, pyranose oxidase, and the proprietary enzyme Oxyrase, the latter two of which are notable for not leading to acidification of the imaging solution.256–259 Oxygen may be depleted nonenzymatically, as well, for instance by using a solution containing methylene blue and a thiol.260 Similarly, a range of thiols have been used, including 2-mercaptoethanol, 2-mercaptoethylamine, cysteine-methyl-ether, and others.76,77,128,141 Live-cell SMLM with photoswitchable organic fluorophores requires additional care, particularly in formulation of oxygen scavenger systems, because the high concentrations of glucose (~10%) used with the popular glucose oxidase and catalase oxygen scavenger system can cause osmotic shock. For example, Jones et al. and Shim et al. used a modified oxygen scavenger system containing ~2% glucose for live-cell SMLM.216,261 Conveniently, reduced glutathione is present in living mammalian cells at a concentration of ~0.5-10 mM262 and provides a thiol for photoswitching.
Many other imaging cocktails have been used with photoswitchable SMLM fluorophores. Olivier et al. showed that addition of the triplet state quencher cyclooctatetraene to deoxygenated solutions containing a thiol could boost the number of photons detected per localization for Alexa Fluor 647 by up to 3.5×, although it was not clear whether the low duty cycle was maintained under these conditions. In a separate report, Olivier et al. also showed that the proprietary mounting agent VECTASHIELD, alone, was able to produce good photoswitching from several cyanine dyes, including Alexa Fluor 647, without requiring additional reagents (e.g., enzymes or thiols), although its use is apparently incompatible with 405 nm activation of dark-state fluorophores due to background fluorescence from the violet-colored solution.263 Lee et al. and Klehs et al. found that heavy water (D2O) moderately boosted photon yields or emission rates for a range of cyanine and oxazine fluorophores in SMLM.264,265 The use of TCEP, together with an oxygen scavenger system and a mixture of methyl viologen and ascorbic acid, was described above for the switching of the cyanine dyes Cy5 and Cy7 (Section 3.1.1, Figure 10A, Figure 9C).135
While fluorescent proteins tend to be less sensitive to their chemical environment than small organic fluorophores, several imaging cocktails have been reported in the literature in order to tune fluorescent protein switching properties for SMLM, including: the boosting of photon yields of several fluorescent proteins by up to ~50% by imaging in heavy water266; the chemical caging of mCherry and some other red fluorescent proteins using a thiol and its subsequent imaging (uncaging) with or without the thiol present267; and the photoconversion of eGFP to a red absorbing/emitting form when illuminating eGFP in the presence of riboflavin.268 Although specialized imaging cocktails are not widely used in SMLM with fluorescent proteins at present, this area seems ripe for further exploration given the wide range of fluorescent proteins and associated sensors that have been created over the past two decades.
In summary, a deoxygenated solution containing thiol and buffer works well with SMLM for a broad variety of photoswitchable organic fluorophores while specialized imaging cocktails are not generally required for SMLM with fluorescent proteins, spontaneously-blinking switches such as HMSiR, or binding-based switches used in DNA-PAINT (see Section 4). Live-cell imaging, on the other hand, requires additional care.
4. Fluorophore switching by binding-based methods (PAINT, DNA-PAINT, etc.)
The visualization of sequential binding of fluorescent probes is known as PAINT, or Point Accumulation for Imaging by Nanoscale Topography, is a SMLM method with unique properties compared to those in which the fluorescent probes begin bound. PAINT was first reported in 2006 by Sharonov et al.29 and was based on related work published during the prior year by the same group.97,269,270 In its initial implementation, PAINT was used to image large unilamellar vesicles (LUVs) with the fluorophore Nile red, which is weakly fluorescent in water but becomes much brighter in a lipid environment. Transient binding events of Nile red to LUVs created brief flashes similar to blinking behavior in photoswitchable or photoactivatable fluorophores and over time were able to provide super-resolution information on the vesicle positions and diameters (Figure 27). Nile red has also been used to study supported lipid bilayers and cell membranes,29,119 while another turn-on probe, merocyanine 540, has been used to study phase segregation in supported lipid bilayers.271
Figure 27.
SMLM imaging using PAINT (Point Accumulation for Imaging by Nanoscale Topography). A) Conventional fluorescence image and B) corresponding SMLM image of vesicles bound to a coverglass acquired by imaging flashes of light caused by binding of individual molecules of the fluorophore Nile red. Reproduced with permission from Ref. 29, Copyright 2006 National Academy of Sciences, U. S. A.
The PAINT approach for SMLM, in which fluorophores are initially diffusing in solution and bind stochastically to a specimen during acquisition of a stack of single molecule images, has been extended to a wide range of probes and specimens. In universal PAINT, or uPAINT, small proteins of interest were probed with nonblinking fluorophores bound to a ligand or antibody with affinity for the target protein.272 This allowed the tracking of membrane receptors over a relatively long time in live cells and resembles sptPALM experiments performed using photoactivatable or photoswitchable fluorescent proteins.111 In interface PAINT, or iPAINT, Aloi et al.273 coupled a photoactivatable rhodamine fluorophore167 to polyethylene glycol in order to develop an interface-binding probe for visualization of solid/liquid, liquid/liquid, and liquid/air interfaces at nanometer scale resolution. Kiuchi et al. developed small, protein-based PAINT probes, which they called IRIS probes, in order to avoid the use of bulky antibodies which tend to broaden labeled structures of interest.274 For instance, the actin-binding probe LifeAct275 was coupled to ATTO 488 and was used to image actin in permeabilized cells, and a number of other protein fragments were created for use as IRIS probes starting from a careful review of known protein-protein interactions.274 Similar sequential binding approaches have been used for nonbiological specimens including to record images of polymeric nanogratings and to map out electric field hot spots near nanoparticles.276,277
The SMLM variant DNA-PAINT uses reversible binding of diffusing fluorescently labeled single-stranded DNA probes to image structures that are decorated with complementary DNA strands.278 Unbound probes diffuse in solution, appearing as a background haze, while immobilized probes exhibit a bright spot. The rates of binding and unbinding are easily tuned, where the binding rate is primarily controlled by adjusting the imager strand concentration and the unbinding rate is primarily controlled by the length and base pair content of double-stranded DNA formed upon hybridization. Virtually any bright, photostable fluorophore may be used in DNA-PAINT, since it doesn’t require fluorophore photoswitching or photoactivation, and the number of localizations attainable is only limited in principle by the amount of time the user is willing to wait. DNA-PAINT has been used to image DNA origami structures as well as cellular specimens labeled with single-stranded DNA-antibody conjugates (Figure 28, Figure 39).279–285
Figure 28.
SMLM imaging using DNA-PAINT. A) DNA origami structure designed to spell ‘Wyss!’ within an area of ~50 nm × 75 nm. B) Representative SMLM image of a single Wyss! DNA origami and C) a class average obtained from a collection of about 80 imaged origami. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 281, Copyright 2016.
Figure 39.
Four-color Exchange-PAINT image of a cell that has been immunostained for tubulin, mitochondria, peroxisomes, and the Golgi apparatus. Scale bar is 5 μm. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 279, Copyright 2014.
PAINT, and several variations, thereof, have delivered promising results for SMLM but have typically faced a tradeoff between speed of imaging and high background signal. The binding rate in PAINT, or kon, is a diffusion-limited process whose rate primarily controlled through choice of concentration of the probe, while the unbinding rate, or koff, is controlled through the strength of the interaction of the probe with its target. Speeding up the data acquisition rate requires an increase in the diffusing probe concentration along with a decrease in the affinity of the probe for its target (e.g., for DNA-PAINT, a shorter imager strand). However, doing so provides less time during which to acquire signal from the bound probe, while also increasing the background signal resulting from the higher concentration of diffusing, unbound probes. In many implementations of PAINT, acquisition frame rates of ~20 Hz were used, which can be prohibitive when many frames of data are desired.
One strategy to decrease background signal with PAINT and DNA-PAINT is to utilize illumination of a thin volume. When a specimen is situated very close to the coverglass (<200 nm), total internal reflection illumination may be used to confine the excitation volume to be limited to a 100-200 nm thick layer above the coverglass. Imaging of specimens which are located substantially higher than 200 nm requires widefield illumination of a much larger thickness which in turn leads to more background signal. Legant et al.126 reported a potential solution to the imaging of thick specimens by combining PAINT with lattice light sheet (LLS) microscopy286 using commercially available and novel fluorescent probes (Figure 29).286 In this arrangement, the LLS excited an approximately 1 μm thick plane while repeatedly scanning thick samples (~5-20 μm thick) back and forth along the axial dimension of the detection path. A frame acquisition rate of ~40 Hz was used, although it required relatively long overall acquisition times of several days per image; shorter acquisition times would be possible with thinner specimens. This work used both commercially available fluorescent probes (BODIPY TR methyl ester) as well as custom-synthesized rhodamine probes (AzepRh, JF646-Hoechst) that had suitable affinity for membranes (BODIPY TR, AzepRh) or DNA (JF646-Hoechst) to enable PAINT-based SMLM.
Figure 29.
A) Volume rendering of an entire dividing LLCPK1 cell imaged by LLS-PAINT. The plasma membrane was labeled with a wheat germ agglutinin-Alexa Fluor 555 probe and intracellular membranes were labeled with BODIPY TR methyl ester. Bounding box is 50 μm × 27 μm × 20 μm. B) Cutaway section through the cell in A revealing various labeled membrane structures and histones (mCherry-H2B, imaged by diffraction-limited imaging prior to LLS-PAINT). Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 126, Copyright 2016.
A second strategy to decrease background signal would be to utilize turn-on probes that emit little to no signal when freely diffusing but which become much brighter when transiently bound. Nile red and merocyanine 540 exhibit behavior of this sort but are limited to very specific membrane targets. In DNA-PAINT, it was reported that the fluorescence of ATTO 655 on unbound imager strands is partly quenched relative to the bound state and exhibits 70% increase of fluorescence brightness upon binding.278 If a quenching strategy could be developed in DNA-PAINT that would enable much higher turn on ratios of, for instance 1:100, it would help reduce background levels and/or speed up data acquisition rates. Turn on ratios of more than 1:10,000 were achieved with the fluorogen malachite green when bound to engineered fluorogen-activating single chain antibody tags,287 and with further engineering these288 or similar approaches may help improve background and or frame rates in PAINT. Auer et al. recently implemented DNA-PAINT with a FRET-based scheme designed to reduce background signal in which an antibody-tethered acceptor fluorophore would only emit signal when a diffusing donor-labeled oligonucleotide bound to a complementary DNA sequence in proximity to the acceptor fluorophore.283
5. Fluorophore switching by catalysis, electrochemistry, or complexation
In addition to (photo)chemical reactions and binding/unbinding processes, stochastic switching of fluorophores can be achieved using catalytic reactions (enzymatic or otherwise), electrochemical reactions, or metal ion complexation.118,289–296 The timescale of switching can be tuned by controlling kinetic rates of the enzymatic, catalytic or electrochemical reactions, or by changing the concentration of the metal ion for complexation. These switching methods are compatible with SMLM of biological specimens and can be extended to other regimes including the mapping of reactivity patterns on single nano-catalysts.
A variety of synthetic substrates based on fluorescein,297 rhodamine,298 rhodol,261 and DCDHF123 fluorophores have been used to develop novel enzymatic fluorogenic reactions, due to the convenient incorporation of enzyme substrate groups on these fluorophores. For instance, Lee and co-workers123 developed a SMLM method based on enzymatic activation of DCDHF fluorophores which had been caged by a nitro group. In this scheme, nitroreductase (NTR) enzymes300 could be targeted to structures of interest such that diffusing nonfluorescent DCDHF substrate molecules could become reduced, activated, and imaged with high intensity excitation light that was used to localize and photobleach activated DCDHF fluorophores before they diffused very far from the NTR enzyme (Figure 30).123 Adjusting the concentration of nitro-DCDHF or slowing the enzymatic rate using inhibitors allows tuning of the rate of activation to produce a higher or lower density of fluorophores. A similar approach has been used in live eukaryotic cells by Halabi et al. who developed novel, non-fluorescent diazoindanone probes and imaged their activation to fluorescent rhodol fluorophores by endogenous cellular esterases in order to map enzymatic activity using SMLM.301
Figure 30.
A) Fluorogenic reactions of nitro-DCDHF based on enzyme-catalyzed conversion of caging groups. B) Conventional fluorescence image (and corresponding bright-field image, inset) of B. subtilis labeled with nitro-DCDHF. C) SMLM image of blue box area in B, showing localization information of the membrane and punctate spots. D) Temporal progression was revealed for the localizations in the punctate spots (blue dashed box area in C). E) SMLM image of green-boxed region in B. F) Enzyme diffusing was demonstrated in green dashed box area of E. Scale bars: 1 μm (B, C, E); 0.5 μm (D, F). Adapted from Ref. 123 with permission of The Royal Society of Chemistry.
The redox conversion between resazurin and resorufin has been used for a variety of catalytic or electrochemical switching experiments.293,296,302–305 Resazurin is a small nonfluorescent, organic molecule that can be reduced to resorufin, which is fluorescent, on an electrode or via N-deoxygenation by NH2OH in the presence of a gold catalyst. Xu et al. have used gold nanoparticles to reduce diffusing resazurin molecules to their fluorescent resorufin form and were able to show that the catalytic activity at sharp corners of gold nanoparticles was higher than that of edge regions which in turn was higher than surface facets (Figure 31).304
Figure 31.
A) Catalyst-enhanced conversion of nonfluorescent resazurin to fluorescent resorufin. B) SMLM image showing a density map of activated resorufin molecules relative to a triangular gold nanoplate. Adapted with permission from Ref. 304, Copyright 2013 American Chemical Society.
Rather than use a turn-on strategy, reversible binding of the fluorescence-quenching ion Cu2+ in close proximity to a fluorophore has been used to modulate emission of the fluorophore between emissive and nonemissive states.118,289,306 Tuning of the concentration of Cu2+ affords some control over the duty cycle of the fluorophore. In addition to Cu2+, other paramagnetic metal ions such as Ni2+, Co2+ or Mn2+ may also be candidates for metal ion complexation switching.
6. Sample labeling
One of the key advantages of fluorescence microscopy is the ability to visualize structures of interest with high molecular specificity and low background signal. This is achieved through a variety of schemes for labeling biological specimens that span the range of covalently or noncovalently binding small molecules, fluorescent proteins, antibodies, oligonucleotides, proteins or peptides. All of these approaches have been used with good results in SMLM, and in many cases, distinct advantages of the various labeling schemes have been powerful tools in enabling key experiments. In this section, we provide an overview of methods for sample labeling. We explicitly focus here on strategies to label specimens with organic fluorophores since labeling with fluorescent proteins is relatively standardized.
6.1. Immunostaining
Immunostaining is perhaps the most widespread technique used to label proteins with organic fluorescent dyes for SMLM in fixed specimens. The basic steps of immunostaining consist of fixation to preserve the structure(s) of interest, labeling of the specimen with dye-conjugated antibodies, and removal of unbound antibodies through washing.78 In practice, a great deal of optimization is often required to produce good immunostained specimens including the choice of fixative, antibodies, detergent, blocking agents, wash stringency, and a host of optional, but often important additional steps, such as pre-fixation extraction by treatment with detergent for studies of the cytoskeleton, reduction with sodium borohydride to quench (glutaraldehyde) fixation-induced fluorescence background, and postfixation with glutaraldehyde to help avoid antibody dissociation from the specimen. Practitioners most commonly use indirect immunofluorescence in which nonfluorescent “primary antibodies” bind a target on the specimen and are then labeled with a fluorescent “secondary antibody” that binds the primary antibody. While some labs conjugate fluorophores to (secondary) antibodies in house in order to achieve a specific dye/antibody ratio or another key property, many commercial sources of antibodies also work well. The fluorescent secondary antibodies can be mixed and matched for multichannel imaging137,138 and many practical guides are available.14,15,17,307–309 The use of indirect immunofluorescence with primary/secondary antibodies often limits the maximum number of channels to ~3 due to the requirement that each primary antibody is antigenically distinct for unique binding by secondary antibodies.
Importantly, the finite size of antibodies (~10 nm) leads to a spatial broadening of structures of interest that is measurable by SMLM. For instance, the binding of primary and secondary antibodies to microtubules, which have an outer diameter of ~25 nm, produces a broadened structure by SMLM that has a full-width at half maximum of ~50-60 nm and contains a ‘hollow’ structure due to the primary antibody and microtubule itself which are surrounded by the fluorescent secondary antibody layer.135–137,310 In some cases, particularly when there is bright signal from fluorescently-labeled primary antibodies without the need for amplification by secondary antibodies, the secondary antibody can be omitted and the amount of label-induced broadening may be reduced.310 The use of direct immunofluorescence with very small antibodies, such as camelid nanobodies which are ~2-3 nm in size, can further reduce label-induced broadening to a negligible amount (Figure 32)310,311 and nanobodies against an increasing variety of endogenous proteins and have been used in SMLM to probe, for example, tubulin,311 several components of the nuclear pore complex,312 and vimentin.313 Pleiner et al. reported anti-immunoglobulin G (IgG) nanobodies as compact secondary antibodies which also help to minimize antibody-induced broadening compared to full IgG-based secondary antibody labeling.314
Figure 32.
Antibody broadening of features for immunostained microtubules in fixed Ptk2 cells expressing tubulin-YFP. A) Indirect immunostain of tubulin (left), direct immunostain with anti-GFP nanobody on cells expressing tubulin-YFP (middle), and direct immunostain of YFP on cells expressing tubulin-YFP using an anti-GFP antibody (right). B) Cross-sectional profiles for boxed regions in A showing substantially narrower profile using the nanobody. C) Illustration of linkage error using either indirect immunofluorescence with full immunoglobulin G antibodies (left) or direct immunofluorescence with nanobody-based probes (right). Scale bars in A-C are 500 nm. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 310, Copyright 2012. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 309, Copyright 2015.
6.2. Protein fusion tags for labeling with organic fluorophores
Rather than use bulky antibodies to target organic dyes to proteins of interest, it is possible to express the protein of interest labeled with a protein tag that can covalently or noncovalently bind a small organic group.24,315–319 These methods also have the potential advantage of compatibility with living specimens, unlike antibody staining which in generally is only used for fixed specimens. Many tags and techniques have been developed for labeling of specific proteins with organic fluorophores, such as FlAsH-tag,320 TMP-tag,321 SNAP-tag,322 CLIP-tag,323 and Halo-tag324 (Figure 33). These protein tags are genetically encodable and have a small size, as do fluorescent protein tags, but allow researchers to harness the (often) superior optical properties of organic fluorophores for live- or fixed-cell imaging. Each has their own challenges regarding specificity, efficiency of labeling, speed of labeling, and their potential for perturbation of the specimen.
Figure 33.
Schematic illustration of representative techniques for targeting organic fluorophores to a tagged protein of interest, including A) FlAsH-tag (tetra-Cys tag); B) TMP-tag; C) SNAP-tag, CLIP-tag, and Halo-tag (Nu=nucleophile, F=fluorophore).
Biarsenical probes that bind tetracysteine motifs genetically introduced into proteins (Figure 33A) have been used in SMLM, both with the original fluorescein bisarsenic “FlAsH” probes320,325 as well as novel bisarsenical cyanine probes326. Figure 33B illustrates the interaction between E. coli dihydro-folate reductase (eDHFR) and trimpethoprim (TMP), which binds eDFHR with high affinity (KD ~1 nM) and high selectivity. This enables the use of TMP-linked fluorophores, in which a protein of interest that is fused with the eDHFR tag is labeled with a TMP-fluorophore conjugate.321 Various fluorophores have been used with TMP-tags, including the cell-permeable fluorophore ATTO 655 for labeling of eDHFR-tagged human histone H2B to study histone core protein dynamics by SMLM in live HeLa cells.327
Several covalent protein labeling strategies based on protein tags that react with specific small molecule ligands have been used in SMLM, including SNAP-tag,322 CLIP-tag,323 and Halo-tag (Figure 33C).324 SNAP-tag and CLIP-tag are modified alkylguanine transferase DNA repair proteins which can react specifically and efficiently with benzylguanine (BG) or chloropyrimidine substrates (SNAP-tag) or with O2-benzylcytosine substrates (CLIP-tag) by forming a stable, covalent thioether bond. These have been used in several live-cell super-resolution microscopy studies including by SMLM and STED.154,178,261,328 For instance, Jones et al. performed live-cell SMLM on clathrin coated pits using SNAP-tagged clathrin together with Alexa Fluor 647-BG, a top dye for SMLM (Figure 40).261 Importantly, since Alexa Fluor 647 is cell-impermeant, Jones et al. used electroporation to deliver the fluorophore to cells,261 although special measures are not necessary when using cell-permeant fluorophores. Halo-tag is another broadly used protein tag for specific labeling and imaging of proteins of interest. With HaloTag, the terminal chloride on the HaloTag ligand undergoes nucleophilic attack by Asp106 of the HaloTag protein and results in the covalent linkage of the HaloTag to the HaloTag ligand and any associated cargo (Figure 33C).324 Like SNAP-tag, HaloTag has been used in a number of live-cell SMLM studies on organelle or protein function, particularly with cell-permeable fluorophores that require no electroporation or other special procedures for intracellular delivery.154,213,328,329
Figure 40.
Live-cell SMLM reveals nanoscale features and dynamics. A) 3D SMLM images of clathrin-coated pits (CCPs) labeled with Alexa Fluor 647 in live BS-C-1 cells that express clathrin light chain tagged with SNAP (see Section 6.2). B) Two-channel, 3D SMLM image of a CCP with clathrin light chain (magenta) labeled as above and with its cargo transferrin (green) directly labeled with Alexa Fluor 568. C) 2D SMLM image of a cultured hippocampal neuron labeled with Dil showing changes to the dendritic structure including a growing spine or filopodium (green arrowhead), an extending filopodium (blue arrowhead), and a retracting filopodium (magenta arrowhead). Scale bars 100 nm (A, B), 1 μm (C). Panels A-B were acquired in 30 sec, while the snapshots in panel C were obtained in 10 sec. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 261, Copyright 2011. Adapted with permission from Ref. 216, Copyright 2012 National Academy of Sciences, U. S. A.
Small peptide tags, together with enzymatic labeling techniques or reactive proteins, have also been applied nicely in SMLM. Howarth et al. have engineered biotin ligase to covalently label a 15-mer amino acid tag (named an acceptor peptide) with biotin for downstream detection via readout with fluorescent streptavidin.330 This general procedure has been refined with the use of monomeric streptavidin, produced recombinantly and fluorescently labeled in vitro, for studies of synaptic adhesion proteins by a variety of super-resolution microscopy methods, including SMLM.331 In a different approach, Zakeri et al. adapted the Streptococcus pyogenes adhesion protein FbaB from to form a new tag system termed SpyTag/SpyCatcher that is based on FbaB’s spontaneous ability to form an intramolecular isopeptide covalent bond.332 In this system, FbaB was split into a small 13-mer amino acid peptide named SpyTag and a 128-mer amino acid protein domain named SpyCatcher that are able to rapidly form a covalent bond in solution. Chamma et al. demonstrated the use of SpyTag/SpyCatcher with SMLM for imaging of SpyTagged actin, keratin, and clathrin proteins in fixed specimens that were labeled with a SpyCatcher-Alexa Fluor 647 conjugate.333
Finally, tagging of proteins of interest with classic, small epitope tags such as HA, myc, FLAG, and V5 provides a convenient means for downstream antibody labeling, although these are generally limited to use with fixed cells. Epitope tagging can be particularly useful when working with proteins for which good antibodies are unavailable or when densely tagging structures for efficient downstream detection. Viswanathan et al. have further advanced this capability by engineering highly antigenic GFP-like tags which utilize GFP as a protein scaffold on which to display 10-15 copies of a classic epitope tag; the resulting tags, termed ‘spaghetti monster’ fluorescent proteins, have enabled bright immunolabeling of key proteins and structures with antibodies for SMLM and other fluorescence-based experiments.334 While classic short epitope tags are generally read out using IgG-based antibodies due to a lack of available nanobodies, a recent report described a nanobody against BC2-tag, a new epitope tag originally derived from beta-Catenin, and its use with SMLM on BC2-tagged actin, tubulin, and vimentin in fixed cells.335–337
6.3. Small molecule affinity probes
Small molecules are another important class of probes that can label specific organelles, structures, or (in some cases) proteins, by virtue of the intrinsic affinity of the fluorophore or through affinity conferred by an appended group. For instance, cell permeant, cationic fluorophores label mitochondria in living cells due to their negative potential, octadecane-conjugated fluorophores target the plasma membrane due to their hydrophobicity, and phalloidin-fluorophore conjugates bind filamentous actin. While such affinity-based probes lack the generality of genetically encoded protein tags, they are nonetheless important tools in SMLM due to their sometimes very intense stains (e.g., MitoTrackers) or their specificity for one protein conformation over another (e.g., phalloidin probes bind polymerized, not monomeric, actin).
Table 3 provides a list of some affinity-based probes used for SMLM with some example images in Figure 34. For example, the staining of filamentous actin can be achieved much more densely with a fluorescent phalloidin conjugate than with bulkier anti-actin antibodies or actin-affinity-based proteins such as LifeAct, and Xu et al. used phalloidin-Alexa Fluor 647 staining of actin together with a novel dual-objective microscope to capture highly detailed images of actin in fixed specimens.343 In live specimens, Shim et al. used various established organelle tracker fluorophores to visualize membrane bound organelles in live cells by SMLM, including the use of MitoTracker Red to monitor mitochondrial fusion and fission events and ER-Tracker Red to monitor dynamics of the endoplasmic reticulum.216
Table 3.
Example affinity stain probes used in SMLM.
| Target | Basis of Affinity | Recognition Mechanism | Examples used in SMLM |
|---|---|---|---|
| Mitochondria | Positive charge of dye and/or ligand | Mitochondria negative potential in live cells attracts cationic, membrane permeable dyes | MitoTracker Orange,216 MitoTracker Red,216,338 MitoTracker Deep Red,216 Rhodamine 123,338 TMRE338, RhoB-HMSiR339 |
| Plasma Membrane | Appended hydrophobic group (e.g., octadecane) | Hydrophobic group partitions dye to plasma membrane; uptake may follow | DiI/DiD/DiR216, DiI-C6-HMSiR339 |
| Endoplasmic Reticulum (ER) | Lipophilic framework or glibenclamide (containing sulfonylurea group) | Lipophilic affinity or binding between sulfonylurea and its potassium channel receptor enriched on ER | ER-Tracker Red,216 Nile Red338,340, Cer-HMSiR339 |
| Golgi | Appended ceramide or sphingomyelin group | Interaction with endogenous lipids and cholesterol | Cer-HMSiR339 |
| Lysosomes | Appended weakly basic amine linker | Protonation-induced accumulation in acidic environment of lysosomes | LysoTracker Red,216 LysoTracker Green338 |
| DNA | Polycyclic, aromatic, and planar | Intercalation, π- π stacking | Picogreen,120,341 YOYO-1120,342 |
| Filamentous actin | Appended phalloidin group | Phalloidin binds interface of polymerized actin subunits | Phalloidin-Alexa Fluor 647343 |
Figure 34.
Use of affinity-based small molecule labels for SMLM. A) 3D SMLM image of axons in fixed, cultured neurons where filamentous actin is labeled with phalloidin-Alexa Fluor 647 and reveals a periodic membrane-associated cytoskeletal structure. B) SMLM image of mitochondria and the endoplasmic reticiulum in a live BS-C-1 cell labeled using MitoTracker Red and ER-Tracker Red. C) SMLM image of DNA in a live U2OS cell labeled with Picogreen. Scale bars are 500 nm for B, and 2.5 μm for C. From Ref. 344, adapted with permission from AAAS. Adapted with permission from Ref. 216, Copyright 2012 National Academy of Sciences, U. S. A. Adapted with permission from Ref. 341, Copyright 2012 John Wiley and Sons.
6.4. FISH probes
A key sub-class of non-genetically-encoded affinity-based probes are oligonucleotide probes used in various FISH (fluorescence in situ hybridization) experiments. FISH probes are designed to hybridize with nucleic acids for the detection of specific sequences of DNA or RNA using a range of fluorophores including the popular photoswitchable fluorophore Alexa Fluor 647. These probes are typically comprised of DNA oligonucleotides but sometimes higher affinity variations are used such as peptide nucleic acids or locked nucleic acids. For example, fluorescent oligonucleotides have been used to label: Xist RNA in mouse embryonic fibroblast nuclei for the study of X chromosome inactivation (Figure 35A-C)345; telomere conformation and factors impacting formation of T-loop structures in which the 3’ terminal overhang of telomeres invades the double-stranded region of the telomere (Figure 35D)346; chromosome conformations347,348; and mRNA distributions in yeast for simultaneous detection of 32 different gene transcripts.349 FISH is an important method for SMLM and will no doubt remain so for years to come.
Figure 35.
SMLM using labeling of nucleic acids by fluorescence in situ hybridization (FISH). A) Conventional fluorescence image of a tetraploid mouse embryonic fibroblast nucleus stained with the general nuclear stain Hoechst 33342 (blue) and FISH probes against Xist RNA (magenta), revealing two inactivated X-chromosomes (zoom-in views, right). B-C) 3D SMLM images of Xist RNA from zoom-in views of A, where distinct Xist puncta are evident at a range of heights (encoded in color). D) SMLM image of telomeres detected using FISH in chromatin from a mouse splenocyte. Adapted with permission from Ref. 345, Copyright 2015 National Academy of Sciences, U. S. A. Adapted from Ref. 346, Copyright 2013, with permission from Elsevier.
6.5. Metabolically incorporated probes
Rather than utilize probes that bind covalently or noncovalently to targets on a biological specimen, some labeling strategies utilize a cell’s metabolic machinery to incorporate exogenous probes. Many of these strategies utilize a two-step procedure in which small molecules containing a “click chemistry” group such as an azide or alkyne can be incorporated into specific molecules or structures and are then incubated with a fluorophore containing a ligand that can specifically react with the click chemistry group on the metabolically introduced label.
Jiang and co-workers350 introduced to living cells monosaccharides functionalized with azide or alkyne groups, conjugated to them a cyanine dye bearing an alkyne or azide group, and studied their dynamics on the membranes of live cancer cells by single-molecule tracking and by SMLM. They were able to observe differences in the mobility of N-linked sialic acids and O-linked N-acetyl galactosamine and they also captured SMLM images of membrane nanotubes in various cancer cell lines.
In order to label actively replicating nucleic acids, 5-ethynyl-2-deoxyuridine (also known as EdU, an alkyne modified nucleobase) can be incubated with a cell and later coupled to a fluorophore using copper-catalyzed azide alkyne cycloaddition.351–353 Zessin et al. utilized EdU labeling of cells to incorporate Alexa Fluor 647 onto chromosomal DNA, revealing a detailed chromatin map.352
An expanded genetic code (Figure 36) has been used to incorporate fluorophores directly to non-canonical amino acids on a specific residue of a protein of interest.354 For example, Uttamapinant et al. incorporated an amino acid bearing a bio-orthogonal reactive group BCNK (bicycle[6.1.0]nonyne-lysine) into various proteins of interest (actin or tubulin) and labeled it with the fluorophore SiR using inverse-electron-demand Diels-Alder cycloaddition with a tetrazine-SiR conjugate; the resulting labeled specimens were imaged by live-cell SMLM.355
Figure 36.
Use of an expanded genetic code to introduce a non-canonical amino acid into a specific protein of interest at a specific site. Blue box and orange arrow indicate complementary functional groups for biorthogonal reactions and red box indicates the fluorophore (F). Adapted with permission from Ref. 356, Copyright 2012 John Wiley and Sons.
7. Multichannel SMLM
Many clever strategies have been utilized to date to image distinct structures or molecules in SMLM, although most researchers either use spectrally distinct fluorophores or use sequential imaging of the same fluorophore. Some may use a combination of these two basic approaches and many exploit unique switching properties of the fluorophores.
The most conceptually straightforward approach for multichannel imaging is to utilize fluorophores which are spectrally well-resolved from each other. For instance, the particularly good SMLM fluorophore Cy5 (or its close relative Alexa Fluor 647) has been paired with many fluorophores for two-color imaging, including ATTO 488, ATTO 520, Cy3B, Cy7 (and their close relatives), mEos2, rsFastLime, and others.128,135,138,139,261,357,358 This type of multichannel approach is commonly used by researchers, but requires a careful sub-pixel alignment of the two or more spectral channels and the use of an instrument with the necessary laser lines at a sufficient power for imaging.
Bates et al. introduced activator-reporter fluorophore pairs for multichannel imaging137 which evolved naturally from their earlier work in developing the Cy5 photoswitch and in developing SMLM.26,76 In this approach, a photoswitchable “reporter” fluorophore such as Cy5 was paired with a bluer-absorbing “activator” fluorophore (e.g., Alexa Fluor 405 or Cy3) on antibody molecules, where distinct sets of antibody molecules were labeled with different activator molecules but the same reporter. The activator-reporter phenomenon has not been investigated mechanistically but appears to be unique to pentamethine and heptamethine cyanine reporters with thiol-based (but not phosphine-based) switching and works with a wide range of blue-shifted activator dyes.135,137 Labeling with activator-reporter pairs is complemented by use of a custom illumination series as follows. Intense red light is used to initially switch reporter molecules to a nonfluorescent ‘off’ state, then illumination with weak light on resonance with one set of activator dyes is used to activate a subset of reporter fluorophores, and then the activated fluorophores are imaged and switched off again with intense red light. The process is then repeated with different activator wavelengths and then the entire cycle is repeated hundreds or thousands of times to build up a large number of localizations. During analysis, the reporter molecules are assigned to a specific channel based on the known activation wavelength that preceded the localization. Nonspecific activation may lead to substantial crosstalk in this activator-reporter approach, particularly when one activator-reporter pair is much more abundant than another, but the effect can be minimized through use of a crosstalk subtraction algorithm.359 Although activator-reporter pairs are not widely used at present, they have nonetheless enabled many important studies (Figure 37).344,359–361
Figure 37.
Conventional image (A) and corresponding SMLM image (B) of synapses in mouse accessory olfactory bulb imaged using activator-reporter pairs. Scale bars are 1 μm for A and B. Adapted from Ref. 359, Copyright 2010, with permission from Elsevier.
Fluorescent reporter molecules with overlapping, but distinct emission spectra are also used for multicolor SMLM in two or more channels. In one approach, the fluorescent image is split by a dichroic mirror into two different spectral bands such that the fluorescence emitted by a given fluorophore is split between two bands with a characteristic ratio.162 When individual fluorophores are imaged in this way, the detected ratio of fluorescence in the two bands can be analyzed and compared to control data to determine the fluorophore’s identity. Up to four spectral channels have been imaged in this way, with good discrimination between fluorophores whose emission spectra may differ by as little as 20 nm.147,216,362–364 In a second approach, a dual-objective SMLM instrument was used to capture images of single molecules in one detection path together with the measurement of single-molecule spectra in the other detection path; up to four-channel imaging was demonstrated, with excellent discrimination of fluorophores whose emission spectra was separated by as little as 10 nm (Figure 38).365 The concurrent detection of a spectrally dispersed image with a non-dispersed image may also be performed using a single objective lens with a custom detection path that splits the image into two beams, one of which is dispersed spectrally.340,366–368 Spectrally-resolved SMLM is also useful in combination with sensor fluorophores like Nile Red, whose emission spectrum reports on local hydrophobicity, to measure membrane polarity in living specimens.340,368
Figure 38.
SMLM using A) spectrally-resolved detection for four-channel imaging. The fluorophores Dyomics 634, DyLight 650, CF660C, and CF680 were used to immunolabel peroxisomes, vimentin filaments, microtubules and the outer mitochondrial membrane, respectively, in a fixed PtK2 cell. Although the fluorescence emission spectra overlapped for the four fluorophores B), concurrent spatial and spectral detection allowed discrimination of each fluorophore from the others for multichannel imaging. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 365, Copyright 2015.
The activator-reporter approach and the overlapping emission spectra approaches each have several benefits: only one powerful laser is required for the instrument; the distinct channels in single detection path instruments are easily registered, not requiring careful sub-pixel alignment. A notable drawback of these approaches, however, is that they divide up the maximum number of achievable localizations among all the channels, often resulting in a reduced density per channel. Consider the scenario where the duty cycle of a particular fluorophore is sufficient to support ~200 fluorophores in a diffraction-limited region without excessive double-localization artifacts. In this case, the fluorophores can be used to label one structure very densely, or four structures with lower density per structure and thus worse resolution per channel than for a single channel. Because spectrally overlapping fluorophores must also be detected individually within a diffraction-limited region, they also face the challenge of finite fluorophore density; additionally, for experiments with different types of fluorophores, researchers should use the lowest duty cycle fluorophores on the densest structures to avoid being limited to the properties of the fluorophore with the highest duty cycle.
Sequential imaging of different proteins or structures using the same fluorophore has been implemented in a variety of ways. For instance, Tam et al.369, Valley et al.370, and Yi et al.371 developed procedures consisting of multiple cycles consisting of: labeling the specimen with the fluorophore Alexa Fluor 647 (or Cy5); imaging by SMLM; and removal of signal through elution, chemical reduction, and/or photobleaching. The SMLM images obtained in each round may then be assigned to a given channel of a composite image. These approaches nicely leverage the robust photoswitching behavior of Alexa Fluor 647 but require a lengthy overall procedure which includes multiple rounds of immunolabeling. The method dubbed Exchange-PAINT from Jungmann et al.279 (Figure 39) minimized these delays by using a single round of immunolabeling with DNA-conjugated antibodies prior to imaging followed by imaging rounds consisting of sequential rounds of incubation with fluorescently labeled (single-stranded) imager DNA strands. Sequential imaging approaches using DNA-conjugated antibodies and imager DNA strands enable scaling to many channels by exchange or to larger depths by refreshing photobleached probes and are an area of active development.284,372,373
8. Live SMLM
Imaging of live specimens by SMLM faces the key challenges of labeling the living specimen, recording a series of frames rapidly enough to avoid blurring due to sample or molecular motion, and avoiding perturbation of the sample. In this section, we will review these key challenges and then we will highlight a few studies which have pushed the envelope for live-cell SMLM beyond those which have already been mentioned, above.
The labeling of living specimens is relatively straightforward using fluorescent proteins. Many early live-cell SMLM experiments utilized various photoactivatable or photoconvertible FPs and focused on the study of relatively slow processes, including: the study of the nanoscale distribution and motion of viral membrane proteins in eukaryotic cells using PA-GFP and EosFP;111,374 the study of focal adhesion dynamics using EosFP;125 and the study of bacterial cytoskeletal proteins using YFP (under conditions that lead to photoswitching of YFP).375 The labeling of living specimens with cell-permeant small molecule probes such as MitoTracker or ER-Tracker has been enabling with experiments for which switchable, specific small molecule probes are available.216,338 A broader set of SMLM experiments in live cells has been enabled by the use of transgenically expressed (reactive) protein tags such as TMP, SNAP, and Halo that can target a specific protein, together with the use of cell-permeant fluorophores such as ATTO 655, tetramethyl rhodamine, and SiR .154,327,362 The development of photoactivatable and cell-permeable organic fluorophores such as PA-JF549 and PA-JF646 has been particularly useful for SMLM-style photoactivated particle tracking experiments (sptPALM)111 due to the high brightness and high photostability of these probes, and they have been used in, for instance, several studies of chromatin and transcription factor dynamics.168,376,377
The second challenge with live-cell imaging is that many localizations must be rapidly recorded in order to observe the cell with a spatial and temporal resolution that captures the features and dynamics of interest. This leads to an inevitable tradeoff between spatial resolution, which is limited in part by the number of frames in an image stack, and temporal resolution, which is limited by the time required to record the image stack. Rapid charge coupled device and scientific complementary metal on semiconductor cameras have achieved frame rates of ~500-3200 Hz for SMLM,216,261,378 which is impressive but still restricts imaging to relatively slow processes.
A third challenge with live imaging is avoiding perturbation of the specimen. In the case of SMLM, where researchers typically use relatively high illumination intensity (≥1 kW/cm2), phototoxicity due to illumination of the sample is a major concern, particularly when imaging with relatively short wavelengths (<500 nm).379 Some fluorophores, upon illumination, can produce toxic substances such as reactive oxygen species, although SMLM fluorophores have not been systematically tested in this regard, and other modes of phototoxicity, for instance due to excitation endogenous chromophores, also likely occur. Finally, for experiments in which an imaging cocktail is used, researchers must exercise care to avoid or minimize toxicity (see Section 3.4). At a minimum, researchers should perform control measurements to make sure their phenotypes of interest are not perturbed by the experimental conditions.
We would like to highlight four recent studies employing live-cell SMLM. Jones et al.261 used live-cell 3D SMLM to resolve the nanoscopic structure of clathrin-coated pits (Figure 40 A-B). Living cells expressing clathrin light chain fused to a SNAP tag (see Section 6.2) were labeled with Alexa Fluor 647, where electroporation was used to deliver the otherwise cell-impermeant fluorophore to the cytosol of eukaryotic cells. In some experiments the cells were also incubated with Alexa Fluor 568-labeled transferrin. Using these specimens, the researchers were able to capture a series of snapshots that revealed the nanoscale morphology of the clathrin coat enclosing the transferrin cluster—features unobservable by conventional fluorescence microscopy methods (Figure 40B). In another study, Shim et al.216 used various commercially available organelle or membrane tracker fluorophores such as Dil (see Section 6.3) to label the plasma membrane of a living, cultured hippocampal neuron, and clearly resolved the extension and retraction of filopodia or dendritic spines (Figure 40C).
Takakura et al.380 used novel conjugates of the spontaneously-blinking silicon rhodamine fluorophore HMSiR164 (Figure 12, Figure 15A) for long time-lapse SMLM of membrane-bound organelles in live cells. They found that the environmentally-sensitive fluorophore HMSiR, when localized to hydrophobic environments such as the membranes of the endoplasmic reticulum (ER), or the cell plasma membrane, exhibits spontaneous blinking with a very low duty cycle (up to a 50-fold improvement over that of HMSiR in a hydrophilic environment) while still emitting ~1,500 detected photons per localization. Additionally, they used a powerful two-step labeling procedure to attain highly dense labeling: in step 1, cells were incubated with a targeting group that localized to a particular region of the cell (e.g., endoplasmic reticulum, plasma membrane) and that was functionalized with a trans-cyclooctene group; in step 2, cells were incubated with a HMSiR fluorophore that is functionalized with tetrazine, where the tetrazine and trans-cyclooctene groups react rapidly and selectively with one another in order to generate the probe, in situ. Together, this approach has enabled hundreds of snapshots to be recorded over tens of minutes in living specimens with a high spatial resolution (Figure 41). The extension of these so-called HIDE (high density, environment-sensitive) probes to multiple channels may benefit from a recently reported381 green-absorbing, spontaneously-blinking fluorophore (HEtetTFER, spectrally orthogonal to HMSiR), although further developments may be required to optimize a second orthogonal coupling strategy to efficiently and selectively couple two different fluorophores with their respective targeting groups.
Figure 41.
High density environmentally sensitive (HIDE) probes for live cell SMLM based on the spontaneously-switching fluorophore HMSiR (red, see also Figure 12). A) HIDE probes are synthesized in vivo in a two-step reaction between a trans-cyclooctene-functionalized targeting ligand R that labels an organelle or region of a cell and the tetrazine-functionalized fluorophore HMSiR. The targeting ligand R and labeling conditions can be chosen to label the Golgi, endoplasmic reticulum (ER), mitochondria, or the plasma membrane. B) A SMLM image of the ER in a live HeLa cell acquired in 2 sec using the HIDE probe Cer-HMSiR with partial conventional fluorescence overlay (lower left). This is one snapshot from a series of ~750 spanning ~25 min. Features of the ER are indicated in lowercase including ER sheets (s), ER tubules (t), and fenestrated ER sheets (f). C-D) Several 2 sec snapshots across 25 minutes of acquisition (2 sec per frame) for the boxed regions in B showing dynamic behavior of the ER including budding (yellow arrowheads), looping (green arrowhead), fusion (blue and magenta arrowheads), and reversible transition between tubular and sheet-like morphologies (white arrowheads). E) Kymograph along purple line in B showing the line profile (vertical) as a function of time. Scale bar 500 nm. Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 380, Copyright 2017.
9. Potential artifacts and pitfalls of SMLM
Despite its strong capabilities, SMLM faces several potential artifacts and pitfalls which require special care by practitioners.382,383 Some of these are common to all fluorescence modalities such as inadequate fixation, nonspecific labeling,384 or overexpression artifacts, while others are more specific to super-resolution methods or SMLM, including double localizations and label linkage errors. In this section we will highlight these concerns.
9.1. Fixation and immunolabeling artifacts
In immunostaining procedures, specimens are typically processed using a protocol consisting of fixation, permeabilization, and antibody staining steps, with optional pre-fixation extraction and post-stain fixation steps, and each may require optimization.384,385 The choice of fixative often represents a tradeoff between preservation of structure and an ability to label the specimen, in two key ways. For instance, fixation with cold methanol can nicely preserve the microtubule cytoskeleton but will destroy many membrane-bound organelles and tends to flatten cellular specimens. Alternatively, fixation with a mixture of formaldehyde and glutaraldehyde can preserve the structure of cytoskeletal components together with subcellular organelles but the use of glutaraldehyde may inhibit antibody penetration into thick tissues and can severely reduce antibody recognition of antigens for some antigens/antibodies. Formaldehyde-only fixation does a better job preserving antigenicity and penetration into thick tissues but does not preserve some structures sufficiently, such as the microtubule cytoskeleton, which may depolymerizes under these conditions, and the choice of detergent in the permeabilization step must be made carefully to avoid damaging less robustly fixed structures such as mitochondria.14,15,383 Glyoxal was recently demonstrated as a potential alternative to formaldehyde and/or glutaraldehyde, fixation, and was reported to generally have good penetration and good structure preservation.386 An extraction step, typically consisting of a brief, pre-fixation treatment with detergent, is extremely useful when studying the cytoskeleton in cultured cells since it permeabilizes the membrane and ‘spills’ unpolymerized cytoskeletal proteins that lead to background signal when interested in studying the assembled cytoskeleton.14,15,383 Many membrane-bound organelles are destroyed or perturbed by extraction, however.
Assuming that the structures of interest are preserved, that the epitopes of interest are preserved, and that antibody penetration into the specimen is sufficient, there still exist several potential pitfalls in antibody labeling including low affinity, low specificity, lot variability, and crosstalk with secondary antibodies. Antibody labeling is idiosyncratic and requires attention to detail.387,388 Some antibodies seem to provide excellent stains in a wide range of conditions while others can lead to high background signals in some situations and others may never seem to work to target the structure indicated by the manufacturer or publications. When co-staining two different proteins in a cell, sometimes the only effective antibodies available are raised in the same host animal, which would lead to severe crosstalk with indirect (primary/secondary) immunofluorescence; workarounds such as the use of direct immunofluorescence (i.e., fluorescently labeled primaries), sequential immunostaining, or isotype-specific secondary antibodies may remedy the situation but also require careful optimization. Secondary antibodies that are fluorescently labeled and that are designed to detect primary antibodies from different, but closely species (e.g. mouse and rat) may have unwanted crosstalk unless highly cross-adsorbed secondary antibodies are used; when available, fluorescently labeled primary antibodies may be utilized.
Hopefully these brief remarks have emphasized the key point that there is no universal immunostaining protocol. SMLM researchers routinely spend substantial time exploring this parameter space when new systems or proteins are being studied.
9.2. Artifacts with protein fusion tags and fluorescent proteins
Fusing a fluorescent protein, epitope tag for high-affinity immunolabeling, or reactive protein tags such as SNAP or Halo to a protein of interest is an elegant and powerful approach for sample labeling that avoids many of the pitfalls of antibody labeling of endogenous proteins but introduces its own set of risks. Fusion tags have the potential to disturb protein function through the tag itself, through tag-mediated dimerization or aggregation, or through perturbation of protein expression levels for instance by overexpression.7,25,63,389 As with antibodies, fusion protein labeling is highly idiosyncratic and requires care when working with and validating new constructs and procedures. For many systems, expression of fusion proteins has been typically achieved by transformation with plasmid DNA bearing the fusion protein, but this has the potential problems that high expression levels may be unphysiological or toxic or that without concurrent knockdown such as with siRNA the procedure is unable to report on endogenous, unlabeled proteins.319 Some of these problems should be addressable through the use of improved genome editing tools390 which can allow for knockout or knock-in cells with fusion proteins under endogenous promoters.
9.3. SMLM-specific artifacts
The unique point-by-point approach of SMLM lends the method to artifacts.9,382,391,392 Besides channel crosstalk incurred through the use of various multi-channel SMLM methods described above, SMLM is particularly sensitive to ‘double localization’ artifacts where two (or more) localizations occur within a diffraction-limited region. As shown in Figure 42, double/multiple localizations can introduce spurious features or substantial distortions and may be easy to overlook for samples containing isolated puncta or filamentous structures where spurious localizations would partly overlay with the true structures.
Figure 42.
Double-localization artifacts in SMLM. A-C) Cartoon illustration, with actual fluorophore distributions shown in black, measured single-localizations shown in magenta and measured double or multiple localizations shown in green. A) Punctate structures produce double localizations toward the center of each punctum and between puncta that are separated by approximately the diffraction limit or less (~250 nm). B) Filaments show double localizations along the length of the filaments, and are easily mistaken for valid, single localizations. Filaments in close proximity, however, reveal spurious localizations where the two filaments approach each other and/or cross. C) A single filamentous structure with abundant double/multiple localizations will appear artificially narrowed. D) Experimental SMLM image of microtubules in a fixed and immunolabeled BS-C-1 cell recorded under conditions that produced many double/multiple localizations; localizations with large spot widths (more likely to be double/multiple localizations) have been rendered in green while others have been rendered in magenta.
Statistically speaking, double/multiple localizations are unavoidable in SMLM, and we can straightforwardly derive a simple estimate for their likelihood. Consider a diffraction-limited region containing N idealized fluorophores which all exhibit perfect, binary switching with the same on/off rates and duty cycle, where duty cycle (also known as the “on fraction”) is defined as DC=ton/(ton+toff). The likelihood that a given fluorophore is off is just 1-DC. From this, it follows that when one of the N fluorophores is on, the likelihood that all of the other N-1 fluorophores will be off is (1-DC)N-1. Finally, the multiple localization likelihood (MLL), when more than one fluorophore in a diffraction-limited region is on at the same time, is then just MLL=1-(1-DC)N-1. For example, with DC=0.01 and N=100, we get MLL≈0.63, meaning that 100 fluorophores with a duty cycle of 0.01 in a diffraction-limited region would have multiple localizations ~63% of the time. For DC=0.001 and N=100, the situation is far better with MLL=0.09, or 9% multiple localizations. The use of a low duty cycle fluorophore can help to reduce this effect, as can filtering on the basis of localization brightness and/or shape (e.g., elliptical), or through the use of more sophisticated data analysis algorithms that can detect more than one emitter per fluorescent spot.343,393–396 Dai et al. have performed related calculations and simulations on multiple localization artifacts that also include the effect of discrete exposure times281 and Sinkó et al. have developed a software tool that can simulate SMLM results including the effects of double/multiple localizations.397
Beyond double/multiple localization artifacts, the short length scales being probed in SMLM or other super-resolution microscopy methods raises additional possible artifacts. First, the linkage between fluorophore and target protein or structure is often many nanometers and this linkage error may result in substantial broadening effects or distortions (see also Section 6.1). Second, multichannel SMLM imaging requires careful sub-pixel registration especially for multicolor approaches where different detection paths are used and aberrations can be substantial.107,398,399 Third, hindered molecular rotation can induce localization errors due to fixed dipole effects, particularly when using high numerical aperture objective lenses as is standard in SMLM,136,400–403 however, custom hardware and has been developed which can mitigate or eliminate these effects.404–407
9.4. “Resolution”
The assessment of resolution in SMLM is not straightforward, and without delving into a rigorous discussion on quantitative metrics of resolution, we wish to make three points not yet raised in this review.126,392,408–410 First, “resolution” in optical microscopy is generally a measure of the minimum distance at which two objects may be distinguished in an image. This basic definition is somewhat at odds with SMLM’s mode of operation, however, since in SMLM, fluorescent labels are already detected individually and are therefore distinguished from one another as a function of both space and time. Second, a high density of localizations is in general sought to support a high spatial resolution in a Nyquist-limit sense (Figure 4), although caveats exist for this sort of analysis. For instance, a sparse distribution of molecules in a sample that is labeled perfectly with fluorophores would have appear to have a low spatial resolution in a Nyquist-limit sense, strictly based on consideration of the density, even if the fluorophore and imaging conditions could have supported a much higher density. Alternatively, a densely-labeled but small object that doesn’t fill a diffraction-limited volume may appear to have a high spatial resolution, but the fluorophore duty cycle may not support that resolution for a larger object of equal density that fills more of the diffraction-limited volume. This apparent dependence of the attainable resolution on the specimen itself is sometimes a source of confusion in the field. Third, localization uncertainty, which also impacts achievable resolution (see Section 2.5), is reported in different ways in the literature. While some researchers refer to the standard deviation (SD) of the localization uncertainty, others refer to the full width at half maximum (FWHM) of the localization uncertainty, where a Gaussian distribution has a FWHM that is 2.35× higher than its SD (Figure 43), and yet others have simply estimated the theoretical localization uncertainty based on the number of detected photons and a few other measured or estimated parameters. Whenever making comparisons of localization uncertainty, it is therefore important to keep in mind which quantity is being considered.
Figure 43.
SMLM studies use different metrics to characterize uncertainty in localization, including full-width at half maximum (FHWM) and standard deviation (SD), and it is important to determine which metric is used when comparing performance in different studies. For a Gaussian distribution, FWHM=2.35×SD.
10. Concluding remarks
SMLM has developed rapidly since its inception in 2006 and within several years has been used by researchers in a wide range of disciplines to gain insights into the nanoscale organization of biological systems. Given the rapid progress in SMLM over the past 12 years, it is tempting to consider its prospects for the future.
Could SMLM one day be used to read out dense distributions of on the few-nm length scale? Recent experiments are promising. Two reports studying DNA origami model systems demonstrated successful resolution of 2D arrays of ~10-50 fluorophores in which adjacent fluorophores were separated by only 5-6 nm. In the first report, Dai et al.281 achieved this by detecting a very large number of photons from bound fluorescent probes using DNA-PAINT with optimized imaging conditions (Figure 28). In the second report, Balzarotti et al.407 combined photoswitching with a novel patterned excitation approach (MINFLUX) to achieve a similar result with far fewer photons detected per molecule. Pushing to even shorter length scales, Weisenburger et al.411 imaged the fluctuating emission from ~4 spectrally identical fluorophores within a diffraction-limited area in a novel cryogenic fluorescence microscope to attain sub-nanometer localization precision (SD) with purified protein specimens. It is worth noting that fluorophores behave quite differently in vitreous ice at low temperatures (~4 K) compared to their behavior in solution near room temperature (~300 K), including a dramatically lower photobleaching rate; if robust photoswitching or photoactivation could be achieved in this state, it could extend the sub-nanometer localization methodology to larger numbers of fluorophores per diffraction-limited spot. Advancing these ultrahigh-resolution methodologies beyond proof of principle may require further developments, perhaps including new fluorescent probe labeling technology, but there are nonetheless several clear paths forward. Alternatively, multimodal experiments involving SMLM and electron microscopy have made good progress in recent years and potentially offer the benefits of each method with few drawbacks as recently reviewed by Hauser et al. and Kopek et al.412,413
The prospects for live-cell SMLM are also good based on recent reports. For instance, new probes such as HMSiR probes for tracking the ER (Figure 41) have already substantially advanced live-cell SMLM experiments to enable several hundreds of SMLM snapshots at a time, although the methodology to date has been only shown on select targets. In terms of speed, it is likely that SMLM faces a hard, fundamental limit, which we can briefly consider. An individual localization consisting of an emission burst of ~1000 detected photons would require at least 10,000 fluorescence emission events (only some of which reach the camera) such that a localization could not be detected in a shorter period than about 10,000× the fluorescence lifetime, or ~0.01 ms, assuming a 1 ns fluorescence lifetime.78 If 1000 frames are required in order to reconstruct a SMLM image, then doing so would require at least ~10 ms, but probably a factor of 10 or more times longer per SMLM image (~100 ms) given the highly optimistic estimated values and several disregarded complications. Achieving high photon emission rates in such a short period of time will of course require very high laser illumination, raising concerns for phototoxicity that may ultimately impose stricter limitations than any consideration of fluorophore (or hardware) rates.
Beyond pushing the frontiers of spatial and temporal resolution in SMLM, important work remains to be done in making SMLM a workhorse technique that biologists can easily use in their research. Two- or three-channel SMLM with FPs in fixed specimens is still difficult due to a lack of robustly switching, spectrally orthogonal fluorophores, and even one-channel SMLM with FPs would benefit greatly from the development of a switchable FP with a substantial boost in photons. Multichannel SMLM in fixed specimens with organic fluorophores still requires a great deal of optimization in sample preparation and skill to execute well, and it would benefit greatly from improved spectrally orthogonal fluorophores and fluorescent labeling strategies.
This leads to the question of how improve fluorophores for SMLM. Fluorescent protein developers have tended to use a combination of structure-guided design and high-throughput screens to assess the properties of new variants. These have yielded a great many switchable (and non-switchable) FPs, but the screens typically have limited ability to report on single-molecule properties. Recent work from Emanuel et al.414 may help to move the field forward. They measured the performance of a library of ~60,000 FP mutants using fluorescence microscopy, with one FP mutant expressed per cell, and after sample fixation, read out unique oligonucleotide barcodes that had been previously associated with each mutant FP DNA sequence. This allowed the association of time-resolved measurements by fluorescence microscopy with a FP DNA sequence that could later be re-expressed in cells and assessed in detail. While ensemble FP properties were selected in the work by Emanuel et al., the approach could potentially be adapted to enable evaluation of single-molecule switching properties for libraries of many thousands of FP mutants.
Photoswitchable organic fluorophores have been in some ways slower to develop than FPs, in part since the underlying photochemistry is in many cases not well understood and switching properties are difficult to predict. Consider that Cy5 (and its close structural relative Alexa Fluor 647) was synthesized as a general-purpose fluorophore long before the advent of SMLM, and that it fortuitously possessed superior photoswitching capabilities that on the whole have yet to be surpassed. Substantial effort may be required to better understand the fundamental photochemical processes for the various fluorophore classes used in SMLM in order to enable rationally-guided improvements or computational screening of candidate SMLM fluorophores. A potential alternative strategy to produce improved photoswitchable fluorophores could be to utilize a combinatorial synthetic approach to generate libraries of fluorophores together with a sufficiently high-throughput ensemble and/or single-molecule screen of fluorophore properties.
While SMLM is now enabling biologists to reveal the inner workings of the cell with rich molecular detail at unprecedented resolution, much work remains to be done as the methodology matures into a workhorse tool. In many ways, fluorescent probes have guided researchers to the development of SMLM, and they also hold the key to unlocking SMLM’s full potential.
Supplementary Material
Figure 6.
Comparison of photoswitching properties of three different fluorophores. A) Single-molecule on-off photoswitching of a representative Alexa Fluor 647 molecule in a deoxygenated solution containing a thiol. Under these conditions, Alexa Fluor 647 exhibits B) an average duty cycle of ~0.0012 and C) an average of ~5200 photons/localization. These switching properties are sufficient to produce high-quality images of hollow clathrin-coated pits in fixed, immunolabeled cells D-F). The fluorophore ATTO 655 has the same duty cycle as Alexa Fluor 647 G-H) but emits ~8× fewer photons per burst I), and the resulting larger position uncertainty obscures the hollow clathrin-coated pits J-L). The fluorophore Cy5.5 has a duty cycle that is ~6× larger that of Alexa Fluor 647 M-N), while having a similar number of detected photons O), and as a result the achievable density of localizations on clathrin-coated pits is insufficient to confidently trace out the hollow structure P-R). Adapted by permission from Springer Customer Service Centre GmbH: Springer Nature, Ref. 128, Copyright 2011.
ACKNOWLEDGEMENTS
This work was supported by the University of Washington, the University of Washington Royalty Research Fund, a Burroughs-Wellcome Award at the Scientific Interface, and NIH grant R01MH115767.
Biography
Joshua C. Vaughan is an Assistant Professor of Chemistry at the University of Washington, Seattle. He grew up in the San Francisco bay area and received a B.A. degree in Chemistry from Reed College in Portland, OR, where he learned to love the Pacific Northwest. He earned a Ph.D. in physical chemistry at MIT where he worked in the lab of Prof. Keith Nelson on applications of adaptive optics to ultrafast spectroscopy. For his postdoctoral research in Prof. Xiaowei Zhuang’s lab at Harvard he developed tools for super-resolution fluorescence microscopy and studied virus transport. His group at the University of Washington is developing new tools for super-resolution fluorescence microscopy and they are using these tools to understand the molecular scale organization of biological systems in their native cellular or tissue contexts. Outside of work, he enjoys music, sailing, and spoiling his children.
Honglin Li received his B.S. degree in Fine Chemistry at Dalian University of Technology in 2005, and then earned his Ph.D. degree in 2011 under the supervision of Prof. Xiaojun Peng. From 2011 to 2013, he worked as a postdoctoral fellow in Dr. Qian Wang’s group at the University of South Carolina, and then he relocated to Seattle to joined Joshua C. Vaughan’s group at the University of Washington. His research interest is focusing on the development of colorimetric and fluorescent materials for bio-sensing and bio-imaging.
Footnotes
Notes
The authors declare no competing financial interests.
REFERENCES
- (1).Abbe E Beiträge zur Theorie des Mikroskops und der mikroskopischen Wahrnehmung: I. Die Construction von Mikroskopen auf Grund der Theorie. Arch. Für Mikrosk. Anat. 1873, 9, 413–418. [Google Scholar]
- (2).Hell SW Nobel Lecture: Nanoscopy with Freely Propagating Light. Rev. Mod. Phys. 2015, 87, 1169–1181. [Google Scholar]
- (3).Moerner WE (William E . Nobel Lecture: Single-Molecule Spectroscopy, Imaging, and Photocontrol: Foundations for Super-Resolution Microscopy. Rev. Mod. Phys. 2015, 87, 1183–1212. [DOI] [PubMed] [Google Scholar]
- (4).Betzig E Nobel Lecture: Single Molecules, Cells, and Super-Resolution Optics. Rev. Mod. Phys. 2015, 87, 1153–1168. [DOI] [PubMed] [Google Scholar]
- (5).Hell SW Far-Field Optical Nanoscopy. Science 2007, 316, 1153–1158. [DOI] [PubMed] [Google Scholar]
- (6).Hell SW Microscopy and Its Focal Switch. Nat. Methods 2009, 6, 24–32. [DOI] [PubMed] [Google Scholar]
- (7).Patterson G; Davidson M; Manley S; Lippincott-Schwartz J Superresolution Imaging Using Single-Molecule Localization. Annu. Rev. Phys. Chem. 2010, 61, 345–367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Huang B; Babcock H; Zhuang X Breaking the Diffraction Barrier: Super-Resolution Imaging of Cells. Cell 2010, 143, 1047–1058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (9).Sauer M; Heilemann M Single-Molecule Localization Microscopy in Eukaryotes. Chem. Rev. 2017, 117, 7478–7509. [DOI] [PubMed] [Google Scholar]
- (10).Thompson MA; Lew MD; Moerner WE Extending Microscopic Resolution with Single Molecule Imaging and Active Control. Annu. Rev. Biophys. 2012, 41, 321–342. [DOI] [PubMed] [Google Scholar]
- (11).Sengupta P; van Engelenburg SB; Lippincott-Schwartz J Superresolution Imaging of Biological Systems Using Photoactivated Localization Microscopy. Chem. Rev. 2014, 114, 3189–3202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (12).Eggeling C; Willig KI; Sahl SJ; Hell SW Lens-Based Fluorescence Nanoscopy. Q. Rev. Biophys. 2015, 48, 178–243. [DOI] [PubMed] [Google Scholar]
- (13).Liu Z; Lavis LD; Betzig E Imaging Live-Cell Dynamics and Structure at the Single-Molecule Level. Mol. Cell 2015, 58, 644–659. [DOI] [PubMed] [Google Scholar]
- (14).Dempsey GT A User’s Guide to Localization-Based Super-Resolution Fluorescence Imaging In Methods in Cell Biology; Elsevier, 2013; Vol. 114, pp 561–592. [DOI] [PubMed] [Google Scholar]
- (15).Halpern AR; Howard MD; Vaughan JC Point by Point: An Introductory Guide to Sample Preparation for Single-Molecule, Super-Resolution Fluorescence Microscopy In Current Protocols in Chemical Biology; Mahal L, Romesberg F, Shah K, Shamu C, Strano MS, Thomas C, Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; pp 103–120. [DOI] [PubMed] [Google Scholar]
- (16).Gould TJ; Verkhusha VV; Hess ST Imaging Biological Structures with Fluorescence Photoactivation Localization Microscopy. Nat. Protoc. 2009, 4, 291–308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (17).van de Linde S; Löschberger A; Klein T; Heidbreder M; Wolter S; Heilemann M; Sauer M Direct Stochastic Optical Reconstruction Microscopy with Standard Fluorescent Probes. Nat. Protoc. 2011, 6, 991–1009. [DOI] [PubMed] [Google Scholar]
- (18).Schermelleh L; Heintzmann R; Leonhardt H A Guide to Super-Resolution Fluorescence Microscopy. J. Cell Biol. 2010, 190, 165–175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Nienhaus K; Nienhaus GU Fluorescent Proteins for Live-Cell Imaging with Super-Resolution. Chem Soc Rev 2014, 43, 1088–1106. [DOI] [PubMed] [Google Scholar]
- (20).Chozinski TJ; Gagnon LA; Vaughan JC Twinkle, Twinkle Little Star: Photoswitchable Fluorophores for Super-Resolution Imaging. FEBS Lett. 2014, 588, 3603–3612. [DOI] [PubMed] [Google Scholar]
- (21).Heilemann M; Dedecker P; Hofkens J; Sauer M Photoswitches: Key Molecules for Subdiffraction-Resolution Fluorescence Imaging and Molecular Quantification. Laser Photonics Rev. 2009, 3, 180–202. [Google Scholar]
- (22).van de Linde S; Sauer M How to Switch a Fluorophore: From Undesired Blinking to Controlled Photoswitching. Chem. Soc. Rev. 2014, 43, 1076–1087. [DOI] [PubMed] [Google Scholar]
- (23).Yang Z; Sharma A; Qi J; Peng X; Lee DY; Hu R; Lin D; Qu J; Kim JS Super-Resolution Fluorescent Materials: An Insight into Design and Bioimaging Applications. Chem Soc Rev 2016, 45, 4651–4667. [DOI] [PubMed] [Google Scholar]
- (24).Lukinavičius G; Johnsson K Switchable Fluorophores for Protein Labeling in Living Cells. Curr. Opin. Chem. Biol. 2011, 15, 768–774. [DOI] [PubMed] [Google Scholar]
- (25).Fernández-Suárez M; Ting AY Fluorescent Probes for Super-Resolution Imaging in Living Cells. Nat. Rev. Mol. Cell Biol. 2008, 9, 929–943. [DOI] [PubMed] [Google Scholar]
- (26).Rust MJ; Bates M; Zhuang X Sub-Diffraction-Limit Imaging by Stochastic Optical Reconstruction Microscopy (STORM). Nat. Methods 2006, 3, 793–796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).Betzig E; Patterson GH; Sougrat R; Lindwasser OW; Olenych S; Bonifacino JS; Davidson MW; Lippincott-Schwartz J; Hess HF Imaging Intracellular Fluorescent Proteins at Nanometer Resolution. Science 2006, 313, 1642–1645. [DOI] [PubMed] [Google Scholar]
- (28).Hess ST; Girirajan TPK; Mason MD Ultra-High Resolution Imaging by Fluorescence Photoactivation Localization Microscopy. Biophys. J. 2006, 91, 4258–4272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (29).Sharonov A; Hochstrasser RM Wide-Field Subdiffraction Imaging by Accumulated Binding of Diffusing Probes. Proc. Natl. Acad. Sci. 2006, 103, 18911–18916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (30).Moerner WE; Kador L Optical Detection and Spectroscopy of Single Molecules in a Solid. Phys. Rev. Lett. 1989, 62, 2535–2538. [DOI] [PubMed] [Google Scholar]
- (31).Orrit M; Bernard J Single Pentacene Molecules Detected by Fluorescence Excitation in a p - Terphenyl Crystal. Phys. Rev. Lett. 1990, 65, 2716–2719. [DOI] [PubMed] [Google Scholar]
- (32).Shera EB; Seitzinger NK; Davis LM; Keller RA; Soper SA Detection of Single Fluorescent Molecules. Chem. Phys. Lett. 1990, 174, 553–557. [Google Scholar]
- (33).Ambrose WP; Moerner WE Fluorescence Spectroscopy and Spectral Diffusion of Single Impurity Molecules in a Crystal. Nature 1991, 349, 225–227. [Google Scholar]
- (34).Basché T; Moerner WE Optical Modification of a Single Impurity Molecule in a Solid. Nature 1992, 355, 335–337. [Google Scholar]
- (35).Xie XS; Dunn RC Probing Single Molecule Dynamics. Science 1994, 265, 361–364. [DOI] [PubMed] [Google Scholar]
- (36).Trautman JK; Macklin JJ; Brus LE; Betzig E Near-Field Spectroscopy of Single Molecules at Room Temperature. Nature 1994, 369, 40–42. [Google Scholar]
- (37).Nie S; Chiu DT; Zare RN Probing Individual Molecules with Confocal Fluorescence Microscopy. Science 1994, 266, 1018–1021. [DOI] [PubMed] [Google Scholar]
- (38).Dickson RM; Cubitt AB; Tsien RY; Moerner WE On/off Blinking and Switching Behaviour of Single Molecules of Green Fluorescent Protein. Nature 1997, 388, 355–358. [DOI] [PubMed] [Google Scholar]
- (39).Nirmal M; Dabbousi BO; Bawendi MG; Macklin JJ; Trautman JK; Harris TD; Brus LE Fluorescence Intermittency in Single Cadmium Selenide Nanocrystals. Nature 1996, 383, 802–804. [Google Scholar]
- (40).Bopp MA; Jia Y; Li L; Cogdell RJ; Hochstrasser RM Fluorescence and Photobleaching Dynamics of Single Light-Harvesting Complexes. Proc. Natl. Acad. Sci. 1997, 94, 10630–10635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (41).Bout DA Discrete Intensity Jumps and Intramolecular Electronic Energy Transfer in the Spectroscopy of Single Conjugated Polymer Molecules. Science 1997, 277, 1074–1077. [Google Scholar]
- (42).Brown GH Photochromism; Techniques of chemistry, v. 3; Wiley-Interscience: New York, 1971. [Google Scholar]
- (43).Dürr H; Bouas-Laurent H Photochromism: Molecules and Systems: Molecules and Systems; Gulf Professional Publishing, 2003. [Google Scholar]
- (44).Yokoe H; Meyer T Spatial Dynamics of GFP-Tagged Proteins Investigated by Local Fluorescence Enhancement. Nat. Biotechnol. 1996, 14, 1252–1256. [DOI] [PubMed] [Google Scholar]
- (45).Sawin KE; Nurse P Photoactivation of Green Fluorescent Protein. Curr. Biol. 1997, 7, R606–R607. [DOI] [PubMed] [Google Scholar]
- (46).Marchant JS; Stutzmann GE; Leissring MA; LaFerla FM; Parker I Multiphoton-Evoked Color Change of DsRed as an Optical Highlighter for Cellular and Subcellular Labeling. Nat. Biotechnol. 2001, 19, 645–649. [DOI] [PubMed] [Google Scholar]
- (47).Elowitz MB; Surette MG; Wolf P-E; Stock J; Leibler S Photoactivation Turns Green Fluorescent Protein Red. Curr. Biol. 1997, 7, 809–812. [DOI] [PubMed] [Google Scholar]
- (48).Chattoraj M; King BA; Bublitz GU; Boxer SG Ultra-Fast Excited State Dynamics in Green Fluorescent Protein: Multiple States and Proton Transfer. Proc. Natl. Acad. Sci. 1996, 93, 8362–8367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (49).Patterson GH; Lippincott-Schwartz J A Photoactivatable GFP for Selective Photolabeling of Proteins and Cells. Science 2002, 297, 1873–1877. [DOI] [PubMed] [Google Scholar]
- (50).Ando R; Hama H; Yamamoto-Hino M; Mizuno H; Miyawaki A An Optical Marker Based on the UV-Induced Green-to-Red Photoconversion of a Fluorescent Protein. Proc. Natl. Acad. Sci. 2002, 99, 12651–12656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (51).Chudakov DM; Belousov VV; Zaraisky AG; Novoselov VV; Staroverov DB; Zorov DB; Lukyanov S; Lukyanov KA Kindling Fluorescent Proteins for Precise in Vivo Photolabeling. Nat. Biotechnol. 2003, 21, 191–194. [DOI] [PubMed] [Google Scholar]
- (52).Ando R Regulated Fast Nucleocytoplasmic Shuttling Observed by Reversible Protein Highlighting. Science 2004, 306, 1370–1373. [DOI] [PubMed] [Google Scholar]
- (53).Chudakov DM; Verkhusha VV; Staroverov DB; Souslova EA; Lukyanov S; Lukyanov KA Photoswitchable Cyan Fluorescent Protein for Protein Tracking. Nat. Biotechnol. 2004, 22, 1435–1439. [DOI] [PubMed] [Google Scholar]
- (54).Verkhusha VV; Sorkin A Conversion of the Monomeric Red Fluorescent Protein into a Photoactivatable Probe. Chem. Biol. 2005, 12, 279–285. [DOI] [PubMed] [Google Scholar]
- (55).Tsutsui H; Karasawa S; Shimizu H; Nukina N; Miyawaki A Semi-Rational Engineering of a Coral Fluorescent Protein into an Efficient Highlighter. EMBO Rep. 2005, 6, 233–238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (56).Gurskaya NG; Verkhusha VV; Shcheglov AS; Staroverov DB; Chepurnykh TV; Fradkov AF; Lukyanov S; Lukyanov KA Engineering of a Monomeric Green-to-Red Photoactivatable Fluorescent Protein Induced by Blue Light. Nat. Biotechnol. 2006, 24, 461–465. [DOI] [PubMed] [Google Scholar]
- (57).Wiedenmann J; Ivanchenko S; Oswald F; Schmitt F; Röcker C; Salih A; Spindler K-D; Nienhaus GU EosFP, a Fluorescent Marker Protein with UV-Inducible Green-to-Red Fluorescence Conversion. Proc. Natl. Acad. Sci. U. S. A. 2004, 101, 15905–15910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (58).Lukyanov KA; Chudakov DM; Lukyanov S; Verkhusha VV Photoactivatable Fluorescent Proteins. Nat. Rev. Mol. Cell Biol. 2005, 6, 885–890. [DOI] [PubMed] [Google Scholar]
- (59).Lippincott-Schwartz J; Patterson GH Photoactivatable Fluorescent Proteins for Diffraction-Limited and Super-Resolution Imaging. Trends Cell Biol. 2009, 19, 555–565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (60).Bourgeois D; Adam V Reversible Photoswitching in Fluorescent Proteins: A Mechanistic View. IUBMB Life 2012, 64, 482–491. [DOI] [PubMed] [Google Scholar]
- (61).Zhou XX; Lin MZ Photoswitchable Fluorescent Proteins: Ten Years of Colorful Chemistry and Exciting Applications. Curr. Opin. Chem. Biol. 2013, 17, 682–690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (62).Durisic N; Laparra-Cuervo L; Sandoval-Álvarez Á; Borbely JS; Lakadamyali M Single-Molecule Evaluation of Fluorescent Protein Photoactivation Efficiency Using an in Vivo Nanotemplate. Nat. Methods 2014, 11, 156–162 [DOI] [PubMed] [Google Scholar]
- (63).Wang S; Moffitt JR; Dempsey GT; Xie XS; Zhuang X Characterization and Development of Photoactivatable Fluorescent Proteins for Single-Molecule–Based Superresolution Imaging. Proc. Natl. Acad. Sci. 2014, 111, 8452–8457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (64).Adam V; Berardozzi R; Byrdin M; Bourgeois D Phototransformable Fluorescent Proteins: Future Challenges. Curr. Opin. Chem. Biol. 2014, 20, 92–102. [DOI] [PubMed] [Google Scholar]
- (65).Shcherbakova DM; Sengupta P; Lippincott-Schwartz J; Verkhusha VV Photocontrollable Fluorescent Proteins for Superresolution Imaging. Annu. Rev. Biophys. 2014, 43, 303–329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (66).Uno S; Tiwari DK; Kamiya M; Arai Y; Nagai T; Urano Y A Guide to Use Photocontrollable Fluorescent Proteins and Synthetic Smart Fluorophores for Nanoscopy. Microscopy 2015, 64, 263–277. [DOI] [PubMed] [Google Scholar]
- (67).Betzig E Proposed Method for Molecular Optical Imaging. Opt. Lett. 1995, 20, 237–239. [DOI] [PubMed] [Google Scholar]
- (68).Burns DH; Callis JB; Christian GD; Davidson ER Strategies for Attaining Superresolution Using Spectroscopic Data as Constraints. Appl. Opt. 1985, 24, 154–161. [DOI] [PubMed] [Google Scholar]
- (69).Van Oijen AM; Köhler J; Schmidt J; Müller M; Brakenhoff GJ 3-Dimensional Super-Resolution by Spectrally Selective Imaging. Chem. Phys. Lett. 1998, 292, 183–187. [Google Scholar]
- (70).Gordon MP; Ha T; Selvin PR Single-Molecule High-Resolution Imaging with Photobleaching. Proc. Natl. Acad. Sci. U. S. A. 2004, 101, 6462–6465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (71).Qu X; Wu D; Mets L; Scherer NF Nanometer-Localized Multiple Single-Molecule Fluorescence Microscopy. Proc. Natl. Acad. Sci. U. S. A. 2004, 101, 11298–11303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (72).Lidke KA; Rieger B; Jovin TM; Heintzmann R Superresolution by Localization of Quantum Dots Using Blinking Statistics. Opt Express 2005, 13, 7052–7062. [DOI] [PubMed] [Google Scholar]
- (73).Ha T; Enderle T; Ogletree DF; Chemla DS; Selvin PR; Weiss S Probing the Interaction between Two Single Molecules: Fluorescence Resonance Energy Transfer between a Single Donor and a Single Acceptor. Proc. Natl. Acad. Sci. 1996, 93, 6264–6268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (74).Ha T Single-Molecule Fluorescence Resonance Energy Transfer. Methods 2001, 25, 78–86. [DOI] [PubMed] [Google Scholar]
- (75).Roy R; Hohng S; Ha T A Practical Guide to Single-Molecule FRET. Nat. Methods 2008, 5, 507–516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (76).Bates M; Blosser T; Zhuang X Short-Range Spectroscopic Ruler Based on a Single-Molecule Optical Switch. Phys. Rev. Lett. 2005, 94, 108101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (77).Heilemann M; Margeat E; Kasper R; Sauer M; Tinnefeld P Carbocyanine Dyes as Efficient Reversible Single-Molecule Optical Switch. J. Am. Chem. Soc. 2005, 127, 3801–3806. [DOI] [PubMed] [Google Scholar]
- (78).Handbook of Biological Confocal Microscopy, 3rd ed.; Pawley JB, Ed.; Springer: New York, NY, 2006. [Google Scholar]
- (79).Gelles J; Schnapp BJ; Sheetz MP Tracking Kinesin-Driven Movements with Nanometre-Scale Precision. Nature 1988, 331, 450–453. [DOI] [PubMed] [Google Scholar]
- (80).Yildiz A Myosin V Walks Hand-Over-Hand: Single Fluorophore Imaging with 1.5-Nm Localization. Science 2003, 300, 2061–2065. [DOI] [PubMed] [Google Scholar]
- (81).Thompson RE; Larson DR; Webb WW Precise Nanometer Localization Analysis for Individual Fluorescent Probes. Biophys J 2002, 82, 2775–2783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (82).Yildiz A Kinesin Walks Hand-Over-Hand. Science 2004, 303, 676–678. [DOI] [PubMed] [Google Scholar]
- (83).Pohl DW; Denk W; Lanz M Optical Stethoscopy: Image Recording with Resolution λ/20. Appl. Phys. Lett. 1984, 44, 651–653. [Google Scholar]
- (84).Dürig U; Pohl DW; Rohner F Near-Field Optical-Scanning Microscopy. J. Appl. Phys 1986, 59, 3318–3327. [Google Scholar]
- (85).Betzig E; Lewis A; Harootunian A; Isaacson M; Kratschmer E Near Field Scanning Optical Microscopy (NSOM): Development and Biophysical Applications. Biophys. J. 1986, 49, 269–279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (86).Harootunian A; Betzig E; Isaacson M; Lewis A Super-Resolution Fluorescence near-Field Scanning Optical Microscopy. Appl. Phys. Lett. 1986, 49, 674–676. [Google Scholar]
- (87).Dunn RC Near-Field Scanning Optical Microscopy. Chem. Rev. 1999, 99, 2891–2928. [DOI] [PubMed] [Google Scholar]
- (88).Hecht B; Sick B; Wild UP; Deckert V; Zenobi R; Martin OJF; Pohl DW Scanning Near Field Optical Microscopy with Aperture Probes: Fundamentals and Applications. J. Chem. Phys. 2000, 112, 7761–7774. [Google Scholar]
- (89).Hell SW; Lindek S; Cremer C; Stelzer EH Confocal Microscopy with an Increased Detection Aperture: Type-B 4Pi Confocal Microscopy. Opt. Lett. 1994, 19, 222–224. [DOI] [PubMed] [Google Scholar]
- (90).Gustafsson MGL; Agard DA; Sedat JW; others. I5M: 3D Widefield Light Microscopy with Better than 100nm Axial Resolution. J. Microsc. 1999, 195, 10–16. [DOI] [PubMed] [Google Scholar]
- (91).Hell SW; Wichmann J Breaking the Diffraction Resolution Limit by Stimulated Emission: Stimulated-Emission-Depletion Fluorescence Microscopy. Opt. Lett. 1994, 19, 780–782. [DOI] [PubMed] [Google Scholar]
- (92).Klar TA; Hell SW Subdiffraction Resolution in Far-Field Fluorescence Microscopy. Opt. Lett. 1999, 24, 954–956. [DOI] [PubMed] [Google Scholar]
- (93).Klar TA; Jakobs S; Dyba M; Egner A; Hell SW Fluorescence Microscopy with Diffraction Resolution Barrier Broken by Stimulated Emission. Proc. Natl. Acad. Sci. 2000, 97, 8206–8210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (94).Bailey B; Farkas DL; Taylor DL; Lanni F Enhancement of Axial Resolution in Fluorescence Microscopy by Standing-Wave Excitation. Nature Publishing Group; 1993. [DOI] [PubMed] [Google Scholar]
- (95).Heintzmann R; Cremer CG Laterally Modulated Excitation Microscopy: Improvement of Resolution by Using a Diffraction Grating; Bigio IJ, Schneckenburger H, Slavik J, Svanberg K, Viallet PM, Eds.; 1999; pp 185–196. [Google Scholar]
- (96).Gustafsson MG Surpassing the Lateral Resolution Limit by a Factor of Two Using Structured Illumination Microscopy. J. Microsc. 2000, 198, 82–87. [DOI] [PubMed] [Google Scholar]
- (97).Mei E; Gao F; Hochstrasser RM Controlled Bimolecular Collisions Allow Sub-Diffraction Limited Microscopy of Lipid Vesicles. Phys. Chem. Chem. Phys. 2006, 8, 2077–2082. [DOI] [PubMed] [Google Scholar]
- (98).Herrmann K-H; Krahl D; Rust H-P A Tv System for Image Recording and Processing in Conventional Transmission Electron Microscopy. Ultramicroscopy 1978, 3, 227–235. [DOI] [PubMed] [Google Scholar]
- (99).McMullan G; Clark AT; Turchetta R; Faruqi AR Enhanced Imaging in Low Dose Electron Microscopy Using Electron Counting. Ultramicroscopy 2009, 109, 1411–1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (100).Li X; Mooney P; Zheng S; Booth CR; Braunfeld MB; Gubbens S; Agard DA; Cheng Y Electron Counting and Beam-Induced Motion Correction Enable near-Atomic-Resolution Single Particle Cryo-EM. Nat. Methods 2013, 10, 584–590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (101).Gambhir SS Molecular Imaging of Cancer with Positron Emission Tomography. Nat. Rev. Cancer 2002, 2, 683–693. [DOI] [PubMed] [Google Scholar]
- (102).Viessmann OM; Eckersley RJ; Christensen-Jeffries K; Tang MX; Dunsby C Acoustic Super-Resolution with Ultrasound and Microbubbles. Phys. Med. Biol. 2013, 58, 6447–6458. [DOI] [PubMed] [Google Scholar]
- (103).Desailly Y; Couture O; Fink M; Tanter M Sono-Activated Ultrasound Localization Microscopy. Appl. Phys. Lett. 2013, 103, 174107. [Google Scholar]
- (104).Christensen-Jeffries K; Browning RJ; Tang M-X; Dunsby C; Eckersley RJ In Vivo Acoustic Super-Resolution and Super-Resolved Velocity Mapping Using Microbubbles. IEEE Trans. Med. Imaging 2015, 34, 433–440. [DOI] [PubMed] [Google Scholar]
- (105).Errico C; Pierre J; Pezet S; Desailly Y; Lenkei Z; Couture O; Tanter M Ultrafast Ultrasound Localization Microscopy for Deep Super-Resolution Vascular Imaging. Nature 2015, 527, 499–502. [DOI] [PubMed] [Google Scholar]
- (106).Toprak E; Enderlein J; Syed S; McKinney SA; Petschek RG; Ha T; Goldman YE; Selvin PR Defocused Orientation and Position Imaging (DOPI) of Myosin V; 2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (107).Churchman LS; Ökten Z; Rock RS; Dawson JF; Spudich JA Single Molecule High-Resolution Colocalization of Cy3 and Cy5 Attached to Macromolecules Measures Intramolecular Distances through Time. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 1419–1423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (108).Geisler C; Schönle A; Von Middendorff C; Bock H; Eggeling C; Egner A; Hell SW Resolution of λ/10 in Fluorescence Microscopy Using Fast Single Molecule Photo-Switching. Appl. Phys. Mater. Sci. Process. 2007, 88, 223–226. [Google Scholar]
- (109).Föiling J; Bossi M; Bock H; Medda R; Wurm CA; Hein B; Jakobs S; Eggeling C; Hell SW Fluorescence Nanoscopy by Ground-State Depletion and Single-Molecule Return. Nat. Methods 2008, 5, 943–945. [DOI] [PubMed] [Google Scholar]
- (110).Heilemann M; van de Linde S; Schüttpelz M; Kasper R; Seefeldt B; Mukherjee A; Tinnefeld P; Sauer M Subdiffraction-Resolution Fluorescence Imaging with Conventional Fluorescent Probes. Angew. Chem. Int. Ed. 2008, 47, 6172–6176. [DOI] [PubMed] [Google Scholar]
- (111).Manley S; Gillette JM; Patterson GH; Shroff H; Hess HF; Betzig E; Lippincott-Schwartz J High-Density Mapping of Single-Molecule Trajectories with Photoactivated Localization Microscopy. Nat. Methods 2008, 5, 155–157. [DOI] [PubMed] [Google Scholar]
- (112).Hu D; Tian Z; Wu W; Wan W; Li ADQ Photoswitchable Nanoparticles Enable High-Resolution Cell Imaging: PULSAR Microscopy. J. Am. Chem. Soc 2008, 130, 15279–15281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (113).Lemmer P; Gunkel M; Baddeley D; Kaufmann R; Urich A; Weiland Y; Reymann J; Müller P; Hausmann M; Cremer C SPDM: Light Microscopy with Single-Molecule Resolution at the Nanoscale. Appl. Phys. B Lasers Opt. 2008, 93, 1–12. [Google Scholar]
- (114).Shtengel G; Galbraith JA; Galbraith CG; Lippincott-Schwartz J; Gillette JM; Manley S; Sougrat R; Waterman CM; Kanchanawong P; Davidson MW; et al. Interferometric Fluorescent Super-Resolution Microscopy Resolves 3D Cellular Ultrastructure. Proc. Natl. Acad. Sci. 2009, 106, 3125–3130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (115).Baddeley D; Jayasinghe ID; Cremer C; Cannell MB; Soeller C Light-Induced Dark States of Organic Fluochromes Enable 30 Nm Resolution Imaging in Standard Media. Biophys. J. 2009, 96, L22–L24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (116).Roeffaers MBJ; De Cremer G; Libeert J; Ameloot R; Dedecker P; Bons A-J; Bückins M; Martens JA; Sels BF; De Vos DE; et al. Super-Resolution Reactivity Mapping of Nanostructured Catalyst Particles. Angew. Chem. Int. Ed. 2009, 48, 9285–9289. [DOI] [PubMed] [Google Scholar]
- (117).Thompson MA; Biteen JS; Lord SJ; Conley NR; Moerner WE Molecules and Methods for Super-Resolution Imaging In Methods In Enzymology; Elsevier, 2010; Vol. 475, pp 27–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (118).Schwering M; Kiel A; Kurz A; Lymperopoulos K; Sprödefeld A; Krämer R; Herten D-P Far-Field Nanoscopy with Reversible Chemical Reactions. Angew. Chem. Int. Ed. 2011, 50, 2940–2945. [DOI] [PubMed] [Google Scholar]
- (119).Lew MD; Lee SF; Ptacin JL; Lee MK; Twieg RJ; Shapiro L; Moerner WE Three-Dimensional Superresolution Colocalization of Intracellular Protein Superstructures and the Cell Surface in Live Caulobacter Crescentus. Proc. Natl. Acad. Sci. 2011, 108, E1102–E1110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (120).Schoen I; Ries J; Klotzsch E; Ewers H; Vogel V Binding-Activated Localization Microscopy of DNA Structures. Nano Lett. 2011, 11, 4008–4011. [DOI] [PubMed] [Google Scholar]
- (121).Burnette DT; Sengupta P; Dai Y; Lippincott-Schwartz J; Kachar B Bleaching/Blinking Assisted Localization Microscopy for Superresolution Imaging Using Standard Fluorescent Molecules. Proc. Natl. Acad. Sci. 2011, 108, 21081–21086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (122).Pinaud F; Dahan M Targeting and Imaging Single Biomolecules in Living Cells by Complementation-Activated Light Microscopy with Split-Fluorescent Proteins. Proc. Natl. Acad. Sci. 2011, 108, E201–E210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (123).Lee MK; Williams J; Twieg RJ; Rao J; Moerner WE Enzymatic Activation of Nitro-Aryl Fluorogens in Live Bacterial Cells for Enzymatic Turnover-Activated Localization Microscopy. Chem. Sci. 2013, 4, 220–225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (124).You C; Richter CP; Löchte S; Wilmes S; Piehler J Dynamic Submicroscopic Signaling Zones Revealed by Pair Correlation Tracking and Localization Microscopy. Anal. Chem. 2014, 86, 8593–8602. [DOI] [PubMed] [Google Scholar]
- (125).Shroff H; Galbraith CG; Galbraith JA; Betzig E Live-Cell Photoactivated Localization Microscopy of Nanoscale Adhesion Dynamics. Nat Meth 2008, 5, 417–423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (126).Legant WR; Shao L; Grimm JB; Brown TA; Milkie DE; Avants BB; Lavis LD; Betzig E High-Density Three-Dimensional Localization Microscopy across Large Volumes. Nat. Methods 2016, 13, 359–365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (127).Mortensen KI; Churchman LS; Spudich JA; Flyvbjerg H Optimized Localization Analysis for Single-Molecule Tracking and Super-Resolution Microscopy. Nat. Methods 2010, 7, 377–381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (128).Dempsey GT; Vaughan JC; Chen KH; Bates M; Zhuang X Evaluation of Fluorophores for Optimal Performance in Localization-Based Super-Resolution Imaging. Nat. Methods 2011, 8, 1027–1036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (129).Simonson PD; Rothenberg E; Selvin PR Single-Molecule-Based Super-Resolution Images in the Presence of Multiple Fluorophores. Nano Lett. 2011, 11, 5090–5096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (130).Wang Y; Fruhwirth G; Cai E; Ng T; Selvin PR 3D Super-Resolution Imaging with Blinking Quantum Dots. Nano Lett. 2013, 13, 5233–5241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (131).Cox S; Rosten E; Monypenny J; Jovanovic-Talisman T; Burnette DT; Lippincott-Schwartz J; Jones GE; Heintzmann R Bayesian Localization Microscopy Reveals Nanoscale Podosome Dynamics. Nat. Methods 2011, 9, 195–200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (132).Dertinger T; Heilemann M; Vogel R; Sauer M; Weiss S Superresolution Optical Fluctuation Imaging with Organic Dyes. Angew. Chem. Int. Ed. 2010, 49, 9441–9443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (133).Dertinger T; Colyer R; Iyer G; Weiss S; Enderlein J Fast, Background-Free, 3D Super-Resolution Optical Fluctuation Imaging (SOFI). Proc. Natl. Acad. Sci. 2009, 106, 22287–22292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (134).Munck S; Miskiewicz K; Sannerud R; Menchon SA; Jose L; Heintzmann R; Verstreken P; Annaert W Sub-Diffraction Imaging on Standard Microscopes through Photobleaching Microscopy with Non-Linear Processing. J. Cell Sci 2012, 125, 2257–2266. [DOI] [PubMed] [Google Scholar]
- (135).Vaughan JC; Dempsey GT; Sun E; Zhuang X Phosphine Quenching of Cyanine Dyes as a Versatile Tool for Fluorescence Microscopy. J. Am. Chem. Soc. 2013, 135, 1197–1200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (136).Vaughan JC; Jia S; Zhuang X Ultrabright Photoactivatable Fluorophores Created by Reductive Caging. Nat Methods 2012, 9, 1181–1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (137).Bates M; Huang B; Dempsey GT; Zhuang X Multicolor Super-Resolution Imaging with Photo-Switchable Fluorescent Probes. Science 2007, 317, 1749–1753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (138).Bates M; Dempsey GT; Chen KH; Zhuang X Multicolor Super-Resolution Fluorescence Imaging via Multi-Parameter Fluorophore Detection. ChemPhysChem 2012, 13, 99–107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (139).Bock H; Geisler C; Wurm CA; Von Middendorff C; Jakobs S; Schönle A; Egner A; Hell SW; Eggeling C Two-Color Far-Field Fluorescence Nanoscopy Based on Photoswitchable Emitters. Appl. Phys. B Lasers Opt. 2007, 88, 161–165. [Google Scholar]
- (140).Heilemann M; van de Linde S; Mukherjee A; Sauer M Super-Resolution Imaging with Small Organic Fluorophores. Angew. Chem. Int. Ed. 2009, 48, 6903–6908. [DOI] [PubMed] [Google Scholar]
- (141).Dempsey GT; Bates M; Kowtoniuk WE; Liu DR; Tsien RY; Zhuang X Photoswitching Mechanism of Cyanine Dyes. J. Am. Chem. Soc. 2009, 131, 18192–18193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (142).van de Linde S; Krstić I; Prisner T; Doose S; Heilemann M; Sauer M Photoinduced Formation of Reversible Dye Radicals and Their Impact on Super-Resolution Imaging. Photochem. Photobiol. Sci. 2011, 10, 499–506. [DOI] [PubMed] [Google Scholar]
- (143).Glembockyte V; Cosa G Redox-Based Photostabilizing Agents in Fluorescence Imaging: The Hidden Role of Intersystem Crossing in Germinate Radical Ion Pairs. J. Am. Chem. Soc. 2017, 139, 13227–13233. [DOI] [PubMed] [Google Scholar]
- (144).Kuhn R; Winterstein A Reduktionen Mit Zinkstaub in Pyridin, I. Mitteil.: Umkehrbare Hydrierung Und Dehydrierung Bei Cyanin-Farbstoffen. Berichte Dtsch. Chem. Ges. B Ser. 1932, 65, 1737–1742. [Google Scholar]
- (145).Tiers GVD; Wiese J Reduced Styryl/Cyanine Dye. US3916069 (A), October 28, 1975.
- (146).Kundu K; Knight SF; Willett N; Lee S; Taylor WR; Murthy N Hydrocyanines: A Class of Fluorescent Sensors That Can Image Reactive Oxygen Species in Cell Culture, Tissue, and In Vivo. Angew. Chem. Int. Ed. 2009, 48, 299–303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (147).Lehmann M; Gottschalk B; Puchkov D; Schmieder P; Schwagerus S; Hackenberger CPR; Haucke V; Schmoranzer J Multicolor Caged DSTORM Resolves the Ultrastructure of Synaptic Vesicles in the Brain. Angew. Chem. Int. Ed. 2015, 54, 13230–13235. [DOI] [PubMed] [Google Scholar]
- (148).Michie MS; Götz R; Franke C; Bowler M; Kumari N; Magidson V; Levitus M; Loncarek J; Sauer M; Schnermann MJ Cyanine Conformational Restraint in the Far-Red Range. J. Am. Chem. Soc. 2017, 139, 12406–12409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (149).Ramette RW; Sandell EB Rhodamine B Equilibria. J Am Chem Soc 1956, 78, 4872–4878. [Google Scholar]
- (150).Chen X; Pradhan T; Wang F; Kim JS; Yoon J Fluorescent Chemosensors Based on Spiroring-Opening of Xanthenes and Related Derivatives. Chem. Rev. 2012, 112, 1910–1956. [DOI] [PubMed] [Google Scholar]
- (151).Kim HN; Lee MH; Kim HJ; Kim JS; Yoon J A New Trend in Rhodamine-Based Chemosensors: Application of Spirolactam Ring-Opening to Sensing Ions. Chem. Soc. Rev. 2008, 37, 1465–1472. [DOI] [PubMed] [Google Scholar]
- (152).Grimm JB; Muthusamy AK; Liang Y; Brown TA; Lemon WC; Patel R; Lu R; Macklin JJ; Keller PJ; Ji N; et al. A General Method to Fine-Tune Fluorophores for Live-Cell and in Vivo Imaging. Nat. Methods 2017, 14, 987–994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (153).Belov VN; Bossi ML Photoswitching Emission with Rhodamine Spiroamides for Super-Resolution Fluorescence Nanoscopies. Isr. J. Chem. 2013, 53, 267–279. [Google Scholar]
- (154).Lukinavičius G; Umezawa K; Olivier N; Honigmann A; Yang G; Plass T; Mueller V; Reymond L; Corrêa IR Jr; Luo Z-G; et al. A Near-Infrared Fluorophore for Live-Cell Super-Resolution Microscopy of Cellular Proteins. Nat. Chem. 2013, 5, 132–139. [DOI] [PubMed] [Google Scholar]
- (155).Grimm JB; Lavis LD Synthesis of Rhodamines from Fluoresceins Using Pd-Catalyzed C-N Cross-Coupling. Org. Lett. 2011, 13, 6354–6357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (156).Grimm JB; English BP; Chen J; Slaughter JP; Zhang Z; Revyakin A; Patel R; Macklin JJ; Normanno D; Singer RH; et al. A General Method to Improve Fluorophores for Live-Cell and Single-Molecule Microscopy. Nat. Methods 2015, 12, 244–250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (157).Fölling J; Belov V; Kunetsky R; Medda R; Schönle A; Egner A; Eggeling C; Bossi M; Hell SW Photochromic Rhodamines Provide Nanoscopy with Optical Sectioning. Angew. Chem. Int. Ed. 2007, 46, 6266–6270. [DOI] [PubMed] [Google Scholar]
- (158).Belov VN; Bossi ML; Fölling J; Boyarskiy VP; Hell SW Rhodamine Spiroamides for Multicolor Single-Molecule Switching Fluorescent Nanoscopy. Chem.- Eur. J 2009, 15, 10762–10776. [DOI] [PubMed] [Google Scholar]
- (159).Fölling J; Belov V; Riedel D; Schönle A; Egner A; Eggeling C; Bossi M; Hell SW Fluorescence Nanoscopy with Optical Sectioning by Two-Photon Induced Molecular Switching Using Continuous-Wave Lasers. ChemPhysChem 2008, 9, 321–326. [DOI] [PubMed] [Google Scholar]
- (160).Mitronova GY; Polyakova S; Wurm CA; Kolmakov K; Wolfram T; Meineke DNH; Belov VN; John M; Hell SW Functionalization of the Meso -Phenyl Ring of Rhodamine Dyes Through S N Ar with Sulfur Nucleophiles: Synthesis, Biophysical Characterizations, and Comprehensive NMR Analysis: Functionalization of Rhodamine Dyes. Eur. J. Org. Chem. 2015, 2015, 337–349. [Google Scholar]
- (161).Mitronova GY; Belov VN; Bossi ML; Wurm CA; Meyer L; Medda R; Moneron G; Bretschneider S; Eggeling C; Jakobs S; et al. New Fluorinated Rhodamines for Optical Microscopy and Nanoscopy. Chem.-Eur. J 2010, 16, 4477–4488. [DOI] [PubMed] [Google Scholar]
- (162).Bossi M; Fölling J; Belov VN; Boyarskiy VP; Medda R; Egner A; Eggeling C; Schönle A; Hell SW Multicolor Far-Field Fluorescence Nanoscopy through Isolated Detection of Distinct Molecular Species. Nano Lett. 2008, 8, 2463–2468. [DOI] [PubMed] [Google Scholar]
- (163).Lee MK; Rai P; Williams J; Twieg RJ; Moerner WE Small-Molecule Labeling of Live Cell Surfaces for Three-Dimensional Super-Resolution Microscopy. J. Am. Chem. Soc. 2014, 136, 14003–14006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (164).Uno S; Kamiya M; Yoshihara T; Sugawara K; Okabe K; Tarhan MC; Fujita H; Funatsu T; Okada Y; Tobita S; et al. A Spontaneously Blinking Fluorophore Based on Intramolecular Spirocyclization for Live-Cell Super-Resolution Imaging. Nat. Chem. 2014, 6, 681–689. [DOI] [PubMed] [Google Scholar]
- (165).Belov VN; Wurm CA; Boyarskiy VP; Jakobs S; Hell SW Rhodamines NN: A Novel Class of Caged Fluorescent Dyes. Angew. Chem. Int. Ed. 2010, 49, 3520–3523. [DOI] [PubMed] [Google Scholar]
- (166).Kolmakov K; Wurm C; Sednev MV; Bossi ML; Belov VN; Hell SW Masked Red-Emitting Carbopyronine Dyes with Photosensitive 2-Diazo-1-Indanone Caging Group. Photochem. Photobiol. Sci. 2012, 11, 522–532. [DOI] [PubMed] [Google Scholar]
- (167).Belov VN; Mitronova GY; Bossi ML; Boyarskiy VP; Hebisch E; Geisler C; Kolmakov K; Wurm CA; Willig KI; Hell SW Masked Rhodamine Dyes of Five Principal Colors Revealed by Photolysis of a 2-Diazo-1-Indanone Caging Group: Synthesis, Photophysics, and Light Microscopy Applications. Chem. - Eur. J 2014, 20, 13162–13173. [DOI] [PubMed] [Google Scholar]
- (168).Grimm JB; English BP; Choi H; Muthusamy AK; Mehl BP; Dong P; Brown TA; Lippincott-Schwartz J; Liu Z; Lionnet T; et al. Bright Photoactivatable Fluorophores for Single Molecule Imaging. Nat. Methods 2016, 13, 985–988. [DOI] [PubMed] [Google Scholar]
- (169).Gee KR; Weinberg ES; Kozlowski DJ Caged Q-Rhodamine Dextran: A New Photoactivated Fluorescent Tracer. Bioorg. Med. Chem. Lett. 2001, 11, 2181–2183. [DOI] [PubMed] [Google Scholar]
- (170).Kobayashi T; Urano Y; Kamiya M; Ueno T; Kojima H; Nagano T Highly Activatable and Rapidly Releasable Caged Fluorescein Derivatives. J. Am. Chem. Soc. 2007, 129, 6696–6697. [DOI] [PubMed] [Google Scholar]
- (171).Bley F; Schaper K; Görner H Photoprocesses of Molecules with 2-Nitrobenzyl Protecting Groups and Caged Organic Acids. Photochem. Photobiol. 2008, 84, 162–171. [DOI] [PubMed] [Google Scholar]
- (172).Kobayashi T; Komatsu T; Kamiya M; Campos C; González-Gaitán M; Terai T; Hanaoka K; Nagano T; Urano Y Highly Activatable and Environment-Insensitive Optical Highlighters for Selective Spatiotemporal Imaging of Target Proteins. J. Am. Chem. Soc. 2012, 134, 11153–11160. [DOI] [PubMed] [Google Scholar]
- (173).Mitchison TJ Polewards Microtubule Flux in the Mitotic Spindle: Evidence from Photoactivation of Fluorescence. J. Cell Biol. 1989, 109, 637–652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (174).Wysocki LM; Grimm JB; Tkachuk AN; Brown TA; Betzig E; Lavis LD Facile and General Synthesis of Photoactivatable Xanthene Dyes. Angew. Chem. Int. Ed. 2011, 50, 11206–11209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (175).Grimm JB; Klein T; Kopek BG; Shtengel G; Hess HF; Sauer M; Lavis LD Synthesis of a Far-Red Photoactivatable Silicon-Containing Rhodamine for Super-Resolution Microscopy. Angew. Chem. Int. Ed. 2016, 55, 1723–1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (176).Grimm JB; Sung AJ; Legant WR; Hulamm P; Matlosz SM; Betzig E; Lavis LD Carbofluoresceins and Carborhodamines as Scaffolds for High-Contrast Fluorogenic Probes. ACS Chem. Biol. 2013, 8, 1303–1310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (177).Lavis LD; Chao T-Y; Raines RT Fluorogenic Label for Biomolecular Imaging. ACS Chem. Biol. 2006, 1, 252–260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (178).Banala S; Maurel D; Manley S; Johnsson K A Caged, Localizable Rhodamine Derivative for Superresolution Microscopy. ACS Chem. Biol. 2012, 7, 289–293. [DOI] [PubMed] [Google Scholar]
- (179).Tian Z; Li ADQ Photoswitching-Enabled Novel Optical Imaging: Innovative Solutions for Real-World Challenges in Fluorescence Detections. Acc. Chem. Res. 2013, 46, 269–279. [DOI] [PubMed] [Google Scholar]
- (180).Tian Z; Wu W; Li ADQ Photoswitchable Fluorescent Nanoparticles: Preparation, Properties and Applications. ChemPhysChem 2009, 10, 2577–2591. [DOI] [PubMed] [Google Scholar]
- (181).Seefeldt B; Kasper R; Beining M; Mattay J; Arden-Jacob J; Kemnitzer N; Drexhage KH; Heilemann M; Sauer M Spiropyrans as Molecular Optical Switches. Photochem. Photobiol. Sci. 2010, 9, 213–220. [DOI] [PubMed] [Google Scholar]
- (182).Zhu M-Q; Zhu L; Han JJ; Wu W; Hurst JK; Li ADQ Spiropyran-Based Photochromic Polymer Nanoparticles with Optically Switchable Luminescence. J. Am. Chem. Soc. 2006, 128, 4303–4309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (183).Tian Z; Wu W; Wan W; Li ADQ Single-Chromophore-Based Photoswitchable Nanoparticles Enable Dual-Alternating-Color Fluorescence for Unambiguous Live Cell Imaging. J. Am. Chem. Soc. 2009, 131, 4245–4252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (184).Wan W; Zhu M-Q; Tian Z; Li ADQ Antiphase Dual-Color Correlation in a Reactant–Product Pair Imparts Ultrasensitivity in Reaction-Linked Double-Photoswitching Fluorescence Imaging. J. Am. Chem. Soc. 2015, 137, 4312–4315. [DOI] [PubMed] [Google Scholar]
- (185).Tian Z; Li ADQ; Hu D Super-Resolution Fluorescence Nanoscopy Applied to Imaging Core–Shell Photoswitching Nanoparticles and Their Self-Assemblies. Chem Commun 2011, 47, 1258–1260. [DOI] [PubMed] [Google Scholar]
- (186).Zhu M-Q; Zhang G-F; Hu Z; Aldred MP; Li C; Gong W-L; Chen T; Huang Z-L; Liu S Reversible Fluorescence Switching of Spiropyran-Conjugated Biodegradable Nanoparticles for Super-Resolution Fluorescence Imaging. Macromolecules 2014, 47, 1543–1552. [Google Scholar]
- (187).Yan J; Zhao L-X; Li C; Hu Z; Zhang G-F; Chen Z-Q; Chen T; Huang Z-L; Zhu J; Zhu M-Q Optical Nanoimaging for Block Copolymer Self-Assembly. J. Am. Chem. Soc. 2015, 137, 2436–2439. [DOI] [PubMed] [Google Scholar]
- (188).Putri RM; Fredy JW; Cornelissen JJLM; Koay MST; Katsonis N Labelling Bacterial Nanocages with Photo-Switchable Fluorophores. ChemPhysChem 2016, 17, 1815–1818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (189).Deniz E; Tomasulo M; Cusido J; Yildiz I; Petriella M; Bossi ML; Sortino S; Raymo FM Photoactivatable Fluorophores for Super-Resolution Imaging Based on Oxazine Auxochromes. J. Phys. Chem. C 2012, 116, 6058–6068. [Google Scholar]
- (190).Petriella M; Deniz E; Swaminathan S; Roberti MJ; Raymo FM; Bossi ML Superresolution Imaging with Switchable Fluorophores Based on Oxazine Auxochromes. Photochem. Photobiol. 2013, 89, 1391–1398. [DOI] [PubMed] [Google Scholar]
- (191).Cusido J; Ragab SS; Thapaliya ER; Swaminathan S; Garcia-Amorós J; Roberti MJ; Araoz B; Mazza MMA; Yamazaki S; Scott AM; et al. A Photochromic Bioconjugate with Photoactivatable Fluorescence for Superresolution Imaging. J. Phys. Chem. C 2016, 120, 12860–12870. [Google Scholar]
- (192).Zhang Y; Swaminathan S; Tang S; Garcia-Amorós J; Boulina M; Captain B; Baker JD; Raymo FM Photoactivatable BODIPYs Designed To Monitor the Dynamics of Supramolecular Nanocarriers. J. Am. Chem. Soc. 2015, 137, 4709–4719. [DOI] [PubMed] [Google Scholar]
- (193).Tomasulo M; Sortino S; Raymo FM A New Family of Photochromic Compounds Based on the Photoinduced Opening and Thermal Closing of [1,3]Oxazine Rings. J. Photochem. Photobiol. Chem. 2008, 200, 44–49. [Google Scholar]
- (194).Irie M Diarylethenes for Memories and Switches. Chem. Rev. 2000, 100, 1685–1716. [DOI] [PubMed] [Google Scholar]
- (195).Tian H; Yang S Recent Progresses on Diarylethene Based Photochromic Switches. Chem. Soc. Rev. 2004, 33, 85–97. [DOI] [PubMed] [Google Scholar]
- (196).Irie M; Fukaminato T; Matsuda K; Kobatake S Photochromism of Diarylethene Molecules and Crystals: Memories, Switches, and Actuators. Chem. Rev. 2014, 114, 12174–12277. [DOI] [PubMed] [Google Scholar]
- (197).Zou Y; Yi T; Xiao S; Li F; Li C; Gao X; Wu J; Yu M; Huang C Amphiphilic Diarylethene as a Photoswitchable Probe for Imaging Living Cells. J. Am. Chem. Soc. 2008, 130, 15750–15751. [DOI] [PubMed] [Google Scholar]
- (198).Uno K; Niikura H; Morimoto M; Ishibashi Y; Miyasaka H; Irie M In Situ Preparation of Highly Fluorescent Dyes upon Photoirradiation. J. Am. Chem. Soc. 2011, 133, 13558–13564. [DOI] [PubMed] [Google Scholar]
- (199).Li C; Hu Z; Aldred MP; Zhao L-X; Yan H; Zhang G-F; Huang Z-L; Li ADQ; Zhu M-Q Water-Soluble Polymeric Photoswitching Dyads Impart Super-Resolution Lysosome Highlighters. Macromolecules 2014, 47, 8594–8601. [Google Scholar]
- (200).Li C; Yan H; Zhao L-X; Zhang G-F; Hu Z; Huang Z-L; Zhu M-Q A Trident Dithienylethene-Perylenemonoimide Dyad with Super Fluorescence Switching Speed and Ratio. Nat. Commun. 2014, 5, 5709. [DOI] [PubMed] [Google Scholar]
- (201).Fukaminato T; Doi T; Tamaoki N; Okuno K; Ishibashi Y; Miyasaka H; Irie M Single-Molecule Fluorescence Photoswitching of a Diarylethene–Perylenebisimide Dyad: Non-Destructive Fluorescence Readout. J. Am. Chem. Soc. 2011, 133, 4984–4990. [DOI] [PubMed] [Google Scholar]
- (202).Li C; Gong W-L; Hu Z; Aldred MP; Zhang G-F; Chen T; Huang Z-L; Zhu M-Q Photoswitchable Aggregation-Induced Emission of a Dithienylethene–Tetraphenylethene Conjugate for Optical Memory and Super-Resolution Imaging. RSC Adv. 2013, 3, 8967–8972. [Google Scholar]
- (203).Gu X; Zhao E; Zhao T; Kang M; Gui C; Lam JWY; Du S; Loy MMT; Tang BZ A Mitochondrion-Specific Photoactivatable Fluorescence Turn-On AIE-Based Bioprobe for Localization Super-Resolution Microscope. Adv. Mater. 2016, 28, 5064–5071. [DOI] [PubMed] [Google Scholar]
- (204).Roubinet B; Weber M; Shojaei H; Bates M; Bossi ML; Belov VN; Irie M; Hell SW Fluorescent Photoswitchable Diarylethenes for Biolabeling and Single-Molecule Localization Microscopies with Optical Superresolution. J. Am. Chem. Soc. 2017, 139, 6611–6620. [DOI] [PubMed] [Google Scholar]
- (205).Willets KA; Ostroverkhova O; He M; Twieg RJ; Moerner WE Novel Fluorophores for Single-Molecule Imaging. J. Am. Chem. Soc. 2003, 125, 1174–1175. [DOI] [PubMed] [Google Scholar]
- (206).Willets KA; Nishimura SY; Schuck PJ; Twieg RJ; Moerner WE Nonlinear Optical Chromophores as Nanoscale Emitters for Single-Molecule Spectroscopy. Acc. Chem. Res. 2005, 38, 549–556. [DOI] [PubMed] [Google Scholar]
- (207).Lord SJ; Conley NR; Lee HD; Nishimura SY; Pomerantz AK; Willets KA; Lu Z; Wang H; Liu N; Samuel R; et al. DCDHF Fluorophores for Single-Molecule Imaging in Cells. ChemPhysChem 2009, 10, 55–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (208).Willets KA; Callis PR; Moerner WE Experimental and Theoretical Investigations of Environmentally Sensitive Single-Molecule Fluorophores. J. Phys. Chem. B 2004, 108, 10465–10473. [Google Scholar]
- (209).Lord SJ; Lu Z; Wang H; Willets KA; Schuck PJ; Lee HD; Nishimura SY; Twieg RJ; Moerner WE Photophysical Properties of Acene DCDHF Fluorophores: Long-Wavelength Single-Molecule Emitters Designed for Cellular Imaging. J. Phys. Chem. A 2007, 111, 8934–8941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (210).Soundararajan N; Platz MS Descriptive Photochemistry of Polyflorinated Azide Derivatives of Methyl Benzoate. J. Org. Chem. 1990, 55, 2034–2044. [Google Scholar]
- (211).Lord S; Lee H; Samuel R; Weber R; Liu N; Conley N; Thompson M; Twieg R; Moerner W Azido Push-Pull Fluorogens Photoactivate to Produce Bright Fluorescent Labels. J. Phys. Chem. B 2010, 114, 14157–14167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (212).Lord SJ; Conley NR; Lee HD; Samuel R; Liu N; Twieg RJ; Moerner WE A Photoactivatable Push-Pull Fluorophore for Single-Molecule Imaging in Live Cells. J. Am. Chem. Soc 2008, 130, 9204–9205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (213).Lee H; Lord S; Iwanaga S; Zhan K; Xie H; Williams J; Wang H; Bowman G; Goley E; Shapiro L; et al. Superresolution Imaging of Targeted Proteins in Fixed and Living Cells Using Photoactivatable Organic Fluorophores. J. Am. Chem. Soc. 2010, 132, 15099–15101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (214).Bittel AM; Nickerson A; Saldivar IS; Dolman NJ; Nan X; Gibbs SL Methodology for Quantitative Characterization of Fluorophore Photoswitching to Predict Superresolution Microscopy Image Quality. Sci. Rep. 2016, 6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (215).The Molecular Probes Handbook: A Guide to Fluorescent Probes and Labeling Technologies, 11 ed.; Johnson I, Ed.; Life Technologies: Carlsbad, Calif., 2010. [Google Scholar]
- (216).Shim SH; Xia C; Zhong G; Babcock HP; Vaughan JC; Huang B; Wang X; Xu C; Bi GQ; Zhuang X Super-Resolution Fluorescence Imaging of Organelles in Live Cells with Photoswitchable Membrane Probes. Proc. Natl. Acad. Sci. 2012, 109, 13978–13983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (217).Görner H Photochromism of Nitrospiropyrans: Effects of Structure, Solvent and Temperature. Phys. Chem. Chem. Phys. 2001, 3, 416–423. [Google Scholar]
- (218).Tiwari DK; Nagai T Smart Fluorescent Proteins: Innovation for Barrier-Free Superresolution Imaging in Living Cells. Dev. Growth Differ. 2013, 55, 491–507. [DOI] [PubMed] [Google Scholar]
- (219).Chudakov DM; Matz MV; Lukyanov S; Lukyanov KA Fluorescent Proteins and Their Applications in Imaging Living Cells and Tissues. Physiol. Rev. 2010, 90, 1103–1163. [DOI] [PubMed] [Google Scholar]
- (220).Subach FV; Patterson GH; Manley S; Gillette JM; Lippincott-Schwartz J; Verkhusha VV Photoactivatable MCherry for High-Resolution Two-Color Fluorescence Microscopy. Nat. Methods 2009, 6, 153–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (221).Adam V; Lelimousin M; Boehme S; Desfonds G; Nienhaus K; Field MJ; Wiedenmann J; McSweeney S; Nienhaus GU; Bourgeois D Structural Characterization of IrisFP, an Optical Highlighter Undergoing Multiple Photo-Induced Transformations. Proc. Natl. Acad. Sci. 2008, 105, 18343–18348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (222).van Thor JJ; Gensch T; Hellingwerf KJ; Johnson LN Phototransformation of Green Fluorescent Protein with UV and Visible Light Leads to Decarboxylation of Glutamate 222. Nat. Struct. Biol. 2002, 9, 37–41. [DOI] [PubMed] [Google Scholar]
- (223).Subach FV; Patterson GH; Renz M; Lippincott-Schwartz J; Verkhusha VV Bright Monomeric Photoactivatable Red Fluorescent Protein for Two-Color Super-Resolution SptPALM of Live Cells. J. Am. Chem. Soc. 2010, 132, 6481–6491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (224).Gunewardene MS; Subach FV; Gould TJ; Penoncello GP; Gudheti MV; Verkhusha VV; Hess ST Superresolution Imaging of Multiple Fluorescent Proteins with Highly Overlapping Emission Spectra in Living Cells. Biophys. J. 2011, 101, 1522–1528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (225).Nienhaus K; Nienhaus GU; Wiedenmann J; Nar H Structural Basis for Photo-Induced Protein Cleavage and Green-to-Red Conversion of Fluorescent Protein EosFP. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 9156–9159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (226).Hayashi I; Mizuno H; Tong KI; Furuta T; Tanaka F; Yoshimura M; Miyawaki A; Ikura M Crystallographic Evidence for Water-Assisted Photo-Induced Peptide Cleavage in the Stony Coral Fluorescent Protein Kaede. J. Mol. Biol. 2007, 372, 918–926. [DOI] [PubMed] [Google Scholar]
- (227).McKinney SA; Murphy CS; Hazelwood KL; Davidson MW; Looger LL A Bright and Photostable Photoconvertible Fluorescent Protein. Nat. Methods 2009, 6, 131–133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (228).McEvoy AL; Hoi H; Bates M; Platonova E; Cranfill PJ; Baird MA; Davidson MW; Ewers H; Liphardt J; Campbell RE MMaple: A Photoconvertible Fluorescent Protein for Use in Multiple Imaging Modalities. PLoS ONE 2012, 7, e51314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (229).Zhang M; Chang H; Zhang Y; Yu J; Wu L; Ji W; Chen J; Liu B; Lu J; Liu Y; et al. Rational Design of True Monomeric and Bright Photoactivatable Fluorescent Proteins. Nat. Methods 2012, 9, 727–729. [DOI] [PubMed] [Google Scholar]
- (230).Paez-Segala MG; Sun MG; Shtengel G; Viswanathan S; Baird MA; Macklin JJ; Patel R; Allen JR; Howe ES; Piszczek G; et al. Fixation-Resistant Photoactivatable Fluorescent Proteins for CLEM. Nat. Methods 2015, 12, 215–218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (231).Rosenbloom AB; Lee S-H; To M; Lee A; Shin JY; Bustamante C Optimized Two-Color Super Resolution Imaging of Drp1 during Mitochondrial Fission with a Slow-Switching Dronpa Variant. Proc. Natl. Acad. Sci. 2014, 111, 13093–13098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (232).Shroff H; Galbraith CG; Galbraith JA; White H; Gillette J; Olenych S; Davidson MW; Betzig E Dual-Color Superresolution Imaging of Genetically Expressed Probes within Individual Adhesion Complexes. Proc. Natl. Acad. Sci. 2007, 104, 20308–20313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (233).Stiel AC; Trowitzsch S; Weber G; Andresen M; Eggeling C; Hell SW; Jakobs S; Wahl MC 1.8 Å Bright-State Structure of the Reversibly Switchable Fluorescent Protein Dronpa Guides the Generation of Fast Switching Variants. Biochem. J. 2007, 402, 35–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (234).Andresen M; Stiel AC; Fölling J; Wenzel D; Schönle A; Egner A; Eggeling C; Hell SW; Jakobs S Photoswitchable Fluorescent Proteins Enable Monochromatic Multilabel Imaging and Dual Color Fluorescence Nanoscopy. Nat. Biotechnol. 2008, 26, 1035–1040. [DOI] [PubMed] [Google Scholar]
- (235).Chang H; Zhang M; Ji W; Chen J; Zhang Y; Liu B; Lu J; Zhang J; Xu P; Xu T A Unique Series of Reversibly Switchable Fluorescent Proteins with Beneficial Properties for Various Applications. Proc. Natl. Acad. Sci. 2012, 109, 4455–4460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (236).Habuchi S; Ando R; Dedecker P; Verheijen W; Mizuno H; Miyawaki A; Hofkens J Reversible Single-Molecule Photoswitching in the GFP-like Fluorescent Protein Dronpa. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 9511–9516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (237).Brakemann T; Stiel AC; Weber G; Andresen M; Testa I; Grotjohann T; Leutenegger M; Plessmann U; Urlaub H; Eggeling C; et al. A Reversibly Photoswitchable GFP-like Protein with Fluorescence Excitation Decoupled from Switching. Nat. Biotechnol. 2011, 29, 942–947. [DOI] [PubMed] [Google Scholar]
- (238).Grotjohann T; Testa I; Leutenegger M; Bock H; Urban NT; Lavoie-Cardinal F; Willig KI; Eggeling C; Jakobs S; Hell SW Diffraction-Unlimited All-Optical Imaging and Writing with a Photochromic GFP. Nature 2011, 478, 204–208. [DOI] [PubMed] [Google Scholar]
- (239).Fuchs J; Böhme S; Oswald F; Hedde PN; Krause M; Wiedenmann J; Nienhaus GU A Photoactivatable Marker Protein for Pulse-Chase Imaging with Superresolution. Nat. Methods 2010, 7, 627–630. [DOI] [PubMed] [Google Scholar]
- (240).Chudakov DM; Lukyanov S; Lukyanov KA Tracking Intracellular Protein Movements Using Photoswitchable Fluorescent Proteins PS-CFP2 and Dendra2. Nat. Protoc. 2007, 2, 2024–2032. [DOI] [PubMed] [Google Scholar]
- (241).Jensen NA; Danzl JG; Willig KI; Lavoie-Cardinal F; Brakemann T; Hell SW; Jakobs S Coordinate-Targeted and Coordinate-Stochastic Super-Resolution Microscopy with the Reversibly Switchable Fluorescent Protein Dreiklang. ChemPhysChem 2014, 15, 756–762. [DOI] [PubMed] [Google Scholar]
- (242).Rego EH; Shao L; Macklin JJ; Winoto L; Johansson GA; Kamps-Hughes N; Davidson MW; Gustafsson MG Nonlinear Structured-Illumination Microscopy with a Photoswitchable Protein Reveals Cellular Structures at 50-Nm Resolution. Proc. Natl. Acad. Sci. 2012, 109, E135–E143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (243).Zhou J; Yang Y; Zhang C Toward Biocompatible Semiconductor Quantum Dots: From Biosynthesis and Bioconjugation to Biomedical Application. Chem. Rev. 2015, 115, 11669–11717. [DOI] [PubMed] [Google Scholar]
- (244).Michalet X; Pinaud FF; Bentolila LA; Tsay JM; Doose S; Li JJ; Sundaresan G; Wu AM; Gambhir SS; Weiss S Quantum Dots for Live Cells, in Vivo Imaging, and Diagnostics. Science 2005, 307, 538–544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (245).Stefani FD; Hoogenboom JP; Barkai E Beyond Quantum Jumps: Blinking Nanoscale Light Emitters. Phys. Today 2009, 62, 34–39.20523758 [Google Scholar]
- (246).Lagerholm BC; Averett L; Weinreb GE; Jacobson K; Thompson NL Analysis Method for Measuring Submicroscopic Distances with Blinking Quantum Dots. Biophys. J. 2006, 91, 3050–3060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (247).Watanabe TM; Fukui S; Jin T; Fujii F; Yanagida T Real-Time Nanoscopy by Using Blinking Enhanced Quantum Dots. Biophys. J. 2010, 99, L50–L52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (248).Chien F-C; Kuo CW; Chen P Localization Imaging Using Blinking Quantum Dots. The Analyst 2011, 136, 1608–1613. [DOI] [PubMed] [Google Scholar]
- (249).Hoyer P; Staudt T; Engelhardt J; Hell SW Quantum Dot Blueing and Blinking Enables Fluorescence Nanoscopy. Nano Lett. 2011, 413–468. [DOI] [PubMed] [Google Scholar]
- (250).Tehrani KF; Xu J; Kner PA Multi-Color Quantum Dot Stochastic Optical Reconstruction Microscopy (QSTORM); Enderlein J, Gregor I, Gryczynski ZK, Erdmann R, Koberling F, Eds.; 2015; p 93310C. [Google Scholar]
- (251).Xu J; Tehrani KF; Kner P Multicolor 3D Super-Resolution Imaging by Quantum Dot Stochastic Optical Reconstruction Microscopy. ACS Nano 2015, 9, 2917–2925. [DOI] [PubMed] [Google Scholar]
- (252).Han G; Mokari T; Ajo-Franklin C; Cohen BE Caged Quantum Dots. J. Am. Chem. Soc. 2008, 130, 15811–15813. [DOI] [PubMed] [Google Scholar]
- (253).Irvine SE; Staudt T; Rittweger E; Engelhardt J; Hell SW Direct Light-Driven Modulation of Luminescence from Mn-Doped ZnSe Quantum Dots. Angew. Chem. Int. Ed. 2008, 47, 2685–2688. [DOI] [PubMed] [Google Scholar]
- (254).Zhu L; Zhu M-Q; Hurst JK; Li ADQ Light-Controlled Molecular Switches Modulate Nanocrystal Fluorescence. J. Am. Chem. Soc. 2005, 127, 8968–8970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (255).Harada Y; Sakurada K; Aoki T; Thomas DD; Yanagida T Mechanochemical Coupling in Actomyosin Energy Transduction Studied by in Vitro Movement Assay. J. Mol. Biol. 1990, 216, 49–68. [DOI] [PubMed] [Google Scholar]
- (256).Aitken CE; Marshall RA; Puglisi JD An Oxygen Scavenging System for Improvement of Dye Stability in Single-Molecule Fluorescence Experiments. Biophys. J. 2008, 94, 1826–1835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (257).Crawford DJ; Hoskins AA; Friedman LJ; Gelles J; Moore MJ Visualizing the Splicing of Single Pre-MRNA Molecules in Whole Cell Extract. RNA 2007, 14, 170–179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (258).Swoboda M; Henig J; Cheng H-M; Brugger D; Haltrich D; Plumeré N; Schlierf M Enzymatic Oxygen Scavenging for Photostability without PH Drop in Single-Molecule Experiments. ACS Nano 2012, 6, 6364–6369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (259).Nahidiazar L; Agronskaia AV; Broertjes J; van den Broek B; Jalink K Optimizing Imaging Conditions for Demanding Multi-Color Super Resolution Localization Microscopy. PLOS ONE 2016, 11, e0158884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (260).Schäfer P; van de Linde S; Lehmann J; Sauer M; Doose S Methylene Blue- and Thiol-Based Oxygen Depletion for Super-Resolution Imaging. Anal. Chem. 2013, 85, 3393–3400. [DOI] [PubMed] [Google Scholar]
- (261).Jones SA; Shim S-H; He J; Zhuang X Fast, Three-Dimensional Super-Resolution Imaging of Live Cells. Nat. Methods 2011, 8, 499–505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (262).Meister A; Anderson ME Glutathione. Annu. Rev. Biochem. 1983, 52, 711–760. [DOI] [PubMed] [Google Scholar]
- (263).Olivier N; Keller D; Rajan VS; Gönczy P; Manley S Simple Buffers for 3D STORM Microscopy. Biomed. Opt. Express 2013, 4, 885–899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (264).Lee SF; Vérolet Q; Fürstenberg A Improved Super-Resolution Microscopy with Oxazine Fluorophores in Heavy Water. Angew. Chem. Int. Ed. 2013, 52, 8948–8951. [DOI] [PubMed] [Google Scholar]
- (265).Klehs K; Spahn C; Endesfelder U; Lee SF; Fürstenberg A; Heilemann M Increasing the Brightness of Cyanine Fluorophores for Single-Molecule and Superresolution Imaging. ChemPhysChem 2014, 15, 637–641. [DOI] [PubMed] [Google Scholar]
- (266).Ong WQ; Citron YR; Schnitzbauer J; Kamiyama D; Huang B Heavy Water: A Simple Solution to Increasing the Brightness of Fluorescent Proteins in Super-Resolution Imaging. Chem Commun 2015, 51, 13451–13453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (267).Cloin BM; De Zitter E; Salas D; Gielen V; Folkers GE; Mikhaylova M; Bergeler M; Krajnik B; Harvey J; Hoogenraad CC; et al. Efficient Switching of MCherry Fluorescence Using Chemical Caging. Proc. Natl. Acad. Sci. 2017, 7013–7018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (268).Matsuda A; Shao L; Boulanger J; Kervrann C; Carlton PM; Kner P; Agard D; Sedat JW Condensed Mitotic Chromosome Structure at Nanometer Resolution Using PALM and EGFP-Histones. PLoS ONE 2010, 5, e12768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (269).Mei E; Hochstrasser RM High-Resolution Optical Imaging from Trajectory Time Distributions. J. Phys. Chem. B 2006, 110, 25101–25107. [DOI] [PubMed] [Google Scholar]
- (270).Gao F; Mei E; Lim M; Hochstrasser RM Probing Lipid Vesicles by Bimolecular Association and Dissociation Trajectories of Single Molecules. J. Am. Chem. Soc. 2006, 128, 4814–4822. [DOI] [PubMed] [Google Scholar]
- (271).Kuo C; Hochstrasser RM Super-Resolution Microscopy of Lipid Bilayer Phases. J. Am. Chem. Soc. 2011, 133, 4664–4667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (272).Giannone G; Hosy E; Levet F; Constals A; Schulze K; Sobolevsky AI; Rosconi MP; Gouaux E; Tampé R; Choquet D; et al. Dynamic Superresolution Imaging of Endogenous Proteins on Living Cells at Ultra-High Density. Biophys. J. 2010, 99, 1303–1310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (273).Aloi A; Vilanova N; Albertazzi L; Voets IK IPAINT: A General Approach Tailored to Image the Topology of Interfaces with Nanometer Resolution. Nanoscale 2016, 8, 8712–8716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (274).Kiuchi T; Higuchi M; Takamura A; Maruoka M; Watanabe N Multitarget Super-Resolution Microscopy with High-Density Labeling by Exchangeable Probes. Nat. Methods 2015, 12, 743–746. [DOI] [PubMed] [Google Scholar]
- (275).Riedl J; Crevenna AH; Kessenbrock K; Yu JH; Neukirchen D; Bista M; Bradke F; Jenne D; Holak TA; Werb Z; et al. Lifeact: A Versatile Marker to Visualize F-Actin. Nat. Methods 2008, 5, 605–607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (276).Wu D; Liu Z; Sun C; Zhang X Super-Resolution Imaging by Random Adsorbed Molecule Probes. Nano Lett. 2008, 8, 1159–1162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (277).Cang H; Labno A; Lu C; Yin X; Liu M; Gladden C; Liu Y; Zhang X Probing the Electromagnetic Field of a 15-Nanometre Hotspot by Single Molecule Imaging. Nature 2011, 469, 385–388. [DOI] [PubMed] [Google Scholar]
- (278).Jungmann R; Steinhauer C; Scheible M; Kuzyk A; Tinnefeld P; Simmel FC Single Molecule Kinetics and Super-Resolution Microscopy by Fluorescence Imaging of Transient Binding on DNA Origami. Nano Lett. 2010, 10, 4756–4761. [DOI] [PubMed] [Google Scholar]
- (279).Jungmann R; Avendaño MS; Woehrstein JB; Dai M; Shih WM; Yin P Multiplexed 3D Cellular Super-Resolution Imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 2014, 11, 313–318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (280).Iinuma R; Ke Y; Jungmann R; Schlichthaerle T; Woehrstein JB; Yin P Polyhedra Self-Assembled from DNA Tripods and Characterized with 3D DNA-PAINT. Science 2014, 344, 65–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (281).Dai M; Jungmann R; Yin P Optical Imaging of Individual Biomolecules in Densely Packed Clusters. Nat. Nanotechnol. 2016, 11, 798–807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (282).Jungmann R; Avendaño MS; Dai M; Woehrstein JB; Agasti SS; Feiger Z; Rodal A; Yin P Quantitative Super-Resolution Imaging with QPAINT. Nat. Methods 2016, 13, 439–442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (283).Auer A; Strauss MT; Schlichthaerle T; Jungmann R Fast, Background-Free DNA-PAINT Imaging Using FRET-Based Probes. Nano Lett. 2017, 17, 6428–6434. [DOI] [PubMed] [Google Scholar]
- (284).Agasti SS; Wang Y; Schueder F; Sukumar A; Jungmann R; Yin P DNA-Barcoded Labeling Probes for Highly Multiplexed Exchange-PAINT Imaging. Chem. Sci. 2017, 8, 3080–3091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (285).Schnitzbauer J; Strauss MT; Schlichthaerle T; Schueder F; Jungmann R Super-Resolution Microscopy with DNA-PAINT. Nat. Protoc. 2017, 12, 1198–1228. [DOI] [PubMed] [Google Scholar]
- (286).Chen B-C; Legant WR; Wang K; Shao L; Milkie DE; Davidson MW; Janetopoulos C; Wu XS; Hammer JA; Liu Z; et al. Lattice Light-Sheet Microscopy: Imaging Molecules to Embryos at High Spatiotemporal Resolution. Science 2014, 346, 1257998–1257998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (287).Szent-Gyorgyi C; Schmidt BA; Creeger Y; Fisher GW; Zakel KL; Adler S; Fitzpatrick JAJ; Woolford CA; Yan Q; Vasilev KV; et al. Fluorogen-Activating Single-Chain Antibodies for Imaging Cell Surface Proteins. Nat. Biotechnol. 2007, 26, 235–240. [DOI] [PubMed] [Google Scholar]
- (288).Yan Q; Schwartz SL; Maji S; Huang F; Szent-Gyorgyi C; Lidke DS; Lidke KA; Bruchez MP Localization Microscopy Using Noncovalent Fluorogen Activation by Genetically Encoded Fluorogen-Activating Proteins. ChemPhysChem 2014, 15, 687–695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (289).Kiel A; Kovacs J; Mokhir A; Krämer R; Herten D-P Direct Monitoring of Formation and Dissociation of Individual Metal Complexes by Single-Molecule Fluorescence Spectroscopy. Angew. Chem. Int. Ed. 2007, 46, 3363–3366. [DOI] [PubMed] [Google Scholar]
- (290).De Cremer G; Sels BF; De Vos DE; Hofkens J; Roeffaers MBJ Fluorescence Micro(Spectro)Scopy as a Tool to Study Catalytic Materials in Action. Chem. Soc. Rev. 2010, 39, 4703–4717. [DOI] [PubMed] [Google Scholar]
- (291).Chen P; Zhou X; Shen H; Andoy NM; Choudhary E; Han K-S; Liu G; Meng W Single-Molecule Fluorescence Imaging of Nanocatalytic Processes. Chem. Soc. Rev. 2010, 39, 4560–4570. [DOI] [PubMed] [Google Scholar]
- (292).Zhou X; Andoy NM; Liu G; Choudhary E; Han K-S; Shen H; Chen P Quantitative Super-Resolution Imaging Uncovers Reactivity Patterns on Single Nanocatalysts. Nat. Nanotechnol. 2012, 7, 237–241. [DOI] [PubMed] [Google Scholar]
- (293).Guerrette JP; Percival SJ; Zhang B Fluorescence Coupling for Direct Imaging of Electrocatalytic Heterogeneity. J. Am. Chem. Soc. 2013, 135, 855–861. [DOI] [PubMed] [Google Scholar]
- (294).Brox D; Kiel A; Wörner SJ; Pernpointner M; Comba P; Martin B; Herten D-P Ensemble and Single-Molecule Studies on Fluorescence Quenching in Transition Metal Bipyridine-Complexes. PLoS ONE 2013, 8, e58049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (295).Cordes T; Blum SA Opportunities and Challenges in Single-Molecule and Single-Particle Fluorescence Microscopy for Mechanistic Studies of Chemical Reactions. Nat. Chem. 2013, 5, 993–999. [DOI] [PubMed] [Google Scholar]
- (296).Lu J; Fan Y; Howard MD; Vaughan JC; Zhang B Single-Molecule Electrochemistry on a Porous Silica-Coated Electrode. J. Am. Chem. Soc. 2017, 139, 2964–2971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (297).Yoshioka K; Komatsu T; Nakada A; Onagi J; Kuriki Y; Kawaguchi M; Terai T; Ueno T; Hanaoka K; Nagano T; et al. Identification of Tissue-Restricted Bioreaction Suitable for in Vivo Targeting by Fluorescent Substrate Library-Based Enzyme Discovery. J. Am. Chem. Soc. 2015, 137, 12187–12190. [DOI] [PubMed] [Google Scholar]
- (298).Sakabe M; Asanuma D; Kamiya M; Iwatate RJ; Hanaoka K; Terai T; Nagano T; Urano Y Rational Design of Highly Sensitive Fluorescence Probes for Protease and Glycosidase Based on Precisely Controlled Spirocyclization. J. Am. Chem. Soc. 2013, 135, 409–414. [DOI] [PubMed] [Google Scholar]
- (299).Kamiya M; Asanuma D; Kuranaga E; Takeishi A; Sakabe M; Miura M; Nagano T; Urano Y β-Galactosidase Fluorescence Probe with Improved Cellular Accumulation Based on a Spirocyclized Rhodol Scaffold. J. Am. Chem. Soc 2011, 133, 12960–12963. [DOI] [PubMed] [Google Scholar]
- (300).Roldán MD; Pérez-Reinado E; Castillo F; Moreno-Vivián C Reduction of Polynitroaromatic Compounds: The Bacterial Nitroreductases. FEMS Microbiol. Rev. 2008, 32, 474–500. [DOI] [PubMed] [Google Scholar]
- (301).Halabi EA; Thiel Z; Trapp N; Pinotsi D; Rivera-Fuentes P A Photoactivatable Probe for Super-Resolution Imaging of Enzymatic Activity in Live Cells. j. Am. Chem. Soc 2017, 139, 13200–13207. [DOI] [PubMed] [Google Scholar]
- (302).English BP; Min W; van Oijen AM; Lee KT; Luo G; Sun H; Cherayil BJ; Kou SC; Xie XS Ever-Fluctuating Single Enzyme Molecules: Michaelis-Menten Equation Revisited. Nat. Chem. Biol. 2006, 2, 87–94. [DOI] [PubMed] [Google Scholar]
- (303).Xu W; Kong JS; Yeh Y-TE; Chen P Single-Molecule Nanocatalysis Reveals Heterogeneous Reaction Pathways and Catalytic Dynamics. Nat. Mater. 2008, 7, 992–996. [DOI] [PubMed] [Google Scholar]
- (304).Andoy NM; Zhou X; Choudhary E; Shen H; Liu G; Chen P Single-Molecule Catalysis Mapping Quantifies Site-Specific Activity and Uncovers Radial Activity Gradient on Single 2D Nanocrystals. J. Am. Chem. Soc. 2013, 135, 1845–1852. [DOI] [PubMed] [Google Scholar]
- (305).Ochoa MA; Chen P; Loring RF Single Turnover Measurements of Nanoparticle Catalysis Analyzed with Dwell Time Correlation Functions and Constrained Mean Dwell Times. J. Phys. Chem. C 2013, 117, 19074–19081. [Google Scholar]
- (306).Sprödefeld A; Kiel A; Herten D-P; Krämer R Monitoring Cu 2+ -Binding to a DNA-Clip-Phen Conjugate and Metal-Centered Redox Processes by a Fluorescent Reporter Group. Z. Für Anorg. Allg. Chem. 2013, 639, 1636–1639. [Google Scholar]
- (307).Laevsky GS; O’Connell CB Comparative and Practical Aspects of Localization-Based Super-Resolution Imaging In Current Protocols In Cytometry; Robinson JP, Darzynkiewicz Z, Dobrucki J, Hyun WC, Nolan JP, Orfao A, Rabinovitch PS, Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2013. [DOI] [PubMed] [Google Scholar]
- (308).Allen JR; Ross ST; Davidson MW Sample Preparation for Single Molecule Localization Microscopy. Phys. Chem. Chem. Phys. 2013, 15, 18771–1878. [DOI] [PubMed] [Google Scholar]
- (309).Kaplan C; Ewers H Optimized Sample Preparation for Single-Molecule Localization-Based Superresolution Microscopy in Yeast. Nat. Protoc. 2015, 10, 1007–1021. [DOI] [PubMed] [Google Scholar]
- (310).Ries J; Kaplan C; Platonova E; Eghlidi H; Ewers H A Simple, Versatile Method for GFP-Based Super-Resolution Microscopy via Nanobodies. Nat. Methods 2012, 9, 582–584. [DOI] [PubMed] [Google Scholar]
- (311).Mikhaylova M; Cloin BMC; Finan K; van den Berg R; Teeuw J; Kijanka MM; Sokolowski M; Katrukha EA; Maidorn M; Opazo F; et al. Resolving Bundled Microtubules Using Anti-Tubulin Nanobodies. Nat. Commun. 2015, 6, 7933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (312).Pleiner T; Bates M; Trakhanov S; Lee C-T; Schliep JE; Chug H; Böhning M; Stark H; Urlaub H; Görlich D Nanobodies: Site-Specific Labeling for Super-Resolution Imaging, Rapid Epitope-Mapping and Native Protein Complex Isolation. eLife 2015, 4, e11349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (313).Traenkle B; Rothbauer U Under the Microscope: Single-Domain Antibodies for Live-Cell Imaging and Super-Resolution Microscopy. Front. Immunol. 2017, 8, 1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (314).Pleiner T; Bates M; Görlich D A Toolbox of Anti–Mouse and Anti–Rabbit IgG Secondary Nanobodies. J. Cell Biol. 2018, 217, 1143–1154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (315).Gronemeyer T; Godin G; Johnsson K Adding Value to Fusion Proteins through Covalent Labelling. Curr. Opin. Biotechnol. 2005, 16, 453–458. [DOI] [PubMed] [Google Scholar]
- (316).Jing C; Cornish VW Chemical Tags for Labeling Proteins Inside Living Cells. Acc. Chem. Res. 2011, 44, 784–792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (317).Chen Z; Cornish VW; Min W Chemical Tags: Inspiration for Advanced Imaging Techniques. Curr. Opin. Chem. Biol. 2013, 17, 637–643. [DOI] [PubMed] [Google Scholar]
- (318).Dean KM; Palmer AE Advances in Fluorescence Labeling Strategies for Dynamic Cellular Imaging. Nat. Chem. Biol. 2014, 10, 512–523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (319).Site-Specific Protein Labeling: Methods and Protocols; Gautier A, Hinner MJ, Eds.; Methods in Molecular Biology; Humana Press : Springer: New York, NY, 2015. [DOI] [PubMed] [Google Scholar]
- (320).Griffin BA; Adams SR; Tsien RY Specific Covalent Labeling of Recombinant Protein Molecules Inside Live Cells. Science 1998, 281, 269–272. [DOI] [PubMed] [Google Scholar]
- (321).Miller LW; Cai Y; Sheetz MP; Cornish VW In Vivo Protein Labeling with Trimethoprim Conjugates: A Flexible Chemical Tag. Nat. Methods 2005, 2, 255–257. [DOI] [PubMed] [Google Scholar]
- (322).Keppler A; Gendreizig S; Gronemeyer T; Pick H; Vogel H; Johnsson K A General Method for the Covalent Labeling of Fusion Proteins with Small Molecules in Vivo. Nat. Biotechnol. 2002, 21, 86–89. [DOI] [PubMed] [Google Scholar]
- (323).Gautier A; Juillerat A; Heinis C; Corrêa IR; Kindermann M; Beaufils F; Johnsson K An Engineered Protein Tag for Multiprotein Labeling in Living Cells. Chem. Biol. 2008, 15, 128–136. [DOI] [PubMed] [Google Scholar]
- (324).Los GV; Encell LP; McDougall MG; Hartzell DD; Karassina N; Zimprich C; Wood MG; Learish R; Ohana RF; Urh M; et al. HaloTag: A Novel Protein Labeling Technology for Cell Imaging and Protein Analysis. ACS Chem. Biol. 2008, 3, 373–382. [DOI] [PubMed] [Google Scholar]
- (325).Lelek M; Di Nunzio F; Henriques R; Charneau P; Arhel N; Zimmer C Superresolution Imaging of HIV in Infected Cells with FlAsH-PALM. Proc. Natl. Acad. Sci. 2012, 109, 8564–8569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (326).Fu N; Xiong Y; Squier TC Synthesis of a Targeted Biarsenical Cy3-Cy5 Affinity Probe for Super-Resolution Fluorescence Imaging. J. Am. Chem. Soc. 2012, 134, 18530–18533. [DOI] [PubMed] [Google Scholar]
- (327).Wombacher R; Heidbreder M; van de Linde S; Sheetz MP; Heilemann M; Cornish VW; Sauer M Live-Cell Super-Resolution Imaging with Trimethoprim Conjugates. Nat. Methods 2010, 7, 717–719. [DOI] [PubMed] [Google Scholar]
- (328).Klein T; van de Linde S; Sauer M Live-Cell Super-Resolution Imaging Goes Multicolor. ChemBioChem 2012, 13, 1861–1863. [DOI] [PubMed] [Google Scholar]
- (329).Benke A; Olivier N; Gunzenhäuser J; Manley S Multicolor Single Molecule Tracking of Stochastically Active Synthetic Dyes. Nano Lett. 2012, 12, 2619–2624. [DOI] [PubMed] [Google Scholar]
- (330).Howarth M; Takao K; Hayashi Y; Ting AY Targeting Quantum Dots to Surface Proteins in Living Cells with Biotin Ligase. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 7583–7588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (331).Chamma I; Letellier M; Butler C; Tessier B; Lim K-H; Gauthereau I; Choquet D; Sibarita J-B; Park S; Sainlos M; et al. Mapping the Dynamics and Nanoscale Organization of Synaptic Adhesion Proteins Using Monomeric Streptavidin. Nat. Commun. 2016, 7, 10773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (332).Zakeri B; Fierer JO; Celik E; Chittock EC; Schwarz-Linek U; Moy VT; Howarth M Peptide Tag Forming a Rapid Covalent Bond to a Protein, through Engineering a Bacterial Adhesin. Proc. Natl. Acad. Sci. 2012, 109, E690–E697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (333).Pessino V; Citron YR; Feng S; Huang B Covalent Protein Labeling by SpyTag-SpyCatcher in Fixed Cells for Super-Resolution Microscopy. ChemBioChem 2017, 18, 1492–1495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (334).Viswanathan S; Williams ME; Bloss EB; Stasevich TJ; Speer CM; Nern A; Pfeiffer BD; Hooks BM; Li W-P; English BP; et al. High-Performance Probes for Light and Electron Microscopy. Nat. Methods 2015, 12, 568–576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (335).Traenkle B; Emele F; Anton R; Poetz O; Haeussler RS; Maier J; Kaiser PD; Scholz AM; Nueske S; Buchfellner A; et al. Monitoring Interactions and Dynamics of Endogenous Beta-Catenin With Intracellular Nanobodies in Living Cells. Mol. Cell. Proteomics 2015, 14, 707–723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (336).Braun MB; Traenkle B; Koch PA; Emele F; Weiss F; Poetz O; Stehle T; Rothbauer U Peptides in Headlock – a Novel High-Affinity and Versatile Peptide-Binding Nanobody for Proteomics and Microscopy. Sci. Rep. 2016, 6, No. 19211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (337).Virant D; Traenkle B; Maier J; Kaiser PD; Bodenhöfer M; Schmees C; Vojnovic I; Pisak-Lukáts B; Endesfelder U; Rothbauer U A Peptide Tag-Specific Nanobody Enables High-Quality Labeling for DSTORM Imaging. Nat. Commun. 2018, 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (338).Carlini L; Manley S Live Intracellular Super-Resolution Imaging Using Site-Specific Stains. ACS Chem. Biol. 2013, 8, 2643–2648. [DOI] [PubMed] [Google Scholar]
- (339).Erdmann RS; Takakura H; Thompson AD; Rivera-Molina F; Allgeyer ES; Bewersdorf J; Toomre D; Schepartz A Super-Resolution Imaging of the Golgi in Live Cells with a Bioorthogonal Ceramide Probe. Angew. Chem. Int. Ed. 2014, 53, 10242–10246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (340).Moon S; Yan R; Kenny SJ; Shyu Y; Xiang L; Li W; Xu K Spectrally Resolved, Functional Super-Resolution Microscopy Reveals Nanoscale Compositional Heterogeneity in Live-Cell Membranes. J. Am. Chem. Soc. 2017, 139, 10944–10947. [DOI] [PubMed] [Google Scholar]
- (341).Benke A; Manley S Live-Cell DSTORM of Cellular DNA Based on Direct DNA Labeling. ChemBioChem 2012, 13, 298–301. [DOI] [PubMed] [Google Scholar]
- (342).Flors C; Ravarani CNJ; Dryden DTF Super-Resolution Imaging of DNA Labelled with Intercalating Dyes. ChemPhysChem 2009, 10, 2201–2204. [DOI] [PubMed] [Google Scholar]
- (343).Xu K; Babcock HP; Zhuang X Dual-Objective STORM Reveals Three-Dimensional Filament Organization in the Actin Cytoskeleton. Nat. Methods 2012, 9, 185–188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (344).Xu K; Zhong G; Zhuang X Actin, Spectrin, and Associated Proteins Form a Periodic Cytoskeletal Structure in Axons. Science 2013, 339, 452–456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (345).Sunwoo H; Wu JY; Lee JT The Xist RNA-PRC2 Complex at 20-Nm Resolution Reveals a Low Xist Stoichiometry and Suggests a Hit-and-Run Mechanism in Mouse Cells. Proc. Natl. Acad. Sci. 2015, 112, E4216–E4225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (346).Doksani Y; Wu JY; de Lange T; Zhuang X Super-Resolution Fluorescence Imaging of Telomeres Reveals TRF2-Dependent T-Loop Formation. Cell 2013, 155, 345–356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (347).Beliveau BJ; Boettiger AN; Avendaño MS; Jungmann R; McCole RB; Joyce EF; Kim-Kiselak C; Bantignies F; Fonseka CY; Erceg J; et al. Single-Molecule Super-Resolution Imaging of Chromosomes and in Situ Haplotype Visualization Using Oligopaint FISH Probes. Nat. Commun. 2015, 6, 7147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (348).Boettiger AN; Bintu B; Moffitt JR; Wang S; Beliveau BJ; Fudenberg G; Imakaev M; Mirny LA; Wu C; Zhuang X Super-Resolution Imaging Reveals Distinct Chromatin Folding for Different Epigenetic States. Nature 2016, 529, 418–422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (349).Lubeck E; Cai L Single-Cell Systems Biology by Super-Resolution Imaging and Combinatorial Labeling. Nat. Methods 2012, 9, 743–748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (350).Jiang H; English BP; Hazan RB; Wu P; Ovryn B Tracking Surface Glycans on Live Cancer Cells with Single-Molecule Sensitivity. Angew. Chem. Int. Ed. 2015, 54, 1765–1769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (351).Raulf A; Spahn CK; Zessin PJM; Finan K; Bernhardt S; Heckel A; Heilemann M Click Chemistry Facilitates Direct Labelling and Super-Resolution Imaging of Nucleic Acids and Proteins. RSC Adv 2014, 4, 30462–30466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (352).Zessin PJM; Finan K; Heilemann M Super-Resolution Fluorescence Imaging of Chromosomal DNA. J. Struct. Biol. 2012, 177, 344–348. [DOI] [PubMed] [Google Scholar]
- (353).Spahn C; Endesfelder U; Heilemann M Super-Resolution Imaging of Escherichia Coli Nucleoids Reveals Highly Structured and Asymmetric Segregation during Fast Growth. J. Struct. Biol. 2014, 185, 243–249. [DOI] [PubMed] [Google Scholar]
- (354).Lang K; Chin JW Cellular Incorporation of Unnatural Amino Acids and Bioorthogonal Labeling of Proteins. Chem. Rev. 2014, 114, 4764–4806. [DOI] [PubMed] [Google Scholar]
- (355).Uttamapinant C; Howe JD; Lang K; Beránek V; Davis L; Mahesh M; Barry NP; Chin JW Genetic Code Expansion Enables Live-Cell and Super-Resolution Imaging of Site-Specifically Labeled Cellular Proteins. J. Am. Chem. Soc. 2015, 137, 4602–4605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (356).Kaya E; Vrabel M; Deiml C; Prill S; Fluxa VS; Carell T A Genetically Encoded Norbornene Amino Acid for the Mild and Selective Modification of Proteins in a Copper-Free Click Reaction. Angew. Chem. Int. Ed. 2012, 51, 4466–4469. [DOI] [PubMed] [Google Scholar]
- (357).Loschberger A; van de Linde S; Dabauvalle M-C; Rieger B; Heilemann M; Krohne G; Sauer M Super-Resolution Imaging Visualizes the Eightfold Symmetry of Gp210 Proteins around the Nuclear Pore Complex and Resolves the Central Channel with Nanometer Resolution. J. Cell Sci. 2012, 125, 570–575. [DOI] [PubMed] [Google Scholar]
- (358).Sigal YM; Speer CM; Babcock HP; Zhuang X Mapping Synaptic Input Fields of Neurons with Super-Resolution Imaging. Cell 2015, 163, 493–505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (359).Dani A; Huang B; Bergan J; Dulac C; Zhuang X Superresolution Imaging of Chemical Synapses in the Brain. Neuron 2010, 68, 843–856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (360).Huang B; Jones SA; Brandenburg B; Zhuang X Whole-Cell 3D STORM Reveals Interactions between Cellular Structures with Nanometer-Scale Resolution. Nat Methods 2008, 5, 1047–1052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (361).Chung J-J; Shim S-H; Everley RA; Gygi SP; Zhuang X; Clapham DE Structurally Distinct Ca2+ Signaling Domains of Sperm Flagella Orchestrate Tyrosine Phosphorylation and Motility. Cell 2014, 157, 808–822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (362).Testa I; Wurm CA; Medda R; Rothermel E; von Middendorf C; Fölling J; Jakobs S; Schönle A; Hell SW; Eggeling C Multicolor Fluorescence Nanoscopy in Fixed and Living Cells by Exciting Conventional Fluorophores with a Single Wavelength. Biophys. J. 2010, 99, 2686–2694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (363).Lampe A; Haucke V; Sigrist SJ; Heilemann M; Schmoranzer J Multi-Colour Direct STORM with Red Emitting Carbocyanines. Biol. Cell 2012, 104, 229–237. [DOI] [PubMed] [Google Scholar]
- (364).Baddeley D; Crossman D; Rossberger S; Cheyne JE; Montgomery JM; Jayasinghe ID; Cremer C; Cannell MB; Soeller C 4D Super-Resolution Microscopy with Conventional Fluorophores and Single Wavelength Excitation in Optically Thick Cells and Tissues. PLoS ONE 2011, 6, e20645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (365).Zhang Z; Kenny SJ; Hauser M; Li W; Xu K Ultrahigh-Throughput Single-Molecule Spectroscopy and Spectrally Resolved Super-Resolution Microscopy. Nat. Methods 2015, 1, 935–938. [DOI] [PubMed] [Google Scholar]
- (366).Mlodzianoski MJ; Curthoys NM; Gunewardene MS; Carter S; Hess ST Super-Resolution Imaging of Molecular Emission Spectra and Single Molecule Spectral Fluctuations. PloS One 2016, 11, e0147506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (367).Dong B; Almassalha L; Urban BE; Nguyen T-Q; Khuon S; Chew T-L; Backman V; Sun C; Zhang HF Super-Resolution Spectroscopic Microscopy via Photon Localization. Nat. Commun. 2016, 7, 12290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (368).Bongiovanni MN; Godet J; Horrocks MH; Tosatto L; Carr AR; Wirthensohn DC; Ranasinghe RT; Lee J-E; Ponjavic A; Fritz JV; et al. Multi-Dimensional Super-Resolution Imaging Enables Surface Hydrophobicity Mapping. Nat. Commun. 2016, 7, 13544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (369).Tam J; Cordier GA; Borbely JS; Sandoval Álvarez Á; Lakadamyali M Cross-Talk-Free Multi-Color STORM Imaging Using a Single Fluorophore. PLoS ONE 2014, 9, e101772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (370).Valley CC; Liu S; Lidke DS; Lidke KA Sequential Superresolution Imaging of Multiple Targets Using a Single Fluorophore. PLoS One 2015, 10, e0123941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (371).Yi J; Manna A; Barr VA; Hong J; Neuman KC; Samelson LE MadSTORM: A Superresolution Technique for Large-Scale Multiplexing at Single-Molecule Accuracy. Mol. Biol. Cell 2016, 27, 3591–3600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (372).Schueder F; Strauss MT; Hoerl D; Schnitzbauer J; Schlichthaerle T; Strauss S; Yin P; Harz H; Leonhardt H; Jungmann R Universal Super-Resolution Multiplexing by DNA Exchange. Angew. Chem. Int. Ed. 2017, 56, 4052–4055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (373).Lin D; Gagnon LA; Howard MD; Halpern AR; Vaughan JC Extended-Depth 3D Super Resolution Imaging Using Probe-Refresh STORM. Biophys. J. 2018, 114, 1980–1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (374).Hess ST; Gould TJ; Gudheti MV; Maas SA; Mills KD; Zimmerberg J Dynamic Clustered Distribution of Hemagglutinin Resolved at 40 Nm in Living Cell Membranes Discriminates between Raft Theories. Proc. Natl. Acad. Sci. 2007, 104, 17370–17375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (375).Biteen JS; Thompson MA; Tselentis NK; Bowman GR; Shapiro L; Moerner WE Super-Resolution Imaging in Live Caulobacter Crescentus Cells Using Photoswitchable EYFP. Nat. Methods 2008, 5, 947–949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (376).Hansen AS; Pustova I; Cattoglio C; Tjian R; Darzacq X CTCF and Cohesin Regulate Chromatin Loop Stability with Distinct Dynamics. eLife 2017, 6, e25776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (377).Cho W-K; Spille J-H; Hecht M; Lee C; Li C; Grube V; Cisse II Mediator and RNA Polymerase II Clusters Associate in Transcription-Dependent Condensates. Science 2018, eaar4199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (378).Huang F; Hartwich TMP; Rivera-Molina FE; Lin Y; Duim WC; Long JJ; Uchil PD; Myers JR; Baird MA; Mothes W; et al. Video-Rate Nanoscopy Using SCMOS Camera-Specific Single-Molecule Localization Algorithms. Nat. Methods 2013, 10, 653–658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (379).Wäldchen S; Lehmann J; Klein T; van de Linde S; Sauer M Light-Induced Cell Damage in Live-Cell Super-Resolution Microscopy. Sci. Rep. 2015, 5, No. 15348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (380).Takakura H; Zhang Y; Erdmann RS; Thompson AD; Lin Y; McNellis B; Rivera-Molina F; Uno S; Kamiya M; Urano Y; et al. Long Time-Lapse Nanoscopy with Spontaneously Blinking Membrane Probes. Nat. Biotechnol. 2017, 35, 773–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (381).Uno S; Kamiya M; Morozumi A; Urano Y A Green-Light-Emitting, Spontaneously Blinking Fluorophore Based on Intramolecular Spirocyclization for Dual-Colour Super-Resolution Imaging. Chem. Commun. 2018, 54, 102–105. [DOI] [PubMed] [Google Scholar]
- (382).Endesfelder U; Heilemann M Art and Artifacts in Single-Molecule Localization Microscopy: Beyond Attractive Images. Nat. Methods 2014, 11, 235–238. [DOI] [PubMed] [Google Scholar]
- (383).Whelan DR; Bell TDM Image Artifacts in Single Molecule Localization Microscopy: Why Optimization of Sample Preparation Protocols Matters. Sci. Rep. 2015, 5, No. 7924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (384).Schnell U; Dijk F; Sjollema KA; Giepmans BNG Immunolabeling Artifacts and the Need for Live-Cell Imaging. Nat. Methods 2012, 9, 152–158. [DOI] [PubMed] [Google Scholar]
- (385).Brock R; Hamelers IH; Jovin TM Comparison of Fixation Protocols for Adherent Cultured Cells Applied to a GFP Fusion Protein of the Epidermal Growth Factor Receptor. Cytometry 1999, 35, 353–362. [DOI] [PubMed] [Google Scholar]
- (386).Richter KN; Revelo NH; Seitz KJ; Helm MS; Sarkar D; Saleeb RS; D’Este E; Eberle J; Wagner E; Vogl C; et al. Glyoxal as an Alternative Fixative to Formaldehyde in Immunostaining and Super-resolution Microscopy. EMBO J. 2017, e201695709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (387).Bradbury A; Plückthun A Reproducibility: Standardize Antibodies Used in Research. Nature 2015, 518, 27–29. [DOI] [PubMed] [Google Scholar]
- (388).Baker M Blame It on the Antibodies. Nature 2015, 521, 274–276. [DOI] [PubMed] [Google Scholar]
- (389).Cranfill PJ; Sell BR; Baird MA; Allen JR; Lavagnino Z; de Gruiter HM; Kremers G-J; Davidson MW; Ustione A; Piston DW Quantitative Assessment of Fluorescent Proteins. Nat. Methods 2016, 13, 557–563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (390).Wang H; La Russa M; Qi LS CRISPR/Cas9 in Genome Editing and Beyond. Annu. Rev. Biochem. 2016, 85, 227–264. [DOI] [PubMed] [Google Scholar]
- (391).Lambert TJ; Waters JC Navigating Challenges in the Application of Superresolution Microscopy. J Cell Biol 2017, 216, 53–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (392).Baddeley D; Bewersdorf J Biological Insight from Super-Resolution Microscopy: What We Can Learn from Localization-Based Images. Annu. Rev. Biochem. 2018, 87. [DOI] [PubMed] [Google Scholar]
- (393).Holden SJ; Uphoff S; Kapanidis AN DAOSTORM: An Algorithm for High-Density Super-Resolution Microscopy. Nat. Methods 2011, 8, 279–280. [DOI] [PubMed] [Google Scholar]
- (394).Babcock H; Sigal YM; Zhuang X A High-Density 3D Localization Algorithm for Stochastic Optical Reconstruction Microscopy. Opt. Nanoscopy 2012, 1, 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (395).Mukamel EA; Babcock H; Zhuang X Statistical Deconvolution for Superresolution Fluorescence Microscopy. Biophys. J. 2012, 102, 2391–2400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (396).Sage D; Kirshner H; Pengo T; Stuurman N; Min J; Manley S; Unser M Quantitative Evaluation of Software Packages for Single-Molecule Localization Microscopy. Nat. Methods 2015, 12, 717–724. [DOI] [PubMed] [Google Scholar]
- (397).Sinkó J; Kákonyi R; Rees E; Metcalf D; Knight AE; Kaminski CF; Szabó G; Erdélyi M TestSTORM: Simulator for Optimizing Sample Labeling and Image Acquisition in Localization Based Super-Resolution Microscopy. Biomed. Opt. Express 2014, 5, 778–787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (398).Pertsinidis A; Zhang Y; Chu S Subnanometre Single-Molecule Localization, Registration and Distance Measurements. Nature 2010, 466, 647–651. [DOI] [PubMed] [Google Scholar]
- (399).Pertsinidis A; Mukherjee K; Sharma M; Pang ZP; Park SR; Zhang Y; Brunger AT; Sudhof TC; Chu S Ultrahigh-Resolution Imaging Reveals Formation of Neuronal SNARE/Munc18 Complexes in Situ. Proc. Natl. Acad. Sci. 2013, 110, E2812–E2820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (400).Bartko AP; Dickson RM Imaging Three-Dimensional Single Molecule Orientations. J. Phys. Chem. B 1999, 103, 11237–11241. [Google Scholar]
- (401).Enderlein J; Toprak E; Selvin PR Polarization Effect on Position Accuracy of Fluorophore Localization. Opt Express 2006, 14, 8111–8120. [DOI] [PubMed] [Google Scholar]
- (402).Engelhardt J; Keller J; Hoyer P; Reuss M; Staudt T; Hell SW Molecular Orientation Affects Localization Accuracy in Superresolution Far-Field Fluorescence Microscopy. Nano Lett. 2011, 11, 209–213. [DOI] [PubMed] [Google Scholar]
- (403).Lew MD; Backlund MP; Moerner WE Rotational Mobility of Single Molecules Affects Localization Accuracy in Super-Resolution Fluorescence Microscopy. Nano Lett. 2013, 13, 3967–3972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (404).Backlund MP; Lew MD; Backer AS; Sahl SJ; Grover G; Agrawal A; Piestun R; Moerner WE Simultaneous, Accurate Measurement of the 3D Position and Orientation of Single Molecules. Proc. Natl. Acad. Sci. 2012, 109, 19087–19092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (405).Lew MD; Moerner WE Azimuthal Polarization Filtering for Accurate, Precise, and Robust Single-Molecule Localization Microscopy. Nano Lett. 2014, 14, 6407–6413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (406).Backlund MP; Arbabi A; Petrov PN; Arbabi E; Saurabh S; Faraon A; Moerner WE Removing Orientation-Induced Localization Biases in Single-Molecule Microscopy Using a Broadband Metasurface Mask. Nat. Photonics 2016, 10, 459–462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (407).Balzarotti F; Eilers Y; Gwosch KC; Gynnå AH; Westphal V; Stefani FD; Elf J; Hell SW Nanometer Resolution Imaging and Tracking of Fluorescent Molecules with Minimal Photon Fluxes. Science 2017, 355, 606–612. [DOI] [PubMed] [Google Scholar]
- (408).Nieuwenhuizen RPJ; Lidke KA; Bates M; Puig DL; Grünwald D; Stallinga S; Rieger B Measuring Image Resolution in Optical Nanoscopy. Nat. Methods 2013, 10, 557–562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (409).Fox-Roberts P; Marsh R; Pfisterer K; Jayo A; Parsons M; Cox S Local Dimensionality Determines Imaging Speed in Localization Microscopy. Nat. Commun. 2017, 8, 13558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (410).Culley S; Albrecht D; Jacobs C; Pereira PM; Leterrier C; Mercer J; Henriques R Quantitative Mapping and Minimization of Super-Resolution Optical Imaging Artifacts. Nat. Methods 2018, 15, 263–266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (411).Weisenburger S; Boening D; Schomburg B; Giller K; Becker S; Griesinger C; Sandoghdar V Cryogenic Optical Localization Provides 3D Protein Structure Data with Angstrom Resolution. Nat. Methods 2017, 14, 141–144. [DOI] [PubMed] [Google Scholar]
- (412).Hauser M; Wojcik M; Kim D; Mahmoudi M; Li W; Xu K Correlative Super-Resolution Microscopy: New Dimensions and New Opportunities. Chem. Rev. 2017, 117, 7428–7456. [DOI] [PubMed] [Google Scholar]
- (413).Kopek BG; Paez-Segala MG; Shtengel G; Sochacki KA; Sun MG; Wang Y; Xu CS; van Engelenburg SB; Taraska JW; Looger LL; et al. Diverse Protocols for Correlative Super-Resolution Fluorescence Imaging and Electron Microscopy of Chemically Fixed Samples. Nat. Protoc. 2017, 12, 916–946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (414).Emanuel G; Moffitt JR; Zhuang X High-Throughput, Image-Based Screening of Pooled Genetic-Variant Libraries. Nat. Methods 2017, 14, 1159–1162. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.