Abstract
Background
Enterococcus avium is a Gram-positive pathogenic bacterium belonging to the family Enterobacteriaceae. E. avium can cause bacteremia, peritonitis, and intracranial suppurative infection. However, the mechanism of its pathogenesis and its adaptation to a special niche is still unclear.
Results
In this study, the E. avium strain 352 was isolated from human bile and whole genome sequencing was performed. The E. avium strain 352 consists of a circular 4,794,392 bp chromosome as well as an 87,705 bp plasmid. The GC content of the chromosome is 38.98%. There are 4905 and 99 protein coding sequences in the chromosome and the plasmid, respectively. The genome of the E. avium strain 352 contains number of genes reported to be associated with bile adaption, including bsh, sbcC, mutS, nifI, galU, and hupB. There are also several virulence-associated genes including esp, fss1, fss3, ecbA, bsh, lap, clpC, clpE, and clpP.
Conclusions
This study demonstrates the presence of various virulence factors of the E. avium strain 352, which has the potential to cause infections. Moreover, the genes involved in bile adaption might contribute to its ability to live in bile. Further comparative genomic studies would help to elucidate the evolution of pathogenesis of E. avium.
Keywords: E. avium, Virulence factors, Bile adaption
Background
Enterococcus avium is a Gram-positive bacterium of the genus Enterococcus and is most commonly found in birds. E. avium is also a cause of infectious diseases in humans including bacteremia, peritonitis, intracranial suppurative infection and osteomyelitis [1–5]. It was reported that E. avium is responsible for approximately 1% of infections in humans [3]. However, there is not much known about the mechanism of its pathogenesis.
Enterococcus avium was isolated from blood samples, fecal samples, spinal cords, jeotgals (a Korean fermented seafood), and scallop solutions [1, 3, 6, 7]. Thus, E. avium can adapt to various environments and this might be an important factor for its survival in humans and for subsequent infections. Currently, there are 8 draft genomes of E. avium accessible on NCBI databases. However, no studies have analyzed these genomes for the niche adaptation of E. avium.
Here, we report the first whole genome sequence of E. avium. We also analyzed the virulence-associated genes and bile stress adaptation mechanism of the E. avium strain 352.
Methods
Strain isolation and characterization
The E. avium strain 352 was isolated from a bile sample of a cholelithiasis patient. This strain was cultivated on blood plate agar under anaerobic conditions at 37 °C for 24 h. This strain was identified by 16S rRNA sequencing using the following primers including 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′). The PCR products were subsequently sequenced, and these sequences were compared against the 16S rRNA bacteria sequence database using BLAST from the NCBI website.
Genome sequencing and de novo assembly
The bacterial genomic DNA was extracted from overnight culture of the E. avium 352 using the Bacteria DNA Kit (OMEGA Bio-Tek Inc., Norcross, GA, USA) according to the manufacturer’s instructions, and quality control was subsequently carried out using TBS-380 fluorometer (Turner BioSystems Inc., Sunnyvale, CA). Then, high qualified DNA sample (OD260/280 = 1.8–2.0, > 6 μg) was utilized to construct a fragment library.
Genomic DNA (above 3 μg) was subjected to whole genome sequencing on an Illumina HiSeq Sequencer (PE150 mode) according to the sequencing protocol. Raw sequencing data was generated by Illumina base calling software CASAVA v1.8.2 (Illumina Inc. San Diego, CA, USA). Contamination reads, such as ones containing adaptors or primers were identified by Trimmomatic with default parameters. Clean data obtained by above quality control processes were used to do further analysis. Meanwhile, the whole-genome sequencing of E. avium 352 was also carried out on the single molecule real-time by the PacBio RS Platform (Pacific Biosciences of California, Inc., Menlo Park, CA, USA). A 20 K template library was generated and sequenced using standard methods.
The Illumina data were used to evaluate the complexity of the genome and correct the PacBio long reads. First, we used ABySS to peform genome assembly with multiple-Kmer parameters and obtained optimal results for the assembly [8]. Second, canu (https://github.com/marbl/canu) was used to assemble the PacBio corrected long reads [9]. Finally, GapCloser software was subsequently applied to fill the remaining local inner gaps and correct the single base polymorphism for the final assembly results [10].
Gene annotation was determined by Annotation NCBI Prokaryotic Genome Annotation Pipeline [11]. Ribosomal RNA genes were detected by RNAmer 1.2 [12] and tRNA genes were recognized via tRNAscan SE v. 2.0 [13]. The circular genomic map was produced using CGView Server [14].
Phylogenetic analysis is based on orthologous genes. First, orthologous gene families were identified by the ORTHOMCL v2.0 program (reciprocal all-by-all BLASTP analysis) with an E-value of 10−5 [15]. Second, multiple alignments were generated with the MUSCLE v3.8.31 program, and the alignments were examined visually [16]. Third, the Maximum-likelihood (ML) methods were performed for the phylogenetic analyses using PhyML 3.0, and the model GTR + G was selected for ML analyses with 500 bootstrap replicates to calculate the bootstrap values [17]. The strains used for phylogenetic tree analysis included the E. avium strain ATCC 14025 (GCA000406965.1), the E. faecalis strain ATCC 19433 (GCA000392875.1), the E. faecium strain DO (GCA000174395.2), the E. gilvus strain ATCC BAA-350 (GCA_000394615.1), the E. pseudoavium strain CBA7133 (GCA 003386455.1), the E. sulfureus strain ATCC 49903 (GCA000407025.1), the E. raffinosus strain ATCC 49464 (GCA000393895.1), the E. gallinarum strain FDAARGOS_163 (GCA001558875.2), and the Vagococcus fluvialis strain DSM 5731 (GCA003337315.1). The putative virulence related genes were identified based on the whole genome of the E. avium strain 352 using the VFDB [18].
Quality assurance
A single colony of the E. avium strain 352 was repeatedly transferred to fresh brain heart infusion (BHI) medium to obtain pure cultures. Before DNA extraction, the identity of the strain was verified through 16S rRNA gene sequencing. After the genome sequence was obtained, the 16S rDNA gene was extracted from the genome using the RNAmmer 1.2 server and then confirmed through a BLAST search of the 16S rRNA gene against the NCBI microbial 16S database.
Results and discussion
General genome features of the E. avium strain 352
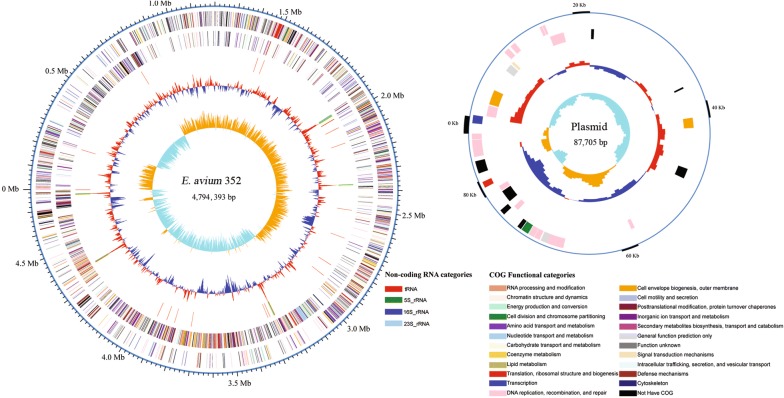
Total of 46,188,978 raw reads were obtained by Illumina HiSeq Sequencer, and 45,357,196 high quality reads were generated after quality control processes. In addition, 168,754 (1.26 Gb) high-quality reads with an average read length of 7500 bp and a 259-fold coverage were generated by PacBio sequencer. These sequences were used to assemble the genome of the E. avium strain 352 and we obtained a circular chromosome without gap. The complete genome is 4.79 Mb in size with a plasmid of 87.7 kb (Fig. 1) and the mean G + C content is 38.98%. This genome contains 4905 predicted genes as well as 18 rRNA and 68 tRNA genes, while there were 99 predicted genes in the plasmid.
Fig. 1.
Genome map of E. avium 352. Circles from the outside to inside showing: (1) DNA coordinates; (2, 3) function-based color-coded mapping of the CDSs predicted on the forward and reverse strands. Functions are color-coding; (4) tRNA genes and rRNA genes; (5) GC plot showing regions above the average (red) and below (blue); (6) GC skew
Phylogenetic analysis
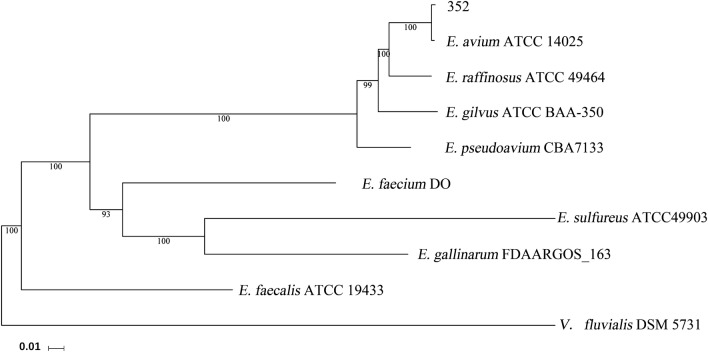
The 16S rRNA gene sequence verified the taxonomic status of the E. avium strain 352 (data not shown). To further elucidate the phylogenetic relationships, whole genome DNA-sequence-based phylogenetic analysis was carried out (Fig. 2). The genome of a highly related and similar type of E. avium strain, E. avium strain ATCC 14025, was selected as standard. The dendrogram of phylogenetic trees illustrated that the E. avium strain 352 was most closely related to the E. avium strain ATCC 14025.
Fig. 2.
Phylogenetic analysis of E. avium 352
Identification of genes related to bile stress
Bile salts have potent antimicrobial activity via damaging membranes and DNA. Thus, bacteria must have intrinsic adapted mechanisms to survive in bile and subsequently cause biliary tract infections [19]. Genomic analysis of the E. avium strain 352 showed the presence of numerous genes that may determine its bile resistance properties (Table 1). The presence of the genes sbcC, mutS and nifI involved in bile resistance in Gram-positive bacteria was identified [19]. It is interesting that there were two bsh genes encoding bile salt hydrolase with a protein sequence identity of 92.9% in the genome of the E. avium strain 352. This result indicated that the BSH might be play an important role in niche-specific adaptation for bile [20]. There were also some genes, including galU and hupB, involved in bile resistance in Gram-negative bacteria [19]. Further studies are needed to verify its genetic properties and evolution traits.
Table 1.
Putative genes for bile adaptation in E. avium 352
| Gene name | Fuction/putative fuction |
|---|---|
| hupB | HU family DNA-binding protein |
| galU | UDP-glucose-pyrophosphorylase |
| sbcC | Exonuclease SbcC |
| mutS | DNA mismatch repair |
| bsh1 | Bile salt hydrolase |
| bsh2 | Bile salt hydrolase |
| nifI | Pyruvate: ferredoxin oxidoreductase |
Analysis of virulence associated genes
Further screening the genome of the E. avium strain 352 for putative virulence-associated genes was conducted by aligning gene sequences to the virulence factor database (Table 2). There are surface protein encoded genes including esp, fss1 and fss3. The E. avium strain 352 also contains the conservative heat shock protein genes clpC, clpE, and clpP [21]. The ecbA gene encoding a collagen binding MSCRAMM (acronym for microbial surface components recognizing adhesive matrix molecules) and gene lap encoding a listeria adhesion protein were found in the genome and might be contribute to adherence to the host tissue [22, 23]. The bsh gene encoding a bile salt hydrolase was also a virulence related factor in Listeria monocytogenes [24]. The clinical significance of this finding warrants further investigation.
Table 2.
Putative virulence associated genes in E. avium 352 predicted by VFDB
| Gene name | Function/putative function | Score | E value |
|---|---|---|---|
| esp | Enterococcal surface protein | 3907 | 0 |
| fss3 | Enterococcus faecalis surface protein Fss3 | 3749 | 0 |
| ecbA | Collagen binding MSCRAMM | 3416 | 0 |
| bsh | Bile salt hydrolase | 274 | 6e−70 |
| lap | Listeria adhesion protein | 129 | 2e−26 |
| clpE | ATP-dependent protease | 127 | 9e−26 |
| clpP | ATP-dependent c1p protease peoteolytic subunit | 117 | 8e−23 |
| fss1 | Enterococcus faecalis surface protein Fss1 | 107 | 8e−20 |
| clpC | Endopeptidase C1p ATP-binding chain C | 76 | 3e−10 |
Authors’ contributions
TY and LXL designed the study; LXL and QLZ isolated and identified the E. avium 352; TY and LXL analyzed data and wrote the manuscript; PW revised the manuscript; XLZ support this study and revised the manuscript. All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing of interests
The authors declare that they have no competing interests.
Availability of data and materials
The completed genome sequence of E. avium 352 has been deposited into GenBank database with accession number CP034169 (chromosome) and CP034168 (plasmid), respectively.
Ethics approval and consent to participate
This study was approved by the ethics committees in Qilu hospital of Shandong University.
Funding
This research was supported by the Key Research and Development Program of Shandong Province (2017CXGC1215) and National Natural Science Foundation of China (81600428).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Tao Yu, Email: yutao@medmail.com.
Lixiang Li, Email: lilixiang@sdu.edu.cn.
Qilin Zhao, Email: qilinandyzhao@163.com.
Peng Wang, Email: wangpeng_qd@163.com.
Xiuli Zuo, Email: zuoxiuli@sdu.edu.cn.
References
- 1.Cottagnoud P, Rossi M. Enterococcus avium osteomyelitis. Clin Microbiol Infect. 1998;4(5):290. doi: 10.1111/j.1469-0691.1998.tb00060.x. [DOI] [PubMed] [Google Scholar]
- 2.Na S, et al. Enterococcus avium bacteremia: a 12-year clinical experience with 53 patients. Eur J Clin Microbiol Infect Dis. 2012;31(3):303–310. doi: 10.1007/s10096-011-1311-1. [DOI] [PubMed] [Google Scholar]
- 3.Okada A, Hangai M, Oda T. Bacteremia with an iliopsoas abscess and osteomyelitis of the femoral head caused by Enterococcus avium in a patient with end-stage kidney disease. Intern Med. 2015;54(6):669–674. doi: 10.2169/internalmedicine.54.3576. [DOI] [PubMed] [Google Scholar]
- 4.Yildirmak T, et al. Community-acquired intracranial suppurative infections: a 15-year report. Surg Neurol Int. 2014;5:142. doi: 10.4103/2152-7806.141891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ugur AR, et al. Enterococcus avium peritonitis in a child on continuous ambulatory peritoneal dialysis. Perit Dial Int. 2014;34(1):127–128. doi: 10.3747/pdi.2012.00241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shin NR, et al. Isolation and characterization of human intestinal Enterococcus avium EFEL009 converting rutin to quercetin. Lett Appl Microbiol. 2016;62(1):68–74. doi: 10.1111/lam.12512. [DOI] [PubMed] [Google Scholar]
- 7.Yang H, et al. Accumulation of gamma-aminobutyric acid by Enterococcus avium 9184 in scallop solution in a two-stage fermentation strategy. Microb Biotechnol. 2016;9(4):478–485. doi: 10.1111/1751-7915.12301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jackman SD, et al. ABySS 2.0: resource-efficient assembly of large genomes using a Bloom filter. Genome Res. 2017;27(5):768–777. doi: 10.1101/gr.214346.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Koren S, et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27(5):722–736. doi: 10.1101/gr.215087.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Luo R, et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience. 2012;1(1):18. doi: 10.1186/2047-217X-1-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tatusova T, et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44(14):6614–6624. doi: 10.1093/nar/gkw569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lagesen K, et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35(9):3100–3108. doi: 10.1093/nar/gkm160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25(5):955–964. doi: 10.1093/nar/25.5.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Grant JR, Stothard P. The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res. 2008;36(Web Server issue):W181–W184. doi: 10.1093/nar/gkn179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li L, Stoeckert CJ, Jr, Roos DS. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 2003;13(9):2178–2189. doi: 10.1101/gr.1224503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Guindon S, et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59(3):307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- 18.Chen L, et al. VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 2016;44(D1):D694–D697. doi: 10.1093/nar/gkv1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Begley M, Gahan CG, Hill C. The interaction between bacteria and bile. FEMS Microbiol Rev. 2005;29(4):625–651. doi: 10.1016/j.femsre.2004.09.003. [DOI] [PubMed] [Google Scholar]
- 20.Bi J, et al. Bile salt tolerance of Lactococcus lactis is enhanced by expression of bile salt hydrolase thereby producing less bile acid in the cells. Biotechnol Lett. 2016;38(4):659–665. doi: 10.1007/s10529-015-2018-7. [DOI] [PubMed] [Google Scholar]
- 21.Cassenego AP, et al. The CtsR regulator controls the expression of clpC, clpE and clpP and is required for the virulence of Enterococcus faecalis in an invertebrate model. Antonie Van Leeuwenhoek. 2016;109(9):1253–1259. doi: 10.1007/s10482-016-0727-0. [DOI] [PubMed] [Google Scholar]
- 22.Yang J, et al. Prevalence of diverse clones of vancomycin-resistant Enterococcus faecium ST78 in a Chinese Hospital. Microb Drug Resist. 2016;22(4):294–300. doi: 10.1089/mdr.2015.0069. [DOI] [PubMed] [Google Scholar]
- 23.Drolia R, et al. Listeria adhesion protein induces intestinal epithelial barrier dysfunction for bacterial translocation. Cell Host Microbe. 2018;23(4):470–484. doi: 10.1016/j.chom.2018.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Larsen N, Jespersen L. Expression of virulence-related genes in listeria monocytogenes grown on danish hard cheese as affected by NaCl content. Foodborne Pathog Dis. 2015;12(6):536–544. doi: 10.1089/fpd.2014.1930. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The completed genome sequence of E. avium 352 has been deposited into GenBank database with accession number CP034169 (chromosome) and CP034168 (plasmid), respectively.