Abstract
We have identified effects of elevated maternal cortisol (induced by maternal infusion 1 mg·kg−1·day−1) on fetal cardiac maturation and function using an ovine model. Whereas short-term exposure (115–130-day gestation) increased myocyte proliferation and Purkinje fiber apoptosis, infusions until birth caused bradycardia with increased incidence of arrhythmias at birth and increased perinatal death, despite normal fetal cortisol concentrations from 130 days to birth. Statistical modeling of the transcriptomic changes in hearts at 130 and 140 days suggested that maternal cortisol excess disrupts cardiac metabolism. In the current study, we modeled pathways in the left ventricle (LV) and interventricular septum (IVS) of newborn lambs after maternal cortisol infusion from 115 days to birth. In both LV and IVS the transcriptomic model indicated over-representation of cell cycle genes and suggested disruption of cell cycle progression. Pathways in the LV involved in cardiac architecture, including SMAD and bone morphogenetic protein (BMP) were altered, and collagen deposition was increased. Pathways in IVS related to metabolism, calcium signaling, and the actin cytoskeleton were altered. Comparison of the effects of maternal cortisol excess to the effects of normal maturation from day 140 to birth revealed that only 20% of the genes changed in the LV were consistent with normal maturation, indicating that chronic elevation of maternal cortisol alters normal maturation of the fetal myocardium. These effects of maternal cortisol on the cardiac transcriptome, which may be secondary to metabolic effects, are consistent with cardiac remodeling and likely contribute to the adverse impact of maternal stress on perinatal cardiac function.
Keywords: cardiac, fetus, maturation, microarray, pregnancy
INTRODUCTION
Chronic maternal stress, including exposure to violence or loss of a relative (34, 78), as well as Cushing’s syndrome (15, 79) during pregnancy are associated with increased risk of adverse obstetrical outcomes, including premature delivery and stillbirth. In previous studies we have found an unexpectedly high incidence of fetal and neonatal demise at term in an ovine model of pregnancy in which the ewe is chronically infused with 1 mg·kg−1·day−1 cortisol (CORT) in late gestation (32). This infusion doubles maternal cortisol concentrations, producing concentrations similar to those during ovine maternal stress (31, 84, 85).
In sheep and human fetuses, cardiomyocytes become terminally differentiated and transition in metabolic activity during the last 20% of gestation and the first days of postnatal life (4, 8, 30). The fetal heart requires glucocorticoid action for normal development (64), but our results suggest that the heart may also be susceptible to remodeling by corticosteroid overexposure (1, 16, 59). We have previously used systems biology to model statistically the transcriptomic changes in hearts of fetuses exposed to CORT to discover key pathways involved in the pathological alterations. In our previous studies, after 15 days of CORT (from 115 to 130 days of gestation), the genes differentially regulated (DR) indicated alterations in pathways relating to muscle cell apoptosis and cellular calcium homeostasis, consistent with the histological findings from these hearts (16, 60). When the treatment was extended until near term (~15 additional days), over representation of genes in pathways related to cardiac metabolism and mitochondrial function were present. Approximately one-fifth of the genes DR in CORT fetuses were associated with the mitochondrial function, and further analysis revealed significant decreases in cardiac mitochondrial content (60). The majority of the DR genes were not changed in the same direction as in normal maturation, suggesting that the exposure to excess maternal cortisol altered the trajectory of normal maturation (60).
More recently, we have found that CORT decreases fetal aortic pressure and heart rate and prolongs the duration of the P wave and PR interval in the hour before birth; we also found increased wall thickness in both the left ventricle (LV) and interventricular septum (IVS) (1). We have also found that amino acid and tricarboxylic acid cycle metabolism as well as lipid and glycerophospholipid metabolism are altered in the newborn cardiac tissue in this model (81). Thus, our data have revealed that the transition from in utero to ex utero life is a particularly vulnerable period in the CORT-overexposed fetal heart. Interestingly, these effects at birth occur despite a normalization of plasma CORT in the fetuses after 130 days, as the fetal adrenal matures. We posit that systems modeling of differentially expressed genes in the newborns would reveal the adverse cellular and molecular events occurring during this transition as a consequence of the prior cortisol overexposure or as a result of continuing maternal cortisol excess and metabolic effects. We tested the hypothesis that the models of biological responses and molecular pathways predicted from the patterns of change in gene expression in the LV and IVS would reveal mechanisms underlying changes in cardiac metabolism and enlargement at birth. We focused in this study on the IVS and the LV because of the observed changes in wall dimension and our previous studies in which we reported effects on septal Purkinje apoptosis and changes in genes involved in mitochondrial metabolism in the septum.
MATERIALS AND METHODS
Animal treatment.
All animal use in this study was approved by the Institutional Animal Care and Use Committee of the University of Florida. Cardiac tissue was collected at necropsy of newborn lambs whose mothers were either continuously infused with 1 mg·kg−1·day−1 CORT (cortisol sodium succinate; Pfizer, New York, NY) beginning at day 115 of gestation until the birth of the lamb) or were not infused (Control). All ewes and their fetuses underwent surgical and sampling procedures as previously described (1, 60); this included placement of fetal arterial catheters for blood sampling and a telemetry device for continuous recording of fetal blood pressure, amniotic pressure, and fetal ECG. Blood pressure and heart rate data in these fetuses have been previously reported. For this transcriptomic analysis tissues from 5 CORT and 5 control lambs were used. Maternal plasma cortisol concentrations averaged 17.5 ± 1.8 ng/ml in these CORT ewes and 8.0 ± 0.9 ng/ml in the Control ewes. Fetal plasma cortisol concentrations were higher in the CORT group at 125 days (CORT: 13.7 ± 4.6 ng/ml, Controls: 9.2 ± 3.3 ng/ml) but were not different between the groups at later time points; on the day of birth CORT was similar in both groups (CORT: 88 ± 28 ng/ml; Control: 62 ± 18 ng/ml). Age at delivery was also not different (CORT: 144 ± 2 days; Control: 142 ± 1 day gestational age). Although fetal heart rate and mean arterial pressure were significantly reduced on the day of birth in the CORT fetuses, fetal aortic arterial pressure was not different in the CORT fetuses over the entire 14 days before delivery (mean systolic aortic pressure over 14 days before birth CORT: 53.4 ± 3.8 mmHg, Control: 59.5 ± 3.0 mmHg; mean diastolic aortic pressure CORT: 36.2 ± 5.0 mmHg, Control: 39.6 ±2.9 mmHg) (1). In this cohort, all fetuses survived delivery; one CORT lamb died 10 min after delivery, and necropsy was immediately performed. Two other CORT lambs were weak when found, and two were standing and appeared normal. Four of five Control lambs were also standing and healthy when found; one was weak and not standing in the first hour. Overall CORT lambs were euthanized at 133 ± 89 min after birth (range 10–480 min), and Control lambs were euthanized at 171 ± 66 min after birth (range 45–375 min); the difference in timing of euthanasia is not significantly different.
The newborn lambs were euthanized via intravenously administered Euthasol solution (Euthasol; Fort Worth, TX). The heart was quickly removed, weighed, and dissected. A cross-section of heart including both ventricles was cut for histology, and then samples of the IVS and LV free wall were dissected and snap-frozen in liquid nitrogen and stored at −80°C until extraction of the RNA. Total RNA was extracted from homogenized samples of each LV and IVS using TRIzol and purified, including on-column DNase digestion (Qiagen RNeasy Plus kits; Qiagen Sciences, Germantown, MD). RNA integrity numbers for the RNA samples were 7.2–9.0 for both IVS and LV. RNA (100 ng) was labeled with Cy3 using the Agilent Low Input QuickAmp Labeling kit to generate cRNA, hybridized to the Agilent-019921 Sheep Gene Expression Microarray 8×15k (G4813A GPL14112), and scanned at the Interdisciplinary Center for Biotechnology Research core at the University of Florida. Our group has described the annotation for this array platform in previous publications (58), and the annotation has been updated as more probes have been identified.
Microarray data analysis.
Raw microarray data were processed using the Limma package in R software (57a, 62, 69), to perform background correction and data normalization using the quantile method. Probes with low expression (less than 10% brighter than the negative control probes) were excluded in the analysis. The remaining probes were retained for further statistical analysis using a moderated t-test that employs an empirical Bayes method for small sample size (P ≤ 0.05) (68). The effect of CORT was analyzed by comparing the CORT group to the Control group. This analysis identified 386 DR genes in the LV and 187 DR genes in the IVS by the maternal CORT infusion. The raw normalized array data for this experiment have been deposited in the National Center for Biotechnology Information’s Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE110470 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE110470).
Network and enrichment analysis.
Gene Set Enrichment Analysis was used to identify over-represented biological processes, molecular functions, and cellular components, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, as well as to predict transcription factor (TF) binding sites associated with the DR genes by CORT treatment in the newborn lambs. These tests used P < 0.05 as the criterion for statistical significance after Benjamini-Hochberg multiple test correction and accepting at least two genes per category (83, 87). The ClueGO (v2.5.2) plugin was used in Cytoscape software to perform a detailed network inference of the DR genes and gene ontology analysis (7, 47, 67); P ≤ 0.05 was used as the criterion for this analysis.
To assess whether CORT is upregulating or downregulating genes normally associated with the process of birth, the DR genes at birth with CORT overexposure in utero were compared with the genes whose expression is altered in Control fetuses in the last days of life, during the normal preparturient surge in plasma cortisol concentrations (i.e., control fetuses at ~142 days gestation compared with the newborn control lambs). The gene expression in the control fetuses at ~142 days gestation and at birth have been previously reported, and the raw data are available [GEO Series accession no. GSE117343; (82)]. The overlap of DR genes between the two data sets was analyzed using Venny (52); pathways for genes changing in the same or opposite direction were then analyzed using WebGestalt.
Immunoblot analysis.
Frozen tissue samples from the IVS and LV (~50 mg) were pulverized and homogenized in protein extraction buffer (50 mM Tris, 150 mM NaCl, 1% Nonidet P-40, 1 mM Na3VO4, phosphatase inhibitor cocktail 2 and 3, and protease inhibitor cocktail, cat. nos. P-5726, P-0044, and P-8340; Sigma-Aldrich). The protein concentration of the homogenate was estimated (Pierce 660 reagents, ThermoFisher Scientific) using bovine serum albumin (cat. no. 23209, ThermoFisher Scientific) as the standard. Aliquots of protein homogenates were stored at −80°C until use. Immunoblotting analysis was carried out using a Li-COR Odyssey system (LICOR Biosciences). The quantity of protein loaded was within the linear range of expression of the protein of interest. Expression of mitochondrial proteins was quantified using primary antibodies against electron transport chain antibody cocktail (cat. no. ab-110413 total OXPHOS cocktail, Abcam; monoclonal antibodies to NADH ubiquinone oxidoreductase subunit B8 of Complex I, NDUFB8; succinate dehydrogenase complex subunit 2 of Complex II, SDHB; ubiquinol cytochrome c reductase core protein 2 of Complex III, UQCRC2; cytochrome c oxidase subunit 1 of Complex IV, MTCO1; ATP synthase subunit α, ATP5A), cytochrome c oxidase subunit 4 (COX4, monoclonal antibody; cat. no. 4850, Cell Signaling), cytochrome c (monoclonal antibody, cat. no. ab-13575, Abcam), pyruvate dehydrogenase E1a (PDHE1a, monoclonal antibody, cat. no. ab-110334, Abcam), and phosphorylated pyruvate dehydrogenase E1a (pPDHE1A, polyclonal antibody, cat. no. ab-92696, Abcam) with a fluorochrome‐conjugated secondary antibody. Detection of fluorescence was performed with the Odyssey Infrared Imaging System (LICOR Biosciences). This method allows the correction of the protein densities for specific proteins using the total abundance of protein in the same lane (measured using Revert total protein stain; LICOR Biosciences). All antibodies were validated for use in sheep by testing for the appearance of antibody at the correct molecular weight as a single band, and linearity of the signal over a range of protein concentrations from 0.3 to 1.3 μg/μl. The effect of CORT on protein expression was determined using Student’s t-test. The criterion for significance was P ≤ 0.05.
Histology and immunohistofluorescent analysis.
At necropsy a section ~6-mm thick was cut along the short axis of the fetal heart and fixed in 4% phosphate-buffered paraformaldehyde for 12–24 h. The tissue was postfixed in 70% reagent alcohol, then embedded in paraffin. Five-micron-thick sections were cut for histological and immunohistological analysis. Collagen deposition in the hearts was determined using picrosirius red staining as previously described (59), imaged using a circular polarized light, and analyzed using Image J (66). Five images were recorded from the LV of each animal at ×10 magnification on an Olympus BX 41 microscope and DP 71 camera. The collagen spectra of the circular polarized light images were isolated and measured using the color threshold and binary tools in Image J. The total tissue area was measured after enhancing the image contrast by 5% and creating an 8-bit binary image. All images were subjected to the same threshold and correction criteria. Collagen density was expressed as collagen area/total tissue area. Proliferation was quantified as Ki-67 staining of nuclei (1:100 dilution, cat. no. ab-15580, Abcam); nuclei were stained with Hoechst (1:1,000 dilution, cat. no. H-1399, Invitrogen). Five images collected at ×10 magnification from the LV and IVS of each animal were imaged, and the mean percentage of Ki-67-stained nuclei in LV and IVS were calculated in each animal.
Assessment of mitochondrial area was performed using an antibody against COX4 (cat. no. 4850, Cell Signaling) as previously described using a red fluorescent (Cy5)-labeled secondary antibody (60), with the addition of wheat germ agglutinin (Wheat Germ Agglutinin, Oregon Green 488 Conjugate; Invitrogen no. W-6748) and DAPI (Invitrogen no. D-1306). Five images were taken from the IVS of each animal; images were collected through blue, green, and red filters (350, 488, and 647 nm) at ×40 magnification. The merged images were adjusted to decrease the blue intensity by 50% to correct for the difference in intensity of DAPI staining. The mitochondrial stained area was measured by converting the images taken at 647 nm to an RGB stack in Image J, with measurement in the red channel using the Triangle thresholding parameter. The nuclear area was measured by converting the images taken at 350 nm to an RGB stack in Image J. The “B” (blue) channel was then measured using the Renyi Entropy thresholding parameter. The cell area was measured by converting the images taken at 488 nm to an RGB stack in Image J. The “G” (green) channel was then measured using the Huang thresholding parameter. The nuclear (DAPI) and cell (wheat germ agglutinin) areas were combined as a measure for tissue area. All images were subjected to the same threshold and correction criteria. The COX4-positive area/tissue area was calculated for each animal as the mean of the five images analyzed.
The average areas of collagen and COX IV and Ki-67-stained nuclei were analyzed to determine the effect of fetal exposure to excess maternal cortisol using Student’s t-test. The criterion for significance was P ≤ 0.05.
RESULTS
CORT induced significant changes in gene expression in both the LV and IVS of the newborn lambs. Three hundred and eighty-six genes were differentially regulated in IVS of CORT newborns: 166 genes were upregulated, and 220 genes were downregulated (Supplemental Table S1; Supplemental Material for this article is available online at the Journal website). In the IVS, 187 genes were differentially regulated in CORT newborns: 62 genes were upregulated and 125 genes were downregulated (Supplemental Table S2).
Patterns of change in gene expression in the newborn lamb LV after excess CORT exposure in utero.
Gene ontology analysis of the genes upregulated in CORT newborn LV revealed biological processes relating to general terms such as cellular component organization of biogenesis (60 genes, adj. P = 3.31E-02), regulation of cell death [23 genes, adjusted (adj.) P = 4.31E-02] response to organic substance (36 genes; adj. P = 1.08E-02), positive regulation of cell process (46 genes, adj. P = 4.31E-02), positive regulation of metabolic process (33 genes; adj. P = 3.31E-02), response to chemical stimulus (48 genes; adj. P = 1.08E-02), and response to stress (46 genes, adj. P = 3.31E-02). More specifically, the upregulated pathways included response to steroid hormones, negative regulation of cell proliferation and negative regulation of apoptotic processes, and extracellular matrix organization (Table 1). The upregulated genes were associated with molecular processes such as SMAD binding and histone demethylase activity. The triglyceride-rich lipoprotein particle, with APOE and APOA1, was also identified as were genes for coatomer proteins involved in Golgi to endoplasmic reticulum transport (COPA, COPB2). KEGG pathways significantly associated with the genes upregulated in CORT newborn LV were metabolic pathways, cancer, cell cycle, Wnt signaling pathway, hedgehog signaling, insulin signaling, melanogenesis, the transforming growth factor (TGF)-β signaling pathway, and protein processing in the endoplasmic reticulum (Table 2). Significant pathways with two to three genes in them included amino sugar and nucleotide metabolism (CYB5R1 and UGDH), PPAR signaling (PPARD and APOA1), adherens junction (SMAD2 and SMAD4), phosphatidylinositol signaling (CALM1, CALM2, and PI4K2B), and glutathione metabolism (GPX3, GSTZ1). Although most of the identified cell components were nonspecific, including cytoplasm, nucleoplasm, nucleolus, Golgi, and endoplasmic reticulum, the model also identified differences in gene expression in striated muscle and the troponin complex pathways, notably the troponin I and T genes, TNNI2 and TNNT2, which were upregulated by CORT.
Table 1.
Biological processes and molecular functions overrepresented by the upregulated differentially regulated genes in neonatal left ventricle with in utero exposure to increased maternal cortisol
| Processes or Functions | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Biological process | |||
| Positive regulation of DNA-dependent DNA replication | 2 | 3.31E-02 | BMP4, BMP6 |
| Negative regulation of apoptotic process | 15 | 3.31E-02 | GLI3, NRAS, AATF, CXCR7M, FOXS1, ITCH, MTDH, HTATIP2, APOE, CCL21, BMP4, ARNT2, CD44, NOL3, PRNP |
| SMAD protein signal transduction | 3 | 3.31E-02 | BMP4, BMP6, SMAD4 |
| Gastrulation | 7 | 3.31E-02 | BMP4, SMAD2, CD44, APOA1, FOXS1, SMAD4, UGDH |
| Ureteric bud development | 6 | 3.31E-02 | BMP4, GLI3, SMAD2, CD44, FOXS1, SMAD4 |
| Bone development | 6 | 3.31E-02 | BMP4, BMP6, GLI3, PAPSS2, GHR, SULF2 |
| Negative regulation of cell proliferation | 14 | 3.79E-02 | PPARD, GLI3, CTLA4, SMAD2, BMP4, SMAD4, SFRP4, LEPREL2, APOE, WFDC1, ITCH, OGN, COL18A1, PRNP |
| Cellular response to chemical stimulus | 28 | 4.06E-02 | NRAS, GPX3, PAPSS2, CYP2C8, NR3C1, CXCR7, FOXS1, UGDH, KDELR3, FZD2, MT1E, PTPN2, SMAD2, PLOD2, ELMO2, GHR, MYOD1, CCL21, APOE, BMP4, CD44, COL4A6, APOA1, EIF4E, PRKAR2A, SULF2, CDC37, CYP1B1 |
| Response to steroid hormone stimulus | 10 | 4.31E-02 | GPX3, SFRP4, NR3C1, WFDC1, GHR, MYOD1, BMP4, ARNT2, ARSB, LOX |
| Histone H3-K4 demethylation | 2 | 4.31E-02 | C12ORF169, KDM1B |
| UDP-glucose metabolic process | 2 | 4.31E-02 | UGGT1, UGDH |
| De novo' posttranslational protein folding | 4 | 4.31E-02 | HSPH1, UGGT1, VBP1, CCT4 |
| Regulation of transmembrane receptor protein serine/threonine kinase signaling pathway | 7 | 4.31E-02 | SKIL, BMP4, SMAD2, NR3C1, BMP6, SMAD4, UCHL5 |
| Extracellular matrix organization | 8 | 4.31E-02 | PLOD2, LEPREL2, FOXS1, COL18A1, SERPINH1, COL4A6, SULF2, LOX |
| High-density lipoprotein particle assembly | 2 | 4.31E-02 | APOE, APOA1 |
| Negative regulation of T-cell differentiation | 3 | 4.31E-02 | BMP4, GLI3, CTLA4 |
| Peripheral nervous system axon regeneration | 2 | 4.31E-02 | APOE, CHST3 |
| Cartilage development | 7 | 4.31E-02 | BMP4, GLI3, FBXW4, CD44, BMP6, GHR, SULF2 |
| Molecular function | |||
| Iron ion binding | 8 | 1.44E-02 | FADS1, CYP2C8, PLOD2, LEPREL2, P4HA2, C14ORF169, CYP1B1, CH25H |
| Glutathione peroxidase activity | 3 | 2.16E-02 | GSTZ1, GPX3, GPX8 |
| l-ascorbic acid binding | 3 | 2.70E-02 | PLOD2, LEPREL2, P4HA2 |
| Phosphatidylcholine-sterol O-acyltransferase activator activity | 2 | 2.70E-02 | APOE, APOA1 |
| Histone demethylase activity (H3-K4-specific) | 2 | 2.81E-02 | C14ORF169, KDM1B |
| Unfolded protein binding | 6 | 2.81E-02 | DNAJC8, UGGT1, SERPINH1, VBP1, CDC37, CCT4 |
| Procollagen-proline dioxygenase activity | 2 | 3.53E-02 | LEPREL2, P4HA2 |
| Copper ion binding | 4 | 4.17E-02 | MT1E, CUTC, LOX, PRNP |
| Tau protein binding | 2 | 4.17E-02 | APOE, AATF |
| TGF-β receptor cytoplasmic mediator activity | 2 | 4.32E-02 | SMAD2, SMAD3 |
| Electron carrier activity | 6 | 4.32E-02 | CYP2C8, DHRS3, P4HA2, SH3BGRL3, UGDH, CYP1B1 |
| I-SMAD binding | 2 | 4.59E-02 | SMAD2, SMAD4 |
| Cis-trans isomerase activity | 3 | 4.59E-02 | GSTZ1, PPIH, FKBP11 |
| Arylsulfatase activity | 2 | 4.75E-02 | ARSB, SULF2 |
TGF, transforming growth factor.
Table 2.
KEGG pathways overrepresented by the upregulated DR genes in neonatal left ventricle with in utero exposure to increased maternal cortisol
| Pathways | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Pathways in cancer | 10 | 2.64E-05 | RALA, PPARD, GLI3, SMAD2, NRAS, BMP4, ARNT2, FZD2, COL4A6, SMAD4 |
| Wnt signaling pathway | 6 | 6.00E-04 | PPARD, SMAD2, FZD2, SFRP4, SMAD4, DAAM1 |
| Insulin signaling pathway | 5 | 3.30E-03 | CALM2, NRAS, EIF4E, PRKAR2A, CALM1 |
| TGF-β signaling pathway | 4 | 3.70E-03 | BMP4, SMAD2, BMP6, SMAD4 |
| Melanogenesis | 4 | 5.00E-03 | CALM2, NRAS, FZD2, CALM1 |
| Metabolic pathways | 12 | 6.50E-03 | NTPCR, PI4K2B, GSTZ1, POLR2I, CYP2CB, PAPSS2, P4HA2, UGDH, STT3B, ARSB, DHRS3, PANK3 |
| Basal cell carcinoma | 3 | 6.50E-03 | BMP4, GLI3, FZD2 |
| Hedgehog signaling pathway | 3 | 6.50E-03 | BMP4, GLI3, BMP6 |
| Cell cycle | 4 | 6.50E-03 | SMAD2, CDK7, CDC26, SMAD4 |
Pathways identified as over-represented by DR genes in CORT but not in the comparison of 142 days to newborn control hearts are shown in bold. CORT, cortisol; DR, differentially regulated; TGF, transforming growth factor.
Analysis of the genes downregulated in CORT newborn LV only identified COOH-terminal protein amino acid modification as a significantly downregulated biological processes by CORT (Table 3). ATPase activity coupled to transmembrane movement of substances, cysteine-type peptidase activity, and glycosylphosphatidylinositol (GPI)-anchor transamidase activity were the molecular functions associated with the genes downregulated by CORT. Among the ATPase activity-dependent proteins whose genes were downregulated include the sarcoplasmic reticulum Ca-ATPase (SERCA; ATP2A2), two ATPases associated with the lysosome (ATP6V06 and ATP6V1H), and three mitochondrial ATP binding cassette (ABC) transport family members. Pathways identified in KEGG analysis associated with the downregulated genes included metabolic pathways, GPI anchor synthesis, the cell cycle, lysosome, hypertrophic cardiomyopathy, and phosphatidylinositol signaling (Table 4); among the genes implicated in the hypertrophic cardiomyopathy pathway are PRKAB1 and PRKAG2, which encode for two of the subunits of AMP activated protein kinase involved in energy sensing in the cardiac cell. Ubiquitin-mediated proteolysis, Wnt signaling, spliceosome, phagosome, and leukocyte transendothelial migration all also contained at least four genes that were downregulated. The cellular components that reflect the downregulated genes include nucleus, as well as the mitochondrial outer membrane, vacuolar membrane, the preautophagosomal structure, vacuolar membrane, microtubule basal body, the GPI anchor transamidase complex, and the adherens junction. Among the adherens junction-related genes were those encoding palladin (PALLD), catenin delta 1 (CTNND1), and tyrosine kinase nonreceptor 2 (TNK2). Outer mitochondrial membrane associated genes include caspase 8 (CASP8) and carnitine palmitoyl transferase (CPT1B). Although cell cycle also appears as an overrepresented pathway in both the up- and downregulated DR genes, the pattern suggests that there may be interruption of cell cycle progression in the LV.
Table 3.
Biological processes and molecular functions over-represented by the downregulated differentially regulated genes in neonatal left ventricle with in utero exposure to increased maternal cortisol
| Pathways, Processes, Functions | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Biological process | |||
| COOH-terminal protein amino acid modification | 5 | 2.92E-02 | PIGB, PIGU, PIGK, PIGV, ICMT |
| Molecular function | |||
| Cysteine-type peptidase activity | 8 | 3.96E-02 | PIGK, SENP3, CASP8, USP11, USP30, ESPL1, SENP2, USP36 |
| ATPase activity, coupled to transmembrane movement of substances | 6 | 3.96E-02 | ABCB6, ATP6V0C, ABCB10, ATP2A2, ATP6V1H, ABCC5 |
| GPI-anchor transamidase activity | 2 | 3.96E-02 | PIGU, PIGK |
GPI, glycosylphosphatidylinositol.
Table 4.
KEGG Pathways over-represented by the downregulated differentially regulated genes in neonatal left ventricle with in utero exposure to increased maternal cortisol
| Pathways | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Glycosylphosphatidylinositol(GPI)-anchor biosynthesis | 4 | 2.00E-04 | PIGB, PIGU, PIGK, PIGV |
| Metabolic pathways | 19 | 2.00E-04 | GAA, CDS2, DGKZ, PIGK, MAN1C1, ATPV0A1, PIGU, PHGDH, INPP5A, GALNS, INPP4B, DHFR, MBOAT1, ATP6VOC, PIGB, ATP6V1H, ACADB, ST6GAL2, PIGV |
| Cell cycle | 6 | 6.00E-04 | HDAC1, CUL1, CHEK2, RBX1, ESPL1, TFDP1 |
| Circadian rhythm–mammal | 3 | 1.80E-03 | CUL1, FBXW11, RBX1 |
| NOD-like receptor signaling pathway | 4 | 1.80E-03 | SUGT1, ERBB2IP, CASP8, NLRP1 |
| Collecting duct acid secretion | 3 | 2.30E-03 | SLC12A7, ATPV0A1, ATP6V0C |
| Lysosome | 5 | 2.60E-03 | ATP6V0A1, ATP6V0C, GAA, ATP6V1H, GALNS |
| Phosphatidylinositol signaling system | 4 | 3.40E-03 | CDS2, DGKZ, INPP5A, INPP4B |
| Glycerophospholipid metabolism | 4 | 3.60E-03 | MBOAT1, LPCAT3, CDS2, DGKZ |
| Hypertrophic cardiomyopathy | 4 | 3.70E-03 | PRKAB1, PRKAG2, ATP2A2, ITGA4 |
Pathways identified as over-represented by differentially regulated genes in cortisol but not in the comparison of 142 days to newborn control hearts are shown in bold; pathways in italics are those that are normally upregulated from 142 days to birth.
Analysis of the up- and downregulated DR genes using ClueGO revealed similar results, albeit with more descriptive terms (Fig. 1A). Three hundred and sixty-eight genes were recognized as having functional annotations by the ClueGO software. Two hundred and six significant nonredundant gene ontology terms and pathways were assigned to 66 of these genes. This analysis indicated several clusters of DR pathways and identified hub genes associated with these clusters (Fig. 1A). CALM1 and CALM2 (49 terms), SMAD2 and SMAD4 (42 terms), CUL1 and RBX1 (27 terms), APOA1 and APOE (13 terms), and PRKAB1 and PRKAG2 (8 terms) were identified as genes with the most associated terms. CUL1, RBX1, PRKAB1, and PRKAG2 were downregulated, whereas CALM1, CALM2, SMAD2, SMAD4, APOA1, and APOE were upregulated by CORT overexposure in utero. The analysis also identified the downregulated hub genes SLC25A10 and SLC25A11; these are involved in transport of malate and oxaloacetate into mitochondria.
Fig. 1.
Enriched ClueGO gene networks from the left ventricle (A) and the septum (B) in newborn lambs. Gene ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways associated with the differentially expressed genes involved in the network (genes named in red). Green lines with arrows represent gene activation interactions; red lines with bars represent interactions involved in gene repression.
The results of TF target analysis from WebGestalt revealed significant association of genes upregulated in CORT newborns in the LV with 160 TF binding sites, and significant association of genes downregulated in CORT newborn LV with 128 TF binding sites. The glucocorticoid receptor (GR) was associated with only seven genes, all downregulated genes; other steroid receptor TF associated with downregulated genes were estrogen receptor (ER; 4 genes), thyroid hormone receptor (T3R; 4 genes), progesterone receptor (PR; 3 genes), and vitamin D receptor (VDR; 4 genes). PR (6 genes), VDR (5 genes), and estrogen-related receptor (ERR1, 12 genes) were also associated with upregulated genes. The 10 TFs with the highest significance associated with the upregulated genes were lymphoid enhancer binding protein 1 (LEF1), nuclear factor of activated T cells (NFAT1), specificity protein 1 (SP1), MYC-associated zinc finger protein (MAZ), E2A immunoglobulin enhancer binding factors (E12), hepatocyte nuclear factor 1 homeobox (HNF1), and several unknown TFs. Other TFs associated with downregulated genes in the LV included nuclear respiratory factor 1 (NRF1), ETS Transcription Factor (ELK1), forkhead box 04 (FOXO4), Ying yang 1 (YY1), and GA binding protein (GABP), as well as SP1, E12, and MAZ.
The modeling of the patterns of change in gene expression in CORT suggested changes to pathways involved in metabolism and collagen deposition. The upregulation in the LV of several genes (SMAD4, BMP4, BMP6) in the TGF-β signaling pathway, a major regulator of collagen deposition (24) prompted us to investigate changes in collagen deposition within the newborn lamb LV using histological methods. CORT significantly increased collagen deposition in the LV of the newborn as identified by picrosirius red staining (1.5 ± 0.3% stained area in LV of control newborns versus 3.6 ± 0.9% stained area in CORT newborns) (Fig. 2). We found no overall proliferative effect of CORT in these hearts, in either LV or IVS, despite the observation that the LV free wall and the septal walls were thicker relative to tibial length (1). As expected for the ovine heart at birth, there were very low levels of Ki-67 staining, indicating low levels of proliferative activity, and no increase in Ki-67 with excess cortisol exposure in the ewe (% nuclei Ki-67-stained LV Control: 0.21 ± 0.11%, LV CORT: 0.11 ± 0.05%; IVS Control: 0.23 ± 0.11%, IVS CORT: 0.11 ± 0.03%). These results suggest the increase in wall thickness reflects hypertrophy rather than hyperplasia and is also consistent with interruption of cell cycle progression.
Fig. 2.

Representative images of left ventricle stained with picrosirius red (depicted as bright areas) and imaged using circular polarized light from control (A) and cortisol (CORT) (B) newborn lambs. Quantification of stained area relative to tissue area (C) is shown. *Statistical difference (P ≤ 0.05) between control and CORT fetuses.
Patterns of change in gene expression in the newborn lamb IVS after excess CORT exposure in utero.
Gene ontology analysis of the genes upregulated in the IVS identified cellular macromolecular complex subunit organization (8 genes, adj. P = 4.56E-02), protein localization (9 genes, adj. P = 1.84E-02), and cytoskeleton-protein binding (8 genes, adj. P = 2.55E-02) processes as biological processes altered by maternal cortisol exposure. The analysis more specifically identified cellular protein complex assembly and actin filament assembly, as well as skeletal muscle proteins as biological processes altered in CORT newborns (Table 5). ER retention sequence binding and actin binding were the molecular functions altered. The cytoplasm and cytoskeleton were cell components related to the DR genes; more specifically the cell leading edge, the apical junction complex, and cytoskeleton were identified. Genes associated with cytoplasm and actin binding included TNNI2 of the troponin complex; TNNI2 was the most highly upregulated gene (Supplemental Table S2). The pathways identified by KEGG analysis that were associated with the upregulated genes in IVS were the insulin signaling pathway, which included hexokinase (HK1) and UDP-glucose pyrophosphatase 2 (UGP2) genes relating to galactose and sucrose metabolism pathways, neurotrophin signaling, and genes in pathway for regulation of the actin cytoskeleton (RHOA, DIAPH1, SCIN) (Table 6). Other pathways identified with significant representation by upregulated genes were the TGF-β signaling pathway (TGFBR1 and ras family member RHOA), tight junctions (RHOA and claudin 1, CLDN1), focal adhesion (RHOA, DIAPH1), and Vibrio cholera infection pathway, based on two genes important in ER retention (Lys-Asp-Glu-Leu ER retention protein receptors, KDEL2 and KDEL3).
Table 5.
Biological processes and molecular functions over-represented by the upregulated differentially regulated genes in neonatal septum with in utero exposure to increased maternal cortisol
| Processes and Functions | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Biological Process | |||
| Negative regulation of protein complex disassembly | 3 | 2.70–02 | MID1IP1, CAPZA2, SCIN |
| Protein retention in ER lumen | 2 | 2.70E-02 | KDELR2, KDELR3 |
| Acute-phase response | 3 | 2.70E-02 | ITIH4, REG3G, TSC2 |
| Negative regulation of hydrolase activity | 5 | 3.27E-02 | ITIH4, ZCCHC9, SERPINH1, APOA1, UCHL5 |
| Heparin sulfate proteoglycan metabolic processes | 2 | 4.86E-02 | DSE, GPC |
| Skeletal muscle contraction | 2 | 4.86E-02 | TNNI2, TNNT1 |
| Regulation of actin cytoskeleton organization | 4 | 4.86E-02 | CAPZA2, RHOA, SCIN, ARPC5L |
| Molecular Function | |||
| ER retention sequence binding | 2 | 5.10E-03 | KDELR3, KDELR2 |
| Actin binding | 6 | 2.55E-02 | TNNI2, CAPZA2, SCIN, ARPC5L, DIAPH1, UXT |
Table 6.
Pathways over-represented by the upregulated differentially regulated genes in neonatal septum with in utero exposure to increased maternal cortisol
| Pathways | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Insulin signaling pathway | 4 | 4.00E-04 | HK1, TSC2, CALM2, CALM1 |
| Pathogenic Escherichia coli infection | 4 | 3.50E-05 | CLDN1, YWHAZ, RHOA, ARPC5L |
| Neurotrophin signaling pathway | 4 | 4.00E-04 | YWHAZ, RHOA, CALM2, CALM1 |
| Regulation of actin cytoskeleton | 4 | 1.70E-03 | SCIN, RHOA, ARPC5L, DIAPH1 |
| Phototransduction | 2 | 3.00E-03 | CALM2, CALM1 |
| Vascular smooth muscle contraction | 3 | 2.80E-03 | CALM2, RHOA, CALM1 |
| Oocyte meiosis | 3 | 2.80E-03 | CALM2, YWHAZ, CALM1 |
| Galactose metabolism | 2 | 2.90–03 | HK1, UGP2 |
| Amino sugar and nucleotide sugar metabolism Starch and sucrose metabolism | 2 | 7.30E-03 | HK1, UGP2 |
| Vibrio cholera infection | 2 | 7.40E-03 | KDELR3, KDELR2 |
There were no significant biological processes associated with the genes downregulated in CORT newborn IVS. The only molecular function identified was protein binding (Table 7). Nucleus cytoplasm and organelles were identified as major cellular components involved in the effects of cortisol, but interestingly, preautophagosomal structure, the sarcomere, Golgi membrane, histone acetyltransferase complex, focal adhesion, and Z-disk were identified. Among the genes in sarcomere were actinin α2 (ACTN2), paladin (PALLD), and matrix metallo protease 2 (MMP2); ACTN2 and PALLD were also among the genes identified in focal adhesion, adherens junction, and the Z-disk. Chromatin and the spindle pole were also identified as cellular components related to downregulated genes; the chromatin-related genes included three also related to histone acyltransferase: CREBBP, BRD1, and ING4. Pathways identified from downregulated genes in CORT newborn IVS were lysosome, insulin, hedgehog, and adipokine signaling pathways (Table 8). Adherens junction and cardiac muscle contraction were also identified, consistent with the gene ontology, although only two genes were identified in each pathway (COX4I1 and ATP1B1 for cardiac muscle contraction and CREBBP and ACTN2 for adherens junction).
Table 7.
Biological processes and molecular functions over-represented by the downregulated differentially regulated genes in neonatal septum with in utero exposure to increased maternal cortisol
| Analysis | Processes and Functions | No. of Genes Involved | Adjusted P Value | Genes |
|---|---|---|---|---|
| Molecular function | ||||
| Protein binding | 67 | 1.57E-02 | SPTBN2, FBXW11, WBP11, PRKAB1, PRKACB, MAPK14, MRFAP1, ACACB, AKAP9, C12orf44, CXXC1, MED6, RPS19BP1, F3, MBD4, CREBBP, PDE3B, EPB41L3, EIF2AK2, EIF4A3, UBA2, DNAP2, CASP2, ISCU, COG3, CPSF3, TNK2, RUNX1T1, ING4, ATP1B1, BRD1, FANCG, SPPL3, NUMA1, ANKZF1, ARHGAP21, NBR1, MAU2, NCOA1, USP11, LPHN2, RPS26, TH1L, FBLIM1, PKP4, ARHGEF7, TRIO, ACTN2, ARF1, FHOD3, SMC3, CTSC, MMP2, ELF2, SRSF11, IGF2R, TELO2, ZC3H11A, FOXN3, DEK, ARID4B, PALLD, RBM5, KIAA0141, ZFYVE26, YY1, RABEP1 | |
Table 8.
Pathways overrepresented by the downregulated differentially regulated genes in neonatal septum with in utero exposure to increased maternal cortisol
| Pathways | No. of Genes Involved | Adjusted P Values | Genes |
|---|---|---|---|
| Insulin signaling pathway | 4 | 1.44E-02 | PRKAB1, ACACB, PDE3B, PRKACB |
| Adipocytokine signaling pathway | 3 | 1.44E-02 | CPT1B, PRKAB1, ACACB |
| Progesterone-mediated oocyte maturation | 3 | 1.92E-02 | PDE3B, MAPK14, PRKACB |
| Oocyte meosis | 3 | 2.15E-02 | FBXW11, SMC3, PRKACB |
| Leukocyte transendothelial migration | 3 | 2.15E-02 | ACTN2, MAPK14, MMP2 |
| GnRH signaling pathway | 3 | 2.15E-02 | MAPK14, PRKACB, MMP2 |
| Lysosome | 3 | 2.15E-02 | IGF2R, GAA, CTSC |
| Wnt signaling | 3 | 2.99E-02 | CREBBP, FBXW11, PRKACB |
| N-Glycan biosynthesis | 2 | 2.99E-02 | MAN1C1, ALG1 |
| Hedgehog signaling pathway | 2 | 3.05E-02 | FBXW11, PRKACB |
| Vibrio cholera infection | 2 | 3.44E-02 | ARF1, PRKACB |
ClueGO software identified 183 genes as having functional annotations in the septum (Fig. 1B). Thirty genes were assigned to 62 significant nonredundant gene ontology terms and pathways. NUMA1 (nuclear apparatus protein 1, 11 terms) NCOA1 (nuclear receptor coactivator, 4 terms), and RHOA (Ras homolog protein A, 4 terms) were associated with the most terms in the septum. NUMA1 and NCOA1 were both downregulated whereas RHOA was upregulated in the septum by cortisol exposure. NUMA1 is associated with mitotic spindle formation and RHOA with actin cytoskeleton.
The results of TF target analysis from WebGestalt revealed significant association of genes upregulated in CORT newborn IVS with 41 TF binding sites and significant association of genes downregulated in CORT newborn IVS with 107 TF binding sites. The top 10 TFs associated with at least 5 upregulated genes were SP1, activator protein 4 (AP4), general transcription factor (GTF2A1), MAZ, ELK1, E4F1, LEF1, paired box 4 (PAX4), NF1, and myogenic differentiation 1 (MYOD1). The top 10 TFs associated with downregulated genes in IVS included GR, FOXO4, SP1, LEF1, E12, NFMUE1, AP1, CREB, E2F, and ELK1. In addition to GR (with a total of 18 genes associated with 3 different sites), PR (4 genes) and ER (5 genes) were also implicated in downregulated genes, and ERR1 was also implicated (4 genes).
In the cardiac muscle contraction pathway, COX4I1, the cytochrome c oxidase subunit IV subunit 1, and ATPB1, the Na-K transporting ATPase β1 polypeptide, were significantly suppressed in the IVS after CORT. In the IVS CORT also significantly reduced COX4-stained area relative to total tissue area (23.3 ± 7.0% stained area in control newborns IVS vs. 5.3 ± 0.8% stained area in CORT newborn IVS) (Fig. 3). Similarly, COX4 protein concentration was significantly reduced by CORT (2.38 ± 0.62 arbitrary fluorescent units in Control newborns vs. 0.99 ± 0.29 arbitrary fluorescent units in CORT newborns) (Table 9). Expression of the other mitochondrial proteins were not different between groups in either IVS or LV (Table 9) nor was COX 4 protein or density altered in the LV.
Fig. 3.

Representative images of septum from control (A) and cortisol (CORT) (B) newborn lambs stained with anti-COX4 antibody (red). Nuclei are stained in blue (DAPI) and myocytes in green (wheat germ agglutinin). Quantification of stained area relative to tissue area (C) is shown. *Statistical difference (P ≤ 0.05) between control and CORT fetuses. COX, cytochrome c oxidase.
Table 9.
Abundance of proteins in left ventricle or septum from fetuses of cortisol infused or control ewes
| Left Ventricle |
Septum |
|||
|---|---|---|---|---|
| Protein | Cortisol | Control | Cortisol | Control |
| NADH:ubiquinone oxidoreductase subunit B8 | 12.4 ± 1.74 | 11.8 ± 2.65 | 28.5 ± 8.77 | 35.5 ± 9.81 |
| Succinate dehydrogenase | 36.9 ± 3.24 | 39.3 ± 2.23 | 33.6 ± 10.3 | 38.2 ± 11.9 |
| Ubiquinol-cytochrome c reductase core protein II | 42.9 ± 2.72 | 43.2 ± 3.28 | 87.3 ± 33.2 | 98.5 ± 36.3 |
| Cytochrome c oxidase subunit I | 67.8 ± 8.39 | 71.5 ± 10.5 | 85.6 ± 30.4 | 90.4 ± 31.2 |
| Cytochrome c oxidase subunit IV | 88.0 ± 11.8 | 104 ± 6.77 | 0.99 ± 0.29* | 2.38 ± 0.62 |
| ATP synthase subunit α | 75.3 ± 5.81 | 85.6 ± 4.82 | 188 ± 69.6 | 193 ± 84.9 |
| Cytochrome c | 24.6 ± 5.65 | 29.3 ± 8.57 | 24.6 ± 5.65 | 29.3 ± 8.53 |
| Pyruvate dehydrogenase E1a | 72.6 ± 5.06 | 74.0 ± 5.39 | 24.7 ± 7.88 | 51.1 ± 9.88 |
| Phosphorylated pyruvate dehydrogenase E1a | 67.7 ± 12.2 | 74.5 ± 12.2 | 26.9 ± 12.2 | 48.1 ± 13.6 |
Data are expressed as means ± SE of relative density units corrected for total protein loaded.
Statistically significant differences between groups, P < 0.05.
Comparison of effects of CORT to normal maturational changes in gene expression.
The normal CORT surge at the end of gestation is responsible for changes in expression of many critical genes. We compared the DR genes in CORT newborn LV with the DR genes from the comparison of Control newborns to Control fetuses at ~142 days of gestation (Fig. 4). In this reanalysis of the effects of CORT on cardiac gene expression, only 20% of the genes in the LV were regulated in the same direction by the exposure to excess CORT as by normal maturational changes from 142 days to birth. However, 122 (73%) of the genes upregulated in CORT newborn LV were genes that were not changed during normal late gestation maturation; similarly, 162 (74%) of the genes downregulated in CORT newborns were not significantly changed during normal late gestation maturation. There were 32 genes that were upregulated in CORT newborns that are normally upregulated between 142 days and birth and 45 genes downregulated in CORT newborns that are normally downregulated from 142 days to birth. The common upregulated genes were identified as associated with UDP glucose metabolism and protein folding, and among the KEGG pathways were insulin signaling (CLAM1, CLAM2, NRAS, EIF4E) and protein processing in endoplasmic reticulum (HSPH1, UGGT1, SEC24D). The common downregulated genes were in WW domain binding, GTPase regulator activity, and the adherens junction (EZR, TNK2, TMEM204, PALLD, ARHGEF7), and the only KEGG pathways were N-glycan biosynthesis (MAN1C1, ST6GAL2) and regulation of the actin cytoskeleton (EZR, ARHGEF7). There were 13 genes downregulated in CORT newborns that are normally upregulated between 142 days and birth. There were 12 genes were upregulated in CORT newborns that are normally downregulated from 142 days to birth. There were no KEGG pathways or biological processes modeled by genes upregulated in CORT newborns and normally downregulated in late gestation. In contrast, the 13 genes downregulated in CORT newborns and normally upregulated from 142 days to birth were modeled by GPI anchor biosynthesis (PIGB, PIGU, PIGK) and include phosphoglycerate dehydrogenase (PHGDH); the three phosphatidylinositol glycan anchor biosynthesis genes were also represented in most of the identified common biological processes, including COOH-terminal protein lipidation, glycolipid biosynthesis, and GPI anchor biosynthesis; carboxylic acid metabolic process (FABP3, LPCAT3, MARS, PHGDH), cellular response to oxidative stress (TMEM161A, PNPT1), and translation were also identified in the biological processes with significant representation by these 13 genes.
Fig. 4.
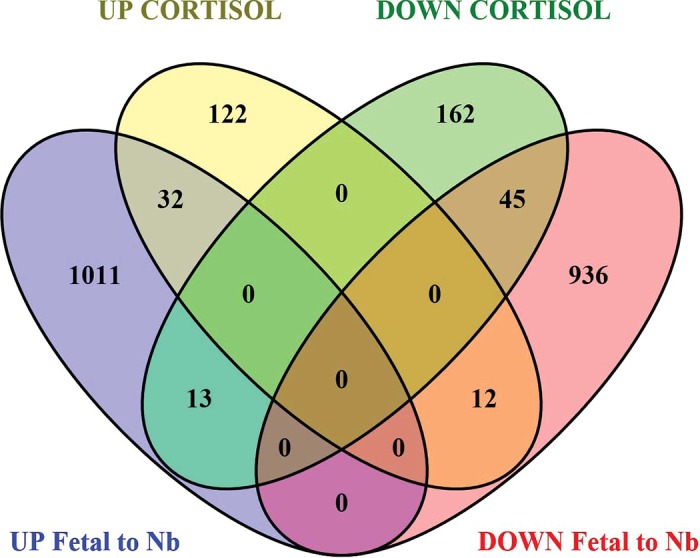
Venn diagram showing overlap of differentially regulated (DR) genes in left ventricle (LV) up (yellow) and down (green) regulated in cortisol (CORT) newborns and DR genes up (purple) and down (pink) regulated in the control newborns compared with control 142-day fetuses, as a reflection of normal maturation. There were 32 genes upregulated in both sets of LV data, and 45 genes downregulated in both sets of LV data. There were 13 genes downregulated in CORT newborn LV but downregulated from 142 days to newborn in the LV, and 12 genes upregulated in CORT newborn LV but upregulated in newborn LV compared with the 142-day fetus LV. There were 122 upregulated and 162 downregulated genes in the CORT newborns LV that were not significantly changed in LV with the transition between 142 days of fetal life and the first day of life.
Of the 122 genes upregulated in CORT but not significantly changed in the control fetuses/lambs from 142 days to birth, the major KEGG pathways were cancer, cell cycle, metabolic pathways, hedgehog signaling, and Wnt signaling (Table 2). Other pathways identified in this analysis of these 122 DR genes were excision repair (CDK7, RAD23B), PPAR signaling (PPARD, APOA1), P450 metabolism (CYP2C8, CYP1B1, GSTZ1), protein digestion and absorption (COL18A1, Col4A6), and extracellular matrix receptor interaction (CD44, COL4A6). Of the 162 genes downregulated in CORT but not normally changed from 142 day to birth, the over-represented pathways included cell cycle, lysosomal pathway, NOD-like signaling, circadian rhythm, phosphatidylinositol signaling, glycerolphopholipid metabolism, collecting duct secretion, and cardiac hypertrophy (Table 4). Also identified were metabolic pathways; the genes for PIGB, PIGK, PIGU, and PIGV were missing from those in the metabolic pathway in this comparison as those were normally upregulated during the peripartum period (Table 4).
DISCUSSION
Chronically elevated maternal cortisol concentrations in late gestation resulted in significant changes in gene expression within pathways involved in maintaining normal cardiac architecture and metabolism in the newborn lamb heart. The modeling indicates that the specific molecular mechanisms underlying the increase in peripartum morbidity and mortality differ in the LV and IVS, although in both regions the changes in gene expression suggest that pathways for cardiac remodeling are altered. The modeling also indicates that the transcriptomic effects of elevated maternal cortisol differ from the pattern of change in gene expression occurring with the normal surge in fetal cortisol production in the peripartal period. This supports the hypothesis that the effects of chronic exposure to low levels of CORT alter the normal developmental trajectory, producing changes in cardiac growth and metabolism that are not simply exaggerations of the effects of the much greater increase in fetal cortisol normally occurring in the days preceding delivery.
Glucocorticoids are essential for the maturational development of fetal tissues in preparation for ex utero survival, and in the fetal sheep the prepartum surge in fetal production of cortisol is critical for lung development as well as maturation of other organs, including the liver and gastrointestinal tract (19, 36). Glucocorticoids are also critical for normal development and maturation of cardiomyocytes (63, 64). Cortisol is needed for the normal preparturient increase in fetal arterial blood pressure (77), but higher levels of cortisol result in elevated fetal arterial pressure (17, 41, 49). Increased exposure to maternal glucocorticoids in utero increases blood pressure in childhood and adulthood (6, 13, 26, 37), suggesting possible programming of cardiac development. Indeed, infusion of cortisol to the fetus at a rate that raises CORT to much higher levels than achieved in our study causes cardiac hypertrophy in association with the relative fetal hypertension and altered cardiac angiotensin gene expression (41). However, other studies have shown that CORT can have proliferative effects when infused directly into the heart (21) or directly applied to cultured fetal cardiomyocytes (22).
This current study extends our understanding of the effects of glucocorticoids on the fetal heart to a model in which a more modest chronic increase in maternal cortisol does not alter fetal blood pressure but in which we have found a slowing of heart rate and an increase in arrhythmias at the time of birth. Taken together, our results suggest that the chronic elevation of maternal cortisol over a period of cardiac maturation in late gestation causes progressive shifts in gene expression that in turn shift the normal ongoing maturational processes in the heart, including both structural and metabolic changes, and these are functionally evident during the challenge of labor and delivery. Our study period, with exposure starting at 80% of gestation and continuing to birth, encompasses a period in which the heart undergoes terminal differentiation, shifting from a highly proliferative tissue to one with little or no capacity for proliferation, and with the majority of the cardiomyocytes binucleated (30). Over this period, mitochondria aggregate between myofibrils (8), and the heart matures metabolically, so that in postnatal life fatty acids rather than glucose and lactate are the primary metabolic substrates (3, 4). The Purkinje fibers, which are present in greater proportions in the IVS, also mature and are thought to continue to proliferate or be recruited during the early postnatal period (45).
The results of this study, when considered in the context of our prior studies in this model, indicate that the increased maternal cortisol produces different patterns of gene expression and different effects on cardiac wall architecture at different time points in the exposure. Although the initial 10–15 days of exposure was found to increase heart weight and wall thickness (27, 28, 59) (Fig. 5), longer exposure does not result in a heavier heart, although specifically the left heart walls are relatively thicker and contain more collagen. The exposure to higher than normal levels of cortisol in the fetus initially induce greater myocyte proliferation, leading to a modest cardiac enlargement from 115 to 130 days relative to controls of the same age (16); in the IVS at 130 days apoptotic markers were increased in Purkinje fibers, suggesting these fibers were remodeled by CORT. The changes were modeled in the IVS by changes in genes associated with proliferation as well as with apoptosis. Continued CORT exposure to 140 days of gestation failed to result in larger hearts, nor were genes in proliferation-related pathways increased in expression (Fig. 5). However, the relative number of mitochondria was reduced, as reflected by mitochondrial DNA, as was gene expression in pathways relating to cardiac metabolism (60). Whereas after 15 days of exposure, there were relatively few “metabolic” genes altered in the heart (60) (Fig. 5) these were overrepresented in the models of differential gene expression at 140 days (60) and at birth. Our present results indicate that the pattern of change in gene expression is further modified at birth, with changes in genes that result in structural remodeling of the LV and IVS. These later changes may be an adaptive response to the further exposure to glucocorticoids or to altered supply of metabolites.
Fig. 5.
Summary of the effects of increased maternal cortisol, produced by infusion of 1 mg·kg−1·day−1 from 115 days to birth, on fetal cardiac phenotype, and the changes in pathways modeled using transcriptomics. Data from hearts at 130 days and 140 days have been previously reported (59, 60). LV, left ventricle; IVS, septum; RV, right ventricle; TGF, transforming growth factor.
It is important to note that although maternal CORT stays elevated throughout the period of CORT infusion, fetal CORT concentrations are only elevated in the first 10–15 days of infusion (1) before the maturation of the fetal adrenal. The elevated maternal CORT does not appear to alter adrenal maturation, as the fetal CORT concentrations in late gestation are not significantly different than those in Control fetuses (1). This suggests either that the increased fetal CORT during 115–130 days reprograms some aspects of cardiac maturation or that effects of the persistently higher maternal CORT have effects through changes in maternal or placental metabolism that have secondary effects on the fetal heart. Indeed, the proliferation we observed at 130 days was cortisol-dependent, as a mineralocorticoid receptor blocker was able to block this effect (16, 59). However, effects of maternal CORT on placental metabolism may contribute to longer-term effects, as changes in the newborn cardiac metabolome appear to more closely parallel changes in the fetal serum metabolome than in the maternal metabolome (81). Valine, glutamine, succinate, and both diglycerides and triglycerides are elevated in CORT cardiac tissue, and dimethylamine, glycerophosphocholine, betaine, and glutamate are diminished in CORT; these appear to parallel changes in fetal (but not maternal) serum metabolites, as branched chain amino acids, choline, and glycine were significantly elevated in serum from CORT fetuses, as is the ketone 3-hydroxybutyrate.
In the term lambs before birth, the gene that was most upregulated by CORT in the fetal septum was PDK4, a glucocorticoid target and key negative regulator of pyruvate metabolism (11), suggesting that oxidative metabolism for glucose and lactate is reduced. Others have shown that chronic glucocorticoid exposure in adults significantly reduces the activities of mitochondrial electron transport complexes I and III, increases reactive oxygen species production, decreases antioxidant levels and activity, and increases susceptibility to proapoptotic signals (14, 42, 46, 72). However, in the term fetuses there were no changes in the expression of the nuclear-encoded genes of the electron transport chain in LV; in IVS only MT-ND1, UQCC3 (of Complex III), and ATPIF1 (of Complex V) were significantly downregulated. We had predicted that at birth there may be more dramatic effects on mitochondrial function, reflected in altered expression of mitochondrial electron transport chain genes; however, this was not the case. Although many genes involved in carbohydrate or lipid metabolism were altered, PDK4 expression in the IVS and LV at birth was not changed by prior exposure to maternal hypercortisolemia. Analysis of pyruvate dehydrogenase E1a (the target for PDK4) protein and its phosphorylated form also indicate that this pathway is not affected in these newborn lambs. This may reflect the increased PDK4 activity at birth in both control and CORT newborns secondary to the normal surge in fetal plasma cortisol. At birth, the only protein of the electron transport chain that was significantly changed by prior CORT was COX4 in the IVS and cytochrome c1 (CYC1, of Complex III) in LV. Assessment of mitochondrial density using COX4 indicated less area with COX4-positive staining in the IVS; COX4 protein expression in the IVS was also reduced. Cytochrome c oxidase is the terminal complex in the electron transport chain and is composed of nuclear and mitochondrial-derived proteins. COX1 is a mitochondrial DNA-derived protein and the catalytic core of the complex; COX4 is a critical assembly protein encoded by nuclear DNA (2, 9, 39). Decreased levels of COX4 in vitro have been shown to compromise severely the assembly of COX, leading to impairment of oxidative phosphorylation, which reduces ATP production and sensitizes the cell to apoptosis (39, 61). These findings are particularly interesting given the results from our studies showing changes in the fetal ECG at birth (1) and implicates mitochondrial dysfunction in the development of this cardiac phenotype.
In both LV and IVS, the transcriptomic analysis indicates that in addition to disruption of normal metabolic transitions at birth, there are changes in the architecture of the ventricular walls as a consequence of the chronic overexposure to maternal CORT. Both cell cycle/proliferative pathways and apoptotic pathways were overrepresented in the DR genes, leading to an overall model of alterations in terminal differentiation with altered cell cycle progression, reduced AMP activated protein kinase signaling, apoptosis, lysosomal and phagosomal activity, and response to DNA damage. In LV we found that SMAD protein signaling was significantly upregulated in CORT newborns, although not normally upregulated with the normal perinatal surge in cortisol. BMP4 and SMAD4 are essential for cardiac development, but enhanced expression has been implicated as a major factor in pathological remodeling of the adult myocardium (29, 48, 57). BMPs are part of the TGF-β superfamily of proteins and signal through SMAD-dependent and -independent pathways. SMAD-dependent pathways converge at the SMAD4 protein as a common mediator of BMP signaling, which regulates target gene transcription (25). BMP signaling is highly conserved and GR knockdown reduces the expression of several BMPs in zebrafish embryos, including BMP4 (50, 51). In contrast, maternal treatment with the synthetic glucocorticoid dexamethasone upregulates the expression of BMP4 in the fetal mouse kidney (12). BMP4 is also upregulated in pathological cardiac hypertrophy, acute myocardial infarction, chronic ischemic heart disease, and cardiomyocyte apoptosis (53, 71, 86). Increased SMAD4 protein expression and collagen deposition occurs in the border and scar tissue following myocardial infarction (24). Our histological results support the transcriptomic modeling, as collagen was increased in the LV. The amount of collagen deposition is less than what is observed in hearts following myocardial infarction (10), as might be expected because there is no functional or histological evidence of LV myocardial infarct or necrosis. These findings suggest that the prior exposure to excessive corticosteroids potentiates BMP-SMAD signaling in the fetal heart, leading to an increase in cardiac collagen deposition; this effect could contribute to pathophysiologies later in life. Because GRs inhibit SMAD signaling (70), this increase also suggests that this is not directly GR-mediated, and rather an adaptive response secondary to prior CORT exposure. Consistent with cardiac hypertrophy is the increased expression with the fetal exposure to CORT of genes encoding for components of the troponin complex. There was increased expression of the genes TNNT1, encoding the tropomyosin-binding subunit of the troponin complex, and TNNI2, encoding the inhibitory subunit. In the adult these genes encode skeletal muscle isoforms of these proteins. However, in the fetus and newborn TnT1 and TnT4, along with TnI2, are the major protein components of the complex binding TnC (23, 65). TNNT4 is not on the array; TNNC1 is on the array and is highly expressed in the fetal heart, but was not significantly upregulated by cortisol exposure. The upregulation of these genes may indicate a similar adaptive response in the fetal heart. Troponin I expression increases in the fetal heart at term in parallel with increased binucleation (56) but may also rise with hypertrophy of LV cardiomyocytes. Increased in expression of the slow skeletal muscle form of TnT is found in end-stage heart failure (5) and may be an adaptive mechanism to increase sensitivity to calcium in states of increased LV wall stress (55). The analysis also suggests that disturbances in gene expression in the GPI anchor pathway and in vacuolar-type H+-ATPases (V-ATPases) may contribute to the pathology. These pathways are essential for normal protein glycosylation and processing by ER and Golgi; defects in other genes in this pathway are associated with congenital diseases with cardiac manifestations that include arrhythmias and myopathies (43). GPI anchoring brings proteins to the plasma membrane and lipid rafts and so is involved in cell adhesion and some receptor function. Taken together the results in the LV suggest several normal mechanisms are disrupted, which potentially contribute to the modest increase in wall thickness and fibrosis. Although we have not found an increase in fetal blood pressure in this model (1, 59), it is possible that there is an increase in wall tension for example if chamber radius is altered. Further studies are necessary to understand more fully the underlying mechanism for the change in architecture of the LV walls in this model.
In the IVS there were also significant changes in gene expression in pathways involved in the Z-disk of the myocyte, focal adhesion, and in actin microfilaments and the cell cytoskeleton. This included upregulation of the gene CAPZA1 encoding the actin filament capping protein. Actin filament capping occurs in the Z-disk of the myocyte; in Purkinje fibers the microfilaments appear to be particularly important in maintaining normal cell structure (76). CAPZ also plays a role in β-adrenergic receptor signaling in cardiac muscle (20) and mitochondrial membrane potential (80). Capping of actin filaments limits actin assembly and therefore may also contribute to the terminal differentiation of cardiomyocytes; posttranslational modification of CAPZ has been shown to allow for actin assembly in models of hypertrophy in the adult heart (38). In the IVS the genes for calmodulins (CALM1 and CALM2) were both upregulated. In adult cardiac hypertrophy, small-conductance Ca2+-activated K+ channels are activated during hypoxia via the Ca2+/calmodulin-dependent protein kinase II (CaMKII)-dependent pathway and may thus increase risk of ventricular arrhythmias (74). Calmodulin activity modifies activity of many ion channels in the atrium and ventricle, including the ryanodine receptor and the L-type calcium channel, as well as playing a role in sensing of reactive oxygen species, and in-fibrosis (reviewed in Ref. 44).
Although studies in rodents suggest that GR-mediated actions are important for cardiac maturation (63), TF target analysis from downregulated genes in the LV and IVS implicated GRs acting at glucocorticoid response elements (GREs) in some genes; however, this only accounts for a small number of DR genes in this study. This is not surprising as the plasma CORT concentrations in the fetuses were not different from 130 days until birth, and cardiac 11 β-hydroxysteroid dehydrogenase gene expression (11BHSD2 and 11BSHD2) was not changed in the LV or IVS at birth. Glucocorticoids also normally drive thyroid hormone-mediated mechanisms important to cardiac maturation through induction of thyroid deiodinase (18, 54, 75). Our analysis of transcription factors associated with the altered pathways did not indicate that thyroid response elements have been identified in the promoters of genes either over- or under expressed after exposure to excess maternal CORT. We hypothesize that the modest increase in maternal and fetal CORT in this model does not alter the normally timing of the induction of the deiodinase by the normal preparturient rise in cortisol, consistent with the similar fetal plasma cortisol concentrations near the time of birth in the two groups of fetuses. Our data suggest that T3R or GR association with retinoid X receptor binding sites is also not critical to the majority of the effects of CORT. Interestingly, the analysis indicated that the nuclear respiratory factor 1 (NRF1) was significantly associated with genes downregulated in CORT newborn LV, and IVS NRF1 has been shown to regulate indirectly nuclear-encoded components of electron transport chain via modulation of mitochondrial gene expression (35). E12/E47 was another transcription factor significantly associated with genes both up and downregulated in CORT newborn LV. E12/E47 is a member of basic helix-loop-helix (bHLH) proteins; heterodimerization of MyoD with E12/E47 is required for initiation of terminal differentiation in myocytes (33, 40). MYOD1 is upregulated in CORT newborn LV, and troponin I2 (TNNI2), one of the genes upregulated by CORT, is an E12/E47-regulated gene.
The use of transcriptomic modeling indicates that the effects of excess maternal CORT disrupt the normal trajectory of cardiac gene expression and therefore cardiac maturation. Although the normal pattern of CORT increase near birth is not disrupted, the chronic elevation in maternal CORT alters the normal gene expression profile, rather than simply amplifying the normal pattern of gene expression. The model indicates that increased expression in the BMP-SMAD pathway results in increased relative thickness and increased collagen deposition in the LV. The transcriptomic analysis also predicts changes in mitochondrial function, in lysosomal and phagosome action, and in protein trafficking. Although the changes in cardiac gene expression result in only a modest change in cardiac phenotype before birth, this remodeling may have on consequences for postnatal cardiac health. Transcriptomic modeling of CORT effects in the IVS suggest that IVS remodeling is more closely related to altered actin filament and the cell cytoskeleton and to metabolic remodeling; this may contribute to the development of arrhythmias, including the ST segment abnormalities that we observed at birth in these newborn lambs (1). Our model indicates that the chronic elevation of maternal CORT, as occurs in chronic maternal stress, has persistent effects on fetal cardiac maturation, which impact neonatal cardiac architecture and function, despite normalization of the fetal cortisol concentrations in the period after fetal adrenal maturation.
GRANTS
This study was funded by National Institutes of Health Grants HD-087306 and HD-057871 (to M. Keller-Wood) and American Heart Association Grant 14-GRNT-20420048 (to M. Keller-Wood).
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the authors.
AUTHOR CONTRIBUTIONS
A.A., E.M.R., C.E.W., and M.K.-W. conceived and designed research; A.A., C.W.C., and M.K.-W. performed experiments; A.A., M.L., E.M.R., C.W.C., C.E.W., and M.K.-W. analyzed data; A.A., M.L., E.M.R., C.E.W., and M.K.-W. interpreted results of experiments; A.A., C.W.C., and M.K.-W. prepared figures; A.A. drafted manuscript; A.A., M.L., E.M.R., C.W.C., C.E.W., and M.K.-W. edited and revised manuscript; A.A., M.L., E.M.R., C.W.C., and C.E.W. approved final version of manuscript.
Supplemental Data
Differentially expressed genes in Left Ventricle (unadjusted p≤0.05) in LV of CORT newborn lambs as compared to Control lambs. - .docx (60 KB)
Differentially expressed genes in Septum (unadjusted p≤0.05) of CORT newborn lambs as compared to Control lambs. - .docx (35 KB)
REFERENCES
- 1.Antolic A, Wood CE, Keller-Wood M. Chronic maternal hypercortisolemia in late gestation alters fetal cardiac function at birth. Am J Physiol Regul Integr Comp Physiol 314: R342–R352, 2018. doi: 10.1152/ajpregu.00296.2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Barrientos A, Barros MH, Valnot I, Rötig A, Rustin P, Tzagoloff A. Cytochrome oxidase in health and disease. Gene 286: 53–63, 2002. doi: 10.1016/S0378-1119(01)00803-4. [DOI] [PubMed] [Google Scholar]
- 3.Bartelds B, Gratama JW, Knoester H, Takens J, Smid GB, Aarnoudse JG, Heymans HS, Kuipers JR. Perinatal changes in myocardial supply and flux of fatty acids, carbohydrates, and ketone bodies in lambs. Am J Physiol Heart Circ Physiol 274: H1962–H1969, 1998. doi: 10.1152/ajpheart.1998.274.6.H1962. [DOI] [PubMed] [Google Scholar]
- 4.Bartelds B, Knoester H, Smid GB, Takens J, Visser GH, Penninga L, van der Leij FR, Beaufort-Krol GC, Zijlstra WG, Heymans HS, Kuipers JR. Perinatal changes in myocardial metabolism in lambs. Circulation 102: 926–931, 2000. doi: 10.1161/01.CIR.102.8.926. [DOI] [PubMed] [Google Scholar]
- 5.Barton PJ, Felkin LE, Koban MU, Cullen ME, Brand NJ, Dhoot GK. The slow skeletal muscle troponin T gene is expressed in developing and diseased human heart. Mol Cell Biochem 263: 91–97, 2004. doi: 10.1023/B:MCBI.0000041851.53074.72. [DOI] [PubMed] [Google Scholar]
- 6.Benediktsson R, Lindsay RS, Noble J, Seckl JR, Edwards CR. Glucocorticoid exposure in utero: new model for adult hypertension. Lancet 341: 339–341, 1993. doi: 10.1016/0140-6736(93)90138-7. [DOI] [PubMed] [Google Scholar]
- 7.Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pagès F, Trajanoski Z, Galon J. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25: 1091–1093, 2009. doi: 10.1093/bioinformatics/btp101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Brook WH, Connell S, Cannata J, Maloney JE, Walker AM. Ultrastructure of the myocardium during development from early fetal life to adult life in sheep. J Anat 137: 729–741, 1983. [PMC free article] [PubMed] [Google Scholar]
- 9.Capaldi RA. Structure and function of cytochrome c oxidase. Annu Rev Biochem 59: 569–596, 1990. doi: 10.1146/annurev.bi.59.070190.003033. [DOI] [PubMed] [Google Scholar]
- 10.Cleutjens JP, Verluyten MJ, Smiths JF, Daemen MJ. Collagen remodeling after myocardial infarction in the rat heart. Am J Pathol 147: 325–338, 1995. [PMC free article] [PubMed] [Google Scholar]
- 11.Connaughton S, Chowdhury F, Attia RR, Song S, Zhang Y, Elam MB, Cook GA, Park EA. Regulation of pyruvate dehydrogenase kinase isoform 4 (PDK4) gene expression by glucocorticoids and insulin. Mol Cell Endocrinol 315: 159–167, 2010. doi: 10.1016/j.mce.2009.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dickinson H, Walker DW, Wintour EM, Moritz K. Maternal dexamethasone treatment at midgestation reduces nephron number and alters renal gene expression in the fetal spiny mouse. Am J Physiol Regul Integr Comp Physiol 292: R453–R461, 2007. doi: 10.1152/ajpregu.00481.2006. [DOI] [PubMed] [Google Scholar]
- 13.Dodic M, May CN, Wintour EM, Coghlan JP. An early prenatal exposure to excess glucocorticoid leads to hypertensive offspring in sheep. Clin Sci (Lond) 94: 149–155, 1998. doi: 10.1042/cs0940149. [DOI] [PubMed] [Google Scholar]
- 14.Du J, Wang Y, Hunter R, Wei Y, Blumenthal R, Falke C, Khairova R, Zhou R, Yuan P, Machado-Vieira R, McEwen BS, Manji HK. Dynamic regulation of mitochondrial function by glucocorticoids. Proc Natl Acad Sci USA 106: 3543–3548, 2009. doi: 10.1073/pnas.0812671106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fayol L, Masson P, Millet V, Simeoni U. Cushing’s syndrome in pregnancy and neonatal hypertrophic obstructive cardiomyopathy. Acta Paediatr 93: 1400–1402, 2004. doi: 10.1111/j.1651-2227.2004.tb02943.x. [DOI] [PubMed] [Google Scholar]
- 16.Feng X, Reini SA, Richards E, Wood CE, Keller-Wood M. Cortisol stimulates proliferation and apoptosis in the late gestation fetal heart: differential effects of mineralocorticoid and glucocorticoid receptors. Am J Physiol Regul Integr Comp Physiol 305: R343–R350, 2013. doi: 10.1152/ajpregu.00112.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Forhead AJ, Broughton Pipkin F, Fowden AL. Effect of cortisol on blood pressure and the renin-angiotensin system in fetal sheep during late gestation. J Physiol 526: 167–176, 2000. doi: 10.1111/j.1469-7793.2000.00167.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Forhead AJ, Fowden AL. Thyroid hormones in fetal growth and prepartum maturation. J Endocrinol 221: R87–R103, 2014. doi: 10.1530/JOE-14-0025. [DOI] [PubMed] [Google Scholar]
- 19.Fowden AL, Li J, Forhead AJ. Glucocorticoids and the preparation for life after birth: are there long-term consequences of the life insurance? Proc Nutr Soc 57: 113–122, 1998. doi: 10.1079/PNS19980017. [DOI] [PubMed] [Google Scholar]
- 20.Gaikis L, Stewart D, Johnson R, Pyle WG. Identifying a role of the actin capping protein CapZ in β-adrenergic receptor signalling. Acta Physiol (Oxf) 207: 173–182, 2013. doi: 10.1111/j.1748-1716.2012.02470.x. [DOI] [PubMed] [Google Scholar]
- 21.Giraud GD, Faber JJ, Jonker S, Davis L, Anderson DF. Intravascular infusions of plasma into fetal sheep cause arterial and venous hypertension. J Appl Physiol (1985) 99: 884–889, 2005. doi: 10.1152/japplphysiol.01429.2004. [DOI] [PubMed] [Google Scholar]
- 22.Giraud GD, Louey S, Jonker S, Schultz J, Thornburg KL. Cortisol stimulates cell cycle activity in the cardiomyocyte of the sheep fetus. Endocrinology 147: 3643–3649, 2006. doi: 10.1210/en.2006-0061. [DOI] [PubMed] [Google Scholar]
- 23.Gomes AV, Venkatraman G, Davis JP, Tikunova SB, Engel P, Solaro RJ, Potter JD. Cardiac troponin T isoforms affect the Ca2+ sensitivity of force development in the presence of slow skeletal troponin I: insights into the role of troponin T isoforms in the fetal heart. J Biol Chem 279: 49579–49587, 2004. doi: 10.1074/jbc.M407340200. [DOI] [PubMed] [Google Scholar]
- 24.Hao J, Ju H, Zhao S, Junaid A, Scammell-La Fleur T, Dixon IM. Elevation of expression of Smads 2, 3, and 4, decorin and TGF-β in the chronic phase of myocardial infarct scar healing. J Mol Cell Cardiol 31: 667–678, 1999. doi: 10.1006/jmcc.1998.0902. [DOI] [PubMed] [Google Scholar]
- 25.Heldin CH, Miyazono K, ten Dijke P. TGF-β signalling from cell membrane to nucleus through SMAD proteins. Nature 390: 465–471, 1997. doi: 10.1038/37284. [DOI] [PubMed] [Google Scholar]
- 26.Huh SY, Andrew R, Rich-Edwards JW, Kleinman KP, Seckl JR, Gillman MW. Association between umbilical cord glucocorticoids and blood pressure at age 3 years. BMC Med 6: 25, 2008. doi: 10.1186/1741-7015-6-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jensen EC, Gallaher BW, Breier BH, Harding JE. The effect of a chronic maternal cortisol infusion on the late-gestation fetal sheep. J Endocrinol 174: 27–36, 2002. doi: 10.1677/joe.0.1740027. [DOI] [PubMed] [Google Scholar]
- 28.Jensen E, Wood CE, Keller-Wood M. Chronic alterations in ovine maternal corticosteroid levels influence uterine blood flow and placental and fetal growth. Am J Physiol Regul Integr Comp Physiol 288: R54–R61, 2005. doi: 10.1152/ajpregu.00149.2004. [DOI] [PubMed] [Google Scholar]
- 29.Jiao K, Kulessa H, Tompkins K, Zhou Y, Batts L, Baldwin HS, Hogan BL. An essential role of Bmp4 in the atrioventricular septation of the mouse heart. Genes Dev 17: 2362–2367, 2003. doi: 10.1101/gad.1124803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jonker SS, Zhang L, Louey S, Giraud GD, Thornburg KL, Faber JJ. Myocyte enlargement, differentiation, and proliferation kinetics in the fetal sheep heart. J Appl Physiol (1985) 102: 1130–1142, 2007. doi: 10.1152/japplphysiol.00937.2006. [DOI] [PubMed] [Google Scholar]
- 31.Keller-Wood M. Inhibition of stimulated and basal ACTH by cortisol during ovine pregnancy. Am J Physiol Regul Integr Comp Physiol 271: R130–R136, 1996. doi: 10.1152/ajpregu.1996.271.1.R130. [DOI] [PubMed] [Google Scholar]
- 32.Keller-Wood M, Feng X, Wood CE, Richards E, Anthony RV, Dahl GE, Tao S. Elevated maternal cortisol leads to relative maternal hyperglycemia and increased stillbirth in ovine pregnancy. Am J Physiol Regul Integr Comp Physiol 307: R405–R413, 2014. doi: 10.1152/ajpregu.00530.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lassar AB, Davis RL, Wright WE, Kadesch T, Murre C, Voronova A, Baltimore D, Weintraub H. Functional activity of myogenic HLH proteins requires hetero-oligomerization with E12/E47-like proteins in vivo. Cell 66: 305–315, 1991. doi: 10.1016/0092-8674(91)90620-E. [DOI] [PubMed] [Google Scholar]
- 34.László KD, Svensson T, Li J, Obel C, Vestergaard M, Olsen J, Cnattingius S. Maternal bereavement during pregnancy and the risk of stillbirth: a nationwide cohort study in Sweden. Am J Epidemiol 177: 219–227, 2013. doi: 10.1093/aje/kws383. [DOI] [PubMed] [Google Scholar]
- 35.Leigh-Brown S, Enriquez JA, Odom DT. Nuclear transcription factors in mammalian mitochondria. Genome Biol 11: 215, 2010. doi: 10.1186/gb-2010-11-7-215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liggins GC. The role of cortisol in preparing the fetus for birth. Reprod Fertil Dev 6: 141–150, 1994. doi: 10.1071/RD9940141. [DOI] [PubMed] [Google Scholar]
- 37.Lindsay RS, Lindsay RM, Edwards CR, Seckl JR. Inhibition of 11β-hydroxysteroid dehydrogenase in pregnant rats and the programming of blood pressure in the offspring. Hypertension 27: 1200–1204, 1996. doi: 10.1161/01.HYP.27.6.1200. [DOI] [PubMed] [Google Scholar]
- 38.Lin YH, Warren CM, Li J, McKinsey TA, Russell B. Myofibril growth during cardiac hypertrophy is regulated through dual phosphorylation and acetylation of the actin capping protein CapZ. Cell Signal 28: 1015–1024, 2016. doi: 10.1016/j.cellsig.2016.05.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li Y, Park JS, Deng JH, Bai Y. Cytochrome c oxidase subunit IV is essential for assembly and respiratory function of the enzyme complex. J Bioenerg Biomembr 38: 283–291, 2006. doi: 10.1007/s10863-006-9052-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Londhe P, Davie JK. Gamma interferon modulates myogenesis through the major histocompatibility complex class II transactivator, CIITA. Mol Cell Biol 31: 2854–2866, 2011. doi: 10.1128/MCB.05397-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lumbers ER, Boyce AC, Joulianos G, Kumarasamy V, Barner E, Segar JL, Burrell JH. Effects of cortisol on cardiac myocytes and on expression of cardiac genes in fetal sheep. Am J Physiol Regul Integr Comp Physiol 288: R567–R574, 2005. doi: 10.1152/ajpregu.00556.2004. [DOI] [PubMed] [Google Scholar]
- 42.Madrigal JL, Olivenza R, Moro MA, Lizasoain I, Lorenzo P, Rodrigo J, Leza JC. Glutathione depletion, lipid peroxidation and mitochondrial dysfunction are induced by chronic stress in rat brain. Neuropsychopharmacology 24: 420–429, 2001. doi: 10.1016/S0893-133X(00)00208-6. [DOI] [PubMed] [Google Scholar]
- 43.Marques-da-Silva D, Francisco R, Webster D, Dos Reis Ferreira V, Jaeken J, Pulinilkunnil T. Cardiac complications of congenital disorders of glycosylation (CDG): a systematic review of the literature. J Inherit Metab Dis 40: 657–672, 2017. doi: 10.1007/s10545-017-0066-y. [DOI] [PubMed] [Google Scholar]
- 44.Mesubi OO, Anderson ME. Atrial remodelling in atrial fibrillation: CaMKII as a nodal proarrhythmic signal. Cardiovasc Res 109: 542–557, 2016. doi: 10.1093/cvr/cvw002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Meysen S, Marger L, Hewett KW, Jarry-Guichard T, Agarkova I, Chauvin JP, Perriard JC, Izumo S, Gourdie RG, Mangoni ME, Nargeot J, Gros D, Miquerol L. Nkx2.5 cell-autonomous gene function is required for the postnatal formation of the peripheral ventricular conduction system. Dev Biol 303: 740–753, 2007. doi: 10.1016/j.ydbio.2006.12.044. [DOI] [PubMed] [Google Scholar]
- 46.Mitsui T, Azuma H, Nagasawa M, Iuchi T, Akaike M, Odomi M, Matsumoto T. Chronic corticosteroid administration causes mitochondrial dysfunction in skeletal muscle. J Neurol 249: 1004–1009, 2002. doi: 10.1007/s00415-002-0774-5. [DOI] [PubMed] [Google Scholar]
- 47.Mlecnik B, Galon J, Bindea G. Comprehensive functional analysis of large lists of genes and proteins. J Proteomics 171: 2–10, 2018. doi: 10.1016/j.jprot.2017.03.016. [DOI] [PubMed] [Google Scholar]
- 48.Morrell NW, Bloch DB, ten Dijke P, Goumans MJ, Hata A, Smith J, Yu PB, Bloch KD. Targeting BMP signalling in cardiovascular disease and anaemia. Nat Rev Cardiol 13: 106–120, 2016. doi: 10.1038/nrcardio.2015.156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mulder EJ, de Heus R, Visser GH. Antenatal corticosteroid therapy: short-term effects on fetal behaviour and haemodynamics. Semin Fetal Neonatal Med 14: 151–156, 2009. doi: 10.1016/j.siny.2008.10.003. [DOI] [PubMed] [Google Scholar]
- 50.Nesan D, Kamkar M, Burrows J, Scott IC, Marsden M, Vijayan MM. Glucocorticoid receptor signaling is essential for mesoderm formation and muscle development in zebrafish. Endocrinology 153: 1288–1300, 2012. doi: 10.1210/en.2011-1559. [DOI] [PubMed] [Google Scholar]
- 51.Nesan D, Vijayan MM. The transcriptomics of glucocorticoid receptor signaling in developing zebrafish. PLoS One 8: e80726, 2013. doi: 10.1371/journal.pone.0080726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Oliveros JC. VENNY. An Interactive Tool for Comparing Lists with Venn Diagrams (Online). http://bioinfogp.cnb.csic.es/tools/venny/index.html.
- 53.Pachori AS, Custer L, Hansen D, Clapp S, Kemppa E, Klingensmith J. Bone morphogenetic protein 4 mediates myocardial ischemic injury through JNK-dependent signaling pathway. J Mol Cell Cardiol 48: 1255–1265, 2010. doi: 10.1016/j.yjmcc.2010.01.010. [DOI] [PubMed] [Google Scholar]
- 54.Parikh SS, Blackwell DJ, Gomez-Hurtado N, Frisk M, Wang L, Kim K, Dahl CP, Fiane A, Tønnessen T, Kryshtal DO, Louch WE, Knollmann BC. Thyroid and glucocorticoid hormones promote functional T-tubule development in human-induced pluripotent stem cell-derived cardiomyocytes. Circ Res 121: 1323–1330, 2017. doi: 10.1161/CIRCRESAHA.117.311920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Pinto JR, Gomes AV, Jones MA, Liang J, Nguyen S, Miller T, Parvatiyar MS, Potter JD. The functional properties of human slow skeletal troponin T isoforms in cardiac muscle regulation. J Biol Chem 287: 37362–37370, 2012. doi: 10.1074/jbc.M112.364927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Posterino GS, Dunn SL, Botting KJ, Wang W, Gentili S, Morrison JL. Changes in cardiac troponins with gestational age explain changes in cardiac muscle contractility in the sheep fetus. J Appl Physiol (1985) 111: 236–243, 2011. doi: 10.1152/japplphysiol.00067.2011. [DOI] [PubMed] [Google Scholar]
- 57.Qi X, Yang G, Yang L, Lan Y, Weng T, Wang J, Wu Z, Xu J, Gao X, Yang X. Essential role of Smad4 in maintaining cardiomyocyte proliferation during murine embryonic heart development. Dev Biol 311: 136–146, 2007. doi: 10.1016/j.ydbio.2007.08.022. [DOI] [PubMed] [Google Scholar]
- 57a.R Core Team R: A Language and Environment for Statistical Computing. Vienna, Austria: R Foundation for Statistical Computing, 2008. https://www.r-project.org/ [Google Scholar]
- 58.Rabaglino MB, Richards E, Denslow N, Keller-Wood M, Wood CE. Genomics of estradiol-3-sulfate action in the ovine fetal hypothalamus. Physiol Genomics 44: 669–677, 2012. doi: 10.1152/physiolgenomics.00127.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Reini SA, Dutta G, Wood CE, Keller-Wood M. Cardiac corticosteroid receptors mediate the enlargement of the ovine fetal heart induced by chronic increases in maternal cortisol. J Endocrinol 198: 419–427, 2008. doi: 10.1677/JOE-08-0022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Richards EM, Wood CE, Rabaglino MB, Antolic A, Keller-Wood M. Mechanisms for the adverse effects of late gestational increases in maternal cortisol on the heart revealed by transcriptomic analyses of the fetal septum. Physiol Genomics 46: 547–559, 2014. doi: 10.1152/physiolgenomics.00009.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Richter-Dennerlein R, Oeljeklaus S, Lorenzi I, Ronsör C, Bareth B, Schendzielorz AB, Wang C, Warscheid B, Rehling P, Dennerlein S. Mitochondrial protein synthesis adapts to influx of nuclear-encoded protein. Cell 167: 471–483.e10, 2016. doi: 10.1016/j.cell.2016.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43: e47, 2015. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Rog-Zielinska EA, Craig MA, Manning JR, Richardson RV, Gowans GJ, Dunbar DR, Gharbi K, Kenyon CJ, Holmes MC, Hardie DG, Smith GL, Chapman KE. Glucocorticoids promote structural and functional maturation of foetal cardiomyocytes: a role for PGC-1α. Cell Death Differ 22: 1106–1116, 2015. doi: 10.1038/cdd.2014.181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Rog-Zielinska EA, Thomson A, Kenyon CJ, Brownstein DG, Moran CM, Szumska D, Michailidou Z, Richardson J, Owen E, Watt A, Morrison H, Forrester LM, Bhattacharya S, Holmes MC, Chapman KE. Glucocorticoid receptor is required for foetal heart maturation. Hum Mol Genet 22: 3269–3282, 2013. doi: 10.1093/hmg/ddt182. [DOI] [PubMed] [Google Scholar]
- 65.Sasse S, Brand NJ, Kyprianou P, Dhoot GK, Wade R, Arai M, Periasamy M, Yacoub MH, Barton PJ. Troponin I gene expression during human cardiac development and in end-stage heart failure. Circ Res 72: 932–938, 1993. doi: 10.1161/01.RES.72.5.932. [DOI] [PubMed] [Google Scholar]
- 66.Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9: 671–675, 2012. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498–2504, 2003. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: e3, 2004. doi: 10.2202/1544-6115.1027. [DOI] [PubMed] [Google Scholar]
- 69.Smyth GK, Michaud J, Scott HS. Use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics 21: 2067–2075, 2005. doi: 10.1093/bioinformatics/bti270. [DOI] [PubMed] [Google Scholar]
- 70.Song CZ, Tian X, Gelehrter TD. Glucocorticoid receptor inhibits transforming growth factor-beta signaling by directly targeting the transcriptional activation function of Smad3. Proc Natl Acad Sci USA 96: 11776–11781, 1999. doi: 10.1073/pnas.96.21.11776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Sun B, Huo R, Sheng Y, Li Y, Xie X, Chen C, Liu HB, Li N, Li CB, Guo WT, Zhu JX, Yang BF, Dong DL. Bone morphogenetic protein-4 mediates cardiac hypertrophy, apoptosis, and fibrosis in experimentally pathological cardiac hypertrophy. Hypertension 61: 352–360, 2013. doi: 10.1161/HYPERTENSIONAHA.111.00562. [DOI] [PubMed] [Google Scholar]
- 72.Tang VM, Young AH, Tan H, Beasley C, Wang JF. Glucocorticoids increase protein carbonylation and mitochondrial dysfunction. Horm Metab Res 45: 709–715, 2013. doi: 10.1055/s-0033-1345119. [DOI] [PubMed] [Google Scholar]
- 74.Tenma T, Mitsuyama H, Watanabe M, Kakutani N, Otsuka Y, Mizukami K, Kamada R, Takahashi M, Takada S, Sabe H, Tsutsui H, Yokoshiki H. Small-conductance Ca2+-activated K+ channel activation deteriorates hypoxic ventricular arrhythmias via CaMKII in cardiac hypertrophy. Am J Physiol Heart Circ Physiol 315: H262–H272, 2018. doi: 10.1152/ajpheart.00636.2017. [DOI] [PubMed] [Google Scholar]
- 75.Thornburg K, Jonker S, O’Tierney P, Chattergoon N, Louey S, Faber J, Giraud G. Regulation of the cardiomyocyte population in the developing heart. Prog Biophys Mol Biol 106: 289–299, 2011. doi: 10.1016/j.pbiomolbio.2010.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Thornell LE, Eriksson A. Filament systems in the Purkinje fibers of the heart. Am J Physiol Heart Circ Physiol 241: H291–H305, 1981. doi: 10.1152/ajpheart.1981.241.3.H291. [DOI] [PubMed] [Google Scholar]
- 77.Unno N, Wong CH, Jenkins SL, Wentworth RA, Ding XY, Li C, Robertson SS, Smotherman WP, Nathanielsz PW. Blood pressure and heart rate in the ovine fetus: ontogenic changes and effects of fetal adrenalectomy. Am J Physiol Heart Circ Physiol 276: H248–H256, 1999. doi: 10.1152/ajpheart.1999.276.1.H248. [DOI] [PubMed] [Google Scholar]
- 78.Valladares E, Peña R, Ellsberg M, Persson LA, Högberg U. Neuroendocrine response to violence during pregnancy—impact on duration of pregnancy and fetal growth. Acta Obstet Gynecol Scand 88: 818–823, 2009. doi: 10.1080/00016340903015321. [DOI] [PubMed] [Google Scholar]
- 79.Vilar L, Freitas MC, Lima LH, Lyra R, Kater CE. Cushing’s syndrome in pregnancy: an overview. Arq Bras Endocrinol Metabol 51: 1293–1302, 2007. doi: 10.1590/S0004-27302007000800015. [DOI] [PubMed] [Google Scholar]
- 80.Viola HM, Hool LC. Cross-talk between L-type Ca2+ channels and mitochondria. Clin Exp Pharmacol Physiol 37: 229–235, 2010. doi: 10.1111/j.1440-1681.2009.05277.x. [DOI] [PubMed] [Google Scholar]
- 81.Walejko JM, Antolic A, Koelmel JP, Garrett TJ, Edison AS, Keller-Wood M. Chronic maternal cortisol excess during late gestation leads to metabolic alterations in the newborn heart. Am J Physiol Endocrinol Metab 316: E000–E000, 2019. doi: 10.1152/ajpendo.00386.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Walejko JM, Koelmel JP, Garrett TJ, Edison AS, Keller-Wood M. Multiomics approach reveals metabolic changes in the heart at birth. Am J Physiol Endocrinol Metab 315: E1212–E1223, 2018. doi: 10.1152/ajpendo.00297.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Wang J, Duncan D, Shi Z, Zhang B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res 41: W77-W83, 2013. doi: 10.1093/nar/gkt439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Wood CE. Are fetal adrenocorticotropic hormone and renin secretion suppressed by maternal cortisol secretion? Am J Physiol Regul Integr Comp Physiol 255: R412–R417, 1988. doi: 10.1152/ajpregu.1988.255.3.R412. [DOI] [PubMed] [Google Scholar]
- 85.Wood CE. Negative-feedback inhibition of fetal ACTH secretion by maternal cortisol. Am J Physiol Regul Integr Comp Physiol 252: R743–R748, 1987. doi: 10.1152/ajpregu.1987.252.4.R743. [DOI] [PubMed] [Google Scholar]
- 86.Wu X, Sagave J, Rutkovskiy A, Haugen F, Baysa A, Nygård S, Czibik G, Dahl CP, Gullestad L, Vaage J, Valen G. Expression of bone morphogenetic protein 4 and its receptors in the remodeling heart. Life Sci 97: 145–154, 2014. doi: 10.1016/j.lfs.2013.12.030. [DOI] [PubMed] [Google Scholar]
- 87.Zhang B, Kirov S, Snoddy J. WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res 33: W741-W748, 2005. doi: 10.1093/nar/gki475. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Differentially expressed genes in Left Ventricle (unadjusted p≤0.05) in LV of CORT newborn lambs as compared to Control lambs. - .docx (60 KB)
Differentially expressed genes in Septum (unadjusted p≤0.05) of CORT newborn lambs as compared to Control lambs. - .docx (35 KB)




