Through a statistical meta-analysis of published data on antimicrobial efficacy against biofilms formed by two common bacterial species, it was concluded that the particular experimental method used is the most important factor determining the outcome of the test. An expected dose-response relationship (greater killing with higher doses or longer treatment times) was observed for data sets derived from a single method but was not observed when data from multiple studies using diverse methods were pooled.
KEYWORDS: antimicrobial agents, biofilm, quantitative methods, statistics
ABSTRACT
Through a statistical meta-analysis of published data on antimicrobial efficacy against biofilms formed by two common bacterial species, it was concluded that the particular experimental method used is the most important factor determining the outcome of the test. An expected dose-response relationship (greater killing with higher doses or longer treatment times) was observed for data sets derived from a single method but was not observed when data from multiple studies using diverse methods were pooled. Method-specific properties such as the surface area/volume ratio, areal biofilm cell density, and microbial species were shown to influence quantitative measurements of biofilm killing. A better appreciation of the method characteristics that affect antibiofilm efficacy tests could aid decision-making related to investment in research and development and regulatory approvals for biofilm control strategies. The following recommendations are offered to those working in research and development related to biofilm control: (i) report the log reduction, surface area/volume ratio, and biofilm areal cell density; (ii) include data for a benchmark agent, making sure that this agent performs competitively at the dose tested; (iii) measure the dose-response relationship, i.e., make measurements at multiple treatment concentrations or dose durations; and (iv) use a standardized method in addition to research methods.
INTRODUCTION
Over the past several years, there has been a flood of literature reports of new antimicrobial chemistries and strategies with efficacy against biofilms. These are important contributions, because established biofilms are highly tolerant to conventional antimicrobial agents (1, 2) and there is pressing demand for better approaches to controlling detrimental biofilms in medicine, dentistry, industry, and consumer environments. Molecules with antibiofilm activity span the chemical and biochemical spectrum and include novel synthetics, plant-derived agents, antimicrobial peptides, and a wide variety of nanotechnologies, photoactivated compounds, quorum-sensing inhibitors, and other creative strategies. The examples are too numerous to cite completely; here we reference a few recent reviews (3–7) and a few primary research articles (8–12) to convey the possibilities and excitement generated by this work.
Which of these approaches should be invested in? How can claims of antimicrobial activity against biofilms be evaluated and compared? Reliable and well-characterized in vitro methods are a critical enabling technology for answering these questions (13–16). There are few standardized methods for evaluating biofilm susceptibility to antimicrobials, and it is not clear how the different methods in the literature compare or which factors govern outcomes. The purpose of the work reported in this article is to analyze literature data on antimicrobial efficacy against biofilms for patterns, to determine the characteristics of the methods being used that are most important for the test outcome, and to make suggestions and recommendations for researchers and developers turning to a biofilm antimicrobial test method.
The analysis pursued here was organized around the concept of an antimicrobial dose-response relationship (17, 18). In this simple version, an increase in biofilm killing (measured as a log reduction [LR] in viable cell counts) is anticipated when the dose of antimicrobial agent (calculated as the product of dose concentration and dose duration [Ctd]) is increased.
RESULTS AND DISCUSSION
Antimicrobial dose-response relationships with biofilms are method dependent.
When biofilm killing (measured as a LR in viable cell counts) was plotted as a function of Ctd, no coherent overall dose-response relationship was observed (Fig. 1A). The coefficient of determination (R2) for a least-squares linear regression of the data set graphed in Fig. 1A (both microorganisms and all antimicrobial agents) was 0.106. One way to interpret this value is that just 10.6% of the variation in LR could be attributed to the linear effect of the antimicrobial dose. Separating the data for the two microorganisms changed the correlations only slightly (Pseudomonas aeruginosa, R2 = 0.0378; Staphylococcus aureus, R2 = 0.2561) (Fig. 1B and C).
FIG 1.
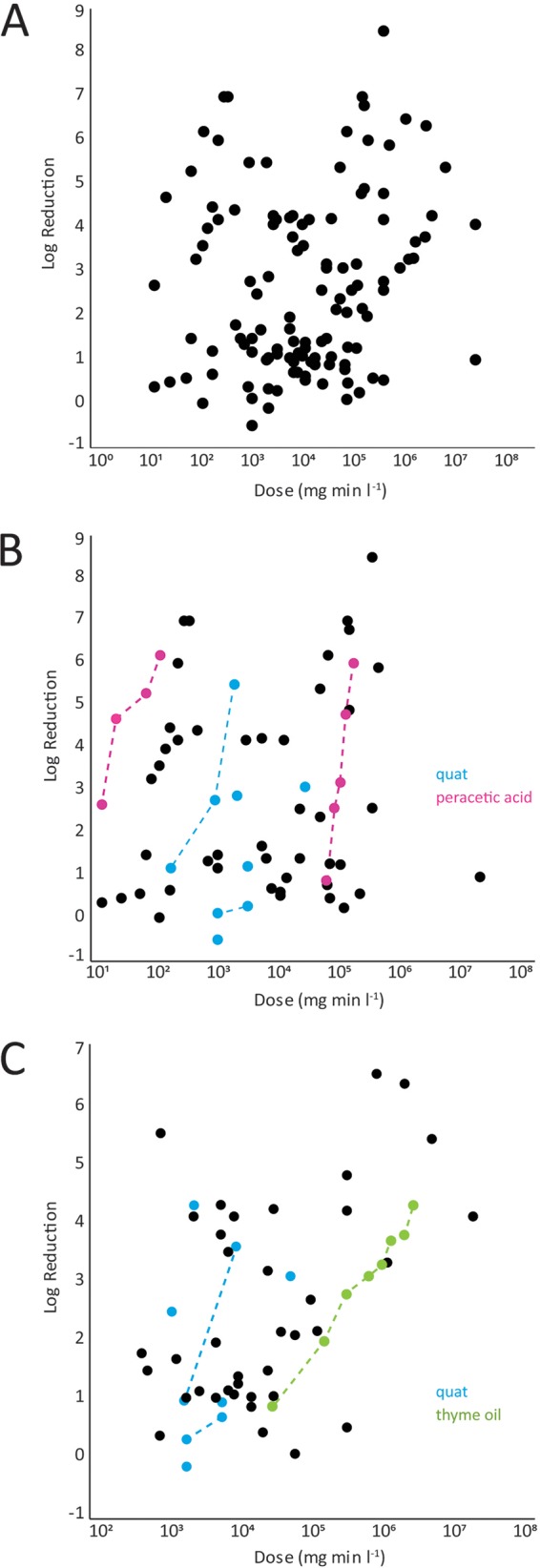
Lack of overall dose-response relationship for biofilm killing by antimicrobial agents. (A) Combined data for P. aeruginosa and S. aureus. (B) Data for P. aeruginosa. (C) Data for S. aureus. Dashed lines connect data points from a single study.
The dose-response relationship is expected to be strongly influenced by the choice of antimicrobial agent, and it is reasonable to begin by inspecting the data for specific antimicrobial agents. Consider the data for quaternary ammonium compounds (QAC) for the P. aeruginosa data set (Fig. 1B, blue symbols). No overall trend is evident in the 9 data points from 4 studies (R2 = 0.2957 [P = 0.1745]). Only when the data points from within an individual study (employing exactly the same method) are connected (Fig. 1B, dotted lines) does the expected dose-response relationship (larger LR with increasing dose) become visually evident. The two highlighted studies differ quantitatively, however; at an exposure of approximately 1,000 mg · min/liter, the study by Campanac et al. (19) found a reduction about 17 times larger than that in the study by Buckingham-Meyer et al. (20). Similar findings and a similar conclusion are arrived at by examining the S. aureus QAC data set over 5 studies (R2 = 0.4197) (Fig. 1C, blue symbols). When data sets from a single study are inspected, they always display the expected dose-response behavior (for example, see the data for thyme oil against S. aureus biofilm in Fig. 1C [green symbols]). Unfortunately, when independent studies of the same agent are compared, quantitative results can vary widely. To underscore this discrepancy, consider two investigations of the action of peracetic acid against P. aeruginosa biofilms (Fig. 1B, magenta symbols). The two studies have internally consistent dose-response behaviors; they differ dramatically, however, in their quantitative determinations of efficacy. To achieve a 3-log-unit reduction, the study by Blanchard et al. (21) indicates an exposure of 16.5 mg · min/liter, whereas an exposure of 84,000 mg · min/liter is suggested by the data of Spoering and Lewis (22), which is a 5,000-fold difference (3 orders of magnitude).
What this qualitative analysis by inspection suggests is that the overwhelming factor determining a dose-response relationship for an antimicrobial biofilm test is the particular method used to make the measurements. This interpretation is borne out by statistical analyses of data for individual antimicrobial agents (Table 1). Within an individual study, LR correlates positively and strongly with Ctd. When data from ≥2 studies are combined, the correlation usually vanishes. The loss of dose-response correlation when data from studies using different methods are combined is apparent in Fig. 2. We conclude that antimicrobial dose-response relationships with biofilms are highly method dependent.
TABLE 1.
Dose-response relationship statistics for antimicrobial agents tested against biofilmsa
| Species and agent | Reference | Mb | Nc | Slope (LR/mg · min) |
P | R2 | AIC | SD(LR)d |
|---|---|---|---|---|---|---|---|---|
| P. aeruginosa | ||||||||
| QAC | 19 | 1 | 3 | 3.946 | 0.237 | 0.8676 | 11.9 | 1.12 |
| Bismuth thiol | 46 | 1 | 3 | 3.316 | 0.0599 | 0.9912 | 0.04 | 0.16 |
| Peracetic acid | 22 | 1 | 5 | 12.0 | 0.0009 | 0.9839 | 5 | 0.29 |
| Peracetic acid | 21 | 1 | 4 | 3.263 | 0.0692 | 0.8664 | 11.3 | 0.66 |
| Chlorine dioxide | 56 | 1 | 11 | 5.525 | <0.0001 | 0.8712 | 33.9 | 0.95 |
| Monochloramine | 58 | 1 | 3 | 13.2 | 0.305 | 0.7881 | 13.5 | 1.46 |
| Peracetic acid | 22, 46 | 2 | 9 | 4.3 | 0.1969 | 0.000 | 37.7 | 11.4 |
| Hydrogen peroxide | 32, 34, 35, 54, 55 | 4 | 4 | −0.3125 | 0.643 | 0.1278 | 9 | 0.51 |
| Chlorine | 20, 46, 55 | 4 | 7 | 1.9377 | 0.0049 | 0.8213 | 27 | 1.06 |
| QAC | 19, 20, 52 | 4 | 9 | 1.72 | 0.1745 | 0.2957 | 36 | 1.99 |
| Fluoroquinolone | 30–32, 46, 53 | 5 | 5 | −0.375 | 0.718 | 0.0498 | 23 | 1.51 |
| Aminoglycoside | 30–32, 37, 46, 53 | 6 | 6 | 0.013 | 0.994 | 1.5 × 10−5 | 31 | 2.23 |
| All | 19–22, 30–32, 34, 35, 37, 46, 52–58 | 18 | 63 | 0.195 | 0.4867 | 0.0378 | 284 | 2.31 |
| S. aureus | ||||||||
| Thyme oil | 42 | 1 | 8 | 1.7163 | <0.0001 | 0.9873 | −5.9 | 0.13 |
| Bismuth thiol | 46 | 1 | 3 | 3.897 | 0.117 | 0.9667 | 7.8 | 0.57 |
| Biguanide/pyridine | 40, 44, 45 | 3 | 3 | 0.1609 | 0.937 | 0.0098 | 15.8 | 2.15 |
| Vancomycin | 25, 33, 49 | 3 | 4 | 2.4771 | 0.1356 | 0.9237 | 13 | 0.94 |
| Chlorine | 20, 40, 41 | 4 | 6 | 0.0686 | 0.8765 | 0.000 | 13 | 0.55 |
| QAC | 19, 20, 36, 39, 40 | 5 | 8 | 2.280 | 0.0263 | 0.4197 | 23 | 1.85 |
| All | 19, 20, 25, 26, 33, 36, 38–51 | 21 | 55 | 0.7825 | <0.0001 | 0.2561 | 200 | 1.48 |
The statistics show strong correlations for data from a single study and mostly poor correlations when data from multiples studies are aggregated. This analysis used data sets when ≥3 data points from a single study, ≥5 points combined for 2 studies, or ≥3 points for ≥3 studies were available. Bold type indicates a negative slope for the regressed line, contrary to the expected positive dose-response relationship.
M indicates the number of studies; the study by Buckingham-Meyer et al. (20) counts as 2 (1 for the CDC biofilm reactor system and 1 for the drip flow biofilm reactor system).
N indicates the total number of data points.
SD(LR) indicates the study reproducibility SD of LRs for M values of >1; for single studies (M = 1), the column indicates only the within-study SD of the LR.
FIG 2.
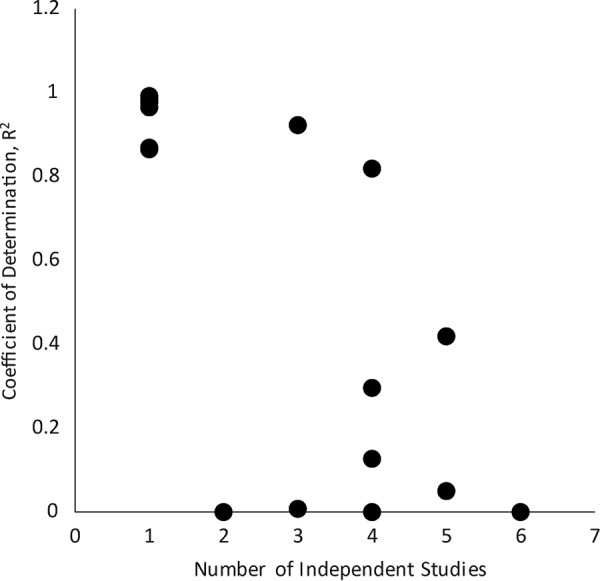
Dose-response correlations for biofilm killing by specific antimicrobial agents diminish as data from multiple studies are aggregated.
Predictors influencing outcomes of antimicrobial biofilm tests.
We can gain some insight into the method features that are most important by briefly digressing to consider the methodological differences between the systems used by Blanchard et al. (21) and by Spoering and Lewis (22). Again, these studies both investigated the effects of peracetic acid on P. aeruginosa biofilms but arrived at quite different results. Blanchard et al. (21) used a modified Robbins device under continuous flow. Biofilm-covered coupons grown in this device were transferred to a clean device and treated with a once-through flow, which resulted in a surface area/volume ratio of approximately 0.002 cm−1. Spoering and Lewis (22) employed the Calgary biofilm device (now marketed as the Innovotech minimum biofilm eradication concentration [MBEC] assay), in which biofilm grows on pegs immersed in the wells of a 96-well plate (23). This is a static system with a much higher surface area/volume ratio of approximately 2 cm−1. The surface density of bacteria was approximately 25 times greater in the system used by Spoering and Lewis (22) (108 CFU/cm2) than in the apparatus used by Blanchard et al. (21) (4 × 106 CFU/cm2). The biofilms in these two systems were challenged at similar biofilm ages. This comparison suggests that flow conditions, biofilm areal density, or surface area/volume ratio might be critical parameters for an antimicrobial biofilm test.
We identified the following five model-dependent parameters that could influence the outcome of a biofilm antimicrobial efficacy test: microbial species, surface area/volume ratio, biofilm areal cell density, static or flow conditions, and biofilm age. The choice of antimicrobial agent also influences the outcome, as discussed above. Linear regression analyses were performed to assess the relative importance of these factors individually in determining the killing rate log10(LR/Ctd); results are tabulated in Table 2, with additional statistical results shown in Table S2 in the supplemental material.
TABLE 2.
Dependence of rates of biofilm killing [log10(LR/Ctd)] on surface area/volume ratio, microbial species, and biofilm areal cell densitya
| Biofilm parameter | Combined |
P. aeruginosa |
S. aureus |
||||||
|---|---|---|---|---|---|---|---|---|---|
| R | P | Slope | R | P | Slope | R | P | Slope | |
| Species | 0.568 | 0.0995 | 0.304 | ||||||
| log10(A/V) | 0.611 | 0.0002 | −0.825 | 0.673 | 0.0081 | −0.882 | 0.329 | 0.016 | −0.754 |
| log10(Xo) | 0.560 | 0.4681 | −0.091 | 0.627 | 0.4536 | −0.186 | 0.304 | 0.053 | −0.299 |
| Flow vs static | 0.569 | 0.0935 | 0.648 | 0.637 | 0.1503 | 0.966 | 0.246 | 0.951 | 0.026 |
| Age | 0.559 | 0.612 | −0.001 | 0.624 | 0.7294 | 0.0011 | 0.253 | 0.462 | −0.002 |
| log10(XoA/V) | 0.582 | 0.011 | −0.277 | 0.672 | 0.0052 | −0.562 | 0.335 | 0.016 | −0.291 |
See Table S2 in the supplemental material for additional statistical results of this analysis. For the categorical factor species, the difference in means between P. aeruginosa and S. aureus is reported. For the categorical factor flow versus static, the difference in means between flow and static conditions is reported.
The method parameter that most consistently and strongly influenced the rate of antimicrobial killing in a biofilm [log10(LR/Ctd)] was the surface area/volume ratio of the test system (for the data set with data for both species, R2 = 0.611, Akaike information criterion [AIC] = 253 [P = 0.0002]) (Table 2). For the combined data set and also for the two species individually (P. aeruginosa, R2 = 0.673, AIC = 127 [P = 0.0081]; S. aureus, R2 = 0.329, AIC = 132 [P = 0.016]), the slopes of the regressed lines were close to −1, indicating a reciprocal dependence. In other words, doubling the surface area/volume ratio approximately halved the killing rate. The effects of flow rate and species were the next most important factors for the combined data set. Age and initial biofilm areal cell density did not have large effects on the killing rate for either species.
Although biofilm age did not emerge in this analysis as an influential predictor, multiple groups have reported clear effects of biofilm age within a given study (1). The absence of correlation with other method-dependent parameters noted does not prove that these parameters are not important. The inherent variability associated with diverse experimental methods makes it difficult to detect the contributions of individual method features. Further research to understand the additional factors that influence the antimicrobial susceptibility of biofilms is warranted. Some of the potentially important factors that come to mind include biofilm age, biofilm thickness (as a proxy for diffusive path length), growth rate, extracellular polymeric substances content, and temperature. The number of factors involved is a reminder to scientists working in the biofilm field to report as many of these factors as possible when describing their work (24).
For each of the P. aeruginosa and S. aureus data sets, we discovered that a combination of surface area/volume ratio and biofilm areal cell density, XoA/V, which was termed biomass density, yielded the strongest effect. This relationship for P. aeruginosa is graphed in Fig. 3A (R2 = 0.672, AIC = 132 [P = 0.005]); the relationship for S. aureus is graphed in Fig. 3B (R2 = 0.335, AIC = 129 [P = 0.017]).
FIG 3.
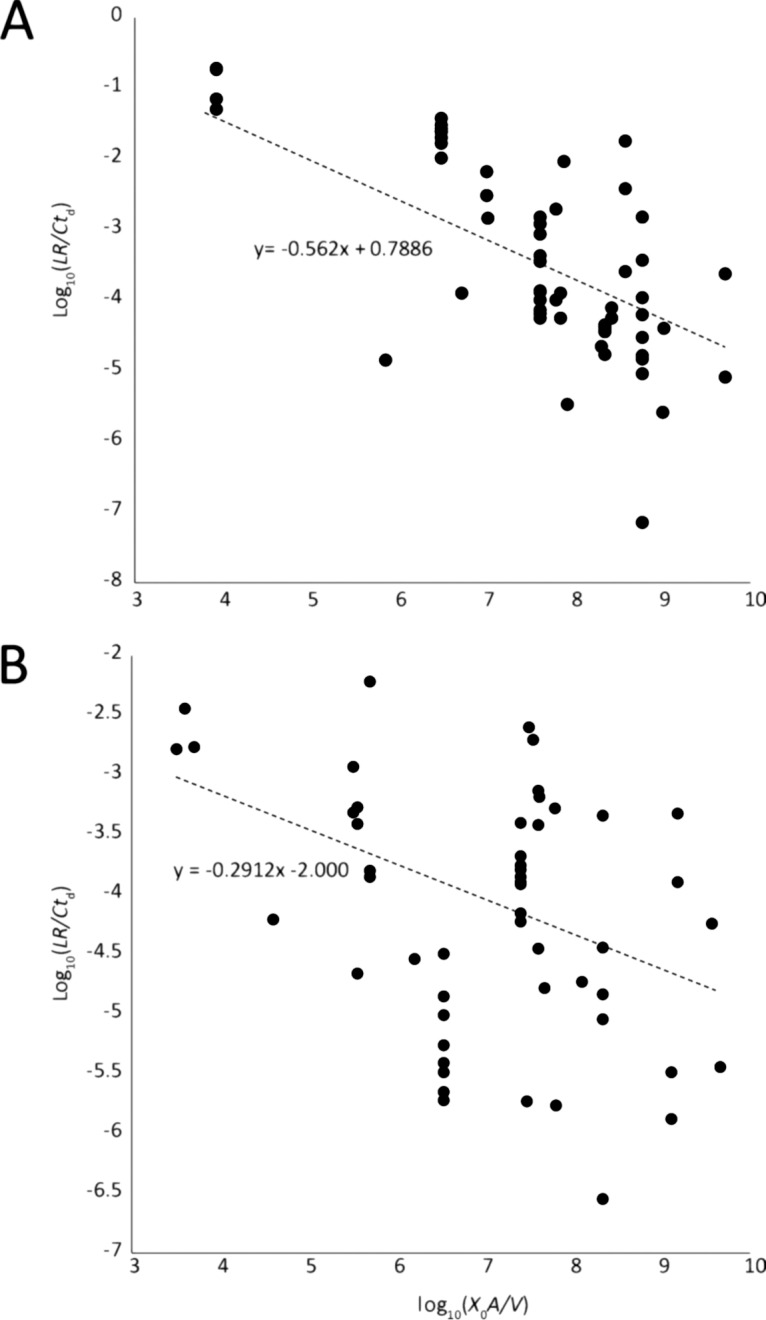
(A) Rates of killing of P. aeruginosa biofilms depend on the biomass density, XoA/V, in the test system. (B) Rates of killing of S. aureus biofilms depend on the surface area/volume ratio, A/V, in the test system.
Identifying superior antibiofilm agents.
We return now to the question of how to compare agents to identify the most effective antimicrobial for a biofilm application. It will be helpful to consider some specific papers to illuminate the issues.
Consider first the synthetic antistaphylococcal pyrrolomycins reported by Yang et al. (8). Biofilm killing was measured in a microtiter plate assay and compared to that observed with vancomycin, a well-known antibiotic. Killing was reported in the article as percent reduction in viable cell numbers. Because it is preferable to work in terms of LR, we made this conversion. A 48-h treatment of S. aureus biofilm with 32 μg/ml vancomycin (22 μM) produced a 0.2-log-unit reduction, whereas a 48-h exposure to 4 μg/ml pyrrolomycin 4 (11 μM) produced a 2.0-log-unit reduction. Thus, the new compound outperforms the benchmark antibiotic. Examining the dose-response relationship for vancomycin (Fig. 4), it can be seen that vancomycin is consistently ineffective (LR against biofilm cells of <0.5 log units) when relatively short exposures are used. Vancomycin works well when prolonged exposures are evaluated (25). In other words, outperforming vancomycin in the Ctd range of 104 to 105 mg · min/liter is not a particularly impressive outcome. In a comparison of the performance of pyrrolomycin 4 to that of other antimicrobials, its efficacy was respectable but not outstanding (Fig. 4). This leads to the suggestion to choose a benchmark agent that exhibits competitive efficacy in the dose range of interest.
FIG 4.
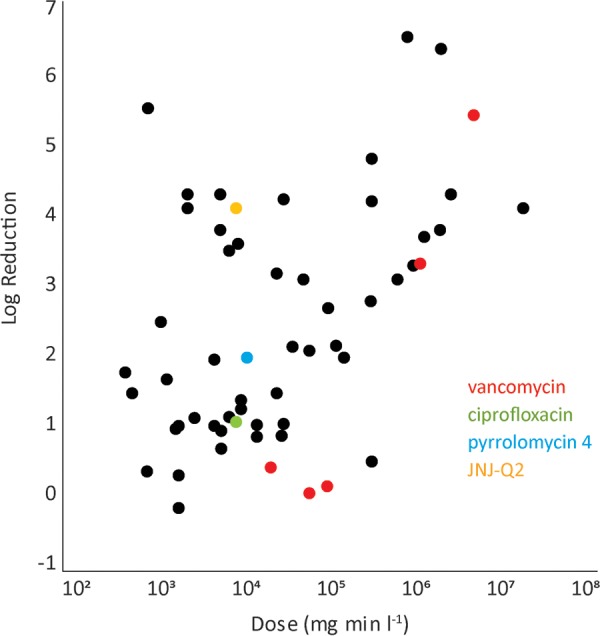
Appropriate benchmark agents facilitate comparisons of antimicrobial activity against biofilms.
Another novel antibiofilm drug was reported by Morrow et al. (26). There the benchmark agent was the antibiotic ciprofloxacin, which falls in the middle of the group in the overall dose-response plot for S. aureus (Fig. 4), with a LR of 1.1 log units at a Ctd value of 8,640 mg · min/liter. The new fluoroquinolone JNJ-Q2 clearly outperforms the benchmark at the same dose (Fig. 4), with a LR of 4.1 log units. The model used by Morrow et al. (26) was robust, in that it involved a 4-day-old biofilm, a high biofilm areal cell density, and a surface area/volume ratio above the median.
Next, we undertook to rank the potency of oxidizing biocides against P. aeruginosa biofilm. We took biomass density into account because we know this parameter is important. As displayed graphically in Fig. 5, biofilm killing rates were ranked in the following order: monochloramine > chlorine and chlorine dioxide > peracetic acid > hydrogen peroxide. Such a ranking of biofilm potency based on measurements from multiple investigations has not been reported previously.
FIG 5.
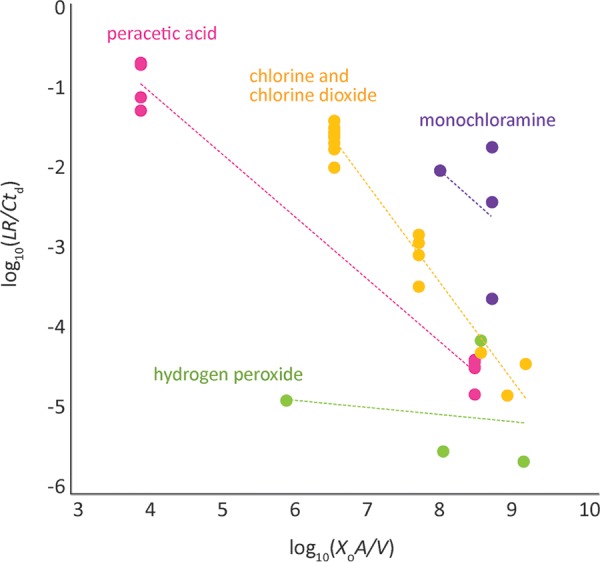
Relative potencies of oxidizing biocides against P. aeruginosa biofilms become apparent when biomass density is taken into account.
It is worth remembering that relative potency is by no means the only or most important property of an antimicrobial agent. In practical applications, cost, environmental impact, material compatibility, cytotoxicity, and safety are all important considerations in the selection of an antimicrobial agent.
One solution to the problem of comparative testing of antibiofilm agents would be for the research community to adopt a common method. Ideally, this common method would have undergone a rigorous standardization process, such as that afforded by the American Society for Testing Materials (ASTM). There are two candidate standard methods for measuring biofilm killing. In 2017, the U.S. Environmental Protection Agency (EPA) issued guidance for testing the efficacy of antimicrobial products against Pseudomonas aeruginosa or Staphylococcus aureus biofilms grown in the CDC biofilm reactor (27, 28). Mature biofilms grown in this high-shear system are used to test antimicrobials using the single-tube method (ASTM method E2871). By design, the single-tube method measures only the LR in viable biofilm bacteria; in other words, removal is not assessed. The second candidate method is the MBEC biofilm disinfection efficacy assay. Formerly known as the Calgary biofilm device, the MBEC system grows biofilm in a low-shear environment, on pegs molded to the top of a microtiter plate lid (23). The mature biofilm is then exposed simultaneously to multiple concentrations of biocide by placing the lid on a new microtiter plate bottom. The MBEC biofilm disinfection test has been standardized and validated as ASTM method E2799 (29). Both of these methods have been validated in interlaboratory studies.
In summary, we performed a meta-analysis of published data on antimicrobial efficacy against biofilms formed by two common bacterial species. We conclude that the particular experimental method used is the most important factor determining the outcome of the test and different methods can produce extremely different results even for the same antimicrobial agent. Method-specific properties such as the surface area/volume ratio, areal biofilm cell density, and microbial species can influence quantitative measurements of biofilm killing. The following recommendations are offered to those working in research and development related to biofilm control: (i) report the LR, surface area/volume ratio, and biofilm areal cell density; (ii) include data for a benchmark agent, making sure that this agent performs competitively at the dose tested; (iii) measure the dose-response relationship, i.e., make measurements at multiple treatment concentrations or dose durations; and (iv) use a standardized method in addition to research methods.
MATERIALS AND METHODS
Meta-analysis.
(i) Data compilation and exclusions. Quantitative data on biofilm killing by antimicrobial agents were gathered from 35 published reports (19–22, 25, 26, 30–58). These papers were identified by searching in the PubMed and Web of Science databases using keywords such as biofilm, antimicrobial, disinfectant, biocide, and log reduction. Approximately one-third of the hits initially identified as containing quantitative data on killing also contained sufficient documentation of other parameters. The following criteria were required for a study to be included in the data set: (i) single-species biofilm of either Pseudomonas aeruginosa or Staphylococcus aureus; (ii) biofilm killing reported as a quantitative LR in viable cell numbers (or LR was readily calculable); (iii) antimicrobial chemistry defined; (iv) quantitative dose concentration (C) and dose duration (td) reported; (v) biofilm areal cell density (Xo) reported for untreated control in CFU per square centimeter (or density was readily calculable); (vi) surface area/volume ratio (A/V) of the biofilm treatment system reported or calculable (specifically, the biofilm-covered area divided by the total volume of antimicrobial treatment solution applied); and (vii) biofilm age at the time of treatment reported. Data for any wild-type strain of P. aeruginosa or S. aureus were used. The data represent 13 different strains for P. aeruginosa and 16 different strains for S. aureus. Data from studies published until early 2017 were included. The resulting collection of data might not be comprehensive for the applied criteria. The data sets are tabulated in Table S1 in the supplemental material.
(ii) Statistical analyses. Least-squares linear regression was used to examine the data from each study separately for the presence of a dose-response relationship and for the influence of individual method-specific parameters on the rate of biofilm killing. This analysis assumed that the rate of decrease of viable cells was proportional to both the antimicrobial agent concentration and the concentration of viable cells, dX/dt = −kCX, where X denotes the biofilm areal cell density, t is time, k is a biofilm killing rate coefficient, and C is the antimicrobial concentration. The solution to this equation, when C is constant, is −ln(X/Xo) = 2.303LR = kCtd, where Xo is the initial biofilm areal cell density and LR = log10(X/Xo). This result predicts that LR should increase linearly with Ctd. To analyze for this dependence, LR values were regressed against Ctd values on a linear scale and the coefficient of determination (R2) was calculated.
To evaluate the dependence of biofilm killing on features of the method used, the specific rate of killing, calculated as log10(LR/Ctd), was regressed against the common logarithm of identified methodological parameters (e.g., surface area/volume ratio), which yielded the coefficient of determination (R2) and the slope. The slope of the log-log plot corresponds to the exponent in a power law dependence. For species and flow versus static conditions, an analysis of variance was fit.
Data across multiple studies were analyzed by linear mixed-effects models (LMMs). These LMMs included a single predictor (as outlined above) and a random effect for study, to estimate the variability of results across studies. All LMMs were fit using the software R (59), package lme4 (60). Model fits were assessed by residual plots. Conditional AIC values were used to compare different LMMs (61), with smaller values indicating better models. Also reported were the study reproducibility standard deviation (SD) (calculated as the square root of the sum of the among-study variance and the residual variance [16]) and the conditional R2, which is a measure of the variability of the response explained by both the predictor and the random effect in the model (e.g., Ctd) (62, 63).
Supplementary Material
ACKNOWLEDGMENTS
Jill Story assisted with the preparation of figures.
This work was supported by NIH/NIGMS grant R01GM109452.
We declare no conflicts of interest.
P.S.S. conceived the meta-analysis, compiled data, performed preliminary statistical analyses, and wrote the manuscript. A.E.P. designed the statistical models, performed the statistical analyses, and wrote and edited sections of the manuscript.
Footnotes
Supplemental material for this article may be found at https://doi.org/10.1128/AAC.00020-19.
REFERENCES
- 1.Stewart PS. 2015. Antimicrobial tolerance in biofilms. Microbiol Spectr doi: 10.1128/microbiolspec.MB-0010-2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hall CW, Mah TF. 2017. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol Rev 41:276–301. doi: 10.1093/femsre/fux010. [DOI] [PubMed] [Google Scholar]
- 3.Stewart PS. 2015. Prospects for anti-biofilm pharmaceuticals. Pharmaceuticals (Basel) 8:504–511. doi: 10.3390/ph8030504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brackman G, Coenye T. 2015. Quorum sensing inhibitors as anti-biofilm agents. Curr Pharm Des 21:5–11. [DOI] [PubMed] [Google Scholar]
- 5.Borges A, Saavedra MJ, Simões M. 2015. Insights on antimicrobial resistance, biofilms and the use of phytochemicals as new antimicrobial agents. Curr Med Chem 22:2590–2614. doi: 10.2174/0929867322666150530210522. [DOI] [PubMed] [Google Scholar]
- 6.Yang L, Givskov M. 2015. Chemical biology strategies for biofilm control. Microbiol Spectr doi: 10.1128/microbiolspec.MB-0019-2015. [DOI] [PubMed] [Google Scholar]
- 7.Moreno MG, Lombardi L, Di Luca M. 2017. Antimicrobial peptides for the control of biofilm formation. Curr Top Med Chem 17:1965–1986. doi: 10.2174/1568026617666170105144830. [DOI] [PubMed] [Google Scholar]
- 8.Yang Z, Liu Y, Ahn J, Qiao Z, Endres JL, Gautam N, Huang Y, Li J, Zheng J, Alnouti Y, Bayles KW, Li R. 2016. Novel fluorinated pyrrolomycins as potent anti-staphylococcal biofilm agents: design, synthesis, pharmacokinetics and antibacterial activities. Eur J Med Chem 124:129–137. doi: 10.1016/j.ejmech.2016.08.017. [DOI] [PubMed] [Google Scholar]
- 9.Basak A, Abouelhassan Y, Zuo R, Yousaf H, Ding Y, Huigens RW. 2017. Antimicrobial peptide-inspired NH125 analogues: bacterial and fungal biofilm-eradicating agents and rapid killers of MRSA persisters. Org Biomol Chem 15:5503–5512. doi: 10.1039/c7ob01028a. [DOI] [PubMed] [Google Scholar]
- 10.Gill RK, Kumar V, Robijns SCA, Steenackers HPL, Van der Eycken EV, Bariwal J. 2017. Polysubstituted 2-aminoimidazoles as anti-biofilm and antiproliferative agents: discovery of potent lead. Eur J Med Chem 138:152–169. doi: 10.1016/j.ejmech.2017.06.043. [DOI] [PubMed] [Google Scholar]
- 11.Kaiser SJ, Mutters NT, Blessing B, Günther F. 2017. Natural isothiocyanates express antimicrobial activity against developing and mature biofilms of Pseudomonas aeruginosa. Fitoterapia 119:57–63. doi: 10.1016/j.fitote.2017.04.006. [DOI] [PubMed] [Google Scholar]
- 12.Zapotoczna M, Boksmati N, Donohue S, Bahtiar B, Boland A, Somali HA, Cox A, Humphreys H, O'Gara JP, Brennan M, O'Neill E. 2017. Novel anti-staphylococcal and anti-biofilm properties of two anti-malarial compounds: MMV665953 {1-(3-chloro-4-fluorophenyl)-3-(3,4-dichlorophenyl)urea} and MMV665807 {5-chloro-2-hydroxy-N-[3-(trifluoromethyl)phenyl]benzamide}. J Med Microbiol 66:377–387. doi: 10.1099/jmm.0.000446. [DOI] [PubMed] [Google Scholar]
- 13.Azeredo J, Azevedo NF, Briandet R, Cerca N, Coenye T, Costa AR, Desvaux M, Di Bonaventura G, Hébraud M, Jaglic Z, Kačániová M, Knøchel S, Lourenço A, Mergulhão F, Meyer RL, Nychas G, Simões M, Tresse O, Sternberg C. 2017. Critical review on biofilm methods. Crit Rev Microbiol 43:313–351. doi: 10.1080/1040841X.2016.1208146. [DOI] [PubMed] [Google Scholar]
- 14.Malone M, Goeres DM, Gosbell I, Vickery K, Jensen S, Stoodley P. 2017. Approaches to biofilm-associated infections: the need for standardized and relevant biofilm methods for clinical applications. Expert Rev Anti Infect Ther 15:147–156. doi: 10.1080/14787210.2017.1262257. [DOI] [PubMed] [Google Scholar]
- 15.Coenye T, Goeres D, Van Bambeke F, Bjarnsholt T. 2018. Should standardized susceptibility testing for microbial biofilms be introduced in clinical practice? Clin Microbiol Infect 24:570–572. doi: 10.1016/j.cmi.2018.01.003. [DOI] [PubMed] [Google Scholar]
- 16.Parker AE, Hamilton MA, Goeres DM. 2018. Reproducibility of antimicrobial test methods. Sci Rep 8:12531. doi: 10.1038/s41598-018-30282-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Axelsen PH. 2002. Essentials of antimicrobial pharmacology: a guide to fundamentals for practice. Humana Press, Totowa, NJ. [Google Scholar]
- 18.Drusano GL. 2004. Antimicrobial pharmacodynamics: critical interactions of ‘bug and drug.’ Nat Rev Microbiol 2:289–300. doi: 10.1038/nrmicro862. [DOI] [PubMed] [Google Scholar]
- 19.Campanac C, Pineau L, Payard A, Baziard-Mouysset G, Roques C. 2002. Interactions between biocide cationic agents and bacterial biofilms. Antimicrob Agents Chemother 46:1469–1474. doi: 10.1128/AAC.46.5.1469-1474.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Buckingham-Meyer K, Goeres DM, Hamilton MA. 2007. Comparative evaluation of biofilm disinfectant efficacy tests. J Microbiol Methods 70:236–244. doi: 10.1016/j.mimet.2007.04.010. [DOI] [PubMed] [Google Scholar]
- 21.Blanchard AP, Bird MR, Wright SJL. 1998. Peroxygen disinfection of Pseudomonas aeruginosa biofilms on stainless steel discs. Biofouling 13:233–253. doi: 10.1080/08927019809378383. [DOI] [Google Scholar]
- 22.Spoering AL, Lewis K. 2001. Biofilms and planktonic cells of Pseudomonas aeruginosa have similar resistance to killing by antimicrobials. J Bacteriol 183:6746–6751. doi: 10.1128/JB.183.23.6746-6751.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ceri H, Olson ME, Stremick C, Read RR, Morck D, Buret A. 1999. The Calgary biofilm device: new technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J Clin Microbiol 37:1771–1776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lourenço A, Coenye T, Goeres DM, Donelli G, Azevedo AS, Ceri H, Coelho FL, Flemming HC, Juhna T, Lopes SP, Oliveira R, Oliver A, Shirtliff ME, Sousa AM, Stoodley P, Pereira MO, Azevedo NF. 2014. Minimum information about a biofilm experiment (MIABiE): standards for reporting experiments and data on sessile microbial communities living at interfaces. Pathog Dis 70:250–256. doi: 10.1111/2049-632X.12146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Post V, Wahl P, Richards RG, Moriarty TF. 2017. Vancomycin displays time-dependent eradication of mature Staphylococcus aureus biofilms. J Orthop Res 35:381–388. doi: 10.1002/jor.23291. [DOI] [PubMed] [Google Scholar]
- 26.Morrow BJ, Abbanat D, Baum EZ, Crespo-Carbone SM, Davies TA, He W, Shang W, Queenan AM, Lynch AS. 2011. Antistaphylococcal activities of the new fluoroquinolone JNJ-Q2. Antimicrob Agents Chemother 55:5512–5521. doi: 10.1128/AAC.00470-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Environmental Protection Agency. 2017. Method and proposed guidance to assess the efficacy of antimicrobial pesticide products intended to control public health biofilms. EPA public docket EPA-HQ-OPP-2016-0357 https://www.regulations.gov/docket?D=EPA-HQ-OPP-2016-0357.
- 28.Goeres D, Loetterle L, Hamilton M, Murga R, Kirby D, Donlan R. 2005. Statistical assessment of a laboratory method for growing biofilms. Microbiology 151:757–762. doi: 10.1099/mic.0.27709-0. [DOI] [PubMed] [Google Scholar]
- 29.Parker A, Walker D, Goeres D, Allan N, Olson M, Omar A. 2014. Ruggedness and reproducibility of the MBEC biofilm AM efficacy test. J Microbiol Methods 102:55–64. doi: 10.1016/j.mimet.2014.04.013. [DOI] [PubMed] [Google Scholar]
- 30.Walters MC, Roe F, Bugnicourt A, Franklin MJ, Stewart PS. 2003. Contributions of antibiotic penetration, oxygen limitation, and low metabolic activity to tolerance of Pseudomonas aeruginosa biofilms to ciprofloxacin and tobramycin. Antimicrob Agents Chemother 47:317–323. doi: 10.1128/AAC.47.1.317-323.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Folsom JP, Richards L, Pitts B, Roe F, Ehrlich GD, Parker A, Mazurie A, Stewart PS. 2010. Physiology of Pseudomonas aeruginosa in biofilms as revealed by transcriptome analysis. BMC Microbiol 10:294. doi: 10.1186/1471-2180-10-294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Liao J, Sauer K. 2012. The MerR-like transcriptional regulator BrlR contributes to Pseudomonas aeruginosa biofilm tolerance. J Bacteriol 194:4823–4836. doi: 10.1128/JB.00765-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Okuda K, Zendo T, Sugimoto S, Iwase T, Tajima A, Yamada S, Sonomoto K, Mizunoe Y. 2013. Effects of bacteriocins on methicillin-resistant Staphylococcus aureus biofilm. Antimicrob Agents Chemother 57:5572–5579. doi: 10.1128/AAC.00888-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stewart PS, Roe F, Rayner J, Elkins JG, Lewandowski Z, Ochsner UA, Hassett DJ. 2000. Effect of catalase on hydrogen peroxide penetration into Pseudomonas aeruginosa biofilms. Appl Environ Microbiol 66:836–838. doi: 10.1128/AEM.66.2.836-838.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wood P, Jones M, Bhakoo M, Gilbert P. 1996. A novel strategy for control of microbial biofilms through generation of biocide at the biofilm-surface interface. Appl Environ Microbiol 62:2598–2602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Luppens SB, Reij MW, van der Heijden RW, Rombouts FM, Abee T. 2002. Development of a standard test to assess the resistance of Staphylococcus aureus biofilm cells to disinfectants. Appl Environ Microbiol 68:4194–4200. doi: 10.1128/AEM.68.9.4194-4200.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Anwar H, Dasgupta M, Lam K, Costerton JW. 1989. Tobramycin resistance of mucoid Pseudomonas aeruginosa biofilm grown under iron limitation. J Antimicrob Chemother 24:647–655. doi: 10.1093/jac/24.5.647. [DOI] [PubMed] [Google Scholar]
- 38.Anwar H, Strap JL, Costerton JW. 1992. Eradication of biofilm cells of Staphylococcus aureus with tobramycin and cephalexin. Can J Microbiol 38:618–625. doi: 10.1139/m92-102. [DOI] [PubMed] [Google Scholar]
- 39.Cabeça TK, Pizzolitto AC, Pizzolitto EL. 2012. Activity of disinfectants against foodborne pathogens in suspension and adhered to stainless steel surfaces. Braz J Microbiol 43:1112–1119. doi: 10.1590/S1517-838220120003000038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ueda S, Kuwabara Y. 2007. Susceptibility of biofilm Escherichia coli, Salmonella enteritidis and Staphylococcus aureus to detergents and sanitizers. Biocontrol Sci 12:149–153. doi: 10.4265/bio.12.149. [DOI] [PubMed] [Google Scholar]
- 41.Agostinho AM, Hartman A, Lipp C, Parker AE, Stewart PS, James GA. 2011. An in vitro model for the growth and analysis of chronic wound MRSA biofilms. J Appl Microbiol 111:1275–1282. doi: 10.1111/j.1365-2672.2011.05138.x. [DOI] [PubMed] [Google Scholar]
- 42.Vázquez-Sánchez D, Cabo ML, Rodríguez-Herrera JJ. 2015. Antimicrobial activity of essential oils against Staphylococcus aureus biofilms. Food Sci Technol Int 21:559–570. doi: 10.1177/1082013214553996. [DOI] [PubMed] [Google Scholar]
- 43.Nostro A, Marino A, Blanco AR, Cellini L, Di Giulio M, Pizzimenti F, Sudano Roccaro A, Bisignano G. 2009. In vitro activity of carvacrol against staphylococcal preformed biofilm by liquid and vapour contact. J Med Microbiol 58:791–797. doi: 10.1099/jmm.0.009274-0. [DOI] [PubMed] [Google Scholar]
- 44.Amalaradjou MA, Venkitanarayanan K. 2014. Antibiofilm effect of octenidine hydrochloride on Staphylococcus aureus, MRSA and VRSA. Pathogens 3:404–416. doi: 10.3390/pathogens3020404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Widodo A, Spratt D, Sousa V, Petrie A, Donos N. 2016. An in vitro study on disinfection of titanium surfaces. Clin Oral Implants Res 27:1227–1232. doi: 10.1111/clr.12733. [DOI] [PubMed] [Google Scholar]
- 46.Folsom JP, Baker B, Stewart PS. 2011. In vitro efficacy of bismuth thiols against biofilms formed by bacteria isolated from human chronic wounds. J Appl Microbiol 111:989–996. doi: 10.1111/j.1365-2672.2011.05110.x. [DOI] [PubMed] [Google Scholar]
- 47.Kirker KR, Fisher ST, James GA. 2015. Potency and penetration of telavancin in staphylococcal biofilms. Int J Antimicrob Agents 46:451–455. doi: 10.1016/j.ijantimicag.2015.05.022. [DOI] [PubMed] [Google Scholar]
- 48.Wolcott RD, Rumbaugh KP, James G, Schultz G, Phillips P, Yang Q, Watters C, Stewart PS, Dowd SE. 2010. Biofilm maturity studies indicate sharp debridement opens a time-dependent therapeutic window. J Wound Care 19:320–328. doi: 10.12968/jowc.2010.19.8.77709. [DOI] [PubMed] [Google Scholar]
- 49.Hall Snyder AD, Vidaillac C, Rose W, McRoberts JP, Rybak MJ. 2015. Evaluation of high-dose daptomycin versus vancomycin alone or combined with clarithromycin or rifampin against Staphylococcus aureus and S. epidermidis in a novel in vitro PK/PD model of bacterial biofilm. Infect Dis Ther 4:51–65. doi: 10.1007/s40121-014-0055-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Chuard C, Vaudaux P, Waldvogel FA, Lew DP. 1993. Susceptibility of Staphylococcus aureus growing on fibronectin-coated surfaces to bactericidal antibiotics. Antimicrob Agents Chemother 37:625–632. doi: 10.1128/AAC.37.4.625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lee JY, Ko KS, Peck KR, Oh WS, Song JH. 2006. In vitro evaluation of the antibiotic lock technique (ALT) for the treatment of catheter-related infections caused by staphylococci. J Antimicrob Chemother 57:1110–1115. doi: 10.1093/jac/dkl098. [DOI] [PubMed] [Google Scholar]
- 52.Abdallah M, Benoliel C, Ferreira-Theret P, Drider D, Dhulster P, Chihib NE. 2015. Effect of culture conditions on the resistance of Pseudomonas aeruginosa biofilms to disinfecting agents. Biofouling 31:49–59. doi: 10.1080/08927014.2014.993390. [DOI] [PubMed] [Google Scholar]
- 53.Goto T, Nakame Y, Nishida M, Ohi Y. 1999. In vitro bactericidal activities of beta-lactamases, amikacin, and fluoroquinolones against Pseudomonas aeruginosa biofilm in artificial urine. Urology 53:1058–1062. doi: 10.1016/S0090-4295(98)00649-9. [DOI] [PubMed] [Google Scholar]
- 54.Elkins JG, Hassett DJ, Stewart PS, Schweizer HP, McDermott TR. 1999. Protective role of catalase in Pseudomonas aeruginosa biofilm resistance to hydrogen peroxide. Appl Environ Microbiol 65:4594–4600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.DeQueiroz GA, Day DF. 2007. Antimicrobial activity and effectiveness of a combination of sodium hypochlorite and hydrogen peroxide in killing and removing Pseudomonas aeruginosa biofilms from surfaces. J Appl Microbiol 103:794–802. doi: 10.1111/j.1365-2672.2007.03299.x. [DOI] [PubMed] [Google Scholar]
- 56.Behnke S, Camper AK. 2012. Chlorine dioxide disinfection of single and dual species biofilms, detached biofilm and planktonic cells. Biofouling 28:635–647. doi: 10.1080/08927014.2012.700705. [DOI] [PubMed] [Google Scholar]
- 57.Sanderson SS, Stewart PS. 1997. Evidence of bacterial adaptation to monochloramine in Pseudomonas aeruginosa biofilms and evaluation of biocide action model. Biotechnol Bioeng 56:201–209. doi: 10.1002/(SICI)1097-0290(19971020)56:2<201::AID-BIT9>3.0.CO;2-T. [DOI] [PubMed] [Google Scholar]
- 58.Chen C-I, Griebe T, Characklis WG. 1993. Biocide action of monochloramine on biofilm systems of Pseudomonas aeruginosa. Biofouling 7:1–17. doi: 10.1080/08927019309386240. [DOI] [Google Scholar]
- 59.R Core Team. 2017. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria: https://www.R-project.org. [Google Scholar]
- 60.Bates D, Maechler M, Bolker B, Walker W. 2015. Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48. [Google Scholar]
- 61.Saefken B, Ruegamer D. 2018. cAIC4: conditional Akaike information criterion for lme4, R package version 0.3. https://CRAN.R-project.org/package=cAIC4.
- 62.Barton K. 2017. MuMIn: multi-model inference, R package version 1.40.0. https://CRAN.R-project.org/package=MuMIn.
- 63.Nakagawa S, Schielzeth H. 2013. A general and simple method for obtaining R2 from generalized linear mixed-effects models. Methods Ecol Evol 4:133–142. doi: 10.1111/j.2041-210x.2012.00261.x. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


