Abstract
The uterine microenvironment during pre-implantation presents a pro-survival milieu and is essential for embryo elongation in ruminants. The European roe deer (Careolus capreolus) pre-implantation embryo development is characterised by a 4-month period of reduced development, embryonic diapause, after which the embryo rapidly elongates and implants. We investigated the uterine fluid proteome by label-free liquid chromatography tandem mass spectrometry at four defined stages covering the phase of reduced developmental pace (early diapause, mid-diapause and late diapause) and embryo elongation. We hypothesised that embryo development during diapause is halted by the lack of signals that support progression past the blastocyst stage. Three clusters of differentially abundant proteins were identified by a self-organising tree algorithm: (1) gradual reduction over development; (2) stable abundance during diapause, followed by a sharp rise at elongation; and (3) gradual increase over development. Proteins in the different clusters were subjected to gene ontology analysis. ‘Cellular detoxification’ in cluster 1 was represented by alcohol dehydrogenase, glutathione S-transferase and peroxiredoxin-2. ATP-citrate synthase, nucleolin, lamin A/C, and purine phosphorylase as cell proliferation regulators were found in cluster 2 and ‘cortical cytoskeleton’, ‘regulation of substrate adhesion-dependent cell spreading’ and ‘melanosome’ were present in cluster 3. Cell cycle promoters were higher abundant at elongation than during diapause, and polyamines presence indicates their role in diapause regulation. This study provides a comprehensive overview of proteins in the roe deer uterine fluid during diapause and forms a basis for studies aiming at understanding the impact of the lack of cell cycle promoters during diapause.
Introduction
The uterine environment changes constantly during early embryo development to support survival and the establishment of pregnancy. Until the blastocyst stage, the embryo develops relatively autonomous and independently of the milieu of the maternal tract (Munoz et al. 2012). Its adopting to different culture conditions with a relatively high degree of flexibility substantiates this. Once the embryo hatches, proteins in the uterine fluid are essential for embryo development past the blastocyst stage. A number of proteins are hypothesised to support embryo elongation (Bazer 1975), which has to date not been shown to occur in vitro. By employing the uterine gland-knockout sheep model, Gray et al. confirmed that glandular secretions are a prerequisite for embryo elongation and establishment of pregnancy (Gray et al. 2001, 2002). The composition of the uterine fluid presents a mixture of embryonic and endometrial secretions. Morphological changes of the embryo coincide with changes of the uterine fluid composition such as proteins, amino acids, lipids and ions (Forde et al. 2014).
Embryonic diapause describes a reproductive strategy in which fertilisation and subsequent embryo implantation are decoupled. In species displaying diapause, the uterine environment supports embryo survival for a prolonged period of time without negatively affecting embryo survival (Renfree & Fenelon 2017). It has been suggested that the uterine secretions supply sufficient nutrients for the embryo to maintain it in a viable state during diapause, but might lack stimulatory factors to initiate resumption of embryonic development (Mead 1993).
The European roe deer (Capreolus capreolus) is one of the species in which embryonic diapause is characterised by very slow intrauterine growth at the hatched blastocyst stage (Renfree & Fenelon 2017). A substantial number of studies explored diapause in roe deer, but until now, the key signals causing entry of diapause and reactivation of embryo development have not been described. In 1854, Bischoff suggested that the reproductive tract of the mother had to be the primary cause of diapause (Bischoff 1854). His large observational study showed that the roe deer buck provides functional semen in June, July and August, and that rut, copulation and fertilisation take place between end of July and end of August (Bischoff 1854). The fertilised egg stays in the oviduct until mid-August and rests in the uterus from mid-August until the end of December (Bischoff 1854). After diapause, roe deer embryonic development resembles that of other ruminants (Bischoff 1854, Keibel 1902). Like in other ruminants, embryo elongation has been associated with the release of secretions from the endometrial glands (Aitken et al. 1973, Lambert 1999). These secretions were found to be up to 1.5-fold increased during implantation compared to diapause (Aitken et al. 1973, Lambert 1999). A first exploratory analysis of the biochemical nature of the uterine fluid during elongation showed a rise in hexose, fructose, total protein, α-amino nitrogen and calcium concentration (Aitken 1974a ,b , 1976). However, there was no association between elongation and secretion of zinc, glucose and prostaglandin F2α (Aitken 1974a ,b ). A more recent study from Lambert (2005) identified a specific secreted protein from the roe deer trophoblast cells, which has the same molecular weight and isoelectric point (pI) as pregnancy-associated glycoproteins (PAGs) (Lambert 2005). The abundance of PAG, which Lambert found elevated in maternal serum after implantation, was hypothesised to induce a rapid increase in maternal serum oestradiol concentrations, which has been linked to endometrial preparation for implantation (Lambert 2005). To our knowledge, no further studies have been conducted to examine the composition of the roe deer uterine fluid. Other mammalian species that undergo diapause show a distinct presence of factors implicated in the regulation of diapause such as leukaemia-inhibiting factor (LIF), insulin-like growth factors (IGF), epidermal growth factor (EGF), platelet-derived growth factor (PDGF), fibroblast growth factor (FGF) and transforming growth factor beta (TGFβ) (Renfree & Shaw 2000). In the European mink, the inhibition of ornithine decarboxylase 1 (ODC1), an enzyme involved in the synthesis of polyamines, has been demonstrated to induce diapause and an increased synthesis of polyamines has been shown to reactivate embryo development (Lefevre et al. 2011). Specifically, it has been proven that polyamine deprivation results in mitotic arrest in embryos (Lefevre et al. 2011) and that hormonal stimulation can regulate the expression of ODC1 via pSTAT1 and mTOR (Fenelon et al. 2016). More recently, and in line with the importance of polyamines, embryonic MYC depletion and mTOR inhibition have been confirmed to induce diapause in mice (Bulut-Karslioglu et al. 2016, Scognamiglio et al. 2016).
Collectively, this emphasises the importance of proteins for embryo survival and indicates a possible role in controlling embryonic diapause. We hypothesise that the roe deer uterine fluid is a pro-survival environment that halts embryo development during the prolonged period of diapause between September and December and that it lacks signals to support embryonic progression past the blastocyst stage. We analysed the protein abundance at four defined developmental stages covering the period of diapause and embryo elongation to gain insight into prolonged survival during embryonic diapause and its regulation.
Methods
Sample collection
The reproductive tract of 225 female roe deer was obtained during regular huntings from September 2015 to January 2016 and September 2016 to January 2017. Ethical approval was neither required nor available as field sampling was performed over the course of regular huntings, where no animals were killed for research purposes. After opening the carcass at the gathering place, the samples were kept on ice until flushing. The average time elapsed between death and uterus removal was 3 h, with a maximum of 4 h. Samples were generally obtained within 1 h after uterus removal. The uterus was freed from connective tissue and flushed with a volume of 2.5 mL phosphate buffered saline (PBS). The embryos were visualised under a stereo microscope (SteREO Discovery Microscope V8, 1:8 Zoom rate, Zeiss), and pictures were taken with the Olympus SC50 camera. The embryo diameter was measured and the embryos were snap frozen in liquid N2 after washing them twice in fresh PBS. The uterine fluid was centrifuged for 10 min at 4°C at 800 × g to pellet any remaining cells. The supernatant was then snap frozen and stored at −80°C until further analysis.
Isolation of embryonic nucleic acids and experimental groups
Embryonic DNA and total RNA were extracted with the Qiagen AllPrep DNA/RNA micro kit (Qiagen, cat.# 80284) according to manufacturers’ instructions. The genomic DNA content was determined with the Promega Quantus and the QuantiFluor® ONE dsDNA System (Promega, cat.# E4870) according to manufacturers’ instructions. Total RNA was stored at −80°C for further analyses. The total genomic DNA content was calculated and used for the estimation of the total number of cells as a proxy for the developmental stage of each embryo. Four groups were defined based on the embryonic genomic DNA content and morphological characteristics (blastocyst or elongated), and n = 5 animals/group were randomly assigned as representative samples for analysis. The four groups referred to as early, mid and late diapause and elongated embryos had a mean ± s.e.m. number of cells of 951 ± 63, 2254 ± 70, 3670 ± 171 and 325,132 ± 40,194, respectively. Early diapause samples were collected between October 21st and November 12th, mid-diapause samples were collected between November 21st and December 3th, late diapause samples were collected between December 5th and December 28th, and elongated samples were collected between November 25th and January 6th.
Quantification of total protein content
Quantification of total protein content in uterine flushing was performed with the Pierce™ BCA Protein Assay Kit (ThermoScientific, prod # 23225) according to manufacturers’ instructions. The standard curve had a range from 0 to 2000 µg/mL BSA. Ten microliter standard and 10 µL undiluted sample were mixed with 200 µL of 50:1 reagent A:B. The plate was shaken for 10 s, followed by 30 min of incubation at 37°C, after which the absorbance was measured at 540 nm.
Generation of the roe deer proteome database
Data processing was performed on our locally installed Galaxy system (version 18.09) (Blankenberg et al. 2010). The protein sequences were obtained by using TransDecoder (Haas et al. 2013, Douglas 2018) with a minimum protein length of 100 amino acids. Identified proteins were annotated using BLASTp (Altschul et al. 1990, Cock et al. 2015) against the human (GRCh38.p12) and bovine (Bos_taurus_UMD_3.1.1) proteome available at NCBI (ftp://ftp.ncbi.nlm.nih.gov/genomes/Homo_sapiens/protein/protein.fa.gz, ftp://ftp.ncbi.nlm.nih.gov/genomes/Bos_taurus/ARCHIVE/ANNOTATION_RELEASE.105/protein/protein.fa.gz).
LC-MS/MS analysis for protein identification and quantification
Twenty-five micrograms of total protein diluted in 45 µL 50 mM NH4HCO3 were reduced using dithioerythritol (DTE) at a final concentration of 5 mM for 30 min at 37°C. Cysteins were blocked using iodoacetamide (final concentration 15 mM) at room temperature for 30 min in the dark. Proteins were first digested for 4 h at 37°C using 250 ng LysC (FUJIFILM Wako Pure Chemicals, Osaka, Japan) and then overnight at 37°C with 500 ng porcine trypsin (Promega). Five micrograms of peptides diluted in 0.1% formic acid (FA) were injected into an Ultimate 3000 (Thermo Scientific) nano-chromatography system and transferred to a trap column (PepMap 100 C18, 100 µm × 2 cm, 5 µM particles, Thermo Scientific) at a flow rate of 20 µL/min of solvent A (0.1% FA). Peptides were separated at 250 nL/min (column: PepMap RSLC C18, 75 µm × 50 cm, 2 µm particles, Thermo Scientific) with a 160 min gradient from 5% solvent A to 25% solvent B (0.1% FA in acetonitrile) and a subsequent 10 min gradient from 25 to 40% solvent B. For MS acquisition a Q Exactive HFX (Thermo Scientific) instrument and a top 15 data-dependent method was used. Ion spray voltage was set to 2.2 kV and MS spectra were acquired at a resolution 60,000 (mass-range: 350–1600). The resolution for MS/MS was set to 15,000.
Data analysis and statistics
The uterine fluid protein concentrations were compared by a one-way ANOVA with Bonferroni post hoc in IBM® SPSS® statistics version 23. Acquired raw proteome files were used for protein identification with the MaxQuant Software by matching the ion spectra against the roe deer proteome. Perseus (Tyanova et al. 2016) was used for subsequent statistical analysis. In brief, the label-free quantification (LFQ) values were log2 transformed and rows were filtered based on valid values, i.e., at least three out of five samples per group in at least one group had valid LFQ values. Subsequently, the imputation feature of Perseus was used to handle missing values. Statistically significant differences in relative abundance of proteins were assessed by a Student’s t-test, where different developmental stages were compared. Proteins were considered differentially abundant if the Benjamini–Hochberg adjusted P value was ≤0.05 (q value) and the fold-change was ≥1.5. A principle component analysis (PCA), protein abundance visualisation and self-organising tree algorithm (SOTA) were performed in R with prcomp (Wickham 2016), UpSet (Lex et al. 2014) and clValid (Brock et al. 2008), respectively. Gene ontology (GO) (Ashburner et al. 2000) and pathway analysis was conducted using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) with the standard parameter set (Dennis et al. 2003). Functional networks integrating the GO-Biological Process, GO-Cellular Component and GO-Molecular Function were graphically displayed using the ClueGO plug-in (Bindea et al. 2009, 2013) in Cytoscape 3.6.1 (Shannon et al. 2003). A list of all identified proteins including LFQ values can be found in Supplementary Table 1 (see section on supplementary data given at the end of this article).
Results
Protein quantification
The uterine fluid protein concentration was not statistically significantly different between the four defined developmental stages. Mean ± s.e.m. protein concentrations for early, mid and late diapause and elongated were 1495 ± 342, 1694 ± 386, 1323 ± 116 and 2039 ± 613 µg/mL, respectively.
Distinctive developmental stage protein abundancies
A total of 819 proteins were identified and quantified with a false discovery rate (FDR) of <1%. The mean ± s.e.m. number of quantified proteins was 733 ± 39, 589 ± 49, 534 ± 75 and 637 ± 66 at early, mid, late diapause and elongation. The most abundant protein was serum albumin precursor. A total of 107 differentially abundant proteins (DAP) was identified between either one of the six comparisons, while the remaining 712 proteins remained stable irrespective of embryonic developmental stage.
A PCA of the DAP showed clustering of the early and mid-diapause, and of two separate clusters representing the late diapause and elongated protein profile (Fig. 1A). Most differentially abundant proteins were identified in the uterine fluid comparing early diapause versus elongated embryos (94 DAP, 54 uniquely for early vs elongated), followed by mid-diapause vs elongated (44 DAP, 7 uniquely for mid vs elongated), late diapause vs elongated (18 DAP, 5 uniquely for late vs elongated) and early vs late (1 DAP) and mid vs late diapause (1 uniquely DAP) (Fig. 1B). The single DAP between early and late diapause was IST1 homolog isoform X1, and the single DAP between mid and late diapause was dynactin subunit 3. A full list of DAP with log2 fold-changes is show in Table 1.
Figure 1.
(A) Principle component analysis of differentially abundant proteins in the roe deer uterine fluid. Principle components 1 and 2 explain 61.7 and 8.5% of the variation, respectively. (B) Distribution of the number of DAP between different developmental stages. The dots indicate the DAP for a given comparison, and dots connected with lines indicate that the developmental stage comparisons share this number of DAP.
Table 1.
Protein list of differentially abundant proteins with bovine RefSeq protein accession number, proteins name, log2 fold-change (FC) and q-value.
| Developmental stage | Protein ID | Protein name | log2 FC | q-value |
|---|---|---|---|---|
| Early – Late Diapause | XP_005218560.1 | IST1 homolog isoform X1 | 2.98 | 0.022 |
| Mid – Late Diapause | NP_001069128.1 | Dynactin subunit 3 | −1.90 | 0.005 |
| Late Diapause – Elongated | NP_001015600.1 | Keratin, type I cytoskeletal 19 | −5.70 | 0.0375 |
| NP_776393.1 | Thioredoxin | −5.59 | 0.0003 | |
| NP_001068616.1 | 78 kDa glucose-regulated protein precursor | −5.28 | 0.0140 | |
| XP_002690791.3 | Purine nucleoside phosphorylase | −4.64 | 0.0205 | |
| XP_015331087.1 | Ig heavy chain Mem5-like | −4.57 | 0.0121 | |
| NP_776771.1 | 10 kDa heat shock protein, mitochondrial | −4.41 | 0.0139 | |
| XP_005225065.1 | Malate dehydrogenase, mitochondrial isoform X1 | −4.38 | 0.0207 | |
| NP_001039876.1 | Keratin, type II cytoskeletal 7 | −4.38 | 0.0179 | |
| NP_001179024.1 | Keratin, type I cytoskeletal 18 | −4.37 | 0.0403 | |
| NP_001019738.1 | Cystathionine gamma-lyase | −4.06 | 0.0136 | |
| NP_001193589.1 | Nucleolin | −3.33 | 0.0421 | |
| NP_001035610.1 | 60S ribosomal protein L7a | −2.87 | 0.0205 | |
| XP_005219340.1 | Protein arginine N-methyltransferase 1 isoform X1 | −1.69 | 0.0116 | |
| NP_001028795.1 | Phosphatidylethanolamine-binding protein 1 | 0.87 | 0.0381 | |
| XP_010814868.1 | Growth factor receptor-bound protein 2 isoform X1 | 1.36 | 0.0218 | |
| NP_777040.1 | Superoxide dismutase (Cu-Zn) | 1.40 | 0.0403 | |
| NP_776917.2 | 14-3-3 protein eta | 1.85 | 0.0222 | |
| NP_001091040.1 | Biliverdin reductase A | 2.33 | 0.0413 | |
| Mid-Diapause – Elongated | NP_001028782.1 | Keratin, type II cytoskeletal 8 | −5.41 | 0.0339 |
| NP_001068616.1 | 78 kDa glucose-regulated protein precursor | −5.16 | 0.0113 | |
| XP_002690791.3 | Purine nucleoside phosphorylase | −5.05 | 0.0113 | |
| NP_776771.1 | 10 kDa heat shock protein, mitochondrial | −4.91 | 0.0070 | |
| NP_001179024.1 | Keratin, type I cytoskeletal 18 | −4.82 | 0.0073 | |
| XP_005225065.1 | Malate dehydrogenase, mitochondrial isoform X1 | −4.52 | 0.0141 | |
| NP_001193589.1 | Nucleolin | −4.33 | 0.0042 | |
| XP_010819218.1 | Protein SET-like | −4.26 | 0.0087 | |
| XP_015313986.1 | 60 kDa heat shock protein, mitochondrial isoform X1 | −4.12 | 0.0477 | |
| NP_001019738.1 | Cystathionine gamma-lyase | −3.89 | 0.0087 | |
| XP_002687260.2 | Early endosome antigen 1 isoform X2 | −3.87 | 0.0109 | |
| NP_001178987.1 | Annexin A10 | −3.69 | 0.0181 | |
| NP_777125.1 | Endoplasmin precursor | −3.12 | 0.0307 | |
| XP_005220772.1 | ATP-citrate synthase isoform X2 | −2.97 | 0.0324 | |
| NP_776993.1 | Polyadenylate-binding protein 1 | −2.72 | 0.0069 | |
| XP_005211893.1 | Kinectin isoform X5 | −2.48 | 0.0322 | |
| NP_001039500.1 | Tissue alpha-L-fucosidase precursor | −2.37 | 0.0391 | |
| NP_786984.1 | Isocitrate dehydrogenase (NADP), mitochondrial precursor | −2.22 | 0.0197 | |
| XP_005219340.1 | Protein arginine N-methyltransferase 1 isoform X1 | −2.06 | 0.0081 | |
| NP_001091521.1 | Alpha-actinin-4 | −2.06 | 0.0336 | |
| XP_005209627.1 | Dihydropyrimidinase-related protein 3 isoform X1 | −1.92 | 0.0397 | |
| NP_001263282.1 | Lamin-B2 | −1.72 | 0.0409 | |
| NP_001030360.2 | Transaldolase | −1.57 | 0.0319 | |
| XP_005208438.1 | Heterogeneous nuclear ribonucleoprotein H isoform X2 | −1.42 | 0.0325 | |
| XP_005222963.1 | Filamin-B isoform X3 | −1.00 | 0.0461 | |
| NP_001029487.1 | Adenosylhomocysteinase | −0.87 | 0.0509 | |
| NP_776404.2 | Actin, cytoplasmic 1 | −0.80 | 0.0313 | |
| XP_005226540.1 | Vinculin isoform X3 | −0.65 | 0.0276 | |
| NP_776770.2 | Heat shock cognate 71 kDa protein | −0.43 | 0.0184 | |
| NP_001029421.1 | Alcohol dehydrogenase class-3 | 0.84 | 0.0324 | |
| XP_010814868.1 | Growth factor receptor-bound protein 2 isoform X1 | 1.21 | 0.0144 | |
| NP_777040.1 | Superoxide dismutase (Cu-Zn) | 1.36 | 0.0319 | |
| XP_015328655.1 | Talin-2 | 1.37 | 0.0387 | |
| NP_001180044.1 | Ras-related protein Rab-6A | 1.41 | 0.0190 | |
| XP_010805021.1 | Peroxiredoxin-2 isoform X1 | 1.51 | 0.0311 | |
| NP_001030358.1 | Prostaglandin reductase 1 | 1.77 | 0.0180 | |
| XP_010806674.1 | Unconventional myosin-VI isoform X7 | 2.12 | 0.0115 | |
| NP_776403.1 | Low–molecular-weight phosphotyrosine protein phosphatase | 2.14 | 0.0192 | |
| NP_001103431.1 | glutamine-fructose-6-phosphate aminotransferase (isomerising) 1 | 2.20 | 0.0420 | |
| XP_005217265.1 | Regulatory solute carrier protein family 1 member 1 isoform X1 | 2.27 | 0.0500 | |
| XP_010806860.1 | Band 4.1-like protein 2 isoform X4 | 2.57 | 0.0262 | |
| NP_001091040.1 | Biliverdin reductase A | 2.98 | 0.0052 | |
| NP_001070444.2 | Uteroglobin precursor | 4.31 | 0.0111 | |
| NP_001095620.1 | Phosphoserine aminotransferase | 5.18 | 0.0201 | |
| Early Diapause – Elongated | NP_001028782.1 | Keratin, type II cytoskeletal 8 | −5.65 | 0.0414 |
| XP_002690791.3 | Purine nucleoside phosphorylase | −5.21 | 0.0056 | |
| NP_776393.1 | Thioredoxin | −4.92 | 0.0427 | |
| NP_776771.1 | 10 kDa heat shock protein, mitochondrial | −4.78 | 0.0044 | |
| XP_005225065.1 | Malate dehydrogenase, mitochondrial isoform X1 | −4.75 | 0.0074 | |
| NP_001068616.1 | 78 kDa glucose-regulated protein precursor | −4.68 | 0.0238 | |
| NP_001179024.1 | Keratin, type I cytoskeletal 18 | −4.42 | 0.0228 | |
| XP_015313986.1 | 60 kDa heat shock protein, mitochondrial isoform X1 | −4.33 | 0.0283 | |
| NP_001019738.1 | Cystathionine gamma-lyase | −3.82 | 0.0117 | |
| NP_001193589.1 | Nucleolin | −3.67 | 0.0325 | |
| XP_005203678.1 | Lamin isoform X1 | −3.63 | 0.0472 | |
| NP_776993.1 | Polyadenylate-binding protein 1 | −3.22 | 0.0019 | |
| NP_001178987.1 | Annexin A10 | −3.19 | 0.0258 | |
| XP_005217320.1 | Glutamine synthetase isoform X1 | −3.18 | 0.0136 | |
| XP_005220772.1 | ATP-citrate synthase isoform X2 | −3.16 | 0.0139 | |
| NP_777125.1 | Endoplasmin precursor | −3.10 | 0.0129 | |
| XP_005211893.1 | Kinectin isoform X5 | −2.95 | 0.0128 | |
| XP_002687260.2 | Early endosome antigen 1 isoform X2 | −2.95 | 0.0419 | |
| XP_004008889.1 | Stress-70 protein, mitochondrial isoform X1 | −2.80 | 0.0421 | |
| NP_001069978.2 | Beta-hexosaminidase subunit beta preproprotein | −2.64 | 0.0413 | |
| XP_012023387.1 | Ribosome-binding protein 1 isoform X2 | −2.42 | 0.0142 | |
| XP_005214667.2 | Microtubule-associated protein RP/EB family member 1 isoform X1 | −2.40 | 0.0398 | |
| XP_010811610.1 | Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 isoform X1 | −2.38 | 0.0261 | |
| XP_005219340.1 | Protein arginine N-methyltransferase 1 isoform X1 | −2.35 | 0.0011 | |
| NP_777231.1 | Aspartate aminotransferase, mitochondrial | −2.28 | 0.0487 | |
| NP_991353.1 | Staphylococcal nuclease domain-containing protein 1 | −2.26 | 0.0400 | |
| NP_786984.1 | Isocitrate dehydrogenase [NADP], mitochondrial precursor | −2.21 | 0.0168 | |
| NP_001263282.1 | Lamin-B2 | −2.20 | 0.0321 | |
| NP_001095381.2 | Protein RCC2 | −2.04 | 0.0349 | |
| NP_001091521.1 | Alpha-actinin-4 | −1.89 | 0.0125 | |
| NP_001020505.1 | Adenine phosphoribosyltransferase | −1.87 | 0.0365 | |
| NP_001095811.1 | Galectin-3 | −1.74 | 0.0049 | |
| NP_001030396.1 | Ribonuclease inhibitor | −1.61 | 0.0342 | |
| NP_001029666.1 | Proliferating cell nuclear antigen | −1.56 | 0.0417 | |
| XP_010806399.1 | Heterogeneous nuclear ribonucleoprotein K isoform X2 | −1.39 | 0.0279 | |
| XP_005208438.1 | Heterogeneous nuclear ribonucleoprotein H isoform X2 | −1.33 | 0.0137 | |
| NP_001015565.1 | Poly(rC)-binding protein 1 | −1.19 | 0.0058 | |
| NP_001030360.2 | Transaldolase | −1.18 | 0.0411 | |
| NP_001029487.1 | Adenosylhomocysteinase | −1.12 | 0.0477 | |
| NP_776706.2 | Coatomer subunit beta | −1.08 | 0.0500 | |
| NP_001030524.1 | Actin-related protein 2/3 complex subunit 5 | −1.07 | 0.0319 | |
| NP_776404.2 | Actin, cytoplasmic 1 | −1.05 | 0.0166 | |
| NP_001029885.1 | Actin-related protein 2/3 complex subunit 2 | −0.97 | 0.0243 | |
| XP_005222963.1 | Filamin-B isoform X3 | −0.96 | 0.0064 | |
| XP_005226540.1 | Vinculin isoform X3 | −0.65 | 0.0071 | |
| NP_776770.2 | Heat shock cognate 71 kDa protein | −0.55 | 0.0318 | |
| NP_001028795.1 | Phosphatidylethanolamine-binding protein 1 | 0.85 | 0.0429 | |
| XP_010815961.1 | Microtubule-associated protein 4 isoform X10 | 1.06 | 0.0428 | |
| NP_001073766.2 | Switch-associated protein 70 | 1.15 | 0.0410 | |
| XP_010813464.1 | Echinoderm microtubule-associated protein-like 2 isoform X3 | 1.24 | 0.0144 | |
| NP_001180044.1 | Ras-related protein Rab-6A | 1.25 | 0.0450 | |
| XP_010814868.1 | Growth factor receptor-bound protein 2 isoform X1 | 1.27 | 0.0073 | |
| NP_001291895.1 | Phosphoglucomutase-2 | 1.31 | 0.0386 | |
| NP_777040.1 | Superoxide dismutase (Cu-Zn) | 1.36 | 0.0250 | |
| XP_010805021.1 | Peroxiredoxin-2 isoform X1 | 1.43 | 0.0304 | |
| NP_001094668.1 | Chromobox protein homolog 3 | 1.43 | 0.0472 | |
| NP_001039517.1 | Cytoplasmic dynein 1 light intermediate chain 1 | 1.44 | 0.0289 | |
| NP_803470.2 | Glycylpeptide N-tetradecanoyltransferase 1 | 1.53 | 0.0420 | |
| NP_776866.1 | 26S protease regulatory subunit 8 | 1.56 | 0.0338 | |
| NP_001178296.1 | cAMP-dependent protein kinase type II-alpha regulatory subunit | 1.58 | 0.0478 | |
| XP_002686439.1 | Protein FAM151A isoform X1 | 1.66 | 0.0354 | |
| XP_010804133.2 | Tetratricopeptide repeat protein 38 isoform X1 | 1.72 | 0.0426 | |
| NP_001015647.1 | 60S ribosomal protein L10a | 1.81 | 0.0426 | |
| XP_015330819.1 | Arfaptin-1 isoform X3 | 1.83 | 0.0308 | |
| NP_776619.1 | Tubulin-specific chaperone D | 1.91 | 0.0303 | |
| NP_001029871.1 | Proliferation-associated protein 2G4 | 1.91 | 0.0032 | |
| NP_001193081.1 | Cytoplasmic dynein 1 light intermediate chain 2 | 1.92 | 0.0430 | |
| NP_776403.1 | Low-molecular-weight phosphotyrosine protein phosphatase | 1.93 | 0.0472 | |
| NP_001076083.1 | UV excision repair protein RAD23 homolog A | 1.95 | 0.0403 | |
| NP_776850.1 | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-1 | 1.97 | 0.0140 | |
| NP_001039888.1 | 4-trimethylaminobutyraldehyde dehydrogenase | 1.99 | 0.0396 | |
| NP_001014855.1 | 26S protease regulatory subunit 4 | 2.02 | 0.0302 | |
| XP_010803703.1 | Branched-chain-amino-acid aminotransferase, cytosolic isoform X3 | 2.04 | 0.0427 | |
| NP_001073091.1 | Eukaryotic translation initiation factor 3 subunit G | 2.07 | 0.0407 | |
| XP_015330995.1 | Density-regulated protein | 2.11 | 0.0484 | |
| NP_001019739.1 | 40S ribosomal protein S11 | 2.15 | 0.0228 | |
| NP_001180149.1 | ATP-dependent 6-phosphofructokinase, platelet type | 2.15 | 0.0292 | |
| NP_001029452.1 | Leukotriene A-4 hydrolase | 2.15 | 0.0252 | |
| XP_002690482.2 | Ras GTPase-activating-like protein IQGAP2 isoform X1 | 2.23 | 0.0412 | |
| NP_001069351.1 | Cytoplasmic dynein 1 intermediate chain 2 | 2.27 | 0.0239 | |
| NP_001094683.1 | Isochorismatase domain-containing protein 1 | 2.35 | 0.0254 | |
| XP_005218560.1 | IST1 homolog isoform X1 | 2.37 | 0.0044 | |
| NP_001030358.1 | Prostaglandin reductase 1 | 2.37 | 0.0062 | |
| NP_001103431.1 | glutamine-fructose-6-phosphate aminotransferase (isomerising) 1 | 2.42 | 0.0230 | |
| NP_001040025.1 | Glutathione S-transferase Mu 3 | 2.60 | 0.0502 | |
| XP_010802603.1 | Secernin-1 isoform X1 | 2.66 | 0.0384 | |
| NP_001030548.1 | Spermine synthase | 2.78 | 0.0476 | |
| NP_001039509.1 | UMP-CMP kinase | 2.85 | 0.0376 | |
| XP_010806674.1 | Unconventional myosin-VI isoform X7 | 2.90 | 0.0056 | |
| NP_001091040.1 | Biliverdin reductase A | 3.10 | 0.0028 | |
| XP_015316025.1 | Stathmin isoform X1 | 3.43 | 0.0113 | |
| NP_001015613.1 | Creatine kinase B-type | 3.72 | 0.0394 | |
| NP_001070444.2 | Uteroglobin precursor | 3.88 | 0.0403 | |
| NP_001095620.1 | Phosphoserine aminotransferase | 5.34 | 0.0133 |
A positive or negative log2 FC indicate higher abundance at the first or second developmental stage, respectively. Comparisons are as follows: Early – Late Diapause, Mid – Late Diapause, Late Diapause – Elongated, Mid-Diapause – Elongated and Early Diapause – Elongated.
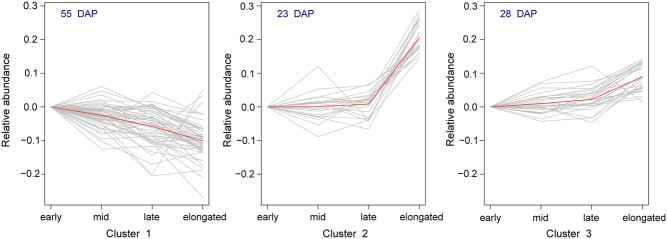
To gain insight into the dynamic changes over development, a SOTA was used. The analysis revealed three clusters (Fig. 2), where the DAP showed one of the following abundance patterns: (1) a gradual decrease over development (55 proteins); (2) stable abundance during the phases of diapause and sharp increase at elongation (23 proteins); and (3) gradual increase over development (28 proteins).
Figure 2.
Self-organising tree algorithm on differentially abundant proteins reveals three clusters, namely clusters showing (1) a gradual decrease over development (55 DAP); (2) a stable abundance during the phases of diapause and sharp increase at elongation (23 DAP) and (3) a gradual increase over development (28 DAP).
Biological functions
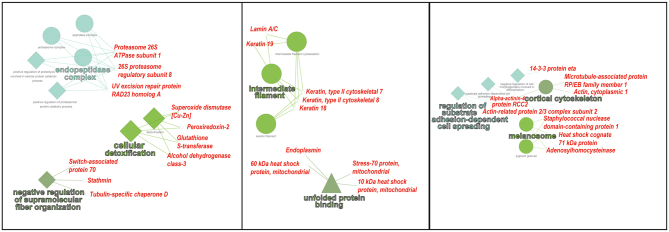
The proteins that were not statistically significantly different between the developmental stages were subjected to GO to identify enriched pathways in proteins that support embryo development during a prolonged period of decelerated embryo development. The top five enriched clusters were GO:0070062~extracellular exosome, GO:0005912~adherens junction, GO:0005913~cell-cell adherens junction, GO:0000502~proteasome complex, and GO:0044445~cytosolic part. A full overview of all enriched clusters can be found in Supplementary Table 2. The biological function of the differentially expressed proteins in the three SOTA-derived clusters was further analysed by subjecting the DAP to ClueGO and CluePedia to visualise the functional interactions (Table 2 and Fig. 3). Proteins in cluster 1, where the relative abundancy of proteins decreased over embryo development, were in the GO categories ‘endopeptidase complex’ (GO:1905369), ‘cellular detoxification’ (GO:1990748) and ‘negative regulation of supramolecular fibre organisation’ (GO:1902904). Proteins in cluster 2, where the abundance is stable during diapause and increase at elongation, were in GO categories ‘intermediate filament’ (GO:0005882) and ‘unfolded protein binding’ (GO:0051082). Lastly, proteins in cluster 3, where protein abundance increases over development, were in GO categories ‘cortical cytoskeleton’ (GO:0030863), ‘regulation of substrate adhesion-dependent cell spreading’ (GO:1900024) and ‘melanosome’ (GO:0042470).
Table 2.
Gene ontology of differentially abundant proteins in cluster 1, 2 and 3 derived from the SOTA analysis.
| Cluster | GO category | GO term | Proteins |
|---|---|---|---|
| Cluster 1 | Endopeptidase complex | GO:1905369 | Proteasome 26S ATPase subunit 1 |
| 26S proteasome regulatory subunit 8 | |||
| UV excision repair protein RAD23 homolog A | |||
| Cellular detoxification | GO:1990748 | Alcohol dehydrogenase class-3 | |
| Glutathione S-transferase | |||
| Peroxiredoxin-2 | |||
| Superoxide dismutase [Cu-Zn] | |||
| Negative regulation of supramolecular fibre organisation | GO:1902904 | Stathmin | |
| Switch-associated protein 70 | |||
| Tubulin-specific chaperone D | |||
| Cluster 2 | Intermediate filament | GO:0005882 | Keratin 18 |
| Keratin 19 | |||
| Keratin, type II cytoskeletal 7 | |||
| Keratin, type II cytoskeletal 8 | |||
| Lamin A/C | |||
| Unfolded protein binding | GO:0051082 | Endoplasmin | |
| 10 kDa heat shock protein, mitochondrial | |||
| Stress-70 protein, mitochondrial | |||
| 60 kDa heat shock protein, mitochondrial | |||
| Cluster 3 | Cortical cytoskeleton | GO:0030863 | Actin, cytoplasmic 1 |
| Alpha-actinin-4 | |||
| Microtubule-associated protein RP/EB family member 1 | |||
| Regulation of substrate adhesion-dependent cell spreading | GO:1900024 | Alpha-actinin-4 | |
| Actin-related protein 2/3 complex subunit 2 | |||
| protein RCC2 | |||
| Melanosome | GO:0042470 | Adenosyl homocysteinase | |
| Heat shock cognate 71 kDa protein | |||
| Staphylococcal nuclease domain-containing protein 1 |
Figure 3.
CluePedia of differentially abundant proteins in the three categories derived from the SOTA analysis. GO-BiologicalProcess, GO-CellularProcess and GO-MolecularFunction are displayed by a square, circle and triangle, respectively. The proteins are displayed in red and the GO terms are displayed in coloured text.
Specific proteins related to diapause and reactivated embryo development
Several DAPs were specifically related to embryonic growth repression and reactivation of embryo development. In cluster 1, there was a high abundance of 4-trimethylaminobutyraldehyde dehydrogenase (ALDH9A1) and spermine synthase (SMS) during early diapause, which were significantly lower abundant during elongation. Both proteins have been implicated in the amine and polyamine biosynthesis pathway. In cluster 2, several proteins including ATP-citrate synthase (ACLY), nucleolin (NCL), lamin A/C (LMNA) and purine phosphorylase (PNP), which are involved in cell proliferation, were higher abundant at elongation than during diapause. Lastly, in cluster 3, protein arginine N-methyltransferase 1 (PRMT1), heterogeneous nuclear ribonucleoprotein K (HNRNPK), aspartate aminotransferase (AST) and tissue alpha-L-fucosidase (FUCA1) were higher abundant at elongation than during diapause.
Discussion
The uterine environment notably changes and adapts to the embryos’ needs to support early development and survival. There was only a limited overlap of the DAP in the present study with DAP reported in other species analysed so far. As field sampling does not allow adequate determination of the day of pregnancy, a direct interspecies comparison was not conducted. Data from different published studies in various species was used to gain insight into potentially important factors for the regulation of diapause and reactivation of embryo development in the roe deer. The total number of 819 identified and quantified uterine fluid proteins falls within the observed range (between 299 and 1359) in various species including cows, mouse, pig and horse (Kayser et al. 2006, Mullen et al. 2012, Forde et al. 2014, 2015, Jalali et al. 2015, Kelleher et al. 2016, Martin et al. 2016, Smits et al. 2018).
We hypothesised to find different abundance profiles, including a gradual increase or decrease during pregnancy, as well as a stable abundance during diapause followed by a sharp increase or decrease at the onset of elongation. In line with the lack of embryo elongation in the uterine gland knockout sheep model (Gray et al. 2002), most DAP in this study were identified in the uterine fluid between early diapause and elongation. Surprisingly, a cluster with stable abundance during diapause, but sharp decrease at elongation did not appear in the present dataset. This potentially indicates an active reactivation rather than an active repression of embryo development.
GO was used to gain insight into the functions of the proteins in the three different clusters. In cluster 1, ‘endopeptidase complex’, ‘cellular detoxification’, and ‘negative regulation of supramolecular fibre organisation’ were represented. The proteins in this clusters showed a significantly lower abundance at elongation than during diapause. Proteins in the ‘endopeptidase complex’ are involved in breaking peptide bonds within proteins. Proteins involved in ‘cellular detoxification’ are recruited to remove toxic substances, whereas proteins involved in ‘negative regulation of supramolecular fibre organisation’ are involved in processes that stop, prevent or reduce the frequency, rate or extent of fibril organisation. None of the proteins representing the ‘endopeptidase complex’ was quantified in cattle on day 10, 13, 16 and 19 of pregnancy (Forde et al. 2014). The proteins alcohol dehydrogenase class-3, glutathione S-transferase and peroxiredoxin-2 in the category ‘cellular detoxification’ and the protein stathmin in the category ‘negative regulation of supramolecular fibre organisation’ decreased like in day 10 to 19 pregnancy in cattle (Forde et al. 2014). In addition, alcohol dehydrogenase class-3 and superoxide dismutase (Cu-Zn) showed a non-significant but gradual decrease from day 3 to day 4 of pregnancy in mice (Kelleher et al. 2016), indicating a species-independent gradual decrease of proteins involved in cellular detoxification before implantation. Moreover, the abundance of peroxiredoxin-2 was significantly higher in pregnant versus cyclic cattle on day 8 of pregnancy (Munoz et al. 2012), the abundance of glutathione S-transferase was significantly higher in pregnant versus cyclic horses on day 13 (Smits et al. 2018), and the abundance of superoxide dismutase (Cu-Zn) increased in uterine fluid from cycle day 10 to 13 in cattle (Mullen et al. 2012). This highlights that these proteins are altered by the presence of an embryo and thus are likely to play an important role during early pregnancy.
In cluster 2, ‘intermediate filament’ and ‘unfolded protein binding’ were represented. The proteins in this clusters showed a sharp increase at elongation and were significantly higher abundant at elongation than during diapause. The proteins in ‘intermediate filament’ are involved in the cytoskeletal structure in eukaryotic cells and keratin 18, keratin, type II cytoskeletal 7, keratin, type II cytoskeletal 8, but not keratin 19 and lamin A/C showed an increased abundance at elongation like in cattle (Forde et al. 2014). All identified keratin proteins consisted of at least one roe deer specific peptide. In the porcine endometrial tissue, the abundance of keratin, type II cytoskeletal 8 was significantly decreased on day 12 versus day 9 of pregnancy (Kayser et al. 2006). A similar abundance pattern was found in mice for keratin 19, which was high on day 3, significantly lower on day 4, but not different between day 4 and 5 in mice (Kelleher et al. 2016). Nevertheless, keratin 19 abundance seems important for ruminant pregnancy, as it was significantly more abundant in uterine fluid of pregnant versus cyclic cattle on day 8 (Munoz et al. 2012). The proteins involved in ‘unfolded protein binding’ are interacting selectively and non-covalently with an unfolded protein, but none of the proteins in that category followed the relatively stable abundance as observed in bovine (Forde et al. 2014). However, it has been shown that the 10 kDa heat shock protein, mitochondrial is produced by the bovine embryo and can be detected in uterine fluid from day 16 onwards, which is in line with the sharp increase in abundance at elongation in the roe deer uterine fluid (Forde et al. 2015).
In cluster 3, proteins involved in ‘cortical cytoskeleton’, ‘regulation of substrate adhesion-dependent cell spreading’ and ‘melanosome’ were represented. The proteins in these clusters showed a significantly higher abundance at elongation than during diapause. The proteins in ‘cortical cytoskeleton’ are proteins that lie just beneath the plasma membrane and actin, cytoplasmic 1, but not alpha-actinin-4, and microtubule-associated protein RP/EB family member 1 showed a gradual increase in cattle like observed in roe deer (Forde et al. 2014). Like in roe deer, actin, cytoplasmic 1 significantly increased from day 9 to day 12 of pregnancy in pigs (Kayser et al. 2006). The proteins in ‘regulation of substrate adhesion-dependent cell spreading’ are proteins that are involved in any process that modulates the frequency, rate or extent of substrate adhesion-dependent cell spreading. Lastly, proteins in the category ‘melanosome’, which is a membrane-bound cytoplasmic organelle within which melanin pigments are synthesised and stored, did not follow the gradual increase observed in roe deer in either cattle or mice (Forde et al. 2014, Kelleher et al. 2016).
The function of individual proteins that were not categorised by GO were assessed to gain insight into their potential role in the regulation of growth repression and/or reactivation of embryo development. The proteins 4-trimethylaminobutyraldehyde dehydrogenase (ALDH9A1) and spermine synthase have been implicated in the amine and polyamine biosynthesis. In roe deer, both proteins decreased in abundance at elongation compared to diapause. In cattle, ALDH9A1 decreased from day 10 to 13 of pregnancy and stayed fairly stable from day 16 to day 19 of pregnancy (Forde et al. 2014). The spermine synthase decreased both from day 10 to 13 and day 16 to 19 of pregnancy in cattle (Forde et al. 2014) and decreased from day 9 to day 12 of pregnancy in the porcine endometrium (Jalali et al. 2015). In light of diapause regulation, the presence of polyamines is particularly interesting, as inhibition of polyamine synthesis causes entry of the mouse (Fenelon & Murphy 2017), hamster (Galliani et al. 1983) and rat (Reddy & Rukmini 1981) blastocyst into embryonic diapause. An increase in putrescine, a polyamine, has been found to be increased at reactivation (Lefevre et al. 2011) and can even cause reactivation of embryo development in mink blastocysts (Fenelon et al. 2016). Ornithine decarboxylase (ODC1), which converts ornithine into polyamine, has a higher abundance in endometrial luminal, glandular epithelium and subepithelial stroma upon embryo reactivation in mink (Lefevre et al. 2011). ODC1 was not identified in the uterine fluid of roe deer, but polyamine synthesis could be rescued by the arginine-agmatine-putrescine (ADC/AGMAT-dependent) pathway, as has previously been confirmed to be the compensatory pathway in ODC1-knockdown sheep embryos (Wang et al. 2014). Spermine synthase catalyses the conversion of spermidine into spermine (both metabolites of putrescine) and ALDH9A1 catalyses the conversion of 4-aminobutanal (a putrescine metabolite) into GABA (Ippolito & Piwnica-Worms 2014). The role of polyamines in the regulation of diapause in roe deer remains speculative. However, putrescine is one of the main drivers of cell proliferation (Fenelon et al. 2016) and can thereby cause embryo reactivation after embryonic diapause. The reduced expression of ALDH9A1 and spermine synthase in roe deer could imply an accumulation of putrescine at the end of diapause, which promotes reactivation of embryo development.
In line with the reactivation of embryo development, proteins involved in cell proliferation including ATP-citrate synthase, nucleolin, lamin A/C and purine phosphorylase were higher abundant at elongation than during diapause in the roe deer. The ATP-citrate synthase, which is the enzyme that synthesises cytosolic acetyl CoA, did not increase at elongation in cattle as observed in roe deer (Forde et al. 2014). However, acetyl CoA has been shown to increase after blastulation in Xenopus laevis and is known to play a role in the regulation of cell proliferation by affecting protein acetylation reactions (Tsuchiya et al. 2014). Nucleolin abundance stayed stable in mice (Kelleher et al. 2016), but has been implicated to play a role in progression through the S-phase of the cell cycle and is thus important for cell proliferation (Xiao et al. 2014). Although important for stem cell differentiation (Sehgal et al. 2013), lamin A/C decreased at elongation in cattle (Forde et al. 2014), whereas it sharply increased at elongation in the roe deer. Lastly, purine nucleoside phosphorylase, an enzyme involved in metabolism of nucleotides with stimulatory effects on cell proliferation (Boehncke et al. 1994) decreased from day 10 to 13 and day 16 to 19 of pregnancy in cattle (Forde et al. 2014).
Proteins with a gradual increase during diapause and elongation included arginine N-methyltransferase 1, heterogeneous nuclear ribonucleoprotein K, aspartate aminotransferase and tissue alpha-L-fucosidase. These proteins play a role in amino acid homeostasis, DNA damage repair and fructose synthesis and are thus important for cell proliferation (Pawlak et al. 2000, Ezawa et al. 2016, Zhang et al. 2016, Feist et al. 2018). In cattle, pigs or mice, none of these proteins has been reported to show a rise prior to implantation as observed in roe deer. Supported by the findings in tammar wallaby (Martin et al. 2016), roe deer embryo development may be reactivated by the presence of positive regulators of the cell cycle. In the tammar wallaby, the high abundance of cell cycle inhibition proteins during diapause switches to increased abundance of growth factors once the embryo resumes development (Martin et al. 2016).
We provide evidence that the abundance of the proteins in the uterine environment of the pre-implantation roe deer embryo remain stable during the period of diapause, while changes occur upon embryo elongation. The observed pattern of DAP, concurrent with the morphological change from the diapausing blastocyst to the elongated embryo, indicates that the uterine fluid proteome during diapause has a high antioxidant capacity and that the embryo is kept in diapause in the absence of polyamines. In addition, embryo development may be actively resumed through the presence of cell cycle promoters at elongation.
Supplementary Material
Declaration of interest
The authors declare that there is no conflict of interest that could be perceived as prejudicing the impartiality of the research reported.
Funding
The study was funded by the Swiss National Science Foundation SNSF (159734).
Author contribution statement
V A v d W performed sample collection, coordinated the experiments, analysed the data and wrote the manuscript. J T B performed the proteome assembly. S B supervised the proteome assembly. T F and G A conducted the proteome experiment and supervised the data analysis. B D rendered possible and coordinated field sampling and performed sample collection. S E U conceptualised the study, coordinated and supervised the project and revised the manuscript. All authors read, edited and approved the final manuscript.
Acknowledgements
The samples were collected in collaboration with local hunters in Switzerland and Southern Germany to which we express our gratitude. The authors thank Alba Rudolf Vegas and Vladimir Milojevic for their valuable help in sample collection and Miwako Kösters for technical assistance. The authors are active participants of the COST Action CA16119 (In vitro 3D total cell guidance and fitness).
References
- Aitken RJ. 1974a. Calcium and zinc in the endometrium and uterine flushings of the roe deer (Capreolus capreolus) during delayed implantation. Journal of Reproduction and Fertility 40 333–340. ( 10.1530/jrf.0.0400333) [DOI] [PubMed] [Google Scholar]
- Aitken RJ. 1974b. Delayed implantation in roe deer (Capreolus capreolus). Journal of Reproduction and Fertility 39 225–233. ( 10.1530/jrf.0.0390225) [DOI] [PubMed] [Google Scholar]
- Aitken RJ. 1976. Uterine secretion of fructose in the roe deer. Journal of Reproduction and Fertility 46 439–440. ( 10.1530/jrf.0.0460439) [DOI] [PubMed] [Google Scholar]
- Aitken RJ, Burton J, Hawkins J, Kerr-Wilson R, Short RV, Steven DH. 1973. Histological and ultrastructural changes in the blastocyst and reproductive tract of the roe deer, Capreolus capreolus, during delayed implantation. Journal of Reproduction and Fertility 34 481–493. ( 10.1530/jrf.0.0340481) [DOI] [PubMed] [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. Journal of Molecular Biology 215 403–410. ( 10.1016/S0022-2836(05)80360-2) [DOI] [PubMed] [Google Scholar]
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT. et al. 2000. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics 25 25–29. ( 10.1038/75556) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bazer FW. 1975. Uterine protein secretions: relationship to development of the conceptus. Journal of Animal Science 41 1376–1382. ( 10.2527/jas1975.4151376x) [DOI] [PubMed] [Google Scholar]
- Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pages F, Trajanoski Z, Galon J. 2009. ClueGO: a cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25 1091–1093. ( 10.1093/bioinformatics/btp101) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bindea G, Galon J, Mlecnik B. 2013. CluePedia cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 29 661–663. ( 10.1093/bioinformatics/btt019) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bischoff TLW. 1854. Die Entwicklungsgeschichte des Rehes. Giessen: J. Bicker’sche Buchhandlung. [Google Scholar]
- Blankenberg D, Gordon A, Von Kuster G, Coraor N, Taylor J, Nekrutenko A. & Galaxy Team 2010. Manipulation of FASTQ data with Galaxy. Bioinformatics 26 1783–1785. ( 10.1093/bioinformatics/btq281) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boehncke WH, Gilbertsen RB, Hemmer J, Sterry W. 1994. Evidence for a pathway independent from 2’-deoxyguanosine and reversible by IL-2 by which purine nucleoside phosphorylase inhibitors block T-cell proliferation. Scandinavian Journal of Immunology 39 327–332. ( 10.1111/j.1365-3083.1994.tb03379.x) [DOI] [PubMed] [Google Scholar]
- Brock G, Pihur V, Datta S, Datta S. 2008. clValid: an R Package for Cluster Validation Journal of Statistical Software 25 22. ( 10.18637/jss.v025.i04) [DOI] [Google Scholar]
- Bulut-Karslioglu A, Biechele S, Jin H, Macrae TA, Hejna M, Gertsenstein M, Song JS, Ramalho-Santos M. 2016. Inhibition of mTOR induces a paused pluripotent state. Nature 540 119–123. ( 10.1038/nature20578) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cock PJA, Chilton JM, Grüning B, Johnson JE, Soranzo N. 2015. NCBI Blast+ integrated into Galaxy. GigaScience 4 39. ( 10.1186/s13742-015-0080-7) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dennis G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA. 2003. DAVID: database for annotation, visualization, and integrated Discovery. Genome Biology 4 P3. [PubMed] [Google Scholar]
- Douglas P. 2018. TransDecoder (find coding regions within transcripts) [Online) (available at: https://github.com/TransDecoder/TransDecoder/wiki; Accessed on 13 January 2019). [Google Scholar]
- Ezawa I, Sawai Y, Kawase T, Okabe A, Tsutsumi S, Ichikawa H, Kobayashi Y, Tashiro F, Namiki H, Kondo T. et al. 2016. Novel p53 target gene FUCA1 encodes a fucosidase and regulates growth and survival of cancer cells. Cancer Science 107 734–745. ( 10.1111/cas.12933) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feist M, Schwarzfischer P, Heinrich P, Sun X, Kemper J, Von Bonin F, Perez-Rubio P, Taruttis F, Rehberg T, Dettmer K. et al. 2018. Cooperative STAT/NF-kappaB signaling regulates lymphoma metabolic reprogramming and aberrant GOT2 expression. Nature Communications 9 1514. ( 10.1038/s41467-018-03803-x) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fenelon JC, Murphy BD. 2017. Inhibition of polyamine synthesis causes entry of the mouse blastocyst into embryonic diapause. Biology of Reproduction 97 119–132. ( 10.1093/biolre/iox060) [DOI] [PubMed] [Google Scholar]
- Fenelon JC, Banerjee A, Lefevre P, Gratian F, Murphy BD. 2016. Polyamine-mediated effects of prolactin dictate emergence from mink obligate embryonic diapause. Biology of Reproduction 95 6. ( 10.1095/biolreprod.116.139204) [DOI] [PubMed] [Google Scholar]
- Forde N, Mcgettigan PA, Mehta JP, O’Hara L, Mamo S, Bazer FW, Spencer TE, Lonergan P. 2014. Proteomic analysis of uterine fluid during the pre-implantation period of pregnancy in cattle. Reproduction 147 575–587. ( 10.1530/REP-13-0010) [DOI] [PubMed] [Google Scholar]
- Forde N, Bazer FW, Spencer TE, Lonergan P. 2015. ‘Conceptualizing’ the endometrium: identification of conceptus-derived proteins During early pregnancy in cattle. Biology of Reproduction 92 156. ( 10.1095/biolreprod.115.129296) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galliani G, Colombo G, Luzzani F. 1983. Contragestational effects of DL-alpha-difluoro-methylornithine, an irreversible inhibitor of ornithine decarboxylase, in the hamster. Contraception 28 159–170. ( 10.1016/0010-7824(83)90015-X) [DOI] [PubMed] [Google Scholar]
- Gray CA, Taylor KM, Ramsey WS, Hill JR, Bazer FW, Bartol FF, Spencer TE. 2001. Endometrial glands are required for preimplantation conceptus elongation and survival. Biology of Reproduction 64 1608–1613. ( 10.1095/biolreprod64.6.1608) [DOI] [PubMed] [Google Scholar]
- Gray CA, Burghardt RC, Johnson GA, Bazer FW, Spencer TE. 2002. Evidence that absence of endometrial gland secretions in uterine gland knockout ewes compromises conceptus survival and elongation. Reproduction 124 289–300. ( 10.1530/rep.0.1240289) [DOI] [PubMed] [Google Scholar]
- Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M. et al. 2013. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nature Protocols 8 1494–1512. ( 10.1038/nprot.2013.084) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ippolito JE, Piwnica-Worms D. 2014. A fluorescence-coupled Assay for gamma aminobutyric acid (GABA) reveals metabolic stress-induced modulation of GABA content in neuroendocrine cancer. PLoS ONE 9. ( 10.1371/journal.pone.0088667) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jalali BM, Bogacki M, Dietrich M, Likszo P, Wasielak M. 2015. Proteomic analysis of porcine endometrial tissue during peri-implantation period reveals altered protein abundance. Journal of Proteomics 125 76–88. ( 10.1016/j.jprot.2015.05.003) [DOI] [PubMed] [Google Scholar]
- Kayser JP, Kim JG, Cerny RL, Vallet JL. 2006. Global characterization of porcine intrauterine proteins during early pregnancy. Reproduction 131 379–388. ( 10.1530/rep.1.00882) [DOI] [PubMed] [Google Scholar]
- Keibel F. 1902. Die Entwicklung Des Rehes bis zur Anlage Des Mesoblast. Anatomia Physiologia Supplement 292. [Google Scholar]
- Kelleher AM, Burns GW, Behura S, Wu G, Spencer TE. 2016. Uterine glands impact uterine receptivity, luminal fluid homeostasis and blastocyst implantation. Scientific Reports 6 38078. ( 10.1038/srep38078) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambert RT. 1999. Conceptus-endometrial interactions and reproductive hormone profiles during embryonic diapause and reactivation of the blastocyst in the European roe deer (Capreolus capreolus). Rangifer 19 41. ( 10.7557/2.19.1.294) [DOI] [PubMed] [Google Scholar]
- Lambert RT. 2005. A pregnancy-associated glycoprotein (Pag) unique to the roe deer (Capreolus capreolus) and its role in the termination of embryonic diapause and maternal recognition of pregnancy. Israel Journal of Zoology 51 1–11. ( 10.1560/NF9R-4CYL-41L0-1577) [DOI] [Google Scholar]
- Lefevre PL, Palin MF, Chen G, Turecki G, Murphy BD. 2011. Polyamines are implicated in the emergence of the embryo from obligate diapause. Endocrinology 152 1627–1639. ( 10.1210/en.2010-0955) [DOI] [PubMed] [Google Scholar]
- Lex A, Gehlenborg N, Strobelt H, Vuillemot R, Pfister H. 2014. UpSet: visualization of intersecting sets. IEEE Transactions on Visualization and Computer Graphics 20 1983–1992. ( 10.1109/TVCG.2014.2346248) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin FC, Ang CS, Gardner DK, Renfree MB, Shaw G. 2016. Uterine flushing proteome of the tammar wallaby after reactivation from diapause. Reproduction 152 491–505. ( 10.1530/REP-16-0154) [DOI] [PubMed] [Google Scholar]
- Mead RA. 1993. Embryonic diapause in vertebrates. Journal of Experimental Zoology 266 629–641. ( 10.1002/jez.1402660611) [DOI] [PubMed] [Google Scholar]
- Mullen MP, Elia G, Hilliard M, Parr MH, Diskin MG, Evans AC, Crowe MA. 2012. Proteomic characterization of histotroph during the preimplantation phase of the estrous cycle in cattle. Journal of Proteome Research 11 3004–3018. ( 10.1021/pr300144q) [DOI] [PubMed] [Google Scholar]
- Munoz M, Corrales FJ, Caamano JN, Diez C, Trigal B, Mora MI, Martin D, Carrocera S, Gomez E. 2012. Proteome of the early embryo-maternal dialogue in the cattle uterus. Journal of Proteome Research 11 751–766. ( 10.1021/pr200969a) [DOI] [PubMed] [Google Scholar]
- Pawlak MR, Scherer CA, Chen J, Roshon MJ, Ruley HE. 2000. Arginine N-methyltransferase 1 is required for early postimplantation mouse development, but cells deficient in the enzyme are viable. Molecular and Cellular Biology 20 4859–4869. ( 10.1128/MCB.20.13.4859-4869.2000) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy PR, Rukmini V. 1981. Alpha-difluoromethylornithine as a postcoitally effective antifertility agent in female rats. Contraception 24 215–221. ( 10.1016/0010-7824(81)90094-9) [DOI] [PubMed] [Google Scholar]
- Renfree MB, Fenelon JC. 2017. The enigma of embryonic diapause. Development 144 3199–3210. ( 10.1242/dev.148213) [DOI] [PubMed] [Google Scholar]
- Renfree MB, Shaw G. 2000. Diapause. Annual Review of Physiology 62 353–375. ( 10.1146/annurev.physiol.62.1.353) [DOI] [PubMed] [Google Scholar]
- Scognamiglio R, Cabezas-Wallscheid N, Thier MC, Altamura S, Reyes A, Prendergast ÁM, Baumgartner D, Carnevalli LS, Atzberger A, Haas S. et al. 2016. Myc depletion induces a pluripotent dormant state mimicking diapause. Cell 164 668–680. ( 10.1016/j.cell.2015.12.033) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sehgal P, Chaturvedi P, Kumaran RI, Kumar S, Parnaik VK. 2013. Lamin A/C haploinsufficiency modulates the differentiation potential of mouse embryonic stem cells. PLoS ONE 8 e57891. ( 10.1371/journal.pone.0057891) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Research 13 2498–2504. ( 10.1101/gr.1239303) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smits K, Willems S, Van Steendam K, van de Velde M, de Lange V, Ververs C, Roels K, Govaere J, van Nieuwerburgh F, Peelman L. et al. 2018. Proteins involved in embryo-maternal interaction around the signalling of maternal recognition of pregnancy in the horse. Scientific Reports 8 5249. ( 10.1038/s41598-018-23537-6) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchiya Y, Pham U, Hu W, Ohnuma S-I, Gout I. 2014. Changes in acetyl CoA levels during the early embryonic development of Xenopus laevis. PLoS ONE 9 e97693. ( 10.1371/journal.pone.0097693) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, Mann M, Cox J. 2016. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nature Methods 13 731–740. ( 10.1038/nmeth.3901) [DOI] [PubMed] [Google Scholar]
- Wang X, Ying W, Dunlap KA, Lin G, Satterfield MC, Burghardt RC, Wu G, Bazer FW. 2014. Arginine decarboxylase and agmatinase: an alternative pathway for de novo biosynthesis of polyamines for development of mammalian conceptuses. Biology of Reproduction 90 84. ( 10.1095/biolreprod.113.114637) [DOI] [PubMed] [Google Scholar]
- Wickham H. 2016. ggplot2: Elegant Graphics for Data Analysis: Springer-Verlag New York. [Google Scholar]
- Xiao S, Caglar E, Maldonado P, Das D, Nadeem Z, Chi A, Trinite B, Li X, Saxena A. 2014. Induced expression of nucleolin phosphorylation-deficient mutant confers dominant-negative effect on cell proliferation. PLoS ONE 9 e109858. ( 10.1371/journal.pone.0109858) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang P, Wang N, Lin X, Jin L, Xu H, Li R, Huang H. 2016. Expression and localization of heterogeneous nuclear ribonucleoprotein K in mouse ovaries and preimplantation embryos. Biochemical and Biophysical Research Communications 471 260–265. ( 10.1016/j.bbrc.2016.02.003) [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.



 This work is licensed under a
This work is licensed under a