Abstract
Background
Recently, gut microbiota have been studied extensively for health promotion, disease prevention, disease treatment, and exercise performance. It is recommended that athletes avoid dietary fiber and resistant starch to promote gastric emptying and reduce gastrointestinal distress during exercise, but this diet may reduce microbial diversity and compromise the health of the athlete’s gut microbiota.
Objective
This study compared fecal microbiota characteristics using high-throughput sequencing among healthy sedentary men (as controls), bodybuilders, and distance runners, as well as the relationships between microbiota characteristics, body composition, and nutritional status.
Methods
Body composition was measured using DXA, and physical activity level was assessed using IPAQ. Dietary intake was analyzed with the computerized nutritional evaluation program. The DNA of fecal samples was extracted and it was sequenced for the analysis of gut microbial diversity through bioinformatics cloud platform.
Results
We showed that exercise type was associated with athlete diet patterns (bodybuilders: high protein, high fat, low carbohydrate, and low dietary fiber diet; distance runners: low carbohydrate and low dietary fiber diet). However, athlete type did not differ in regard to gut microbiota alpha and beta diversity. Athlete type was significantly associated with the relative abundance of gut microbiota at the genus and species level: Faecalibacterium, Sutterella, Clostridium, Haemophilus, and Eisenbergiella were the highest (p < 0.05) in bodybuilders, while Bifidobacterium and Parasutterella were the lowest (p < 0.05). At the species level, intestinal beneficial bacteria widely used as probiotics (Bifidobacterium adolescentis group, Bifidobacterium longum group, Lactobacillus sakei group) and those producing short chain fatty acids (Blautia wexlerae, Eubacterium hallii) were the lowest in bodybuilders and the highest in controls. In addition, aerobic or resistance exercise training with an unbalanced intake of macronutrients and low intake of dietary fiber led to similar diversity of gut microbiota. Specifically, daily protein intake was negatively correlated with operation taxonomic unit (r = − 0.53, p < 0.05), ACE (r = − 0.51, p < 0.05), and Shannon index (r = − 0.64, p < 0.01) in distance runners..
Conclusion
Results suggest that high-protein diets may have a negative impact on gut microbiota diversity for athletes, while athletes in resistance sports that carry out the high protein low carbohydrates diet demonstrate a decrease in short chain fatty acid-producing commensal bacteria.
Keywords: Gut microbiota, Body builder, Distance runner, Dietary fiber
Introduction
Regular exercise offers a beneficial effect on health and a preventive effect on non-communicable diseases by challenging systemic homeostasis [1]. Practically, exercise is recommended as a useful tool to prevent disease and improve the prognosis when an athlete becomes sick or injured. Diseases in which exercise produces a preventive and treatment effect include colon and breast cancer [2, 3], type 2 diabetes [4], sarcopenia [5], cardiovascular diseases [6], and stress-related disorders such as anxiety and depression [7]. Recently, gut microbiota are being studied extensively to understand their effects on health promotion, disease prevention and treatment, and how exercise can modulate these effects [8, 9]. For instance, animal studies have indicated that exercise-induced changes in gut microbiota may be involved in the modulation of high fat-induced obesity [10, 11], polychlorinated biphenyls-induced dysbiosis [12], metabolic syndrome [13], experimental diabetes [14], and chemically-induced colitis [15]. Human studies have also reported that regular exercise plays a beneficial role in host health by affecting the structure and diversity of gut microbiota. For example, subjects with higher cardiorespiratory fitness showed high gut microbiota diversity and a relative abundance of butyrate-producing bacteria, which are important in gut microbiota homeostasis [16, 17]. In particular, high-protein intake with exercise training increased the diversity of gut microbiota in a study comparing the gut microbiota of male rugby players and healthy controls [18].
Clark and Mach (2016) reported that diets recommended for athletes likely influence gut microbiota by reducing diversity because many athletes’ diets have insufficient dietary fiber [19]. It is recommended that athletes consume a high amount of monosaccharides to maximize glycogen storage and sustain blood glucose during exercise training, as well as minimize intake of dietary fiber and resistant starch to prevent gastrointestinal disturbances [20]. Low intake of dietary fiber and resistant starch may lead to decreased bowel movements resulting in decreased bowel function, and also decrease the diversity of gut microbiota [21]. In addition, athletes consume more animal protein than non-athletes to satisfy muscle accretion needs [22]. Excessive protein ingestion leads to an excess of nitrogen substrates in the intestinal microbes, producing putrefactive fermentation products such as ammonia, hydrogen sulfide, amines, phenols, thiols, and indoles [23]. As digesta moves through the intestines, the carbohydrate content decreases, and putrefactive fermentation becomes more harmful [24]. In fact, high protein intake is reported to lead to DNA damage in the colon mucosa when dietary levels of fermentable carbohydrates are low [25–27].
Moreover, high-intensity exercise stimulates redistribution of blood from the intestinal organs to the muscles while they actively undergo cellular respiration [28]. The frequent redistribution of blood could potentially disturb gut microbiota by splanchnic hypo-perfusion and ischemia and subsequent reperfusion [29]. Therefore, to investigate the long-term effects of a specific exercise type and athletes’ diets on gut microbiota, we compared the gut microbiota characteristics, dietary intake, and body composition of healthy men in their twenties who did not have previous exercise habits (control group) with those of athletes in their twenties (bodybuilders and distance runners) who adhered to specific exercise training regimes and diets.
Materials and methods
Subject characteristics and sample treatments
Bodybuilders (n = 15), elite distance runners (n = 15), and healthy men in their twenties without regular exercise habits (n = 15) were recruited for this study. All subjects were male. Bodybuilders were 25 (±3) years old on average and had athletic careers of 7.6 (±3.7) years. Their mean body mass index (BMI) was 28.1 (±2.6) kg/m2. Distance runners were 20 (±1) years old on average and had been runners for 7.5 (±2.1) years. Their mean BMI was 20.5 (±0.8) kg/m2. Healthy men without exercise habits were 26 (±2) years old on average and had a mean BMI of 25.9 (±4.2) kg/m2.
All subjects provided written informed consent prior to beginning the study. This study was conducted after approval was obtained from the Institutional Review Board of Kyung Hee University. Exclusion criteria were prescribed antibiotics within 6 months, immune diseases, digestive tract disorders, acute or chronic cardiovascular diseases, and metabolic disorders. Fecal samples were collected from all participants as the first process of the stuidy. DNA was extracted from fresh stool samples stored on ice, and samples were frozen at − 80 °C immediately after that.
Body composition and physical activity level
Body composition of all subjects was measured using dual-energy X-ray absorptiometry (DXA: Hologic, QDR-4500 W, USA) (Table 1). The DXA was calibrated daily with a phantom, and the coefficient of variance was maintained at less than 1.5%. Each participant wore comfortable clothes without any metal. Whole body scanning was performed for 7 min and the results were analyzed by a technician.
Table 1.
Subject characteristics
| Control (n = 15) |
Bodybuilders (n = 15) |
Distance runners (n = 15) |
|
|---|---|---|---|
| Age (years) | 26.3 ± 2.0 | 24.9 ± 2.7 | 19.8 ± 0.8 |
| Career (years) | 7.6 ± 3.7 | 7.5 ± 2.1 | |
| BMI (kg/m2) | 25.9 ± 4.2+ | 28.1 ± 2.6*** | 20.5 ± 0.9¥¥¥ |
| Lean tissue (kg) | 56.5 ± 4.6+++ | 70.4 ± 9.2*** | 51.8 ± 4.1¥¥ |
| Fat tissue (kg) | 19.4 ± 7.9++ | 11.9 ± 4.5*** | 5.5 ± 1.1¥¥¥ |
| Body fat % | 23.9 ± 6.9+++ | 14 ± 5.1*** | 9.2 ± 1.6¥¥¥ |
Data shown as mean ± SD, controls versus bodybuilders: +p < 0.05 or ++p < 0.01 or +++p < 0.001, bodybuilders versus distance runners: ***p < 0.001, distance runners versus controls: ¥¥p < 0.01 or ¥¥¥p < 0.001
BMI body mass index
Physical activity level was assessed using the International Physical Activity Questionnaire (IPAQ). The Korean version of the IPAQ questionnaire (http://www.ipaq.ki.se) was used and physical activity level was calculated by Metabolic Equivalent of Task (MET) as described in a previous study [30]. In our study, the average physical activity of the subjects in control sedentary group was 860 (±979) METs during their previous 6 months.
Dietary intake data collection
Dietary intake information was obtained from each individual based on a 3-day food diary (2 weekdays and 1 weekend day) that reflected habitual dietary intake. Although self-recorded estimates of food intake in food diaries may not provide accurate or unbiased estimates of a person’s energy intake, the participants in our study were supervised by a specialist to ensure that accurate information was provided. Furthermore, macronutrient and micronutrient supplements were recorded. Daily nutrient intake was analyzed using the nutritional evaluation program CANPro 4.0 and Dietary Reference Intakes for Koreans developed by the Korean Nutrition Society [31].
DNA extraction and high-throughput amplicon sequencing
The DNA of fecal samples was extracted from feces using a Fast DNA™ SPIN extraction kit (MP Biomedicals, Solon, Ohio, USA). The first PCR amplification was performed using a T100 thermal cycler (Bio-Rad, Hercules, CA, USA) to amplify the V3 and V4 regions of 16S rRNA. Primers used were 341F (5′-TCGTCGGCAGCGTC-AGATGTGTATAAGAGACAG-CCTACGGGNGGCWGCAG-3′, the underlined sequence indicates the target region primer) and 805R (5′-GTCTCGTGGGCTCGG-AGATGTGTATAAGAGACAG-GACTACHVGGGTATCTAATCC-3′). The first PCR amplification was carried out under the following conditions: initial denaturation at 95 °C for 3 min followed by 25 cycles of denaturation at 95 °C for 30 s, primer annealing at 55 °C for 30 s, and extension at 72 °C for 30 s, with a final elongation at 72 °C for 5 min. The second PCR amplification to attach the Illumina NexTera barcodes was performed with the i5 forward primer (5′-AATGATACGGCGACCACCGAGATCTACAC-XXXXXXXX-TCGTCGGCAGCGTC-3′, X indicates the barcode region) and the i7 reverse primer (5′-CAAGCAGAAGACGGCATACGAGAT-XXXXXXXX-AGTCTCGTGGGCTCxGG-3′). Conditions used for the second amplification reaction were the same as those described for the first reaction except only eight amplification cycles were performed. Amplification was confirmed by 2% agarose gel electrophoresis and visualization of the PCR products using a Gel Doc system (BioRad). PCR amplification products were purified using a QIAquick PCR purification kit (Qiagen, Valencia, CA, USA). Equal concentrations of purified products were pooled together and short fragments (non-target products) were removed with Ampure beads (Agencourt Bioscience, MA, USA). The size and quality of the amplified product were assessed on a Bioanalyzer 2100 (Agilent, Palo Alto, CA, USA) using a DNA 7500 chip. Mixed amplicons were pooled and sequencing was performed by ChunLab, Inc. (Seoul, Korea) using the Illumina MiSeq Sequencing system (Illumina, USA).
Bioinformatics analysis
Raw reads were quality checked and low-quality reads (<Q25) were filtered out using trimmomatic 0.32 [32]. After the quality control (QC) process, paired-end sequence data were combined using PandaSeq [33]. Primers were then trimmed using a proprietary program of ChunLab using a similarity cut-off of 0.8. Sequences were denoised using mothur’s pre-clustering algorithm, which combines sequences and extracts distinct sequences with two or more differences [34]. The EzTaxon database was blasted using BLAST 2.2.22 for taxonomic assignment [35], and pairwise alignment [36] was utilized to detect chimeras for reads that contained a lower than 97% best hit similarity rate [37]. Sequence data were then clustered by CD-Hit [38] and UCLUST [39], and alpha and beta diversity analysis expressed with OTUs [40], Chao1 [41], ACE [42], Shannon [43] were carried out using BIOiPLUG, which is ChunLab’s bioinformatics cloud platform.
Statistical methods
All data obtained in this study were analyzed using SPSS version 22.0 for Windows (SPSS Inc., Chicago, USA). Dietary and relative abundance of gut microbiota data were visualized with R statistical package, version 3.4.4. The characteristics, body composition, and gut microbiota data of the participants are presented as means and standard deviations, and dietary intake data are presented as medians and inter-quartile ranges. Kruskal-Wallis tests with Bonferroni post hoc tests were used to determine the significance of the differences in gut microbiota composition, alpha diversity, body composition, and dietary intake among groups. The correlations between characteristics of the gut microbiota, dietary intake, and body composition were analyzed using Spearman’s rank correlation analysis. The significance level (α) of all statistical analyses was set to 0.05.
Results
Body composition
There were significant differences in body composition according to exercise type and dietary habits. The body compositions of all subjects are presented in Table 1. Lean tissue was the highest in bodybuilders (70.4 ± 9.2 kg) and lowest in distance runners (51.8 ± 4.1 kg) (bodybuilders versus distance runners: P < 0.001). Fat tissue was the highest in controls (19.4 ± 7.9 kg) and lowest in distance runners (5.5 ± 1.1 kg) (controls versus distance runners: P < 0.001). Body fat percentage was the highest in controls (23.9 ± 6.9%) and lowest in distance runners (9.2 ± 1.6%) (controls versus distance runners: P < 0.001). The body composition characteristics and career of each group confirmed that both bodybuilders and distance runners were adapted to specific exercise stimuli for a long time.
Dietary intake
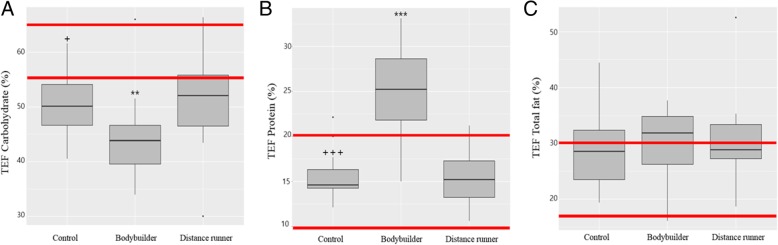
Macronutrient intake of the subjects is shown in Table 2. Athletes consumed more energy than controls; specifically, carbohydrate and lipid intake was significantly higher in athletes than controls. Protein intake was not significantly different between the controls and the distance runners, but bodybuilders consumed significantly more protein than the other two groups. None of the groups ate the recommended daily amount of dietary fiber (> 25 g). Regarding the protein:carbohydrate ratio, the bodybuilders were twice as high as the other groups (P < 0.001), and regarding the dietary fiber:carbohydrate ratio, the controls were 1.4-times higher than the distance runners (P < 0.05). The nutrient uptake ratios showed that the bodybuilders had a high protein diet pattern and the distance runners had a low dietary fiber diet pattern. The energy ratio of macronutrients of the three groups and dietary reference intakes for Koreans are shown in Fig. 1 (carbohydrate: 55–65%; protein: 7–20%; total fat: 15–30%). The energy contribution ratio of carbohydrates of all groups was lower than that of dietary reference intakes for Koreans, and the bodybuilder group was significantly lower than controls (P < 0.05) and distance runners (P < 0.01). The energy contribution ratio of protein and total fat of controls and distance runners fulfilled dietary reference intakes for Koreans, and bodybuilders exceeded it. The percent of energy from protein among the bodybuilders was significantly higher than the other groups (P < 0.001) but the percent of energy from total fat did not differ between groups.
Table 2.
Composition of the macronutrients and dietary fiber intake of controls, bodybuilders, and distance runners obtained from 3-d food diaries
| DRIK | Controls (n = 15) |
Bodybuilders (n = 15) |
Distance runners (n = 15) |
|
|---|---|---|---|---|
| Energy (kcal) | 2600 | 1757+++/2046–1306 | 3362 /5392–2488 | 2787¥¥¥/3756–2371 |
| Carbohydrates (g) | 219+++/ 264–160 | 411 / 547–319 | 369¥¥¥/ 408–325 | |
| Protein (g) | 71+++/ 84–51 | 236***/ 314–175 | 103/ 111–79 | |
| Total fat (g) | 49+++/ 85–39 | 131 / 188–76 | 88¥¥/ 120–65 | |
| Saturated fat (g) | < 20 g | 10+++/ 11–8 | 4** / 7–3 | 14 / 18–9 |
| Monounsaturated fat (g) | 11++/ 13–10 | 5** /12–5 | 16 / 18–9 | |
| Polyunsaturated fat (g) | 9 /11–7 | 5 /12–5 | 9 / 14–5 | |
| Dietary fiber (g) | ≥25 g | 14+/ 19–11 | 19 / 36–14 | 17 / 20–15 |
| Protein: Carbohydrate ratio | 0.3+++ | 0.6+++ | 0.3 | |
| Dietary fiber: Carbohydrate ratio | 0.07 | 0.06 | 0.05¥ |
Data shown as median/IQR (Q3-Q1), controls versus bodybuilders: +p < 0.05, ++p < 0.01 or +++p < 0.001, bodybuilders versus distance runners: **p < 0.01, ***p < 0.001, distance runners versus controls: ¥¥p < 0.01, ¥¥¥p < 0.001. DRIK: Dietary Reference Intakes for Koreans
Fig. 1.
Comparison of the percentage of energy from each macronutrient category. The percentages were calculated by dividing the available energy from the macronutrients by the total energy. a The percentage of the energy obtained from carbohydrates; b The percentage of the energy obtained from protein; c The percentage of the energy obtained from total fat. The red line represents acceptable macronutrient distribution ranges (AMDR). The AMDR for carbohydrates is 55 to 65%, for total fat is 15 to 30%, and for protein is 7 to 20% of the energy intake for Korean adults. Controls versus bodybuilders: +p < 0.05 or +++p < 0.001, bodybuilders versus distance runners: **p < 0.01 or ***p < 0.001
Microbial taxonomy
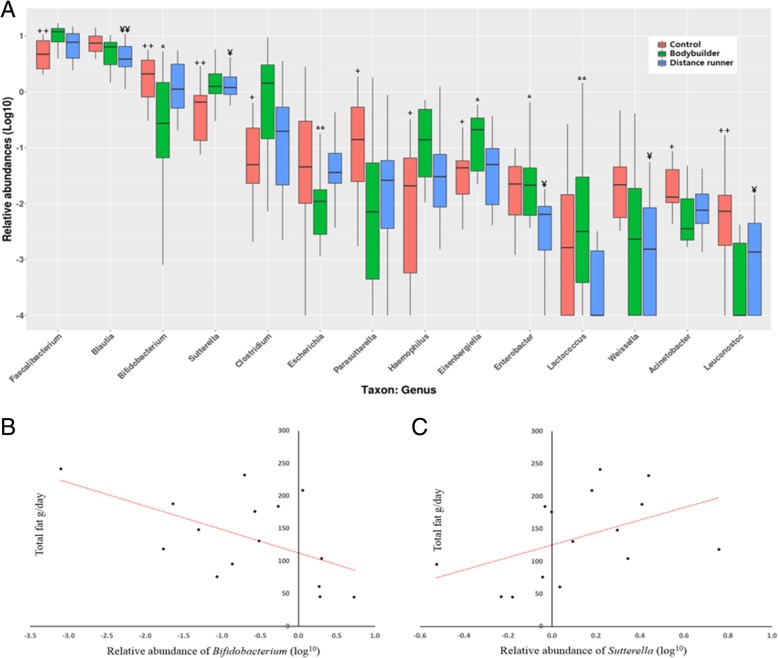
Type of exercise training and athlete diet influenced relative abundance of gut microbiota at the genus and species levels. Faecalibacterium, Sutterella, Clostridium, Haemophilus, and Eisenbergiella were the highest (P < 0.05) in bodybuilders while Bifidobacterium and Parasutterella were the lowest (P < 0.05) in bodybuilders (Fig. 2a). In particular, the high fat intake by bodybuilders was related to the relative abundance of Bifidobacterium and Sutterella. The relative abundance of Bifidobacterium in bodybuilders was negatively correlated with fat intake (r = − 0.52, p = 0.048) (Fig. 2b), while the relative abundance of Sutterella in bodybuilders was positively correlated with fat intake (r = 0.58, p = 0.023) (Fig. 2c). At the species level, intestinal beneficial bacteria, which are widely used as probiotics (Bifidobacterium adolescentis group, Bifidobacterium longum group, Lactobacillus sakei group), and those producing short chain fatty acids (Blautia wexlerae, Eubacterium hallii) were the lowest (P < 0.05) in bodybuilders and the highest (P < 0.05) in controls (Table 3).
Fig. 2.
Certain types of exercise training and athlete diet affected the relative abundance of some microorganisms. a Comparison of gut microbiota relative abundance at the genus level in the three groups. Relative abundance represent log of percentage as whole microbiota. For example, when the relative abundance of a particular genus is 1, it means 10% of whole microbiota, and when it is 0, it means 1% and when it is −1, it means 0.1%. Controls versus bodybuilders: +p < 0.05 or ++p < 0.01, bodybuilders versus distance runners: *p < 0.05 or **p < 0.01, distance runners versus controls: ¥p < 0.05 or ¥¥p < 0.01. b Total fat intake negatively correlated with relative abundance of Bifidobacterium in bodybuilders (Correlation coefficient: −0.52, p-value: 0.048). c Total fat intake positively correlated with relative abundance of Sutterella in bodybuilders (Correlation coefficient: 0.58, p-value: 0.023)
Table 3.
Gut microbiota composition at species level
| Controls (n = 15) |
Bodybuilders (n = 15) |
Distance runners (n = 15) |
|
|---|---|---|---|
| Blautia wexlerae | 4.9 ± 2.6+ | 3.2 ± 1.9* | 2.1 ± 1.1¥¥ |
| Faecalibacterium prausnitzii | 2.3 ± 2.0 | 4.9 ± 3.8 | 2.8 ± 2.4 |
| Eubacterium hallii | 2.0 ± 1.1+ | 1.0 ± 0.7 | 1.2 ± 0.9¥ |
| Bacteroides stercoris | 1.9 ± 2.6++ | 0.4 ± 0.9 | 0.8 ± 1.8 |
| B. adolescentis group | 1.3 ± 1.6 | 0.4 ± 1.1** | 1.1 ± 0.9 |
| Bacteroides caccae | 0.15 ± 0.2 | 0.2 ± 0.2** | 0.9 ± 1.0¥¥ |
| Ruminococcus callidus | 0.3 ± 0.4++ | 0.07 ± 0.2* | 0.4 ± 0.7 |
| B. longum group | 0.5 ± 0.5++ | 0.1 ± 0.1 | 0.3 ± 0.3 |
| Escherichia coli group | 0.4 ± 0.8+ | 0.2 ± 0.8** | 0.3 ± 0.7 |
| Faecalibacterium_uc | 0.1 ± 0.06+++ | 0.2 ± 0.1 | 0.2 ± 0.1 |
| Sutterella_uc | 0.03 ± 0.03++ | 0.07 ± 0.05 | 0.07 ± 0.06¥ |
| Bifidobacterium_uc | 0.08 ± 0.03+ | 0.03 ± 0.04* | 0.07 ± 0.06 |
| Clostridium innocuum group | 0.1 ± 0.3 | 0.007 ± 0.02 | 0.002 ± 0.004¥ |
| Weissella confusa group | 0.07 ± 0.1 | 0.03 ± 0.1 | 0.001 ± 0.002¥¥ |
| Enterobacter cloacae group | 0.03 ± 0.02 | 0.08 ± 0.2** | 0.005 ± 0.005¥¥ |
| Lactobacillus sakei group | 0.05 ± 0.06+ | 0.01 ± 0.03 | 0.03 ± 0.05 |
Data shown as mean ± SD, controls versus bodybuilders: +p < 0.05 or ++p < 0.01 or +++p < 0.001, bodybuilders versus distance runners: *p < 0.05 or **p < 0.01, distance runners versus controls: ¥p < 0.05 or ¥¥p < 0.01 or ¥¥¥p < 0.001
Microbial diversity
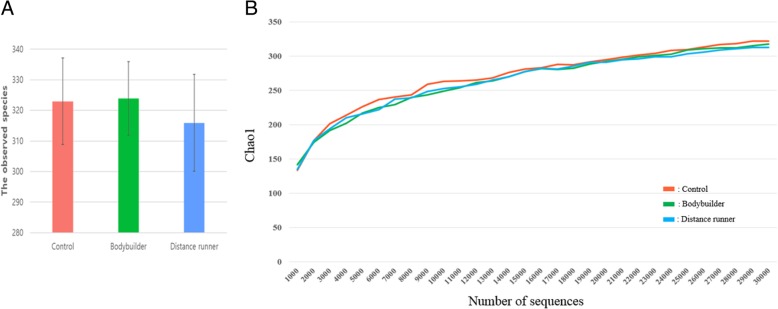
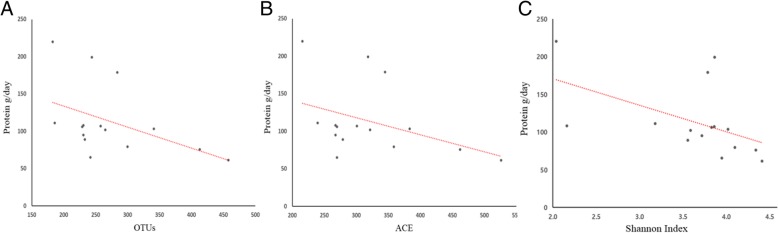
Aerobic or resistance exercise training accompanied by an unbalanced intake of macronutrients and low intake of dietary fiber did not lead to increased diversity of gut microbiota (Fig. 3a and b). Specifically, for distance runners, a negative correlation was found between protein intake and gut microbiota diversity indices (Fig. 4). Daily protein intake showed a negative correlation with operation taxonomic unit (r = − 0.53, p = 0.04), ACE (r = − 0.51, p = 0.05), and Shannon index (r = − 0.64, p = 0.01) in distance runners. Despite differences in exercise type, body composition, and nutrient intake, the gut microbiota beta diversity of healthy men in the control group and the athlete groups did not differ (Fig. 5).
Fig. 3.
There was no difference in the gut microbiota diversity between the controls, bodybuilders, and distance runners. a Comparison of observed species of controls, bodybuilders, and distance runners obtained from 30,000 sequences per sample. b Estimation of the abundance of unique operational taxonomic units (OTUs) using Chao1. Phylogenetic diversity was estimated using the average values for the Chao1 plot of the gut microbiota in the controls, bodybuilders, and distance runners. Data are based on 30,000 sequences per sample from the study subjects
Fig. 4.
Protein intake negatively correlated with alpha diversity in distance runners. a Protein intake negatively correlated with OTUs (Correlation coefficient: − 0.53, p value: 0.04). b Protein intake negatively correlated with ACE (Correlation coefficient: − 0.51, p-value: 0.05). c Protein intake negatively correlated with Shannon index (Correlation coefficient: − 0.63, p-value: 0.012)
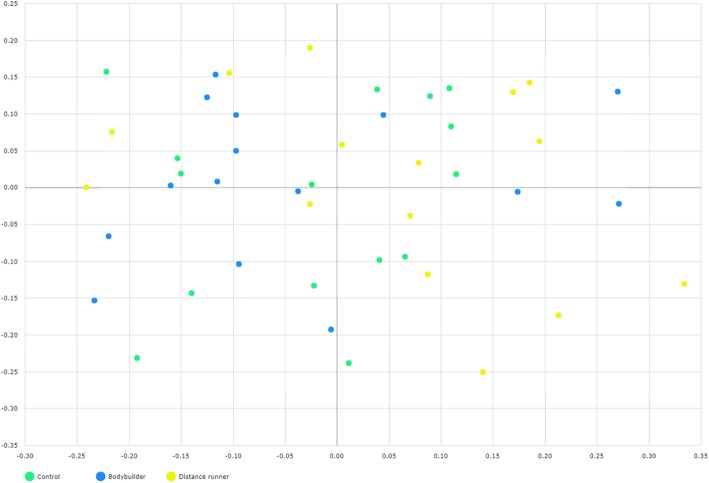
Fig. 5.
The plot was generated using a generalized UniFrac principal coordinate analysis (PCoA) of fecal microbiota from 45 subjects. Generalized UniFrac PCoA analyzed genus rank level and included unclassified OTUs. Subject color coding: green, controls; blue, bodybuilders; and yellow, distance runners
Discussion
There are several reports regarding the effects of various nutrients and diet patterns on human gut microbiota [23, 44, 45] but, recently, physical exercise was disclosed as yet another factor affecting the composition, diversity, and metabolic activity of the gut microbiota [15, 17, 18]. However, the impact of physical exercise associated with diet pattern and type of exercise training on the gut microbiota is not fully understood. Our findings, in contrast to recent studies, indicate that type of exercise training and the diet pattern associated with specific sports did not make a difference in the beta diversity of gut microbiota, but they did affect the relative abundance of certain intestinal microbes. In particular, bodybuilders’ high fat intake made Sutterella more abundant, while significantly reducing the abundance of Bifidobacterium. Cani et al. (2007) reported that a high fat diet caused a decrease in Bifidobacterium [46], and Bifidobacterium has a negative correlation with the concentration of lipopolysaccharide (LPS), an endotoxin, in the blood [46]. A high protein, low carbohydrate (HP-LC) diet in conjunction with a high fat diet causes a decrease in Bifidobacterium [47, 48]. The lack of Bifidobacterium resulting from an HP-LC diet could be due to a shortage of carbohydrate-based substrates, the detrimental effects of protein-fermentative metabolites, or competitive exclusion by protein-fermenting microbes in the gut [49]. Several reports have accentuated the significance of Bifidobacterium in modulating intestinal homeostasis, regulating local and systemic immune responses, and defending against inflammatory diseases and infections [50, 51]. In addition, acetate-producing bacteria (e.g., Blautia wexlerae, Bifidobacterium adolescentis group, and Bifidobacterium longum group) and lactate-producing bacteria (e.g., Lactobacillus sakei group) appeared less in bodybuilders, which may have led to the observed decrease in butyrate-producing Eubacterium hallii, which use acetate and lactate as substrates. Eubacterium hallii is one of the most abundant butyrate-producing bacteria in the human intestine (followed by Faecalibacterium prausnitzii, E. rectale, E. hallii, and Anaerostipes hadrus) [52, 53]; it uses glycerol to produce reuterin, an antimicrobial substance that regulates the homeostasis of intestinal microbial metabolism and inhibits pathogens [54]. As a result, the HP-LC and high fat diet of the bodybuilders is expected to lower the relative abundance of Bifidobacterium and, in particular, to lower the relative abundance of acetate-producing bacteria and lactate-producing bacteria, thus influencing the substrate supply of butyrate-producing bacteria. Sutterella, highly expressed in athletes, is a genus of Gram-negative, anaerobic, nonspore-forming bacteria that is associated with autism, Down’s syndrome, and inflammatory disease [55, 56]. Sutterella is augmented by a high-fat diet as well as by a low-fiber diet [57]. The mechanism of the increase of Sutterella may due to the decrease in mucosal thickness caused by the lack of luminal butyrate, because it can adhere to epithelial cells [58]. Therefore, the increase of Sutterella in athletes may be associated with the lack of butyrate-producing bacteria such as Bifidobacterium adolescentis group, Bifidobacterium longum group, Blautia wexlerae, Lactobacillus sakei group, and Eubacterium hallii.
Athletes who had adapted to certain exercise training practices and diet regimens for long time periods did not have the high gut microbiota diversity found in controls who did not engage in regular exercise training. Specifically, in the case of low carbohydrate and dietary fiber intake of distance runners, gut microbiota diversity tended to decrease as protein intake increased. However, Clarke et al. (2014) reported that as protein intake in rugby athletes increased, the gut microbiota diversity also increased. This inconsistency between our results and those of Clarke et al. seems to be caused by the differences in nutrition status of the athletes [18]. The rugby athletes of Clarke et al.’s study met all of the recommended intake requirements, while the athletes of our study had insufficient carbohydrate and dietary fiber intake. Carbohydrates and dietary fiber are the main nutrients that provide carbon and energy to the intestinal microorganisms. In particular, adequate intake of dietary fiber increases the diversity of gut microbiota [59, 60]. Therefore, inadequate intake of carbohydrates and dietary fiber by athletes seem to counteract the benefits of exercise and a high protein diet that tend to increase gut microbiota diversity. Taken together, our results suggest that high-protein diets may have a negative impact on gut microbiota diversity for athletes in endurance sports who consume low carbohydrates and low dietary fiber, while athletes in resistance sports that carry out the HP-LC and high fat diet demonstrate a decrease in short chain fatty acid-producing commensal bacteria. Additional studies should be conducted to determine the effects of external stimuli on the gut microbiota characteristics, exercise performance, and physical condition in athletes.
Acknowledgements
Not applicable.
Funding
This study was supported by the Ministry of Education of the Republic of Korea and the National 347 Research Foundation of Korea (NRF-2016S1A5A2A01023587).
Availability of data and materials
The data sets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- BMI
Body Mass Index
- DRIK
Dietary Reference Intakes for Koreans
- DXA
Dual-energy X-ray Absorptiometry
- HP-LC
High Protein, Low Carbohydrate
- IPAQ
International Physical Activity Questionnaire
- LPS
lipopolysaccharide
- MET
Metabolic Equivalent of Task
Authors’ contributions
LJ and HP proposed the study for the funding and performed most of the measurements and writing. GC and SK helped to manage all subjects on collecting samples. BK and SL analyzed gut microbiota profiles and offered bioinformatics station. All authors read and approved the final manuscript.
Ethics approval and consent to participate
All subjects provided written informed consent prior to beginning the study. This study was conducted after approval was obtained from the Institutional Review Board of Kyung Hee University.
Consent for publication
We have used our Institutional Consent Form and ready to submit under your request any time.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Lae-Guen Jang, Email: francis@khu.ac.kr.
Geunhoon Choi, Email: gunhoon22@khu.ac.kr.
Sung-Woo Kim, Email: kswrha@khu.ac.kr.
Byung-Yong Kim, Email: bykim@chunlab.com.
Sunghee Lee, Email: slee@ildong.com.
Hyon Park, Email: hpark@khu.ac.kr.
References
- 1.Hawley JA, Hargreaves M, Joyner MJ, Zierath JR. Integrative biology of exercise. Cell. 2014;159(4):738–749. doi: 10.1016/j.cell.2014.10.029. [DOI] [PubMed] [Google Scholar]
- 2.Wolin KY, Yan Y, Colditz GA, Lee I. Physical activity and colon cancer prevention: a meta-analysis. Br J Cancer. 2009;100(4):611. doi: 10.1038/sj.bjc.6604917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Mehnert A, Veers S, Howaldt D, Braumann KM, Koch U, Schulz KH. Effects of a physical exercise rehabilitation group program on anxiety, depression, body image, and health-related quality of life among breast cancer patients. Onkologie. 2011;34(5):248–253. doi: 10.1159/000327813. [DOI] [PubMed] [Google Scholar]
- 4.Zanuso S, Jimenez A, Pugliese G, Corigliano G, Balducci S. Exercise for the management of type 2 diabetes: a review of the evidence. Acta Diabetol. 2010;47(1):15–22. doi: 10.1007/s00592-009-0126-3. [DOI] [PubMed] [Google Scholar]
- 5.Cartee GD, Hepple RT, Bamman MM, Zierath JR. Exercise promotes healthy aging of skeletal muscle. Cell Metab. 2016;23(6):1034–1047. doi: 10.1016/j.cmet.2016.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li J, Siegrist J. Physical activity and risk of cardiovascular disease—a meta-analysis of prospective cohort studies. Int J Environ Res Public Health. 2012;9(2):391–407. doi: 10.3390/ijerph9020391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Carek PJ, Laibstain SE, Carek SM. Exercise for the treatment of depression and anxiety. The International Journal of Psychiatry in Medicine. 2011;41(1):15–28. doi: 10.2190/PM.41.1.c. [DOI] [PubMed] [Google Scholar]
- 8.Cerdá B, Pérez M, Pérez-Santiago JD, Tornero-Aguilera JF, González-Soltero R, Larrosa M. Gut microbiota modification: another piece in the puzzle of the benefits of physical exercise in health? Front Physiol. 2016;7:51. doi: 10.3389/fphys.2016.00051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Codella R, Luzi L, Terruzzi I. Exercise has the guts: how physical activity may positively modulate gut microbiota in chronic and immune-based diseases. Dig Liver Dis. 2018;50(4):331–341. doi: 10.1016/j.dld.2017.11.016. [DOI] [PubMed] [Google Scholar]
- 10.Kang SS, Jeraldo PR, Kurti A, Miller MEB, Cook MD, Whitlock K, et al. Diet and exercise orthogonally alter the gut microbiome and reveal independent associations with anxiety and cognition. Mol Neurodegener. 2014;9(1):36. doi: 10.1186/1750-1326-9-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Evans CC, LePard KJ, Kwak JW, Stancukas MC, Laskowski S, Dougherty J, et al. Exercise prevents weight gain and alters the gut microbiota in a mouse model of high fat diet-induced obesity. PLoS One. 2014;9(3):e92193. doi: 10.1371/journal.pone.0092193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Choi JJ, Eum SY, Rampersaud E, Daunert S, Abreu MT, Toborek M. Exercise attenuates PCB-induced changes in the mouse gut microbiome. Environ Health Perspect. 2013;121(6):725–730. doi: 10.1289/ehp.1306534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Feng X, Uchida Y, Koch L, Britton S, Hu J, Lutrin D, et al. Exercise Prevents enhanced Postoperative neuroinflammation and cognitive Decline and rectifies the gut Microbiome in a rat Model of Metabolic syndrome. Frontiers in immunology. 2017;8:1768. [DOI] [PMC free article] [PubMed]
- 14.Lambert JE, Myslicki JP, Bomhof MR, Belke DD, Shearer J, Reimer RA. Exercise training modifies gut microbiota in normal and diabetic mice. Appl Physiol Nutr Metab. 2015;40(7):749–752. doi: 10.1139/apnm-2014-0452. [DOI] [PubMed] [Google Scholar]
- 15.Allen J, Mailing L, Cohrs J, Salmonson C, Fryer JD, Nehra V, et al. Exercise training-induced modification of the gut microbiota persists after microbiota colonization and attenuates the response to chemically-induced colitis in gnotobiotic mice. Gut Microbes. 2018;9(2):115–130. doi: 10.1080/19490976.2017.1372077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Estaki M, Pither J, Baumeister P, Little JP, Gill SK, Ghosh S, et al. Cardiorespiratory fitness as a predictor of intestinal microbial diversity and distinct metagenomic functions. Microbiome. 2016;4(1):42. doi: 10.1186/s40168-016-0189-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Barton W, Penney NC, Cronin O, Garcia-Perez I, Molloy MG, Holmes E, et al. The microbiome of professional athletes differs from that of more sedentary subjects in composition and particularly at the functional metabolic level. Gut. 2018;67(4):625–633. doi: 10.1136/gutjnl-2016-313627. [DOI] [PubMed] [Google Scholar]
- 18.Clarke SF, Murphy EF, O'Sullivan O, Lucey AJ, Humphreys M, Hogan A, et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut. 2014;63(12):1913–1920. doi: 10.1136/gutjnl-2013-306541. [DOI] [PubMed] [Google Scholar]
- 19.Clark A, Mach N. Exercise-induced stress behavior, gut-microbiota-brain axis and diet: a systematic review for athletes. Journal of the International Society of Sports Nutrition. 2016;13(1):43. doi: 10.1186/s12970-016-0155-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Rodriguez NR, DiMarco NM, Langley S. American dietetic association, dietitians of Canada, American College of Sports Medicine: nutrition and athletic performance. Position of the American dietetic association, dietitians of Canada, and the American College of Sports Medicine: nutrition and athletic performance. J Am Diet Assoc. 2009;109(3):509–527. doi: 10.1016/j.jada.2009.01.005. [DOI] [PubMed] [Google Scholar]
- 21.David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature. 2014;505(7484):559. doi: 10.1038/nature12820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hughes R, Magee E, Bingham S. Protein degradation in the large intestine: relevance to colorectal cancer. Curr Issues Intestinal Microbiol. 2000;1(2):51–58. [PubMed] [Google Scholar]
- 23.Macfarlane GT, Macfarlane S. Bacteria, colonic fermentation, and gastrointestinal health. J AOAC Int. 2012;95(1):50–60. doi: 10.5740/jaoacint.sge_macfarlane. [DOI] [PubMed] [Google Scholar]
- 24.Toden S, Bird AR, Topping DL, Conlon MA. Resistant starch attenuates colonic DNA damage induced by higher dietary protein in rats. Nutr Cancer. 2005;51(1):45–51. doi: 10.1207/s15327914nc5101_7. [DOI] [PubMed] [Google Scholar]
- 25.Toden S, Bird AR, Topping DL, Conlon MA. Differential effects of dietary whey, casein and soya on colonic DNA damage and large bowel SCFA in rats fed diets low and high in resistant starch. Br J Nutr. 2007;97(3):535–543. doi: 10.1017/S0007114507336817. [DOI] [PubMed] [Google Scholar]
- 26.Russell WR, Gratz SW, Duncan SH, Holtrop G, Ince J, Scobbie L, et al. High-protein, reduced-carbohydrate weight-loss diets promote metabolite profiles likely to be detrimental to colonic health. Am J Clin Nutr. 2011;93(5):1062–1072. doi: 10.3945/ajcn.110.002188. [DOI] [PubMed] [Google Scholar]
- 27.Shaughnessy DT, Gangarosa LM, Schliebe B, Umbach DM, Xu Z, MacIntosh B, et al. Inhibition of fried meat-induced colorectal DNA damage and altered systemic genotoxicity in humans by crucifera, chlorophyllin, and yogurt. PLoS One. 2011;6(4):e18707. doi: 10.1371/journal.pone.0018707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gutekunst K, Krüger K, August C, Diener M, Mooren F. Acute exercises induce disorders of the gastrointestinal integrity in a murine model. Eur J Appl Physiol. 2014;114(3):609–617. doi: 10.1007/s00421-013-2791-8. [DOI] [PubMed] [Google Scholar]
- 29.Wang F, Li Q, Wang C, Tang C, Li J. Dynamic alteration of the colonic microbiota in intestinal ischemia-reperfusion injury. PLoS One. 2012;7(7):e42027. doi: 10.1371/journal.pone.0042027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Craig CL, Marshall AL, Sjorstrom M, Bauman AE, Booth ML, Ainsworth BE, et al. International physical activity questionnaire: 12-country reliability and validity. Med Sci Sports Exerc. 2003;35(8):1381–1395. doi: 10.1249/01.MSS.0000078924.61453.FB. [DOI] [PubMed] [Google Scholar]
- 31.The Korean Nutrition Society [Internet]. Sejong : The Korean Nutrition Society; 2016 [updated 2016 January 18; cited 2018 August 14]. Available from: http://www.kns.or.kr/FileRoom/FileRoom.asp?BoardID=Kdr.
- 32.Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114–2120. doi: 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics. 2012;13(1):31. doi: 10.1186/1471-2105-13-31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75(23):7537–7541. doi: 10.1128/AEM.01541-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 36.Myers EW, Miller W. Optimal alignments in linear space. Bioinformatics. 1988;4(1):11–17. doi: 10.1093/bioinformatics/4.1.11. [DOI] [PubMed] [Google Scholar]
- 37.Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27(16):2194–2200. doi: 10.1093/bioinformatics/btr381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Fu L, Niu B, Zhu Z, Wu S, Li W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics. 2012;28(23):3150–3152. doi: 10.1093/bioinformatics/bts565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26(19):2460–2461. doi: 10.1093/bioinformatics/btq461. [DOI] [PubMed] [Google Scholar]
- 40.Blaxter M, Mann J, Chapman T, Thomas F, Whitton C, Floyd R, et al. Defining operational taxonomic units using DNA barcode data. Philos Trans R Soc Lond Ser B Biol Sci. 2005;360(1462):1935–1943. doi: 10.1098/rstb.2005.1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chao A. Nonparametric estimation of the number of classes in a population. Scand J Stat. 1984:265–70.
- 42.Chao A, Yang MC. Stopping rules and estimation for recapture debugging with unequal failure rates. Biometrika. 1993;80(1):193–201. [Google Scholar]
- 43.Shannon CE. A mathematical theory of communication. Bell system technical journal. 1948;27(3):379–423. [Google Scholar]
- 44.Wu GD, Chen J, Hoffmann C, Bittinger K, Chen Y, Keilbaugh SA, et al. Linking long-term dietary patterns with gut microbial enterotypes. Science. 2011;334(6052):105–108. doi: 10.1126/science.1208344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.DeFilippo C, Cavalieri D, DiPaola M, Ramazzotti M, Poullet JB, Massart S, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. PNAS. 2010;107(33):14691–6. [DOI] [PMC free article] [PubMed]
- 46.Cani PD, Neyrinck AM, Fava F, Knauf C, Burcelin RG, Tuohy KM, et al. Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia. 2007;50(11):2374–2383. doi: 10.1007/s00125-007-0791-0. [DOI] [PubMed] [Google Scholar]
- 47.Moreira APB, Texeira TFS, Ferreira AB. Peluzio, Maria do Carmo Gouveia, Alfenas, Rita de Cássia Gonçalves. Influence of a high-fat diet on gut microbiota, intestinal permeability and metabolic endotoxaemia. Br J Nutr. 2012;108(5):801–809. doi: 10.1017/S0007114512001213. [DOI] [PubMed] [Google Scholar]
- 48.Lubbs D, Vester B, Fastinger N, Swanson K. Dietary protein concentration affects intestinal microbiota of adult cats: a study using DGGE and qPCR to evaluate differences in microbial populations in the feline gastrointestinal tract. J Anim Physiol Anim Nutr. 2009;93(1):113–121. doi: 10.1111/j.1439-0396.2007.00788.x. [DOI] [PubMed] [Google Scholar]
- 49.Hooda S, Boler BMV, Kerr KR, Dowd SE, Swanson KS. The gut microbiome of kittens is affected by dietary protein: carbohydrate ratio and associated with blood metabolite and hormone concentrations. Br J Nutr. 2013;109(9):1637–1646. doi: 10.1017/S0007114512003479. [DOI] [PubMed] [Google Scholar]
- 50.Lomax A, Calder P. Probiotics, immune function, infection and inflammation: a review of the evidence from studies conducted in humans. Curr Pharm Des. 2009;15(13):1428–1518. doi: 10.2174/138161209788168155. [DOI] [PubMed] [Google Scholar]
- 51.Salminen S, Nybom S, Meriluoto J, Collado MC, Vesterlund S, El-Nezami H. Interaction of probiotics and pathogens—benefits to human health? Curr Opin Biotechnol. 2010;21(2):157–167. doi: 10.1016/j.copbio.2010.03.016. [DOI] [PubMed] [Google Scholar]
- 52.Tap J, Mondot S, Levenez F, Pelletier E, Caron C, Furet J, et al. Towards the human intestinal microbiota phylogenetic core. Environ Microbiol. 2009;11(10):2574–2584. doi: 10.1111/j.1462-2920.2009.01982.x. [DOI] [PubMed] [Google Scholar]
- 53.Walker AW, Ince J, Duncan SH, Webster LM, Holtrop G, Ze X, et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. The ISME journal. 2011;5(2):220. doi: 10.1038/ismej.2010.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Engels C, Ruscheweyh H, Beerenwinkel N, Lacroix C, Schwab C. The common gut microbe Eubacterium hallii also contributes to intestinal propionate formation. Front Microbiol. 2016;7:713. doi: 10.3389/fmicb.2016.00713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Williams BL, Hornig M, Parekh T, Lipkin WI. Application of novel PCR-based methods for detection, quantitation, and phylogenetic characterization of Sutterella species in intestinal biopsy samples from children with autism and gastrointestinal disturbances. MBio. 2012;3(1). 10.1128/mBio.00261-11 Print 2012. [DOI] [PMC free article] [PubMed]
- 56.Wang L, Christophersen CT, Sorich MJ, Gerber JP, Angley MT, Conlon MA. Increased abundance of Sutterella spp. and Ruminococcus torques in feces of children with autism spectrum disorder. Molecular autism. 2013;4(1):42. doi: 10.1186/2040-2392-4-42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hwang N, Eom T, Gupta SK, Jeong S, Jeong D, Kim YS, et al. Genes and gut Bacteria involved in luminal butyrate reduction caused by diet and Loperamide. Genes. 2017;8(12):350. doi: 10.3390/genes8120350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zeng B, Han S, Wang P, Wen B, Jian W, Guo W, et al. The bacterial communities associated with fecal types and body weight of rex rabbits. Sci Rep. 2015;5:9342. doi: 10.1038/srep09342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Martínez I, Lattimer JM, Hubach KL, Case JA, Yang J, Weber CG, et al. Gut microbiome composition is linked to whole grain-induced immunological improvements. The ISME journal. 2013;7(2):269. doi: 10.1038/ismej.2012.104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tap J, Furet J, Bensaada M, Philippe C, Roth H, Rabot S, et al. Gut microbiota richness promotes its stability upon increased dietary fiber intake in healthy adults. Environ Microbiol. 2015;17(12):4954–4964. doi: 10.1111/1462-2920.13006. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data sets used and/or analyzed during the current study are available from the corresponding author on reasonable request.