Abstract
Brachydactyly type E, which can be an isolated finding or part of a syndrome in combination with other clinical anomalies, involves metacarpals and metatarsals with or without short phalanges. Herein we report two unrelated Turkish females who presented with brachydactyly type E and vitamin D deficiency in the absence of marked alterations in serum calcium, phosphate, and parathyroid hormone. After excluding disease-causing variants in two candidate genes, PTHLH and PDE4D, we identified different pathogenic variants in TRPS1, the gene mutated in patients with tricho-rhino-phalangeal syndrome (TRPS). In one of the patients, who displayed severe brachydactyly and short stature, we identified a novel heterozygous missense pathogenic variant in exon 6 (c.2783A>G, p.Tyr928Cys), located within the GATA DNA-binding domain. The second patient, who had relatively milder brachydactyly and was of normal height, carried a heterozygous nonsense pathogenic variant in exon 4 (c. 1870C>T, p.Arg624Ter), which has been previously described. Both pathogenic variants segregated in affected family members. The patients additionally showed sparse hair and a bulbous nose, consistent with the clinical features of TRPS. Our findings, in addition to identifying the genetic cause of brachydactyly in two unrelated kindreds, emphasize the role of pathogenic TRPS1 variants in the development of brachydactyly type E and highlight the GATA DNA-binding region of TRPS1 protein with respect to phenotype-genotype correlation.
Keywords: Brachydactyly type E, Tricho-rhino-phalangeal syndrome (TRPS), TRPS1, GATA DNA-binding domain, skeletal dysplasia
Introduction
Congenital malformations of the long bones of the hands or feet are seen in many skeletal dysplasias, including brachydactyly, which refers to disproportionate shortening of digits [1]. Brachydactyly type E (BDE) involves the shortening of one or more metacarpals with normal or short phalanges, combined occasionally with short metatarsals [MIM# 113300]. Some individuals with BDE may also be of moderately short stature. BDE is inherited as an autosomal dominant trait with variable expressivity and results from pathogenic variants of genes important during bone development [2]. HOXD13 pathogenic variants are responsible for isolated BDE, but most patients display features overlapping with brachydactyly type D, characterized by shortness of the distal phalanx of the thumb; some patients can also display syndactyly or synpolydactylyl [3, 4]. Pathogenic variants in PTHLH are associated with BDE with short stature [5, 6], and pathogenic variants in PDE3A are responsible for BDE with hypertension (Bilginturan Disease) [7]. Deletions of chromosome 2q37 involving HDAC4 lead to BDE with intellectual disability [8, 9]. In some cases, the phenotype is additionally complicated by multihormone resistance, such as in Albright’s hereditary osteodystrophy and pseudohypoparathyroidism, which are caused by pathogenic variants in Gsα-coding exons of the GNAS complex locus [10, 11]. Acrodysostosis, a disorder of severe BDE, short stature, and facial dysmorphism, result from pathogenic variants in PRKAR1A [12], particularly when associated with hormone resistance, or PDE4D [13, 14]. Pathogenic variants in TRPS1 also lead to BDE, as seen in patients with Tricho-rhino-phalangeal syndrome (TRPS), a rare autosomal dominant disorder of skeletal abnormalities primarily affecting distal bones, abnormal facial features with a distinctive nose shape, and sparse scalp hair [15, 16].
In this report, we present clinical and genetic characterization of two unrelated patients and their family members with BDE in whom we discovered one novel and one previously described pathogenic variants in TRPS1.
Patients and Methods
Patients and healthy family members
We investigated two unrelated females presented at different times with varying severity of brachydactyly. The affected and unaffected family members were also studied via molecular genetic analyses. DNA samples, as well as clinical and biochemical information, were available for family A – two affected (I-2, III-1) and four unaffected individuals (I-1, II-1, II-2, II-3; Fig. 1A), and for family B–three affected family members (I-1, II-1, II-2; Fig.1B). All biochemical measurements related to clinical management were performed at Ankara Training and Research Hospital, Ankara, Turkey. The study involving molecular genetic analyses was approved by the Institutional Review Board of the Massachusetts General Hospital. Informed consent was obtained from each investigated individual.
Figure 1. Pedigrees of family A and B.
Affected individuals are indicated by filled symbols and probands by arrows. Individuals whose DNA samples were available for the analysis are shown by Arabic numerals. In Family A, I-2 and III-1 were found to have a novel pathogenic missense variant in TRPS1 exon 6; this variant was not present in individuals II-2 and II-3. In Family B, I-1, II-1, and II-2 were found to have a nonsense variant in TRPS1 exon 4.
DNA sequence analysis
Blood samples of two probands and their family members were collected, and genomic DNA was extracted from peripheral blood leukocytes using standard methods. The DNA sequence analysis was performed by PCR amplification of exons and flanking intronic regions, followed by direct Sanger sequencing of the purified PCR products. PCR primer sequences are listed in Table 1. Sequencing was performed at the Center for Computational and Integrative Biology DNA Core Facility, Massachusetts General Hospital. Sequencing results were analyzed by using Sequencher 5.4.6 (Gene Codes Corporation, USA). Multiple sequence alignment was obtained from ENSEMBL Compara (http://useast.ensembl.org/info/genome/compara/index.html). SWISS-MODEL [17] was utilized for the peptide modeling, using TRPS1 residues 887–945 (NP_054831) and the human transcription factor GATA-4 (SMTL ID: 2m9w.1) as template.
Table 1.
Primers used in the amplification of PTHLH, PDE4D, and TRPS1
| Gene | Exon | Forward (5′−3′) | Reverse (5′−3′) |
|---|---|---|---|
| PTHLH | |||
| 3 | gctgagaggctccagagaaat | ggttagggagagggaccatc | |
| 4 | tacccatccagcttgtgtctta | ctgtcctccagcaccatagaga | |
| PDE4D | |||
| 4 | acgctaaatgctttttgaaaatatg | gtgcactcaggttaaatctaccatt | |
| 5 | tttaatattgaacgacactgagcac | gcttgaatgaaccaagaatttgtat | |
| 6 | tctcttaacaaaccattttcaagtgt | ggctcaatcaagttgagaaaactta | |
| 8 | gctagcggacccgtgaag | ggggctatgggtaccagtg | |
| 9 | ggtgttttcccctcattgaa | ctcagaggcggatagtgtcg | |
| 12 | tcactagacatgtgagagtgcagtt | cattgaaaacccttcagataaaagt | |
| 13 | agactctcccttgacacattgtaac | aactttagggccaatacaactttct | |
| 15 | gcccccaaggagtagtaagg | gctgcttaagtggctggttc | |
| 16 | tatttttaagtgttgcagtcatcca | aagtaaccaaatgctaaagcggta | |
| 17 | ccatgatctaattaagcttttccat | cagatgacagtgaggtgtgacc | |
| TRPS1 | |||
| 2 | ctggggagggagaggatct | gcacgatgttttactgtgtgc | |
| 3 | tttttcctagaggaagcatttga | tggagtgtgtccatctcttctg | |
| 4 | tcacctgttactttggcctaca | ttccccaagtcattggaatc | |
| 5 | tcagaacgctgtctttgcag | tgcacagcacacacaaacac | |
| 6 | ctcctgggttgatttggtct | agccagggaatgggacttat | |
| 7 | gcagaaagcctatccctgct | tgttggataaggcaggctct | |
Clinical and laboratory findings
Patient 1 (Family A, III-1)
The first patient, a 17-year old female (Fig. 2A, B), presented to our endocrinology outpatient clinic with short stature an asymmetric brachydactyly. Her height was 152 cm (−2SD) [18], and her weight 55 kg (BMI: 23.8). She was single with no children. All her metacarpals were short, with the exception of the third one in her left hand. Additionally, proximal phalanges of both index fingers were short, and X-rays demonstrated cone-shaped epiphyses in the middle phalanges (Fig. 2A, B). All metatarsals of both feet were also short, except for the second metatarsals (Fig. 2C, D).
Figure 2. Brachydactyly in Family A. A,B (Individual III-1).
Asymmetric short metacarpals of the proband, with cone-shaped epiphyses at middle phalanges (arrows). C,D (Individual III-1). Short metatarsals in both feet, except for the 2nd metatarsals. E (Individual I-2). Symmetric short metacarpals of the maternal grandmother, with residual angulated deformity of proximal middle phalanges (arrows) related to fusion of prior cone-shaped epiphyses.
She was born small for gestational age (1700g) at term to unrelated parents and stayed in the Neonatal Unit for a week. According to her father, she had a normal development and her brachydactyly was noticed at adolescence. She also had normal puberty, and her periods were regular. Laboratory findings at the initial presentation showed normal levels of corrected calcium (9.55mg/dl; normal: 8.8–10.6), phosphate (3.8mg/dl; normal: 2.5–4.5) and PTH (68.6pg/ml; normal: 14–72). TSH and fT4 were within normal limits, with negative TPO antibodies and normal liver and kidney functions. The 25(OH)vitamin D level was markedly low at 8.2 ng/ml (normal: 20–80, measured in winter), while 1,25(OH)2D and alkaline phosphatase measured within normal limits at 36pg/ml (normal: 16–65) and 78U/L (normal: 30–120), respectively. She had no evidence of pseudofractures or osteoporosis by X-rays and a dual-energy X-ray absorptiometry (DXA) scan. Regarding the etiology of vitamin D deficiency, all possible causes were ruled out, including coeliac disease, prolactinemia, and hepatic or renal diseases. She did not have history of medications that may affect vitamin D metabolism. However, she was wearing a hijab for several years and living indoors after graduation from high school, suggesting limited sunlight exposure as a cause of vitamin D deficiency in this patient. Vitamin D treatment according to the Endocrine Society guidelines [19] was commenced: 50,000IU vitamin D3 once a week for total of 8 weeks to achieve 25(OH)vitamin D levels greater than 30ng/ml, followed by maintenance therapy of 600IU/day.
The proband’s mother died a year after her delivery due to leukemia; unfortunately, only few details are known about her medical history, and her DNA was not available (Fig. 1A). However, she reportedly had short stature and abnormal-looking hands and feet. The proband’s maternal grandmother was alive and 70 years old at the time of our evaluations. X-rays showed symmetrically short metacarpals and short proximal phalanx of index finger, with cone-shaped epiphyses at the proximal ends of middle phalanges (Fig. 2E). Her height was the same as her granddaughter, and her weight was 72 kg (BMI: 31.2). She reported that she had always been “tiny” in comparison with her siblings and peers, and was told that she took after her grandmother’s looks.
The maternal grandfather of the proband was unaffected and unrelated to his wife (Fig. 1A). The proband’s maternal three uncles also lacked any skeletal abnormalities. One of her maternal aunts was reportedly short and had a clinical phenotype similar to that of the proband; however, her DNA sample could not be obtained for our studies.
Patient 2 (Family B, II-1)
The second patient, a 40-year old female, had symmetric shortening of metacarpal bones, which was noticed by a physician to whom she went due to pain in her hand joints and back pain. Hand X-rays showed bilateral short fifth metacarpals and cone-shaped epiphyses in middle phalanges of all her fingers except for the thumb (Fig. 3A). She stood 156-cm tall and weighed 53.5 kg (BMI: 22). She was reportedly not short compared to her siblings. She was married and had three children at ages 10, 6, and 4 years.
Figure 3. Brachydactyly in Family B. A (Individual II-1).
Short fifth metacarpals and cone-shaped epiphyses at proximal middle phalanges of the proband (arrows). B (Individual II-2). Short third, fourth, and fifth metacarpals of the proband’s sister, with cone-shaped epiphyses at proximal phalanges (arrows).
The patient also suffered from pain in her back and thighs with generalized muscle pain, which disturbed her sleep at night and could be relieved with the use of analgesics. Her laboratory exam showed serum corrected calcium level of 8.74mg/dl (normal: 8.8–10.6), phosphate level of 4mg/dl (normal: 2.5–4.5), and PTH level of 74pg/ml (normal: 14–72). The level of 25(OH)vitamin D was 19.3 ng/ml (normal: 20–80, measured in summer), 1,25(OH)2 vitamin D was 26pg/ml (normal: 16–65), and alkaline phosphatase was 94U/L (normal: 30–120). A DXA scan revealed decreased bone mineral density consistent with premenopausal osteoporosis (Lumbar spine, Z-score = −2.5 and T-score = −2.7; femoral neck, Z-score: −0.6 and T-score = −0.8). All possible other causes for vitamin D deficiency were ruled out as in the previous patient. The patient did not have any underlying pathology or medication history that could explain the low vitamin D level, except for spending most of her time indoors and wearing a hijab nearly all the time. Treatment for osteomalacia was initiated with 1000mg/d calcium supplements and vitamin D in compliance with the Endocrine Society guidelines [18].
The proband’s parents were unrelated. She had four sisters three of whom had similar skeletal phenotypes consistent with BDE. Her two brothers were reportedly unaffected. X-rays of one of her sisters (II-2) showed short third, fourth, and fifth metacarpals and severe cone-shaped epiphyses of middle phalanges (Fig. 3B). Her biochemical parameters were normal with respect to calcium, phosphate, and PTH. She had the same height as the proband. Additional family members who displayed BDE included the proband’s 68-year old mother and the proband’s two sons (Fig. 1B).
Genetic analysis of the candidate genes
Since the patients and their affected family members did not have evidence of hormone resistance, hypertension, or intellectual disability, we considered PTHLH, PDE4D, and TRPS1 as primary candidate genes. First, we screened exons 3 and 4 of PTHLH (NM_198965), because most disease-causing alterations have been identified in these exons, which comprise all of the coding sequence of the gene with the exception of the last eight nucleotides. We found no potentially disease-causing variants, but Patient 1 had a heterozygous single nucleotide polymorphism (SNP) in intron 2 (rs6242, Supplementary Table 1).
Next, we sequenced PDE4D, as described [20]. We found no variants that were potentially disease-causing in the exons and flanking intronic regions (NM_001165899). However, both patients shared the same previously identified SNPs in introns 7, 8, and 15, i.e. rs2279737, rs7724713, and rs10051847, respectively. Additionally, patient 1 had a heterozygous synonymous SNP (rs7736186) in exon 17 (Supplementary Table 1).
Finally, we examined all the coding exons and flanking intronic regions of TRPS1 (NM_014112.4). In patient 1 we found a novel heterozygous missense variant in exon 6 c.2783A>G (p.Thyr928Cys) (Fig. 4A, Table 2). The substitution was located in the region giving rise to the putative GATA DNA-binding zinc-finger domain, which is indispensable for the transcription repression activity of TRPS1 [21]. This pathogenic variant was not present in large population databases, including Genome Aggregation Database (gnomAD, http://gnomad.broadinstitute.org). In silico tools, including Mutation Taster, SIFT, and Polyphen-2, predicted this missense variant to have an adverse effect on protein function (Table 2). DNA sequence analysis of family members revealed that the same single nucleotide change was also present in the proband’s affected maternal grandmother (I-2), but not in the unaffected maternal grandfather or the two unaffected uncles of the proband (I-1, II-2 and II-3). Thus, according to the current guidelines of the American College of Medical Genetics and Genomics [22], the p.Thyr928Cys pathogenic variant can be classified as “likely pathogenic”.
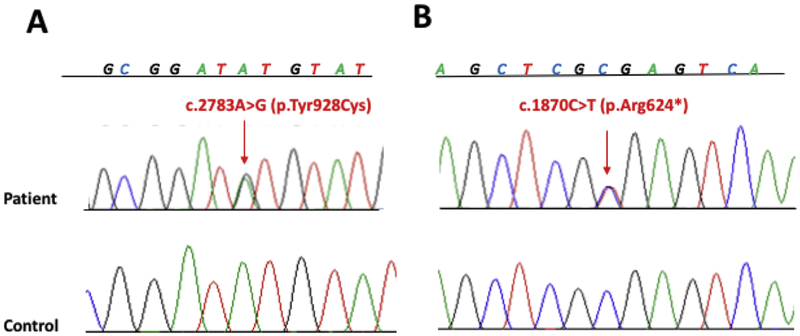
Figure 4. Sequence traces showing the disease-causing nucleotide changes in patients compared to controls.
A. Missense variant in exon 6, p.Tyr928Cys in patient 1. B. Nonsense variant in exon 4, p.Arg624* in patient 2.
Table 2.
The pathogenic TRPS1 variants identified in affected individuals of Family A and B.
| Family A | Family B | |
|---|---|---|
| Variant | c.2783A>G, p.Tyr928Cys | c.1870C>T, p.Arg624* |
| Genomic position (hg19) | chr8:116,430,599 | Chr8: 116616326 |
| Exon (according to NM_014112) | 6 | 4 |
| Protein domain | Zinc finger GATA-DNA binding | Not applicable |
| dbSNP | - | rs121908431 |
| MAF (gnomAD) | - | 0.000008027 |
| Previous report | No | Momeni P et al.2000, Maas SM et al.2015, 2017 |
| MutationTaster | Disease causing | Disease causing |
| SIFT | Damaging | Not available |
| PolyPhen-2 | Probably damaging | Not available |
MAF, minor allele frequency as reported in Genome Aggregation Database (gnomAD; http://gnomad.broadinstitute.org/)
MutationTaster: http://www.mutationtaster.org/
PolyPhen-2 http://genetics.bwh.harvard.edu/pph2/
In patient 2 we identified a heterozygous nonsense pathogenic variant in exon 4, c.1870C>T (p.Arg624*) (Fig. 4B, Table 2). We subsequently performed sequence analysis of genomic DNA from the proband’s affected mother (I-1) and one of her affected sisters (II-2), thus confirming the presence of the same nucleotide substitution in those individuals. The p.Arg624* variant has been previously described in multiple patients with TRPS [23, 24]. This variant is in ClinVar and considered to be pathogenic in multiple entries (https://www.ncbi.nlm.nih.gov/clinvar/variation/5570). The TRPS1 transcript (NM_014112.4) has seven exons. As the p.Arg624* variant is located in exon 4, it is predicted to lead to nonsense-mediated decay of the transcript and, thereby, loss of protein.
Both patient 1 and patient 2, as well as their affected family members, had additional polymorphisms located in TRPS1 exon 2 corresponding to the 5’-untranslated regions (Supplementary Table 1).
Discussion
In this study, we report two families with severe BDE, in whom we identified heterozygous pathogenic variants in the TRPS1 gene. These pathogenic variants likely explain the brachydactyly phenotype observed in the probands and their affected family members. The skeletal abnormalities associated with pathogenic TRPS1 variants are quite variable, and the most typical radiographic findings are cone-shaped epiphyses, which predominantly involve the middle phalanges, but can also be found at other tubular bones [25–27]. The X-rays of our patients clearly demonstrated cone-shaped epiphyses, particularly in the middle phalanges. In addition, consistent with the variable expressivity, the involvement of the metacarpal bones and the severity of shortening varied significantly, even among affected individuals of the same family.
Patients with TRPS1 pathogenic variants have typical facial features including a pear-shaped nose, underdeveloped alae nasi, and long flat philtrum [15, 24, 27]. These patients also have sparse scalp hair and thin lateral eye brows. Our probands were not originally recognized to have those features. On further evaluation, however, each patient, as well as her affected family members, appeared to have a bulbous nose with a broad ridge and tip, including underdeveloped alae nasi and broad septum (Fig. 5). These features were more prominent in patient 1, who also showed a long flat philtrum, and thin lateral eyebrows (Fig. 5A, B). In addition, both our patients had thin sparse and slow-growing hair, more prominently at temporal regions of the head, consistent with ectodermal dysplasia. Thus, combined with the severe BDE, the phenotypes of our patients match TRPS.
Figure 5. Facial features of the probands with TRPS1-associated BDE.
Photographs of individuals III-1 (A,B) from Family A (Patient 1) and II-1 (C) from Family B (Patient 2). Each patient had a bulbous nose, long flat philtrum, and thin upper lip, although the latter two features were more pronounced in Patient 1.
Vitamin D deficiency is the most common cause of osteomalacia, which is usually characterized by bone pain and reduced bone mineral density. The low vitamin D levels could be with or without hypocalcemia and hyperphosphatemia. Patient 1 had strikingly low levels of vitamin D, but her serum calcium, phosphate, and PTH levels were within the normal range. She also lacked pseudofractures or reduced bone mineral density. Patient 2 had modestly low vitamin D levels but also showed a mild decrease of serum calcium coupled with a mild increase of serum PTH. These biochemical changes were associated with symptoms and findings of osteomalacia in Patient 2. These differences in presentation between the two probands may reflect differences in age and gravidity-parity. In the literature, biochemical alterations, particularly in vitamin D levels, have not been commonly addressed in TRPS patients. We found reports of only two unrelated TRPS patients who displayed vitamin D deficiency, which, in both cases, was accompanied by hyperparathyroidism [28, 29]. It remains currently unknown whether this disorder specifically affects vitamin D metabolism. In contrast, TRPS1 protein has been investigated with respect to its role in bone remodeling and mineralization. Molecular studies have indicated that it modulates mineralized bone matrix formation in proliferating and differentiating osteoblast cells [30]. And polymorphisms in TRPS1 have been associated with bone mineral density in mice and humans [31–33]. Moreover, osteopenia and osteoporosis have been reported in several patients with TRPS [34–36]. Hence, it is possible that TRPS pathophysiology may include defective bone mineralization and/or reduced bone mass, which could further be complicated by vitamin D deficiency in some cases.
Three TRPS subtypes have been described: TRPS I [MIM# 190350], caused by pathogenic variants in the TRPS1 on chromosome 8; TRPS II [MIM# 150230], a contiguous gene deletion syndrome affecting the TRPS1, RAD21, and EXT1 genes; and TRPS III [MIM# 190351], a form with severe BDE and severe short stature. Lüdecke et al. [27] reported that TRPS III is the severe end of the TRPS spectrum and can be caused by a specific class of pathogenic variants in TRPS1. The TRPS1 protein is a zinc-finger transcription factor consisting of 1294 amino acids (NP_054831). It contains an unusual combination of nine zinc-finger motifs of eight different types. Thus far, the function of only the last three motifs has been identified, including the GATA DNA-binding zinc-finger, which is flanked by two nuclear localization signals. This region, harboring the novel p.Tyr928Cys variant identified in Patient 1 and her affected family members, shows strong evolutionary conservation (Fig. 6) and is critical for the transcription repressor activity of TRPS1 [37]. All identified missense pathogenic variants in this region have been reported to cause a severe phenotype including short-stature (i.e. TRPS III). The severe phenotype is proposed to result from a dominant-negative effect induced by those missense pathogenic variants as opposed to loss-of-function variants, such as nonsense or frameshift, which are likely to yield no more than a 50% reduction in activity, i.e. haploinsufficiency [27, 36]. These missense TRPS1 variants likely impair DNA-binding, and this is probably the case for the tyrosine-to-cysteine substitution at position 928, which could disrupt zinc ion-binding and/or the folding of the entire domain (Fig. 6). The ability of TRPS1 to form complexes with other proteins, however, may be preserved. Given that TRPS1 form homodimers for its activity [38], it is tempting to speculate that the transcriptionally inactive TRPS1 mutant dimerizes with and hinders the activity of the wild-type TRPS1 protein, thereby leading to a severe functional impairment. Consistent with this phenotype-genotype correlation, the phenotype in our patients carrying the p.Tyr928Cys variant was more severe than the phenotype in those carrying the p.Arg624* variant. Nonetheless, it seems that some loss-of-function pathogenic variants can also present with a severe phenotype, as a frameshift variant in exon 4 has been reported to cause a mild TRPS I-like phenotype in one kindred and, unexpectedly, a severe TRPS III-like phenotype in another [39]. Thus, the genetic and molecular mechanisms governing the effects of pathogenic TRPS1 variants remain incompletely understood.
Figure 6. Protein sequence conservation at the TRPS1 GATA DNA-binding domain and the modeling of the region comprising Try928.
The multiple alignment of the primary amino acid sequence with orthologs (top) indicate the evolutionary conservation; asterisks, cysteine residues predicted to interact with Zn2+. The tyrosine residue at position 928 is delineated on both sides. The peptide modeling (bottom) was performed by using SWISS-MODEL with residues 887–945 (template: human GATA-4). The Zn2+ ion and the four interacting cysteine residues are highlighted as ball-and-stick figures, as well as the native tyrosine residue at position 928.
In conclusion, we identified a novel missense pathogenic variant within the GATA DNA-binding region and a previously described nonsense pathogenic variant in TRPS1 as the genetic causes of BDE in two families. Our findings explain the clinical phenotypes of the affected individuals and contribute further to the phenotype-genotype correlation in TRPS.
Supplementary Material
Highlights.
Clinical features of two unrelated patients with brachydactyly type E and their family members were investigated.
A novel missense (p.Tyr928Cys) and a previously described nonsense variant (p.Arg624*) in TRPS1 were identified in the affected individuals
Affected individuals had additional features consistent with tricho-rhino-phalangeal syndrome (TRPS), caused by pathogenic TRPS1 variants.
p.Tyr928Cys is located in the GATA DNA-binding domain and associated with severe brachydactyly and short-stature.
The findings explain the brachydactyly E in two kindreds and provide insights into the phenotype-genotype correlation in TRPS.
Acknowledgments
This work was supported in part by a research grant from the National Institutes of Health/the National Institute of Diabetes and Digestive and Kidney Diseases (RO1 DK073911 to M.B.). Karaca A was supported by a grant from The Society of Endocrinology and Metabolism of Turkey.
Funding source: NIH/NIDDK RO1 DK073911
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflict of interests
The authors have no conflict of interest.
References
- 1.Temtamy SA and Aglan MS. Brachydactyly. Orphanet J Rare Dis. 2008; 3:15. doi: 10.1186/1750-1172-3-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pereda A et al. Brachydactyly E: isolated or as a feature of a syndrome. Orphanet J Rare Dis. 2013. September 12;8:141. doi: 10.1186/1750-1172-8-141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Johnson D, Kan SH, Oldridge M et al. Missense mutations in the homeodomain of HOXD13 are associated with brachydactyly types D and E. Am J Hum Genet. 2003. April;72(4):984–97 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brison N, Tylzanowski P, Debeer P. Limb skeletal malformations - what the HOX is going on? Eur J Med Genet. 2012. January;55(1):1–7. doi: 10.1016/j.ejmg.2011.06.003 [DOI] [PubMed] [Google Scholar]
- 5.Maass PG, Wirth J, Aydin A et al. A cis-regulatory site downregulates PTHLH in translocation t(8;12)(q13;p11.2) and leads to Brachydactyly Type E. Hum Mol Genet. 2010. March 1;19(5):848–60. doi: 10.1093/hmg/ddp553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Klopocki E, Hennig BP, Dathe K et al. Deletion and point mutations of PTHLH cause brachydactyly type E. Am J Hum Genet. 2010. March 12;86(3):434–9. doi: 10.1016/j.ajhg.2010.01.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Maass PG, Aydin A, Luft FC et al. PDE3A mutations cause autosomal dominant hypertension with brachydactyly. Nat Genet. 2015. June;47(6):647–53. doi: 10.1038/ng.3302 [DOI] [PubMed] [Google Scholar]
- 8.Chaabouni M, Le Merrer M, Raoul O et al. Molecular cytogenetic analysis of five 2q37 deletions: refining the brachydactyly candidate region. Eur J Med Genet. 2006. May-Jun;49(3):255–63. [DOI] [PubMed] [Google Scholar]
- 9.Williams SR, Aldred MA, Der Kaloustian VM et al. Haploinsufficiency of HDAC4 causes brachydactyly mental retardation syndrome, with brachydactyly type E, developmental delays, and behavioral problems. Am J Hum Genet. 2010. August 13;87(2):219–28. doi: 10.1016/j.ajhg.2010.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Weinstein LS, Gejman PV, Friedman E et al. Mutations of the Gs alpha-subunit gene in Albright hereditary osteodystrophy detected by denaturing gradient gel electrophoresis. Proc Natl Acad Sci U S A. 1990. November;87(21):8287–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Patten JL, Johns DR, Valle D et al. Mutation in the gene encoding the stimulatory G protein of adenylate cyclase in Albright’s hereditary osteodystrophy. N Engl J Med. 1990. May 17;322(20):1412–9. [DOI] [PubMed] [Google Scholar]
- 12.Linglart A, Menguy C, Couvineau A et al. Recurrent PRKAR1A mutation in acrodysostosis with hormone resistance. N Engl J Med. 2011. June 9;364(23):2218–26. doi: 10.1056/NEJMoa1012717. [DOI] [PubMed] [Google Scholar]
- 13.Michot C, Le Goff C, Goldenberg A et al. Exome sequencing identifies PDE4D mutations as another cause of acrodysostosis. Am J Hum Genet. 2012. April 6;90(4):740–5. doi: 10.1016/j.ajhg.2012.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lee H, Graham JM Jr, Rimoin DL et al. Exome sequencing identifies PDE4D mutations in acrodysostosis. Am J Hum Genet. 2012. April 6;90(4):746–51. doi: 10.1016/j.ajhg.2012.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Böni R, Huch Böni R, Tsambaos D Trichorhinophalangeal Syndrome. Dermatology 1995;190:152–155 doi: 10.1159/000246667 [DOI] [PubMed] [Google Scholar]
- 16.Maas S, Shaw A, Bikker H, Hennekam RCM. Trichorhinophalangeal Syndrome In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJH, Stephens K, Amemiya A, editors. GeneReviews®[Internet]. Seattle (WA): University of Washington, Seattle; 1993–2018. [PubMed] [Google Scholar]
- 17.Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46(W1), W296–W303 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Günöz H, Bundak R, Furman A et al. Z-score reference values for height in Turkish children aged 6 to 18 years. J Clin Res Pediatr Endocrinol. 2014;6(1):28–33. doi: 10.4274/Jcrpe.1260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Michael F Holick Neil C Binkley Heike A et al. Weaver Evaluation, Treatment, and Prevention of Vitamin D Deficiency: an Endocrine Society Clinical Practice Guideline The Journal of Clinical Endocrinology & Metabolism, Volume 96, Issue 7, 1 July 2011, Pages 1911–1930, 10.1210/jc.2011-0385 [DOI] [PubMed] [Google Scholar]
- 20.Elli FM, Bordogna P, de Sanctis L et al. Screening of PRKAR1A and PDE4D in a Large Italian Series of Patients Clinically Diagnosed With Albright Hereditary Osteodystrophy and/or Pseudohypoparathyroidism. J Bone Miner Res. 2016. June;31(6):1215–24. doi: 10.1002/jbmr.2785. [DOI] [PubMed] [Google Scholar]
- 21.Malik TH, Shoichet SA, Latham P et al. Transcriptional repression and developmental functions of the atypical vertebrate GATA protein TRPS1. EMBO J 2001; 20: 1715–1725. doi: 10.1093/emboj/20.7.1715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Richards S, Aziz N, Bale S et al. , of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015. 17: 405–423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Momeni P, Glöckner G, Schmidt O et al. Mutations in a new gene, encoding a zinc-finger protein, cause tricho-rhino-phalangeal syndrome type I. Nat Genet. 2000. January;24(1):71–4. doi: 10.1038/71717. [DOI] [PubMed] [Google Scholar]
- 24.Maas SM, Shaw AC, Bikker H et al. Phenotype and genotype in 103 patients with tricho-rhino-phalangeal syndrome. Eur J Med Genet. 2015. May;58(5):279–92. doi: 10.1016/j.ejmg.2015.03.002. [DOI] [PubMed] [Google Scholar]
- 25.Naselli A, Vignolo M, Di Battista E et al. Trichorhinophalangeal syndrome type I in monozygotic twins discordant for hip pathology: report on the morphological evolution of cone-shaped epiphyses and the unusual pattern of skeletal maturation. Pediatr Radiol 28:851–855. doi: 10.1007/s002470050481. [DOI] [PubMed] [Google Scholar]
- 26.Momeni P, Glöckner G, Schmidt O, et al. Mutations in a new gene, encoding a zinc-finger protein, cause tricho-rhino-phalangeal syndrome type I. Nature Genetics 24, 71–74 (2000), doi: 10.1038/71717. [DOI] [PubMed] [Google Scholar]
- 27.Lüdecke HJ, Schaper J, Meinecke P, et al. Genotypic and phenotypic spectrum in tricho-rhino-phalangeal syndrome types I and III. Am J Hum Genet. 2000;68(1):81–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ye D, Fei Y, Sheng et a Y‐E. Analysis of a Chinese pedigree with trichorhinophalangeal syndrome derived from a missense mutation in the TRPS1 gene. Clin Exp Dermatol. 2017. doi: 10.1111/ced.13082 [DOI] [PubMed] [Google Scholar]
- 29.Crippa M, Bestetti I, Perotti M et al. New case of trichorinophalangeal syndrome-like phenotype with a de novo t(2;8)(p16.1;q23.3) translocation which does not disrupt the TRPS1 gene. BMC Med Genet. 2014. May 2;15:52. doi: 10.1186/1471-2350-15-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Piscopo DM, Johansen EB, Derynck R et al. Identification of the GATA factor TRPS1 as a repressor of the osteocalcin promoter. J Biol Chem. 2009;284(46):31690–703. doi: 10.1074/jbc.M109.052316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ackert-Bicknel CL, Demissie S, Tsaih SW et al. Genetic variation in TRPS1 may regulate hip geometry as well as bone mineral density. Bone. 2012. May;50(5):1188–95. doi: 10.1016/j.bone.2012.01.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ishimori N, Stylianou IM, Korstanje R et al. Quantitative trait loci for BMD in an SM/J by NZB/BlNJ intercross population and identification of Trps1 as a probable candidate gene. J Bone Miner Res. 2008. September;23(9):1529–37. doi: 10.1359/jbmr.080414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang L, Lu W, Zhang L et al. Trps1 differentially modulates the bone mineral density between male and female mice and its polymorphism associates with BMD differently between women and men. PLoS One. 2014. January 8;9(1):e84485. doi: 10.1371/journal.pone.0084485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stagi S, Bindi G, Galluzzi F et al. Partial growth hormone deficiency and changed bone quality and mass in type I trichorhinophalangeal syndrome. Am J Med Genet A. 2008. June 15;146A(12):1598–604. doi: 10.1002/ajmg.a.32348. [DOI] [PubMed] [Google Scholar]
- 35.Shao C, Tian J, Shi DH et al. A novel mutation in TPRS1 gene caused tricho-rhino-phalangeal syndrome in a Chinese patient with severe osteoporosis. Chin Med J (Engl). 2011. May;124(10):1583–5. [PubMed] [Google Scholar]
- 36.Macchiaiolo M, Mennini M, Digilio MC et al. Thricho-rhino-phalangeal syndrome and severe osteoporosis: a rare association or a feature? An effective therapeutic approach with bisphosphonates. Am J Med Genet A. 2014. March;164A(3):760–3. doi: 10.1002/ajmg.a.36327. [DOI] [PubMed] [Google Scholar]
- 37.Malik TH, Shoichet SA, Latham P et al. Transcriptional repression and developmental functions of the atypical vertebrate GATA protein TRPS1. EMBO J. 2001. April 2;20(7):1715–25. doi: 10.1093/emboj/20.7.1715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.McCarty AS, Kleiger G, Eisenberg D, Smale ST. Selective dimerization of a C2H2 zinc finger subfamily. Mol Cell. 2003. February;11(2):459–70. [DOI] [PubMed] [Google Scholar]
- 39.Itoh M, Kittaka Y, Niida Y, Saikawa Y et al. A novel frameshift mutation in the TRPS1 gene caused Tricho-rhino-phalangeal syndrome type I and III in a Japanese family. Clin Pediatr Endocrinol. 2016;25(3):115–8. doi: 10.1297/cpe.25.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.