Abstract
Recruitment of the Polycomb repressive complexes PRC1 and PRC2 by Xist RNA is an important paradigm for chromatin regulation by long non-coding RNAs. Here we show that the non-canonical PCGF3/5-PRC1 complex initiates recruitment of both PRC1 and PRC2 in response to Xist RNA expression. PCGF3/5-PRC1 mediated ubiquitylation of histone H2A signals recruitment of other non-canonical PRC1 complexes, and of PRC2, leading to deposition of histone H3 lysine 27 methylation chromosome wide. Pcgf3/5 gene knockout results in female specific embryo lethality, and abrogates Xist mediated gene repression, highlighting a key role for Polycomb in Xist dependent chromosome silencing. Our findings overturn existing models for Polycomb recruitment by Xist RNA, and moreover establish precedence for H2AK119u1 in initiating Polycomb domain formation in a physiological context.
Polycomb proteins play an important role in developmental gene regulation, functioning primarily by establishing stable repressive chromatin states. Accordingly, the two major Polycomb repressor complexes, PRC1 and PRC2, catalyse post-translational modifications of core histone proteins, mono-ubiquitylation of histone H2A lysine 119 (H2AK119u1), and methylation of histone H3 lysine 27 (H3K27me3) respectively (reviewed in (1)). In mammalian cells, the non-coding RNA Xist recruits PRC1 and PRC2 to the inactive X chromosome (Xi) (2–4). Previous studies have proposed that Polycomb recruitment is initiated by direct binding of PRC2 to Xist RNA (5), although some evidence is difficult to reconcile with this model (6,7).
Non-canonical PRC1 complexes are recruited to Xi via an alternative pathway (8–10), and recent studies have demonstrated that PRC2 complexes can bind to PRC1 mediated H2AK119u1 (11–14), indicating that PRC1 could have a role in initiating Polycomb recruitment on Xi. To test this possibility we developed a mouse embryonic stem cell (mESC) model in which Xist RNA could be induced in the absence of RING1A/B, the core catalytic subunit of PRC1 (Note S1, Fig S1A, B). As shown in Fig 1A-C, deletion of RING1A/B largely abolished Xist dependent PRC2 (EZH2) recruitment, and H3K27me3 deposition. Formation of Xist RNA domains was unaltered (Fig. 1C, D).
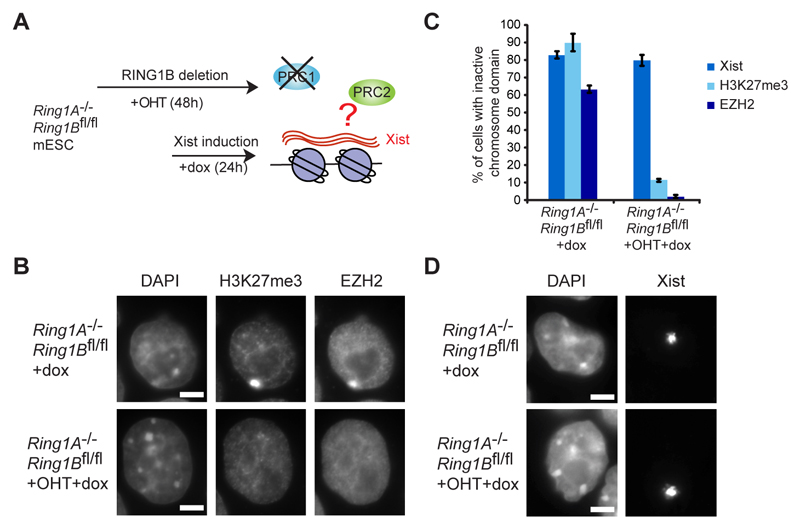
Fig. 1. H2AK119u1 mediates PRC2 recruitment by Xist RNA.
A. Schematic of the experiment. Tamoxifen (+OHT). Doxycycline (+dox).
B. Examples of H3K27me3 and EZH2 immunofluorescence following 24h Xist induction.
C. Percentage of cells with domains for Xist, H3K27me3 and EZH2 in tamoxifen treated and untreated cells after Xist induction for 24h. A minimum of 800 cells were counted in 4 biological repeats. Error bars indicate standard deviation.
D. Xist RNA FISH illustrating deletion of Ring1A/B has no effect on Xist RNA domain formation.
We confirmed these findings by depleting H2AK119u1 using MG132, a proteasome inhibitor (15), in 3E mESCs that carry an inducible Xist transgene (16). H2AK119u1 was extensively depleted 6h after MG132 treatment (Fig S1C, D), as reported previously (9). Xist dependent recruitment of PRC2 (EED and H3K27me3 domains), was abolished (Fig S1E,F). Together, these findings indicate that H2AK119u1 deposition is required for Xist dependent recruitment of PRC2.
It has been proposed that the A-repeat region of Xist RNA, and/or a 4kb region immediately downstream in Xist exon 1, designated XN and encompassing the F, B and C repeats, mediates PRC2 recruitment in X inactivation (5,7). In light of our findings we analysed the role of these regions in Xist dependent H2AK119u1 deposition. Thus, we established cell lines with inducible Xist transgenes carrying deletions of either the A-repeat or the XN region (Fig. S2A). Immuno-FISH analysis of Xist RNA together with either H2AK119u1 or H3K27me3 revealed that deletion of the A-repeat had no effect, whereas deletion of the XN region entirely abolished Xist dependent deposition of H2AK119u1 and H3K27me3 (Fig S2B-E). These findings implicate the XN region in recruitment of PRC1 complex(es) required to initiate H2AK119u1 deposition on Xi.
In vertebrates, the catalytic core of PRC1 comprises RING1A/B and one of six homologues of the Polycomb group RING finger (PCGF) protein. Unique characteristics of PCGF1-6 define distinct subunit assemblies, subdivided into canonical and non-canonical PRC1 complexes (9, 17) (Fig. S3A). PCGF2 and PCGF4 are close homologues, as are PCGF3 and PCGF5 (Fig S3B). To determine which of the different PRC1 complexes are recruited to Xist RNA domains, we stably expressed N-terminal enhanced green fluorescent protein (eGFP) fusion proteins for PCGF1, PCGF2 (which together with PCGF4, contributes to either canonical or non-canonical PRC1 complexes), PCGF3, PCGF5 and PCGF6 in 3E-H inducible Xist mESCs (18) (Fig S3C). All of the eGFP-PCGF fusion proteins were assembled into PRC1 complexes, as judged by co-immunoprecipitation analysis of the RING1B subunit (Fig S3D). Live cell confocal imaging revealed Xist dependent concentration of each eGFP-PCGF fusion protein within a single discrete sub-nuclear region, referred to henceforth as Xist RNA domains (Fig. 2A, Movie S1). Localisation of eGFP-PCGF fusion proteins to Xist RNA domains was also observed using immunofluorescence (IF), albeit to varying degrees (Fig S3E, F). The variable efficiency of detection by IF likely reflects the dynamic behaviour of different PRC1 complexes (Note S2).
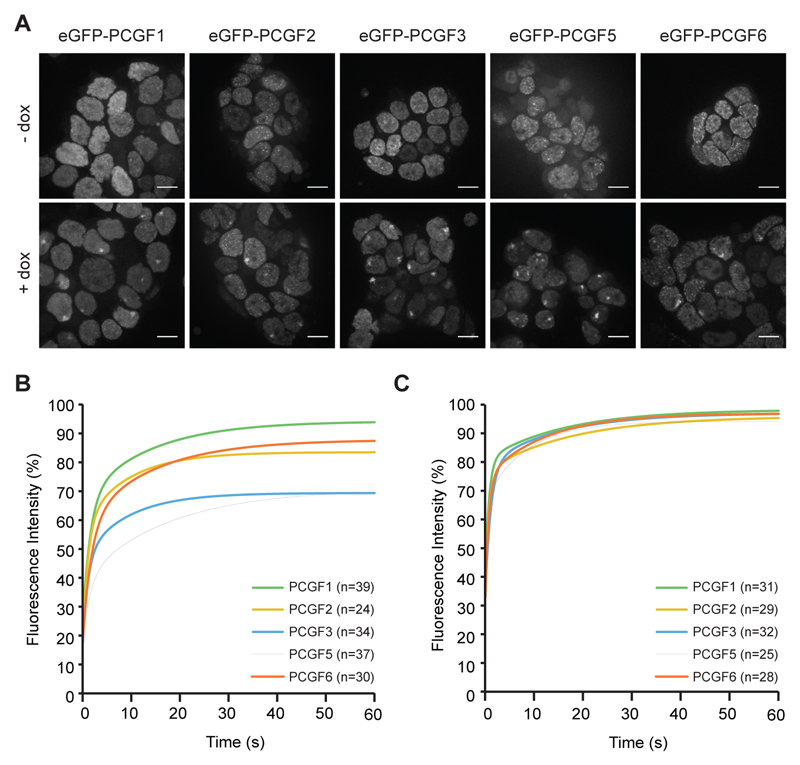
Fig. 2. Xist dependent localisation and dynamics of non-canonical PRC1 complexes.
A. Images illustrating focal enrichment of eGFP-PCGF fusion proteins following induction of Xist RNA for 24h in 3E-H mESCs sublines (+ dox). Focal enrichment is not seen in uninduced cells (- dox). Images are maximum intensity projections of six consecutive z-stacks. Scale bar is 10μm.
B. FRAP of eGFP-PCGF fusion proteins within Xist RNA domains.
C. FRAP of eGFP-PCGF fusion proteins within randomly selected regions of the nucleoplasm. In (B) and (C) data from single cells was fitted with a bi-exponential equation. Curves represent average of the fit from a number (n) of cells.
The localisation of eGFP-PCGF fusion proteins over Xist RNA domains enabled us to compare the dynamic behaviour of different PRC1 complexes using fluorescence recovery after photobleaching (FRAP). Thus, for each eGFP-PCGF fusion protein, we took FRAP measurements both within Xist RNA domains, and also in a randomly selected nucleoplasmic region. For eGFP-PCGF1, eGFP-PCGF2 and eGFP-PCGF6 the average recovery within Xist RNA domains was in the order of a few seconds. In contrast, for eGFP-PCGF3 and eGFP-PCGF5 a significant immobile fraction, retained over the time-course of the experiment (60s), indicated a relatively stable interaction (Fig. 2B, Fig S4A,B,D). We refer to these interaction modes as transient (<60s) and stable (>60s) respectively. In the nucleoplasm, all of the eGFP-PCGF proteins showed similar transient dynamic behaviour, fully recovering over a timescale of seconds (Fig. 2C, Fig S4B-D). This suggests that the distinct behaviour of eGFP-PCGF3 and eGFP-PCGF5 reflects differences in their interactions with the Xist RNA domain, rather than systematic differences between the eGFP- fusion proteins or the cell lines. These conclusions were further supported by analysis of the rate of turnover (t1/2), determined by fitting recovery curves with a bi-exponential equation (Note S3, Fig S4C). The FRAP experiments thus reveal a qualitative difference between complexes formed by PCGF1/2/6, which interact transiently with the Xist RNA domain, and eGFP-PCGF3/5-PRC1 which additionally form stable interactions.
The non-canonical PRC1 subunit RYBP, and/or its homologue YAF2, can bind to H2AK119u1, mediated by an NZF domain (9,17,19, 20). To test if this interaction could contribute to localisation of PRC1 complexes within Xist RNA domains, we generated Rybp-/- Yaf2-/- mESCs (Note S4, Fig S5A,B). Recruitment of PRC1 (and PRC2) by Xist RNA was retained, as indicated by accumulation of H2AK119u1 and H3K27me3 (Fig S5C,E). However, quantification of domain size and time-course analysis revealed that deposition of H2AK119u1 and H3K27me3 is impaired relative to wild-type mESCs (Note S5, Fig S5D,E). The time-course analysis (Fig S5E) further illustrates that H2AK119u1 deposition within Xist RNA domains precedes H3K27me3, supporting that PRC1 initiates PRC2 recruitment.
Interestingly, although PRC1 activity over Xist RNA domains is evident in Rybp-/- Yaf2-/- mESCs, we were unable to detect enrichment of eGFP-PCGF fusion proteins using either live cell imaging or conventional IF (Fig S6A, Movies S2-6), likely reflecting their reduced levels relative to background. Consistent with this idea, we were able to detect localisation of eGFP-PCGF3 and eGFP-PCGF5, using a modified IF protocol (Note S6, Fig S6B).
We went on to test if reduced Xist RNA domain enrichment of eGFP-PCGF proteins in Rybp-/-Yaf2-/- mESCs is linked to the ability of RYBP to bind H2AK119u1. Thus, we complemented Rybp-/-Yaf2-/- mESCs using constructs that express mCherry tagged wild-type RYBP or a mutant protein, RYBP TF-AA, which has three point mutations in the NZF domain that abrogates the interaction with H2AK119u1 (20). As illustrated in Fig 3, wild-type, but not TF-AA RYBP, restored Xist RNA domain enrichment of eGFP-PCGF fusion proteins. We conclude that RYBP/YAF2 binding to H2AK119u1 likely underpins the transient interaction of non-canonical PRC1 complexes with Xist RNA domains, but does not account for stable interaction of PCGF3/5-PRC1 complexes.
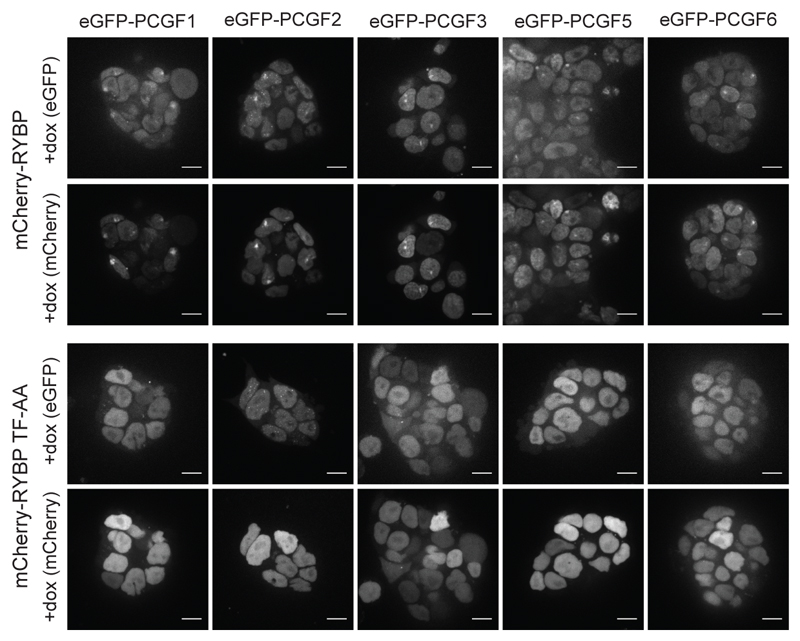
Fig. 3. RYBP/YAF2 interaction with H2AK119u1 contributes to Xist dependent enrichment of non-canonical PRC1 complexes.
Images illustrating distribution of eGFP-PCGF proteins in Rybp-/-Yaf2-/- mESC lines complemented with wild-type mCherry-RYBP, or mCherry-RYBP TF-AA mutant after Xist induction (+dox) for 24 h. Images are maximum intensity projections of six consecutive z-stacks. Scale bar is 10 μm.
We next tested the role of PCGF3/5-PRC1 complexes in initiating Polycomb recruitment by Xist RNA. Thus, we engineered a series of ESC lines with doxycycline inducible Xist transgenes in which we could investigate the importance of PCGF3 and PCGF5 either alone or together (Fig S7A,B,D,E). A conditional knockout mESC line for Pcgf1 was used as a negative control (Note S7, Fig S7A-C). We then tested if H2AK119u1 and H3K27me3 deposition occur over Xist RNA domains 24 h after induction of the Xist transgene (Fig. 4A). Deletion of either Pcgf1 or Pcgf5 alone had no effect on Xist dependent H2AK119u1 and H3K27me3 deposition (Fig S8A-D). Deletion of Pcgf3 alone resulted in a small but significant decrease of both H2AK119u1, and H3K27me3 positive domains (Fig. Fig S8C,D). Strikingly, deletion of both Pcgf3 and Pcgf5 together strongly reduced Xist dependent H2AK119u1 and H3K27me3 deposition (Fig. 4B,C, Fig S8C,D,E). Moreover, in Pcgf3-/-Pcgf5-/- mESCs that were cultured with tamoxifen for 96 h and then expanded prior to inducing Xist expression, H2AK119u1 and/or H3K27me3 domains were entirely abolished (Fig. 4C). Global levels of H2AK119u1, H3K27me3, and PRC1 subunits were unaffected (Fig S8F), indicating that the observed deficiency is specific to Polycomb recruitment by Xist RNA.
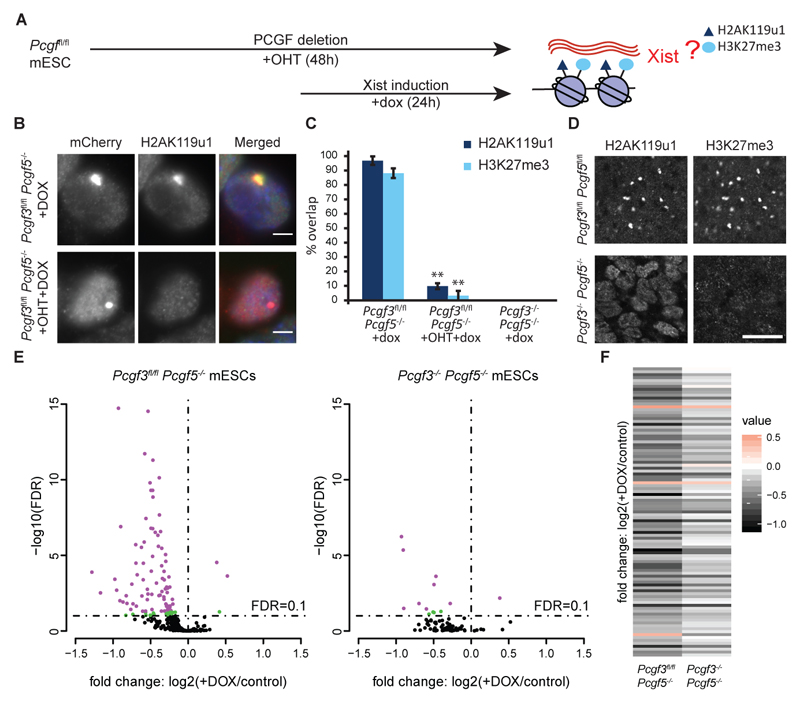
Fig. 4. PCGF3/5-PRC1 initiates Polycomb recruitment to facilitate chromosome silencing by Xist RNA.
A. Schematic representation of PCGF knockout experiments. Tamoxifen (+OHT). Doxycycline (+dox).
B. Images illustrating accumulation of H2AK119u1 over Xist RNA domains in the presence and absence of PCGF proteins.
C. Scoring data representing a minimum of 300 cells in three replicates is summarised in the bar chart. Error bars show standard deviation. Significant differences (Student t-test) are indicated (**p<0.001).
D. Immunofluorescence analysis of H2AK119u1 and H3K27me3 domains in Pcgf3-/- :Pcgf5-/- and Pcgf3fl/fl:Pcgf5fl/fl female embryos at E7.5. Images show representative single focal plane scans through epiblast. Scale bar is 20 μm.
E. Genes differentially expressed between Xist induced (+dox) and uninduced (control) in Pcgf3fl/flPcgf5-/- (left panel) and Pcgf3-/-Pcgf5-/- (right panel) mESCs on Xist transgene bearing chromosome 16. Genes with FDR< 0.05 are depicted in violet; FDR >0.05 and <0.1 are in green.
F. Heatmap showing expression change between Xist induced (+dox) and uninduced (control) mESCs for all genes significantly misregulated on chromosome 16 in Pcgf3fl/flPcgf5-/- (FDR<0.1) together with expression changes for the same genes in Pcgf3-/-Pcgf5-/- mESCs.
Our results suggest that PCGF3/5-PRC1 mediated H2AK119u1 initiates Xist dependent recruitment both of other non-canonical PRC1 complexes and of PRC2. In support of this model, Xist dependent recruitment of eGFP-PCGF3 or eGFP-PCGF5 occurs in the absence of PRC2 and H3K27me3 (Note S8, Fig S9A), and is abolished in cells expressing the ΔXN Xist RNA (Note S9, Fig S9B,C). Moreover, analysis by 3D-structured illumination super-resolution microscopy revealed that PCGF3/5-PRC1 is significantly closer to Xist RNA than are other non-canonical PRC1 complexes (Note S10, Fig S10A, B).
To verify our results in the context of X inactivation in vivo, we went on to establish Pcgf3 and Pcgf5 conditional knockout mouse lines (Fig S7D, FigS11A). Analysis of E7.5 XX Pcgf3-/-Pcgf5-/- embryos revealed complete absence of H2AK119u1 and H3K27me3 on Xi in embryonic and extraembryonic regions (Note S11, Fig. 4D, Fig S11B-E). We further analysed progeny from Pcgf3+/-:Pcgf5+/- heterozygote crosses. Pcgf3-/-Pcgf5-/- were not recovered amongst live-born progeny. At mid-gestation stages (E9.5 and E10.5) we recovered male Pcgf3-/-Pcgf5-/- conceptuses in which the embryos and placentas appeared grossly normal. However, in female Pcgf3-/-Pcgf5-/- conceptuses, the embryo was extensively degraded (Note S12, Fig S12A). Histological analysis revealed some defects in placentas from male embryos, but more strikingly, female placentas lack trophoblasts altogether and as a consequence fail to form a labyrinth (Note S12, Fig. S12B). Other genotypes, including Pcgf3 and Pcgf5 single homozygous embryos, were found at approximately the expected Mendelian ratio (Fig S12C).
Female specific phenotypes in Pcgf3-/-Pcgf5-/- embryos suggest that Xist mediated silencing may be compromised. To test this directly we performed RNA-Seq analysis, comparing silencing in the presence or absence of Xist dependent Polycomb recruitment in mESCs (Pcgf3fl/flPcgf5-/- compared with Pcgf3-/-Pcgf5-/-) (Note S13). The results demonstrate strong abrogation of Xist mediated gene silencing affecting genes across the entire chromosome (Fig 4E,F, Fig S13A-C). We conclude that loss of both PRC1 and PRC2 significantly compromises Xist mediated silencing.
This study redefines the molecular pathway for Polycomb recruitment by Xist RNA, as summarised in Fig. S14. Our findings further provide proof of principle that non-canonical PRC1 and associated H2AK119u1 can function to initiate the formation of Polycomb domains in a physiological context. In future studies it will be important to understand how PCGF3/5-PRC1 interacts with the Xist XN region, and to determine if the same pathway is utilised by other lncRNAs.
Supplementary Material
Acknowledgements
We thank the Brockdorff and Koseki labs for comments, A. Otte and F. Barr for antibodies, R. Klose for plasmids and antibodies, A. Bassett for facilitating CRISPR/Cas9 mutagenesis, D. Moralli and C. Green for transgene mapping, E. Hookway for sequencing. Work in N.B. lab was funded by grants from the Wellcome Trust (103768 and 091911) and European Research Council (340081). M.A. was funded by a PhD fellowship from Fundação para a Ciência e Tecnologia, Portugal (SFRH/BD/51706/2011). H.K. was funded by CREST from JST-AMED. AM was funded by Wellcome Trust (102811). Accession number: ArrayExpress E-MTAB-5642. DB was funded by a Wellcome Trust PhD studentship (092926).
References
- 1.Schwartz YB, Pirrotta V. A new world of Polycombs: unexpected partnerships and emerging functions. Nat Rev Genet. 2013;14:853–864. doi: 10.1038/nrg3603. published online EpubDec (nrg3603 [pii]) [DOI] [PubMed] [Google Scholar]
- 2.de Napoles M, Mermoud JE, Wakao R, Tang YA, Endoh M, Appanah R, Nesterova TB, Silva J, Otte AP, Vidal M, Koseki H, et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Developmental cell. 2004;7:663–676. doi: 10.1016/j.devcel.2004.10.005. published online EpubNov. [DOI] [PubMed] [Google Scholar]
- 3.Silva J, Mak W, Zvetkova I, Appanah R, Nesterova TB, Webster Z, Peters AH, Jenuwein T, Otte AP, Brockdorff N. Establishment of histone h3 methylation on the inactive X chromosome requires transient recruitment of Eed-Enx1 polycomb group complexes. Developmental cell. 2003;4:481–495. doi: 10.1016/s1534-5807(03)00068-6. published online EpubApr. [DOI] [PubMed] [Google Scholar]
- 4.Plath K, Fang J, Mlynarczyk-Evans SK, Cao R, Worringer KA, Wang H, de la Cruz CC, Otte AP, Panning B, Zhang Y. Role of histone H3 lysine 27 methylation in X inactivation. Science. 2003;300:131–135. doi: 10.1126/science.1084274. published online EpubApr 4. [DOI] [PubMed] [Google Scholar]
- 5.Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science. 2008;322:750–756. doi: 10.1126/science.1163045. published online EpubOct 31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cerase A, Smeets D, Tang YA, Gdula M, Kraus F, Spivakov M, Moindrot B, Leleu M, Tattermusch A, Demmerle J, Nesterova TB, et al. Spatial separation of Xist RNA and polycomb proteins revealed by superresolution microscopy. Proc Natl Acad Sci U S A. 2014;111:2235–2240. doi: 10.1073/pnas.1312951111. published online EpubFeb 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.da Rocha ST, Boeva V, Escamilla-Del-Arenal M, Ancelin K, Granier C, Matias NR, Sanulli S, Chow J, Schulz E, Picard C, Kaneko S, et al. Jarid2 Is Implicated in the Initial Xist-Induced Targeting of PRC2 to the Inactive X Chromosome. Molecular cell. 2014;53:301–316. doi: 10.1016/j.molcel.2014.01.002. published online EpubJan 23. [DOI] [PubMed] [Google Scholar]
- 8.Schoeftner S, Sengupta S, Kubicek S, Mechtler K, Spahn L, Koseki H, Jenuwein T, Wutz A. Recruitment of PRC1 function at the initiation of X inactivation independent of PRC2 and silencing. EMBO J. 2006;25:3110–3122. doi: 10.1038/sj.emboj.7601187. published online EpubJul 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tavares L, Dimitrova E, Oxley D, Webster J, Poot R, Demmers J, Bezstarosti K, Taylor S, Ura H, Koide H, Wutz A, et al. RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell. 2012;148:664–678. doi: 10.1016/j.cell.2011.12.029. published online EpubFeb 17 (S0092-8674-12)00027-X [pii] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chu C, Zhang QC, da Rocha ST, Flynn RA, Bharadwaj M, Calabrese JM, Magnuson T, Heard E, Chang HY. Systematic discovery of Xist RNA binding proteins. Cell. 2015;161:404–416. doi: 10.1016/j.cell.2015.03.025. published online EpubApr 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blackledge NP, Farcas AM, Kondo T, King HW, McGouran JF, Hanssen LL, Ito LL, Cooper S, Kondo K, Koseki Y, Ishikura T, et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell. 2014;157:1445–1459. doi: 10.1016/j.cell.2014.05.004. published online EpubJun 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cooper S, Dienstbier M, Hassan R, Schermelleh L, Sharif J, Blackledge NP, De Marco V, Elderkin S, Koseki H, Klose R, Heger A, et al. Targeting polycomb to pericentric heterochromatin in embryonic stem cells reveals a role for H2AK119u1 in PRC2 recruitment. Cell reports. 2014;7:1456–1470. doi: 10.1016/j.celrep.2014.04.012. published online EpubJun 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kalb R, Latwiel S, Baymaz HI, Jansen PW, Muller CW, Vermeulen M, Muller J. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nature structural & molecular biology. 2014;21:569–571. doi: 10.1038/nsmb.2833. published online EpubJun. [DOI] [PubMed] [Google Scholar]
- 14.Cooper S, Grijzenhout A, Underwood E, Ancelin K, Zhang T, Nesterova TB, Anil-Kirmiztas B, Bassett A, Kooistra SM, Agger K, Helin K, et al. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nature communications. 2016;7 doi: 10.1038/ncomms13661. 13661; published online EpubNov 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Materials and methods are available as supplementary materials at the Science website.
- 16.Tang YA, Huntley D, Montana G, Cerase A, Nesterova TB, Brockdorff N. Efficiency of Xist-mediated silencing on autosomes is linked to chromosomal domain organisation. Epigenetics Chromatin. 2010;3:10. doi: 10.1186/1756-8935-3-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gao Z, Zhang J, Bonasio R, Strino F, Sawai A, Parisi F, Kluger Y, Reinberg D. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Molecular cell. 2012;45:344–356. doi: 10.1016/j.molcel.2012.01.002. published online EpubFeb 10 (S1097-2765-12)00035-4 [pii] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Moindrot B, Cerase A, Coker H, Masui O, Grijzenhout A, Pintacuda G, Schermelleh L, Nesterova TB, Brockdorff N. A Pooled shRNA Screen Identifies Rbm15, Spen, and Wtap as Factors Required for Xist RNA-Mediated Silencing. Cell reports. 2015;12:562–572. doi: 10.1016/j.celrep.2015.06.053. published online EpubJul 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Garcia E, Marcos-Gutierrez C, del Mar Lorente M, Moreno JC, Vidal M. RYBP, a new repressor protein that interacts with components of the mammalian Polycomb complex, and with the transcription factor YY1. The EMBO journal. 1999;18:3404–3418. doi: 10.1093/emboj/18.12.3404. published online EpubJun 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Arrigoni R, Alam SL, Wamstad JA, Bardwell VJ, Sundquist WI, Schreiber-Agus N. The Polycomb-associated protein Rybp is a ubiquitin binding protein. FEBS Lett. 2006;580:6233–6241. doi: 10.1016/j.febslet.2006.10.027. published online EpubNov 13. [DOI] [PubMed] [Google Scholar]
- 21.Seibler J, Zevnik B, Kuter-Luks B, Andreas S, Kern H, Hennek T, Rode A, Heimann C, Faust N, Kauselmann G, Schoor M, et al. Rapid generation of inducible mouse mutants. Nucleic acids research. 2003;31:e12. doi: 10.1093/nar/gng012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sakai K, Miyazaki J. A transgenic mouse line that retains Cre recombinase activity in mature oocytes irrespective of the cre transgene transmission. Biochemical and biophysical research communications. 1997;237:318–324. doi: 10.1006/bbrc.1997.7111. [DOI] [PubMed] [Google Scholar]
- 23.Endoh M, Endo TA, Endoh T, Fujimura Y, Ohara O, Toyoda T, Otte AP, Okano M, Brockdorff N, Vidal M, Koseki H. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development. 2008;135:1513–1524. doi: 10.1242/dev.014340. published online EpubApr. [DOI] [PubMed] [Google Scholar]
- 24.Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 2013;154:1380–1389. doi: 10.1016/j.cell.2013.08.021. published online EpubSep 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wutz A, Rasmussen TP, Jaenisch R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nature genetics. 2002;30:167–174. doi: 10.1038/ng820. published online EpubFeb. [DOI] [PubMed] [Google Scholar]
- 26.Mueller F, Karpova TS, Mazza D, McNally JG. Monitoring dynamic binding of chromatin proteins in vivo by fluorescence recovery after photobleaching. Methods in molecular biology. 2012;833:153–176. doi: 10.1007/978-1-61779-477-3_11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sprague BL, Pego RL, Stavreva DA, McNally JG. Analysis of binding reactions by fluorescence recovery after photobleaching. Biophysical journal. 2004;86:3473–3495. doi: 10.1529/biophysj.103.026765. published online EpubJun. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ball G, Demmerle J, Kaufmann R, Davis I, Dobbie IM, Schermelleh L. SIMcheck: a Toolbox for Successful Super-resolution Structured Illumination Microscopy. Scientific reports. 2015;5 doi: 10.1038/srep15915. 15915; published online EpubNov 03. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Dignam JD, Lebovitz RM, Roeder RG. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic acids research. 1983;11:1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nojima T, Gomes T, Grosso AR, Kimura H, Dye MJ, Dhir S, Carmo-Fonseca M, Proudfoot NJ. Mammalian NET-Seq Reveals Genome-wide Nascent Transcription Coupled to RNA Processing. Cell. 2015;161:526–540. doi: 10.1016/j.cell.2015.03.027. published online EpubApr 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. published online EpubJan 01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Huber W, Carey VJ, Gentleman R, Anders S, Carlson M, Carvalho BS, Bravo HC, Davis S, Gatto L, Girke T, Gottardo R, et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nature methods. 2015;12:115–121. doi: 10.1038/nmeth.3252. published online EpubFeb. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Schmiedeberg L, Skene P, Deaton A, Bird A. A temporal threshold for formaldehyde crosslinking and fixation. PLoS One. 2009;4:e4636. doi: 10.1371/journal.pone.0004636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Smeets D, Markaki Y, Schmid VJ, Kraus F, Tattermusch A, Cerase A, Sterr M, Fiedler S, Demmerle J, Popken J, Leonhardt H, et al. Three-dimensional super-resolution microscopy of the inactive X chromosome territory reveals a collapse of its active nuclear compartment harboring distinct Xist RNA foci. Epigenetics & chromatin. 2014;7:8. doi: 10.1186/1756-8935-7-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.