Abstract
The aim of this study was to systematically appraise the existing literature on the yet-unclear heritability of gingivitis and periodontitis. This review was conducted following the PRISMA guidelines. A search was conducted through the electronic databases Medline, Embase, LILACS, Cochrane Library, Open Grey, Google Scholar, and Research Gate, as complemented by a hand search, for human studies reporting measures of heritability of gingivitis and periodontitis. A total of 9,037 papers were initially identified from combined databases and 10,810 on Google Scholar. After full-text reading, 28 articles met the inclusion criteria and were carried forward to data abstraction. The reviewed data included information from >50,000 human subjects. Meta-analyses were performed by grouping studies based on design and outcome. Heritability (H2) of periodontitis was estimated at 0.38 (95% CI, 0.34 to 0.43; I2 = 12.9%) in twin studies, 0.15 (95% CI, 0.06 to 0.24; I2 = 0%) in other family studies, and 0.29 (95% CI, 0.21 to 0.38; I2 = 61.2%) when twin and other family studies were combined. Genome-wide association studies detected a lower heritability estimate of 0.07 (95% CI, −0.02 to 0.15) for combined definitions of periodontitis, increasing with disease severity and when the interaction with smoking was included. Furthermore, heritability tended to be lower among older age groups. Heritability for the self-reported gingivitis trait was estimated at 0.29 (95% CI, 0.22 to 0.36; I2 = 37.6%), while it was not statistically significant for clinically measured gingivitis. This systematic review brings forward summary evidence to confirm that up to a third of the periodontitis variance in the population is due to genetic factors. This seems consistent across the different studied populations and increases with disease severity. In summary, up to a third of the variance of periodontitis in the population is due to genetic factors, with higher heritability for more severe disease.
Keywords: genetics, periodontal disease, meta-analysis, heredity, GWAS, gingivitis
Background
Heritability measures the ratio of the genetic component to the total phenotypic variance or, in other words, how much genetics can account for the difference in the distribution of disease seen in a population (Visscher et al. 2008). Therefore, this is a concept applied to a “population” rather than to specific individuals. Knowledge of disease heritability can help set boundaries on the potential of genetic variants to explain health and disease traits, inform genetic risk estimation, and estimate familial disease risk (Smith 1971).
Heritability has traditionally been estimated by conducting twin or family studies with different analytic approaches (Tenesa and Haley 2013) and, more recently, by genome-wide association studies (GWASs). GWASs have been used to estimate the overall genetic contribution to physical characteristics (e.g., human height; Yang et al. 2010) and diseases (e.g., glioma; Kinnersley et al. 2015). However, most GWASs have sample sizes that allow them to effectively examine only common variants (>1% or 5% minor allele frequency), providing a lower bound for narrow-sense heritability (Kinnersley et al. 2015).
The heritability of periodontal diseases, including gingivitis and periodontitis, was estimated in animal models (Miley et al. 2011; Hiyari et al. 2015) and in humans. Notably, studies of twins reported periodontitis heritability estimates that vary from around 0.3 to 0.5 (30% to 50%; Michalowicz, Aeppli, Virag, et al. 1991; Corey et al. 1993; Mucci et al. 2005). In contrast, other studies rejected the hypothesis of substantial heritability for periodontitis (Dowsett et al. 2002).
Overall, there is still a high degree of uncertainty regarding how much of the observed variance in gingivitis and periodontitis is attributable to genetics. In the era of precision medicine (Divaris 2017), with a continuous effort to derive individual disease susceptibility assessments to inform prevention and treatment, it is important to try to clarify this aspect. With that in mind, we aimed to systematically appraise the existing evidence base to estimate the heritability of gingivitis and periodontitis.
Materials and Methods
A systematic review protocol was written in the planning stages, and the PRISMA checklist (Appendix; Moher et al. 2009) was followed in the planning and reporting of the review. The protocol was registered with PROSPERO (CRD42018091892).
Focused Question
What is the heritability (contribution of host genetic variants) of gingivitis and periodontitis in humans?
PECO Outline
Population: subjects with measures of periodontal disease or periodontal health
Exposure: analysis of host genetic risk factors
Comparisons: subjects with periodontal health
Outcomes: presence of different degrees of periodontitis (and/or continuous measures of periodontitis or tooth loss, when available)
Eligibility Criteria
Twin studies, including monozygotic (MZ) twins and dizygotic (DZ) twins, and family studies reporting measures of heritability of periodontal outcomes (clinical, radiographic or patient-reported outcomes of gingivitis and periodontitis including tooth loss) or concordance or correlation rates, which allow the calculation of heritability estimates
GWASs (case-control, cohort, or cross-sectional design) or other case-control or cross-sectional studies of humans that report heritability estimates to periodontal outcomes (as before)
Animal studies reporting heritability of periodontal disease outcomes (clinical or radiographic)
Any paper not fulfilling these criteria was excluded. The present paper reports data relative only to studies of humans.
Choice of Main Outcomes
The main outcome for the review was the estimate of “heritability” identified in the different papers by different methods.
Information Sources
The search was conducted with no language restriction through the electronic databases Medline, Embase, LILACS, Cochrane Library, Open Grey, Google Scholar, and Research Gate up to February 2, 2018, and was complemented by a search through the reference lists of included studies. Narrative or systematic reviews on the topic were searched to identify suitable papers. The search strategy is described in the Appendix. A hand search was conducted of the Journal of Dental Research, Journal of Periodontal Research, Periodontology 2000, Journal of Clinical Periodontology, Journal of Periodontology, and American Journal of Epidemiology. Potentially soon-to-be-published papers were sought by editorial contacts.
Study Selection
Study selection was carried out by 2 independent reviewers (J.B.C. and S.A.A.) in the following stages:
Initial screening of potentially suitable titles and abstracts against the inclusion criteria to identify potentially relevant papers, resulting in a complete database by merging studies included by at least 1 reviewer
Screening of the full papers identified as possibly being relevant in the initial screening
Studies were excluded if they did not meet the inclusion criteria.
The level of agreement between the reviewers was calculated with kappa statistics for the first- and second-stage screening.
Data Collection Process and Data Items
Data were extracted according to the general study characteristics, study design, number of patients included, patients’ demographics, definition and diagnosis of periodontal disease/health, methods for estimation of periodontal disease heritability (phenotypic variance attributable to genetic factors), and method used for genetic analysis (if any).
Risk of Bias in Individual Studies
We aimed to assess the risk of bias of the included studies, as this could have an impact on the overall results and conclusions. The Newcastle-Ottawa Scale was used for case-control studies (twin and family studies), while a scale designed for periodontal genetic association studies (Nibali 2013) was used for GWASs.
Summary Measures, Synthesis of Results, and Statistical Methods
A meta-analysis of the reported or recalculated estimates of heritability from each study was performed separately for twin and family studies with metafor in R. For studies that did not report measures of standard error (SE) of heritability estimates, their confidence intervals were recalculated as previously described with a “combined variance method” (Li et al. 2003). This method calculated the “weighted mean standard deviation” for studies that reported SEs, and it used this mean to estimate SEs for all other studies. A random effects model was used, which accounts for interstudy heterogeneity. In these analyses, weights are assigned according to the inverse of the total variance, comprising the individual study variance and the residual between-study variance. To investigate potential explanations for heterogeneity in estimates across twin studies, random effects metaregression analyses were conducted with R (version 3.3.3). Funnel plots were used to assess publication bias, and Egger’s test was used to test small study effects (P values <0.05 were considered significant).
Results
Study Selection
The study selection and inclusion flowchart are included in the Appendix. The initial search resulted in the following numbers of articles/entries per database: 3,436, PubMed; 7,050, Embase; 190, LILACS; 30, Cochrane Library; 148, Open Grey; and 1, Research Gate. A total of 9,037 remained after elimination of duplicates. A further 10,810 were found by Google Scholar. Following first-stage screening of titles and abstracts, 210 articles qualified for full-text screening (considered potentially suitable by at least 1 reviewer). A further 18 were identified by hand search, and an additional soon-to-be-published paper was found by editorial contact (Kurushima et al. 2018), giving a total of 229 entries to undergo the second screening. After full-text reading, 28 articles met the defined inclusion criteria, and 201 were excluded (reported in the Appendix). The reasons for exclusion were as follows: 148 did not report any estimates of heritability or any measure to calculate it based on the requested study outcomes; 44 were reviews; 5 were case reports; 3 were editorials; and 1 was a letter to the editor. After exclusion from this report of animal studies and a paper reporting only edentulism (Kurushima et al. 2017), 21 papers qualified for inclusion. Every effort was made to obtain any relevant missing data by contacting the authors by email. In particular, corresponding authors of all published GWASs detected in the review search (and reported in Appendix Table 1) were contacted to explore if unpublished heritability estimates were available. Only 2 were able to provide unpublished data on heritability (Feng et al. 2014; Sanders et al. 2017) and were thus included in the study, resulting in a final total of 23 papers.
The kappa value for interreviewer agreement was 0.61 at title and abstract screening and 1.0 at full-text reading, showing substantial agreement between the reviewers.
Study Characteristics
All 23 included papers were written in English. Twelve were conducted in the United States; 2 each in Sweden and India; and 1 each in Canada, Iraq, Guatemala, Japan, Germany, United Kingdom, and Finland. In terms of study design, there were 13 twin studies, 6 family studies, and 4 GWASs. Publication years ranged from 1969 to 2018.
Synthesis of Results
Twin Studies
Table 1 reports the summary of methods and results of twin studies. The number of subjects in the 13 twin studies ranged from 52 (Ciancio et al. 1969) to 21,156 (Mucci et al. 2005). There was some overlap of participants between 2 couples of publications (Michalowicz, Aeppli, Virag, et al. 1991; Michalowicz, Aeppli, Kuba, et al. 1991; Mucci et al. 2005; Mucci et al. 2009). Periodontal disease diagnosis was established by clinical or radiographic examination in most studies, while others relied on self-report (Corey et al. 1993; Mucci et al. 2005; Mucci et al. 2009; Rintakoski et al. 2010; Kurushima et al. 2018). All studies used comparisons between DZ and MZ twins to calculate concordance rates and/or correlations. One of the studies also included comparisons between MZ twins reared together or reared apart (Michalowicz, Aeppli, Virag, et al. 1991; Michalowicz, Aeppli, Kuba, et al. 1991). The concordance and/or correlation was calculated per categorical variables (e.g., self-reported diagnosis of periodontal disease) or continuous variables (e.g., number of probing pocket depths >4 mm). Heritability estimates were usually calculated from concordance rates with Holzinger’s formula, (concordance MZ – concordance DZ) / (1 – concordance DZ), or from correlation rates with Falconer’s (1965) formula, (correlation MZ – correlation DZ) × 2, although some studies reported fully adjusted estimates only (Mucci et al. 2009). We assumed that all studies except GWASs measured broad-sense heritability (H2; Michalowicz et al. 2000), which takes into account dominant as well as additive genetic effects, rather than narrow-sense heritability (h2), which measures only additive effects (Zaitlen and Kraft 2012). When this was not provided, we extrapolated crude heritability estimates based on the data given in the original papers via Falconer’s formula: H2 = 2(rmz – rdz), where H2 is the broad sense heritability, rmz is the identical twin (MZ) correlation, and rdz is the fraternal twin (DZ) correlation.
Table 1.
Summary of Included Human Twin Studies.
| Author (Year): Source | Zygosity Assessment | Method of Diagnosis | Subjects, n | Mean ± SD Age (Range), y | PD Definition | Method for Heritability Estimation | Exclusions | Main Results: Heritability (95% CI) |
|---|---|---|---|---|---|---|---|---|
| Al-Bayaty (2011): Different cities in Iraq | List criteria, blood typing, fingerprint ridge counts | Clinical examination (PPD and AL) on mb surface of Ramfjord teeth for each individual | 75 same-sex twin pairs reared together | 33.7 (16 to 45) | NR | Concordance rates | Systemic diseases or drug intake | PPD concordance: MZ, 60.5%; DZ, 37.8%. AL concordance: MZ, 63.2%; DZ 35.1% |
| Ciancio (1969): NR | 9 blood factors | Clinical examination (6 Ramfjord teeth, 6 sites) | 52 | 15 (12 to 17) | PPD >3 mm | Concordance rates | NR | PPD >3 mm: MZ, 14%; DZ, 21% |
| Corey (1993): Virginia Twin Registry | Zygosity questionnaire | Self-report (questionnaire) | 11,178 (total), 4,908 (PD data available) | MZ, 32.6; DZ, 35.3 | “Have you been affected by periodontal (gum) disease?” | Concordance rates | Incomplete periodontal data | MZ concordance: pairwise, 0.23; proband wise, 0.38. DZ concordance: pairwise, 0.08; proband wise, 0.16 |
| Kurushima (2015): Osaka Twin Research Registry | 15 short-tandem repeat markers | Clinical and radiographic examination (6 sites) | 132 twin pairs (116 MZ and 16 DZ) | 66.1 | PPD, tooth loss, radiographic bone loss | Intraclass correlation, log-likelihood method | NR | Bone resorption: MZ 0.63 (0.48 to 0.74); DZ, 0.68 (0.28 to 0.88). Maximum PPD: MZ, 0.10 (–0.12 to 0.30); DZ, 0.11 (–0.48 to 0.65). All nonsignificant |
| Kurushima (2018): TwinsUK registry | Questionnaire and DNA short-tandem repeat fingerprinting | Self-report (questionnaire) | 1,188 MZ and 934 DZ twin pairs (for GI outcome, slightly less for PD) | 57.2 (20 to 91) | Five questionsa | Intraclass correlation | Male sex | Gingivitis heritability, 24%; periodontitis heritability, 30% |
| Lovelina (2012): Ragas Dental College, India | Medical records, dermatoglyphics, chorionicity, and number of placental cords | Clinical examination (WHO Oral Health Assessment) | 30 twin pairs (9 MZ, 21 DZ) reared together | 18.5 ± 5.5 (13 to 24) | Community Periodontal Index | Pearson’s correlation coefficient | Smoking, MH, ongoing orthodontic treatment, malocclusion | Correlation: 0.89 for MZ, 0.14 for DZ |
| Michalowicz, Aeppli, Virag, (1991): Minnesota Twin Family Registry + Minnesota Study of Twins Reared Apart | 4 red blood cell antigens, 6 serum proteins, and 4 red blood cell enzymes; anthropomorphic measures | Clinical examination (mb or ml surface of the Ramfjord teeth) | 110 (63 MZ reared together, 14 MZ reared apart, 33 same-sex DZ pairs) | 40.3 (16 to 70) | AL, PPD, GI, PI | Intraclass correlation | 1 twin with <5 Ramfjord teeth; zygosity not confirmed | Heritability: PPD, 0.51 (0.12 to 0.68); AL, 0.48 (0.21 to 0.71); GI, 0.40 (–0.17 to 0.66) |
| Michalowicz, Aeppli, Kuba, (1991): Minnesota Twin Family Registry + Minnesota Study of Twins Reared Apart | 14 blood markers and 3 anthropomorphic measures. | Radiographic measurements of BL | 120 pairs (62 MZ reared together, 33 MZ reared apart, 25 same-sex DZ pairs) | 40.4 (16 to 80) | BL | Intraclass correlation | <10 teeth could be evaluated radiographically | Heritability for BL: 0.36 (–0.15 to 0.77); MZ reared apart, 0.55 (0.35 to 0.74) |
| Michalowicz (2000): Virginia Twin Registry | Questionnaire (+ DNA fingerprinting for 87 pairs) | Clinical examination | 117 pairs (64 MZ, 53 DZ) | MZ, 41.2; DZ, 42.3 | Percentage of teeth with ≥1 sites with AL, 2 to 3 mm; PPD, 4 to 5 mm | Intraclass correlation | <10 teeth could be evaluated radiographically | Heritability: AL (fully adjusted), 43%; AL (≥2 mm), 59%; AL (≥3 mm), 50%; PD, 45%; PPDs ≥ 4 mm, 50%; GI, 0% |
| Mucci (2005): Swedish Twin Registry (1998 and 2002) | Questionnaire | Telephone interviews | 10,578 pairs (2,747 MZ, 7,831 DZ) | Minimum, 42 | PD: Mobile teeth or diagnosed by dentist. Gingivitis: bleeding gums | Tetrachoric correlation, concordance rates, log-likelihood method | Edentulous twins | PD, 0.39 (0.31 to 0.47); gingivitis, 0.36 (0.28 to 0.43) |
| Mucci (2009): Swedish Twin Registry | Questionnaire | Self-report | 15,273 | 35 (then followed up for 35) | Mobility in at least half of the teeth | Cholesky’s decomposition | Prevalent CVD in 1963, missing follow-up data or information on smoking or covariates | Risk of PD: h2 = 0.42 |
| Rintakoski (2010): Fourth wave, FinnTwin16 Study (2000 to 2002) | Questionnaire | Oral Health questionnaire | 3,065 pairs | 24.6 (23 to 27) | Gingival bleeding | Polychoric correlations | NR | Additive genetic effect for bleeding: 32% (0.22 to 0.41) |
| Sharma (2007): NR | A1A2BO, Rh MN, Kell, and Duffy blood groups; ABH secretion, PTC tasting ability, dermatoglyphics, and photographs | Clinical examination (WHO probe) | 57 pairs: 26 MZ and 31 DZ twin pairs | 18.2 ± 4.2, MZ; 16.9 ± 4.1, DZ (11 to 28) | WHO 1987 | Holzinger’s 1929 Heritability Estimate | NR | Heritability for gingivitis: 0.82 |
Where not provided, heritability was calculated by Falconer’s formula: Hb2 = 2(rmz – rdz), where Hb2 is the broad-sense heritability, rmz is the identical (MZ) twin correlation, and rdz is the fraternal (DZ) twin correlation.
AL, attachment loss; BL, bone loss; CVD, cardiovascular disease; DZ, dizygous; F, female; GI, gingival index; M, male; mb, mesiobuccal; MH, medical history; ml, mesiolingual; MZ, monozygous; NR, not reported; NS, not statistically significant; PD, periodontitis; PI, plaque index; PPD, probing pocket depth; PTC, Phenylthiocarbamide; WHO, World Health Organization.
“Have you had gum bleeding?” “Do your gums bleed when you brush your teeth?” “Have you ever had gum decay or loose teeth?” “Do you have any loose teeth?” “Have you ever been told by a dentist or hygienist that you have gum disease?”
Family Studies
Table 2 reports the summary of family studies and GWASs. The number of subjects in the 6 family studies ranged between 57 and 959. Two papers reported data from the same population (Chung et al. 1977; Rao et al. 1979), so only 1 was used for meta-analyses. There was possible overlap between 2 more publications (Diehl et al. 2003; Diehl et al. 2005). Periodontal disease diagnosis was established by clinical examination in all studies, and continuous measures (e.g., average clinical attachment loss [CAL]) were usually reported. In 2 studies, this was restricted to aggressive periodontitis (Diehl et al. 2003; Diehl et al. 2005). Heritability estimates were calculated by comparing affected and unaffected family members and analyzing data with various statistical methods.
Table 2.
Summary of Included Family Studies and Genome-wide Association Studies.
| Author (Year): Country | Method of Diagnosis | Subjects, n | Age, y | PD Definition | PD Prevalence | Method for Heritability Estimation | Heritability Estimate |
|---|---|---|---|---|---|---|---|
| Human family studies | |||||||
| Beaty (1993): US | Clinical examination (4 sites per tooth) | 178 (75 families) | <40, 69%; <20, M21 + F26; 21 to 40, M24 + F52; 41 to 60, M16 + F30; >60, M4 + F5 | Continuous variable (measured PI, GI, AL) | NR | Analysis of variance + standard familial correlations, log-likelihood method | 31% of variance for GI, 29% for AL, and 34% for PI due to familial factors (including genetics) |
| Chung (1977), Rao (1979): US | Clinical examination on Ramfjord teeth | 939 (241 families) | Range, 20 to ≥61: 20 (n = 404), 21 to 30 (n = 38), 31 to 40 (n = 132), 41 to 50 (n = 268), 51 to 60 (n = 76), ≥61 (n = 21) | Index including gingivitis, PPD, calculus, contact, attrition, mobility, and plaque | NR | Path analysis | 0.22 (SE, 0.20). In Rao paper: 0.22 ± 0.14 in children, 0.06 ± 0.09 in adults |
| Diehl (2003): US | Clinical examination (4 sites per tooth) | 274 (60 families) | NR | AAP 1999 | 79 healthy, 51 LAgP, 83 GAgP, 55 CP, 6 edentulous | Quantitative genetic analysis: SOLAR | All teeth 0.17 (0.11). First molars, 0.29 (0.14). Smoking with AgP, 0.29 (0.11) |
| Diehl (2005): US | Clinical examination (4 sites per tooth) | 959 (heritability data based on 288 individuals) | NR | AAP 1999 | 610 total: 237 healthy, 169 LAgP, 204 GAgP (others not classified as <13 y, >35 y, or had CP) | Quantitative genetic analysis: SOLAR | All teeth 0.16 |
| Dowsett (2002): Guatemala | Clinical examination (6 sites per tooth) | 113 (40 families) | Mean, 48.2 (range, 35 to 60) | PD: AL ≥5 mm; GI, >50% sites with BOP | Subjects with ≥50% bleeding = 84% (gingivitis). Subjects with AL ≥5 mm = 84% | Intraclass correlation | AL = 0.31 (−0.09 to 0.78). GI = 0.03 (−0.20 to −0.48) |
| Genome-wide association studies | |||||||
| Divaris (2013): US | Clinical assessment | 4,504 | Mean, 62 (range, 53 to 74) | AAP 2007 criteria | 43%, moderate CP; 17%, severe CP | GCTA: first, all available SNPs used to estimate a genetic relationship matrix among all study participants. Then, genetic relationship matrix used in restricted maximum likelihood analyses to estimate the proportion of variance explained by all SNPs | Moderate CP, 0.00 (SE, 0.14); severe CP, 0.22 (SE, 0.19); moderate CP–smoking, 0.36 (SE, 0.26); severe CP–smoking, 0.51 (SE, 0.35) (all adjusted for age and sex) |
| Teumer (2013): Germany | Clinical assessment | 4,032 | Mean, 46 and 50 in the 2 independent samples | AAP 2007 criteria | 35.3% and 37.0%, moderate CP; 19.9% and 16.6%, severe CP | GCTA (as above) | 0.14 (SE, 0.09) for moderate-severe CP, 0.23 (SE, 0.08) for AL (continuous variable) (all adjusted for age, sex, and smoking); including smoking interaction, 0.20 (SE, 0.17) for moderate-severe CP and 0.30 (SE, 0.16) for AL |
| Feng (2014)a: US | Dental records | 866 + 1,819 for replication | Mean, 41 | ≥30% of teeth with AL ≥5 mm | 11.4% of original sample | GCTA (as above) | 0.24 (SE, 1.80) adjusted for sex |
| Sanders (2017)a: US | Clinical assessment | 10,935 | Mean, 45 | Mean interproximal AL | NR | Haseman-Elston regression method | 0.04 (95% CI, 0 to 0.11) for mean interproximal AL |
AAP, American Academy of Periodontology; AgP, aggressive periodontitis; AL, attachment loss; BOP, bleeding on probing; CP, chronic periodontitis; F, female; GAgP, generalized aggressive periodontitis; GCTA, genome-wide complex trait analysis; GI, gingival index; LAgP, localized aggressive periodontitis; M, male; NR, not reported; PD, periodontitis; PI, plaque index; PPD, probing pocket depth; SNP, single-nucleotide polymorphism.
Heritability not found in the published paper but separately provided by the authors.
Genome-wide Association Studies
The number of subjects in the 4 included studies ranged from 866 (Feng et al. 2014) to 10,935 (Sanders et al. 2017). In 2 GWASs, the heritability estimate was reported in the published paper (Divaris et al. 2013; Teumer et al. 2013). Both studies used the same genome-wide complex trait analysis (GCTA) software and the same disease classification (Page and Eke 2007) for estimation of heritability. However, 1 study reported heritability estimates separately for moderate and severe periodontitis (Divaris et al. 2013) and the other for moderate and severe periodontitis combined versus health and for mean proximal attachment loss (Teumer et al. 2013). For the other 2 studies, heritability estimates were obtained from the authors, using the GCTA software (Feng et al. 2014) and the Haseman-Elston regression method (Sanders et al. 2017) as described before (Sofer 2017).
Heritability Estimates
Heritability estimates (H2) across studies for the different parameters analyzed ranged widely from 0 (concordance rates higher in DZ than MZ twins; Kurushima et al. 2015) to 1 (concordance rates more than double for MZ than DZ twins; Lovelina et al. 2012). Studies were subgrouped for meta-analyses of the heritability estimate.
Gingivitis
Heritability for the self-reported gingivitis trait, including 7 papers of twins (Mucci et al. 2005; female and male results reported separately), was estimated at 0.29 (95% CI, 0.21 to 0.38) with moderate heterogeneity (I2 = 37.6%, P = 0.13). However, when only clinically measured gingivitis was included (3 studies), heritability was not statistically significant (0.32; 95% CI, −0.24 to 0.87), with moderate heterogeneity (I2 = 42.5%, P = 0.19; see forest plots in Fig. 1).
Figure 1.
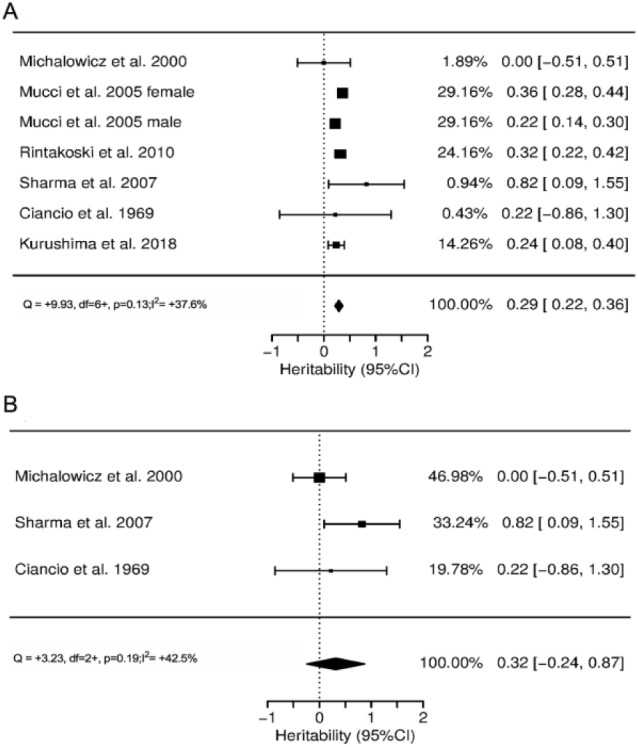
Forest plots of heritability estimates for the gingivitis trait: studies with (a) self-report and (b) clinical measurement.
Periodontitis
The forest plots for heritability of the periodontitis trait in twin and other family studies are presented in Figure 2. In twin studies, heritability for periodontitis (various definitions, 9 studies) was 0.38 (95% CI, 0.34 to 0.43) with low heterogeneity (I2 = 12.9%, P = 0.54; Fig. 2a). This estimate remained relatively unchanged when only studies with confirmed periodontitis were included (excluding self-reported periodontitis, 6 studies): the H2 was 0.34 (95% CI, 0.10 to 0.59) with low heterogeneity (I2 = 14.2%, P = 0.32; Fig. 2b). In family studies, a meta-analysis was conducted for periodontitis (4 studies) showing an H2 of 0.15 (95% CI, 0.06 to 0.24) with low heterogeneity (I2 = 0%, P = 0.77; Fig. 2c). When twin and family studies were combined, heritability for periodontitis was estimated at 0.29 (95% CI, 0.21 to 0.38) with high heterogeneity (I2 = 61.2%, P = 0.01; Appendix Figure 1).
Figure 2.
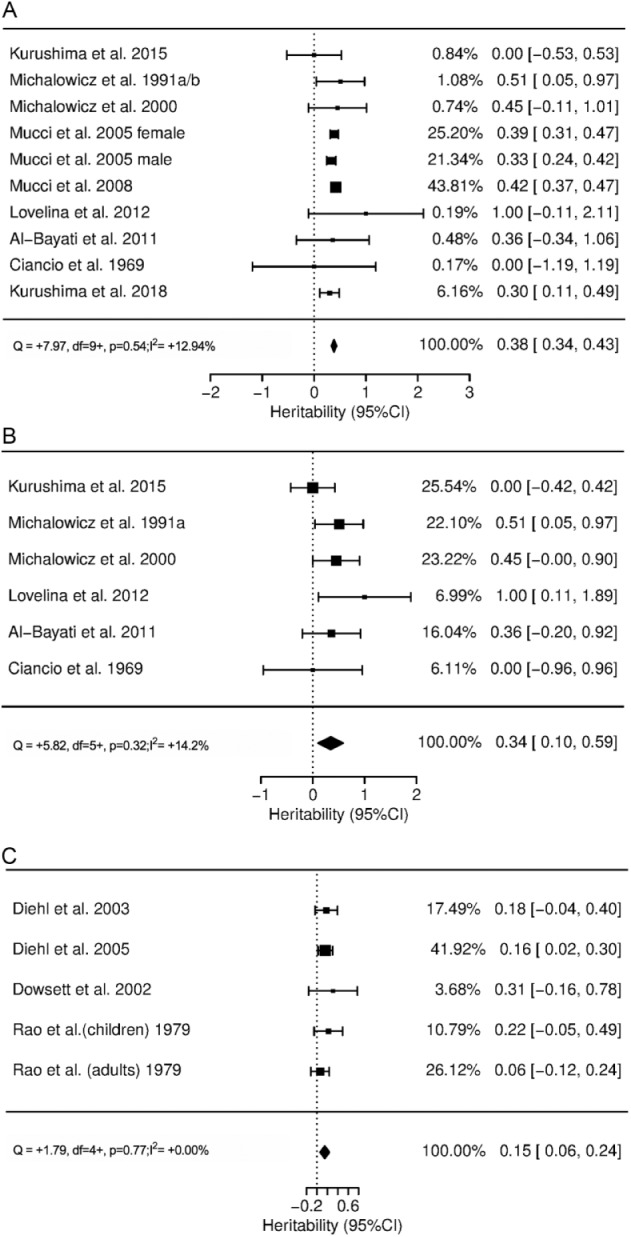
Forest plots of heritability estimates for the periodontitis trait: (a) all twin studies combined; (b) twin studies, excluding those where periodontitis was self-reported; and (c) other family studies.
When mean attachment loss of twins was considered as the outcome (3 studies), H2 was 0.45 (95% CI, 0.25 to 0.64) with low heterogeneity (I2 = 0%, P = 0.97; see Appendix Figure 2).
In the GWAS context, Divaris et al. (2013) reported adjusted estimates of 0.00 (SE, 0.14) and 0.22 (SE, 0.19) for moderate and severe chronic periodontitis (CP), respectively. The German study reported adjusted estimates of 0.14 (SE, 0.09) for moderate-severe CP and 0.23 (SE, 0.08) for attachment loss (continuous variable; Teumer et al. 2013). Both papers reported higher heritability estimates when the interaction with smoking was included in the model: 0.36 (SE, 0.26) for moderate CP and 0.51 (SE, 0.35) for severe CP in the US study (Divaris et al. 2013) and 0.20 (SE, 0.17) for moderate-severe CP and 0.30 (SE, 0.16) for attachment loss in the German study (Teumer et al. 2013). Heritability for mean interproximal attachment loss was calculated as 0.04 (95% CI, 0 to 0.11) by Sanders et al. (2007). Feng et al. (2014) obtained a heritability value of 0.24 (SE, 1.80), adjusted for sex, for periodontitis defined as ≥30% of teeth with attachment loss ≥5 mm. In the absence of common definitions of periodontitis in the 4 included GWASs, meta-analysis was conducted for continuous measures of mean attachment loss (Teumer et al. 2013; Sanders et al. 2017) combined with severe/generalized periodontitis (Divaris et al. 2013; Feng et al. 2014; see Fig. 3), resulting in an h2 of 0.07 (95% CI, −0.02 to 0.15) with low heterogeneity (I2 = 14.7%, P = 0.57).
Figure 3.
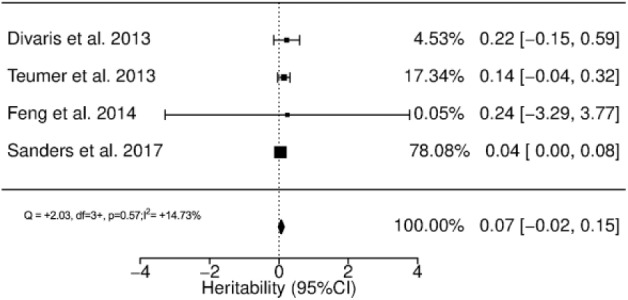
Forest plot of heritability estimates for genome-wide association studies.
Metaregression for age and h2 showed that the association between age and heritability was not statistically significant, although a trend for an inverse association was observed between heritability and age for periodontitis and gingivitis (Appendix Figure 3).
Risk-of-Bias Assessment
Risk-of-bias analyses are presented in Appendix Table 2. Twin studies scored from 4 to 9 and family studies from 6 to 9 in the Newcastle-Ottawa Scale (maximum, 9). GWASs scored 15 to 17 in the specific genetic periodontal scale (maximum, 20; Nibali 2013). Funnel plots of meta-analysis (Appendix Figure 4) appeared generally symmetrical, showing no evidence of publication bias. This was confirmed by Egger’s test (all tests P > 0.05).
Discussion
This systematic search of the literature, including twin and family studies, yielded a total reviewed sample of >50,000 participants and showed that approximately up to one-third of the variance of periodontitis trait in human populations is attributable to heritable factors (0.38 for all twin studies, 0.34 for twin studies excluding self-reported periodontitis outcomes, 0.15 for family studies, and 0.29 for all twin and family studies combined but with high heterogeneity). The estimates from twin studies are derived by comparing the prevalence of disease between MZ twins and DZ twins. This approach aims to reduce confounding due to environmental factors (including intrauterine factors) and assumes that MZ twins share 100% of the genome while DZ twins share approximately half the genome. Therefore, heritability of disease should show higher concordance (e.g., both twins being diseased or healthy) between MZ and DZ twins. Only 2 studies included the different model of MZ twins reared together and reared apart (Michalowicz, Aeppli, Virag, et al. 1991; Michalowicz, Aeppli, Kuba, et al. 1991), which could give more information on the effect of environmental factors on identical subjects, since phenotypic similarities in reared-apart MZ twins are thought to be exclusively attributable to shared genes as well as to prenatal environment but not to postnatal environmental factors. The heritability estimates for reared-apart MZ twins in the studies by Michalowicz et al. (0.38 for CAL, 0.55 for alveolar bone loss, and 0.82 for gingival index) confirm the important role of genetic factors in periodontal disease but also suggest that genetic-environmental interactions probably enhance the similarity of MZT twins.
The lower heritability found in GWASs versus other study designs, ranging from 0 for moderate CP (Divaris et al. 2013) to 0.14 for moderate-severe CP (Teumer et al. 2013) and 0.23 to 0.24 for severe CP (Divaris et al. 2013; Feng et al. 2014), was close to the lower-bound confidence interval of the twin studies. Meta-analysis for this outcome resulted in an h2 of 0.07. This lower heritability value may have been affected by the highly heterogeneous definitions of periodontitis across studies. These ranged from continuous measures of mean attachment loss in the largest study with stronger weight in the meta-analysis results (Sanders et al. 2017) to categorical measures of periodontitis, such as ≥30% of teeth with attachment loss ≥5 mm, which resulted in higher heritability estimates (Feng et al. 2014). This was expected owing to the so-called missing heritability, including the role of rare variants, gene-environment interactions, and epigenetics, which are not captured in GWASs but may be represented in the heritability obtained in twin studies. In particular, GWASs usually do not account for rare variants (>1% or 5% minor allele frequency) and may only reflect the effect of a limited range of gene variants as compared with twin studies (Zaitlen and Kraft 2012). In other words, GWASs may estimate “bottom-up,” narrow-sense (rather than broad-sense) heritability, ignoring dominance and interaction effects (Zaitlen and Kraft 2012). The fact that severe periodontitis showed higher heritability than moderate periodontitis is also expected. Heritability in family studies is lower than in twin studies, probably due to the methods employed and to the generally smaller sample size—and hence larger confidence intervals—in family studies. The figure of one-third heritability is similar to what was reported in animal studies (Nashef et al. 2018). Heritability studies alone are not sufficient to demonstrate a genetic component in periodontitis, because shared behavior and other environmental factors can contribute to covariance among relatives and mimic genetic correlation. However, these results may suggest that the shared household has limited impact on the measured periodontitis heritability in humans.
Interestingly, inclusion of interactions with smoking considerably increased the obtained heritability estimates in the GWAS context: inclusion of a genome-smoking interaction term increased the accounted variance of severe periodontitis to half (Divaris et al. 2013) and all periodontitis to 0.30 (Teumer et al. 2013). Although these results can be interpreted as the consequence of a gene-environment interaction model, one could also interpret them as a compound heritability model, in which the heritability of being dependent to cigarette smoking is added to the heritability of severe periodontitis.
Experimental gingivitis models have shown different degrees and timing of gingival inflammation upon microbial colonization (Loe et al. 1965), suggesting an element of individual susceptibility to the gingivitis trait. Heritability of gingivitis in humans was estimated at 0.29 in the current meta-analysis. However, a substantial proportion of this figure was derived by 2 studies (Mucci et al. 2005; Rintakoski et al. 2010) where gingivitis was defined by self-report of “bleeding gums,” which may lack reliability and may be affected by oral hygiene regimes, smoking, and the potential presence of periodontitis (rather than just gingivitis). When self-reported studies were excluded, heritability of gingivitis was no longer statistically significant, largely due to a study where heritability of gingivitis was 0 after adjustment for confounders (Michalowicz et al. 2000). Therefore, heritability of gingivitis requires further investigation.
Different settings, sample sizes, study designs, and methods of examination and definition of the periodontal phenotype likely explain some of the differences among studies. However, very low heterogeneity was detected in most meta-analyses performed, suggesting that these data can be accepted with reasonable confidence. The only exception was when twin and other family studies were combined. The average age of the examined individuals seemed to show a nonstatistically significant inverse trend with heritability estimates, consistent with a postulated greater role played by environmental factors among those who develop periodontitis late in life, as previously observed for diabetes (Almgren et al. 2011). This is supported by the higher heritability detected in GWASs when only younger subjects were included (Teumer et al. 2013).
Although studies were carried out in the United States, Europe, and Asia, few reported on the racial/ethnic makeup of the participating individuals. Therefore, no inference can be made on the heritability among different ethnic groups. Some studies reported differences in heritability between male and female subcohorts. However, while Mucci et al. (2005) reported higher heritability rates for females for gingivitis, Corey et al. (1993) reported higher heritability for periodontitis in males versus females. Therefore, again, no clear conclusion can be made on sex-associated heritability estimates. The risk of bias across studies was low overall, with no specific areas found to be associated with high risk of bias.
This systematic review confirms that
a substantial proportion of the phenotypic variance of periodontitis in the population is due to genetics;
heritability tends to be higher for severe early-onset traits and younger individuals; and
heritability is significantly increased when genome-wide interactions with smoking are accounted for.
The limitations of this review are the potential ascertainment bias of some of the studies (especially self-reported) and the nonexact assumptions made to estimate heritability in the different studies. Strengths include the novelty, the low heterogeneity of the meta-analyses, and the inclusion of a large sample size.
These findings could help to set boundaries on the potential of genetic variants to explain disease, to inform genetic risk estimation, and to ultimately lead to a better understanding of disease pathogenesis. Heritability for periodontal disease may have common pathways to other inflammatory/chronic diseases. Hence, associations between periodontitis and diseases of other systems (e.g., cardiovascular disease; see Mucci et al. 2009) may be detected in cohorts that happen to share similar inflammatory responses. Twin studies performed of other chronic diseases suggest a considerable heritability of type 2 diabetes (Almgren et al. 2011) and a range of heritability, from weak in rheumatoid arthritis and systemic sclerosis to high in celiac disease and psoriasis (Generali et al. 2017). This suggests possible pleiotropy, with certain genes appearing to affect ≥2 unrelated phenotypic traits (Loos et al. 2015; Chapple et al. 2017).
It is important to stress that heritability is a concept related to relative contribution in a population, meaning that if certain environmental factors in a population considerably increase the risk of disease, then the relative contribution of genetics would be smaller. As such, no direct inference can be derived in terms of treatment of the single patient. However, although no clear evidence for this has been produced yet, it is reasonable to assume that, as with other diseases, such as breast cancer (Escala-Garcia et al. 2019), as well as increased susceptibility to periodontitis, genetic variants may affect treatment response. For example, individuals who respond less favorably to treatment (due to their accentuated genetic predisposition) could benefit from more intense therapeutic regimes or more frequent recalls, which may require changes in policy related to reimbursements and will affect important stakeholder practices. Since the search for the specific genomic loci where this heritability lies has so far proved quite elusive (Nibali et al. 2017), collaborative efforts, including consortium meta-analyses of several studies and hundreds of thousands of individuals, and examination of “refined” biologically informed traits (which may be more homogeneous) can help elucidate the biological basis and heritability of periodontal disease.
Author Contributions
L. Nibali, contributed to conception, design, data analysis, and interpretation, drafted and critically revised the manuscript; J. Bayliss-Chapman, S.A. Almofareh, contributed to design, data acquisition, and interpretation, critically revised the manuscript; Y. Zhou, contributed to data analysis and interpretation, critically revised the manuscript; K. Divaris, A.R. Vieira, contributed to design, data analysis, and interpretation, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
Supplemental material, DS_10.1177_0022034519842510 for What Is the Heritability of Periodontitis? A Systematic Review by L. Nibali, J. Bayliss-Chapman, S.A. Almofareh, Y. Zhou, K. Divaris and A.R. Vieira in Journal of Dental Research
Footnotes
The authors received no financial support and declare no potential conflicts of interest with respect to the authorship and/or publication of this article.
A supplemental appendix to this article is available online.
References
- Al-Bayaty FHMH, AL-Fatlawi ZM, Mirza KB, Abdulla MA. 2011. Prevalence and severity of periodontal disease among Iraqi twin population. Sci Res Essays. 6(5):1034–1038. [Google Scholar]
- Almgren P, Lehtovirta M, Isomaa B, Sarelin L, Taskinen MR, Lyssenko V, Tuomi T, Groop L; Botnia Study Group. 2011. Heritability and familiality of type 2 diabetes and related quantitative traits in the Botnia Study. Diabetologia. 54(11):2811–2819. [DOI] [PubMed] [Google Scholar]
- Beaty TH, Colyer CR, Chang YC, Liang KY, Graybeal JC, Muhammad NK, Levin LS. 1993. Familial aggregation of periodontal indices. J Dent Res. 72(2):544–551. [DOI] [PubMed] [Google Scholar]
- Chapple IL, Bouchard P, Cagetti MG, Campus G, Carra MC, Cocco F, Nibali L, Hujoel P, Laine ML, Lingstrom P, et al. 2017. Interaction of lifestyle, behaviour or systemic diseases with dental caries and periodontal diseases: consensus report of group 2 of the joint EFP/ORCA workshop on the boundaries between caries and periodontal diseases. J Clin Periodontol. 44 Suppl 18:S39–S51. [DOI] [PubMed] [Google Scholar]
- Chung CS, Kau MC, Chung SS, Rao DC. 1977. A genetic and epidemiologic study of periodontal disease in Hawaii: II. Genetic and environmental influence. Am J Hum Genet. 29(1):76–82. [PMC free article] [PubMed] [Google Scholar]
- Ciancio SG, Hazen SP, Cunat JJ. 1969. Periodontal observations in twins. J Periodontal Res. 4(1):42–45. [DOI] [PubMed] [Google Scholar]
- Corey LA, Nance WE, Hofstede P, Schenkein HA. 1993. Self-reported periodontal disease in a Virginia twin population. J Periodontol. 64(12):1205–1208. [DOI] [PubMed] [Google Scholar]
- Diehl SR, Wu T, Burmeister JA, Califano JV, Brooks CN, Tew JG, Schenkein HA. 2003. Evidence of a substantial genetic basis for IgG2 levels in families with aggressive periodontitis. J Dent Res. 82(9):708–712. [DOI] [PubMed] [Google Scholar]
- Diehl SR, Wu T, Michalowicz BS, Brooks CN, Califano JV, Burmeister JA, Schenkein HA. 2005. Quantitative measures of aggressive periodontitis show substantial heritability and consistency with traditional diagnoses. J Periodontol. 76(2):279–288. [DOI] [PubMed] [Google Scholar]
- Divaris K. 2017. Fundamentals of precision medicine. Compend Contin Educ Dent. 38(8 Suppl):30–32. [PMC free article] [PubMed] [Google Scholar]
- Divaris K, Monda KL, North KE, Olshan AF, Reynolds LM, Hsueh WC, Lange EM, Moss K, Barros SP, Weyant RJ, et al. 2013. Exploring the genetic basis of chronic periodontitis: a genome-wide association study. Hum Mol Genet. 22(11):2312–2324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dowsett SA, Archila L, Foroud T, Koller D, Eckert GJ, Kowolik MJ. 2002. The effect of shared genetic and environmental factors on periodontal disease parameters in untreated adult siblings in Guatemala. J Periodontol. 73(10):1160–1168. [DOI] [PubMed] [Google Scholar]
- Escala-Garcia M, Guo Q, Dörk T, Canisius S, Keeman R, Dennis J, Beesley J, Lecarpentier J, Bolla MK, Wang Q, et al. 2019. Genome-wide association study of germline variants and breast cancer-specific mortality. Br J Cancer. 120(6):647–657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falconer DS. 1965. The inheritance of liability to certain diseases estimated from incidence among relatives. Ann Hum Genet. 29(1):51–76. [Google Scholar]
- Feng P, Wang X, Casado PL, Küchler EC, Deeley K, Noel J, Kimm H, Kim JH, Haas AN, Quinelato V, et al. 2014. Genome wide association scan for chronic periodontitis implicates novel locus. BMC Oral Health. 14:84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Generali E, Ceribelli A, Stazi MA, Selmi C. 2017. Lessons learned from twins in autoimmune and chronic inflammatory diseases. J Autoimmun. 83:51–61. [DOI] [PubMed] [Google Scholar]
- Hiyari S, Atti E, Camargo PM, Eskin E, Lusis AJ, Tetradis S, Pirih FQ. 2015. Heritability of periodontal bone loss in mice. J Periodontal Res. 50(6):730–736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinnersley B, Mitchell JS, Gousias K, Schramm J, Idbaih A, Labussière M, Marie Y, Rahimian A, Winchmann HE, Schreiber S, et al. 2015. Quantifying the heritability of glioma using genome-wide complex trait analysis. Sci Rep. 5:17267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurushima Y, Ikebe K, Matsuda K, Enoki K, Ogata S, Yamashita M, Murakami S, Hayakawa K, Maeda Y. 2015. Influence of genetic and environmental factors on oral diseases and function in aged twins. J Oral Rehabil. 42(1):49–56. [DOI] [PubMed] [Google Scholar]
- Kurushima Y, Silventoinen K, Dokkedal U, Skytthe A, Mucci LA, Christensen K, Hjelmborg JVB. 2017. Heritability of the number of teeth in middle-aged and older Danish twins. J Dent Res. 96(13):1513–1517 [DOI] [PubMed] [Google Scholar]
- Kurushima Y, Bowyer R, Ide M, Hughes FJ, Steves CJ. 2018. Genetic and environmental contributions to the association between mood disorder and periodontal disease: a cross-sectional study amongst female twins in the UK. J Clin Periodontol. 46(1):40–50. [DOI] [PubMed] [Google Scholar]
- Li Ming D, Ma JZ, Swan GE. 2003. A meta-analysis of estimated genetic and environmental effects on smoking behavior in male and female adult twins. Addiction. 98(1):23–31. [DOI] [PubMed] [Google Scholar]
- Loe H, Theilade E, Jensen SB. 1965. Experimental gingivitis in man. J Periodontol. 36:177–187. [DOI] [PubMed] [Google Scholar]
- Loos BG, Papantonopoulos G, Jepsen S, Laine ML. 2015. What is the contribution of genetics to periodontal risk? Dent Clin North Am. 59(4):761–780. [DOI] [PubMed] [Google Scholar]
- Lovelina FD, Shastri SM, Kumar PD. 2012. Assessment of the oral health status of monozygotic and dizygotic twins—a comparative study. Oral Health Prev Dent. 10(2):135–139. [PubMed] [Google Scholar]
- Michalowicz BS, Aeppli D, Virag JG, Klump DG, Hinrichs JE, Segal NL, Bouchard TJ, Jr, Pihlstrom BL. 1991. Periodontal findings in adult twins. J Periodontol. 62(5):293–299. [DOI] [PubMed] [Google Scholar]
- Michalowicz BS, Aeppli DP, Kuba RK, Bereuter JE, Conry JP, Segal NL, Bouchard TJ, Jr, Pihlstrom BL. 1991. A twin study of genetic variation in proportional radiographic alveolar bone height. J Dent Res. 70(11):1431–1435. [DOI] [PubMed] [Google Scholar]
- Michalowicz BS, Diehl SR, Gunsolley JC, Sparks BS, Brooks CN, Koertge TE, Califano JV, Burmeister JA, Schenkein HA. 2000. Evidence of a substantial genetic basis for risk of adult periodontitis. J Periodontol. 71(11):1699–1707. [DOI] [PubMed] [Google Scholar]
- Miley DD, Baumgartner MH, Cheverud JM, Roseman CC, Rogers J, McLeod DE, Reyes E, Hildebolt CF. 2011. Heritability of alveolar bone loss from periodontal disease in a baboon population: a pilot study. J Periodontol. 82(4):575–580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moher D, Liberati A, Tetzlaff J, Altman DG. 2009. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Ann Int Med. 151(4):264–269, W264. [PMC free article] [PubMed] [Google Scholar]
- Mucci LA, Bjorkman L, Douglass CW, Pedersen NL. 2005. Environmental and heritable factors in the etiology of oral diseases: a population-based study of Swedish twins. J Dent Res. 84(9):800–805. [DOI] [PubMed] [Google Scholar]
- Mucci LA, Hsieh CC, Williams PL, Arora M, Adami HO, de Faire U, Douglass CW, Pedersen NL. 2009. Do genetic factors explain the association between poor oral health and cardiovascular disease? A prospective study among Swedish twins. Am J Epidemiol. 170(5):615–621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nashef A, Qabaja R, Salaymeh Y, Botzman M, Munz M, Dommisch H, Krone B, Hoffmann P, Wellmann J, Laudes M, et al. 2018. Integration of murine and human studies for mapping periodontitis susceptibility. J Dent Res. 97(5):537–546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nibali L. 2013. Suggested guidelines for systematic reviews of periodontal genetic association studies. J Clin Periodontol. 40(8):753–756. [DOI] [PubMed] [Google Scholar]
- Nibali L, Di Iorio A, Tu YK, Vieira AR. 2017. Host genetics role in the pathogenesis of periodontal disease and caries. J Clin Periodontol. 44 Suppl 18:S52–S78. [DOI] [PubMed] [Google Scholar]
- Page RC, Eke PI. 2007. Case definitions for use in population-based surveillance of periodontitis. J Periodontol. 78(7s):1387–1399. [DOI] [PubMed] [Google Scholar]
- Rao DC, Chung CS, Morton NE. 1979. Genetic and environmental determinants of periodontal disease. Am J Med Genet. 4(1):39–45. [DOI] [PubMed] [Google Scholar]
- Rintakoski K, Kaprio J, Murtomaa H. 2010. Genetic and environmental factors in oral health among twins. J Dent Res. 89(7):700–704. [DOI] [PubMed] [Google Scholar]
- Sanders AE, Sofer T, Wong Q, Kerr KF, Agler C, Shaffer JR, Beck JD, Offenbacher S, Salazar CR, North KE, et al. 2017. Chronic periodontitis genome-wide association study in the Hispanic Community Health Study/Study of Latinos. J Dent Res. 96(1):64–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma K. 2007. Genetic susceptibility to periodontal diseases: a twin study. In: Sharma K, Pathak RK, Shalina, et al., editors. Genes, environment and health: anthropological perspectives. New Delhi (India): Serials Pub, p. 198–203. [Google Scholar]
- Smith C. 1971. Recurrence risks for multifactorial inheritance. Am J Hum Genet. 23(6):578–588. [PMC free article] [PubMed] [Google Scholar]
- Sofer T. 2017. Confidence intervals for heritability via Haseman-Elston regression. Stat Appl Genet Mol Biol. 16(4):259–273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tenesa A, Haley CS. 2013. The heritability of human disease: estimation, uses and abuses. Nat Rev Genet. 14(2):139–149. [DOI] [PubMed] [Google Scholar]
- Teumer A, Holtfreter B, Völker U, Petersmann A, Nauck M, Biffar R, Volzke H, Kroemer HK, Meisel P, Homuth G, et al. 2013. Genome-wide association study of chronic periodontitis in a general German population. J Clin Periodontol. 40(11):977–985. [DOI] [PubMed] [Google Scholar]
- Visscher PM, Hill WG, Wray NR. 2008. Heritability in the genomics era—concepts and misconceptions. Nature Rev Genet. 9(4):255–266. [DOI] [PubMed] [Google Scholar]
- Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, Madden PA, Heath AC, Martin NG, Montgomery GW, et al. 2010. Common SNPs explain a large proportion of the heritability for human height. Nat Genet. 42(7):565–569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaitlen N, Kraft P. 2012. Heritability in the genome-wide association era. Hum Genet. 131(10):1655–1664. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material, DS_10.1177_0022034519842510 for What Is the Heritability of Periodontitis? A Systematic Review by L. Nibali, J. Bayliss-Chapman, S.A. Almofareh, Y. Zhou, K. Divaris and A.R. Vieira in Journal of Dental Research


