Abstract
Purpose: EGFR and anaplastic lymphoma kinase (ALK) alterations have been regarded as oncogenic drivers and incorporated into clinical practices to manage nonsmall cell lung cancer (NSCLC). Alterations of these two genes were traditionally considered to be mutually exclusive, but recent studies have suggested that they can occur concomitantly. Here, we investigated the prevalence, clinical features and outcomes in response to the treatment of NSCLC patients who harbor EGFR and ALK co-alterations.
Methods: We reviewed the genomic profiles of 419 ALK-rearranged NSCLC patients with the intent of investigating the EGFR kinase domain (exon 18–21) and ALK co-alterations. The genomes of these patients were sequenced in a Clinical Laboratory Improvement Amendments-certified laboratory.
Results: The overall frequency of concomitant EGFR (exon 18–21) and ALK alterations was 5.01% (21/419) in ALK-rearranged NSCLC patients. The concomitant rate of EGFR alterations in patients with EML4-ALK co-alterations (3.06%, 11/359) was dramatically lower than that in patients with non-EML4-ALK co-alterations (16.67%, 10/60, p<0.01). EML4-ALK/EGF R co-alterations were more prone to occur in females than in males, and non-EML4-ALK/EGFR co-alterations were more common in males than in females (p=0.02). Before the detection of EGFR-ALK co-alterations, some patients were treated with EGFR-TKIs (n=16) according to previously detected EGFR alterations; Kaplan–Meier analysis revealed that EML4-ALK/EGFR co-altered patients (n=7) had a significantly shorter progression-free survival (PFS) after EGFR-TKI treatment than that of non-EML4-ALK/EGFR co-altered patients (n=8; mPFS, 6.0 vs 15.0 months, p=0.046). In addition, we demonstrated the subsequent clinical outcomes of co-altered patients after previous EGFR-TKI treatment. Five EGFR/ALK co-altered patients treated with single TKIs (EGFR-TKIs or ALK-TKIs) displayed diverse clinical outcomes. Three patients who received dual-TKI treatment (EGFR-TKI plus ALK-TKI) all achieved a PFS of more than 5 months (8.4 months, 8.6 months, >5.2 months).
Conclusion: EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alterations displayed distinct clinical features and responses to EGFR-TKIs, suggesting that non-EML4-ALK co-alterations are likely to occur as a resistance mechanism to EGFR-TKI. In addition, dual-TKI therapy might be a better choice than single-TKI treatments for these co-altered patients. To the best of our knowledge, this is the largest dual-positive EGFR/ALK cohort study in People's Republic of China.
Keywords: EGFR alteration, ALK rearrangement, nonsmall cell lung cancer, EML4-ALK, tyrosine kinase inhibitor
Introduction
EGFR and anaplastic lymphoma kinase (ALK) gene alterations have both been regarded as oncogenic driver mutations and drug targets in nonsmall cell lung cancers (NSCLCs).1–3 EGFR and ALK alterations were conventionally considered to be mutually exclusive4–6 and as mutual causes of resistance to ALK-TKIs or EGFR-TKIs.7,8 However, co-alterations of EGFR and ALK exist in a subset of NSCLCs and challenge the previous dogma.9–11
In ALK-rearranged NSCLC patients, echinoderm microtubule-associated protein-like 4 (EML4) is the main fusion partner of ALK. EML4-ALK, in which the N-terminal of EML4 is fused to the intracellular kinase domain of ALK, displays constitutive ALK activation and generates oncogenic activity.4,5 In addition to EML4, other ALK 5ʹ-partners have also been identified, including kinesin family member 5B, TRK-fused gene, and kinesin light chain 1.12–14 The frequency of non-EML4-ALK alterations is approximately 10–20% in ALK-positive lung cancers,15,16 and the clinical significance of these alterations still under investigation.
In previous studies about patients with EGFR and ALK co-alterations, researchers often combined patients with EGFR/ALK co-alterations as a single group, regardless of the ALK fusion partner, for clinical features or drug efficacy investigations. Little is known about the difference in clinical features and drug efficacy between the EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alteration subgroups. Here, we interrogated the distinct concurrent alterations rate, clinical features, and clinical outcomes during EGFR-TKI treatment in both the EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alteration subgroups. In addition, we sought to evaluate the clinical activity of these co-altered patients in response to single-TKI or dual-TKI treatments.
Materials and methods
Patient information
We retrospectively reviewed the genomic profiling data of 7,661 lung cancer patients, whose tissue or plasma samples were sequenced in a Clinical Laboratory Improvement Amendments-certified clinical molecular diagnostic laboratory using next-generation sequencing (NGS) between September 2015 and January 2018. Among them, 419 patients were identified as harboring ALK rearrangements. The clinical characteristics of patients harboring dual-positive EGFR and ALK alterations were collected. All patients had a histologically confirmed diagnosis of advanced-stage NSCLC. Progression-free survival (PFS) after EGFR-TKI treatment and survival information were assessed for the cohorts. The study was approved by the institutional review board of Peking University Shenzhen Hospital. All other centers were covered by this protocol. All patients whose tissue and medical data were used in this research provided written informed consent, in accordance with the Declaration of Helsinki.
Tissue DNA and plasma cfDNA preparation
The tissue DNA was extracted from all tissue samples using the QIAamp DNA FFPE tissue Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. Circulating cfDNA was recovered from 4 to 5 mL plasma by using the QIAamp Circulating Nucleic Acid kit (Qiagen). DNA was quantified with the Qubit 2.0 fluorimeter (Thermo Fisher Scientific, Waltham, MA, USA).
Targeted DNA sequencing
The genomic DNA was profiled by using capture-based targeted sequencing panel that consisted of 8, 56, 168 or 295 cancer-related genes (Burning Rock Biotech, Guangzhou, People's Republic of China). Alterations of eight well-established driver genes, EGFR, ERBB2, ALK, ROS1, RET, KRAS, BRAF and MET, were all included in these sequencing panels. The NGS library was prepared as previously described.17 In brief, DNA was sheared using Covaris M220 (Covaris, Inc., Woburn, MA, USA), which was then followed by end-repair, phosphorylation and adaptor ligation. Fragments that were 200–400 bp in size were selected by beads (Agencourt AMPure XP Kit, Beckman Coulter, Brea, CA, USA), followed by hybridization with capture probes or baits, hybrid selection with magnetic beads and PCR amplification. A high-sensitivity DNA assay was performed using a bioanalyzer to evaluate the DNA quality and size. Indexed samples were then sequenced on a NextSeq 500 (Illumina, Inc., San Diego, CA, USA) with pair-end reads.
The sequencing data were in a FASTQ format and mapped to the human genome (hg19) using BWA aligner 0.7.10.18,19 Local alignment optimization, mark duplication and variant calling were performed using the Genome Analysis ToolKit 3.2,20 Picard (http://picard.sourceforge.net/) and VarScan.21 Gene translocations were called with FACTERA,22 and the copy number variation was analyzed with an in-house algorithm based on sequencing depth.
Statistical analysis
Statistical analysis was conducted using R software version 3.3.3. Between the EML4-ALK and non-EML4-ALK subgroups, differences in sex and mutation rate were calculated and presented using Fisher’s exact tests, while differences in age were calculated using paired, two-tailed Student’s t-tests. For all statistical tests, p<0.05 was considered statistically significant.
Results
Patient characteristics
The genomic sequencing results of 419 ALK-rearranged NSCLC patients were retrospectively reviewed, including 359 (85.7%) EML4-ALK fusions and 60 (14.3%) non-EML4-ALK fusions. Among the 419 ALK-rearranged lung cancers, a total of 21 patients (5.01%) were detected to harbor concurrent ALK and EGFR (exon 18–21) genomic alterations. The concomitant rate of EGFR alterations in patients harboring EML4-ALK co-alterations (3.06%, 11/359) was dramatically lower than that in non-EML4-ALK co-altered patients (16.67%, 10/60, p<0.01, Fisher’s exact test, Table 1).
Table 1.
Characteristics of patients harboring dual-positive ALK/EGFR alterations
| Patient Characteristics | EML4-ALK | Non-EML4-ALK | P-value |
|---|---|---|---|
| Total | 11 | 10 | |
| Co-occurrence rate in ALK-rearranged NSCLC | 3.06% (11/359) | 16.67% (10/60) | p<0.01 |
| Gender | p=0.02 | ||
| Male | 4 | 9 | |
| Female | 7 | 1 | |
| Age (year) | p=0.31 | ||
| Median | 53 | 59.5 | |
| Range | 42–74 | 44–81 | |
| Histological types | |||
| Adenocarcinoma | 11 | 10 |
Abbreviations: ALK, anaplastic lymphoma kinase; NSCLC, nonsmall cell lung cancer; EML4, echinoderm microtubule-associated protein-like 4.
All the 21 ALK/EGFR co-altered cases were diagnosed as adenocarcinomas. In the EML4-ALK/EGFR co-altered subgroup, 4 (36%) patients were male, and 7 (64%) patients were female. In contrast, the non-EML4-ALK/EGFR co-altered subgroup comprised 9 (90%) males and 1 (10%) female (p=0.02, Fisher’s exact test). This difference suggested that EML4-ALK/EGFR co-alterations were more prone to occur in females than males, and non-EML4-ALK/EGFR co-alterations were more common in males than in females. The median age of the EML4-ALK/EGFR and non-EML4-ALK/EGFR co-altered subgroups were 53.0 and 59.5 years, respectively (p=0.31, t-test). The patient characteristics are summarized in Table 1.
Identification of molecular patterns of co-altered ALK/EGFR
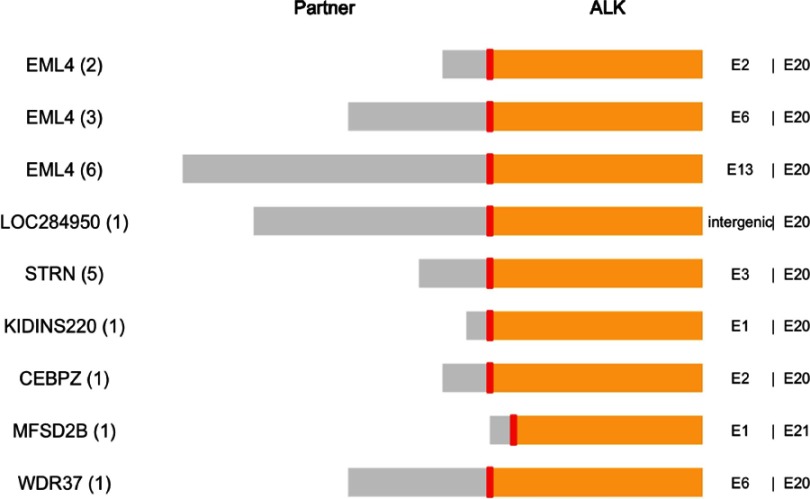
In the 11 EML4-ALK/EGFR co-altered patients, capture-based sequencing identified different EML4-ALK variants, including 6 with E13;A20 (V1), 3 with E6;A20 (V3), and 2 with E2;A20 (Figure 1). As for the 10 non-EML4-ALK/EGFR co-altered patients, 6 unique ALK fusion partners were detected. STRN was the most common fusion partner in non-EML4-ALK co-alterations and was identified in five patients (50%). Apart from those five patients, another two STRN-ALK positive patients were identified in the whole cohort of 419 ALK-rearranged NSCLC patients. One patient harbored a co-alteration of STRN-ALK and EGFR exon 15 V592I, and another patient was identified as being STRN-ALK-positive only. This observation suggested that co-alterations with EGFR were a common feature of STRN-ALK fusions. Before these five patients were detected to have STRN-ALK/EGFR (exon 18–21) co-alterations, all of them had previously been detected to have EGFR alterations and had been treated with EGFR-TKIs but not with ALK-TKIs, indicating that STRN-ALK might be one of the resistance mechanisms for EGFR-TKIs. In addition, five novel ALK fusion partners were identified including WDR37-ALK, CEBPZ-ALK, KIDINS220-ALK,LOC284950-ALK and MFSD2B-ALK.
Figure 1.
Structure and breakpoints of the 21 ALK fusions in EGFR-mutated nonsmall cell lung cancer patients identified by next-generation sequencing. E2:E20 indicates that exon 2 of EML4 is fused to exon 20 of ALK. The number in the brackets indicates the number of patients with that respective fusion variant.Abbreviations: ALK, anaplastic lymphoma kinase; EML4, echinoderm microtubule-associated protein-like 4.
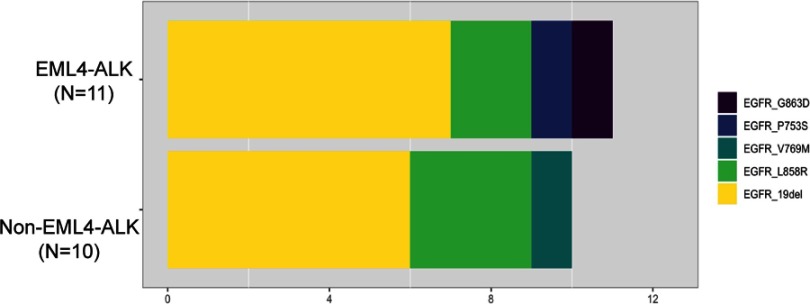
Of the 21 patients who harbored co-altered ALK/EGFR, the following 11 patients with EML4-ALK/EGFR co-alterations were identified to have diverse EGFR variants: 7 patients had exon 19 deletions, 2 had exon 21 L858R point mutations, 1 had a P753S point mutation and 1 had a G863D point mutation. Meanwhile, the 10 non-EML4-ALK/EGFR-altered patients included 6 EGFR exon 19 deletions, 3 L858R point mutations and 1 V769M point mutation (Figure 2).
Figure 2.
EGFR mutation variants in the 21 EGFR/ALK co-altered NSCLC patients. Different colors indicate different variants of EGFR.Abbreviations: ALK, anaplastic lymphoma kinase; EML4, echinoderm microtubule-associated protein-like 4.
Is concomitant non-EML4-ALK co-alterations a de novo or acquired rearrangement?
The distribution of EML4-ALK and non-EML4-ALK co-alterations in the “de novo ALK”, “acquired ALK” and “uncertain” groups are summarized in Table 2. Patients identified with co-altered EGFR/ALK at the time of the first diagnosis were classified as “de novo ALK” since they had not received TKI treatment before. This group contained 4 patients (4/5, 80%) with EGFR/EML4-ALK co-alterations and 1 patient (1/5, 20%) with an EGFR/non-EML4-ALK co-alteration. Patients who were identified as EGFR (+)/ALK (-) at a previous diagnosis and later detected to have co-altered EGFR/ALK after developing EGFR-TKI resistance were classified as “acquired ALK”; this group consisted of 1 patient with (1/4, 25%) an EGFR/EML4-ALK co-alteration and 3 patients (3/4, 75%) with EGFR/non-EML4-ALK co-alterations. In addition, patients who were detected to have EGFR/ALK co-alterations after developing EGFR-TKI resistance but whose previous ALK status was unavailable were defined as “uncertain”; this group included 6 patients with EGFR/EML4-ALK co-alterations and 6 patients with EGFR/non-EML4-ALK co-alterations. The observation that EML4-ALK co-alterations accounted for most of the rearrangements in the de novo ALK patients (80%) while non-EML4-ALK co-alterations accounted for most of the alterations in the acquired ALK patients (75%) after EGFR-TKI treatment indicated that EML4-ALK co-alterations were likely to be de novo alterations and non-EML4-ALK co-alterations might be likely to appear as a drug resistance mechanism of EGFR-TKIs.
Table 2.
Distribution of EML4-ALK and non-EML4-ALK in de novo and acquired ALK rearrangements
| ALK fusions | NGS before baselinea | Treatment before baseline | EML4-ALK | Non-EML4-ALK |
|---|---|---|---|---|
| De novo ALK (n=5) | N/A | Treatment-naive | 4/5 (80%) | 1/5 (20%) |
| Acquired ALK (n=4) | EGFR (+), ALK (-) | EGFR-TKI | 1/4 (25%) | 3/4 (75%) |
| Uncertain (n=12) | EGFR (+), ALK (unknown) | EGFR-TKI | 6/12 (50%) | 6/12 (50%) |
Notes: aThe timepoint of positive-detection of co-altered EGFR/ALK was defined as baseline.
Abbreviations: ALK, anaplastic lymphoma kinase; N/A, not applicable; NGS, next generation sequencing; EML4, echinoderm microtubule-associated protein-like 4.
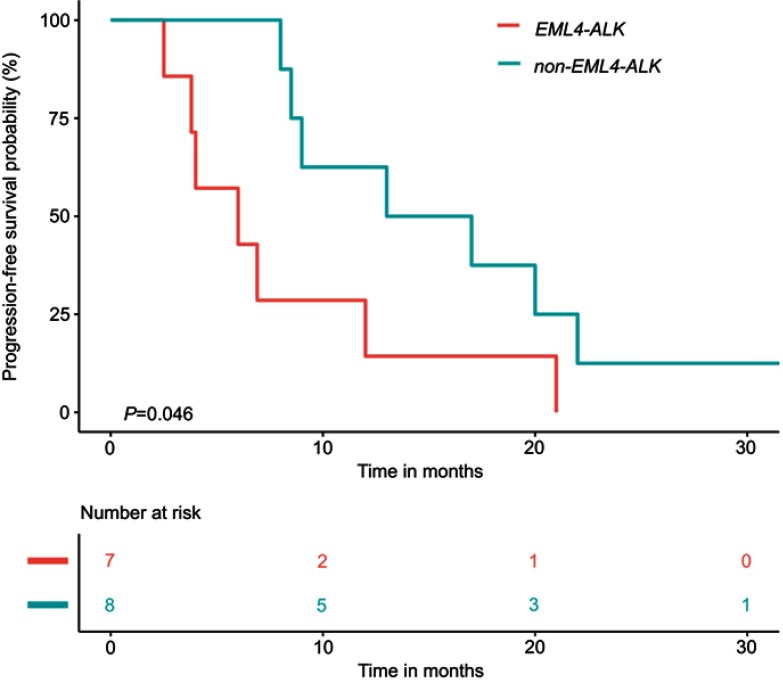
Next, for patients progressed on previous EGFR-TKI treatment (“acquired ALK” plus “uncertain” subgroups), we interrogated the correlation of PFS in response to EGFR-TKIs before the detection of ALK rearrangements and the distribution of ALK fusion partners after EGFR-TKI treatment. The Kaplan–Meier analysis revealed that after EGFR-TKI treatment, patients with EML4-ALK/EGFR co-alterations (n=7) commonly had a significantly shorter median PFS than that of patients with non-EML4-ALK/EGFR co-alterations (n=8, PFS information of one patient was unavailable; mPFS, 6.0 vs 15.0 months, p=0.046, Figure 3). The discrepancy in PFS between the two subgroups has never been reported before and could suggest that EML4-ALK co-alterations were likely to be induced in the early stages of EGFR-TKI treatment, whereas non-EML4-ALK co-alterations might be acquired at relatively late stages after EGFR-TKI treatment.
Figure 3.
Kaplan–Meier curve demonstrates the correlation of PFS in response to EGFR-TKIs before ALK detection and the distribution of ALK fusion partners after EGFR-TKI treatment. The risk table (below) illustrates the number of patients included per time point. Abbreviations: ALK, anaplastic lymphoma kinase; EML4, echinoderm microtubule-associated protein-like 4; PFS, progression-free survival.
Subsequent TKI efficacy after previous EGFR-TKI treatment
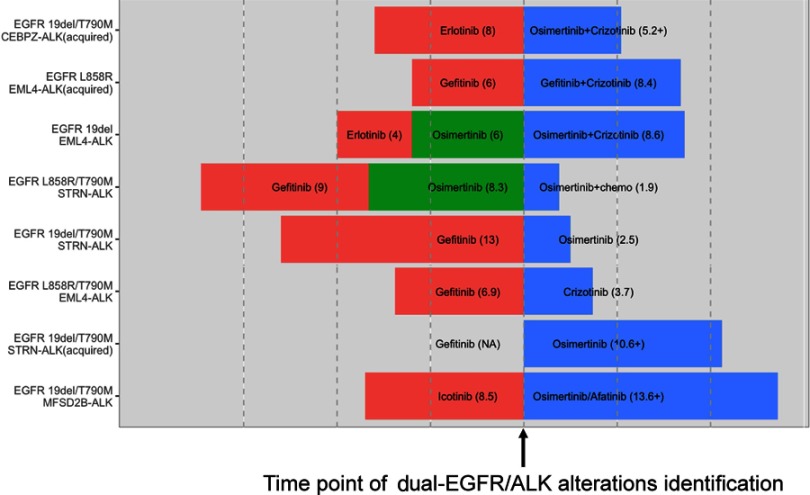
The clinical data of eight patients with EGFR/ALK co-alterations who received subsequent single- or dual-TKI treatment after already receiving previous EGFR-TKI treatment were collected and are presented in Figure 4. Of the eight cases, six (75%) of them were detected to have the first-generation EGFR-TKI resistant mutation T790M, indicating that ALK rearrangements might occur concomitantly with EGFR-T790M after EGFR-TKI therapy. Five patients were treated with subsequent single TKIs (EGFR-TKI or ALK-TKI) after the detection of EGFR/ALK co-alterations, and the clinical efficacy of the single TKI varied greatly. Among those with the T90M mutation, three patients who received subsequent osimertinib treatment had a PFS of 2.5, 2.9 and >10.6 months. One patient received combined afatinib and osimertinib and had a PFS of more than 13.6 months. One patient was treated with osimertinib plus chemotherapy and only had a PFS of 1.9 months, and one patient received crizotinib and achieved a PFS of 3.7 months. In addition, three patients received subsequent dual-TKI treatment (EGFR-TKI plus ALK-TKI), and the PFS of all three patients were longer than 5 months (8.4 months, 8.6 months, >5.2 months), indicating that patients might benefit from the combination therapy of ALK-TKIs and EGFR-TKIs.
Figure 4.
Presentation of the detailed treatment history (before and after baseline) and subsequent clinical outcomes (after baseline) of eight patients with EGFR/ALK co-alterations. The timepoint of the positive detection of the EGFR/ALK co-alteration was defined as baseline. After baseline, five patients were treated with subsequent single TKIs (EGFR-TKIs or ALK-TKIs), and three patients received subsequent dual-TKI treatment (EGFR-TKI plus ALK-TKI).Abbreviation: ALK, anaplastic lymphoma kinase.
Discussion
EGFR mutations and ALK rearrangements were commonly considered to be mutually exclusive in previous studies. However, there is increasing evidence to support the notion that they can be concomitantly mutated in cancers. In this study, we retrospectively reviewed the genomic profiling data of 419 ALK-positive Chinese NSCLC patients and identified 21 patients who harbored concurrent EGFR and ALK gene alterations. We investigated and compared the prevalence, clinical features, and different responses to tyrosine kinase inhibitors in subgroups of patients harbored EML4-ALK/EGFR or non-EML4-ALK/EGFR co-alterations. We found that EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alterations showed different clinical characteristics and clinical outcomes to EGFR-TKIs, suggesting that non-EML4-ALK co-alterations might be likely to emerge as a drug resistance mechanism for EGFR-TKIs. In addition, dual-TKI therapy might be a better choice for patients with co-altered ALK/EGFR than single-TKI therapy. To the best of our knowledge, this is the largest cohort study of ALK rearrangements for dual EGFR/ALK co-alterations in People's Republic of China. In addition, this is the largest series of ALK-positive patients reported, and five novel ALK fusion partners were identified in this study.
There were inconsistencies about the clinical features of EGFR/ALK co-mutated patients. Yang et al reported that the median age in EGFR/ALK co-altered patients was 59 years, and the number of patients who were female and male were 8 (62%) and 5 (38%), respectively.9 A European study revealed that there were two males and one female in their study of three EGFR/ALK co-altered patients.23 We also interrogated the clinical demographics of EGFR/ALK comutated patients in this study. We found that the median age of the co-altered patients was 58 years old. Among them, 8 (38.1%) patients were female and 13 (61.9%) were male. This difference in sex was probably induced by different factors and the limited cohort size of each study. Moreover, none of the previous studies compared the clinical characteristics between patients with EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alterations. We found that EML4-ALK/EGFR co-alterations were more prone to occur in females than in males, and non-EML4-ALK/EGFR co-alterations were more likely to occur in males than in females (p=0.02). The median age of EML4-ALK/EGFR co-altered patients was slightly younger than that of non-EML4-ALK/EGFR co-altered patients (53.0 vs 59.5 years, p=0.31). This clinically relevant observation may be important for further investigations of treatment strategies.
Several previous studies have reported that acquired ALK rearrangements could occur as a resistance mechanism after EGFR-TKI treatment. For example, Liang et al reported that ALK fusions were detected in the new metastatic lesion of an EGFR-mutated NSCLC patient.8 In our study, we found that after EGFR-TKI treatments EML4-ALK co-alterations accounted for most of the 5 de novo ALK-rearranged patients (80%, 4/5), while the non-EML4-ALK co-alterations accounted for most of the 4 acquired ALK-positive patients (75%, 3/4). We also observed that EML4-ALK/EGFR co-altered patients had a significantly inferior median PFS in response to EGFR-TKI compared to that of non-EML4-ALK/EGFR co-altered patients (mPFS, 6.0 vs 15.0 months, p=0.046). These results indicated that EML4-ALK co-alterations were likely to be a de novo alteration and non-EML4-ALK co-alterations might be an acquired resistance mechanism induced by EGFR-TKIs. Moreover, EML4-ALK co-alterations were likely to be induced in the early stages during EGFR-TKI treatment, and non-EML4-ALK co-alterations might emerge at a relatively late stage. Although the evidence of non-EML4/ALK co-alterations as a resistance mechanism to EGFR-TKIs has not been solidified, our study has greatly expanded the current knowledge of the underlying association of EGFR and ALK alterations and might be helpful for understanding the co-alteration mechanism and guiding clinical decision-making. In addition, we proposed that for patients developing first-line EGFR-TKI treatment resistance, genomic profiling and sequencing analysis should be performed rather than testing for EGFR-T790M solely since the EGFR-T790M mutation and ALK rearrangements may be detected simultaneously.
A limited number of studies have reported about the clinical efficacy of TKIs in patients who harbor EGFR/ALK co-alterations and have reached diverse conclusions. A Caucasian patient with lung adenosquamous carcinoma who harbored such a co-alteration displayed resistance to erlotinib treatment.24 Ten NSCLC patients in a Chinese cohort achieved a response rate of 80% to the first-line EGFR-TKIs.9 In a Korean cohort with 14 co-altered patients, 3 patients who were treated with gefitinib showed poor response, and 8 patients who received ALK inhibitors showed good response with a response rate of 87.5%.11 Previous studies with targeted therapy on EGFR/ALK co-altered patients commonly focused on single TKIs or the sequential use of EGFR- and ALK-TKIs. No studies have compared the clinical outcomes of single TKIs with combinatorial therapies of EGFR-TKIs and ALK-TKIs in EGFR/ALK co-altered patients. Here, we demonstrated the clinical activities of single TKIs and dual TKIs in EGFR/ALK co-altered NSCLC patients who progressed after EGFR-TKIs. We found that the clinical efficacy of single TKIs in five patients varied greatly. We also observed that patients who harbor dual alterations might benefit from the combination therapy of ALK-TKIs and EGFR-TKIs, and the PFS of all three cases with dual alterations in this study were longer than 5 months.
However, there were several limitations in this study. First, 21 (1.2%) NSCLC patients were identified to have EGFR/ALK co-alterations from a total of 419 ALK-positive patients. Based on the small number of patients, it was difficult to draw clear conclusions on the clinical characteristics and therapeutic outcomes of the patients. Second, since not all samples were collected at baseline and since there was previously identified targetable mutations and treatment records were lacking, we proposed several concepts from our analysis, and further investigations are still needed to validate our ideas.
Conclusion
In summary, we demonstrated that NSCLC patients with EML4-ALK/EGFR and non-EML4-ALK/EGFR co-alterations displayed distinct rates of concurrent alterations, clinical demographicsand survival in response to EGFR-TKI treatment. We proposed that non-EML4-ALK alterations are likely to be an acquired resistance mechanism later during EGFR-TKI treatment. The underlying molecular mechanisms of the two alterations and potential of combination treatments with dual TKIs require further investigation with this co-altered subgroup.
Acknowledgments
The authors wish to thank Burning Rock Biotech for their technical and writing assistance. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. The abstract of this paper was presented at the 2018 World Conferences on Lung Cancer (Asia) as a poster presentation with interim findings. The poster’s abstract was published in “Poster Abstracts” in 2018 Journal of Thoracic Oncology (DOI: https://doi.org/10.1016/j.jtho.2018.10.035).
Disclosure
The authors report no conflicts of interest in this work.
References
- 1.Kwak EL, Bang YJ, Camidge DR, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med. 2010;363(18):1693–1703. doi: 10.1056/NEJMoa1006448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Croegaert K, Kolesar JM. Role of anaplastic lymphoma kinase inhibition in the treatment of non-small-cell lung cancer. Am J Health Syst Pharm. 2015;72(17):1456–1462. doi: 10.2146/ajhp140836 [DOI] [PubMed] [Google Scholar]
- 3.Kobayashi S, Boggon TJ, Dayaram T, et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med. 2005;352(8):786–792. doi: 10.1056/NEJMoa044238 [DOI] [PubMed] [Google Scholar]
- 4.Shaw AT, Yeap BY, Mino-Kenudson M, et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J Clin Oncol. 2009;27(26):4247–4253. doi: 10.1200/JCO.2009.22.6993 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Soda M, Choi YL, Enomoto M, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007;448(7153):561–566. doi: 10.1038/nature05945 [DOI] [PubMed] [Google Scholar]
- 6.Horn L, Pao W. EML4-ALK: honing in on a new target in non-small-cell lung cancer. J Clin Oncol. 2009;27(26):4232–4235. doi: 10.1200/JCO.2009.23.6661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sasaki T, Koivunen J, Ogino A, et al. A novel ALK secondary mutation and EGFR signaling cause resistance to ALK kinase inhibitors. Cancer Res. 2011;71(18):6051–6060. doi: 10.1158/0008-5472.CAN-11-1340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liang W, He Q, Chen Y, et al. Metastatic EML4-ALK fusion detected by circulating DNA genotyping in an EGFR-mutated NSCLC patient and successful management by adding ALK inhibitors: a case report. BMC Cancer. 2016;16:62. doi: 10.1186/s12885-016-2088-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yang JJ, Zhang XC, Su J, et al. Lung cancers with concomitant EGFR mutations and ALK rearrangements: diverse responses to EGFR-TKI and crizotinib in relation to diverse receptors phosphorylation. Clin Cancer Res. 2014;20(5):1383–1392. doi: 10.1158/1078-0432.CCR-13-0699 [DOI] [PubMed] [Google Scholar]
- 10.Ulivi P, Chiadini E, Dazzi C, et al. Nonsquamous, non-small-cell lung cancer patients who carry a double mutation of EGFR, EML4-ALK or KRAS: frequency, clinical-pathological characteristics, and response to therapy. Clin Lung Cancer. 2016;17(5):384–390. doi: 10.1016/j.cllc.2015.11.004 [DOI] [PubMed] [Google Scholar]
- 11.Won JK, Keam B, Koh J, et al. Concomitant ALK translocation and EGFR mutation in lung cancer: a comparison of direct sequencing and sensitive assays and the impact on responsiveness to tyrosine kinase inhibitor. Ann Oncol. 2015;26(2):348–354. doi: 10.1093/annonc/mdu530 [DOI] [PubMed] [Google Scholar]
- 12.Togashi Y, Soda M, Sakata S, et al. KLC1-ALK: a novel fusion in lung cancer identified using a formalin-fixed paraffin-embedded tissue only. PLoS One. 2012;7(2):e31323. doi: 10.1371/journal.pone.0031323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Choi YL, Lira ME, Hong M, et al. A novel fusion of TPR and ALK in lung adenocarcinoma. J Thorac Oncol. 2014;9(4):563–566. doi: 10.1097/JTO.0000000000000093 [DOI] [PubMed] [Google Scholar]
- 14.Shan L, Jiang P, Xu F, et al. BIRC6-ALK, a novel fusion gene in ALK break-apart FISH-negative lung adenocarcinoma, responds to crizotinib. J Thorac Oncol. 2015;10(6):e37–e39. doi: 10.1097/JTO.0000000000000467 [DOI] [PubMed] [Google Scholar]
- 15.Li Y, Zhang T, Zhang J, et al. Response to crizotinib in advanced ALK-rearranged non-small cell lung cancers with different ALK-fusion variants. Lung Cancer. 2018;118:128–133. doi: 10.1016/j.lungcan.2018.01.026 [DOI] [PubMed] [Google Scholar]
- 16.Ou S-HI, Schrock AB, Gowen K, et al. Association of ALK resistance mutations by EML4-ALK variant (v3 vs. non-v3) in ALK+ non-small cell lung cancer (NSCLC). J Clin Oncol. 2017;35(15_suppl):9010. doi: 10.1200/JCO.2017.35.15_suppl.9010 [DOI] [Google Scholar]
- 17.Mao X, Zhang Z, Zheng X, et al. Capture-based targeted ultradeep sequencing in paired tissue and plasma samples demonstrates differential subclonal ctDNA-releasing capability in advanced lung cancer. J Thorac Oncol. 2017;12(4):663–672. doi: 10.1016/j.jtho.2016.11.2235 [DOI] [PubMed] [Google Scholar]
- 18.Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114–2120. doi: 10.1093/bioinformatics/btu170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–1760. doi: 10.1093/bioinformatics/btp324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–1303. doi: 10.1101/gr.107524.110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Koboldt DC, Zhang Q, Larson DE, et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22(3):568–576. doi: 10.1101/gr.129684.111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Newman AM, Bratman SV, Stehr H, et al. FACTERA: a practical method for the discovery of genomic rearrangements at breakpoint resolution. Bioinformatics. 2014;30(23):3390–3393. doi: 10.1093/bioinformatics/btu549 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lee T, Lee B, Choi YL, Han J, Ahn MJ, Um SW. Non-small cell lung cancer with concomitant EGFR, KRAS, and ALK mutation: clinicopathologic features of 12 cases. J Pathol Transl Med. 2016;50(3):197–203. doi: 10.4132/jptm.2016.03.09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tiseo M, Gelsomino F, Boggiani D, et al. EGFR and EML4-ALK gene mutations in NSCLC: a case report of erlotinib-resistant patient with both concomitant mutations. Lung Cancer. 2011;71(2):241–243. doi: 10.1016/j.lungcan.2010.11.014 [DOI] [PubMed] [Google Scholar]