Abstract
Transcriptional changes in the hippocampus are required for memory formation, and these changes are regulated by numerous post-translational modifications of chromatin-associated proteins. One of the epigenetic marks that has been implicated in memory formation is histone 3 lysine 4 trimethylation (H3K4me3), and this modification is found at the promoters of actively transcribed genes. The total levels of H3K4me3 are increased in the CA1 region of the hippocampus during memory formation, and genetic perturbation of the K4 methyltransferases and demethylases interferes with forming memories. Previous chromatin immunoprecipitation followed by deep sequencing (ChIP-seq) analyses failed to detect changes in H3K4me3 levels at the promoters of memory-linked genes. Since the breadth of H3K4me3 marks was recently reported to be associated with the transcriptional outcome of a gene, we re-analyzed H3K4me3 ChIP-seq data sets to identify the role of H3K4me3 broad domains in CA1 neurons, as well as identify differences in breadth that occur during contextual fear conditioning. We found that, under baseline conditions, broad H3K4me3 peaks mark important learning and memory genes and are often regulated by super-enhancers. The peaks at many learning-associated genes become broader during novel environment exposure and memory formation. Furthermore, the important learning- and memory-associated lysine methyltransferases, Kmt2a and Kmt2b, are involved in maintaining H3K4me3 peak width. Our findings highlight the importance of analyzing H3K4me3 peak shape, and demonstrate that breadth of H3K4me3 marks in neurons of the hippocampus is regulated during memory formation.
Graphical Abstract

1. Introduction
Analyzing dynamic regulation of modifications on histone tails is emerging as a way to understand the molecular underpinnings of neuronal plasticity during memory formation. Active transcription of genes with RNA polymerase II (Pol II) is required to form a memory (Agranoff et al., 1965), and transcription is regulated by posttranslational modifications of histones (Lubin et al., 2011). Lysine methylation is one such modification that adorns the unstructured N-terminal tails of histones. Trimethylation of histone 3 at lysine 4 (H3K4me3) near gene promoters is correlated with high levels of transcriptional activity (Barski et al., 2007; Santos-Rosa et al., 2002), and has been implicated as a regulator of memory formation (Collins et al., 2019). During contextual fear conditioning (CFC), a behavioral test that elicits robust memory formation, global levels of H3K4me3 in the hippocampus are increased (Gupta et al., 2010).
Further leading to the conclusion that H3K4me3 is important in memory formation, all the known methyltransferases and demethylases that alter methylation of H3K4 have been linked to human diseases and disabilities affecting cognition (Collins et al., 2019). The most-studied H3K4 methyltransferases in the context of memory formation are lysine methyltransferase 2a (Kmt2a) and 2b (Kmt2b). Perturbation of both isoforms in rodent models leads to disruptions in memory formation (Jakovcevski et al., 2015; Kerimoglu et al., 2013; Kerimoglu et al., 2017). Moreover, mutations of KMT2A and KMT2B in humans are associated with Weidmann-Steiner syndrome and Dystonia 28, respectively, and both conditions are associated with intellectual disability (Collins et al., 2019). While related, there are distinctions between Kmt2a and Kmt2b since they affect H3K4 methylation at different regions of the genome (Kerimoglu et al., 2017).
Most actively transcribed genes harbor enrichment of H3K4me3 in a symmetric 1–2 kilobase (kb) region on either side of the transcription start site (TSS), but a subset of genes has been shown to have broader enrichment of H3K4me3 that is several kb in length and skewed downstream into the gene body. Genes that are marked by broad domains of H3K4me3 are important for cell type-specific functions across a variety of organisms and tissue types (Benayoun et al., 2014). We are beginning to understand that H3K4me3 peak shape is an important aspect of this modification, and broad domains are linked to various functions including higher transcription levels (Chen et al., 2015), a poised-for transcriptional activation state (Grandy et al., 2016), transcriptional consistency (Benayoun et al., 2014), and enhancer regulation (Chen et al., 2015).
Enhancer elements are DNA sequences that regulate transcription from a promoter, despite being located far from the promoter. Recent evidence shows that enhancers regulate the transcription of important learning- and memory-associated genes (Joo et al., 2016; Kim et al., 2010). Furthermore, some enhancers are classified as super-enhancers, which, like regular enhancers, activate transcription, but are broad intergenic regions that serve as docking sites for transcription factors and transcriptional regulators (Whyte et al., 2013). Super-enhancers, like H3K4me3 broad domains, are important for defining cell identity (Hnisz et al., 2013). H3K4me3 modifiers have been linked to regulation of super-enhancer activity (Dhar et al., 2018), and H3K4me3 itself has been found at enhancer elements (Core et al., 2014; Henriques et al., 2018).
The hippocampus is essential for memory formation, and the CA1 subregion of this brain structure is particularly important for memory retrieval (Ji et al., 2008). In previous studies, labs did CFC, then isolated CA1 neurons and conducted chromatin immunoprecipitation followed by sequencing (ChIP-seq) of several epigenetic marks, including H3K4me3 (Halder et al., 2016). Despite seeing a global increase in H3K4me3 across promoters, the study found very few (<100) site-specific increases during learning, and the increases they observed were not predominantly at memory-associated genes. However, the newly emerging chromatin metric, peak breadth, was not assessed.
In this study, we analyzed previously archived ChIP-seq datasets to examine H3K4me3 broad peaks in CA1 neurons. We find that broad domains are correlated with basal transcription level, transcriptional induction during learning, and super-enhancer regulation, but not transcriptional consistency. We found that genes exhibiting increases in H3K4me3 breadth during memory formation are transcriptionally activated, and have functions associated with learning. This was not true for genes that increased in intensity, but maintained a constant width. This demonstrates that breadth may be more relevant than peak intensity in regards to transcriptional induction following learning. Furthermore, using conditional knockout datasets, we find that learning-associated genes Kmt2a and Kmt2b are major regulators of H3K4me3 peak breadth in CA1 neurons. Our findings indicate that H3K4me3 peak breadth is important for understanding the regulation of this mark during memory formation.
2. Methods
2.1. Data Sources
Raw H3K4me3 ChIP- and RNA-sequencing datasets were downloaded from the Gene Expression Omnibus (GEO) at accession numbers GSE74971 (Halder et al., 2016) and GSE99250 (Kerimoglu et al., 2017). 3 month-old male C56BL/6 mice were used to generate data in the GSE74971 data set. 4–6 month-old male and female C57BL/6J mice were used to generate data in the GSE99250 data set. In the GSE99250 (Kmt2a and Kmt2b cKO) data set, control mice had the loxP sites, but did not express Cre transgene. Also, animals used for ChIP-seq in this data set were not fear conditioned. Raw data was downloaded as Sequence Read Archive (SRA) files from GEO using the SRA Toolkit (v2.8.2) command fastq-dump (Leinonen et al., 2011). Lists of enhancers and super-enhancers identified in human hippocampus were taken from (Hnisz et al., 2013), accession number GSM916035. Mouse gene IDs were extracted from these lists of human gene IDs using the online interface of Ensembl BioMart (Zerbino et al., 2018).
2.2. H3K4me3 ChIP-seq Data Alignment, Analysis, and Peak Calling
Reads were aligned to the mouse NCBI genome version 38 (mm10) using Bowtie2 (v2.3.3.1) in local mode, which permits base pair clipping at either or both read ends to optimize alignment score (Langmead et al., 2012). The resulting Sequence Alignment Map (SAM) files were converted to Binary Alignment Map (BAM) files and indexed using the SAMtools suite (v1.9) (Li et al., 2009). H3K4me3 peaks were called on individual replicate BAM files with macs2 (v2.1.1) using broad peak calling with a q-value cutoff of 0.1 (Zhang et al., 2008). Peaks within 500 base pairs of each other were stitched together using the merge command of the bedtools suite (v2.27.1) (Quinlan et al., 2010). Peaks were then filtered to retain only those that overlap promoter regions (defined as −100 to +500 base pairs relative to the TSS) and assigned to RefSeq-annotated genes (Haeussler et al., 2019; Karolchik et al., 2004). For the GSE74971 data set, H3K4me3 breadth was defined as base pair distance downstream of the annotated gene TSS to the downstream edge of the H3K4me3 peak and averaged over replicates. The top 5% broadest peaks were defined as H3K4me3 broad domains, as had been done previously (Benayoun et al., 2014; Dincer et al., 2015). The top 25% broadest peaks were subject to analysis of changes in H3K4me3 breadth with CFC (broad). The other 75% of peaks were subject to analysis of changes in H3K4me3 intensity levels with CFC (sharp). For the GSE99250 data set, peak breadth was determined as described above for each replicate, and breadth was subjected to QL F-Test using EdgeR (Robinson et al., 2010). Peaks with p < 0.05 were identified as significantly changed in breadth. All control replicates were considered together. To analyze peak intensity, reads over H3K4me3 peak coordinates were counted using a custom script employing the multicov command of bedtools. Normalized read counts were used to define H3K4me3 intensity levels.
Gene tracks were generated from WIG files with bins of 20 base pairs visualized using the Integrative Genome Browser (IGB). Two replicate datasets were averaged into one track for each condition. Opacity of the tracks of the different conditions were decreased and overlaid. Heatmaps were constructed using the annotatePeaks.pl function of Homer (v4.9.1) (Heinz et al., 2010) in TSS mode and visualized using the heatmap.2 function of R.
Our WIG file generating script, Homer, and macs2 all normalize ChIP-seq reads to total mapped reads for all sequencing libraries.
2.3. RNA-seq Data Alignment and Analysis
Reads were aligned to the mouse NCBI genome version 38 (mm10) using gapped alignment with Hisat2 (v2.1.0) using default parameters (Kim et al., 2015). The resulting SAM files were converted to BAM files and indexed using the SAMtools suite. Reads were summarized using FeatureCounts (v1.6.2) at the meta-feature (gene) level (Liao et al., 2014). Multi-overlapping reads, or reads that align to multiple meta-features, were attributed to each matched meta-feature in order to retain reads mapping to multiple isoforms of one gene. Reads Per Kilobase of transcript per Million mapped reads (RPKM) was determined using the rpkm() function of edgeR (Robinson et al., 2010). Transcriptional variance was defined as the standard deviation of RNA-seq replicate RPKM values, normalized to maximum replicate RPKM value. This normalization step was taken to remove bias across genes of varying basal expression levels.
Significantly activated and repressed genes in CFC were taken from the supplementary table of Halder et al., 2016. Ensembl transcript identifiers were converted to Refseq ones, and breadth changes at these genes were calculated.
2.4. Functional Annotation
Gene ontology (GO) analysis was carried out using the functional annotation tool DAVID (Database for Annotation, Visualization, and Integrated Discovery, v6.8) (Huang da et al., 2009a; b). Significance of GO BP_DIRECT terms was evaluated by DAVID’s inbuilt hypergeometric test, using p-values with Bonferroni correction for multiple hypothesis testing.
2.5. Statistical Analyses
Wilcoxon rank sum was implemented in R for analysis of differences in gene expression, transcriptional variance, H3K4me3 breadth, and H3K4me3 intensity levels. If more than one comparison was made, a Kruskal-Wallace test was run. If p < 0.05, a post hoc pairwise Wilcoxon rank sum with Bonferroni correction was done and the p value of this test is reported.
3. Results
3.1. H3K4me3 broad domains in CA1 hippocampal neurons
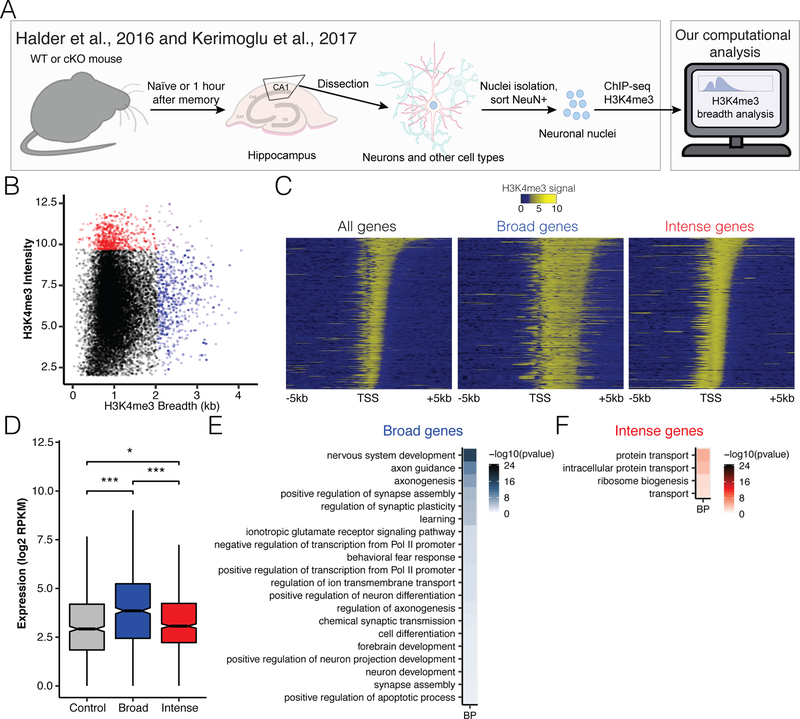
Prior studies have analyzed H3K4me3 peak intensity, while the breadth at gene promoters in neurons has not been thoroughly investigated. Therefore, we sought to understand if broad domains were present in neurons in the CA1 region of the hippocampus and identify the genes they demarcate. We utilized already available datasets of ChIP-seq data from CA1 neurons (Halder et al., 2016; Kerimoglu et al., 2017). In both of the studies we used as source data, the CA1 region of mice was dissected, and the tissue was subjected to fluorescence-activated nuclei sorting to isolate chromatin from neurons within this brain region. ChIP with an H3K4me3 antibody was conducted, and the resulting DNA fragments were subjected to sequencing (Figure 1A). After aligning the data, we conducted peak calling and stitched together adjacent peaks to prevent a single broad peak from being identified as multiple smaller peaks. We first looked at H3K4me3 ChIP-seq in wild-type (WT) mice that had not gone through any behavioral assay, which we sourced from naïve mice of the Halder et al., 2016 study.
Figure 1. H3K4me3 broad domains in neurons of the hippocampus mark promoters of genes that are cell type-specific and highly expressed.
A) Data collection strategy for source data. CA1 region of hippocampus from cKO mice or mice that went through CFC was dissected, nuclei were isolated, and neuronal nuclei were sorted using NeuN antibody (Halder et al., 2016; Kerimoglu et al., 2017). Publicly available data from these studies were downloaded, and we conducted breadth analysis of H3K4me3 peaks and assigned regions of enrichment to annotated promoters. B) Scatter plot of H3K4me3 breadth in kilobases (kb) and H3K4me3 intensity for naïve WT mice. Genes uniquely marked by top 5% broadest H3K4me3 peaks are marked in blue, genes uniquely marked by H3K4me3 peaks with top 5% intensity are red, and genes fitting both criteria are in purple. Genes fulfilling neither criterion are marked in black. C) Heatmaps of H3K4me3 across all genes, broad genes, and intense genes. Heatmaps are centered at gene TSS and span +/− 5 kb. Yellow indicates higher H3K4me3 levels. D) Boxplot of RNA expression levels of control gene set, broad genes, and intense genes. Control peaks were a set of genes of the same number as the broad and sharp gene sets that were randomly selected from genes that were neither broad nor sharp but had H3K4me3 at their promoters. Horizontal lines in the middle of boxes indicate median expression level. * = p < 0.05, *** = p < 1e-8 by Wilcoxon rank sum with Bonferroni correction. E) DAVID GO analysis of broad genes. F) DAVID GO analysis of intense genes. Biological process (BP) terms for genes with p < 0.05. Bonferroni-corrected p value shown.
We calculated peak intensity as well as breadth to define subsets of genes with high levels of enrichment of H3K4me3 and broad domains of H3K4me3 at their promoters (Figure 1B). We focused our analysis on H3K4me3 breadth at gene promoters because this is often where this mark is found. Furthermore, our interest was to characterize genes with broad domains using GO analysis, so we needed to associate our broad domains with recognized gene promoters. With a few exceptions marked in purple, broad peaks are not as intense as sharp peaks and define a distinct group of genes. We looked at the distribution of H3K4me3 around the TSSs of all genes and the broadly and intensely marked gene sets (Figure 1C). In the heatmaps, each gene is represented as a horizontal line, with higher signal intensity from H3K4me3 shown in yellow. Genes are sorted by decreasing peak breadth. Like what has been reported previously (Chen et al., 2015; Collins et al., 2019), our broad genes had asymmetric distribution of signal downstream of the TSS into the gene body compared to the more symmetrically distributed intense genes.
To determine the impact of the distribution and intensity of the H3K4me3 signal at the promoter on transcription, we looked at the RNA expression level of genes with broad and intense peaks at their promoters. As would be expected, intense peaks (Figure 1D) had higher expression compared to a set of control peaks. Moreover, broad peaks had even higher expression than control and intense peaks, suggesting it may be more important for predicting expression level than peak intensity in this cell type.
DAVID GO analysis was used to determine the involvement of broadly and intensely marked genes in different biological processes. Broadly marked genes are attributed to pathways involved in nervous system development, consistent with prior studies that show broad domains determine cell identity (Benayoun et al., 2014; Chen et al., 2015; Dincer et al., 2015). Importantly, we also see the terms ‘learning’ and ‘regulation of synaptic plasticity’ appearing in the broad gene set (Figure 1E). By contrast, sharply marked genes are involved in protein transport and ribosome biogenesis (Figure 1F).
3.2. Transcriptional variance in CA1 neurons is not decreased by H3K4me3 breadth
H3K4me3 broad domains have been previously reported to control transcriptional variance, though this was never investigated in neurons. Using ChIP-seq data from naïve WT mice, we found that transcriptional variance and breadth are not correlated in hippocampal neurons (Supplementary Figure 1A). For H3K4me3 intensity, however, there was a slight negative relationship to variance (Supplementary Figure 1B). This was confirmed statistically using our subgroupings of broad and intense genes defined in Figure 1A. Intensely marked genes had lower variance than controls, but broadly marked genes had similar transcriptional variance to control genes (Supplementary Figure 1C). Based on our findings, H3K4me3 intense peaks, but not broad domains, constrain transcriptional variability in the hippocampus.
3.3. Super-enhancer regulated genes are marked by broad H3K4me3 domains in CA1 neurons
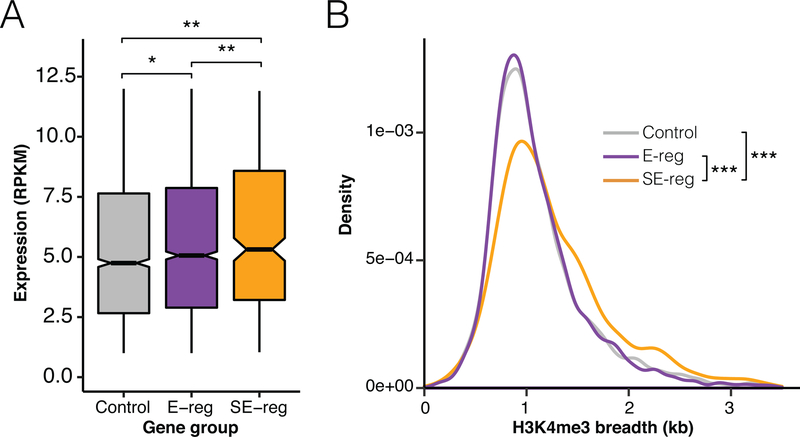
H3K4me3 peak breadth has been linked to regulation by transcriptional enhancer elements. We utilized a list of enhancer regulated (E-reg) and super-enhancer regulated (SE-reg) genes in the hippocampus (Hnisz et al., 2013), and looked at the breadth of H3K4me3 at these genes in the CA1 neuron H3K4me3 ChIP-seq data. First, to predict if these enhancers and super-enhancers activate transcription in the datasets we are utilizing, we looked at the expression levels of these genes in the RNA-seq data. E-reg genes are expressed more than a set of genes that are not predicted to be regulated by an enhancer, and SE-reg genes are even further increased in expression relative to control and E-reg genes (Figure 2A). Since the enhancer and super-enhancer predictions seem likely to be accurately defining E-reg and SE-reg genes, we examined if the width of H3K4me3 peaks from behaviorally naïve WT mice is different at the promoters of genes they regulate. From this analysis, we find that SE-reg genes show increased breadth compared to control genes and E-reg genes (Figure 2B). The median increase in breadth of SE-reg genes relative to control genes is 136 base pairs, so roughly one nucleosome. In CA1 neurons, therefore, genes under the control of super-enhancers have widely distributed H3K4me3 enrichment at their promoters.
Figure 2. Genes regulated by super-enhancers harbor H3K4me3 broad domains in hippocampal neurons.
A) Boxplot of RNA expression levels for a control gene set, E-reg genes, and SE-reg genes. Control peaks were a set of genes of the same number as the E-reg gene set that was randomly selected from genes that were not part of the E-reg or SE-reg gene sets. B) Density plot showing the distribution of H3K4me3 breadth values for control genes, E-reg genes, and SE-reg genes. * = p < 0.05, ** = p < 0.01, *** = p < 1e-10 by Wilcoxon rank sum with Bonferroni correction.
3.4. Dynamic regulation of H3K4me3 in CA1 neurons at transcriptionally activated genes during memory formation
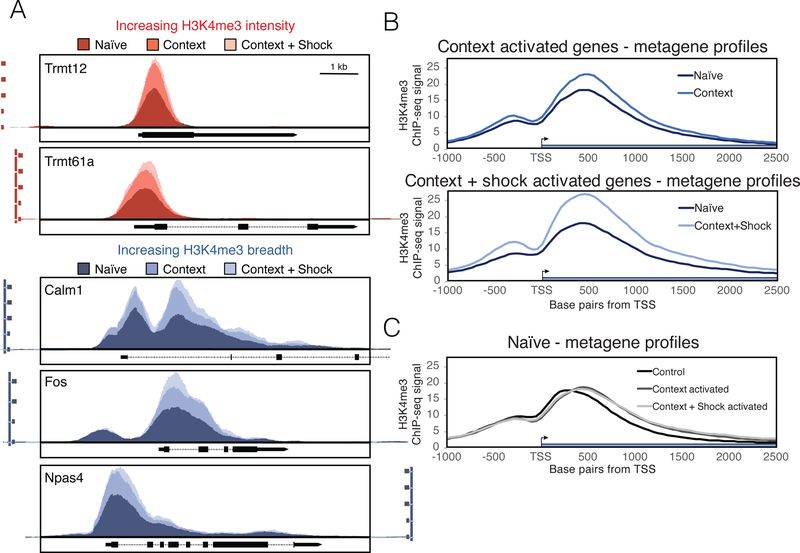
CFC requires memory formation pathways in the hippocampus. During CFC, animals are subdivided into three groups: naïve mice that stay in the home cage, context mice that are moved to the new environment, and context plus shock mice that move to the new environment and receive a foot shock. In our re-analysis of previously published H3K4me3 ChIP-seq data generated from CA1 neurons 1 hour after CFC (Halder et al., 2016), we found examples of genes that had increased H3K4me3 intensity at their promoters, but sustained a constant width throughout the different behavioral treatments, such as Trmt12 and Trmt61a (Figure 3A, top). These two genes were selected because they were some of the genes exhibiting the highest increase in intensity in our genome-wide analysis of H3K4me3 peaks. We also saw many genes involved in memory formation that showed both increased H3K4me3 breadth and intensity in context and context plus shock relative to naïve (Figure 3A, bottom). At these genes, peaks maintain constant boundaries upstream of the promoter, and signal increases in the gene body. An example of a gene showing this pattern is Calm1, which encodes calmodulin, a factor that is involved in memory formation (Limback-Stokin et al., 2004). Fos and Npas4, two well-known immediate early genes (Minatohara et al., 2015; Sun et al., 2016), also show broadening, as indicated by increased H3K4me3 ChIP-seq signal in their gene bodies. Traditional analysis using peak callers failed to detect changes at these genes (Halder et al., 2016).
Figure 3. H3K4me3 breadth is dynamically regulated at gene promoters in CA1 neurons by CFC.
A) Tracks of representative genes increasing in H3K4me3 intensity (top) and breadth (bottom) with context and context plus shock relative to naïve. Scale is normalized across conditions by sequencing library size, and data was binned into 20 base pair windows. Horizontal black line marked 1 kb gives scale for all gene tracks. B) Metagene profiles averaging signal of H3K4me3 across all significantly activated genes relative to naïve from RNA-seq in context (top) or context + shock (bottom). C) Metagene profiles of H3K4me3 signal in naïve mice at genes that become activated in context or context plus shock relative to a control group of genes that was randomly selected from genes with no significant expression change in RNA-seq.
Since our handful of learning- and memory-associated genes that increase in breadth are transcriptionally activated during CFC, we looked at peak distribution of all significantly activated genes, as defined from the RNA-seq data from the same paper (Halder et al., 2016). In context- and context plus shock-activated genes, we see broadening of H3K4me3 into the gene body (Figure 3B), and this is increase is in breadth statistically significant (Supplementary Figure 2A). Intensity at these peaks is significantly increased in context plus shock, but not context relative to controls at these activated genes (Supplementary Figure 2B). We also observed that for genes activated during CFC, their baseline breadth (or breadth in naïve animals) is wider than other genes (Figure 3C and Supplementary Figure 2C), but their baseline intensity is not any different (Supplementary Figure 2D), suggesting that broad domains are poised for activation. In our subsequent analysis, we looked more closely at the functions of genes with increases and decreases in breadth, and corollary transcriptional changes during CFC.
3.5. Broad and broadening genes during CFC have increased expression and are involved in neuroplasticity
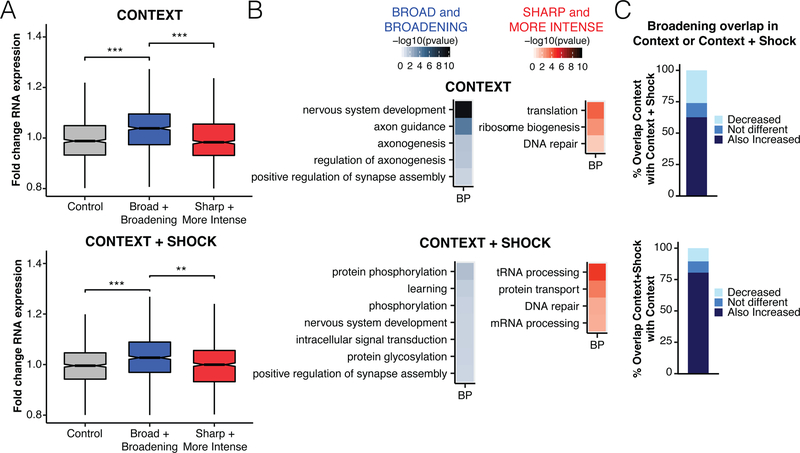
To analyze the impact of peak shape and intensity on expression, we looked at genes that are broad and broaden further during CFC and compared them to sharp and more intensely marked genes. We did this in addition to the analysis of transcriptionally activated genes because the RNA-seq experiment was not cell-type specific, so looking at genes solely based on the ChIP-seq data gives a better picture of what is happening in neurons. Genes we identified by looking for increases in H3K4me3 breadth at already broad genes after exposure to context or context plus shock also increase in RNA expression (Figure 4A). Genes with sharp peaks that are more intensely marked by H3K4me3 are not transcriptionally increased during CFC. Furthermore, the biological roles of the genes that gain H3K4me3 in these different patterns are distinct. Broadening genes in context vs. naïve seem to have almost the exact functions to the top most-broad genes in general (Figure 4B top left vs. Figure 1E), whereas context plus shock broadening genes are most highly associated with protein phosphorylation and learning (Figure 4B bottom left). Genes that become more intensely marked are involved in translation, DNA repair, and protein transport in context and context plus shock (Figure 4B right).
Figure 4. Genes exhibiting increased H3K4me3 breadth with CFC are learning-related and highly induced.
A) Boxplots of expression changes relative to naïve for broad and broadening genes, sharp and more intense genes, and a control gene set of the same size. B) GO analysis of broad and broadening and sharp and more intense gene sets for context (top) and context plus shock (bottom) conditions. Biological process (BP) terms meeting significance level of Bonferroni-corrected p < 0.05 are shown. Heatmaps reflect −log10 of Bonferroni-corrected p-value. C) Bar plots of agreement between broad and broadening genes with context or context plus shock relative to naïve. If context or context plus shock peak breadth was less than 25 base pairs different up or downstream of the naïve peak boundary, it was classified as not different. ** = p < 1e-9, *** = p < 1e-15 by Wilcoxon rank sum with Bonferroni correction.
Broadly marked genes increase in breadth and intensity in context plus shock, so we compared broad and broadening genes to broad and more intense genes in CFC. Whether we defined H3K4me3 increases by intensity or breadth, we see increased expression at broadly marked genes (Supplementary Figure 3A). Many of the genes we pull out based on the two different metrics overlap, so like broadening genes, GO analysis of the intensifying broad gene group identifies terms related to synaptic plasticity (Supplementary Figure 3B). Sharp genes with increased breadth, by contrast, do not identify learning-associated GO terms (Supplementary Figure 3C).
Previous studies show that while DNA methylation may be different in context vs. context plus shock conditions, few changes in expression of mRNA are observed between the two conditions (Duke et al., 2017). We see that H3K4me3 breadth corresponds more to mRNA expression level in that we see a high level of similarity in changes in breadth between context and context plus shock (Figure 4C).
3.6. H3K4me3 at repressed genes and genes with decreased breadth during CFC
Despite H3K4me3 being globally increased (Gupta et al., 2010), we wanted to know if decreasing H3K4me3 intensity or breadth is involved in transcriptional repression. Therefore, we looked at H3K4me3 at transcriptionally repressed genes in CFC, and found that breadth either stays the same (in context) or increases (in context plus shock; Supplementary Figure 4A and B), suggesting that narrowing breadth is not associated with decreased transcription. Even repressed genes show increases in H3K4me3 intensity at their promoters (fold change > 1). However, relative to controls, context plus shock shows a smaller fold change increase in peak intensity compared to naïve (Supplementary Figure 4C). Genes that are repressed are slightly wider in naïve than control genes (Supplementary Figure 4D and E), but their baseline intensity is not any different than controls (Supplementary Figure 4F).
We also identified narrowing domains, and looked at their functions (Supplementary Figure 5). We do not observe very large magnitudes of expression changes despite identifying some genes with narrowing or decreased intensity of H3K4me3 at the gene promoter. However, less intense sharp peaks in context plus shock are observed to have significantly decreased expression (Supplementary Figure 5A). Not many GO terms are retrieved in narrowing and less intense genes. Narrowing peaks are predominantly involved in nervous system development in context and context plus shock conditions relative to naïve, and in the context plus shock condition, genes associated with GABAergic synaptic transmission narrowed (Supplementary Figure 5B). Again, we see a high level of agreement between genes that narrow in context and context plus shock conditions (Supplementary Figure 5C).
3.7. Kmt2a and Kmt2b regulate H3K4me3 breadth in CA1 neurons
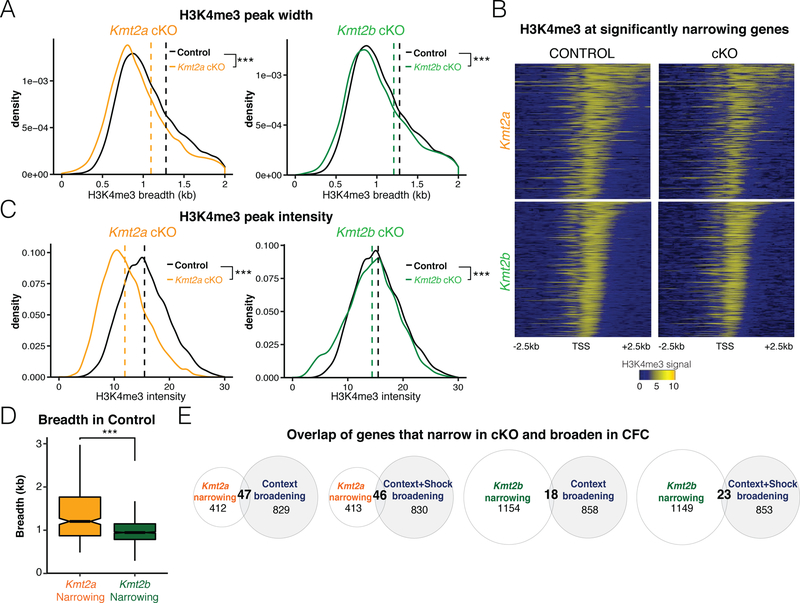
H3K4me3 is deposited by six related methyltransferase isoforms, and little is known about which ones contribute to the regulation of H3K4me3 breadth in CA1 neurons. Conditional knock-out (cKO) mice had been previously generated for Kmt2a and Kmt2b genes using Camkii-cre to delete these genes in the forebrain, and while ChIP-seq of H3K4me3 was conducted on naïve cKO mice, peak breadth was never analyzed (Kerimoglu et al., 2017). Importantly, both lines of these cKO mice exhibit decreased learning in behavioral tasks, including decreased freezing during fear conditioning (Kerimoglu et al., 2013; Kerimoglu et al., 2017). The same CA1 dissection and nuclei sorting methodology as was used as in the fear conditioning experimental datasets (Halder et al., 2016) (Figure 1A). These lysine methyltransferases add methylation to histones, so if they were regulators of peak breadth, we would predict that their knockout would result in narrowing of H3K4me3 peaks. We subjected these data sets to H3K4me3 breadth analysis, and analyzing all genes across the genome with a H3K4me3 peak at their promoters, we observe that peak breadth narrows in both Kmt2a and Kmt2b cKO compared to controls (Figure 5A), and Kmt2a cKO shows a stronger magnitude of narrowing. Looking at the significantly narrowing genes in heatmaps, the Kmt2a cKO shows visible decrease in signal in the gene body portion of the peak (Figure 5B). As previously reported, cKO of both genes decreases overall peak intensity, and our analysis of peaks confirms this finding (Figure 5C). Taken together, this data suggests that KMT2A and KMT2B maintain H3K4me3 intensity and breadth. We analyzed whether peak breadth in control animals is significantly wider at genes that narrow after Kmt2a cKO because of the patterns observed in the heatmaps, and found that Kmt2a cKO narrows H3K4me3 peaks at genes that are normally marked by broader peaks (Figure 5D). Since CFC tends to also cause changes in broad genes, we looked at whether the lysine methyltransferases regulate peak breadth at the same genes that change in breadth during CFC. Knockout of Kmt2a causes narrowing at a larger proportion of genes regulated in CFC than Kmt2b (Figure 5E).
Figure 5. H3K4 methyltransferases Kmt2a and Kmt2b globally regulate H3K4me3 breadth in hippocampal neurons.
A) Density plots of H3K4me3 breadth for all genes in control, Kmt2a cKO, and Kmt2b cKO mice. B) Genes with significant narrowing of H3K4me3 breadth in Kmt2a cKO (top right panel) or Kmt2b cKO (bottom right panel) relative to control mice (left panels). C) Density plots of H3K4me3 intensity in control, Kmt2a cKO, and Kmt2b cKO mice. D) Breadth of peaks in control mice that narrow after cKO of Kmt2a or Kmt2b. E) Overlap of genes that narrow after cKO of Kmt2a or Kmt2b and genes that broaden in context or context plus shock relative to naïve. *** = p < 1e-15 by Wilcoxon rank sum.
4. Discussion
Our results support a model whereby changes to the shape of peaks made by epigenetic marks help neurons encode learned information. In general, studies are only recently recognizing the importance of the distribution width of epigenetic modifications to histones and their implications in different biological processes. H3K4me3 breadth is emerging as one of the most relevant aspects of this mark and has been attributed to determining cell identity and regulation of transcription. Because the aspect of breadth has been long overlooked by studies of H3K4me3 in learning and memory, we were able to identify its importance in marking pertinent genes using archived datasets.
H3K4me3 marks are able to cause transcriptional activation. Increasing total H3K4me3 on nucleosomes increases RNA output in in vitro transcription reactions (Lauberth et al., 2013), and loss of Kmt2a, which deposits H3K4me3, leads to abnormal distribution of Pol II across the genome (Milne et al., 2005). Future studies are needed to see if H3K4me3 intensity or breadth are the cause of increased transcription in neurons, and this could be addressed using genome-targeting technologies like transcription-activator-like effectors (TALE) or CRISPR/nuclease-deficient Cas9 systems to direct H3K4 methyltransferases or demethylases to promoters or gene bodies (Tost, 2016).
From our analysis of H3K4me3 breadth in data from conditional knockout mice, we find that KMT2A and KMT2B regulate H3K4me3 breadth, and their knockout elicited significantly decreased breadth. KMT2A and KMT2B regulated breadth at different genes, and KMT2A regulates breadth at genes with broader H3K4me3 to begin with. The disparities in the genes the two isoforms regulate the breadth of could be why slightly different memory phenotypes are seen in the two knockouts. cKO of both factors affects memory, but CFC tests and Morris water maze tests appear to have more severe impairment in Kmt2a cKO compared to Kmt2b cKO animals relative to controls, though these two cKO mice were not tested side-by-side. Another difference between the cKO animals is that Kmtb2 cKO, but not Kmt2a cKO, show deficits in object recognition memory (Kerimoglu et al., 2013; Kerimoglu et al., 2017). These differences in behavioral phenotype highlight distinct roles of each isoform in proper memory formation. Moreover, we found that Kmt2a cKO changes breadth at more genes regulated during fear conditioning than Kmt2b, but because we only have ChIP-seq of naïve cKO mice for these two methyltransferases, more work is needed to see if the two isoforms are responsible for H3K4me3 broadening during memory formation.
While these Kmt2a and Kmt2b are likely candidates for controlling H3K4me3 breadth in the hippocampus because of their ascribed functions in memory formation, four other methyltransferases with activity towards H3K4me3 are also expressed in this brain region. Indeed, Kmt2d knockout in the brain also leads to decreased H3K4me3 at broad peaks (Dhar et al., 2018). However, the genes it affects are predominantly tumor-suppressor genes, and knocking out Kmt2d leads to increased brain tumor formation. Interestingly, Kmt2d knockout also affected super-enhancer activity, and we likewise see a relationship between H3K4me3 breadth in CA1 neurons and super-enhancer regulation. From this and our data, it seems likely that the other H3K4 methyltransferases could affect H3K4me3 peak breadth at certain promoters. Potential difficulties in discriminating the contribution of each isoform to regulating breadth will be redundancies in their function and possible compensatory changes in other isoforms when one is knocked out.
Mutations in demethylases of H3K4 also lead to intellectual disability (Collins et al., 2019), and we found some genes with associations to plasticity have decreased H3K4me3 breadth after CFC. Therefore, it is likely that demethylases play a role in regulating peak breadth during memory formation, as well. It would also be interesting to address if any of the demethylases contribute to returning the peak breadth to baseline levels in CA1 neurons after the memory is encoded.
Broad H3K4me3 domains demarcate developmental genes in neurons of the CA1 region of the hippocampus, as well as genes involved in learning. We found that these domains are associated with RNA expression level, rather than transcriptional consistency. Furthermore, we show that peak breadth is increased at important learning-associated genes during memory formation, and these changes correlate with changes in RNA expression. During development, broad H3K4me3 domains mark regions that will be activated after differentiation (Grandy et al., 2016). In a similar fashion, we see that broad H3K4me3 domains mark regions that will be acutely activated during memory formation and become broader after stimuli exposure. Broad domains seem to be poised for activation, and this may explain why we see a high level of variance in their expression.
Supplementary Material
Broad domains of the activating H3K4me3 mark are found at many genes in CA1 neurons
These broad domains mark promoters of neurodevelopmental and learning genes
Broad domains are found at super-enhancer regulated genes
3K4me3 widens at learning-associated genes during contextual fear conditioning
Memory-associated genes, Kmt2a and Kmt2b, regulate H3K4me3 breadth
Acknowledgements
We thank Qin Yan for initiating our interest in analyzing peak breadth, and Roger Colbran, Colleen Niswender, and Danny Winder for helpful feedback on the manuscript.
Funding
This work was supported by grants from the National Institutes of Health [grant numbers MH091122, MH057014, T32GM007347, and T32MH065215]. The content is solely the responsibility of the authors and does not necessarily reflect the official views of the National Institutes of Health.
Abbreviations
- H3K4me3
histone 3 lysine 4 trimethylation
- Pol II
RNA polymerase II
- E-reg
enhancer-regulated
- SE-reg
super-enhancer-regulated
- CFC
contextual fear conditioning
- kb
kilobase
- TSS
transcription start site
- ChIP-seq
chromatin immunoprecipitation followed by sequencing
- DAVID
Database for Annotation, Visualization, and Integrated Discovery
- GO
gene ontology
- BP
biological process
- Kmt2a
lysine methyltransferase 2a
- Kmt2b
lysine methyltransferase 2b
- RPKM
Reads Per Kilobase of transcript per Million mapped reads
- FC
fold change
- WT
wild-type
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Competing Interests
The authors have no conflicts of interest to declare.
References
- Agranoff BW, Davis RE, & Brink JJ (1965). Memory fixation in the goldfish. Proc Natl Acad Sci U S A, 54, 788–793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, & Zhao K (2007). High-resolution profiling of histone methylations in the human genome. Cell, 129, 823–837. [DOI] [PubMed] [Google Scholar]
- Benayoun BA, Pollina EA, Ucar D, Mahmoudi S, Karra K, Wong ED, Devarajan K, Daugherty AC, Kundaje AB, Mancini E, Hitz BC, Gupta R, Rando TA, Baker JC, Snyder MP, Cherry JM, & Brunet A (2014). H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell, 158, 673–688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, Chen Z, Wu D, Zhang L, Lin X, Su J, Rodriguez B, Xi Y, Xia Z, Chen X, Shi X, Wang Q, & Li W (2015). Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor-suppressor genes. Nat Genet, 47, 1149–1157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins BE, Greer CB, Coleman BC, & Sweatt JD (2019). Histone H3 lysine K4 methylation and its role in learning and memory. Epigenetics Chromatin, 12, 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Core LJ, Martins AL, Danko CG, Waters CT, Siepel A, & Lis JT (2014). Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat Genet, 46, 1311–1320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhar SS, Zhao D, Lin T, Gu B, Pal K, Wu SJ, Alam H, Lv J, Yun K, Gopalakrishnan V, Flores ER, Northcott PA, Rajaram V, Li W, Shilatifard A, Sillitoe RV, Chen K, & Lee MG (2018). MLL4 Is Required to Maintain Broad H3K4me3 Peaks and Super-Enhancers at Tumor Suppressor Genes. Mol Cell, 70, 825–841 e826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dincer A, Gavin DP, Xu K, Zhang B, Dudley JT, Schadt EE, & Akbarian S (2015). Deciphering H3K4me3 broad domains associated with gene-regulatory networks and conserved epigenomic landscapes in the human brain. Transl Psychiatry, 5, e679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duke CG, Kennedy AJ, Gavin CF, Day JJ, & Sweatt JD (2017). Experience-dependent epigenomic reorganization in the hippocampus. Learn Mem, 24, 278–288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandy RA, Whitfield TW, Wu H, Fitzgerald MP, VanOudenhove JJ, Zaidi SK, Montecino MA, Lian JB, van Wijnen AJ, Stein JL, & Stein GS (2016). Genome-Wide Studies Reveal that H3K4me3 Modification in Bivalent Genes Is Dynamically Regulated during the Pluripotent Cell Cycle and Stabilized upon Differentiation. Mol Cell Biol, 36, 615–627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta S, Kim SY, Artis S, Molfese DL, Schumacher A, Sweatt JD, Paylor RE, & Lubin FD (2010). Histone methylation regulates memory formation. J Neurosci, 30, 3589–3599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haeussler M, Zweig AS, Tyner C, Speir ML, Rosenbloom KR, Raney BJ, Lee CM, Lee BT, Hinrichs AS, Gonzalez JN, Gibson D, Diekhans M, Clawson H, Casper J, Barber GP, Haussler D, Kuhn RM, & Kent WJ (2019). The UCSC Genome Browser database: 2019 update. Nucleic Acids Res, 47, D853–D858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halder R, Hennion M, Vidal RO, Shomroni O, Rahman RU, Rajput A, Centeno TP, van Bebber F, Capece V, Garcia Vizcaino JC, Schuetz AL, Burkhardt S, Benito E, Navarro Sala M, Javan SB, Haass C, Schmid B, Fischer A, & Bonn S (2016). DNA methylation changes in plasticity genes accompany the formation and maintenance of memory. Nat Neurosci, 19, 102–110. [DOI] [PubMed] [Google Scholar]
- Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, & Glass CK (2010). Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell, 38, 576–589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henriques T, Scruggs BS, Inouye MO, Muse GW, Williams LH, Burkholder AB, Lavender CA, Fargo DC, & Adelman K (2018). Widespread transcriptional pausing and elongation control at enhancers. Genes Dev, 32, 26–41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, Hoke HA, & Young RA (2013). Super-enhancers in the control of cell identity and disease. Cell, 155, 934–947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, & Lempicki RA (2009a). Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res, 37, 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang da W, Sherman BT, & Lempicki RA (2009b). Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc, 4, 44–57. [DOI] [PubMed] [Google Scholar]
- Jakovcevski M, Ruan H, Shen EY, Dincer A, Javidfar B, Ma Q, Peter CJ, Cheung I, Mitchell AC, Jiang Y, Lin CL, Pothula V, Stewart AF, Ernst P, Yao WD, & Akbarian S (2015). Neuronal Kmt2a/Mll1 histone methyltransferase is essential for prefrontal synaptic plasticity and working memory. J Neurosci, 35, 5097–5108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji J, & Maren S (2008). Differential roles for hippocampal areas CA1 and CA3 in the contextual encoding and retrieval of extinguished fear. Learn Mem, 15, 244–251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joo JY, Schaukowitch K, Farbiak L, Kilaru G, & Kim TK (2016). Stimulus-specific combinatorial functionality of neuronal c-fos enhancers. Nat Neurosci, 19, 75–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karolchik D, Hinrichs AS, Furey TS, Roskin KM, Sugnet CW, Haussler D, & Kent WJ (2004). The UCSC Table Browser data retrieval tool. Nucleic Acids Res, 32, D493–496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerimoglu C, Agis-Balboa RC, Kranz A, Stilling R, Bahari-Javan S, Benito-Garagorri E, Halder R, Burkhardt S, Stewart AF, & Fischer A (2013). Histone-methyltransferase MLL2 (KMT2B) is required for memory formation in mice. J Neurosci, 33, 3452–3464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerimoglu C, Sakib MS, Jain G, Benito E, Burkhardt S, Capece V, Kaurani L, Halder R, Agis-Balboa RC, Stilling R, Urbanke H, Kranz A, Stewart AF, & Fischer A (2017). KMT2A and KMT2B Mediate Memory Function by Affecting Distinct Genomic Regions. Cell Rep, 20, 538–548. [DOI] [PubMed] [Google Scholar]
- Kim D, Langmead B, & Salzberg SL (2015). HISAT: a fast spliced aligner with low memory requirements. Nat Methods, 12, 357–360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, Markenscoff-Papadimitriou E, Kuhl D, Bito H, Worley PF, Kreiman G, & Greenberg ME (2010). Widespread transcription at neuronal activity-regulated enhancers. Nature, 465, 182–187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, & Salzberg SL (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods, 9, 357–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lauberth SM, Nakayama T, Wu X, Ferris AL, Tang Z, Hughes SH, & Roeder RG (2013). H3K4me3 interactions with TAF3 regulate preinitiation complex assembly and selective gene activation. Cell, 152, 1021–1036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leinonen R, Sugawara H, Shumway M, & International Nucleotide Sequence Database, C. (2011). The sequence read archive. Nucleic Acids Res, 39, D19–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, & Genome Project Data Processing, S. (2009). The Sequence Alignment/Map format and SAMtools. Bioinformatics, 25, 2078–2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Y, Smyth GK, & Shi W (2014). featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics, 30, 923–930. [DOI] [PubMed] [Google Scholar]
- Limback-Stokin K, Korzus E, Nagaoka-Yasuda R, & Mayford M (2004). Nuclear calcium/calmodulin regulates memory consolidation. J Neurosci, 24, 10858–10867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lubin FD, Gupta S, Parrish RR, Grissom NM, & Davis RL (2011). Epigenetic mechanisms: critical contributors to long-term memory formation. Neuroscientist, 17, 616–632. [DOI] [PubMed] [Google Scholar]
- Milne TA, Dou Y, Martin ME, Brock HW, Roeder RG, & Hess JL (2005). MLL associates specifically with a subset of transcriptionally active target genes. Proc Natl Acad Sci U S A, 102, 14765–14770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minatohara K, Akiyoshi M, & Okuno H (2015). Role of Immediate-Early Genes in Synaptic Plasticity and Neuronal Ensembles Underlying the Memory Trace. Front Mol Neurosci, 8, 78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinlan AR, & Hall IM (2010). BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics, 26, 841–842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, & Smyth GK (2010). edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26, 139–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, & Kouzarides T (2002). Active genes are tri-methylated at K4 of histone H3. Nature, 419, 407–411. [DOI] [PubMed] [Google Scholar]
- Sun X, & Lin Y (2016). Npas4: Linking Neuronal Activity to Memory. Trends Neurosci, 39, 264–275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tost J (2016). Engineering of the epigenome: synthetic biology to define functional causality and develop innovative therapies. Epigenomics, 8, 153–156. [DOI] [PubMed] [Google Scholar]
- Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, & Young RA (2013). Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell, 153, 307–319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zerbino DR, Achuthan P, Akanni W, Amode MR, Barrell D, Bhai J, Billis K, Cummins C, Gall A, Giron CG, Gil L, Gordon L, Haggerty L, Haskell E, Hourlier T, Izuogu OG, Janacek SH, Juettemann T, To JK, Laird MR, Lavidas I, Liu Z, Loveland JE, Maurel T, McLaren W, Moore B, Mudge J, Murphy DN, Newman V, Nuhn M, Ogeh D, Ong CK, Parker A, Patricio M, Riat HS, Schuilenburg H, Sheppard D, Sparrow H, Taylor K, Thormann A, Vullo A, Walts B, Zadissa A, Frankish A, Hunt SE, Kostadima M, Langridge N, Martin FJ, Muffato M, Perry E, Ruffier M, Staines DM, Trevanion SJ, Aken BL, Cunningham F, Yates A, & Flicek P (2018). Ensembl 2018. Nucleic Acids Res, 46, D754–D761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, & Liu XS (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biol, 9, R137. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.