Abstract
Background
Habitats colonized by acidophiles as an ideal physical barrier may induce genetic exchange of microbial members within the common communities, but little is known about how species in extremely acidic environments diverge and evolve.
Results
Using the acidophilic sulfur-oxidizer Acidithiobacillus as a case study, taxonomic reclassifications of many isolates provides novel insights into their phylogenetic lineage. Whole-genome-based comparisons were attempted to investigate the intra- and inter-species divergence. Recent studies clarified that functional and structural specificities of bacterial strains might provide opportunities for adaptive evolution responding to local environmental conditions. Acidophilic microorganisms play a key role in the acidification of natural waters and thus the formation of extremely acidic environments, and the feedbacks of the latter might confer the distinct evolutionary patterns of Acidithiobacillus spp. Varied horizontal gene transfer events occurred in different bacterial strains, probably resulting in the expansion of Acidithiobacillus genomes. Gene loss as another evolutionary force might cause the adaptive phenotypic diversity. A conceptual model for potential community-dependent evolutionary adaptation was thus proposed to illustrate the observed genome differentiation.
Conclusions
Collectively, the findings shed light on the phylogeny and divergent evolution of Acidithiobacillus strains, and provided a useful reference for evolutionary studies of other extremophiles.
Electronic supplementary material
The online version of this article (10.1186/s12864-019-5827-6) contains supplementary material, which is available to authorized users.
Keywords: Acidithiobacillus, Comparative genomics, Phylogeny, Divergent evolution
Background
Knowledge about physiological and ecological constraints to organismal dispersal has extended our understanding of geographic barriers that limit gene flow across species in different regions. For macroorganisms, generally, dispersal barriers hinder the genetic exchange between species in different geographical areas, and local evolutionary processes such as genetic drift and econiche adaptation contribute to endemism [1], the equivalent of term ‘precinctive’ that refers to species in the restricted geographic location. To the contrary, microbial life including single-celled organisms are small-size and dispersed easily, and their biogeographical distributions are traditionally regarded to be a result of environmental selection (quoted by the old microbiological tenet ‘Everything is everywhere, but, the environment selects’) rather than dispersal limitation and stochasticity [2]. In recent years, however, considerable efforts have been attempted to elucidate a high degree of microbial endemicity [3–6]. Earlier studies have emphasized the importance of geographical isolation in the diversification and evolution of microorganisms on a global scale [7, 8]. Spatial separation, to a large extent, significantly affects the allopatric speciation in microbial world. In some isolated econiches, microbial species with low capacities to acquire alien genes, such as Buchnera aphidicola, was reported to exhibit an extreme genome stability [9], suggesting the effect of dispersal limitation on microbial speciation and evolution. Numerous studies pertaining to microbial endemicity have been made, yet knowledges about how geography influences divergence, evolution, and speciation of microorganisms remain elusive.
Acidithiobacillus, type genus of the family Acidithiobacillaceae, are recognized as sulfur-oxidizing acidophiles, and ubiquitously found in extremely acidic environments with heavy metal-laden and nutrient-deficient conditions, although these adverse settings are inhospitable to most life forms [10]. According to the public database List of Prokaryotic names with Standing in Nomenclature (LPSN, which is available at http://www.bacterio.net), several validated species of genus Acidithiobacillus have been described, including Acidithiobacillus thiooxidans (type species of this genus; formerly Thiobacillus thiooxidans) [11, 12], Acidithiobacillus albertensis (formerly Thiobacillus albertis) [13], Acidithiobacillus caldus (formerly Thiobacillus caldus) [14], Acidithiobacillus ferrooxidans (formerly Thiobacillus ferrooxidans) [15], Acidithiobacillus ferrivorans [16], Acidithiobacillus ferridurans [17], and Acidithiobacillus ferriphilus [18]. So far, numerous 16S rRNA gene sequences and draft/complete genome sequences of Acidithiobacillus spp. are available in public databases, thanks to the development of high-throughput sequencing technology.
Members of genus Acidithiobacillus are dominant in diverse sulfur-rich environments worldwide, and they are believed to play key roles in the biogeochemical cycle of sulfur and/or iron [19]. The metabolic activities of acidophiles in both pristine and anthropogenic sulfur-laden econiches contribute to the acidification of natural waters and the formation of extremely acidic habitats [20, 21]. Geochemical conditions of hyperacidic environments (pH < 3) are significantly different from that of surrounding regions, it might provide a potential geographic barrier that limits gene dispersal between acidophilic microorganisms, i.e., Acidithiobacillus populations, and other organisms in the surroundings. In other words, extremely acidic environments as a physical barrier might limit the access of indigenous genes to the global microbial gene pool, and hence increase the frequency of gene drift across the common biological communities. Here, we therefore hypothesized that spatial separation and geochemical isolation would cause the formation of separate ecosystems, which were stochastically colonized by various ancestral species, and a potential community-dependent evolutionary model was developed to delineate the allopatric divergence of Acidithiobacillus spp., which probably reflect the species-specific adaption to local environmental conditions.
Methods
Species selection used in this study
In the last several decades, a vast number of Acidithiobacillus strains have been isolated from a diverse range of sulfur-abundant environments worldwide [22], and plentiful 16S rRNA gene sequence dataset has been acquired on the basis of environmental sampling [23]. However, the taxonomic assignments of many of these strains and sequence clones are as yet unclear, which confuses the understanding of evolutionary lineage of Acidithiobacillus spp. Accordingly, it is necessary for these unclassified and cryptic species to conduct a more exhaustive revision of the phylogenetic taxon [21].
The genus (Acidi)thiobacillus is presently composed of seven identified species, dating back to 1922. In this study, the available genomes of Acidithiobacillus strains were downloaded from GenBank database, and 16S rRNA gene sequences of type strains were extracted from individual genomes using the RNAmmer 1.2 Server [24]. As listed in Additional file 1: Table S1, type strains contain A. ferrooxidans ATCC 23270T (NC_011761), A. caldus ATCC 51756T (NZ_CP005986), and A. albertensis DSM 14366T (MOAD00000000). In addition, A. thiooxidans ATCC 19377T (AFOH00000000) was included in this study. Given the lack of genomic sequence of A. ferrivorans DSM 22755T, strain SS3 (NC_015942) was used as a reference. In the absence of genomic sequences of A. ferridurans ATCC 33020T and A. ferriphilus DSM 100412T, the partial 16S rRNA gene sequences were acquired according to LPSN, accession numbers of which were AJ278719 and KR905751, respectively. Apart from type strains, other existing complete/draft genomes of Acidithiobacillus strains deposited in public database were also used for the extraction of 16S rRNA gene sequences and then for the phylogenetic analysis. Referred to the length of above-mentioned sequences (ranging from 1,323 bp to 1,527 bp), 572 additional 16S rRNA gene sequences of Acidithiobacillus spp. were obtained from GenBank database, fitting the sequence length (ranging from 1,300 bp to 1,550 bp) and non-redundancy requirements.
Analyses of phylogeny and taxonomy
Using a broader dataset that comprises up to 602 16S rRNA gene sequences of Acidithiobacillus spp. (Additional file 2: Table S2), sequence alignment was conducted using the online service MAFFT v7.402 [25] with a FFT-NS-2 method. The software Mesquite v3.51 was applied for the conversion of FASTA format to PHYLIP format. Given that saturation in substitutions could lead to incorrect phylogenetic inferences [26], saturation test was thus performed to evaluate the number of transitions and transversions against the Tamura-Nei (TN93) [27] genetic distance using DAMBE v5.2.73 [28], referred to a previous study [29]. Maximum-likelihood phylogeny was constructed using the online PhyML 3.0 [30] with the following settings: nucleotide substitution model, TN93; number of substitution rate categories, 6; and tree search algorithm, subtree pruning and regrafting. Visualization for 16S rRNA gene-based phylogenetic tree was performed using the program FigTree v1.4.3 (available at http://tree.bio.ed.ac.uk/software/figtree/).
As of July 2018, many genomes of Acidithiobacillus spp. have been released and deposited at the GenBank database, including genomes from five defined species and some other unclassified strains. Acidithiobacillus genomes have been automatically processed by NCBI prokaryotic genome annotation pipeline, such as gene prediction and functional annotation. In this study, 28 of bacterial genomes were included. Detailed genome information was illustrated in Additional file 1: Figure S1, such as accession numbers and genome sizes. Alignment-free phylogeny based on concatenated protein sequences of Acidithiobacillus genomes was constructed using the web server CVTree3 [31] with K-tuple length 6. Herein, Achromobacter xylosoxidans A8 was used as outgroup. The phylogenomic tree was then visualized using MEGA5.
Recently, average nucleotide identity (ANI) approach [32] was developed to substitute the conventional laboratory-based DNA-DNA hybridization, a gold standard for prokaryotic species identification. In our study, all 28 complete and/or draft genomes of Acidithiobacillus strains (Additional file 1: Figure S1) were used for the evaluation of their genomic similarity. Pairwise comparisons of bacterial ANI based on BLAST (ANIb) and MUMmer algorithm (ANIm) were performed using the online service JSpeciesWS [33] with default settings.
Identification of orthologous and non-orthologous genes
To identify the putative orthologous and non-orthologous genes among Acidithiobacillus strains, BLASTP all-versus-all comparisons of entire amino acid sequences between pairs of Acidithiobacillus strains were conducted. In order to further explore the intra- and inter-species diversity of Acidithiobacillus spp., the computational tool Bacterial Pan Genome Analysis (BPGA) [34] was then applied to determine the orthologs and non-orthologs, including orthologous genes that were shared by all tested genomes or a subset of bacterial genomes, and strain-specific genes. In this procedure, we implemented the following parameters: E-value cut-off, 1e-5; and sequence identity cut-off, 50%. Giving the lineage-specific expansions of mobile genetic elements [35], these genes were excluded prior to the identification of orthologs. We finally performed the manual inspection and correction of all results. CVTree3 were employed to construct the phylogenetic tree, which was based on orthologous proteins shared by all Acidithiobacillus strains.
Functional assignments of core, accessory, and unique genes among Acidithiobacillus strains were performed via BLASTP alignment against specialized database, i.e., the extended Clusters of Orthologous Groups (COG) [36], with E-value cut-off set to 1e-5. The BLAST results were then manually checked based on the highest hit coverage value. Additionally, KEGG Automatic Annotation Server (KAAS) [37] was implemented to execute the automatically functional annotation of query amino acid sequences, with an assignment method of single-directional best hit. Statistical calculation was performed to identify the relative abundance of genes assigned to COG categories and KEGG pathways.
Model extrapolation for Acidithiobacillus pan-genome
BPGA pipeline [34] was applied to extrapolate the pan-genome and core genome size based on 28 Acidithiobacillus genomes, as described in previous study [38]. To avoid the bias after any new genomes were added, random permutations were performed in the order of addition of genomes and medians were used to estimate the average number of pan-genomes and core genomes. In this procedure, the size of pan-genome and core genome were extrapolated by fitting the empirical power law equation [Ps(n) = κnγ] and exponential equation [Fc(n) = κcexp(-n/τc) + Ω] respectively, where Ps(n) and Fc(n) denote the calculated pan-genome and core genome sizes respectively, n is the number of sequenced genomes, and k, γ, κc, τc, and Ω are the fitting parameters. The exponent γ < 0 and 0 <γ < 1 indicate that Acidithiobacillus pan-genome is ‘closed’ and ‘open’, respectively.
Calculation for gene turnover of Acidithiobacillus genomes
The program OrthoFinder v2.3.1 [39] with Markov cluster algorithm [40] and Diamond was applied to identify the orthogroups (herein referred as gene families) of Acidithiobacillus genomes. To detect the rates of gene gain and death, we performed the program BadiRate v1.35 [41] with a ‘GD-FR-CWP’ model by counting gain/loss events from the minimum number of family members in the phylogenetic nodes (inferred by the Wagner parsimony algorithm). Additionally, the topology of the whole-genome-based phylogeny was used as the reference tree. In the phylogenetic tree, each branch had its own turnover rate.
Prediction of mobile genetic elements
Amino acid sequences of Acidithiobacillus strains were extracted from individual genomes using in-house Perl script, and putative transposable elements including transposon and insertion sequence (IS) were predicted using an updated version of online tool ISfinder [42] with an E-value cut-off of 1e-5. The web server tRNAscan-SE 2.0 [43] was used to search for the putative tRNA genes. Putative phage-associated genes in the sequenced Acidithiobacillus genomes were detected using the tool Prophinder [44] and BLASTP search against the ACLAME database of proteins in viruses and prophages [45] with the E-value threshold of 1e-3, as described previously [46]. Finally, all results above were manually checked.
Results
Phylogeny of Acidithiobacillus spp.
In our study, a set of 16S rRNA gene sequences were retrieved from GenBank database (Additional file 2: Table S2), according to certain criteria (see section Species selection used in this study). Saturation test was performed prior to the analysis of 16S rRNA gene-based phylogeny of Acidithiobacillus spp. As depicted in Additional file 1: Figure S2, transitions outnumbered transversions, suggesting that substitutions were not saturated and this dataset could be suitable for the subsequent construction of phylogenetic tree.
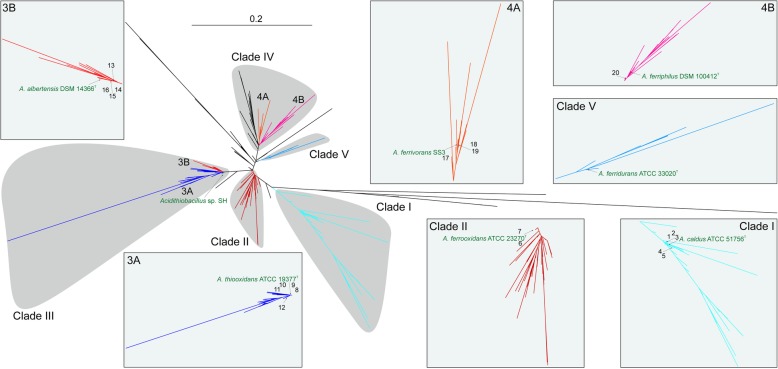
To evaluate the potential evolutionary relationships of all Acidithiobacillus strains and sequence clones, their available 16S rRNA genes as the marker were used for phylogeny. The resulting maximum-likelihood tree presented a coherent picture of Acidithiobacillus lineage (Fig. 1). Obviously, these strains and sequence clones were separately clustered into various distinctive clades, such as A. caldus ATCC 51756T (Clade I), A. ferrooxidans ATCC 23270T (Clade II), A. thiooxidans ATCC 19377T (3A in Clade III), A. albertensis DSM 14366T (3B in Clade III), A. ferrivorans SS3 (4A in Clade IV), A. ferriphilus DSM 100412T (4B in Clade IV), and A. ferridurans ATCC 33020T (Clade V). The finding was slightly different from an earlier study [21] , in which sequences from A. thiooxidans, A. albertensis, and A. ferridurans were gathered into the common clade. Additionally, other sister-clades possibly representing novel species or candidate phylotype, such as subclade within the A. ferrivorans-A. ferriphilus Clade IV (black colors), were shown in the branches of phylogenetic tree, suggesting a considerably underappreciated diversity within the genus Acidithiobacillus. Fascinatingly more, our study provided a revision for the isolates with unclassified and/or conflicting taxonomic assignments (Additional file 2: Table S2) via re-evaluating their phylogenetic relationships. As a result, many Acidithiobacillus spp. (e.g., Acidithiobacillus sp. GGI-221 within A. ferrooxidans Clade II) were assigned to the clades with recognized species, and the potential evolutionary relationships and classifications of several strains and sequence clones with known specific assignment (e.g., A. ferrooxidans BY0502 within A. ferriphilus subclade 4B) were reconfirmed according to our current 16S rRNA gene-based tree. Oddly, A. thiooxidans strains ZBY, DXS-W, CLST, and GD1-3 were apparently clustered into A. albertensis subclade 3B. Similarly, an earlier study revealed a high nucleotide sequence identity of up to 99.9% between A. thiooxidans-A. albertensis pairs [21], which exceeded both typical (97%) and conservative (98.7%) threshold values for species delineation [47, 48].
Fig. 1.
Maximum-likelihood (ML) phylogeny based on 16S rRNA gene sequences of Acidithiobacillus spp. Seven validated species are assigned into five clades, including A. caldus (Clade I), A. ferrooxidans (Clade II), A. thiooxidans (3A in Clade III), A. albertensis (3B in Clade III), A. ferrivorans (4A in Clade IV), A. ferriphilus (4B in Clade IV), and A. ferridurans (Clade V). These species are highlighted in different colors. Several strains of interest in various minor branches are also marked in this phylogenetic tree, including A. caldus strains DX (1), ZJ (2), ZBY (3), SM-1 (4), MTH-04 (5), A. ferrooxidans ATCC 53993 (6), Acidithiobacillus sp. GGI-221 (7), A. thiooxidans strains A01 (8), A02 (9), DMC (10), JYC-17 (11), BY-02 (12), ZBY (13), DXS-W (14), CLST (15), GD1-3 (16), A. ferrivorans strains YL15 (17), PRJEB5721 (18), CF27 (19), A. ferrooxidans BY0502 (20), and Acidithiobacillus sp. SH. Other branches representing unclear species are shown in black color. More details about these 602 16S rRNA gene sequences used in ML tree construction and their clade assignments are listed in Additional file 2: Table S2.
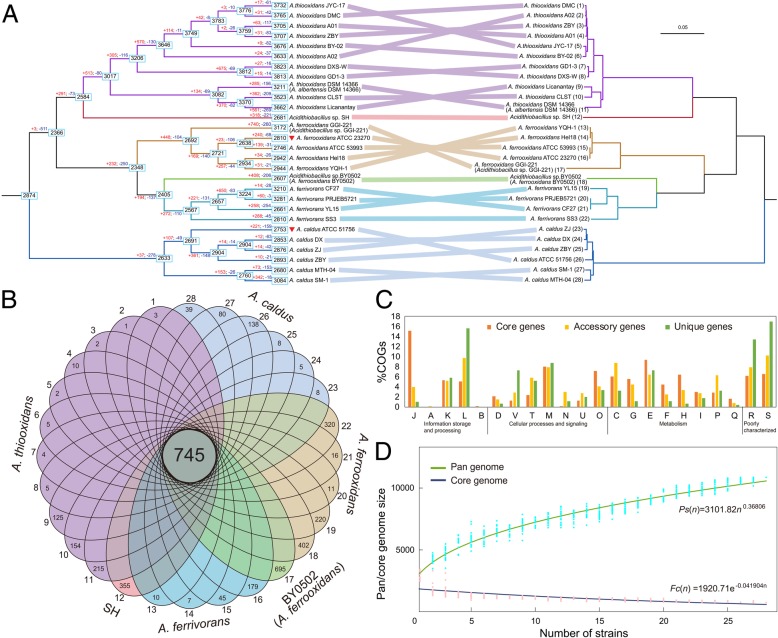
A comprehensive strategy based on Acidthiobacillus genomes was used to improve the phylogenetic resolution and consolidate their evolutionary relationships. Based on 28 Acidthiobacillus genomes, these sequences were expected to be assigned into diverse clades (Fig. 2a). In this phylogenomic tree, strains of each species were clustered but apparently separated from that of the others. Similar to the 16S rRNA gene-based phylogeny, strain GGI-221 was likely to be phylogenetically affiliated to A. ferrooxidans species, and strain BY0502 might be a member of novel species rather than that of the current delineated species, i.e., A. ferrooxidans. Additionally, ANIb and ANIm values further supported the taxonomic reclassifications (Additional file 1: Figure S3). Notably, a striking finding was the affiliation of strain DSM 14366 (formerly belonged to A. albertensis). The values (≥ 96.62%) of ANIb and ANIm between DSM 14366 and each A. thiooxidans strains, accompanied by phylogenomic indicator, strongly indicated that strain DSM 14366 was likely to be affiliated to the type species A. thiooxidans instead of A. albertensis.
Fig. 2.
Comparative genomic analyses of 28 selected Acidithiobacillus strains. a Phylogeny based on concatenated proteins of Acidithiobacillus genomes and orthologous proteins shared by Acidithiobacillus spp. using CVTree3 with a composition vector approach. The topology of phylogenomic tree was shown, and Achromobacter xylosoxidans A8 was used as outgroup. Type strains are shown in red triangles. Specially, the redefined species are shown in the phylogenomic tree, and their former names can be also found in brackets. Numbers at internal nodes represent the number of possible ancestral genes as retrieved under the GD-FR-CWP model. Numbers on the branches and at the end of branches indicate the number of gain (red) and loss (blue) genes, and the extant counts of genes, respectively. For core genome-based phylogeny, the species are numbered in sequence. b Venn diagram representing the orthologous and non-orthologous genes of Acidithiobacillus spp. The number of orthologous genes shared by 28 Acidithiobacillus genomes and the strain-specific genes are shown in the center circle and petals, respectively. c COG assignments of core, accessory, and unique genes. Abbreviations: J, translation, ribosomal structure, and biogenesis; A, RNA processing and modification; K, transcription; L, replication, recombination, and repair; B, chromatin structure and dynamics; D, cell cycle control, cell division, chromosome partitioning; V, defense mechanisms; T, signal transduction mechanisms; M, cell wall/membrane/envelope biogenesis; N, cell motility; U, intracellular trafficking, secretion, and vesicular transport; O, posttranslational modification, protein turnover, chaperones; C, energy production and conversion; G, carbohydrate transport and metabolism; E, amino acid transport and metabolism; F, nucleotide transport and metabolism; H, coenzyme transport and metabolism; I, lipid transport and metabolism; P, inorganic ion transport and metabolism; Q, secondary metabolites biosynthesis, transport and catabolism; R, general function prediction only; S, function unknown. d Mathematical extrapolation for the estimation of size of Acidithiobacillus pan-genome and core genome. Detailed description for calculation is shown in section Model Extrapolation for Acidithiobacillus Pan-Genome.
Identification and functional assignments of orthologs and non-orthologs
Using a total of 28 available genomes, the pipeline BPGA was implemented to systematically explore the hereditary differentiation of Acidthiobacillus isolates. As demonstrated in Fig. 2b, inter- and intra-species diversity could be strongly reflected by the number of orthologs and non-orthologs. Intriguingly, many genes (745) were common in all Acidthiobacillus strains, probably indicating that these acidophiles were likely to share a suit of specialized genes contributing to the basic lifestyle and major phenotypic traits. Furthermore, the strain-specific genes were identified in each Acidthiobacillus genomes, showing the individual difference at the genome level. The rest genes shared by a subset of Acidthiobacillus strains was accessory genome, which was traditionally recognized to be responsible for species diversity and probably confer selective advantages such as econiche adaptation [49, 50].
Functional classifications of core, accessory, and unique genes among these Acidthiobacillus genomes were performed using BLAST algorithm against the COG database. Of these sequences, generally, most have no match in the current database, which was similar to many previous studies [51–54]. Apart from these sequences with no hits, the large proportion of genes (39.33%) in the core genome were predicted to pertain to the metabolic profiles (Fig. 2c), highlighting the potential contribution of metabolism-related genes in mediating the autotrophic lifestyle of Acidthiobacillus spp. KAAS annotation (Additional file 1: Table S3) revealed that genes involved in carbohydrate metabolism (81), amino acid metabolism (80), metabolism of cofactors and vitamins (57), and energy metabolism (55) outnumbered other metabolism-associated genes. Further inspection revealed that the three most abundant genes were assigned to COG categories [J] (translation, ribosomal structure, and biogenesis), [E] (amino acid transport and metabolism), and [M] (cell wall/membrane/envelope biogenesis). As for accessory and unique genes, however, most of them were related to COG categories [L] (replication, recombination, and repair); The finding was frequently found in other similar studies [52, 55].
Mathematical extrapolation for estimating the size of Acidithiobacillus pan-genome
Comparative analysis showed that the relatively small proportion of protein-coding genes (18.6% - 27.5%) were assigned to core genome. Additionally, the inconsistency between these two phylogenies based on whole-genome and core genome was identified (Fig. 2a), which provided an indication that abundant exogenous genes introduced by genetic exchange might contribute to genome plasticity. The inference, to a large extent, could be explained by a well counter-example about B. aphidicola, in which the most extreme genome stability was observed because its genome has undergone few events of chromosome rearrangements, or gene acquisitions in the past 50 to 70 million years [56].
Due to its genome expansion, in theory, novel genes would be introduced into Acidithiobacillus genomes with the new sequenced genome. To evaluate the size of Acidithiobacillus pan-genome, mathematical extrapolation based on the present 28 genomes was performed using the pipeline BPGA (Fig. 2d). Exponential equation [Fc(n) = 1920.71e-0.041904n] indicated that the extrapolated curve following a gentle slope reached a minimum of 745 after the 28th Acidithiobacillus genome was added into the dataset. Furthermore, empirical power law equation [Ps(n) = 3101.82n0.36806] revealed the Acidithiobacillus pan-genome with an average parameter of 0.36806. The exponent 1 > γ > 0, according to the Heaps’ law [57], hinted that its size was increasing and unbounded. Taken together, it was inferred that frequent lateral exchange of genetic material might dramatically expand the gene content of bacterial genomes, and promote the environmental adaptability of Acidithiobacillus isolates.
Potential forces driving genome evolution of Acidithiobacillus strains
Adaptive evolution of microbial genomes is generally regarded to be driven by several genetic events, such as duplication, rearrangement, acquisition, and loss of genes and genome segments [58, 59]. Especially, many studies have discussed the potential roles of gene gain and loss in genome evolution in response to local environmental conditions [46, 52, 55, 60]. Both gene gain and loss are thought to be two key adaptive mechanisms that have great potential to drive genome evolution of microbial species, but their relative importance in shaping the content of microbial genomes and driving microbial evolution and speciation has been unclear yet.
To investigate the potential evolutionary forces, gene turnover rates of Acidithiobacillus genomes were first evaluated, followed by the prediction of mobile genetic elements. A subsequent analysis that relates the comparison of genomic regions of interest (such as genes associated with metabolic profiles) to the presence or absence of mobile genetic elements in the genomic neighborhoods may facilitate the deduction of evolutionary mechanisms that drive adaptive evolution and diversification of microbial genomes.
Gene turnover potentially contributing to genome differentiation
Gene families in Acidithiobacillus genomes were classified as orthogroups and listed in Additional file 1: Table S4. To obtain an overview of the evolutionary dynamics of gene families, the ancestral gene set profiles were constructed at all nodes of the phylogenomic tree among these 28 fully sequenced strains (Fig. 2a). Variation occurred among Acidithiobacillus genomes in terms of the total gene numbers. On a global scale, numerous gain/loss events were frequently identified. Starting from a common ancestor, divergent evolution occurred between A. caldus strains and the others in an earlier time, with a dramatic gene decrease. Nevertheless, there was a global gene increase in many of branches. In short, many divergences among Acidithiobacillus strains were observed in the whole evolutionary process, accompanied by diverse events of both gene gain and loss. The data presented here hinted that frequent gene turnover might confer selective advantages to acidophiles in acid environments, probably contributing to the observed genome differentiation.
Prediction of mobile genetic elements
Gene acquisition is usually accompanied by the events of horizontal gene transfer (HGT), which frequently occur in microorganisms and are beneficial for microbial species to rapidly adapt to changing environments [58]. A significant part of HGT is facilitated by mobile genetic elements, and is generally characterized by some signatures in microbial genomes, such as integration sites often related to tRNA genes, abnormal GC contents, or varied codon usage [61–63]. Accordingly, transposases and integrases were predicted and classified using the platform ISFinder (Additional file 3: Table S5). Results showed that putative transposable elements Tn3, ISL3, IS91, and IS1595 were present and abundant in most of Acidithiobacillus genomes. While many genes were assigned to various IS families, some transposable elements were identified unique in certain Acidithiobacillus strains, such as IS51 in A. thiooxidans Licanantay, and IS605 in several A. thiooxidans strains.
Putative phage-associated genes were identified in all Acidithiobacillus genomes using the server Prophinder, accompanied by BLASTP search against the proteins in viruses and prophages of ACLAME database. In our study, many predicted phage-associated genes were dispersed over certain genomic regions within A. caldus strains SM-1, MTH-04, and ATCC 51756, A. ferrooxidans ATCC 23270, and A. thiooxidans DSM 14366 (Additional file 4: Table S6), probably suggesting that these strains might been targeted by phage via infection. Referred to the previous studies [46, 64], especially, a gene cluster composed of several phage-associated genes in A. caldus ATCC 51756 (chromosome: 100435-143772) might be a fragment of the putative prophage, since some DNA packaging- (phage terminase large subunit and portal protein), head-, tail-, and cell lysis-associated genes were identified. However, genes related to small subunit of the phage terminase and phage DNA replication were absent in this genome. In addition, genes encoding putative prophage DNA integration, phage DNA replication, and some phage with unknown functions were predicted to be scattered throughout the genomic regions of Acidithiobacillus strains via comparison to the ACLAME database.
Comparisons of genomic regions of interest in Acidithiobacillus spp.
Comparative genomics has facilitated the systematical investigation regarding the intra- and inter-species diversity. Comparisons of genomic regions of interest, accompanied by the prediction of putative mobile genetic elements, probably allow the identification of potential driving forces of genome evolution. Since metabolism-related genes accounted for a large proportion of bacterial genomes, we thus focused on the comparison of inferred metabolic profiles.
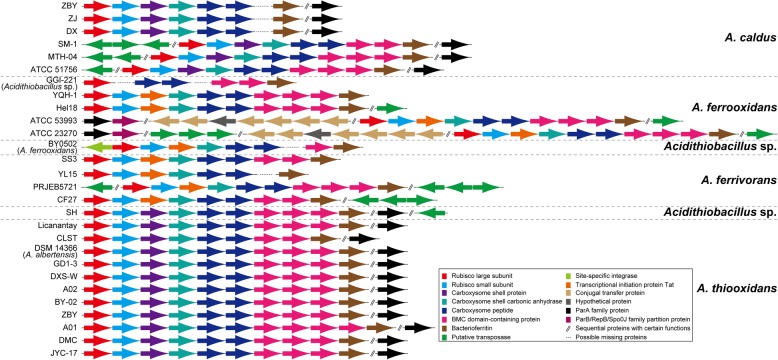
Carboxysome, a cytoplasmic and polyhedral bacterial microcompartment (BMC), has been traditionally thought to be a central part of the carbon-concentrating mechanism, which was greatly useful for increasing the fixation of external carbon in cyanobacteria and some chemoautotrophs [65]. In our study, a gene cluster potentially associated with carboxysome was widespread in the obligate chemolithotroph Acidithiobacillus. Further inspection showed that the genomic organization of carboxysome gene clusters was similar in all Acidithiobacillus strains (Fig. 3 and Additional file 5: Table S7), mainly including large and small subunits of ribulose1,5-bisphosphate carboxylase/oxygenase (RubisCO), carboxysome shell protein (or transcriptional initiation protein Tat in A. ferrooxidans strains, Acidithiobacillus sp. BY0502, and A. ferrivorans strains), carboxysome shell carbonic anhydrase (CA; ε-class) [66], carboxysome peptides A and B, several BMC domain-containing proteins, and bacterioferritin. Despite their similar gene content, order, and orientation, fascinatingly, various putative transposases or site-specific integrases were predicted to be in the genomic neighborhoods (both upstream and downstream; Fig. 3). The similar observations have been documented in some other acidophiles inhabiting the acid environments, such as Ferroplasma acidarmanus [67] and Ferrovum sp. [46]. Other signatures of HGT including phage-associated genes and genes related to the type IV secretion system were further identified. No putative phage-associated genes were found in these genomic neighborhoods, but a set of genes encoding the components of putative type IV secretion system was predicted in A. ferrooxidans strains ATCC 23270 and ATCC 53993 (Additional file 5: Table S7). In addition, other characteristic function associated with plasmid partition system was predicted. Obviously, the ParAB partition system was in the upstream of carboxysome-associated gene clusters within A. ferrooxidans strains ATCC 23270 and ATCC 53993, while a truncated ParAB partition system, which lacked the ParB family partition protein, was found in the downstream of genomic region related to carboxysome within A. caldus strains, Acidithiobacillus sp. SH, and A. thiooxidans strains. As reported in the previous studies, the ParAB partition system was not only responsible for the segregation of low copy number plasmids [46], but also involved in the bacterial chromosome segregation [68]. So far, some Acidithiobacillus strains were documented to harbor at least one plasmid, including A. ferrivorans PRJEB5721, A. caldus strains ATCC 51756, SM-1, and MTH-04, although whether the plasmid existed in the others remains unclear.
Fig. 3.
Genomic organization of the gene clusters related to carboxysomes in Acidithiobacillus strains. As depicted in this figure, the order and orientation of each gene are presented. Varied functional genes are shown in different colors. More details for gene information are shown in Additional file 4: Table S6.
Nitrogenase complex has been recognized to be the key enzyme directing the fixation of molecular nitrogen. In our study, a suit of nif-genes encoding the components of putative nitrogenase complex, such as MoFe cofactor biosynthesis proteins NifXNE and nitrogenase structural subunits NifKDH, was predicted to be present in A. ferrooxidans strains except for strain GGI-221, of which several genes were absent probably due to the potential frameshift (Additional file 5: Table S7). The identical genomic organization were also observed in all A. ferrivorans strains. However, no nif-genes were identified in the genomes of other Acidithiobacillus spp. Interestingly, a gene encoding putative transposase was inserted into the nitrogenase-associated gene cluster of A. ferrivorans strains CF27 and PRJEB5721. Additionally, an integrase-encoding gene was identified in the genomic neighborhood of A. ferrooxidans strains ATCC 23270 and ATCC 53993. In spite of these findings, the conserved genomic segments associated with nitrogenase complex strongly indicated that these nif-genes in all strains of A. ferrooxidans and A. ferrivorans were likely to be vertically inherited from the common ancestral genomes, and these strains with putative mobile genetic elements might undergo potential HGT events to recruit novel hereditable character instead of nitrogen-fixing ability. From another point of view, the absence of nitrogenase-associated gene cluster in other Acidithiobacillus isolates was likely to be the consequence of gene loss in the common ancestor of Acidithiobacillus spp. A similar observation for iron-oxidizing acidophile L. ferriphilum was reported in a previous study, in which homologous genes involved in nitrogenase complex were absent in strains DX and ML-04 [53].
Discussion
The genus Acidithiobacillus has been currently recognized to be composed of seven validated species. Using a full set of 16S rRNA gene sequence data collected from the public database, the sampled species-level phylogeny was constructed to extrapolate the potential evolutionary lineage of Acidithiobacillus spp. Similar to a recent study [21], the current results revealed a significant diversity of genus Acidithiobacillus, suggesting that many underappreciated species within this genus remain to be recognized and redefined. In the past, provisional recognition of a number of Acidithiobacillus species, such as A. concretivorus [69–71] and A. cuprithermicus [72], has occurred, but some of them have often been questioned with regard to the validity of newly proposed ‘species’ [73]. Our current study provided additional evidence to revise the taxon of an A. cuprithermicus strain (Additional file 2: Table S2). In addition, many unclassified Acidithiobacillus spp., such as Acidithiobacillus sp. GGI-221, and strains belonging to identified species were re-evaluated (Fig. 1). Especially, the debatable A. ferrooxidans BY0502 and A. albertensis DSM 14366 were redefined, according to a polyphasic taxonomic study including 16S rRNA gene- and whole-genome-based phylogeny, and ANI approach. The findings presented here indicated that strain BY0502 might be phylogenetically affiliated to A. ferriphilus instead of the currently proposed A. ferrooxidans. However, more robust evidence based on genomic data should be provided to support the taxonomic assignment in the future study. As for strain DSM 14366, it has been previously proposed to be a member of species A. albertensis. Although physiologically similar to A. thiooxidans, this species was reported to be distinguished by harboring the glycocalyx- and flagella-associated genes [12]. And in fact, our previous studies have reported the presence of putative flagella-related genes in A. thiooxidans strains [10, 66]. Accordingly, the limited evidence might not support the controversially proposed A. albertensis. However, subspecies-level analyses remain to be attempted to improve the resolution of Acidithiobacillus phylogeny and further distinguish the ecologically distinct organisms with closely related taxa.
Comparative genomics has yielded insights into the genetic diversity of microbial genomes. In this study, functional assignments highlighted the relatively high proportion of accessory and strain-specific genes associated with COG category [L] (Fig. 2c), suggesting that these genes might confer adaptive advantage to harsh eco-environments, as high concentrations of toxic substances in acid environments, such as heavy metals, easily caused the DNA injury [74]. Additionally, a set of 745 genes common in all 28 Acidithiobacillus strains were identified (Fig. 2b). Although mathematical model showed that extrapolated curve has reached a trough (Fig 2d), this core set of genes might not be equal to the theoretically minimized genome of sulfur-oxidizing Acidithiobacillus populations. As stated by Koonin [75], the definition of ‘minimal gene-set’ should be associated with the local environmental conditions under which these genes were necessary and sufficient for survival and proliferation of any given organisms. Despite this, the presence of a set of core genes in bacterial genomes was testament to the conservative nature of long-term evolution [76]. Given these similar eco-environments where Acidithiobacillus strains inhabit, a backbone of conserved genes might be necessary for sustaining a functional cell. These essential genes might evolve from the last universal common ancestor in multiple ways, and endow bacterial species with opportunities to adapt to environmental perturbations. It might be reasonable that a large proportion of core genes were assigned to COG categories [J], [E], and [M], in light of the following possible explanations: (i) genes involved in translation were conserved in all cellular life forms [75]; (ii) efficient uptake of nutrients was essential for basic lifestyle of microorganisms [77]; and (iii) distinctive structural and functional characteristics were developed to cope with harsh environmental conditions such as extremely acidic settings [10], since specialized cellular structures and functions, e.g., highly impermeable membranes, selective outer membrane porin proteins, reversed membrane potential, and active proton pumping, were benefit for maintaining a stable pH gradient and thus for supporting the growth of acidophilic microorganisms at low pH [78]. In an earlier study, pH value was observed to selectively influence the expression of certain genes involved in key metabolisms in some iron/sulfur-oxidizing microorganisms [79]. Combined with comparative genomics in this study, it should be an interesting prospect to design other omics-oriented studies, such as transcriptomics and proteomics, in the future works to explore the potential pH homeostatic mechanisms of acidophiles.
Since the frequent gene turnover among Acidithiobacillus strains might contribute to their genetic diversity (Fig. 2a), further studies targeted the issues that what and how adaptive evolutionary forces influenced the gene repertoires of bacterial genomes. Focusing on genomic regions related to metabolic profiles, we found that several signatures of mobilization (transposases, integrases, phage-related genes, integration sites) and conjugation (the type IV secretion system) were scattered throughout the genomic regions associated with carboxysome, probably providing a coherent picture of the prevalence of varied HGT events. RubisCO, the key enzyme in carbon fixation, might allow the obligate lithoautotroph Hydrogenovibrio marinus MH-110 to adapt well to different CO2 concentrations [80]. And carboxysome-associated CA was reported to elevate the concentrations of carbon dioxide in the surrounding of Rubisco via effectively converting the accumulated cytosolic bicarbonate into CO2 [81]. Genes arose by HGT coupled with functional recruitments in order to cope with the harsh environmental conditions, indicating the high genome plasticity. However, the findings revealed that the absence of nitrogenase-associated genes in A. thiooxidans strains, A. caldus strains, Acidithiobacillus sp. strains BY0502 and SH might be caused by gene loss in the common ancestral genomes of Acidithiobacillus strains. Currently, the increasing genomic data deluge in public databases has been revealing an unexpected perspective of gene loss as an effective means of hereditary variation that potentially resulted in adaptive phenotypic diversity [82]. Referred to a classical theory, i.e., the Black Queen Hypothesis [83], a community-dependent evolutionary pattern was thus proposed to explain the observed loss of genomic segments. In the context of deficient nitrogen source in acid eco-environments [84], some microbial members in the common communities should make a ‘compromise’ to optimally utilize the limited resources of the entire communities. The fixation of externally-derived nitrogen might be partitioned into a small fraction of co-existing diazotrophic species, such as A. ferrooxidans and A. ferrivorans. The product of their metabolic activities may provide alternative nitrogen compounds, such as ammonium, for supporting the nitrogen supply of other co-occurring members without the nitrogen-fixing abilities.
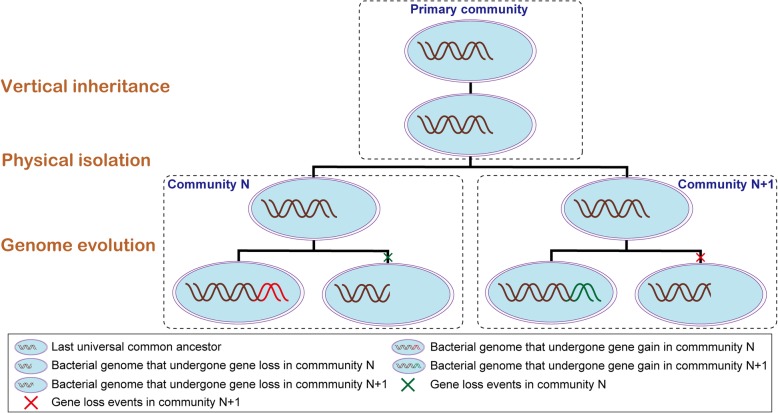
Taken together, a potential evolutionary model for Acidithiobacillus strains was thus proposed to extrapolate the hereditary differentiation patterns in response to the local environmental conditions (Fig. 4). Extremely acidic environments as a possible natural barrier might provide few opportunities for indigenous species to access the gene pool in the surrounding environments, but prompt the gene flow across these acidophilic microorganisms in the relatively isolated regions, thereby indicating that the evolution of given species might depend on other members in a common community. The essential genes in these acidophiles have vertically inherited from ancestral genomes to support their basic lifestyle, and meanwhile evolved the adaptive evolutionary mechanisms responding to the changing environments via various genetic events including gene gain and/or loss. Gene gain events mediated by transfer, transduction, and conjugation may extend the gene repertoires of microbial genomes and potentially recruit novel functionalities, and genome reduction dependent on community functions could economize the nutrient requirements under the resource-deficient conditions, resulting in the observed allopatric speciation.
Fig. 4.
Schematic diagram depicting the potential community-dependent evolutionary model of Acidithiobacillus population. Extremely acidic environments as potential physical barrier might contribute to geographical isolation. Varied gene flow occurs in isolated microbial communities of ecological niches around the world. Gene gain events via various horizontal gene transfer expand the gene repertoires of bacterial genomes, and gain loss events dependent on microbial community increase cellular economization, thereby resulting in the allopatric speciation.
Conclusions
Phylogeny using both 16S rRNA gene and genome sequences expanded our understanding of the genetic diversity of Acidithiobacillus populations. Redefinitions of many questioned species provided new insights into the taxonomic assignments of bacterial species.
A recent study revealed the observation that presented a co-evolution pattern of acidophiles and their acidic habitats [20]. The metabolic activities of acidophilic microorganisms might contribute to the formation of extremely acidic environments, and the feedbacks of these existing harsh eco-environments, in turn, could enhance the exchange frequencies of genetic material within the members of entire microbial communities, thereby resulting in the habitat-driven genome evolution and speciation. Potential evolutionary forces were discussed in this study to further highlight the roles of gene gain and loss in genome evolution, and the donors and recipients of genes were suspected to be derived from the co-occurring members of microbial communities. Gene gain by HGT was doubtlessly recognized as an efficient way to expand the gene repertoire of given species, and gene loss in some ex-existing members might be a good choice to economize the limited resources of the whole community [83, 85]. A case study of genome reduction involved in nitrogen fixation may advance our current knowledge of community-dependent evolutionary adaptation of Acidithiobacillus isolates. Acidophilic microorganisms with a diazotrophic lifestyle act as pioneers to colonize new habitats, the later colonizers might increase cellular economization via gene loss to rationally utilize the limited resources, thereby resulting in adaptive evolution and genome differentiation within the whole community.
Additional files
Figure S1. Geographic distributions and genome attributes of Acidithiobacillus strains used for the construction of genome-based phylogeny and the calculation of average nucleotide identity. Figure S2. Substitution pattern of 16S rRNA genes used for phylogenetic tree. The number of transitions (cross) and transversions (triangle) against the TN93 distance is shown in different colors. Each point in individual sites indicates a pairwise comparison between two of taxa. Figure S3. Calculations of ANIb (upper half) and ANIm (lower half) for Acidithiobacillus strains. The values of ANI above the threshold for species delineation (95%) are highlighted. Table S1. Statistics for the number of RNA (rRNA and tRNA) in Acidithiobacillus (A.) strains with available genomes. Table S3. Functional classifications of the common genes shared by Acidithiobacillus strains using an online platform KAAS. Table S4. Summary for gene families in the genomes of Acidithiobacillus strains, including A. thiooxidans (formerly A. albertensis) DSM 14366 (1), A. caldus strains ATCC 51756 (2), DX (3), MTH-04 (4), SM-1 (5), ZBY (6), ZJ (7), GGI-221 within A. ferrooxidans (formerly Acidithiobacillus sp.) GGI-221 (8), Acidithiobacillus sp. SH (9), A. ferrooxidans strains ATCC 23270 (10), ATCC 53993 (11), Hel18 (12), YQH-1 (13), Acidithiobacillus sp. (formerly A. ferrooxidans) BY0502 (14), A. ferrivorans strains CF27 (15), PRJEB5721 (16), SS3 (17), YL15 (18), A. thiooxidans strains A01 (19), A02 (20), BY-02 (21), CLST (22), DMC (23), DXS-W (24), GD1-3 (25), JYC-17 (25), Licanantay (27), and ZBY (28). (DOC 37024 kb)
Table S2. General properties of 16S rRNA genes of Acidithiobacillus isolates and clones available in the public database. (XLSX 33 kb)
Table S5. Statistics for puative transposases in Acidithiobacillus genomes using the online tool ISFinder. (XLSX 13 kb)
Table S6. Prediction of putative phage-associated genes in the genomes of Acidithiobacillus strains, including A. caldus strains SM-1, MTH-04, and ATCC 51756, A. ferrooxidans ATCC 23270, and A. thiooxidans DSM 14366. (XLSX 38 kb)
Table S7. Gene content of genomic regions associated with carboxysome and nitrogenase in Acidithiobacillus strains. (XLSX 26 kb)
Acknowledgements
Not applicable.
Abbreviations
- ANI
Average nucleotide identity
- ANIb
ANI based on BLAST algorithm
- ANIm
ANI based on MUMmer algorithm
- BMC
Microcompartment
- BPGA
Bacterial Pan Genome Analysis
- CA
Carbonic anhydrase
- COG
Clusters of Orthologous Groups
- HGT
Horizontal gene transfer
- IS
Insertion sequence
- KAAS
KEGG Automatic Annotation Server
- LPSN
List of Prokaryotic names with Standing in Nomenclature
- RubisCO
Ribulose1,5-bisphosphate carboxylase/oxygenase
- TN93
Tamura-Nei
Authors’ contributions
XZ and XL conceived the study. XZ designed the manuscript, interpreted the data, prepared the figures, and drafted and revised the paper. LL, GW, and DZ assisted in analyzing a part of data. YL and BM participated in the discussion to improve the manuscript. All authors have read and approved the final manuscript.
Funding
This study was financially supported by the National Natural Science Foundation of China (31570113). The funding body had no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Availability of data and materials
The raw dataset, including 16S rRNA gene sequences and genomic sequences of Acidithibacillus spp., was available and download from the public database National Center for Biotechnology Information. All data supporting the findings of our study can be found within the manuscript and additional file tables.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Mazaris AD, Gouni MM, Michaloudi E, Bobori DC. Biogeographical patterns of freshwater micro-and macroorganisms: a comparison between phytoplankton, zooplankton and fish in the eastern Mediterranean. J Biogeogr. 2010;37(7):1341–1351. doi: 10.1111/j.1365-2699.2010.02294.x. [DOI] [Google Scholar]
- 2.Baas-Becking L. Geobiologie of inleiding tot de milieukunde. Hague: WP Van Stockum and Zoon; 1934. [Google Scholar]
- 3.Cho J, Tiedje JM. Biogeography and degree of endemicity of fluorescent Pseudomonas strains in soil. Appl Environ Microbiol. 2000;66(12):5448–5456. doi: 10.1128/AEM.66.12.5448-5456.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Papke RT, Ramsing NB, Bateson MM, Ward DM. Geographical isolation in hot spring cyanobacteria. Environ Microbiol. 2003;5(8):650–659. doi: 10.1046/j.1462-2920.2003.00460.x. [DOI] [PubMed] [Google Scholar]
- 5.Ryšánek D, Hrčková K, Škaloud P. Global ubiquity and local endemism of free-living terrestrial protists: phylogeographic assessment of the streptophyte alga Klebsormidium. Environ Microbiol. 2015;17(3):689–698. doi: 10.1111/1462-2920.12501. [DOI] [PubMed] [Google Scholar]
- 6.Whitaker RJ, Grogan DW, Taylor JW. Geographic barriers isolate endemic populations of hyperthermophilic archaea. Science. 2003;301(5635):976–978. doi: 10.1126/science.1086909. [DOI] [PubMed] [Google Scholar]
- 7.Papke RT, Ward DM. The importance of physical isolation to microbial diversification. FEMS Microbiol Ecol. 2004;48(3):293–303. doi: 10.1016/j.femsec.2004.03.013. [DOI] [PubMed] [Google Scholar]
- 8.Hanson CA, Fuhrman JA, Horner-Devine MC, Martiny JBH. Beyond biogeographic patterns: processes shaping the microbial landscape. Nat Rev Microbiol. 2012;10(7):497–506. doi: 10.1038/nrmicro2795. [DOI] [PubMed] [Google Scholar]
- 9.Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R. The microbial pan-genome. Curr Opin Genet Dev. 2005;15(6):589–594. doi: 10.1016/j.gde.2005.09.006. [DOI] [PubMed] [Google Scholar]
- 10.Zhang X, Liu X, Liang Y, Fan F, Zhang X, Yin H. Metabolic diversity and adaptive mechanisms of iron- and/or sulfur-oxidizing autotrophic acidophiles in extremely acidic environments. Environ Microbiol Rep. 2016;8(5):738–751. doi: 10.1111/1758-2229.12435. [DOI] [PubMed] [Google Scholar]
- 11.Waksman SA, Joffe JS. Microorganisms concerned in the oxidation of sulfur in the soil. II. Thiobacillus thiooxidans, a new sulfur-oxidizing organism isolated from the soil. J Bacteriol. 1922;7(2):239–256. doi: 10.1128/jb.7.2.239-256.1922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kelly DP, Wood AP. Reclassification of some species of Thiobacillus to the newly designated genera Acidithiobacillus gen. nov., Halothiobacillus gen. nov. and Thermithiobacillus gen. nov. Int J Syst Evol Microbiol. 2000;50:511–516. doi: 10.1099/00207713-50-2-511. [DOI] [PubMed] [Google Scholar]
- 13.Bryant RD, McGroarty KM, Costerton JW, Laishley EJ. Isolation and characterization of a new acidophilic Thiobacillus species (T. albertis) Can J Microbiol. 1983;29(9):1159–1170. doi: 10.1139/m83-178. [DOI] [Google Scholar]
- 14.Hallberg KB, Lindström EB. Characterization of Thiobacillus caldus sp. nov., a moderately thermophilic acidophile. Microbiology. 1994;140:3451–3456. doi: 10.1099/13500872-140-12-3451. [DOI] [PubMed] [Google Scholar]
- 15.Temple KL, Colmer AR. The autotrophic oxidation of iron by a new bacterium: Thiobacillus ferrooxidans. J Bacteriol. 1951;62(5):605–611. doi: 10.1128/jb.62.5.605-611.1951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hallberg KB, González-Toril E, Johnson DB. Acidithiobacillus ferrivorans, sp. nov.; facultatively anaerobic, psychrotolerant iron-, and sulfur-oxidizing acidophiles isolated from metal mine-impacted environments. Extremophiles. 2010;14(1):9–19. doi: 10.1007/s00792-009-0282-y. [DOI] [PubMed] [Google Scholar]
- 17.Hedrich S, Johnson DB. Acidithiobacillus ferridurans sp. nov., an acidophilic iron-, sulfur-and hydrogen-metabolizing chemolithotrophic gammaproteobacterium. Int J Syst Evol Microbiol. 2013;63(Pt 11):4018–4025. doi: 10.1099/ijs.0.049759-0. [DOI] [PubMed] [Google Scholar]
- 18.Falagán C, Johnson DB. Acidithiobacillus ferriphilus sp. nov., a facultatively anaerobic iron- and sulfur-metabolizing extreme acidophile. Int J Syst Evol Microbiol. 2016;66(1):206–211. doi: 10.1099/ijsem.0.000698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhao H, Zhang Y, Zhang X, Qian L, Sun M, Yang Y, Zhang Y, Wang J, Kim H, Qiu G. The dissolution and passivation mechanism of chalcopyrite in bioleaching: An overview. Miner Eng. 2019;136:140–154. doi: 10.1016/j.mineng.2019.03.014. [DOI] [Google Scholar]
- 20.Colman DR, Poudel S, Hamilton TL, Havig JR, Selensky MJ, Shock EL, Boyd ES. Geobiological feedbacks and the evolution of thermoacidophiles. ISME J. 2018;12:225–236. doi: 10.1038/ismej.2017.162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nuñez H, Moya-Beltrán A, Covarrubias PC, Issotta F, Cárdenas JP, González M, Atavales J, Acuña LG, Johnson DB, Quatrini R. Molecular systematics of the genus Acidithiobacillus: insights into the phylogenetic structure and diversification of the taxon. Front Microbiol. 2017;8:30. doi: 10.3389/fmicb.2017.00030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ni Y, He K, Bao J, Yang Y, Wan D, Li H. Genomic and phenotypic heterogeneity of Acidithiobacillus spp. strains isolated from diverse habitats in China. FEMS Microbiol Ecol. 2008;64(2):248–259. doi: 10.1111/j.1574-6941.2008.00457.x. [DOI] [PubMed] [Google Scholar]
- 23.Huang L, Kuang J, Shu W. Microbial ecology and evolution in the acid mine drainage model system. Trends Microbiol. 2016;24(7):581–593. doi: 10.1016/j.tim.2016.03.004. [DOI] [PubMed] [Google Scholar]
- 24.Lagesen K, Hallin P, Rødland EA, Stærfeldt H, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35(9):3100–3108. doi: 10.1093/nar/gkm160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Katoh K, Rozewicki J, Yamada KD. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform. 2017;30:3059. [DOI] [PMC free article] [PubMed]
- 26.Swofford DL, Olsen G, Waddell PJ, Hillis DM. Phylogenetic inference in molecular systematics. In: Hillis DM, Moritz C, Sunderland MBK, editors. Molecular systematics. Massachusetts: Sinauer Associates Inc; 1996. pp. 407–514. [Google Scholar]
- 27.Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10(3):512–526. doi: 10.1093/oxfordjournals.molbev.a040023. [DOI] [PubMed] [Google Scholar]
- 28.Xia X, Xie Z. DAMBE: software package for data analysis in molecular biology and evolution. J Hered. 2001;92(4):371–373. doi: 10.1093/jhered/92.4.371. [DOI] [PubMed] [Google Scholar]
- 29.Renoult JP, Kjellberg F, Grout C, Santoni S, Khadari B. Cyto-nuclear discordance in the phylogeny of Ficus section Galoglychia and host shifts in plant-pollinator associations. BMC Evol Biol. 2009;9:248. doi: 10.1186/1471-2148-9-248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Guindon S, Dufayard J, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59(3):307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- 31.Zuo G, Hao B. CVTree3 web server for whole-genome-based and alignment-free prokaryotic phylogeny and taxonomy. Genomics, Proteomics & Bioinformatics. 2015;13(5):321–331. doi: 10.1016/j.gpb.2015.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Richter M, Rosselló-Móra R. Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA. 2009;106(45):19126–19131. doi: 10.1073/pnas.0906412106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Richter M, Rosselló-Móra R, Glöckner FO, Peplies J. JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics. 2016;32(6):929–931. doi: 10.1093/bioinformatics/btv681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chaudhari NM, Gupta VK, Dutta C. BPGA- an ultra-fast pan-genome analysis pipeline. Sci Rep. 2016;6(1):24373. doi: 10.1038/srep24373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Carretero-Paulet L, Librado P, Chang T, Ibarra-Laclette E, Herrera-Estrella L, Rozas J, Albert VA. High gene family turnover rates and gene space adaptation in the compact genome of the carnivorous plant Utricularia gibba. Mol Biol Evol. 2015;32(5):1284–1295. doi: 10.1093/molbev/msv020. [DOI] [PubMed] [Google Scholar]
- 36.Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C, et al. STRING v9. 1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41(D1):D808–D815. doi: 10.1093/nar/gks1094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007;35(suppl 2):W182–W185. doi: 10.1093/nar/gkm321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wu X, Wu X, Shen L, Li J, Yu R, Liu Y, Qiu G, Zeng W. Whole genome sequencing and comparative genomics analyses of Pandoraea sp. XY-2, a new species capable of biodegrade tetracycline. Front Microbiol. 2019;10:33. doi: 10.3389/fmicb.2019.00033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Emms DM, Kelly S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 2015;16(1):157. doi: 10.1186/s13059-015-0721-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Enright AJ, Van Dongen S, Ouzounis CA. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002;30(7):1575–1584. doi: 10.1093/nar/30.7.1575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Librado P, Vieira FG, Rozas J. BadiRate: estimating family turnover rates by likelihood-based methods. Bioinformatics. 2012;28(2):279–281. doi: 10.1093/bioinformatics/btr623. [DOI] [PubMed] [Google Scholar]
- 42.Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M. ISfinder: the reference centre for bacterial insertion sequences. NUCLEIC ACIDS RES. 2006;34(S1):D32–D36. doi: 10.1093/nar/gkj014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lowe TM, Chan PP. tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 2016;44(W1):W54–W57. doi: 10.1093/nar/gkw413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lima-Mendez G, Van Helden J, Toussaint A, Leplae R. Prophinder: a computational tool for prophage prediction in prokaryotic genomes. Bioinformatics. 2008;24(6):863–865. doi: 10.1093/bioinformatics/btn043. [DOI] [PubMed] [Google Scholar]
- 45.Leplae R, Lima-Mendez G, Toussaint A. ACLAME: a CLAssification of Mobile genetic Elements, update 2010. Nucleic Acids Res. 2010;38(suppl_1):D57–D61. doi: 10.1093/nar/gkp938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ullrich SR, González C, Poehlein A, Tischler JS, Daniel R, Schlömann M, Holmes DS, Mühling M. Gene loss and horizontal gene transfer contributed to the genome evolution of the extreme acidophile “Ferrovum”. Front Microbiol. 2016;7:797. doi: 10.3389/fmicb.2016.00797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Stackebrandt E, Goebel BM. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Evol Microbiol. 1994;44(4):846–849. doi: 10.1099/00207713-44-4-846. [DOI] [Google Scholar]
- 48.Stackebrandt E, Ebers J. Taxonomic parameters revisited: tarnished gold standards. Microbiol Today. 2006;33:152–155. [Google Scholar]
- 49.Vernikos G, Medini D, Riley DR, Tettelin H. Ten years of pan-genome analyses. Curr Opin Microbiol. 2015;23:148–154. doi: 10.1016/j.mib.2014.11.016. [DOI] [PubMed] [Google Scholar]
- 50.Tettelin H, Riley D, Cattuto C, Medini D. Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol. 2008;11(5):472–477. doi: 10.1016/j.mib.2008.09.006. [DOI] [PubMed] [Google Scholar]
- 51.Zhang X, Feng X, Tao J, Ma L, Xiao Y, Liang Y, Liu X, Yin H. Comparative genomics of the extreme acidophile Acidithiobacillus thiooxidans reveals intraspecific divergence and niche adaptation. Int J Mol Sci. 2016;17(8):1355. doi: 10.3390/ijms17081355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Zhang X, Liu X, Liang Y, Guo X, Xiao Y, Ma L, Miao B, Liu H, Peng D, Huang W, et al. Adaptive evolution of extreme acidophile Sulfobacillus thermosulfidooxidans potentially driven by horizontal gene transfer and gene loss. Appl Environ Microbiol. 2017;83(7):e03098–e03016. doi: 10.1128/AEM.03098-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhang X, Liu X, Yang F, Chen L. Pan-genome analysis links the hereditary variation of Leptospirillum ferriphilum with its evolutionary adaptation. Front Microbiol. 2018;9:577. doi: 10.3389/fmicb.2018.00577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhang X, Liu Z, Wei G, Yang F, Liu X. In silico genome-wide analysis reveals the potential links between core genome of Acidithiobacillus thiooxidans and its autotrophic lifestyle. Front Microbiol. 2018;9:1255. doi: 10.3389/fmicb.2018.01255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zhang X, Liu X, He Q, Dong W, Zhang X, Fan F, Peng D, Huang W, Yin H. Gene turnover contributes to the evolutionary adaptation of Acidithiobacillus caldus: insights from comparative genomics. Front Microbiol. 2016;7:1960. doi: 10.3389/fmicb.2016.01960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tamas I, Klasson L, Canbäck B, Näslund AK, Eriksson A, Wernegreen JJ, Sandström JP, Moran NA, Andersson SGE. 50 million years of genomic stasis in endosymbiotic bacteria. Science. 2002;296(5577):2376–2379. doi: 10.1126/science.1071278. [DOI] [PubMed] [Google Scholar]
- 57.Heaps HS. Information Retrieval, Computational and Theoretical Aspects. Orlando: Academic Press; 1978. [Google Scholar]
- 58.Gogarten JP, Doolittle WF, Lawrence JG. Prokaryotic evolution in light of gene transfer. Mol Biol Evol. 2002;19(12):2226–2238. doi: 10.1093/oxfordjournals.molbev.a004046. [DOI] [PubMed] [Google Scholar]
- 59.Boon E, Meehan CJ, Whidden C, Wong DHJ, Langille MGI, Beiko RG. Interactions in the microbiome: communities of organisms and communities of genes. FEMS Microbiol Rev. 2014;38(1):90–118. doi: 10.1111/1574-6976.12035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Lin W, Zhang W, Zhao X, Roberts AP, Paterson GA, Bazylinski DA, Pan Y. Genomic expansion of magnetotactic bacteria reveals an early common origin of magnetotaxis with lineage-specific evolution. ISME J. 2018;12(6):1508. doi: 10.1038/s41396-018-0098-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Waack S, Keller O, Asper R, Brodag T, Damm C, Fricke WF, Surovcik K, Meinicke P, Merkl R. Score-based prediction of genomic islands in prokaryotic genomes using hidden Markov models. BMC Bioinformatics. 2006;7:142. doi: 10.1186/1471-2105-7-142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Juhas M, van der Meer JR, Gaillard M, Harding RM, Hood DW, Crook DW. Genomic islands: tools of bacterial horizontal gene transfer and evolution. FEMS Microbiol Rev. 2009;33(2):376–393. doi: 10.1111/j.1574-6976.2008.00136.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Acuña LG, Cárdenas JP, Covarrubias PC, Haristoy JJ, Flores R, Nuñez H, Riadi G, Shmaryahu A, Valdés J, Dopson M, et al. Architecture and gene repertoire of the flexible genome of the extreme acidophile Acidithiobacillus caldus. PLoS One. 2013;8(11):e78237. doi: 10.1371/journal.pone.0078237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Canchaya C, Proux C, Fournous G, Bruttin A, Brüssow H. Prophage genomics. Microbiol Mol Biol Rev. 2003;67(2):238–276. doi: 10.1128/MMBR.67.2.238-276.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Klein MG, Zwart P, Bagby SC, Cai F, Chisholm SW, Heinhorst S, Cannon GC, Kerfeld CA. Identification and structural analysis of a novel carboxysome shell protein with implications for metabolite transport. J Mol Biol. 2009;392(2):319–333. doi: 10.1016/j.jmb.2009.03.056. [DOI] [PubMed] [Google Scholar]
- 66.Zhang X, She S, Dong W, Niu J, Xiao Y, Liang Y, Liu X, Zhang X, Fan F, Yin H. Comparative genomics unravels metabolic differences at species and/or strain level and extremely acidic environmental adaptation of ten bacteria belonging to the genus Acidithiobacillus. Syst Appl Microbiol. 2016;39(8):493–502. doi: 10.1016/j.syapm.2016.08.007. [DOI] [PubMed] [Google Scholar]
- 67.Allen EE, Tyson GW, Whitaker RJ, Detter JC, Richardson PM, Banfield JF. Genome dynamics in a natural archaeal population. Proc Natl Acad Sci USA. 2007;104(6):1883–1888. doi: 10.1073/pnas.0604851104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Lioy VS, Volante A, Soberón NE, Lurz R, Ayora S, Alonso JC. ParAB partition dynamics in Firmicutes: nucleoid bound ParA captures and tethers ParB-plasmid complexes. PLoS One. 2015;10(7):e131943. doi: 10.1371/journal.pone.0131943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Parker CD. The corrosion of concrete. I. The isolation of a species of bacterium associated with the corrosion of concrete exposed to atmosphere containing hydrogen sulphide. Aust J Biol Med Sci. 1945;23(2):81–90. doi: 10.1038/icb.1945.13. [DOI] [Google Scholar]
- 70.Parker CD. The corrosion of concrete 2. The function of Thiobacillus concretivorus (nov. spec.) in the corrosion of concrete exposed to atmospheres containing hydrogen sulphide. Aust J Biol Med Sci. 1945;23(2):91–98. doi: 10.1038/icb.1945.14. [DOI] [Google Scholar]
- 71.Parker CD. Species of sulphur bacteria associated with the corrosion of concrete. Nature. 1947;159(4039):439. doi: 10.1038/159439b0. [DOI] [PubMed] [Google Scholar]
- 72.Fernández-Remolar DC, Rodriguez N, Gómez F, Amils R. Geological record of an acidic environment driven by iron hydrochemistry: The Tinto River system. J Geophys Res. 2003;108(E7):5080. doi: 10.1029/2002JE001918. [DOI] [Google Scholar]
- 73.Vishniac W, Santer M. The thiobacilli. Bacteriol Rev. 1957;21(3):195–213. doi: 10.1128/br.21.3.195-213.1957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Silver S, Phung LT. Bacterial heavy metal resistance: new surprises. Annu Rev Microbiol. 1996;50:753–789. doi: 10.1146/annurev.micro.50.1.753. [DOI] [PubMed] [Google Scholar]
- 75.Koonin EV. Comparative genomics, minimal gene-sets and the last universal common ancestor. Nat Rev Microbiol. 2003;1(2):127–136. doi: 10.1038/nrmicro751. [DOI] [PubMed] [Google Scholar]
- 76.Lapierre P, Gogarten JP. Estimating the size of the bacterial pan-genome. Trends Genet. 2009;25(3):107–110. doi: 10.1016/j.tig.2008.12.004. [DOI] [PubMed] [Google Scholar]
- 77.Ji B, Zhang S, Arnoux P, Rouy Z, Alberto F, Philippe N, Murat D, Zhang W, Rioux J, Ginet N, et al. Comparative genomic analysis provides insights into the evolution and niche adaptation of marine Magnetospira sp. QH-2 strain. Environ Microbiol. 2014;16(2):525–544. doi: 10.1111/1462-2920.12180. [DOI] [PubMed] [Google Scholar]
- 78.Baker-Austin C, Dopson M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 2007;15(4):165–171. doi: 10.1016/j.tim.2007.02.005. [DOI] [PubMed] [Google Scholar]
- 79.Peng T, Zhou D, Liu Y, Yu R, Qiu G, Zeng W. Effects of pH value on the expression of key iron/sulfur oxidation genes during bioleaching of chalcopyrite on thermophilic condition. Ann Microbiol. 2019;69(6):627–635. doi: 10.1007/s13213-019-01453-y. [DOI] [Google Scholar]
- 80.Yoshizawa Y, Toyoda K, Arai H, Ishii M, Igarashi Y. CO2-Responsive Expression and Gene Organization of Three Ribulose-1,5-Bisphosphate Carboxylase/Oxygenase Enzymes and Carboxysomes in Hydrogenovibrio marinus Strain MH-110. J Bacteriol. 2004;186(17):5685–5691. doi: 10.1128/JB.186.17.5685-5691.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Price GD, Woodger FJ, Badger MR, Howitt SM, Tucker L. Identification of a SulP-type bicarbonate transporter in marine cyanobacteria. Proc Natl Acad Sci USA. 2004;101(52):18228–18233. doi: 10.1073/pnas.0405211101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Albalat R, Cañestro C. Evolution by gene loss. Nat Rev Genet. 2016;17:379–391. doi: 10.1038/nrg.2016.39. [DOI] [PubMed] [Google Scholar]
- 83.Morris JJ, Lenski RE, Zinser ER. The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. mBio. 2012;3(2):e00036–e00012. doi: 10.1128/mBio.00036-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Parro V, Moreno-Paz M, González-Toril E. Analysis of environmental transcriptomes by DNA microarrays. Environ Microbiol. 2007;9(2):453–464. doi: 10.1111/j.1462-2920.2006.01162.x. [DOI] [PubMed] [Google Scholar]
- 85.Morris JJ, Papoulis SE, Lenski RE. Coexistence of evolving bacteria stabilized by a shared Black Queen function. Evolution. 2014;68(10):2960–2971. doi: 10.1111/evo.12485. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1. Geographic distributions and genome attributes of Acidithiobacillus strains used for the construction of genome-based phylogeny and the calculation of average nucleotide identity. Figure S2. Substitution pattern of 16S rRNA genes used for phylogenetic tree. The number of transitions (cross) and transversions (triangle) against the TN93 distance is shown in different colors. Each point in individual sites indicates a pairwise comparison between two of taxa. Figure S3. Calculations of ANIb (upper half) and ANIm (lower half) for Acidithiobacillus strains. The values of ANI above the threshold for species delineation (95%) are highlighted. Table S1. Statistics for the number of RNA (rRNA and tRNA) in Acidithiobacillus (A.) strains with available genomes. Table S3. Functional classifications of the common genes shared by Acidithiobacillus strains using an online platform KAAS. Table S4. Summary for gene families in the genomes of Acidithiobacillus strains, including A. thiooxidans (formerly A. albertensis) DSM 14366 (1), A. caldus strains ATCC 51756 (2), DX (3), MTH-04 (4), SM-1 (5), ZBY (6), ZJ (7), GGI-221 within A. ferrooxidans (formerly Acidithiobacillus sp.) GGI-221 (8), Acidithiobacillus sp. SH (9), A. ferrooxidans strains ATCC 23270 (10), ATCC 53993 (11), Hel18 (12), YQH-1 (13), Acidithiobacillus sp. (formerly A. ferrooxidans) BY0502 (14), A. ferrivorans strains CF27 (15), PRJEB5721 (16), SS3 (17), YL15 (18), A. thiooxidans strains A01 (19), A02 (20), BY-02 (21), CLST (22), DMC (23), DXS-W (24), GD1-3 (25), JYC-17 (25), Licanantay (27), and ZBY (28). (DOC 37024 kb)
Table S2. General properties of 16S rRNA genes of Acidithiobacillus isolates and clones available in the public database. (XLSX 33 kb)
Table S5. Statistics for puative transposases in Acidithiobacillus genomes using the online tool ISFinder. (XLSX 13 kb)
Table S6. Prediction of putative phage-associated genes in the genomes of Acidithiobacillus strains, including A. caldus strains SM-1, MTH-04, and ATCC 51756, A. ferrooxidans ATCC 23270, and A. thiooxidans DSM 14366. (XLSX 38 kb)
Table S7. Gene content of genomic regions associated with carboxysome and nitrogenase in Acidithiobacillus strains. (XLSX 26 kb)
Data Availability Statement
The raw dataset, including 16S rRNA gene sequences and genomic sequences of Acidithibacillus spp., was available and download from the public database National Center for Biotechnology Information. All data supporting the findings of our study can be found within the manuscript and additional file tables.