Abstract
Therapeutic resistance in metastatic castration resistant prostate cancer (mCRPC) can be accompanied by treatment-emergent small cell neuroendocrine carcinoma (t-SCNC), a morphologically distinct subtype. We performed integrative whole-genome and -transcriptome analysis of mCRPC tumor biopsies including paired biopsies after progression, and multiple samples from the same individual. t-SCNC was significantly less likely to have amplification of AR or an intergenic AR enhancer locus, and demonstrated lower expression of AR and its downstream transcriptional targets. Genomic and transcriptional hallmarks of t-SCNC included biallelic loss of RB1, elevated expression levels of CDKN2A and E2F1, and loss of expression of the AR and AR-responsive genes including TMPRSS2 and NKX3–1. We identified three tumors that converted from adenocarcinoma to t-SCNC and demonstrate spatial and temporal intra-patient heterogeneity of metastatic tumors harboring adenocarcinoma, t-SCNC, or mixed expression phenotypes, with implications for treatment strategies in which dual targeting of adenocarcinoma and t-SCNC phenotypes may be necessary.
Introduction
Metastatic castration resistant prostate cancer (mCRPC) is a clinically and genomically heterogeneous disease entity with widely varying outcomes1,2. Small cell neuroendocrine cancer is a highly lethal subset of prostate cancer that is rare at the time of diagnosis but increasingly common upon emergence of resistance to androgen receptor (AR)-targeted therapies3–6. Targeted and whole exome sequencing of treatment-emergent small cell neuroendocrine prostate cancer (t-SCNC) demonstrates frequent inactivating mutations and/or copy number loss of RB1 and TP533. However the exome represents a small percentage of the genome, and the complete genomic landscape of t-SCNC, along with the impact on downstream transcriptional profile, remain to be elucidated.
Whole genome sequencing methods have identified key structural variants present in mCRPC, including amplification of an upstream enhancer of AR that drives AR expression and contributes to progression of castration-resistant disease7–9. Whether t-SCNC harbors structural variants, and whether this has downstream implications for disease progression from adenocarcinoma to t-SCNC is not known. We previously interrogated the whole genomes and transcriptomes of 101 mCRPC samples7, and have subsequently used unbiased clustering to identify the subgroup of tumors harboring a t-SCNC gene expression signature10. We then contrasted the genomic hallmarks of tumors harboring t-SCNC expression signatures with those that did not, including the presence of AR enhancer amplification, confirming our observations using a previously published mCRPC cohort5. We next extended these results to evaluate the gene expression profile of 14 metastatic paired biopsies obtained at baseline and progression from our recently published cohort6 to identify changes in expression patterns associated with the transition from adenocarcinoma to t-SCNC. Finally, we interrogated a published mCRPC cohort that assayed multiple distinct metastatic sites per individual6 and demonstrated frequent intra-individual heterogeneity in t-SCNC.
Materials and Methods
Case Selection and Sample Preparation
Patient tissue samples were obtained through the Stand Up 2 Cancer/Prostate Cancer Foundation-funded West Coast Prostate Cancer Dream Team project, a prospective, multi-center study that acquired metastatic CRPC biopsies from five investigational sites11. Patients were required to have histologic evidence of prostate adenocarcinoma at the time of diagnosis, with subsequent development of mCRPC and at least one metastatic lesion accessible for image-guided percutaneous biopsy. Human studies were approved and overseen by Institutional Review Board of the participating institutions. All individuals provided written informed consent to participate in the prospective tissue acquisition protocol including molecular profiling of tumor and germline samples.
Details of patient sample preparation have been previously reported7,10. Briefly, samples were obtained using image-guided core needle biopsy of metastatic lesion in the bone or soft tissue. Separate cores were obtained and freshly frozen for DNA/RNA sequencing and formalin-fixed, paraffin-embedded for histologic determination of adenocarcinoma versus t-SCNC morphology, as previously described7,10. Tumor structural variation and copy number data were ascertained by whole genome sequencing (WGS) as previously described7. Data from previously published studies5,6 were downloaded from cBioPortal (http://cbioportal.org).
Statistical Analyses
All statistical analysis was performed using R (v3.3.3). Between-group comparisons of continuous variables were performed with the Wilcoxon rank sum test. Contingency table tests were performed with Fisher’s exact test.
Results
Unbiased clustering of expression profile identifies t-SCNC subset
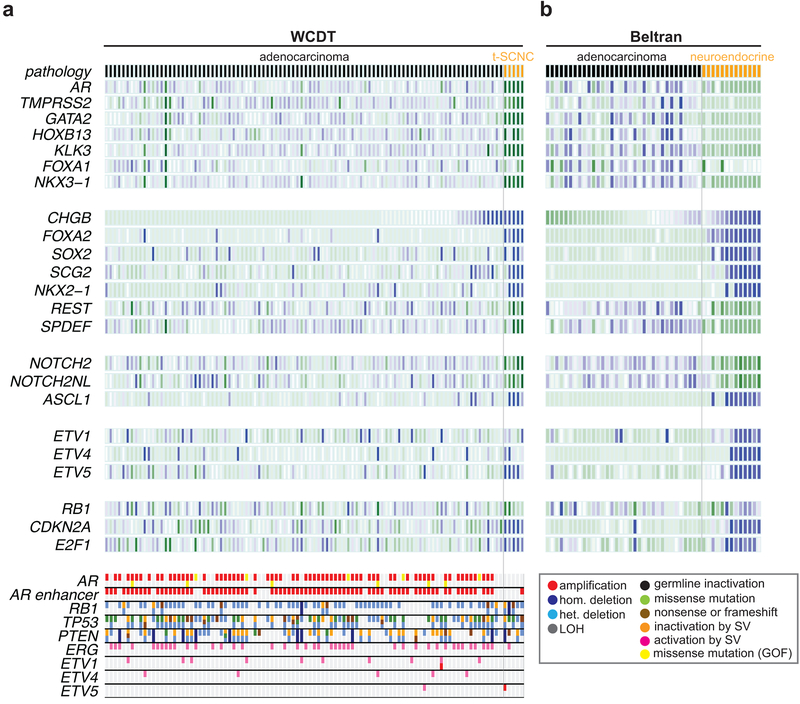
Unbiased gene expression clustering of 101 samples in the WGS cohort identified a previously reported subset of five samples10 that bore a t-SCNC expression signature, with low to absent expression of androgen receptor (AR) signaling, elevated expression of genes associated with small cell morphology, elevated expression of E2F1 and CDKN2A, low expression of NOTCH2/NOTCH2NL, and elevated expression of the ASCL1, a transcription factor essential for neural developement12. (Figure 1A). t-SCNC samples often harbored elevated expression of the ETV1, ETV4, and ETV5 in the absence of activating ETS-family gene fusions; these transcription factors play an important role in neural development13,14. Clinical features of the cohort are shown in Supplementary Table 1. We compared these results to a previously published analysis of mCRPC and neuroendocrine tumors5 (Figure 1B). A gene set reported by Beltran and colleagues to distinguish adenocarcinoma and neuroendocrine tumors separated WGS t-SCNC and neuroendocrine samples5 (Figure S1), confirming the neuroendocrine phenotype of the t-SCNC samples. WGS and Beltran t-SCNC tumors consistently lost AR gene expression. We observed the Beltran tumors exhibited uniform loss of AR signaling but a gradient of RB1 and neuroendocrine marker expression, suggesting neuroendocrine tumors can harbor a heterogeneous expression phenotype.
Figure 1: Expression correlates of t-SCNC status.
A) Top: heatmap showing expression of selected genes in the WGS cohort differentially expressed between t-SCNC and adenocarcinoma expression phenotypes, where darker blue / green indicate higher / lower expression. Adenocarcinoma / t-SCNC status assessed by gene expression profile is indicated by black / orange bar in top row. Bottom: somatic variants detected by WGS in key prostate cancer driver and tumor suppressor genes. The frequency of AR enhancer amplification was lower, and RB1 bi-allelic loss higher, in t-SCNC versus adenocarcinoma samples (P < 0.05 for both comparisons).
B) Heatmap showing expression of genes in panel A in the Beltran data set5. Samples designated CRPC-ADENO / CRPC-NEPC in that publication are indicated by black / orange bar in top row.
Molecular correlates of the t-SCNC phenotype
We then searched genome-wide for associations between the t-SCNC expression phenotype and somatic alterations by WGS. As previously reported5, tumors harboring the t-SCNC expression phenotype were less likely to bear AR gene locus DNA amplification than samples with an adenocarcinoma expression phenotype (2 of 5; 40% vs. 69 of 96; 72%). Additionally, we observed t-SCNC tumors were significantly less likely to harbor AR enhancer amplification (1 of 5; 20% vs. 80 of 96; 83%, P = 0.005). The two WGS t-SCNC samples with AR gene locus DNA amplification had significantly lower AR expression levels than other samples bearing AR amplification, with one sample (DTB-205) showing negligible expression of AR, compatible with complete silencing of AR expression despite the presence of a DNA copy number increase at the AR locus. Samples with the t-SCNC expression phenotype were significantly more likely to harbor bi-allelic RB1 inactivation (3 of 5; 60% vs. 9 of 96; 9.4% P = 0.01). The two t-SCNC samples without bi-allelic RB1 inactivation harbored mono-allelic RB1 inactivation. We observed a significant association between RB1 expression levels and the number of predicted functional RB1 alleles, compatible with a dose effect of allelic loss (Figure 2A). All t-SCNC samples had significantly lower expression of RB1 (P = 0.003) and significantly higher expression of CDKN2A and E2F1 (P = 0.0002, 0.0004 respectively) compared to samples with an adenocarcinoma expression phenotype, consistent with genetic inactivation of RB1 (Figure 2B). Application of a validated RB1 loss gene expression signature6 likewise demonstrated significant enrichment in the t-SCNC samples (mean RB1 loss score of 3.79 in t-SCNC vs. −1.16 in samples with adenocarcinoma expression phenotype, P = 0.009). No other recurrent structural variant was significantly associated with the t-SCNC expression phenotype. Bi-allelic TP53 inactivation was present in three of five tumors with the t-SCNC expression phenotype (60%) vs. 43 of 96 adenocarcinoma samples (45%) (P > 0.05, Figure 1A). Likewise, bi-allelic PTEN inactivation was present in two of five tumors with t-SCNC phenotype (40%) vs. 33 of 96 adenocarcinoma samples (34%).
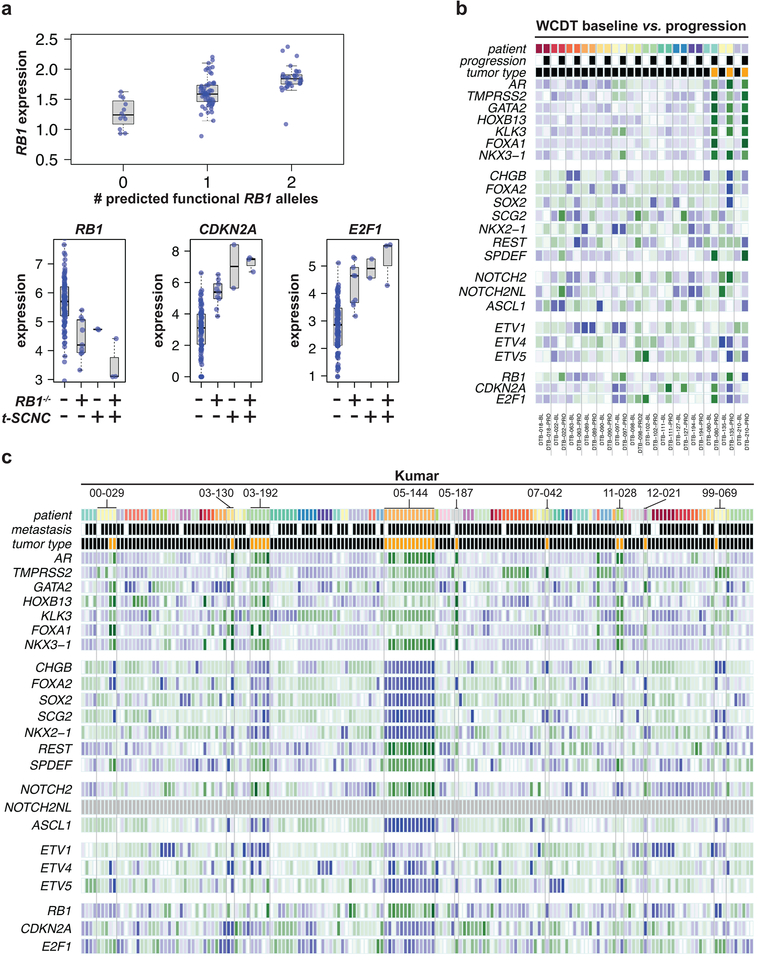
Figure 2: RB pathway expression is associated with t-SCNC status.
A) Top: expression of RB1 is significantly associated with the number of predicted functional alleles. Bottom: expression of RB1, CDKN2A, and E2F1 were significantly different in RB1−/− and t-SCNC tumors compared to RB1+/+ tumors with adenocarcinoma expression phenotype.
B) Heatmap showing expression of NEPC genes in paired WCDT baseline and progression mCRPC samples. Tumors from the same patient are indicated by a common color on the top line and set off by vertical lines. Baseline / progression status is indicated by white / black bar in second row. Adenocarcinoma / t-SCNC status assessed by gene expression profile is indicated by black / orange bar in third row.
C) Heatmap showing expression of NEPC genes in primary, localized, and mCRPC samples from Kumar data set6. All tumors from the same patient are indicated by a common color on the top line. Samples derived from primary / metastatic lesions are indicated by white / black on the second line. Adenocarcinoma / t-SCNC status assessed by gene expression profile is indicated by black / orange bar in third row.
Conversion to t-SCNC may be a late onset event in mCRPC
The timing and mechanism of conversion from adenocarcinoma to t-SCNC during the course of AR-targeting therapies and progression from castration-sensitive to castration-resistant disease remains to be elucidated. To begin to address this question, we identified fourteen patients in our previously published cohort6 of patients who had RNA-seq data available from metastatic CRPC biopsies obtained at two time points (baseline and progression). The clinical characteristics of these patients are shown in Supplementary Table 2. In 11 of the 14 patients, both baseline and progression biopsies had an adenocarcinoma gene expression profile (Figure 2B, left). Three patients (DTB-080, DTB-135, and DTB-210) had an adenocarcinoma expression phenotype in their baseline sample and t-SCNC expression profile at progression (Figure 2B, right). The previously published neuroendocrine gene set readily distinguished the three progression t-SCNC tumors in this analysis from the adenocarcinoma samples (Figure S1). The median duration of androgen signaling inhibitor therapy between baseline and progression biopsies did not differ between those with t-SCNC at progression versus those without (median duration 6.7 months vs. 9.5 months, p = 0.13). In the case of patients DTB-080 and DTB-210, both the baseline and progression biopsy samples were derived from the same lesion (in bone and lymph node, respectively). Paired samples from patient DTB-135 were obtained from adjacent lymph nodes in the same anatomic region. We observed consistent AR signaling loss and a gradient of small cell expression in the progression samples, ranging from very strong (DTB-135) to weak (DTB-080), further evidence of intra-class heterogeneity within t-SCNC.
To further assess intra-patient heterogeneity of the t-SCNC phenotype, we then re-analyzed a previously published cohort of 176 primary and metastatic prostate tumors6. The investigators of this autopsy series, wherein most patients contributed more than one tumor sample, had reported that in most cases a single metastasis was representative of all tumors present in a given patient. We confirmed that, as previously reported, all metastatic tumors from patients 03–192 (N = 5) and 05–144 (N = 12) harbored the NEPC phenotype (Figure 2C). However, we also noted patients with metastases harboring both t-SCNC and adenocarcinoma phenotypes (patients 00–029, 03–130, 05–187, 07–042). The neuroendocrine gene set distinguished t-SCNC and adenocarcinoma tumors in these samples, and confirmed the neuroendocrine expression phenotype of t-SCNC tumors that diverged in phenotype from adenocarcinomas within a given patient (Figure S1). We conclude that both the adenocarcinoma and neuroendocrine phenotypes can be present in distinct metastatic lesions within in the same patient, compatible with this phenotype arising after the independent establishment of physically distinct metastatic lesions.
Discussion
In this report we describe the structural variants of t-SCNC, a highly lethal variant of mCRPC, and how they differ from adenocarcinoma tumors. We undertook an unbiased clustering approach of the transcriptome to independently identify a subset of tumors harboring a t-SCNC expression profile and confirmed this classification using a previously published gene signature of neuroendocrine prostate cancer5. We observed a near mutual exclusivity with AR enhancer amplification, which has recently been reported to be present in up to 80% of mCRPC biopsies that do not harbor t-SCNC7. This finding, coupled with the enrichment of RB1 loss, distinguished t-SCNC from mCRPC without transcriptional features of small cell/neuroendocrine transformation. Using serial biopsies, we demonstrate temporal heterogeneity of t-SCNC differentiation arising in patients with established mCRPC. Based upon independent analysis of a previously published dataset, we demonstrate spatial heterogeneity of t-SCNC differentiation across metastatic lesions within an individual patient.
We and others have previously demonstrated that t-SCNC harbors lower canonical AR transcriptional output, yet with retained AR protein expression10. The results of the current analysis build upon these previous findings by virtue of whole genome interrogation and the finding of lower prevalence of amplification of upstream enhancer of AR. Notably, even in the two cases of t-SCNC with amplification of the AR gene locus or upstream enhancer, respectively, we observed lower expression of AR and its downstream transcriptional targets. This is consistent with a characterization of an AR ‘indifferent state’, and suggests that epigenetic dysregulation is a hallmark of t-SCNC15. Further study of the epigenetic landscape of t-SCNC will be required to ascertain the mechanisms underpinning the putative silencing of canonical AR-driven transcription.
In a pair-wise comparison of the transcriptional profile of biopsies obtained at two time points in patients with mCRPC, we observed the emergence of t-SCNC at progression in three patients. These observations, while preliminary given the limited sample size, are consistent with a model where t-SCNC develops late during progression after the onset of mCRPC and may be enriched following the use of second-generation androgen signaling inhibitors5. Whether the process driving the emergence of t-SCNC is related to transdifferentiation from an adenocarcinoma precursor versus clonal expansion remains to be defined. Further prospective studies in mCRPC patients with serial genomic and transcriptomic assessment at various time points are needed to delineate the kinetics and mechanism underpinning the emergence of t-SCNC.
Previously, our group and others demonstrated that loss of function of tumor suppressors RB1, TP53, and PTEN are hallmarks of small cell neuroendocrine prostate cancer3,6. Mechanistic studies have demonstrated that RB1- and TP53 deficient prostate cancer leads to lineage plasticity and resistance to antiandrogen therapy16–18. Loss of two or more of the three tumor suppressors has been shown to lead to worse treatment outcomes in a prior study and in pre-clinical models3,19. In the current analysis of the whole genome, RB1 loss stands out as the key genomic hallmark of t-SCNC, leading to up-regulation of E2F1 and acceleration of cell cycle progression. TP53 loss may promote a different genotype and phenotype than RB1 loss, characterized by marked frequency of inversions and chromothripsis7. Further interrogation of individual loss of each of these tumor suppressor genes is required to delineate specific effects on pathogenesis of t-SCNC and treatment outcomes.
Our finding of interpatient heterogeneity in the gradient of t-SCNC expression, together with intra-patient spatial and temporal heterogeneity of adenocarcinoma versus t-SCNC differentiation demonstrated in our dataset and others, has implications for the development of therapeutic strategies in this setting. Successful treatment of t-SCNC may require dual targeting with continuation of treatments aimed at blocking adenocarcinoma targets including the androgen receptor. The gradient of t-SCNC expression suggests plasticity in the emergence of this phenotype, and underscores the potential for epigenetically-mediated reversal of t-SCNC differentiation and restoring dependence on the androgen receptor. These strategies warrant prospective evaluation as novel targets and targeted therapies are evaluated in t-SCNC.
Supplementary Material
Implications:
The t-SCNC phenotype is characterized by lack of AR enhancer gain and loss of RB1 function, and demonstrates both inter-individual and intra-individual heterogeneity.
Acknowledgments
Research supported by a Stand Up To Cancer-Prostate Cancer Foundation Dream Team-Prostate Dream Team Translational Research Grant (SU2C-AACR-DT0812). This research grant is made possible by the generous support of the Movember Foundation. Stand Up to Cancer is a division of the Entertainment Industry Foundation. Research grants are administered by the American Association for Cancer Research, the Scientific Partner of SU2C.
U.S. Army Department of Defense Prostate Cancer Program Impact Award (W81XWH-16-1-0495 to E. Small)
Prostate Cancer Foundation Special Challenge Award (15CHAS03 to E. Small)
Prostate Cancer Foundation Young Investigator Award (to D. Quigley)
Footnotes
Conflicts of interest: The authors declare no potential conflicts of interest.
References
- 1.Robinson D et al. Integrative Clinical Genomics of Advanced Prostate Cancer. Cell 161, 1215–1228, doi: 10.1016/j.cell.2015.05.001 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ryan CJ et al. Abiraterone in metastatic prostate cancer without previous chemotherapy. The New England journal of medicine 368, 138–148, doi: 10.1056/NEJMoa1209096 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Aparicio AM et al. Combined Tumor Suppressor Defects Characterize Clinically Defined Aggressive Variant Prostate Cancers. Clin Cancer Res 22, 1520–1530, doi: 10.1158/1078-0432.CCR-15-1259 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Beltran H et al. Molecular characterization of neuroendocrine prostate cancer and identification of new drug targets. Cancer Discov 1, 487–495, doi: 10.1158/2159-8290.CD-11-0130 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Beltran H et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat Med 22, 298–305, doi: 10.1038/nm.4045 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kumar A et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat Med 22, 369–378, doi: 10.1038/nm.4053 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Quigley DA et al. Genomic Hallmarks and Structural Variation in Metastatic Prostate Cancer. Cell 174, 758–769 e759, doi: 10.1016/j.cell.2018.06.039 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Takeda DY et al. A Somatically Acquired Enhancer of the Androgen Receptor Is a Noncoding Driver in Advanced Prostate Cancer. Cell 174, 422–432 e413, doi: 10.1016/j.cell.2018.05.037 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Viswanathan SR et al. Structural Alterations Driving Castration-Resistant Prostate Cancer Revealed by Linked-Read Genome Sequencing. Cell 174, 433–447 e419, doi: 10.1016/j.cell.2018.05.036 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Aggarwal R et al. Clinical and Genomic Characterization of Treatment-Emergent Small-Cell Neuroendocrine Prostate Cancer: A Multi-institutional Prospective Study. J Clin Oncol 36, 2492–2503, doi: 10.1200/JCO.2017.77.6880 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Aggarwal R et al. Targeting Adaptive Pathways in Metastatic Treatment-Resistant Prostate Cancer: Update on the Stand Up 2 Cancer/Prostate Cancer Foundation-Supported West Coast Prostate Cancer Dream Team. European Urology Focus 2, 469–471, doi: 10.1016/j.euf.2016.10.011 (2016). [DOI] [PubMed] [Google Scholar]
- 12.Guillemot F et al. Mammalian achaete-scute homolog 1 is required for the early development of olfactory and autonomic neurons. Cell 75, 463–476 (1993). [DOI] [PubMed] [Google Scholar]
- 13.Paratore C, Brugnoli G, Lee HY, Suter U & Sommer L The role of the Ets domain transcription factor Erm in modulating differentiation of neural crest stem cells. Dev Biol 250, 168–180 (2002). [DOI] [PubMed] [Google Scholar]
- 14.Lin JH et al. Functionally related motor neuron pool and muscle sensory afferent subtypes defined by coordinate ETS gene expression. Cell 95, 393–407 (1998). [DOI] [PubMed] [Google Scholar]
- 15.Chen R, Dong X & Gleave M Molecular model for neuroendocrine prostate cancer progression. BJU Int 122, 560–570, doi: 10.1111/bju.14207 (2018). [DOI] [PubMed] [Google Scholar]
- 16.Ku SY et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science (New York, N.Y.) 355, 78–83, doi: 10.1126/science.aah4199 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Mu P et al. SOX2 promotes lineage plasticity and antiandrogen resistance in TP53- and RB1-deficient prostate cancer. Science (New York, N.Y.) 355, 84–88, doi: 10.1126/science.aah4307 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Park JW et al. Reprogramming normal human epithelial tissues to a common, lethal neuroendocrine cancer lineage. Science (New York, N.Y.) 362, 91–95, doi: 10.1126/science.aat5749 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zou M et al. Transdifferentiation as a Mechanism of Treatment Resistance in a Mouse Model of Castration-Resistant Prostate Cancer. Cancer Discov 7, 736–749, doi: 10.1158/2159-8290.CD-16-1174 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.