Abstract
Background
Evidence is accumulating to characterise the key differences between systemic sclerosis (SSc) and rheumatoid arthritis (RA), which are similar but distinct systemic autoimmune diseases. However, the differences at the genetic level are not yet clear. Therefore, the aim of the present study was to identify key differential genes between patients with SSc and RA.
Methods
The Gene Expression Omnibus database was used to identify differentially expressed genes (DEGs) between SSc and RA biopsies. The DEGs were then functionally annotated using Gene Ontology (GO) terms and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways with the Database for Annotation, Visualization and Integrated Discovery (DAVID) tools. A protein–protein interaction (PPI) network was constructed with Cytoscape software. The Molecular Complex Detection (MCODE) plugin was also used to evaluate the biological importance of the constructed gene modules.
Results
A total of 13,556 DEGs were identified between the five SSc patients and seven RA patients, including 13,465 up-regulated genes and 91 down-regulated genes. Interestingly, the most significantly enriched GO terms of up- and down-regulated genes were related to extracellular involvement and immune activity, respectively, and the top six highly enriched KEGG pathways were related to the same processes. In the PPI network, the top 10 hub nodes and top four modules harboured the most relevant genes contributing to the differences between SSc and RA, including key genes such as IL6, EGF, JUN, FGF2, BMP2, FOS, BMP4, LRRK2, CTNNB1, EP300, CD79, and CXCL13.
Conclusions
These genes such as IL6, EGF, JUN, FGF2, BMP2, FOS, BMP4, LRRK2, CTNNB1, EP300, CD79, and CXCL13 can serve as new targets for focused research on the distinct molecular pathogenesis of SSc and RA. Furthermore, these genes could serve as potential biomarkers for differential diagnoses or therapeutic targets for treatment.
Electronic supplementary material
The online version of this article (10.1186/s41065-019-0091-y) contains supplementary material, which is available to authorized users.
Keywords: Gene expression data, Systemic sclerosis, Rheumatoid arthritis, Microarray, Differentially expressed genes, Key genes
Introduction
Systemic sclerosis (SSc) is an autoimmune disease [1] that is often characterised by joint involvement, especially arthritis [2]. However, the degree of synovial inflammation in SSc has not yet been characterised thoroughly [3]. Rheumatoid arthritis (RA) is another autoimmune disease associated with articular damage and consequent disability, which may lead to several complications [4]. The pathogenesis of RA involves excessive reaction of immune components leading to severe inflammation of the joints. Along with recent progress in the diagnosis of SSc and RA based on updated signs and symptoms, several potential diagnostic and therapeutic targets have been uncovered.
In general, SSc and RA are diagnosed by auxiliary approaches such as clinical manifestations, biochemical indicators, and X-ray findings [5, 6]. Since they are both autoimmune diseases with similar clinical signs and symptoms, especially joint involvement, it is not easy to distinguish between them in some cases with uncharacteristic signs and symptoms. To improve the differential diagnosis and therapy of SSc and RA, it is necessary to identify genetic markers that are sufficiently sensitive and highly specific for the two diseases in order to initiate the correct course of treatment.
Gene expression profiling with microarrays is regarded as a standard method for identifying differentially expressed genes (DEGs) and potential biological pathways associated with SSc [7] and RA [8]. To the best of our knowledge, no specific genomic expression analyses have been conducted to distinguish SSc and RA to date. Therefore, in the present study, we investigated the genomic expression profiles to identify DEGs between SSc and RA using a part of the GSE93698 microarray database, including transcriptome data of five SSc tenosynovial biopsy samples and seven RA synovial biopsy samples. Moreover, Gene Ontology (GO) enrichment analyses and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses were used to perform functional enrichment analysis and identify important biological pathways related to the identified DEGs. In addition, the Retrieval of Interacting Genes (STRING) database was used to construct a protein–protein interaction (PPI) network. Finally, the hub genes of the network were analysed using Cytoscape software, which were used to establish the most significant modules that differentiate SSc and RA.
Material and methods
Data source
Gene expression data of seven SSc tenosynovial biopsy samples and five RA synovial biopsy samples were obtained from the Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo/) database. The GSE93698 data were derived from the GPL570 microarray platform [HG-U133_Plus_2] Affymetrix Human Genome U133A 2.0 Array.
Identification of DEGs
Differentially expressed genes (DEGs), including up- and down-regulated genes, were identified between SSc and RA through the R package limma [9, 10] based on the criteria of a statistically significant difference in expression levels (p < 0.05) through a t-test [11] and a fold change (FC) > 2. Subsequently, DEGs were ultimately selected according to a false discovery rate < 0.05 and |logFC| > 1.
Functional enrichment analysis
Gene functional enrichment analyses included classifying gene functions and identifying gene conversions, which were performed through determining enriched Gene ontology (GO) terms [12] and the Kyoto Encyclopedia of Genes and Genomes (KEGG) [13] pathways with the online tool the Database for Annotation, Visualization and Integrated Discovery (DAVID) [14, 15]. Significantly enriched terms/pathways were those with a P-value < 0.05 and gene number ≥ 2. In addition, GOplot was used for visualisation of detailed information of the molecules in functional enrichments [16].
Construction of the PPI network and module analysis
A PPI network was established using the Search Tool for the Retrieval of Interacting Genes (STRING) database [17] based on the significantly up- and down-regulated DEGs to identify the most crucial genes and modules differentiating SSc and RA. A combined score > 0.4 was selected as the cut-off value to construct the PPI network, which was visualised using Cytoscape software. The Molecular Complex Detection (MCODE) plugin was also used to evaluate the biological importance of the constructed gene modules [18]. The top 10 essential nodes ranked by degree were selected, and modules were selected with an MCODE score > 6 and number of nodes > 6.
Results
DEGs identification by microarray expression profiling
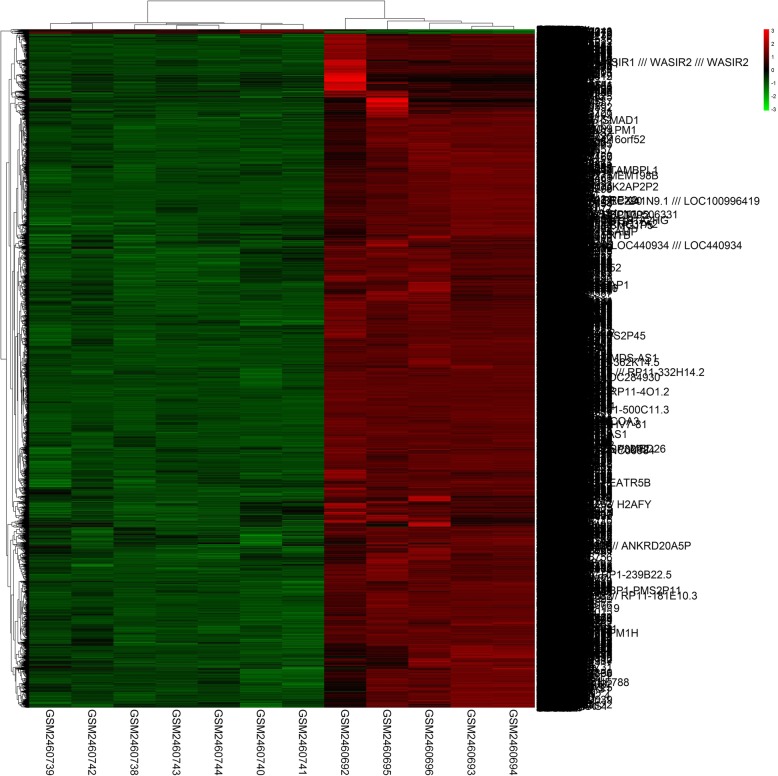
Using the GEO GSE93698 dataset of microarray data, we identified a total of 13,556 DEGs (p < 0.05 and |logFC| > 1) between SSc and RA samples, including 13,465 up-regulated genes and 91 down-regulated genes. Thus, the great majority of genes that are specifically involved in the SSc pathological process were up-regulated compared to those in RA. The heatmap of the DEGs identified is shown in Fig. 1.
Fig. 1.
Heatmap of the 13,556 differentially expressed genes (DEGs) between systemic sclerosis and rheumatoid arthritis
GO functional enrichment
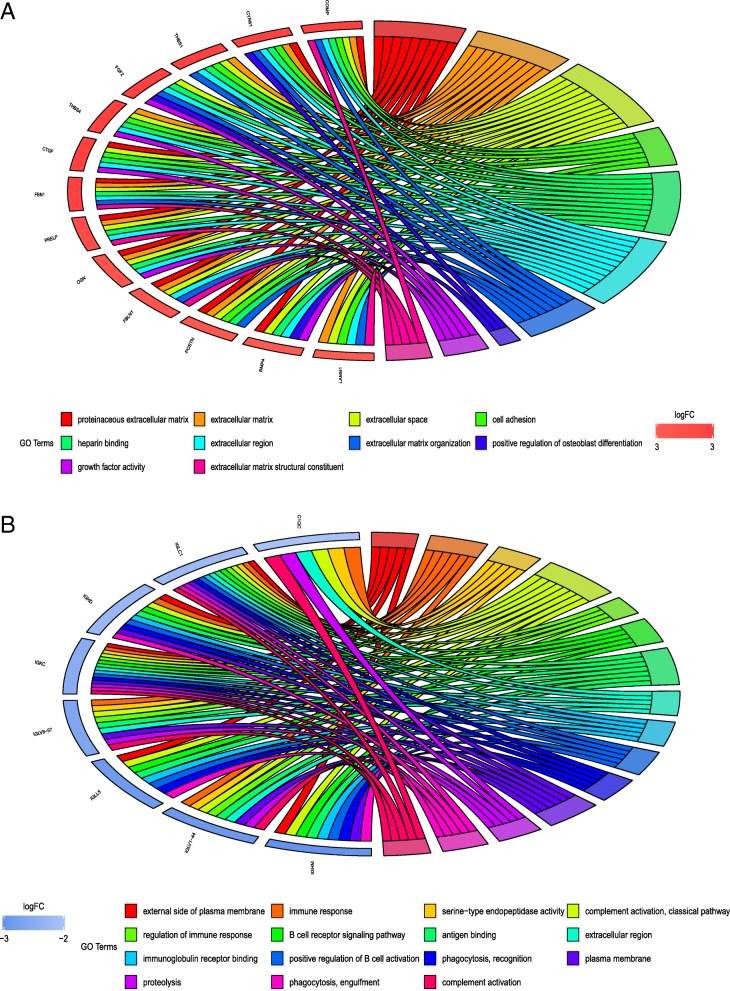
To investigate the functions of the large range of gene signatures obtained, we performed GO enrichment analysis from the GO database [19] including terms of the biological process, molecular function, and cellular component categories for the top 1000 up-regulated genes and 91 down-regulated genes (Fig. 2).
Fig. 2.
Gene Ontology enrichment analyses. a Gene Ontology enrichment of up-regulated genes. b Gene Ontology enrichment of down-regulated genes. The criteria for enrichment were: P-value < 0.05, FDR < 0.05, and fold enrichment > 1. Each gene was assigned to at least six terms
Ten terms were enriched for the up-regulated genes, which were predominantly related to extracellular activities in the biological process, molecular function, and cellular component categories. In the cellular component cluster, representative terms were related to proteinaceous extracellular matrix (ECM), ECM, extracellular space, and extracellular region. In addition, the biological process cluster included terms of cell adhesion, ECM organization, and positive regulation of osteoblast differentiation. The significantly enriched terms for the molecular function cluster were heparin binding, growth factor activity, and ECM structural constituent. The up-regulated genes COMP, CYR61, THBS1, FGF2, THBS4, CTGF, FBN1, PRELP, OGN, FBLN1, POSTN, BMP4, and LAMB1 appeared frequently in these terms (Fig. 2). However, the 15 enrichment terms related to the down-regulated genes were mainly related to immune activities, which could have important clinical implications. These representative down-regulated genes enriched in these terms were C1QC, IGLC1, IGHD, IGKC, IGLV6–57, IGLL5, IGLV1–44, and IGHM.
KEGG pathways analysis
KEGG pathway analysis identified the top six important KEGG pathways of the up- and down-regulated DEGs (Table 1), including ECM–receptor interaction, Wnt signalling pathway, transforming growth factor-beta (TGFβ) signalling pathway, primary immunodeficiency, hematopoietic cell lineage, and cytokine–cytokine receptor interaction.
Table 1.
Top 3 the most significantly enriched KEGG pathways of up and down regulated DEGs respectively
| Regulate | Term | Count | P-value | Genes | Fold Enrichment |
|---|---|---|---|---|---|
| up | hsa04512:ECM-receptor interaction | 16 | 4.35E-06 | COL4A4, ITGA1, ITGA2, ITGA10, CHAD, LAMA2, CD36, COMP, RELN, TNN, THBS1, LAMB1, COL24A1, THBS2, COL11A1, THBS4 | 4.194075 |
| up | hsa04310:Wnt signaling pathway | 19 | 2.80E-05 | VANGL2, MMP7, PPP3R1, FZD7, FZD6, CTNNB1, WNT2, GPC4, PLCB4, EP300, DKK1, PRICKLE1, SFRP2, JUN, SFRP4, PRICKLE2, WNT9A, SOX17, BAMBI | 3.139857 |
| up | hsa04350:TGF-beta signaling pathway | 14 | 6.24E-05 | BMP4, BMP2, FST, BMPR2, SMAD1, ACVR1C, INHBA, EP300, ID1, BAMBI, THBS1, BMPR1B, BMP5, ACVR1 | 3.80088 |
| down | hsa05340:Primary immunodeficiency | 6 | 3.89E-07 | CD19, CD3D, CD8A, IL2RG, CD79A, IL7R | 36.95187 |
| down | hsa04640:Hematopoietic cell lineage | 7 | 2.03E-06 | CD19, CD3D, CD8A, MS4A1, CD2, IL7R, CSF1R | 17.24421 |
| down | hsa04060:Cytokine-cytokine receptor interaction | 6 | 0.003777 | CXCL13, IL21R, TNFRSF17, IL2RG, IL7R, CSF1R | 5.462451 |
KEGG Kyoto encyclopedia of genes and genomes, DEGs differentially expressed genes
Construction of the PPI network and module analysis
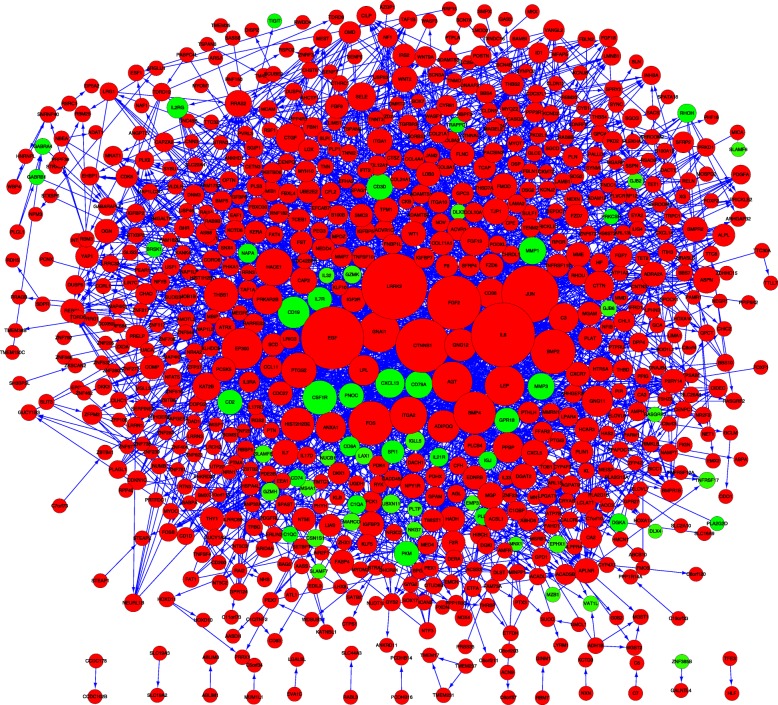
The top 1000 up-regulated genes and 91 down-regulated genes were mapped by the STRING database to establish a PPI network (Fig. 3). Protein pairs with a combined score of > 0.4 were selected. According to the information from STRING, the top 10 hub nodes with high degrees were identified using the Cytoscape tool, including LRRK2, IL6, EGF, JUN, CTNNB1, FGF2, BMP2, FOS, BMP4, and EP300 (Additional file 1: Table S1). The largest node degrees were detected for CXCL5, CXCL13, GPR18, NPY1R, ADRA2A, CXCR7, AGT, GNAI1, HTR5A, HCAR3, GNG11, P2RY14, ANXA1, PPBP, C3, PNOC, GNG12, and APLNR (Additional file 2: Table S2), suggesting that these genes may play an important role in the pathological process.
Fig. 3.
Protein–protein interaction network constituted by differentially expressed genes identified in this study. Red nodes represent up-regulated genes, and green nodes represent down-regulated genes
The top four modules of the PPI network are presented in Fig. 4. Enrichment analyses of these modules (Table 2) showed that genes in module 1 (including PPBP, CXCL13, GNG12, CXCL5, HTR5A, GNAI1, P2RY14, HTR5A, and ADRA2A) were associated with chemokine signalling pathway, serotonergic synapse, and neuroactive ligand-receptor interaction. Module 2 included NEDD4, CDC27, UBE2E2, and TCEB1 associated with ubiquitin-mediated proteolysis and renal cell carcinoma pathways. In addition, pathways in cancer, signalling pathways regulating stem cells, and the Hippo signalling pathway were central components of module 3, which was enriched in the key genes JUN, WNT9A, FGF2, BMP4, BMP2, IL6, COL4A4, and FGF13, among others. Finally, module 4 included the genes PLCB4, MMP1, CTNNB1, EGF, F2R, and LPAR4, associated with pathways in cancer, Rap1 signalling pathway, and phospholipase D signalling pathways.
Fig. 4.
Four significant modules identified from the protein–protein interaction network using the molecular complex detection method with a score > 6.0: Module 1 (MCODE score = 18), Module 2 (MCODE score = 11), Module 3 (MCODE score = 8.035), and Module 4 (MCODE score = 7.048)
Table 2.
KEGG pathway enriched by differentially expressed genes in different modules
| Term | Description | P-value | Genes |
|---|---|---|---|
| Module1 | |||
| hsa04062 | Chemokine signaling pathway | 1.04E-08 | PPBP;CXCL13;GNG12;GNG11;CXCL5;GNAI1 |
| hsa04726 | Serotonergic synapse | 2.68E-06 | HTR5A;GNG12;GNG11;GNAI1 |
| hsa04080 | Neuroactive ligand-receptor interaction | 3.63E-06 | P2RY14;NPY1R;APLNR;HTR5A;ADRA2A |
| Module2 | |||
| hsa04120 | Ubiquitin mediated proteolysis | 6.7E-07 | NEDD4;CDC27;UBE2E2;TCEB1 |
| hsa05211 | Renal cell carcinoma | 0.035716 | TCEB1 |
| Module3 | |||
| hsa05200 | Pathways in cancer | 5.39E-12 | CSF1R;JUN;FZD7;FZD6;PDGFA;WNT9A;FOS;FGF2;BMP4;FGF7;BMP2;IL6;COL4A4;FGF13 |
| hsa04550 | Signaling pathways regulating pluripotency of stem cells | 7.52E-09 | BMP4;BMP2;ACVR1C;FZD7;FZD6;WNT9A;BMPR1B;FGF2 |
| hsa04390 | Hippo signaling pathway | 1.35E-08 | BMP4;BMP2;FZD7;FZD6;WNT9A;BMPR1B;CTGF;BMP5 |
| Module 4 | |||
| hsa05200 | Pathways in cancer | 1.73E-10 | PLCB4;EDNRB;EGF;MMP1;ITGA2;F2R;CTNNB1;LPAR4;PTGS2 |
| hsa04015 | Rap1 signaling pathway | 8.32E-08 | PLCB4;EGF;F2R;CTNNB1;LPAR4;THBS1 |
| hsa04072 | Phospholipase D signaling pathway | 4.31E-07 | DNM3;PLCB4;EGF;F2R;LPAR4 |
Top 3 terms were selected according to P-value when more than 3 terms enriched terms were identified in each category
Discussion
Despite progress in the differential diagnoses between SSc and RA, a more effective and sensitive method for helping to distinguish between these two diseases is needed. The present findings highlight some distinct pathological molecular mechanisms of these two diseases, which may provide further information as therapeutic targets. Of the 13,556 DEGs including 13,465 up-regulated genes and 91 down-regulated genes identified between SSc and RA samples. Interestingly, the number of up-regulated genes were far more than the number of down-regulated genes, which was related to huge different characteristics between these two different diseases. The majority were up-regulated in SSc; these genes were mainly related to extracellular activities, which should be taken into account in further studies on SSc pathology. SSc is an autoimmune rheumatic disease with multisystem fibrosis manifestations. In normal physiological conditions, fibroblasts can be protected by the ECM, whereas the damaged fibroblasts in SSc attached to the ECM are destroyed [20]. Abnormal ECM remodelling mechanisms are linked to the fibrosis that occurs in connective tissue diseases. Excessive ECM, including collagens, hyaluronic acid, fibronectin, and proteoglycans, promotes scarring and sustained fibrosis, leading to excessive scar tissue [21]. Some studies suggested that overexpression of ECM genes may have a central effect in fibrotic cells [22–24]. Pathologically activated fibrosis arises from high accumulation of ECM components, highlighting a novel therapeutic approach in SSc [25]. However, we further showed that many of the down-regulated genes in SSc were related to immune activities, which warrants further investigation to identify new therapeutic targets. Indeed, the pathological process of RA is well known to be related to an excessive or dysregulated immune response, including an abnormal autoimmune response and genetic susceptibility. Immune cells such as dendritic cells, T cells, B cells, and natural killer cells are all related to the development of RA [26]; thus, genes regulating immune activities have been highlighted as potential therapeutic targets for RA [27]. For example, anti-CD79A antibody therapy was shown to enhance immune system recovery against autoimmunity [28]. Thus, our results further indicate that fibrosis and other clinical manifestations may be related to the immune response.
KEGG pathway enrichment analyses of the identified DEGs demonstrated six key significantly enriched pathways, including ECM–receptor interaction and primary immunodeficiency, which is in line with the known pathological mechanisms. However, the other pathways identified, including Wnt signalling pathway, TGFβ signalling pathway, hematopoietic cell lineage, and cytokine–cytokine receptor interaction, deserve new attention. The selective stabilization of β-catenin leads to excessive ECM production [29], and Wnt may induce the fibroblast activation and abundant collagen production related to SSc [30]. Indeed, the Wnt pathway has been proposed to be a core factor involved in the progression of SSc [30]. Moreover, the Wnt cascade tightly interacts with TGFβ signalling, which may be involved in ECM activities [31]. Cells of the hematopoietic lineage such as CD79a-positive B cells are helpful for diagnosing RA [32], and this pathway has deep associations with many immune hematopoietic cells. Similarly, the cytokine–cytokine receptor interaction is another important component of the RA pathological mechanism [33]. Therefore, monitoring these signalling pathways may aid in the prediction of the progression of these two diseases.
The PPI network constructed with the DEGs resulted in 10 hub genes that can be used to differentiate between SSc and RA: LRRK2, IL6, EGF, JUN, CTNNB1, FGF2, BMP2, FOS, BMP4, and EP300. These genes have also been previously highlighted to play a role in the pathogenesis of SSc and RA. IL-6, which is linked to the ECM, can regulate collagen synthesis by fibroblasts in SSc [34, 35], and EGF was shown to up-regulate TGFRII expression in SSc fibroblasts [36]; thus, EGFR signalling was suggested as a therapeutic target in fibrotic diseases [37]. FGF-2 plays a role in the pathogenic process of pulmonary arterial hypertension, which modifies pulmonary vascular remodelling leading to the vascular manifestations in SSc [38]. FGF, which regulates the synthesis of collagen and ECM components, is up-regulated by TGFβ in SSc, resulting in an increase in BMP signalling [39]. The AP-1 family members c-Jun, c-Fos, and JunD are also known to play an important role in SSc [40, 41]. JunD is a downstream mediator of TGFβ signalling [40], and Jun N-terminal kinases are regarded as intracellular mediators that may be affected by TGFβ [41]. Thus, almost all of these genes are associated with TGFβ, which is known to play a key role in fibrotic diseases. LRRK2 [42] and CTNNB1 [43] were also reported to interact with the Wnt pathway in fibromatosis. However, EP300 is mainly known to influence specific T cell states [44]. Although LRRK2, CTNNB1, and EP300 have not been previously clearly associated with SSc, our results suggest that they may be potential biomarkers of this disease, and thus worthy of further investigation.
Module analyses of the PPI network further revealed the differential development of SSc and RA. CXCL13 in module 1, related to the chemokine signalling pathway, has been linked to joint inflammation and the development of autoimmune disorders, including RA [45]. CXCL13 is a proinflammatory cytokine that can serve as a biomarker in early RA and reflects the severity of synovitis [46]. Thus, CXCL13 appears to represent a new direction for RA treatment. FGF2 and IL6 in module 3 are regarded as important factors for SSc development based on the background summarised above, as is EGF, part of module 4, which mediates the up-regulation of TGFβ receptor in SSc. However, many genes identified in this study have not been previously associated with SSc or RA, such as NEDD4, CDC27, and UBE2E2 in module 2. Thus, attention should be focused on these genes in future work as potential biomarkers and therapeutic targets.
Conclusions
This study identified a series of core genes and pathways that differentiate SSc and RA. Compared to RA, the majority of the up-regulated genes in SSc were related to extracellular activities, whereas the down-regulated genes were mainly related to immune activities. These two distinct processes can provide a new direction for methods of clinically distinguishing between these two diseases by focusing on the involvement of extracellular and immune activities. On the one hand, IL6, EGF, JUN, FGF2, BMP2, FOS, and BMP4 should be further studied as they may play an essential and specific role in the SSc pathogenesis. On the other hand, CD79 and CXCL13 might be representative genes for RA. These genes might be used as biomarkers to improve the differential diagnosis and treatment of SSc and RA. However, many of the other genes identified in this study have not been previously reported to be associated with these diseases, such as LRRK2, CTNNB1, and EP300, and should not be ignored. Thus, further studies and clinical trials are needed to verify our findings and establish reliable diagnostic or therapeutic targets.
Additional files
Table S1. Degree of top 10 genes. (DOCX 14 kb)
Table S2. The modules of the PPI network. Four modules from the protein-protein interaction network satisfied the criteria of MCODE scores >6 and number of nodes >6. (DOCX 14 kb)
Acknowledgements
The authors acknowledge the efforts of the group who created the GEO database.
Funding
This study was funded by National Natural Science Foundation of China (81772361).
Abbreviation
- DAVID
the Database for Annotation, Visualization and Integrated Discovery
- DEG
Differentially expressed gene
- ECM
Extracellular matrix
- GO
Gene Ontology
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- MCODE
the Molecular Complex Detection
- PPI
Protein–protein interaction
- RA
Rheumatoid arthritis
- SSc
System sclerosis
- STRING
the Search Tool for the Retrieval of Interacting Genes
- TGFβ
Transforming growth factor-beta
Authors’ contributions
ZS contributed significantly to analysis and manuscript preparation; WW performed the data analyses and wrote the manuscript; DY helped contribute to the conception of the study; YM contributed to the conception of the study. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Degang Yu, Phone: 18019790572, Email: ydg163@126.com.
Yuanqing Mao, Phone: 18019790572, Email: yuanqingmao@163.com.
References
- 1.Denton CP, Khanna D. Systemic sclerosis. Lancet. 2017;390(10103):1685–1699. doi: 10.1016/S0140-6736(17)30933-9. [DOI] [PubMed] [Google Scholar]
- 2.Allali F, Tahiri L, Senjari A, Abouqal R, Hajjaj-Hassouni N. Erosive arthropathy in systemic sclerosis. BMC Public Health. 2007;7:260. doi: 10.1186/1471-2458-7-260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chitale S, Ciapetti A, Hodgson R, Grainger A, O'Connor P, Goodson NJ, et al. Magnetic resonance imaging and musculoskeletal ultrasonography detect and characterize covert inflammatory arthropathy in systemic sclerosis patients with arthralgia. Rheumatology. 2010;49(12):2357–2361. doi: 10.1093/rheumatology/keq268. [DOI] [PubMed] [Google Scholar]
- 4.Ostrowska M, Maslinski W, Prochorec-Sobieszek M, Nieciecki M, Sudol-Szopinska I. Cartilage and bone damage in rheumatoid arthritis. Reumatologia. 2018;56(2):111–120. doi: 10.5114/reum.2018.75523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wasserman A. Rheumatoid arthritis: common questions about diagnosis and management. Am Fam Physician. 2018;97(7):455–462. [PubMed] [Google Scholar]
- 6.Distler O, Allanore Y, Denton CP, Matucci-Cerinic M, Pope JE, Hinzmann B, et al. Factors influencing early referral, early diagnosis and management in patients with diffuse cutaneous systemic sclerosis. Rheumatology (Oxford) 2018;57(5):813–817. doi: 10.1093/rheumatology/kex504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Taroni JN, Greene CS, Martyanov V, Wood TA, Christmann RB, Farber HW, et al. A novel multi-network approach reveals tissue-specific cellular modulators of fibrosis in systemic sclerosis. Genome Med. 2017;9(1):27. doi: 10.1186/s13073-017-0417-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sun L, Chai Y. Bioinformatic analysis to find small molecules related to rheumatoid arthritis. Int J Rheum Dis. 2014;17(1):71–77. doi: 10.1111/1756-185X.12201. [DOI] [PubMed] [Google Scholar]
- 9.Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3. doi: 10.2202/1544-6115.1027. [DOI] [PubMed] [Google Scholar]
- 10.Smyth GK, Michaud J, Scott HS. Use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics. 2005;21(9):2067–2075. doi: 10.1093/bioinformatics/bti270. [DOI] [PubMed] [Google Scholar]
- 11.Baldi P, Long AD. A Bayesian framework for the analysis of microarray expression data: regularized t -test and statistical inferences of gene changes. Bioinformatics. 2001;17(6):509–519. doi: 10.1093/bioinformatics/17.6.509. [DOI] [PubMed] [Google Scholar]
- 12.Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009;10:48. doi: 10.1186/1471-2105-10-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nakaya A, Katayama T, Itoh M, Hiranuka K, Kawashima S, Moriya Y, et al. KEGG OC: a large-scale automatic construction of taxonomy-based ortholog clusters. Nucleic Acids Res. 2013;41(Database issue):D353–D357. doi: 10.1093/nar/gks1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Huang d W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
- 15.Huang d W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13. doi: 10.1093/nar/gkn923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Walter W, Sanchez-Cabo F, Ricote M. GOplot: an R package for visually combining expression data with functional analysis. Bioinformatics. 2015;31(17):2912–2914. doi: 10.1093/bioinformatics/btv300. [DOI] [PubMed] [Google Scholar]
- 17.Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41(Database issue):D808–D815. doi: 10.1093/nar/gks1094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bader Gary D, Hogue Christopher WV. BMC Bioinformatics. 2003;4(1):2. doi: 10.1186/1471-2105-4-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Dennis G, Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4(5):P3. doi: 10.1186/gb-2003-4-5-p3. [DOI] [PubMed] [Google Scholar]
- 20.Leask A. Getting out of a sticky situation: targeting the myofibroblast in scleroderma. Open Rheumatol J. 2012;6:163–169. doi: 10.2174/1874312901206010163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Leask A. Matrix remodeling in systemic sclerosis. Semin Immunopathol. 2015;37(5):559–563. doi: 10.1007/s00281-015-0508-2. [DOI] [PubMed] [Google Scholar]
- 22.Ichiki Y, Smith EA, LeRoy EC, Trojanowska M. Basic fibroblast growth factor inhibits basal and transforming growth factor-beta induced collagen alpha 2(I) gene expression in scleroderma and normal fibroblasts. J Rheumatol. 1997;24(1):90–95. [PubMed] [Google Scholar]
- 23.Sandorfi N, Louneva N, Hitraya E, Hajnoczky G, Saitta B, Jimenez SA. Inhibition of collagen gene expression in systemic sclerosis dermal fibroblasts by mithramycin. Ann Rheum Dis. 2005;64(12):1685–1691. doi: 10.1136/ard.2005.037515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lee CG, Herzog EL, Ahangari F, Zhou Y, Gulati M, Lee CM, et al. Chitinase 1 is a biomarker for and therapeutic target in scleroderma-associated interstitial lung disease that augments TGF-beta1 signaling. J Immunol. 2012;189(5):2635–2644. doi: 10.4049/jimmunol.1201115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Dave AJ, Fiorentino D, Lingala B, Krishnan E, Chung L. Atherosclerotic cardiovascular disease in hospitalized patients with systemic sclerosis: higher mortality than patients with lupus and rheumatoid arthritis. Arthritis Care Res (Hoboken) 2014;66(2):323–327. doi: 10.1002/acr.22152. [DOI] [PubMed] [Google Scholar]
- 26.Chen M, Wang YH, Wang Y, Huang L, Sandoval H, Liu YJ, et al. Dendritic cell apoptosis in the maintenance of immune tolerance. Science. 2006;311(5764):1160–1164. doi: 10.1126/science.1122545. [DOI] [PubMed] [Google Scholar]
- 27.Pappas DA, Geraldino-Pardilla L, Bathon JM. Immune modulation of rheumatoid arthritis. Best Pract Res Clin Rheumatol. 2011;25(6):873–889. doi: 10.1016/j.berh.2011.11.003. [DOI] [PubMed] [Google Scholar]
- 28.Hardy IR, Anceriz N, Rousseau F, Seefeldt MB, Hatterer E, Irla M, et al. Anti-CD79 antibody induces B cell anergy that protects against autoimmunity. J Immunol. 2014;192(4):1641–1650. doi: 10.4049/jimmunol.1302672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Beyer C, Schramm A, Akhmetshina A, Dees C, Kireva T, Gelse K, et al. Beta-catenin is a central mediator of pro-fibrotic Wnt signaling in systemic sclerosis. Ann Rheum Dis. 2012;71(5):761–767. doi: 10.1136/annrheumdis-2011-200568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dees C, Schlottmann I, Funke R, Distler A, Palumbo-Zerr K, Zerr P, et al. The Wnt antagonists DKK1 and SFRP1 are downregulated by promoter hypermethylation in systemic sclerosis. Ann Rheum Dis. 2014;73(6):1232–1239. doi: 10.1136/annrheumdis-2012-203194. [DOI] [PubMed] [Google Scholar]
- 31.Usategui A, del Rey MJ, Pablos JL. Fibroblast abnormalities in the pathogenesis of systemic sclerosis. Expert Rev Clin Immunol. 2011;7(4):491–498. doi: 10.1586/eci.11.39. [DOI] [PubMed] [Google Scholar]
- 32.Mo YQ, Dai L, Zheng DH, Zhu LJ, Wei XN, Pessler F, et al. Synovial infiltration with CD79a-positive B cells, but not other B cell lineage markers, correlates with joint destruction in rheumatoid arthritis. J Rheumatol. 2011;38(11):2301–2308. doi: 10.3899/jrheum.110615. [DOI] [PubMed] [Google Scholar]
- 33.An Q, Yan W, Zhao Y, Yu K. Enhanced neutrophil autophagy and increased concentrations of IL-6, IL-8, IL-10 and MCP-1 in rheumatoid arthritis. Int Immunopharmacol. 2018;65:119–128. doi: 10.1016/j.intimp.2018.09.011. [DOI] [PubMed] [Google Scholar]
- 34.Duncan MR, Berman B. Stimulation of collagen and glycosaminoglycan production in cultured human adult dermal fibroblasts by recombinant human interleukin 6. J. Invest. Dermatol. 1991;97(4):686–692. doi: 10.1111/1523-1747.ep12483971. [DOI] [PubMed] [Google Scholar]
- 35.Sato S, Hasegawa M, Takehara K. Serum levels of interleukin-6 and interleukin-10 correlate with total skin thickness score in patients with systemic sclerosis. J Dermatol Sci. 2001;27(2):140–146. doi: 10.1016/S0923-1811(01)00128-1. [DOI] [PubMed] [Google Scholar]
- 36.Yamane K, Ihn H, Tamaki K. Epidermal growth factor up-regulates expression of transforming growth factor beta receptor type II in human dermal fibroblasts by phosphoinositide 3-kinase/Akt signaling pathway: resistance to epidermal growth factor stimulation in scleroderma fibroblasts. Arthritis Rheum. 2003;48(6):1652–1666. doi: 10.1002/art.11029. [DOI] [PubMed] [Google Scholar]
- 37.Beyer C, Distler JH. Tyrosine kinase signaling in fibrotic disorders: Translation of basic research to human disease. Biochim biophys acta. 2013;1832(7):897–904. doi: 10.1016/j.bbadis.2012.06.008. [DOI] [PubMed] [Google Scholar]
- 38.Shirai Y, Okazaki Y, Inoue Y, Tamura Y, Yasuoka H, Takeuchi T, et al. Elevated levels of pentraxin 3 in systemic sclerosis: associations with vascular manifestations and defective vasculogenesis. Arthritis Rheumatol. 2015;67(2):498–507. doi: 10.1002/art.38953. [DOI] [PubMed] [Google Scholar]
- 39.Gilbane AJ, Derrett-Smith E, Trinder SL, Good RB, Pearce A, Denton CP, et al. Impaired bone morphogenetic protein receptor II signaling in a transforming growth factor-beta-dependent mouse model of pulmonary hypertension and in systemic sclerosis. Am J Respir Crit Care Med. 2015;191(6):665–677. doi: 10.1164/rccm.201408-1464OC. [DOI] [PubMed] [Google Scholar]
- 40.Palumbo K, Zerr P, Tomcik M, Vollath S, Dees C, Akhmetshina A, et al. The transcription factor JunD mediates transforming growth factor {beta}-induced fibroblast activation and fibrosis in systemic sclerosis. Ann Rheum Dis. 2011;70(7):1320–1326. doi: 10.1136/ard.2010.148296. [DOI] [PubMed] [Google Scholar]
- 41.Reich N, Tomcik M, Zerr P, Lang V, Dees C, Avouac J, et al. Jun N-terminal kinase as a potential molecular target for prevention and treatment of dermal fibrosis. Ann Rheum Dis. 2012;71(5):737–745. doi: 10.1136/annrheumdis-2011-200412. [DOI] [PubMed] [Google Scholar]
- 42.Sancho RM, Law BM, Harvey K. Mutations in the LRRK2 roc-COR tandem domain link Parkinson's disease to Wnt signalling pathways. Hum Mol Genet. 2009;18(20):3955–3968. doi: 10.1093/hmg/ddp337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Crago AM, Chmielecki J, Rosenberg M, O'Connor R, Byrne C, Wilder FG, et al. Near universal detection of alterations in CTNNB1 and Wnt pathway regulators in desmoid-type fibromatosis by whole-exome sequencing and genomic analysis. Genes Chromosomes Cancer. 2015;54(10):606–615. doi: 10.1002/gcc.22272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ghosh S, Taylor A, Chin M, Huang HR, Conery AR, Mertz JA, et al. Regulatory T cell modulation by CBP/EP300 Bromodomain inhibition. J Biol Chem. 2016;291(25):13014–13027. doi: 10.1074/jbc.M115.708560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Rao DA, Gurish MF, Marshall JL, Slowikowski K, Fonseka CY, Liu Y, et al. Pathologically expanded peripheral T helper cell subset drives B cells in rheumatoid arthritis. Nature. 2017;542(7639):110–114. doi: 10.1038/nature20810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bugatti S, Manzo A, Benaglio F, Klersy C, Vitolo B, Todoerti M, et al. Serum levels of CXCL13 are associated with ultrasonographic synovitis and predict power Doppler persistence in early rheumatoid arthritis treated with non-biological disease-modifying anti-rheumatic drugs. Arthritis Res Ther. 2012;14(1):R34. doi: 10.1186/ar3742. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. Degree of top 10 genes. (DOCX 14 kb)
Table S2. The modules of the PPI network. Four modules from the protein-protein interaction network satisfied the criteria of MCODE scores >6 and number of nodes >6. (DOCX 14 kb)