Abstract
Background:
The management of hypertrophic cardiomyopathy (HCM) patients requires the knowledge of risk factors associated with sudden cardiac death (SCD). SCD risk factors such as syncope and family history of SCD (FH-SCD) as well as family history of HCM (FH-HCM) are documented in electronic health records (EHRs) as clinical narratives. Automated extraction of risk factors from clinical narratives by natural language processing (NLP) may expedite management workflow of HCM patients. The aim of this study was to develop and deploy NLP algorithms for automated extraction of syncope, FH-SCD, and FH-HCM from clinical narratives.
Methods and Results:
We randomly selected 200 patients from the Mayo HCM registry for development (n=100) and testing (n=100) of NLP algorithms for extraction of syncope, FH-SCD as well as FH-HCM from clinical narratives of EHRs. The clinical reference standard was manually abstracted by 2 independent annotators. Performance of NLP algorithms was compared to aggregation and summarization of data entries in the HCM registry for syncope, FH-SCD, and FH-HCM. We also compared the NLP algorithms with billing codes for syncope as well as responses to patient survey questions for FH-SCD and FH-HCM. These analyses demonstrated NLP had superior sensitivity (0.96 vs 0.39, p < 0.001) and comparable specificity (0.90 vs 0.92, p = 0.74) and PPV (0.90 vs 0.83, p = 0.37) compared to billing codes for syncope. For FH-SCD, NLP outperformed survey responses for all parameters (sensitivity: 0.91 vs 0.59, p = 0.002; specificity: 0.98 vs 0.50, p < 0.001; PPV: 0.97 vs 0.38, p < 0.001). NLP also achieved superior sensitivity (0.95 vs 0.24, p < 0.001) with comparable specificity (0.95 vs 1.0, p-value not calculable) and positive predictive value (PPV) (0.92 vs 1.0, p = 0.09) compared to survey responses for FH-HCM.
Conclusions:
Automated extraction of syncope, FH-SCD and FH-HCM using NLP is feasible and has promise to increase efficiency of workflow for providers managing HCM patients.
Keywords: Hypertrophic cardiomyopathy, sudden cardiac death, natural language processing, electronic health records
1. Introduction
The widespread adoption of electronic health records (EHRs) has enabled more efficient approaches to health care delivery.1, 2 Digital information stored in EHRs may potentially be used to support decision-making by enabling automated retrieval, summarization and analysis of relevant data. Clinical narratives created by providers to describe patient encounters contain rich information.1 Natural language processing (NLP) extracts information from text and can be leveraged for unlocking information embedded in clinical narratives.1, 3, 4 Prior studies have demonstrated the successful use of NLP for extracting information about diabetes, heart failure, peripheral arterial disease, critical limb ischemia and inflammatory bowel disease.5–9
Hypertrophic cardiomyopathy (HCM) is the most common inherited cardiomyopathy and the leading cause of sudden cardiac death (SCD) in young adults.10–12 Established risk factors for SCD in HCM patients include syncope and family history of SCD (FH-SCD) which are routinely documented in clinical narratives.13 Family history of HCM (FH-HCM) may also be documented in clinical narratives and is a risk factor for the diagnosis of HCM.10 These risk factors contained within EHR narratives of HCM patients are potential targets for extraction by NLP.
Billing codes from EHRs have been previously used for identification of a patient cohort with HCM.14 However, there are limitations for use of billing codes for electronic phenotyping of risk factors. For example, syncope cannot be discriminated from pre-syncope by billing codes while FH-SCD or FH-HCM are not captured by billing codes. Moreover, to the best of our knowledge, NLP algorithms have not been previously described for extraction of syncope, FH-SCD, and FHHCM from clinical narratives of patients with HCM. In this study, we present NLP algorithms which extract these risk factors. We tested the hypothesis that NLP algorithms are superior to billing codes, responses to a patient survey, or data entries to a HCM registry for extraction of syncope, FH-SCD, and FH-HCM.
2. Methods
2.1. Study Design
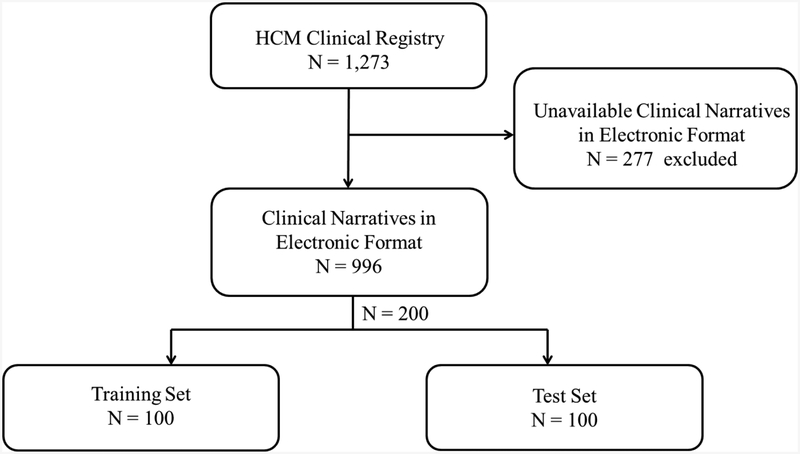
Study subjects were identified from 1,273 patients evaluated at the HCM clinic of the Mayo Clinic, Rochester, Minnesota from 1994 to 2016 who participated in a dedicated registry (Figure 1). Medical records of these individuals were manually reviewed and data entered into a database by registered nurses. Patients from the registry who did not have clinical narratives in electronic format were excluded (n = 277). In this cohort (n= 996 patients) 100 patients had known syncope during the study period. These subjects were matched to 100 HCM patients from the same cohort without syncope. Subsequently subjects with syncope and without syncope were randomly allocated to training and test sets. In each set of 100 subjects approximately half of subjects had a diagnosis of syncope and half did not. The study was approved by the Mayo Clinic Institutional Review Board.
Figure 1.
Study Design
Study subjects were identified from 1,273 patients who participated in a dedicated HCM registry. Patients from the registry who did not have clinical narratives in electronic format were excluded (n = 277). From this cohort (n = 996) 200 subjects were randomly selected and allocated to training and test sets of 100 subjects each. HCM = hypertrophic cardiomyopathy.
2.2. Risk Factor Selection and Definition
Two established risk factors for SCD in HCM patients including syncope and FH-SCD were selected for automated extraction. FH-HCM is a risk factor for the diagnosis of HCM and was also selected for automated extraction. Syncope was defined as any syncopal episode within 5 years prior to the index date for each subject and defined by criteria from practice guidelines.15 FH-SCD was defined as unexpected death of any family member, diagnosed at any age, at any time prior to the index date. FH-HCM was defined as family history of HCM in any family member, diagnosed at any age, at any time prior to the index date.
2.3. Automated Approaches for Risk Factor Extraction
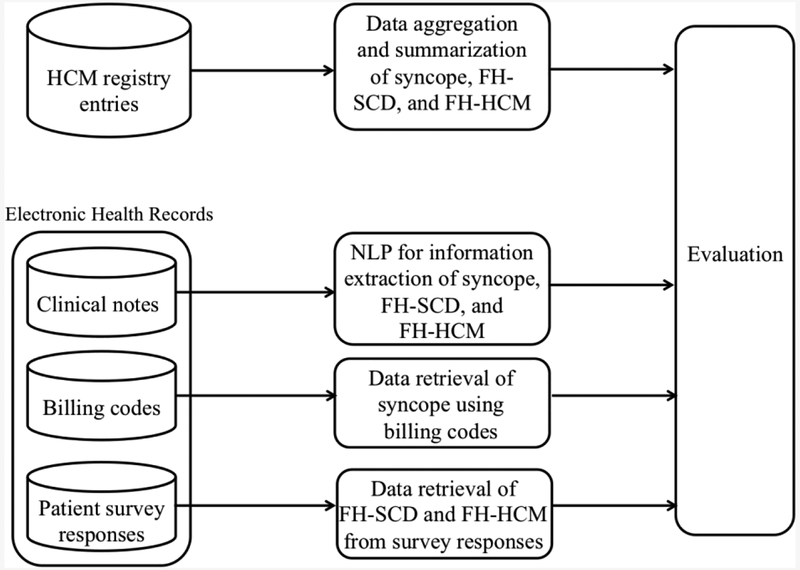
As summarized in Figure 2, the data entries of syncope, FH-SCD, and FH-HCM status in the database of the HCM registry were aggregated and summarized. NLP algorithms were used for information extraction of syncope, FH-SCD, and FH-HCM from clinical narratives. Billing codes were used for retrieval of syncope. Responses to patient survey questions were retrieved for FH-SCD and FH-HCM.
Figure 2.
Extraction of Risk Factors in HCM by Automated Approaches
Diverse automated technologies for risk factor extractions were used for each data type. FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history sudden of cardiac death; HCM = hypertrophic cardiomyopathy; NLP = natural language processing.
2.4. NLP Algorithms
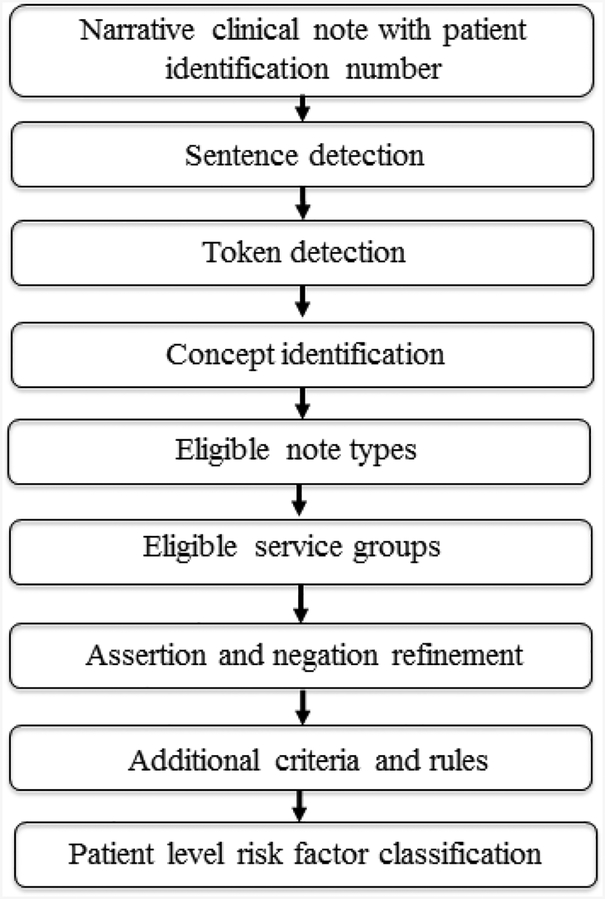
In the present study the NLP task addressed was information extraction. Three rule-based NLP algorithms were developed using Med Tagger,16–18 an open source NLP tool incorporating dictionary look-up, and regular expression pattern detection which has been used in various clinical NLP applications.19 Med Tagger has been adopted by Mayo Clinic enterprise-wide to deliver NLP services for clinical and translational research and healthcare delivery.20 To build NLP algorithms the terms (keywords) used for each concept were collected by 1) manual review of EHR clinical narratives and 2) use of Med Tagger for retrieval of lexical variations and synonyms from the Unified Medical Language System (UMLS) Meta thesaurus, a large biomedical thesaurus organized by concepts which links similar terms for the same concept from nearly 200 different vocabularies.21 In addition, assertion and negations generated by Med Tagger were refined using sentence patterns from clinical narratives. As shown in Figure 3, the Med Tagger Information Extraction (IE) tool processes clinical narratives containing unique identification numbers for each patient as input and generates risk factor status as output (Figure 3). The status of each risk factor is then displayed as “Yes” (present) or “No” (absent).
Figure 3.
Information Extraction by Med Tagger - IE Med Tagger - IE
processes clinical narratives containing unique identification numbers for each patient as input and generates risk factor status as output. The status of each risk factor is displayed as “Yes” (present) or “No” (absent). Sentences used for classification are displayed as evidence. Legend: IE = Information extraction.
The note types, service types and note sections where the sentences of interest were identified were also collected by manual chart review and used to build NLP algorithms (Table 1).
Table 1.
Associated Note Types, Note Sections and Service Groups for NLP Algorithms
| Risk factor | Note types | Note sections | Service groups |
|---|---|---|---|
| Syncope | Admission Progress Consult Subsequent-visit Supervisory |
Chief Complaint Subjective History of Present Illness Past Medical and Surgical History Impression, Report and Plan Procedure Information |
Cardiology Cardiology admission Cardiology floor practice Cardiology-invasive Catheterization laboratory Electrophysiology Cardiology Electrophysiology Consult Heart-Failure Heart-Rhythm Heart-Rhythm Device Community-Cardiology Interventional Cardiology Valve-Structural Cardiology |
| FH-SCD | Admission Progress Consult Supervisory |
History of Present Illness Family History Impression, Report and Plan |
|
| FH-HCM | Admission Progress Consult Subsequent-visit Supervisory |
Chief Complaint History of Present Illness Family History Impression, Report and Plan |
NLP: natural language processing; FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history of sudden cardiac death.
2.5. Structured Data Types for Syncope, FH-SCD, and FH-HCM
Structured data types evaluated included billing codes (International Classification of Diseases (ICD)-9 and ICD-10), and responses to a survey of patient provided information (PPI). Billing codes were retrieved from the institutional data warehouse. The content expert, a cardiologist and board-certified clinical informatician, identified the ICD-9 and ICD-10 billing codes for syncope. There were no billing codes available for FH-SCD or FH-HCM during the study period (see Table 4).
Table 4.
Billing Codes
| Risk factor | Diagnosis codes | Procedural Codes |
|---|---|---|
| Syncope | 780.2 - ICD9 R55 - ICD10 |
None |
| FH-SCD | None | None |
| FH-HCM | None | None |
Procedural Codes
Diagnosis codes
Risk factor
ICD = International Classification of Disease; FH-SCD = family history sudden cardiac death; FH-HCM = family history hypertrophic cardiomyopathy.
The investigators also searched responses to a survey of PPI, another source for structured data available in the EHR data warehouse. The PPI survey is routinely requested of every patient evaluated at Mayo Clinic. The keywords for SCD, HCM, and family history (see Table 2) were used to search the survey questions of the PPI for identification of patients with relatives with FH-SCD and FH-HCM. Reponses to the relevant patient survey questions identified using this approach were used for analysis. The relevant survey questions are listed in Table 5.
Table 2.
Key words used for the NLP algorithms
Table 2 lists the concepts and corresponding key words for the NLP algorithms.
| Concepts | Key words (case insensitive) |
|---|---|
| Syncope | syncope, syncopal spell, syncopal event, syncopal episode, syncopal attack, collapse, pass out, black out, faint, fainted, fainting spells, loss of consciousness |
| SCD | cardiac, sudden, unexplained death, scd |
| HCM | hypertrophic obstructive cardiomyopathy, hcm, hocm |
| Family History |
mom, mother, dad, father, parent, son, daughter, sister, brother, sibling, child, relative, cousin, aunt, uncle, grandfather, grandmother, grandparent, nephew, the family, maternal, family history, family member, half brother, half sister |
NLP: natural language processing; SCD = sudden cardiac death; HCM = hypertrophic cardiomyopathy.
Table 5.
Relevant Patient Survey Questions Identified by Keyword Search
| Potentially relevant questions |
FH-SCD | FH-HCM |
|---|---|---|
| Set #1 | “Which blood relatives have had heart disease?” “Relatives with heart disease?” “Have your relatives had heart attack?” “Have your relatives had other heart problems?” |
|
| Set #2 | “If mother deceased age at death” “If father deceased age at death” “Mother’s age at death” “Father’s age at death” “Relatives with unexplained death” “Sons or daughters - unknown cause of death” “Sisters or brothers - unknown cause of death” |
“Relatives with genetic disorder” “Which blood relatives have had a genetic disorder?” |
FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history of sudden cardiac death.
2.6. Manual Review of Clinical Narratives
Manual review of clinical narratives was performed for identification of each risk factor for each subject by two independent trained abstractors and classified the status of each risk factor as present or absent. A written guideline for abstraction was generated to standardize the process. In the event of disagreement between annotators, a third independent annotator (board-certified cardiologist) reviewed the EHR notes to resolve disagreements. The final status of each risk factor served as the clinical reference standard.
The guidelines for annotation contained all steps needed to find the relevant information in the clinical notes from the EHR, and criteria to classify each patient in the categories (yes, no, or not available). Additionally, the diagnostic criteria for each condition of interest (syncope, FH-SCD and FH-HCM) were also provided (see section 2.2 for risk factor definition).
2.7. Statistical Analysis
True-positives represented correctly identified risk factors whereas true-negatives represented a correctly identified absent risk factor. False-positives represented incorrectly identified present risk factors whereas false-negatives represented incorrectly identified absent risk factors. Sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated to evaluate the performance of each automated method for risk factor identification compared to the clinical reference standard of manual abstraction. Additionally, for each risk factor performance was compared between NLP algorithms with (a) structured data algorithms or (b) registry abstractions. McNemar’s test was used to compare sensitivity and specificity. Generalized score statistics were used to compare PPV and NPV. Analyses were performed using SAS version 9.4 (Cary, NC) and two-sided p ≤ 0.05 was considered statistically significant.
3. Results
200 HCM subjects with median age 61 years, 55% (n = 111) men, and 89% (n = 176) Caucasians were randomly allocated to the training (n=100) or test sets (n=100). There were 409 encounters (training set = 217 and test set = 192), 16,270 clinical narratives (training set = 7,464 and test = 8,806), and 1,201 billing codes or PPI responses (training set = 570 and test set = 631). Risk factor status verified by clinical reference standard (manual review of EHR clinical narratives) is summarized in Table 6. Approximately half of patients had syncope (random selection was in proportion to number of syncope cases identified by billing codes) and approximately one third had FH-SCD or FH-HCM (Table 6).
Table 6.
Risk Factor Status Verified by Clinical Reference Standard
| Clinical reference standard | Training Set n = 100 |
Test Set n = 100 |
|
|---|---|---|---|
| Syncope | Present | 52 | 49 |
| Absent | 48 | 51 | |
| FH-SCD | Present | 33 | 34 |
| Absent | 67 | 66 | |
| FH-HCM | Present | 39 | 38 |
| Absent | 61 | 62 | |
N = number of patients; FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history of sudden cardiac death.
For syncope, the information extracted from clinical narratives by NLP had superior sensitivity and NPV with comparable specificity and PPV compared to billing codes for both data sets (Table 7 and Table 8). For FH-SCD, NLP was compared to PPI survey responses and had superior performance for all parameters (Table 7 and Table 8). For FH-HCM, information extracted from clinical narratives by NLP had superior sensitivity and NPV with comparable specificity and PPV compared to PPI survey responses (Table 7 and Table 8). For all three risk factors NLP had superior sensitivity and NPV with comparable specificity and PPV compared to registry entries (Table 7 and Table 8). However, for syncope in the test set (Table 8) NLP performance was similar to registry entries. without HCM in the training set were correctly identified by survey response as not having HCM.
Table 7.
Performance of Diverse Approaches for Extraction of Risk Factors in Training Set (n = 100 patients)
| Risk factors |
Data Sources | Sensitivity 95% CI |
Specificity 95% CI |
PPV 95% CI |
NPV 95% CI |
|---|---|---|---|---|---|
| Syncope | Clinical Narratives |
0.98 (0.90,1.00) |
0.98 (0.89, 1.00) |
0.98 (0.90, 1.00) |
0.98 (0.89, 1.00) |
| Billing codes | 0.29 (0.17,0.43) p < 0.001 |
0.94 (0.83, 0.99) p = 0.32 |
0.83 (0.59,0.96) p = 0.11 |
0.55 (0.44, 0.66) p < 0.001 |
|
| Registry Entries |
0.87 (0.74,0.94) p = 0.03 |
0.90 (0.77, 0.97) p = 0.10 |
0.90 (0.78,0.97) p = 0.08 |
0.86 (0.73, 0.94) p = 0.02 |
|
| FH-SCD | Clinical Narratives |
0.82 (0.64–0.93) |
0.99 (0.92, 1.00) |
0.96 (0.82 – 1.00) |
0.92 (0.83, 0.97) |
| Survey Responses |
0.61 (0.42,0.77) p = 0.03 |
0.57 (0.44, 0.69) p < 0.001 |
0.41 (0.27,0.56) p < 0.001 |
0.75 (0.60, 0.86) p = 0.002 |
|
| Registry Entries |
0.42 (0.25,0.61) p < 0.001 |
0.99 (0.92, 1.00) p = 0.99 |
0.93 (0.68, 1.00) p = 0.67 |
0.78 (0.67, 0.86) p < 0.001 |
|
| FH-HCM | Clinical Narratives |
0.90 (0.76,0.97) |
1.00 (0.90, 1.00) |
1.00 (0.90,1.00) |
0.94 (0.95, 0.98) |
| Survey Responses |
0.23 (0.11, 0.39) P < 0.001 |
0.97 (0.89, 1.00) p = NA* |
0.82 (0.48,0.98) p = 0.14 |
0.66 (0.55, 0.96) p < 0.001 |
|
| Registry Entries |
0.72 (0.55,0.85) p = 0.03 |
0.98 (0.91, 1.00) p = NA* |
0.97 (0.82, 1.00) p = 0.31 |
0.85 (0.94, 0.92) p = 0.03 |
all p values are for comparison with NLP algorithms. CI = Confidence Interval; FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history of sudden cardiac death; N = number of patients; NA: not applicable; NPV = Negative predictive value; PPV = Positive predictive value;
= unable to calculate a p-value because all subjects
Table 8.
Performance of Diverse Approaches for Identification of Risk Factors in Test Set (n= 100 patients)
| Risk factors |
Data Sources | Sensitivity 95% CI |
Specificity 95% CI |
PPV 95% CI |
NPV 95% CI |
|---|---|---|---|---|---|
| Syncope | Clinical Narratives |
0.96 (0.86,1.00) |
0.90 (0.79, 0.97) |
0.90 (0.79,0.97) |
0.96 (0.86, 0.99) |
| Billing codes | 0.39 (0.25,0.54) p < 0.001 |
0.92 (0.81, 0.98) p = 0.74 |
0.83 (0.61,0.95) p = 0.37 |
0.61 (0.49, 0.72) p < 0.001 |
|
| Registry Entries |
0.88 (0.75,0.95) P = 0.16 |
0.86 (0.74, 0.94) p = 0.56 |
0.86 (0.73,0.94) p = 0.47 |
0.88 0.76, 0.95) p = 0.14 |
|
| FH-SCD | Clinical Narratives |
0.91 (0.76,0.98) |
0.98 (0.92, 1.00) |
0.97 (0.84,1.00) |
0.96 (0.88, 0.99) |
| Survey Responses |
0.59 (0.41,0.75) p = 0.002 |
0.50 (0.37, 0.63) p < 0.001 |
0.38 (0.25,0.52) p < 0.001 |
0.70 (0.55, 0.83) p < 0.001 |
|
| Registry Entries |
0.50 (0.32,0.68) p < 0.001 |
0.97 (0.89, 1.00) p = 0.56 |
0.89 (0.67,0.99) p = 0.33 |
0.79 (0.69, 0.87) p < 0.001 |
|
| FH-HCM | Clinical Narratives |
0.95 (0.82,0.99) |
0.95 (0.87, 0.99) |
0.92 (0.79,0.98) |
0.97 (0.98, 1.00) |
| Survey Responses |
0.24 (0.11,0.40) p < 0.001 |
1.00 (0.94, 1.00) p = NA* |
1.00 (0.66,1.00) p = 0.09 |
0.68 (0.58, 0.78) p < 0.001 |
|
| Registry Entries |
0.76 (0.60,0.89) p = 0.008 |
0.98 (0.91, 1.00) p = 0.32 |
0.97 (0.83,1.00) p = 0.41 |
0.87 (0.77, 0.94) p = 0.006 |
|
| FH-HCM | Clinical Narratives |
0.95 (0.82,0.99) |
0.95 (0.87, 0.99) |
0.92 (0.79,0.98) |
0.97 (0.98, 1.00) |
| Survey Responses |
0.24 (0.11,0.40) p < 0.001 |
1.00 (0.94, 1.00) p = NA* |
1.00 (0.66,1.00) p = 0.09 |
0.68 (0.58, 0.78) p < 0.001 |
|
| Registry Entries |
0.76 (0.60,0.89) p = 0.008 |
0.98 (0.91, 1.00) p = 0.32 |
0.97 (0.83,1.00) p = 0.41 |
0.87 (0.77, 0.94) p = 0.006 |
all p values are for comparison with NLP algorithms. CI = Confidence Interval; FH-HCM = family history of hypertrophic cardiomyopathy; FH-SCD = family history of sudden cardiac death; N = number of patients; NA: not applicable; NPV = Negative predictive value; PPV = Positive predictive value;
= unable to calculate a p-value because all subjects without HCM in the training set were correctly identified by survey response as not having HCM.
4. Discussion
This study made the novel observations that NLP algorithms deployed to clinical narratives had superior sensitivity for extraction of risk factors for SCD (syncope and FH-SCD) as well as FHHCM in HCM patients compared to data entries in an HCM registry, as well as billing codes and responses to a patient survey. We also observed that NLP algorithms had similar PPV compared to registry entries. Hence, the study herein is the first to report successful extraction of syncope, FH-SCD and FH-HCM in HCM patients from clinical narratives by NLP systems. The general lessons from these investigations were: 1) development and internal validation of approaches described requires a multidisciplinary team composed of clinicians, clinical informaticians, data scientists, NLP scientists as well as abstractors and 2) clear understanding of clinical workflow and written guidelines for review of medical records were necessary for development and validation of these methodologies.
In the present study we developed and internally validated automated algorithms for extraction of syncope, FH-SCD, and FH-HCM using a cohort of patients with the diagnosis of HCM confirmed by both review of EHRs by a physician expert and imaging by echocardiography or cardiac magnetic resonance. We then used clinical narratives from EHRs of these confirmed cases for extraction of concepts, terminologies and patterns of risk factors and to validate performance of the NLP algorithms. Clinical narratives had been generated from encounters with patients in routine clinical practice. The NLP algorithms identified relevant terms and sentence patterns from different note types, sections and service groups where information of interest was found by manual chart review, and accounted for the superior performance of NLP.
Our study demonstrated a potential limitation of billing codes compared to NLP for extraction of syncope. Clinical practice guidelines define syncope as “a symptom that presents with an abrupt, transient, complete loss of consciousness, associated with inability to maintain postural tone, with rapid and spontaneous recovery.”15 Hence “syncope and collapse” is used to name both the ICD-10 billing code R55 and ICD-9 code 780–2 (the ICD-9 code 780–2 converts directly to ICD-10 R55). These codes are used for syncope, regardless of the mechanism. In most patients, the diagnosis of syncope is made by medical history and most patients are not monitored during the event and it is not possible to determine the etiology for syncope. Moreover, the same billing codes (ICD-9 780–2 and ICD-10 R55) are used for patients with pre-syncope. However, syncope is a known risk factor for SCD in HCM patients while pre-syncope is not a risk factor for SCD in HCM. Hence the performance of billing code algorithms for syncope was limited while the NLP algorithm discriminated patients with syncope from those with pre-syncope yielding superior performance.
Data entered to the HCM registry was prospectively collected by registered nurses by review of the EHR with manual entry of variables of interest into the dedicated digital dataset. Compared to NLP algorithms for information extraction from clinical narratives the aggregation and summarization of registry entries had similar PPV and specificity but inferior sensitivity (except for syncope in the test set). This may be because the NLP algorithms were deployed to all relevant EHR clinical narratives of each patient using uniform rules22 whereas the nurses reviewed the most recent clinical narratives without the standardized guideline. The use of an automated approach using NLP algorithms may assist and complement human abstraction which may reduce effort required for retrieval of data elements to populate registries.22 The use of an automated approach may also increase efficiency of clinical review for risk factors routinely performed by providers prior to patient encounters. Hence, NLP algorithms may rapidly extract clinically relevant information from clinical narratives thereby optimizing clinical workflow and realizing the vision of the “Roadmap for Innovation” endorsed by the American College of Cardiology.23
The investigators plan to apply the NLP algorithms to clinical narratives of patients evaluated in other settings including primary care and emergency departments for extraction of syncope, FH-SCD, and FH-HCM in the general population. The hypothesis is that if applied to a general electronic clinical narrative corpus the NLP algorithms we reported herein will have excellent performance for identification of individuals who may need referral for further evaluation and may benefit from imaging tests to evaluate for HCM. NLP algorithms for extraction of family history could also be modified for identification of patients with family history of other rare inherited heart conditions (e.g. long QT syndrome) which require identification of family members who may be at increased risk for sudden and unexpected death. Importantly, machine learning combined with NLP has been previously used for automated extraction of risk factors for surgical site infections24 suggesting a similar approach may also be used for identification of rare instances of risk factors for HCM and SCD in large document corpora.
For FH-SCD, there are no available billing codes. The approach to extract structured data for this risk factor used responses to a patient survey containing screening questions offered to all patients evaluated at the Mayo Clinic. However, the performance of questionnaires for identification of risk factors was limited in part because the content of standardized questions were not specifically designed for extraction of FH-SCD in HCM patients. Survey questions were designed only for identification of family history of heart disease. Additionally, responses to these survey questions were provided by patients, without contribution of providers or medical record information. An ICD-10 diagnostic code for FH-HCM became available for billing in October, 2017. The dataset used in the present study was built prior to availability of this code. Future studies may evaluate the performance of the new ICD-10 code for FH-HCM for extraction of this risk factor compared to NLP algorithms.
4.1. Limitations
For this study, NLP algorithms were developed and tested in a single tertiary medical center. In the future it will be important to evaluate these automated approaches in other centers to demonstrate portability. In mitigation, it should be noted that NLP systems are potentially applicable to any practice which uses EHRs and are also agnostic to vendor. However, robust computing and informatics infrastructures are necessary to deploy this technology to patient care.20
5. Conclusions
Our study demonstrates that automated extraction of syncope, FH-SCD, and FH-HCM in HCM patients from clinical narratives by NLP algorithms is feasible. The algorithms developed can be translated to clinical decision support systems to increase efficiency of workflow for providers managing HCM patients and contribute to improved quality of care.
Table 3.
Rules and examples of text spans identified in clinical narratives
Table 3 lists the rules for the NLP algorithms with examples of text spans identified by these rules. At least one key word of the concept of interest (see Table 2) was required by each rule.
| Risk factors | Rules and examples of text spans | |
|---|---|---|
| Syncope | Rule | Patient (experiencer) + any token {0,n}+ key word for syncope |
| Example | She sat on the bed and then had a brief loss of consciousness witnessed by her husband. | |
| FH-SCD | Rule | Key word for family history + any token {0,15} + died + any token {0,5} + key word for SCD |
| Example | A paternal grandfather
died
suddenly and unexpectedly at 50 years of age. |
|
| FH-HCM | Rule | Key word for family history + any token {0,15} + has/have/diagnosed with/of/with + any token {0,15} + key word for HCM |
| Example | The main interval change since his last visit was that his father was recently diagnosed with hypertrophic cardiomyopathy based on echocardiographic imaging. | |
HCM = hypertrophic cardiomyopathy; FH-HCM = family history hypertrophic cardiomyopathy; FH-SCD = family history sudden cardiac death; SCD = sudden cardiac death.
Summary table.
What was already known on the topic:
Risk factors for sudden cardiac death (syncope and FH-SCD) and FH-HCM are documented in clinical narratives. These risk factors contained within EHR narratives of HCM patients may be targets for extraction by NLP. No studies have previously described automated algorithms which extract syncope, FH-SCD and FH-HCM, from clinical narratives.
What this study added to our knowledge:
This study extracted SCD risk factors (syncope and FH-SCD) and FH-HCM from EHRs by automated approaches including NLP.
NLP algorithms had superior sensitivity for extraction of syncope, FH-SCD, and FHHCM information compared to aggregation of data entries in an HCM registry as well as retrieval of billing codes and responses to a patient survey.
NLP may support efficient workflow for providers managing HCM patients by automated extraction of extraction of syncope, FH-SCD, and FH-HCM from clinical narratives to enable prompt and timely review by providers at the point-of-care.
Acknowledgments:
The authors thank Corina Moreno for data abstraction and Rebecca Olson for secretarial support.
Sources of Funding: This study supported by the National Heart, Lung and Blood Institute of the National Institutes of Health (K01HL124045) and by a Mayo Clinic K2R award. The NLP system was developed through NIGMS award R01GM102282. The content is solely the responsibility of the authors and does not necessarily represent official views of the National Institutes of Health.
List of Abbreviations
- EHR
Electronic Health Record
- FH
Family History
- HCM
Hypertrophic Cardiomyopathy
- ICD
International Classification of Diseases
- NLP
Natural Language Processing
- NPV
Negative Predictive Value
- PPI
Patient Provided Information
- PPV
Positive Predictive Value
- SCD
Sudden Cardiac Death
- UMLS
Unified Medical Language System
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Jensen PB, Jensen LJ and Brunak S. Mining electronic health records: towards better research applications and clinical care. Nat Rev Genet. 2012;13:395–405. [DOI] [PubMed] [Google Scholar]
- 2.Maddox TM, Albert NM, Borden WB, Curtis LH, Ferguson TB Jr., Kao DP, Marcus GM, Peterson ED, Redberg R, Rumsfeld JS, Shah ND, Tcheng JE, American Heart Association Council on Quality of C, Outcomes R, Council on Cardiovascular Disease in the Y, Council on Clinical C, Council on Functional G, Translational B and Stroke C. The Learning Healthcare System and Cardiovascular Care: A Scientific Statement From the American Heart Association. Circulation. 2017;135:e826–e857. [DOI] [PubMed] [Google Scholar]
- 3.Rosenbloom ST, Denny JC, Xu H, Lorenzi N, Stead WW and Johnson KB. Data from clinical notes: a perspective on the tension between structure and flexible documentation. J Am Med Inform Assoc. 2011;18:181–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Demner-Fushman D, Chapman WW and McDonald CJ. What can natural language processing do for clinical decision support? J Biomed Inform. 2009;42:760–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Afzal N, Mallipeddi VP, Sohn S, Liu H, Chaudhry R, Scott CG, Kullo IJ and Arruda-Olson AM. Natural language processing of clinical notes for identification of critical limb ischemia. Int J Med Inform. 2018;111:83–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Afzal N, Sohn S, Abram S, Scott CG, Chaudhry R, Liu H, Kullo IJ and Arruda-Olson AM. Mining peripheral arterial disease cases from narrative clinical notes using natural language processing. J Vasc Surg. 2017;65:1753–1761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zheng L, Wang Y, Hao S, Shin AY, Jin B, Ngo AD, Jackson-Browne MS, Feller DJ, Fu T, Zhang K, Zhou X, Zhu C, Dai D, Yu Y, Zheng G, Li YM, McElhinney DB, Culver DS, Alfreds ST, Stearns F, Sylvester KG, Widen E and Ling XB. Web-based Real-Time Case Finding for the Population Health Management of Patients With Diabetes Mellitus: A Prospective Validation of the Natural Language Processing-Based Algorithm With Statewide Electronic Medical Records. JMIR Med Inform. 2016;4:e37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wang Y, Luo J, Hao S, Xu H, Shin AY, Jin B, Liu R, Deng X, Wang L, Zheng L, Zhao Y, Zhu C, Hu Z, Fu C, Hao Y, Zhao Y, Jiang Y, Dai D, Culver DS, Alfreds ST, Todd R, Stearns F, Sylvester KG, Widen E and Ling XB. NLP based congestive heart failure case finding: A prospective analysis on statewide electronic medical records. Int J Med Inform. 2015;84:1039–47. [DOI] [PubMed] [Google Scholar]
- 9.Ananthakrishnan AN, Cai T, Savova G, Cheng SC, Chen P, Perez RG, Gainer VS, Murphy SN, Szolovits P, Xia Z, Shaw S, Churchill S, Karlson EW, Kohane I, Plenge RM and Liao KP. Improving case definition of Crohn’s disease and ulcerative colitis in electronic medical records using natural language processing: a novel informatics approach. Inflamm Bowel Dis. 2013;19:1411–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gersh BJ, Maron BJ, Bonow RO, Dearani JA, Fifer MA, Link MS, Naidu SS, Nishimura RA, Ommen SR, Rakowski H, Seidman CE, Towbin JA, Udelson JE, Yancy CWand American College of Cardiology Foundation/American Heart Association Task Force on Practice G. 2011 ACCF/AHA Guideline for the Diagnosis and Treatment of Hypertrophic Cardiomyopathy: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. Developed in collaboration with the American Association for Thoracic Surgery, American Society of Echocardiography, American Society of Nuclear Cardiology, Heart Failure Society of America, Heart Rhythm Society, Society for Cardiovascular Angiography and Interventions, and Society of Thoracic Surgeons. J Am Coll Cardiol. 2011;58:e212–60. [DOI] [PubMed] [Google Scholar]
- 11.O’Mahony C, Jichi F, Pavlou M, Monserrat L, Anastasakis A, Rapezzi C, Biagini E, Gimeno JR, Limongelli G, McKenna WJ, Omar RZ, Elliott PM and Hypertrophic Cardiomyopathy Outcomes I. A novel clinical risk prediction model for sudden cardiac death in hypertrophic cardiomyopathy (HCM risk-SCD). Eur Heart J. 2014;35:2010–20. [DOI] [PubMed] [Google Scholar]
- 12.O’Mahony C, Jichi F, Ommen SR, Christiaans I, Arbustini E, Garcia-Pavia P, Cecchi F, Olivotto I, Kitaoka H, Gotsman I, Carr-White G, Mogensen J, Antoniades L, Mohiddin SA, Maurer MS, Tang HC, Geske JB, Siontis KC, Mahmoud KD, Vermeer A, Wilde A, Favalli V, Guttmann OP, Gallego-Delgado M, Dominguez F, Tanini I, Kubo T, Keren A, Bueser T, Waters S, Issa IF, Malcolmson J, Burns T, Sekhri N, Hoeger CW, Omar RZ and Elliott PM. International External Validation Study of the 2014 European Society of Cardiology Guidelines on Sudden Cardiac Death Prevention in Hypertrophic Cardiomyopathy (EVIDENCE-HCM). Circulation. 2018;137:1015–1023. [DOI] [PubMed] [Google Scholar]
- 13.Ranthe MF, Carstensen L, Oyen N, Jensen MK, Axelsson A, Wohlfahrt J, Melbye M, Bundgaard H and Boyd HA. Risk of Cardiomyopathy in Younger Persons With a Family History of Death from Cardiomyopathy: A Nationwide Family Study in a Cohort of 3.9 Million Persons. Circulation. 2015;132:1013–9. [DOI] [PubMed] [Google Scholar]
- 14.Pujades-Rodriguez M, Guttmann OP, Gonzalez-Izquierdo A, Duyx B, O’Mahony C, Elliott P and Hemingway H. Identifying unmet clinical need in hypertrophic cardiomyopathy using national electronic health records. PLoS One. 2018;13:e0191214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Shen WK, Sheldon RS, Benditt DG, Cohen MI, Forman DE, Goldberger ZD, Grubb BP, Hamdan MH, Krahn AD, Link MS, Olshansky B, Raj SR, Sandhu RK, Sorajja D, Sun BC and Yancy CW. 2017 ACC/AHA/HRS Guideline for the Evaluation and Management of Patients With Syncope: Executive Summary: A Report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines and the Heart Rhythm Society. J Am Coll Cardiol. 2017;70:620–663. [DOI] [PubMed] [Google Scholar]
- 16.Liu H, Bielinski SJ, Sohn S, Murphy S, Wagholikar KB, Jonnalagadda SR, Ravikumar K, Wu ST, Kullo IJ and Chute CG. An Information Extraction Framework for Cohort Identification Using Electronic Health Records. AMIA Jt Summits Transl Sci Proc. 2013:149–153. [PMC free article] [PubMed] [Google Scholar]
- 17.Wang Y, Wang L, Rastegar-Mojarad M, Moon S, Shen F, Afzal N, Liu S, Zeng Y, Mehrabi S, Sohn S and Liu H. Clinical information extraction applications: A literature review. J Biomed Inform. 2018;77:34–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Torii M, Wagholikar K and Liu H. Using machine learning for concept extraction on clinical documents from multiple data sources. J Am Med Inform Assoc. 2011;18:580–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wu S, Liu S, Sohn S, Moon S, Wi CI, Juhn Y and Liu H. Modeling asynchronous event sequences with RNNs. J Biomed Inform. 2018;83:167–177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kaggal VC, Elayavilli RK, Mehrabi S, Pankratz JJ, Sohn S, Wang Y, Li D, Rastegar MM, Murphy SP, Ross JL, Chaudhry R, Buntrock JD and Liu H. Toward a Learning Health-care System - Knowledge Delivery at the Point of Care Empowered by Big Data and NLP. Biomed Inform Insights. 2016;8:13–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pivovarov R and Elhadad N. Automated methods for the summarization of electronic health records. Journal of the American Medical Informatics Association. 2015;22:938–947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Rastegar-Mojarad M, Sohn S, Wang L, Shen F, Bleeker TC, Cliby WA and Liu H. Need of informatics in designing interoperable clinical registries. Int J Med Inform. 2017;108:78–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bhavnani SP, Parakh K, Atreja A, Druz R, Graham GN, Hayek SS, Krumholz HM, Maddox TM, Majmudar MD, Rumsfeld JS and Shah BR. 2017 Roadmap for Innovation-ACC Health Policy Statement on Healthcare Transformation in the Era of Digital Health, Big Data, and Precision Health: A Report of the American College of Cardiology Task Force on Health Policy Statements and Systems of Care. J Am Coll Cardiol. 2017;70:2696–2718. [DOI] [PubMed] [Google Scholar]
- 24.Sohn S, Larson DW, Habermann EB, Naessens JM, Alabbad JY and Liu H. Detection of clinically important colorectal surgical site infection using Bayesian network. J Surg Res. 2017;209:168–173. [DOI] [PMC free article] [PubMed] [Google Scholar]