Abstract
Hypertrophic cardiomyopathy (HCM) has been considered a heterogeneous cardiac disease ascribed solely to single sarcomere gene mutations. Limitations of this hypothesis suggest that sarcomere mutations alone do not adequately explain all HCM clinical and pathobiologic features. Disease-causing sarcomere mutations are absent in ~70% of patients with established disease, and sarcomere gene carriers can live to advanced ages without developing HCM. Some features of HCM are also inconsistent with the single sarcomere gene hypothesis, such as regional left ventricular (LV) hypertrophy and myocardial fibrosis, as well as structurally abnormal elongated mitral valve leaflets and remodeled intramural coronary arterioles, which involve tissue types that do not express cardiomyocyte sarcomere proteins. It is timely to expand the HCM research focus beyond a single molecular event toward more inclusive models to explain this disease in its entirety. We chart paths forward addressing this knowledge gap utilizing novel analytical approaches, particularly network medicine, to unravel the pathobiologic complexity of HCM.
Keywords: Hypertrophic cardiomyopathy, network medicine, genetics
CONDENSED ABSTRACT:
For 3 decades, HCM has been regarded as a genetic heart disease explained solely by the single sarcomere gene mutation hypothesis. However, this is contestable given that pathogenic sarcomere mutations are not identified in 70% of HCM patients, and even when present do not explain all disease characteristics. For example, cardiomyocyte sarcomere proteins are not expressed in elongated mitral valve leaflets or remodeled intramural coronary arterioles.Although sarcomeres are expressed throughout the left ventricle (LV), regional LVH is common with most of the LV wall free of thickening. We suggest novel research initiatives relying on network medicine strategies to clarify pathobiology and move HCM forward.
Introduction
Hypertrophic cardiomyopathy (HCM) is viewed as the most common familial cardiovascular disorder, often inherited in an autosomal dominant pattern and disseminated worldwide with a prevalence of about 1:500 – 1:200 in the general population (1). Over the last 35 years, more than 2,000 variants in at least 11 genes encoding proteins of the cardiac sarcomere or associated structures have been reported in HCM patients (2). For this reason, HCM is often cited as a genetic disease of contractile proteins, and, therefore, as a prime example of how DNA sequencing can potentially clarify the basis of human disease. However, while the HCM genetic era is well into its third decade, critical knowledge gaps remain concerning the contribution of sarcomere gene mutations to the overall pathobiological mechanisms and clinical course of this complex disease.
In this commentary, we consider the limitations inherent in viewing HCM through the narrow prism of a single molecular (genetic) event. We also propose a ‘call to action’ for the HCM research community to provide fresh insights addressing this issue, utilizing contemporary scientific methodologies in biological complexity with an emphasis on network medicine (Central Illustration).
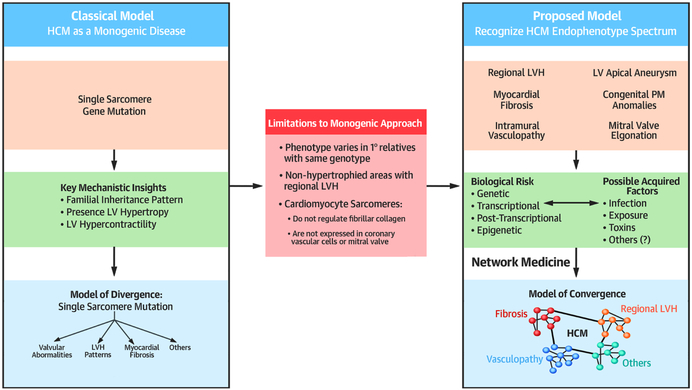
Central Illustration: Contemporizing the approach to understanding HCM pathobiology.
Left panel: The classical model of hypertrophic cardiomyopathy (HCM). In this monogenic approach, a single molecular event involving a mutant sarcomere gene promotes abnormalities in different cardiac structures. Center panel: However, limitations to this paradigm raise the speculation that parallel or alternative mechanisms to a single sarcomere gene mutation may be important for understanding HCM. (Right panel) We propose a path forward that focuses initially on the totality of HCM endophenotypes, explores acquired (non-genetic) determinants of disease expression, and leverages novel analytical methods, particularly network medicine, in reconsidering the pathobiological basis of HCM. This provides a flexible and integrative framework in which each HCM endophenotype is regulated by a unique network focusing on the functional or physical relationship between biological components (e.g., protein-protein interactions. In this figure a circle represents each protein, or network node, and each functional or physical relationship between proteins is represented by a line, or network edge). Protein-protein interactions (two or more connected circles in this figure) may be influenced by genetic variants or environmental cues among other acquired determinants, and may cross-talk between endophenotype networks. Ultimately, this results in a flexible model emphasizing endophenotype convergence for understanding the heterogeneity of HCM encountered in clinical practice. LV, left ventricle; LVH, left ventricular hypertrophy; PM, papillary muscle.
Historical Context
Prior to 1980, understanding of HCM could be considered rudimentary by today’s standards. For example, only at that time did a unifying morphological definition and a standardized name for this condition emerge (3). A familial autosomal dominant inheritance pattern was evident early in selected, large, multigenerational kindreds used in linkage analysis studies (4). Nevertheless, controversy has persisted regarding the existence of non-genetic vs. genetic forms (5), and even whether HCM represents a single disease or several similar and loosely affiliated clinical entities. For example, some previous observers even considered obstructive and nonobstructive forms of HCM to be separate conditions with distinct etiologies.
Greater availability of DNA sequencing in the early 1990s sparked widespread interest in the genetic basis of many cardiovascular diseases (2,6,7). In particular, the discovery of genetic variants was proposed as the critical link required to unravel the genotype-endophenotype-clinical phenotype relationship (Figure 1A). In this proposed model, a gain- or loss-of-function mutation (genotype) underlies the development of a fundamental disease feature (endophenotype, such as hypertrophy), which results in a disease that is diagnosed clinically (phenotype).
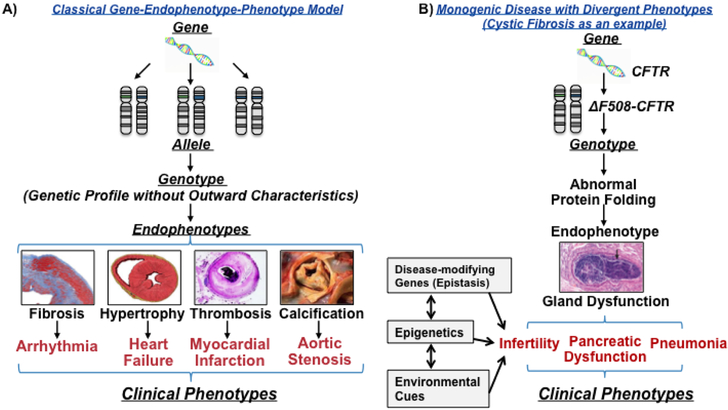
Figure 1. Classical model used to describe clinical disorders based on genetics.
(A) A loss- or gain-of-function gene mutation corresponds to an allele, or abnormal trait, which collectively describe a disease-predisposing genotype. A pathogenic gene variant, in turn, results in the development of an endophenotype, which serves as the fundamental basis for a clinical disorder. This classical model provides a direct link between genotype and clinical disease, and has been proposed to explain many diseases associated with rare genetic variants. (B) For example, in cystic fibrosis a deletion of three nucleotides spanning positions 507 and 508 of the transmembrane conductance regulator (CFTR) gene results in a loss of phenylalanine. In >90% of patients, this functions as the molecular event that generates exocrine gland dysfunction (endophenotype), which, in turn, is responsible for the totality of clinical findings in affected patients. However, truly monogenic diseases are exceedingly rare, since most disease, particularly HCM, involve a combination of endophenotypes that converge to produce the clinical phenotype. Images reproduced with permission from: fibrosis, Creative Commons License ShareAll at https://www.neurologyadvisor.com/epilepsy/cardiac-pathology-in-sudep-compared-with-sudden-arrhythmic-or-traumatic-deaths/article/781382/; hypertrophy, Creative Commons License ShareAlike at https://en.wikipedia.org/wiki/Left_ventricular_hypertrophy; thrombosis, Yale Medical Studies Library at http://medcell.med.yale.edu/histology/blood_vessels_lab/coronary_thrombosis.php; calcification, Sage Publications through Rightslink from Morsy, Y., et al. Int J of Artif Organs, 2004; 27:445-451.; Gland dysfunction, Massachusetts Medical Society from Stoltz, D.A., et al. Origins of Cystic Fibrosis Lung Disease. N Engl J Med 372(4): 1574-1575.
Early on in HCM, it was proposed that variants in genes coding for sarcomere proteins alone accounted for otherwise unexplained left ventricular hypertrophy (LVH) in the absence of chamber dilation, which was also considered in-and-of-itself to represent the full clinical phenotype. In support of this conclusion were pedigree studies with echocardiographic assessment demonstrating that sarcomere mutations track with LVH. Thereafter, commercial genetic testing became a vehicle for next generation (cascade) family screening (2,7). Based on this conventional paradigm, relatives of HCM patients who themselves do not have LVH or harbor HCM gene variants have been considered probably free of risk for the disease.
Single Gene Hypothesis
Thus, the “single gene hypothesis” for HCM emerged almost 3 decades ago in which it was assumed that a single genetic variant in (one of several) cardiomyocyte sarcomere proteins accounts for all aspects of the pathophenotype. HCM has thereby become widely regarded as a monogenic cardiac disease with myriad gene mutations identified in clinically affected family members. This recognition has had two salient effects on the clinical community’s approach to HCM: First, it provided the opportunity to consolidate what is otherwise a clinically heterogeneous disease into a single entity based on the genetic substrate. Second, a paradigm was established that continues today in which sarcomere mutations alone (largely missense and nonsynonymous) are assumed to be primarily, if not exclusively, disease-causing. Importantly, this paradigm continues to garner widespread acceptance despite its inability to explain certain mechanistic aspects of HCM, such as the interplay between dysregulated cardiac energetics and LV hypercontractility, and interstitial and perivascular fibrosis (6).
Overall, however, when considering the entire clinical phenotype the mechanisms by which sarcomere gene mutations cause HCM remain incompletely defined (7). In addition, contemporary data leveraging access to large genetic repositories of normal populations, unavailable in the first era of HCM genetics, demonstrate that some mutations initially considered causative and pathogenic, are, in fact, not observed at a different frequency from that in race-matched non-disease controls (8). Therefore, with the advantage of reflecting on decades of accumulated clinical, pathologic, and genetic observations in HCM, it now seems reasonable to ask the following questions: (1) Should we continue to consider HCM a monogenic disease? (2) Are alternative (genetic and non-genetic) mechanisms important determinants of the HCM phenotype and its natural history?
Evidence Inconsistent with the Single Gene Hypothesis: HCM Structural Features
Classically described monogenic diseases are defined by close mechanistic coupling between the functional consequences of a single gene mutation and specific disease-related pathobiology and clinical presentations. For example, in cystic fibrosis, a mutation involving the loss of 3 nucleotides in the CFTR gene (F508del-CFTR), which is the most common principal genetic event responsible for this disease, causes exocrine duct dysfunction in several organs. Yet, while this mutation (or one of nearly 2000 other primary driver mutations) is required for the development of exocrine gland dysfunction (endophenotype) and, thus the disease to be manifest, its clinical phenotypes vary considerably among patients (divergent phenotypes). This phenomenon can best be explained by genetic context (disease-modifying genes, epistasis) and environmental exposures (Figure 1B) (9).
HCM is also a heterogeneous disease, but comprising a collection of diverse disease features (endophenotypes) confined to the heart that converge to produce the clinically recognized disease phenotype (10). Importantly, each of these disease features can, and often do, coexist within an individual patient. Notwithstanding its narrower organ involvement, as with cystic fibrosis, it is difficult scientifically and intuitively to explain such a complex pathophenotype by a single missense point mutation (the most common type of variant encountered in HCM). Owing to the oversimplified notion that HCM is a more homogeneous phenotype, disease-modifying genes have not been rigorously pursued (11,12).
LV myocardium.
On the basis of a multitude of imaging and pathologic studies over almost 50 years, the morphology of the HCM heart has been interrogated and defined in great detail (10). In nearly all HCM patients, LVH is the sine qua non disease marker and the basis of clinical diagnosis. Nonetheless, the anatomical distribution of cardiac hypertrophy is quite variable in HCM. For example, thickening of the LV wall is often segmental and regionally confined to very small areas and notably with most of the wall non-hypertrophied and virtually normal in thickness and appearance.
Additionally, characteristic are sharp transitions in LV wall thickness between adjacent areas, with regions of hypertrophy separated by regions of normal wall thickness (non-contiguous pattern), and only rarely is hypertrophy of the right ventricular wall observed. Indeed, the diversity in LVH pattern is so common in HCM, that it alone is difficult to reconcile with the single gene hypothesis, given that sarcomere mutations are expressed in cardiomyocytes throughout the left and right ventricular walls (13). Furthermore, dissimilar LVH patterns are typically observed in closely related family members or between families, despite expression of identical HCM genotypes.
Mitral valve leaflets.
Notably, structurally abnormal and elongated mitral valve leaflets are typically observed in a substantial proportion of HCM patients, and are responsible for dynamic outflow obstruction. Mitral leaflet enlargement represents a primary component of HCM disease presentation, largely unrelated to age, degree of LV hypertrophy, or other disease variables (14). The elongated anterior and/or posterior mitral leaflets are comprised of endothelial, interstitial, and fibroblast cell types, as well as collagen and connective tissue elements, but not cardiomyocytes (15).
Intramural coronary arteriopathy.
Structurally remodeled abnormal intramural coronary arterioles are distributed throughout hypertrophied and non-hypertrophied areas of the LV, either within or in close proximity to areas of replacement fibrosis (16). This primary vascular abnormality is characterized by luminal narrowing and prominent thickening of the intima and media, as well as associated perivascular fibrosis. Abnormal intramural arterioles are likely responsible for microvascular ischemia, which, in turn, may contribute to the chest pain reported frequently by HCM patients (10,16).
Myocardial Fibrosis.
Deposition of collagen is a critical component of the heart in HCM. This process includes increased interstitial fibrosis that may expand the LV matrix compartment to as much as 8-fold larger than normal (17). In addition, replacement scar formation, as repair following cardiomyocyte cell death, is predictive of both sudden death and heart failure progression (18).
Congenital malformations.
Several morphologic abnormalities identified in the LV of HCM patients are congenital, not contingent on individual patient genotype, and unrelated to sarcomere protein mutations. For example, muscular mid-cavity obstruction can be caused by direct papillary muscle insertion into the mitral valve in the absence of chordae tendineae owing to embryonic arrest (2). Other examples of likely congenital anomalies relevant to LV outflow tract obstruction include accessory apical-basal muscle bundles, displaced papillary muscles, abnormal chordal connections between the ventricular septum and LV free wall or the papillary muscles, as well as myocardial bridging of the left anterior descending coronary artery.
LV apical aneurysms.
Localized apical aneurysms associated with regional scarring, which often develop later in life, are now recognized as part of the HCM disease spectrum, representing a novel risk factor for sudden death (19). Development of this complex morphologic abnormality would not be expected to result solely from mutant genes encoding sarcomere proteins alone.
Animal experiments.
Transgenic experimental models with myofilament gene mutations have provided some important insights into the translational relevance of sarcomere mutations to HCM pathobiology (7, 10). The majority of data in vivo is from murine models, which largely fail to recapitulate most HCM endophenotypes and clinical features, including markedly increased LV mass, structurally abnormal mitral valves producing outflow tract obstruction, remodeled intramural coronary arteries, or replacement scarring. Results from drosophila and zebra fish studies have characterized the role of troponin-T function in myocardial relaxation and regulatory mechanisms controlling myofilament thickening, respectively (20,21).
Evidence Inconsistent with the Single Gene Hypothesis: HCM Clinical Genetics
Mutation-negative HCM patients.
Only about 20-30% of probands with an established clinical diagnosis of HCM demonstrate a sarcomere mutation classified as pathogenic or believed to be disease-causing (10, 22). Consequently, the vast majority of HCM patients (more than two-thirds) are without a confirmed genetic etiology for their disease according to the standard monogenic paradigm for this pathophenotype. Thus, an important challenge in this field remains for defining the cause of HCM in the majority of affected patients for whom known genetic risk is absent.
Indeed, isolated/sporadic cases, defined by a proband with HCM in the absence of both an HCM mutation and first degree relative with HCM (or sarcomere mutation), is encountered commonly in clinical practice, and accounted for 40% of cases in one study (5). Given that DNA sequencing of the cardiac sarcomere genes and associated structures is essentially complete, it seems somewhat unlikely at this juncture that novel contractile proteins or other gene variants will be identified to provide a monogenic etiology to explain this “nonfamilial HCM” subgroup (5) or, more broadly, genotype-negative HCM patients (10). In a recent study utilizing whole genome sequencing, only one-fifth of families without a known sarcomere mutation were found to have putative pathologic variants in noncoding regions, or in genes not previously recognized as associated with the phenotype, including one mitochondrial genome variant (23). These observations in nonfamilial and familial forms of HCM provide the most compelling evidence in favor of non-sarcomere gene mutations, non-genetic (environmental or epigenetic) contributions, or both in the pathogenesis of HCM.
Incomplete Penetrance.
Identification of pathogenic mutations in family members who do not meet clinical diagnostic criteria for HCM (i.e., LVH) is common. In the largest available series of 203 mainly adult genotype positive-phenotype negative relatives (e.g., gene carriers), none experienced cardiac events or symptoms over the follow-up period, with a substantial proportion achieving an advanced age of up to 70 years (20% >50 years), and only 10% converting to HCM clinically (24). Thus, it would appear that most HCM gene carriers will experience a benign natural history characterized by very low clinical risk or likelihood of developing the pathophenotype over a lifetime. This finding underscores the principle that HCM should not (and perhaps cannot) be viewed solely through the prism of simple genetics, particularly given the substantial uncoupling between the gene variants and the probability of developing clinically important HCM.
Population genetics.
One observation that may shed light on the issue of incomplete penetrance relates to the prevalence of pathogenic sarcomere variants in the general population. In the combined Framingham and Jackson Heart population of 3,600 probands, the prevalence of nonsynonymous sarcomere variants unassociated with clinical/imaging evidence of HCM was 0.5% (25). From these data, it can be estimated that about 1.5 million people in the U.S. would meet the criteria for incompletely penetrant HCM. In turn, this approximation dwarfs the number of individuals with bona fide HCM diagnosed clinically (~100,000 individuals) (2), and is at least 2.5-fold greater than the frequency of HCM in large echocardiographic studies in the general population (1: 500) (1). That many HCM family members with pathogenic sarcomere mutations are without clinical disease suggests that mechanisms critical to disease expression are often independent of sarcomere gene mutations and could include other genetic variants that either offset or promote the adverse consequences of the sarcomere gene mutation, non-genetic or environmental factors, or both.
Variant role in prognosis and treatment.
In HCM, sarcomere gene variants have demonstrated little or no prognostic power, nor a role in effective treatment options, either for end-stage heart failure, atrial fibrillation, or sudden death risk (26). In addition, a cautionary note is appropriate regarding novel treatments proposed to target specific HCM mutations. Personalized strategies based on the monogenicity of HCM that may be advanced to reverse or prevent this disease, such as gene editing (CRISPR-Cas9) and preimplantation genetic diagnosis (PGD), respectively, can be misguided and potentially harmful should the targeted mutation be reclassified later as non-disease-causing, or proves relevant to some (but not all) disease features.
Therefore, organizing management initiatives solely around myofilament mutations may be ineffective or counterproductive, underscoring the importance of developing additional relevant therapeutic targets. In support of this assertion, major reductions in mortality and morbidity in HCM have thus far been attributable solely to initiatives from the clinical arena unrelated to and undirected by knowledge of sarcomere gene mutations (27).
Future Perspectives and Directions
In our view, it is timely to expand the scientific focus in HCM toward a framework that can account for the totality of disease features observed clinically. To be wedded irrevocably to reductionism, which in HCM proposes that multiple disease features can be traced back to a single gene variant, is diminishing in its plausibility and may have several unintended consequences. In particular, this approach is likely to limit understanding of important pathobiological mechanisms and the discovery of novel therapeutic targets. For example, post-transcriptional regulation of proteins is reported widely in the development of many cardiovascular diseases, but has not been well characterized in HCM (28).
Our call to action emphasizes a move in HCM research focus from the single gene hypothesis to alternative and complementary mechanisms. For example, non-sarcomere gene variants may interact with sarcomere gene variants to promote (or mitigate) certain structural (and thereby functional) features of HCM. Characterizing epigenetic mechanisms is also likely to provide insights that explain the highly prevalent genotype positive-phenotype negative phenomenon.
Acquired HCM Risk Factors
Numerous examples throughout the spectrum of medicine implicate the importance of modifiable risk factors for disease incidence and prognosis. Ample precedents support the principle that non-genetic mechanisms are involved in (or cause) cardiomyopathic disease processes even when there is also a genetic variant that indicates or promotes risk (5, 25). What is being proposed here for HCM is, therefore, not dissimilar to dilated cardiomyopathy for which a number of pathogenic sarcomere gene variants have been identified resulting in a dominant inheritance pattern. The phenotype of dilated cardiomyopathy can also be triggered by various non-genetic factors, including alcohol, viral infection, and toxins. It is possible for an individual patient to develop the numerous features of dilated cardiomyopathy (i.e., thin LV wall and enlarged cavity size, systolic dysfunction, fibrosis, small vessel disease) that, together, comprise a unified clinical phenotype that is variably penetrant relative to a particular genotype. In this model, a number of pathobiological mechanisms can be involved, which operate independent of, or synergistic with, a primary pathogenic gene variant.
The Promise of Network Medicine
HCM is characterized by multiple endophenotypes, often present within the same patient (Figure 2, Central Illustration). This heterogeneity implies that unique signaling pathways may be responsible for the pathogenesis of each HCM endophenotype (e.g., fibrosis, hypertrophy). This conceptual framework, already well-established for other cardiovascular diseases (28), recognizes that the same gene mutation is not required (and in fact cannot plausibly be used) to explain all features of a clinical disorder. Indeed, HCM is likely a clinical entity that comprises numerous distinct ‘diseases,’ as defined by unique pathogenicities (29). The alternative---that different molecular signaling pathways lead to a single clinical phenotype with variable morphologic characteristics---seems less likely. Genetic context defines the clinical phenotype—i.e., other coexisting gene mutations or variants that contribute to disease phenotypes only in the presence of the primary disease driver mutation. Alternatively, structural and functional modifications of mRNA and proteins occur commonly in the absence of gene mutations, and when present can contribute directly to the development of specific endophenotypes. This diversity of molecular mechanisms suggests that an individual HCM clinical phenotype may be determined by an integrated pathobiology rather than solely by a single genetic event.
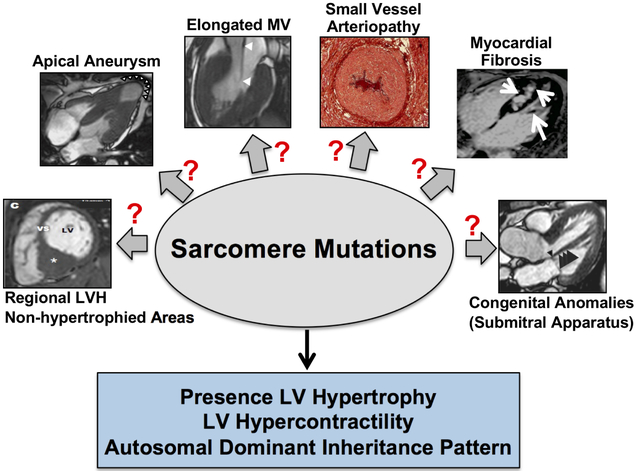
Figure 2. The single sarcomere gene mutation hypothesis suggests HCM is a disease of divergent endophenotypes.
The conventional approach to explaining HCM pathobiology relies on a single cardiomyocyte sarcomere gene mutation to account for multiple endophenotypes. However, many of the HCM endophenotypes involve cell types that do not express cardiomyocyte sarcomeres. Additionally, the single sarcomere gene hypothesis and model corresponds to a collection of divergent endophenotypes, even though multiple endophenotypes are observed within the same patient commonly. LVH, left ventricular hypertrophy; MV, mitral valve. Images reproduced with permission from: long mitral valve, Maron MS, et al. Circulation 2011;124:40.; LV aneurysm, Maron MS, et al. Circulation 2008;118:1541-9.; Anomolous papilary muscle insertion, Rowin EJ, et al. Am J Cardiol. 2013;111:1677-9.; Posterior septum, Rowin EJ, et al. Circulation Cardiovasc Imaging 2017;10(7).
Network medicine focuses on functional and physical associations between biological molecules (e.g., microRNAs, proteins, metabolites) (30), and is likely to offer important insights into understanding HCM for several reasons. First, this approach does not rely on reductionism, and, therefore, does not absolutely require single gene mutational analyses to be exclusively informative. By contrast, network medicine permits utilization of information on epigenetic, post-transcriptional, and post-translational mechanisms to help explain empiric observations. For example, important disease-causing protein-protein signaling pathways regulated by environmental cues (and, therefore, independent of genetic substrate) have already emerged from network analyses in other diseases (28). In addition, network medicine incorporates genetic variant context in interpreting the functional consequences of a primary disease driver mutation. Through the holistic integration of variant function in pathways relevant to the disease phenotype (i.e., disease module in the comprehensive macromolecular interaction network), the consequences of disease-modifying variants can be recognized, and phenotypes distinguished.
Second, with networks it is possible to organize and visualize large amounts of biological information from molecular screens, such as microarrays, in a way that emphasizes the functional significance of the biological data. This integration is important to avoid discovering “red herring” genetic markers, such as variants judged to be pathogenic or benign based primarily on their frequency in populations. Failure to consider this pitfall increases the probability of identifying gene variants that are associated with, but not causative of, disease, and that are often reported from genome-wide population studies (GWAS) in which the frequency of a genetic variant (or other biological component) is alone used to impute functional relevance. A number of evolving analytical strategies enrich the capabilities of network medicine for integrating different datasets, including through machine learning (31). Of particular potential relevance to HCM, these methods have been applied to redefining diseases (32), optimizing the importance of network structure on biological function (33), and drug target identification and drug repurposing (34).
Third, and perhaps most important for HCM, is the opportunity to study and view this disease inclusive of all its clinical, morphologic, and molecular diversity. In this way, it is anticipated that network medicine would provide a unique methodological approach for understanding complex heterogeneous cardiovascular phenotypes, as HCM, not any longer possible by focusing on the single sarcomere gene hypothesis.
Conclusions
There are several credible reasons to challenge conventional wisdom that sarcomere genes are solely responsible for the HCM disease process. Consequently, there is also immense opportunity to re-direct the focus of research toward genetic context and non-genetic mechanisms that underlie HCM (Central Illustration). Therefore, we believe that it is timely to present the argument for reconsidering long-standing assumptions about the cause of HCM, and at the same time opening the possibility of other plausible etiologies. However, this search for the causes of HCM, considering its broad clinical presentation and endophenotypic heterogeneity, requires a more integrative systems-wide, molecular network-guided approach. This call to action suggests that applying the principles of network medicine and expanding considerations for acquired risk factors are necessary for achieving an enhanced understanding of the full biomolecular basis of this complex cardiac disease.
HIGHGLIGHTS.
Single sarcomere gene mutations are insufficient to explain the full clinical and enodphenotypic profile of HCM.
Limitations of the single sarcomere gene hypothesis include failure to identify pathogenic mutations in the vast majority of HCM patients or account for the disease’s mechanistic complexity.
Innovative research methods can be useful for identifying novel determinants of HCM and clarifying the mechanisms responsible for the expression of disease heterogeneity.
Acknowledgments
Funding Sources: B.A.M.: NIH (R56HL131787; 1R01HL139613-01; R21HL145420; 1U01HL125215-01) and National Scleroderma Foundation; JL: NIH (R37HL061795; U54 HL119145; U01 HG007690; P50 GM107618) and AHA (D700382).
Abbreviations:
- HCM
hypertrophic cardiomyopathy
- LVH
left ventricular hypertrophy
Footnotes
Conflicts of Interest: None
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Maron BJ, Gardin JM, Flack JM, Gidding SS, Kurosaki TT, Bild DE. Prevalence of hypertrophic cardiomyopathy in a general population of young adults. Echocardiographic analysis of 4111 subjects in the CARDIA study. Coronary Artery Risk Development in (Young) Adults. Circulation 1995;92:785–9. [DOI] [PubMed] [Google Scholar]
- 2.Alfares AA, Kelly MA, McDermott G, et al. Results of clinical genetic testing of 2,912 probands with hypertrophic cardiomyopathy: expanded panels offer limited additional sensitivity. Genet Med 2015;17:880–8. [DOI] [PubMed] [Google Scholar]
- 3.Maron BJ, Epstein SE. Hypertrophic cardiomyopathy: Discussion of nomenclature. Am J Cardiol 1979;43:1242–4. [DOI] [PubMed] [Google Scholar]
- 4.Jarcho JA, McKenna W, Pare JA, et al. Mapping a gene for familial hypertrophic cardiomyopathy to chromosome 14q1. N Engl J Med 1989;321:1372. [DOI] [PubMed] [Google Scholar]
- 5.Ingles J, Burns C, Bagnall RD, et al. Nonfamilial hypertrophic cardiomyopathy: prevalence, natural history, and clinical implications. Circ Cardiovasc Genet 2017;10:2. [DOI] [PubMed] [Google Scholar]
- 6.Sweeney HL, Feng HS, Yang Z, Watkins H. Functional analysis of troponin T mutations that cause hypertrophic cardiomyopathy: insights into disease pathogenesis and troponin function. Proc Natl Acad Sci USA 1998;95:14406–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Force T, Bonow RO, Houser SR, et al. Research priorities in hypertrophic cardiomyopathy: Report of a working group of the National Heart Lung and Blood Institute. Circulation 2010;122:1130–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Manrai AK, Funke BH, Rehm HL, et al. Genetic misdiagnosis and the potential for health disparities. N Engl J Med 2016;375:655–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gallati S Disease-modifying genes and monogenic disorders: experience in cystic fibrosis. Appl Clin Genet 2014;7:133–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Maron BJ. Clinical course and management of hypertrophic cardiomyopathy. N Engl J Med 2018;379:655–68. [DOI] [PubMed] [Google Scholar]
- 11.Kumar A, Rani B, Sharma R, et al. ACE2, CALM3, and TNNI3K polymorphisms as potential disease modifiers in hypertrophic and dilated cardiomyopathies. Mol Cell Biochem 2018;438:167–174. [DOI] [PubMed] [Google Scholar]
- 12.Moulton JM, van der Merwe L, Goosen L, et al. MYBPH acts as a modifier of cardiac hypertrophy in hypertrophic cardiomyopathy (HCM) patients. Hum Genet 2016;135:477–83. [DOI] [PubMed] [Google Scholar]
- 13.Maron BJ, Wolfson JK, Roberts WC. Relation between extent of cardiac muscle cell disorganization and left ventricular wall thickness in hypertrophic cardiomyopathy. Am J Cardiol 1992;70:785–90. [DOI] [PubMed] [Google Scholar]
- 14.Maron MS, Olivotto I, Harrigan C, et al. Mitral valve abnormalities identified by cardiac magnetic resonance represents a primary phenotype expression of hypertrophic cardiomyopathy. Circulation 2011;124:40–7. [DOI] [PubMed] [Google Scholar]
- 15.Klues HG, Maron BJ, Dollar AL, Roberts WC. Diversity of structural mitral valve alterations in hypertrophic cardiomyopathy. Circulation 1992;85:1651–60 [DOI] [PubMed] [Google Scholar]
- 16.Maron BJ, Wolfson JK, Epstein SE, Roberts WC. Intramural (“small vessel”) coronary artery disease in hypertrophic cardiomyopathy. J Am Coll Cardiol 1986; 8:545–57. [DOI] [PubMed] [Google Scholar]
- 17.Shirani J, Pick R, Roberts WC, Maron BJ. Morphology and significance of the left ventricular collagen network in young patients with hypertrophic cardiomyopathy and sudden cardiac death. J Am Coll Cardiol 2000;35:36–44. [DOI] [PubMed] [Google Scholar]
- 18.Chan RH, Maron BJ, Olivotto I, et al. Prognostic value of quantitative contrast-enhanced cardiovascular magnetic resonance for the evaluation of sudden death risk in patients with hypertrophic cardiomyopathy. Circulation 2014;130:484–95. [DOI] [PubMed] [Google Scholar]
- 19.Rowin EJ, Maron BJ, Haas TS, et al. Hypertrophic cardiomyopathy with left ventricular apical aneurysm: Implications for risk stratification and management. J Am Coll Cardiol 2017;69:761–73. [DOI] [PubMed] [Google Scholar]
- 20.Duncker DJ, Bakkers J, et al. Animal and in silico models for the study of sarcomeric cardiomyopathies. Cardiovasc Res 2015;105:439–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rottbauer W, Wessels G, Dahme T, et al. Cardiac myosin light chain-2: a novel essential component of thick-myofilament assembly and contractility of the heart. Circ Res 2006;99:323–31. [DOI] [PubMed] [Google Scholar]
- 22.Gruner C, Ivanov J, Care M, et al. Toronto hypertrophic cardiomyopathy genotype score for prediction of a positive genotype in hypertrophic cardiomyopathy. Circ Cardiovasc Genet 2013;6:19–26. [DOI] [PubMed] [Google Scholar]
- 23.Bagnall RD, Ingles J, Dinger ME, et al. Whole genome sequencing improves outcome of genetic testing in patients with hypertrophic cardiomyopathy. JACC 2018;72:419–29 [DOI] [PubMed] [Google Scholar]
- 24.Maurizi N, Michels M, Rowin EJ, et al. Clinical course and significance of hypertrophic cardiomyopathy without left ventricular hypertrophy. Circulation 2019; In press. [DOI] [PubMed] [Google Scholar]
- 25.Bick AG, Flannick J, Ito K, et al. Burden of rare sarcomere gene variants in the Framingham and Jackson Heart Study cohorts. Am J Hum Genet 2012;91:513–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Landstrom AP, Ackerman MJ. Mutation type is not clinically useful in predicting prognosis in hypertrophic cardiomyopathy. Circulation 2010;122:2441–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Maron BJ, Rowin EJ, Casey SA, Maron MS. How hypertrophic cardiomyopathy became a contemporary treatable genetic disease with low mortality: Shaped by 50 years of clinical research and practice. JAMA Cardiol 2016;1:98–105. [DOI] [PubMed] [Google Scholar]
- 28.Samokhin AO, Stephens T, Wertheim BM, et al. NEDD9 targets COL3A1 to promote endothelial fibrosis and pulmonary arterial hypertension. Sci Transl Med 2018;10(445). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Loscalzo J, Kohane I, Barabasi A-L. Human disease classification in the postgenomic era: A complex systems approach to human pathobiology. Mol Syst Biol 2007;3:124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Menche J, Sharma A, Kitsak M, et al. Disease networks. Uncovering disease-disease relationships through the incomplete interactome. Science 2015;347:1257601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Eliassi-Rad T, Reducing Networking incompleteness through online learning: A feasibility study (with LaRock T, Sakharov T, and Bhadra S), The 14th International Workshop on Mining and Learning with Graphs (held in conjunction with KDD'18), London, United Kingdon, August, 2018. [Google Scholar]
- 32.Oldham WM, Oliveira RKF, Wang RS, et al. Network analysis to risk stratify patients with exercise intolerance. Circ Res 2018;122:864–76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yang G, Vertes PE, Towlson, et al. Network control principles predict neuron function in the Caenorhabditis elegans connectome. Nature 2017;550(7677):519–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Cheng F, Desai RJ, Handy DE, et al. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat Commun 2018;9:2691. [DOI] [PMC free article] [PubMed] [Google Scholar]