Abstract
Background
A tick-borne segmented RNA virus called Jingmen tick virus (JMTV) was recently identified, variants of which were detected in a non-human primate host and fatal patients with Crimean-Congo haemorrhagic fever. We investigated its infectivity and pathogenicity for humans.
Methods
We obtained skin-biopsy, blood and serum samples from patients with tick bites, and used high-throughput sequencing, in situ hybridisation, and serologic testing to diagnose and ascertain the cases of JMTV infection.
Findings
A JMTV strain was isolated from the tick Amblyomma javanense into an embryo-derived tick cell line. We obtained sustained passage of JMTV, and revealed that it was able to accumulate in salivary glands of experimentally infected ticks. Four JMTV-infected patients were identified by high-throughput sequencing of skin biopsies and blood samples. The virus replication in skin tissue was visualised by in situ hybridisation. The four patients all had an itchy or painful eschar at the site of tick bite, with or without lymphadenopathy. Immunohistochemical examination revealed remarkable local inflammation manifested as infiltration by neutrophils. Eight patients were identified by serological testing and showed more severe clinical manifestations. Two Ixodes persulcatus ticks detached from patients were positive for JMTV. All JMTV strains identified in this study formed a well-supported sub-lineage, distinct from those previously reported in China.
Interpretation
The public significance of JMTV should be highly concerning due to its potential pathogenicity for humans and efficient transmission by potential ticks.
Fund
China Natural Science Foundation, State Key Research Development Programme, and United Kingdom Biotechnology and Biological Sciences Research Council.
Keywords: Jingmen tick virus, Pathogenicity, Human infection, Ticks
Panel: Research in context.
Evidence before this study
We searched PubMed and Embase with the terms “Jingmen Tick Virus (JMTV)” for reports published in English and in Chinese before January 30, 2019. We also searched the China National Knowledge Infrastructure database using synonyms in Chinese. To our knowledge, JMTV was recently identified in Rhipicephalus microplus, seven other tick species, Armigeres sp. (mosquito) and Bos sp. (cattle) from southern or central China. JMTV carries four genome segments, two of which exhibit homology to nonstructural protein NS3 and NS5 sequences of unsegmented RNA viruses in the family Flaviviridae, whereas the other two segments are unique, suggesting that they might have been acquired independently from as-yet-unidentified parental ancestors. JMTV also existed in four predominant tick species in northern China and in R. microplus in Brazil. A variant of JMTV was observed in a non-human primate in Uganda. More recently, JMTV genome sequences were detected in fatal cases of Crimean-Congo haemorrhagic fever (CCHF) in Kosovo by viral metagenomics. However, JMTV could not be maintained in culture, and disappeared after three passages in mosquito (C6/36) and mammalian (DH82) cells, thus their basic biological properties and pathogenicity to humans were unclear.
Added value of this study
In this study, a JMTV strain was isolated from Amblyomma javanense and maintained in the tick embryo-derived cell line BME/CTVM23. Four JMTV-infected patients were identified by high-throughput sequencing of skin biopsies and blood samples. Immunohistochemical examination revealed remarkable local inflammation manifested as infiltration by neutrophils. Eight patients were identified retrospectively by serological testing and showed more severe clinical manifestations. All JMTV strains identified in this study formed a well-supported sub-lineage, distinct from those previously reported in China.
Implications of all the available evidence
JMTV is a newly identified human pathogen. The public health significance of JMTV should be highly concerning due to its potential pathogenicity to humans and potential for efficient transmission by ticks. Physicians and health-care workers in areas with reported tick infection should be aware of possible human infections with this virus in suspected patients.
Alt-text: Unlabelled Box
1. Introduction
A tick-borne segmented RNA virus, called Jingmen tick virus (JMTV), was recently identified in ticks from Hubei Province of middle China but without any report of human infection [1]. The JMTV genome comprises four segments: two of them are related to the non-structural protein NS3 and NS5 sequences of unsegmented RNA viruses in the family Flaviviridae, whereas the other two segments are unique, suggesting that they might have been acquired independently from as-yet-unidentified parental ancestors. Subsequently, several other segmented, flavi-like viruses were reported in arthropods and vertebrates, and have been tentatively named as ‘Jingmenvirus group’ [2].
A variant of JMTV was observed in a non-human primate in Uganda [3]. More recently, JMTV genome sequences were detected in fatal cases of Crimean-Congo haemorrhagic fever (CCHF) in Kosovo by viral metagenomics [4]. These findings indicate the potential pathogenicity of this emerging tick-borne virus for humans. In this paper, we characterise in vitro replication and in vivo tick infection of a JMTV isolated in China, describe the identification of infected patients, and report clinical, serological and immunohistochemical findings of these cases.
2. Materials and methods
2.1. In vitro culture and characterisation of the virus
Amblyomma javanense ticks collected from Manis javanica (pangolins) in Guangxi Autonomous Region of southern China were tested individually by RT-PCR with Flavivirus genus-specific primers targeting the NS5 region (Supplemental Table S1) [5]. The positive amplicons were sequenced and analysed by BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi), revealing that they were genetically close to JMTV with 92·6% identity. A total of 154 A. javanense were then cut vertically into two parts using a sharp blade. The smaller part of each tick was used to extract RNA and tested by a SYBR Green One-Step RealTime RT-PCR (Takara) targeting the segment 2 sequence of JMTV (Supplemental Table S1), and the larger part of each tick was stored at −80 °C for virus isolation.
A total of seven JMTV-positive ticks were used in attempts to isolate the virus. In brief, individual ticks were ground in PBS with a pestle in a porcelain mortar. The resultant suspensions were filtered through a 0·22 μm syringe filter and then aliquots were inoculated onto monolayers of around 60% confluent Vero (African green monkey kidney cell line ATCC CCL-81) cells, DH82 (canine macrophage cell line, ATCC CRL-10389) cells, C6/36 (Aedes albopictus mosquito-derived cell line ATCC CRL-1660) cells and BME/CTVM23 (Rhipicephalus microplus embryo-derived cell line) [6]. After incubation at 28 °C for 1 h, the cells were topped up with the respective growth medium (containing antibiotics) and maintained at the same temperature for seven days. The supernatants from all cell types in which isolation was attempted were then examined by JMTV-specific One-step RealTime RT-PCR as above (Supplemental Table S1). Three successive cell passages were carried out and cultures were tested for virus replication by JMTV-specific PCR at intervals up to one month post inoculation.
We isolated JMTV from ticks using BME/CTVM23. Total RNA was extracted from the culture using the High Pure Viral RNA Kit (Roche, Switzerland) for viral genome sequencing. To determine virus growth, the copy numbers of each viral segment were determined by quantitative RT-PCR (qRT-PCR) (appendix) from each culture at 12-h periods post infection and then daily for 7 days. Fluorescence in-situ hybridisation (FISH) using segment-specific probe sets (Supplemental Table S2) were performed to calculate the viral dose-response curve and visualise replication in BME/CTVM23 cells.
Ticks were experimentally infected using a previously-described method [7]. In brief, adult male A. javanense ticks were exposed to a low relative humidity (47–60%) for 48–72 h before immersion to partially dehydrate them and increase the likelihood that they would imbibe the virus suspension. Ticks were then surface-sterilised and were immersed in a suspension of JMTV in complete L-15 medium [6] and control ticks were immersed in complete L-15 medium, and incubated at 28 °C for 30 min. The tubes were shaken every 10 min to redistribute the ticks in the medium. The tick suspension was then centrifuged at 200 ×g for 30 s and chilled on ice for 5 min. After the suspension was removed, the ticks were washed three times with cold PBS followed by drying, and returned to incubation under the original conditions. JMTV in tick midguts and salivary glands was visualised by FISH (appendix).
2.2. Study design and enrolled patients
We did a retrospective study at Mudanjiang Forestry Central Hospital, Heilongjiang province in northeastern China, where various emerging tick-borne diseases had been reported [[8], [9], [10], [11], [12], [13]]. The patients, from whom skin-biopsy specimens at tick bite sites were collected, were included in the study to detect JMTV. In addition, patients with a history of tick-bite during 2010–2018, from whom paired sera of acute and convalescent phases were available, were enrolled to screen serum IgG antibodies against JMTV. We collected data on demographic information, medical history and tick exposure of each patient from the pre-existing questionnaire. Data for clinical manifestations, underlying conditions, laboratory tests, treatment, and outcomes were retrieved from medical records. Other tick-borne pathogens, such as Anaplasma phagocytophilum, “Anaplasma capra”, Candidatus Neoehrlichia mikurensis, Ehrlichia chaffeensis, Spotted Fever Group Rickettsiae (SFGR), Borrelia burgorferi sensu lato, Borrelia miyamotoi, Babesia, tick-borne encephalitis virus (TBEV) and severe fever with thrombocytopenia syndrome virus were recorded or identified by previous testing [[8], [9], [10], [11], [12], [13], [14], [15], [16]].
The study was approved by the ethics committee of Mudanjiang Forest Central Hospital (2011-No. 03) in accordance with the medical research regulations of China. All the enrolled participants had provided written or verbal informed consent.
2.3. Diagnostic tests
The skin-biopsy specimens were divided into two pieces, and one piece was put into an RNase-free tube containing Sample Protector for RNA/DNA (Takara, Dalian, China) and stored in liquid nitrogen. Total RNA was then extracted using an AllPrep DNA/RNA Mini Kit (Qiagen, USA), and used for high-throughput sequencing (appendix). RNA was also extracted from blood samples (previously stored in Trizol reagent at −80 °C) of patients positive for JMTV in their skin-biopsy specimens, and tested by qRT-PCR and high-throughput sequencing as mentioned above.
The second half of each skin-biopsy was prepared as a Formalin-Fixed Paraffin-Embedded (FFPE) specimen. In situ hybridisation was performed to detect the virus as previous described [17]. A set of probes labeled with Quasar 570 were used. Haematoxylin-eosin staining was conducted following a routine method. Immunohistochemical staining for the T-lymphocyte marker CD3, the polymorphonuclear leucocyte marker CD15, and the macrophage marker CD68 was performed with a peroxidase-based method [18].
We tested serum samples of enrolled patients for IgG antibodies against JMTV by indirect immunofluorescence assay (IFA).In brief, supernatant from the original BME/CTVM23 culture in which the virus was isolated from the tick was inoculated onto fresh BME/CTVM23 cells. We then prepared antigen slides by placing JMTV-infected cells into wells of Teflon-coated slides. The diluted serum samples were added onto slides and incubated at 37 °C for 30 min. After washes, the slides were incubated with FITC-conjugated goat anti-human IgG antibody (1:100 dilution). Evans Blue was used to counterstain host-cell nuclei. A negative control (sterile phosphate-buffered saline) was concurrently included in each slide. The tested sera were screened at a 1:80 dilution and titrated if reactive. To validate the serological assay, we tested 100 serum samples collected from health individuals without tick-bite history, and did not find any positive reaction at a titre of 1:20. Generally speaking, a threshold of 1:40 can be used to screen the samples. To ensure the diagnosis, we chose a higher titre of 1:80 as threshold.
2.4. Tick survey
Ticks feeding on patients who sought treatment at Mudanjiang Forestry Central Hospital were collected and tested. In addition, A. javanense collected from M. javanica in GuangXi were included. The species level and developmental stage were identified by an entomologist (Yi Sun, the 10th author). Most ticks were stored in liquid nitrogen or at −80 °C; a subset of A. javanense collected in July 2017 were reared in an incubator for the tick infection experiment as described above.Total RNA was extracted, and was tested either by RT-PCR or high-throughput sequencing.
2.5. Phylogenetic analyses
Nucleotide sequences in this study were aligned with those of the previously reported JMTV viruses using the E-INS-I algorithm implemented in MAFFT version 7.3 [19], with mean coverages from 73% to 78% in the four genome segments. There were few relatively shorter sequences with lengths between 300 bp to 900 bp. Additional phylogenetic trees excluding these short sequences were conducted, which generated consistent topology with the complete phylogeny presented. Phylogenetic trees were inferred by the Bayesian Markov Chain Monte Carlo (BMCMC) method in MrBayes version 3.2.6 [20], using the WAG+Γ amino acid substitution model or GTR + Γ nucleotide substitution model and 20,000,000 generations sampled every 2000 steps. Maximum clade credibility trees were summarised from the tree samples in TreeAnnotator version 1.8.4 [21], and visualised with ggtree package [22]. The topological support was assessed by, in addition to the posterior clade probability, also the maximum likelihood bootstrap method (1000 replicates) using PhyML version 3 [23], with the same substitution models.
2.6. Role of the funding sources
The sponsors of the study had no role in study design, data collection, data analysis, interpretation, or writing of the report. The corresponding author had full access to all the data in the study and had final responsibility for the decision to submit for publication.
3. Results
3.1. Biological characteristics of JMTV
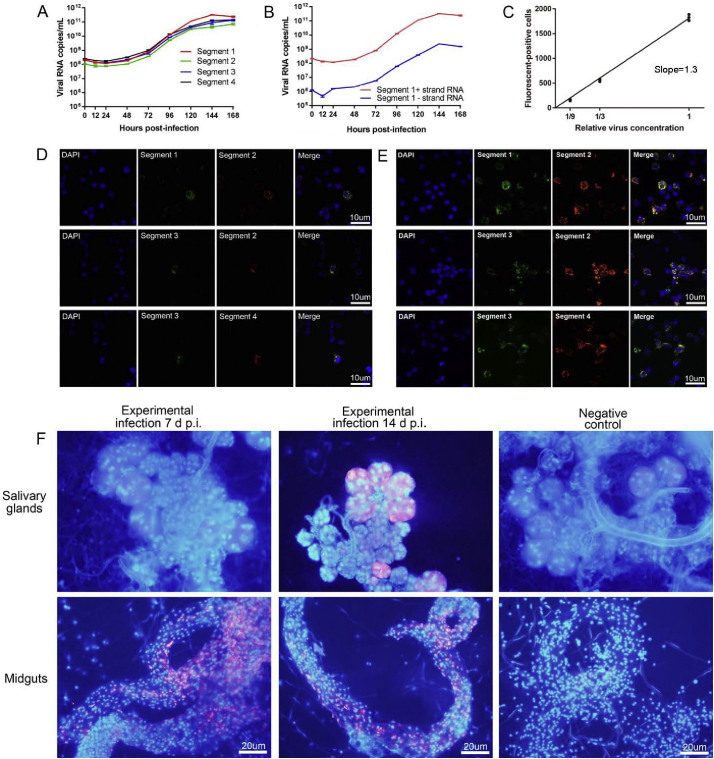
A strain of JMTV was isolated into BME/CTVM23 cells from a RT-PCR-positive adult male A. javanense tick originally collected in October 2016 from M. javanica in Guangxi Autonomous Region of southern China.The complete genome was sequenced and deposited in GenBank with the accession numbers of MG703252- MG703255. The copy number of each JMTV segment simultaneously increased from around 1·0 × 108 to over 1·0 × 1011 copies/ml 7 days post inoculation, showing obvious exponential growth (Fig. 1A). We observed that both positive and negative strands of segment 1 efficiently replicated in BME/CTVM23 cells 48 h post-inoculation. However, the copy number of the negative strand was about 100 times less abundant than that of the positive strand (Fig. 1B), which is a hallmark of single-stranded, positive-sense RNA (ss RNA +) virus replication [24].The number of fluorescent-positive cells per 1·0 × 104 cells was proportional to the first power of the relative concentration of JMTV (Fig. 1 C). Pairs of segments were always visualised together in the individual cells (Fig. 1 D, E, Supplemental Fig. 1S).
Fig. 1.
Biological features of JMTV in BME/CTVM23 cells.
Growth curves of each of the four genome segments of JMTV in BME/CTVM23 cells. The copy numbers of each viral segment were calculated by a standard curve method using a linearised plasmid containing the corresponding segment (Panel A). Detection of positive strand and negative strand of segment 1 by strand-specific quantitative PCR with a tagged primer system during growth in BME/CTVM23 cells (Panel B). The dose-response curve for JMTV. The count of fluorescent-positive cell for representative segment 2 probe sets per 104 cells is shown on the y axis, and the relative viral dilution is shown on the x axis (Panel C). Detection of each genome segment of JMTV in infected BME/CTVM23 cells by fluorescence in-situ hybridisation (FISH). Cells were fixed at 72 h (Panel D) and 144 h (Panel E) post infection respectively. Magnification×200. Detection segment 2 probe sets-JMTV FISH in salivary glands and midguts of male A. javanense experimentally-infected at 7 and 14 days post infection. Nuclei, DAPI (blue), Segment 2 probe, Quasar 570 (red). Magnification×400 (Panel F). Three biological replicates were run for experiments of Panel A to E. Five biological replicates were run for experiment of Panel F. Three technical replicates were run for all experiments. Estimates from the three technical replicates were averaged and error bars indicated standard error across biological replicates. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
We detected replication of the virus in experimentally-infected adult male A. javanense ticks by FISH. The virus was seen in midguts 7 and 14 days after infection, and was also visible in salivary glands 14 days post infection (Fig. 1 F).
3.2. Diagnostic testing
A total of 16 skin-biopsy specimens were tested by high-throughput sequencing, and four presented complete or near complete JMTV genome sequences. The JMTV nucleotide sequences from the four patients showed between 97% and 99% similarity among the different segments. Blood samples from the four patients were all positive by qRT-PCR (viral load 1·26 × 106−3·98 × 107 copies per ml) and were then tested by transcriptome sequencing; two of them (patients 2 and 3) had JMTV sequences (Table 1).
Table 1.
Characteristics of Patients with JMTV Infections in Northeastern China, 2010 through 2018.
| Characteristics | Patients |
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
| Diagnosis method | ||||||||||||
| RNA-seq of skin biopsies (RPM) | 2334 | 287 | 105 | 105 | NA | NA | NA | NA | NA | NA | NA | NA |
| qRT-PCR of blood or serum RNA | + | + | + | + | + | − | − | − | − | − | − | − |
| IgG antibodiesa | ||||||||||||
| Acute phase (days of sample collectionb) | <20 (5) | NA | <20 (2) | NA | <20 (3) | <20 (1) | <20 (1) | <20 (4) | <20 (4) | <20 (4) | <20 (4) | <20 (3) |
| Convalescent phase (days of sample collection) | NA | NA | NA | NA | 80 (41) | 80 (15) | 160 (7) | 160 (14) | 160 (17) | 80 (57) | 80 (28) | 160 (11) |
| Epidemiological feature | ||||||||||||
| Sex | F | F | F | F | M | F | F | M | M | F | F | F |
| Age-yr | 53 | 33 | 69 | 43 | 28 | 61 | 2 | 46 | 25 | 46 | 62 | 41 |
| Tick-bite location | Ear | Scalp | Back/Arm | Back | Abdomen/Back/Axilla | Back | Scalp | Check | Axilla | Scalp | Scalp | Back/Neck |
| Sample collection-yr | 2016 | 2016 | 2016 | 2016 | 2010 | 2012 | 2012 | 2012 | 2012 | 2013 | 2014 | 2018 |
| Clinical manifestation | ||||||||||||
| Elevated temperature (°C) | − | − | − | − | 39 | − | 39 | 39 | 38·5 | − | − | 37·5 |
| Asthenia | − | − | + | − | + | − | − | − | − | + | − | + |
| Headache | − | − | + | − | + | + | − | + | + | + | + | + |
| Anorexia | − | − | − | − | + | − | − | − | − | + | − | − |
| Myalgia | − | − | − | − | + | − | − | − | + | − | − | + |
| Arthralgia | − | − | − | − | − | − | − | − | − | − | − | − |
| Nausea/Vomiting | − | − | − | − | + | + | − | − | − | + | − | − |
| Lymphadenopathy | + | + | − | − | − | − | − | − | − | − | − | − |
| Seizure | − | − | − | − | + | − | − | − | − | − | − | − |
| Skin lesions | ||||||||||||
| Eschar | + | + | + | − | − | − | − | − | + | − | − | − |
| Rash | − | − | − | − | − | − | − | − | − | − | − | + |
| Pruritus | + | + | − | + | − | − | − | − | − | − | − | − |
| Pain or burning | + | − | − | − | − | − | − | − | + | − | − | − |
| Ulceration | + | − | − | − | − | − | − | − | + | − | − | − |
| Laboratory finding | ||||||||||||
| Leucocyte count- × 10−9/l | 4·8 | NA | 4·5 | NA | 5·6 | 6·6 | 5·2 | 4·8 | 5·4 | 5·5 | 5·8 | 6·7 |
| Lymphocyte count- × 10−9/l | 2·0 | NA | 2·7 | NA | 2·0 | 1·3 | 0·75 | 2·4 | 1·2 | 1·7 | 2·9 | 3·2 |
| Neutrophil count- × 10−9/l | 2·6 | NA | 1·6 | NA | 3·0 | 5·1 | 4·3 | 2·2 | 4·0 | 3·7 | 2·7 | 2·8 |
| Platelet count- × 10−9/l | 180 | NA | 230 | NA | 197 | 198 | 378 | 234 | 143 | 231 | 258 | 367 |
| AST-U/l | 59 | NA | 10 | NA | 18 | NA | NA | 28 | NA | NA | NA | 26 |
| ALT-U/l | 71 | NA | 19 | NA | 36 | NA | NA | 45 | NA | NA | NA | 78 |
| Hospitalization-days | 14 | − | − | − | 28 | − | − | 15 | 14 | − | − | 14 |
| Co-infection | − | − | − | − | − | − | − | − | − | SFGR | SFGR | SFGR |
aData is the reciprocal of the serum dilution; bNumber of days after the onset of illness for patients in the acute or the convalescent phase of the illness.
RNA-seq, RNA sequencing; RPM, reads per million; NA, not available (no samples were collected or not reported); AST, aspartate aminotransferase, ALT, alanine aminotransferase; SFGR, spotted fever group Rickettsia.
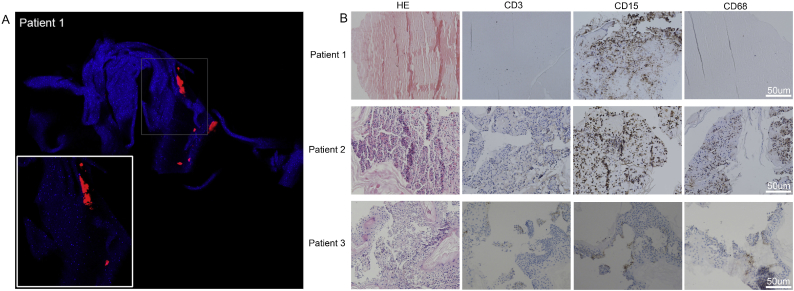
The skin-biopsy specimens were tested by in situ hybridisation, and accumulated viruses were observed in various parts of the slides (Fig. 2 A, Supplemental Fig. 2S). Histopathological examinations of skin-biopsy specimens from the patients showed remarkable coagulative necrosis. Local inflammation was manifested as infiltration mostly by neutrophils (Fig. 2 B).
Fig. 2.
In situ Hybridisation and Immunohistochemical Analysis of Skin-biopsies with JMTV Infection. In situ RNA hybridisation using JMTV-segment 2 probe sets (Quasar 570) on formalin-fixed, paraffin-embedded (FFPE) skin eschar of Patient 1. Nuclei were counterstained with DAPI (blue). Magnification×200, insert×400 (Panel A). Haematoxylin and eosin staining, and immunophenotyping of lymphocytes (CD3), neutrophils (CD15) and macrophages (CD68) on serial sections of Patients 1–3. Magnification×200, counterstained with haematoxylin, peroxidase immunostaining in brown (Panel B). No FFPE specimen was available for Patient 4. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Eight (1·6%) of 509 patients, from whom we obtained paired serum samples, showed seroconversion of IgG antibody against JMTV (Table 1, Supplemental Fig. 3S). The acute phase serum sample from patient 5 was positive for JMTV by qRT-PCR (viral load 2·51 × 107 copies per ml).The eight patients were tested for IgG or IgM antibodies against TBEV. No positive reaction was recorded.
3.3. Patients and clinical manifestations
We identified four patients with JMTV infection by high-throughput sequencing of skin-biopsy and blood samples. The first case, suffering from pulmonary tuberculosis for 10 years, was a 53-year-old woman. She was admitted to the hospital due to painful ulceration and yellow drainage at a tick-bite site on her ear. She reported a tick-bite ten days previously and subsequently removed the tick herself. The abnormal laboratory testing included rises in aspartate aminotransferase (AST) and alanine aminotransferase (ALT), and decrease in prothrombin time. She was administrated with doxycycline at a dosage of 100 mg twice daily for 2 weeks, and discharged after healing of her skin lesion.
Case 2 was a previously healthy woman of 33 years of age. She did not report a tick-bite, but thought that she had a “tumor” on her scalp. Her husband noticed that the “tumor” was movable when touched. After a neighbor told her it might be a feeding tick, she saw a doctor at the sentinel hospital. At the time, the tick was engorged (Supplemental Fig. 4S). She recalled getting the tick bite about one month ago when she was collecting wild vegetables in the mountains. She had cervical lymphadenopathy about 10 days previously and took cephalosporins for a week. The engorged tick was detached and identified as Dermacentor silvarum, and it laid eggs 15 days later. No further symptoms were reported during the follow-up telephone interview.
Case 3 was a 69-year-old woman with tick bites on her right arm and back. She removed the tick from her arm herself, and came to the hospital for the tick on her back to be removed the next day. The tick on her back was identified as Ixodes persulcatus. At the same time, a skin biopsy was collected. Eight days later, she saw a doctor again because of two-day headache and asthenia. Haematologic tests revealed a low neutrophil count. She was prescribed with doxycycline for possible spotted fever group Rickettsia (SFGR) infection.
Case 4, a 43-year-old woman, came to hospital with a tick that had been attached for a day on her hip. She felt pruritus at the site of tick-bite, and inguinal lymphadenopathy was noticed. The feeding tick was detached and identified as I. persulcatus. After a skin biopsy was collected, she went back home. The follow-up telephone interview did not record any further symptoms.
The eight patients identified by retrospective serological testing showed more severe clinical manifestations. One of the 8 cases occurred in 2010, four in 2012, one in 2013, one in 2014, and one in 2018. Median age was 43·5 years (range 2–62 years), and 5 were female. The interval from tick exposure to illness onset ranged from 2 to 16 days with a median of 6·5 days. In general, clinical presentations of the patients included fever (5), headache (7) and myalgia (3). Laboratory abnormalities included increased hepatic aminotransferase concentration (2), and slightly decreased lymphocyte counts (1). Four patients were admitted to hospital because of severe disease with high fever. The median length of hospitalization was 14 days (range 14–28 days). One patient (patient 5) even presented with seizure. This patient was treated with the antivirals acyclovir (0·5 g twice daily i.v) and ribavirin (0·5 g twice daily i.v) for suspected TBE infection, although the TBEV laboratory test was negative. The other three hospitalised patients were treated with doxycycline or penicillin. Two patients (patients 10 and 11) were co-infected with Candidatus Rickettsia tarasevichiae, which was confirmed by specific PCR and sequencing. One patient (patient 12) had a high titre of IgG antibody against R. heilongjiangensis. The three patients co-infected with SFGR did not show more severe manifestations than the other patients (Table 1). The laboratory tests for other co-infections were all negative (Supplemental Table S3).
3.4. Infection of ticks
The two I. persulcatus ticks collected from patients 3 and 4 were positive for JMTV by RT-PCR. The JMTV genome sequence from each tick showed 99·2%-99·9% nucleotide identity to that of the patient from whom the tick was detached. In addition, fifteen tick pools (5–10 ticks per pool) were tested by transcriptome sequencing. JMTV was detected in one pool of I. persulcatus, two pools of D. silvarum, one pool of H. concinna and three pools of A. javanense (Supplemental Fig. 5S). In addition, RT-PCR-based screening identified 6·0% % (32/542) JMTV infection in individually tested A. javanense ticks.
3.5. Phylogenetic analyses
The JMTV sequences from the patients exhibited high similarities with those from ticks in this study and had 90·2%–93% nucleotide identity with JMTV R27 from Uganda, 90·6%–96·1% nucleotide identity with JMTV strains detected in other regions of China, while relatively low nucleotide identity (72·7%-82·6%) with Kosovo strains.
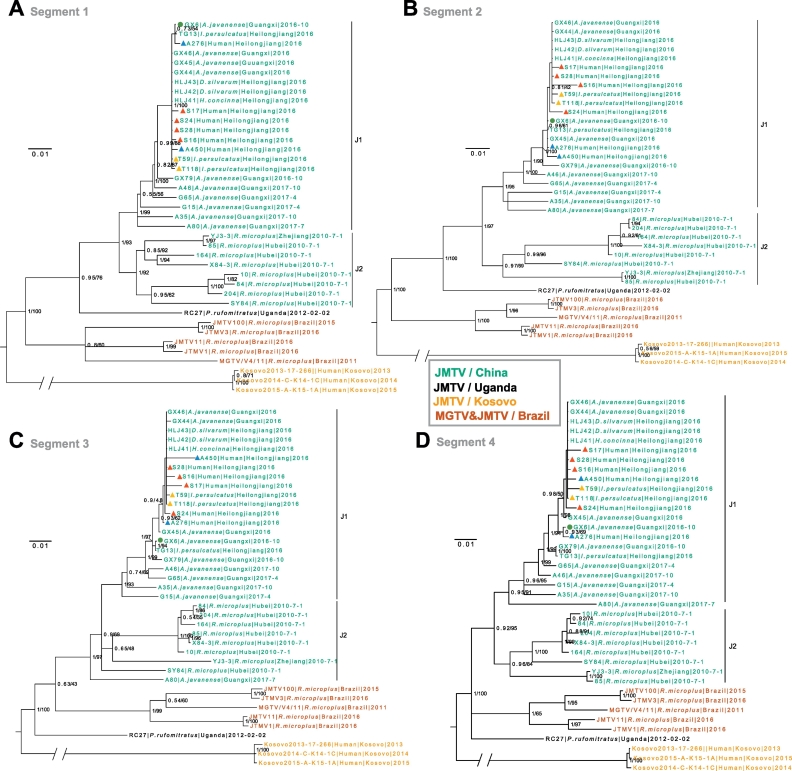
In the phylogenetic trees based on four segments (Fig. 3), all Chinese JMTVs comprised a monophyletic lineage, which was separated from Brazilian MGTV and Ugandan and Kosovan JMTVs. All JMTV strains, including those from patients, identified in this study formed a well-supported sub-lineage (J1), distinct from but close to the clade comprising JMTVs previously reported in China (J2; which is less well resolved in segment 3 tree, Fig. 3C).
Fig. 3.
Phylogenetic Analysis of JMTVs. Bayesian phylogeny of gene segment 1 (Panel A), segment 2 (Panel B), segment 3 (Panel C) and segment 4 (Panel D). The tree is midpoint rooted. Green Dot for isolate strain; Red triangle for human skin samples; Blue triangle for human blood samples; Yellow triangle for ticks detached from patients. The samples relating to four patients are indicated as follows: Case 1: S17; Case2: S24, A276; Case3: S28, A450, T118; Case4: S16, T59. Bayesian posterior probabilities (BPP) and Maximum likelihood (ML) bootstrap values are displayed at major nodes (BPP/ML). Bootstrap testing (1000 replicates) was performed, and the bootstrap values are indicated. Branches are drawn to the scale of substitutions per site. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
4. Discussion
We isolated a strain of JMTV from A. javanense in the tick embryo-derived cell line BME/CTVM23. The sustained passage of the virus in the tick cells allowed us to investigate its biological properties from various aspects. The accumulation and replication of JMTV in salivary glands of experimentally-infected ticks suggests that tick vectors likely transmit the virus to hosts as early as their snouts insert the skin. Therefore, we retrospectively screened for JMTV in human skin biopsy specimens collected from the sites of tick bites through high-throughput sequencing, and obtained whole genome sequences of JMTV from four patients. The virus replication in skin tissue was proved by in situ hybridisation using specific probes (Fig. 2A). Then the viral RNA was detected in blood samples from the four patients. The presence and replication of JMTV in biopsy and blood samples provided the evidence of viral replication and the production of viremia in humans. In addition, eight cases with JMTV infection were identified by IgG antibody seroconversion against JMTV isolated in this study. These findings provide direct evidence that JMTV can be potentially pathogenic to humans.
Human infection with JMTV could be traced to 2010, by retrospective serological diagnosis of a patient. JMTV infection was more frequent in women, as seen with other tick-borne infections such as those associated with lymphadenopathy caused by some species of SFGR [25]. The clinical manifestations of JMTV infection ranged from mild illness to severe disease requiring admission to hospital. Although JMTV infection was difficult to distinguish clinically from other acute febrile illnesses presenting acute onset of fever, headache, and malaise, etc., we did find some distinct clinical presentations. Patients often presented an itchy or painful eschar after a tick bite. Lymphadenopathy was also observed in these patients. A screening bias might exist as patients with more severe disease were more likely to have provided the convalescence phase sample. We cannot describe the full clinical spectrum according to the data from our limited number of patients infected with JMTV, so the clinical features of JMTV infection should be further investigated with clinical data from more cases.
We not only detected viral sequences but also proved multiplication of JMTV by in situ hybridisation in skin-biopsy specimens collected at the sites of tick bites. These findings imply that JMTV might quickly replicate in local tissue following inoculation by a feeding tick, and subsequently lead to skin necrosis that forms an eschar at the site of a tick bite. The local inflammation manifested as infiltration by neutrophilsmight have a role in virus clearance by the immunological response of the host as shown for TBEV (Fig. 2B) [26]. The localised infection and its potential pathogenic features require further investigation. Three patients were co-infected with SFGR. Although the clinical presentation observed in the co-infected patients was no more severe than in singly-infected patients, it is difficult to determine whether the disease was caused either by SFGR or JMTV or both. The tendency for co-infection with JMTV and SFGR deserves further investigation.
Ticks detached from patients were infected with JMTV, which provided further evidence for transmission by ticks. The accumulation and replication of JMTV in salivary glands of experimentally-infected ticks suggests that tick vectors likely transmit the virus to hosts as soon as feeding is initiated. This characteristic has also been observed in other tick-borne viruses, such as the flaviviruses TBE virus and Powassan virus, that accumulate in salivary glands prior to blood-feeding and are immediately transmitted to the host on feeding [[27], [28], [29]]. In contrast, some other tick-borne viruses, such as Thogoto virus and CCHF virus [30,31], are usually present at low or undetectable levels in salivary glands of unfed ticks but accumulate significantly during or after blood-feeding.
Sequences recovered from patients all fell into the subclade J1. It is not known whether viruses in subclade J2 infect humans. In the phylogenies of Jingmen tick virus, JMTV R27 from Uganda was divergent from the monophyletic lineage of JMTVs from China. A possible explanation is that these Chinese JMTVs were derived from a single ancestral virus introduced into China many years ago. The evolutionary timescale of the virus emergence could not be estimated with confidence due to the relatively short sampling span of current data. More samples and sequences of JMTVs in the future might provide a dataset sufficient for molecular clock analysis to obtain the emergence timescale, and will reveal more details of geographical spread.
The study has following limitations. The FISH experiment in this study suggested the four segments of JMTV might be packaged in a single viral particle. Further investigation is required to clarify the replication pattern of the segmented virus. To examine JMTV replication in mammalian tissues, either human or other mammalian cell lines should be used for cultivation. This system would perhaps allow for a plague-based assay of infectivity. To confirm packaging of the viral genome, plaque assay should be conducted to observe dose-response effect [3]. The relative abundance of segments should be taken into account when interpreting dose-response curve, because it could impact the conclusion. If only one segment is present at much lower abundance than the others, and all segments are needed for replication, then the dilution curve dynamics will be driven entirely by the concentration of that segment. Therefore, plaque assay will clarify one-hit dynamics of dose-response effect.
The public health relevance of JMTV should not be neglected. The present study and those of Qin et al. and Meng et al. demonstrated JMTVs existing in multiple common species of ticks, several of which attack humans, in southern, central and northern China [1,32].,Further surveys on infection of ticks and laboratory experiments are required to fully characterise the virus.
In summary, JMTV is a newly identified human pathogen. The public health significance of JMTV should be highly concerning due to its potential pathogenicity for humans and potential for efficient transmission by ticks. Physicians and health-care workers in the areas with reported tick infections should be aware of possible human infections with this virus in suspected patients.
Competing interests
The authors have declared that no competing interests exist.
Contributors
NJ, WCC designed the study and wrote the paper. HBL, YCZ, JLS, JL, BGJ, RW, TTY, YLC, JLY, and JFJ recruited the subjects and collected the data. HBL, JL, QW, YS, RW, LYX, WW, LFL, QYL, XMC, and YGT performed laboratory tests. NJ, HBL, WCC, XBN, YG, TTYL did the data analysis. LBS provided tick cell lines and associated intellectual input. XBN, TTYL and LBS critically reviewed the paper. All authors contributed to review and revision of the paper.
Acknowledgments
The tick cell lines were obtained from the Tick Cell Biobank. This study was supported by Natural Science Foundation of China (81621005, 81773492), the State Key Research Development Program of China (2016YFC 1200301, 2016YFC 1201902), and United Kingdom Biotechnology and Biological Sciences Research Council grant BB/P024270/1.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.ebiom.2019.04.004.
Appendix A. Supplementary data
Supplementary material
References
- 1.Qin X.C., Shi M., Tian J.H. A tick-borne segmented RNA virus contains genome segments derived from unsegmented viral ancestors. Proc Natl Acad Sci U S A. 2014;111:6744–6749. doi: 10.1073/pnas.1324194111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Shi M., Lin X.D., Vasilakis N. Divergent viruses discovered in arthropods and vertebrates revise the evolutionary history of the Flaviviridae and related viruses. J Virol. 2016;90:659–669. doi: 10.1128/JVI.02036-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ladner J.T., Wiley M.R., Beitzel B. A multicomponent animal virus isolated from mosquitoes. Cell Host Microbe. 2016;20:357–367. doi: 10.1016/j.chom.2016.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Emmerich P., Jakupi X., von Possel R. Viral metagenomics, genetic and evolutionary characteristics of Crimean-Congo hemorrhagic fever orthonairovirus in humans, Kosovo. Infect Genet Evol. 2018;65:6–11. doi: 10.1016/j.meegid.2018.07.010. [DOI] [PubMed] [Google Scholar]
- 5.Ayers M., Adachi D., Johnson G. A single tube RT-PCR assay for the detection of mosquito-borne flaviviruses. J Virol Methods. 2016;135:235–239. doi: 10.1016/j.jviromet.2006.03.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Alberdi M.P., Nijhof A.M., Jongejan F. Tick cell culture isolation and growth of Rickettsia raoultii from Dutch Dermacentor reticulatus ticks. Ticks Tick Borne Dis. 2012;3:349–354. doi: 10.1016/j.ttbdis.2012.10.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mitzel D.N., Wolfinbarger J.B., Long R.D. Tick-borne flavivirus infection in Ixodes scapularis larvae: development of a novel method for synchronous viral infection of ticks. Virology. 2007;365:410–418. doi: 10.1016/j.virol.2007.03.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jia N., Jiang J.F., Huo Q.B. Rickettsia sibirica subspecies sibirica BJ-90 as a cause of human disease. N Engl J Med. 2013;369:1176–1178. doi: 10.1056/NEJMc1303625. [DOI] [PubMed] [Google Scholar]
- 9.Jia N., Zheng Y.C., Jiang J.F. Human infection with Candidatus rickettsia tarasevichiae. N Engl J Med. 2013;369:1178–1180. doi: 10.1056/NEJMc1303004. [DOI] [PubMed] [Google Scholar]
- 10.Jiang J.F., Zheng Y.C., Jiang R.R. Epidemiological, clinical, and laboratory characteristics of 48 cases of "Babesia venatorum" infection in China: a descriptive study. Lancet Infect Dis. 2015;15:196–203. doi: 10.1016/S1473-3099(14)71046-1. [DOI] [PubMed] [Google Scholar]
- 11.Li H., Zheng Y.C., Ma L. Human infection with a novel tick-borne Anaplasma species in China: a surveillance study. Lancet Infect Dis. 2015;15:663–670. doi: 10.1016/S1473-3099(15)70051-4. [DOI] [PubMed] [Google Scholar]
- 12.Li H., Jiang J.F., Liu W. Human infection with Candidatus Neoehrlichia mikurensis. China Emerg Infect Dis. 2012;18:1636–1639. doi: 10.3201/eid1810.120594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Jia N., Zheng Y.C., Ma L. Human infections with Rickettsia raoultii, China. Emerg Infect Dis. 2014;20:866–868. doi: 10.3201/eid2005.130995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jiang B.G., Jia N., Jiang J.F. Borrelia miyamotoi infections in humans and ticks, northeastern China. Emerg Infect Dis. 2018;24:236–241. doi: 10.3201/eid2402.160378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Liu H.B., Wei R., Ni X.B. The prevalence and clinical characteristics of tick-borne diseases at one sentinel hospital in northeastern China. Parasitology. 2018;1:1–7. doi: 10.1017/S0031182018001178. [DOI] [PubMed] [Google Scholar]
- 16.Liu W., Li H., Lu Q.B. Candidatus Rickettsia tarasevichiae infection in eastern Central China: a case series. Ann Intern Med. 2016;164:641–648. doi: 10.7326/M15-2572. [DOI] [PubMed] [Google Scholar]
- 17.Wang F., Flanagan J., Su N. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn. 2012;14:22–29. doi: 10.1016/j.jmoldx.2011.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Paris D.H., Phetsouvanh R., Tanganuchitcharnchai A. Orientia tsutsugamushi in human scrub typhus eschars shows tropism for dendritic cells and monocytes rather than endothelium. PLoS Negl Trop Dis. 2012;6 doi: 10.1371/journal.pntd.0001466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Katoh K., Standley D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–780. doi: 10.1093/molbev/mst010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ronquist F., Teslenko M., van der Mark P. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61:539–542. doi: 10.1093/sysbio/sys029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Drummond A.J., Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007;7:214. doi: 10.1186/1471-2148-7-214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yu G.C., Zhu H., Guan Y. GGtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol Evol. 2017;8:28–36. [Google Scholar]
- 23.Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003;52:696–704. doi: 10.1080/10635150390235520. [DOI] [PubMed] [Google Scholar]
- 24.Plaskon N.E., Adelman Z.N., Myles K.M. Accurate strand-specific quantification of viral RNA. PLoS One. 2009;4 doi: 10.1371/journal.pone.0007468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Parola P., Paddock C.D., Socolovschi C. Update on tick-borne rickettsioses around the world: a geographic approach. Clin Microbiol Rev. 2013;26:657–702. doi: 10.1128/CMR.00032-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Thangamani S., Hermance M.E., Santos R.I. Transcriptional immunoprofiling at the tick-virus-host interface during early stages of tick-borne encephalitis virus transmission. Front Cell Infect Microbiol. 2017;7:494. doi: 10.3389/fcimb.2017.00494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Labuda M., Nuttall P.A. Tick-borne viruses. Parasitology. 2004;129(Suppl):S221–S245. doi: 10.1017/s0031182004005220. [DOI] [PubMed] [Google Scholar]
- 28.Grabowski J.M., Tsetsarkin K.A., Long D. Flavivirus infection of Ixodes scapularis (black-legged tick) ex vivo organotypic cultures and applications for disease control. MBio. 2017;8 doi: 10.1128/mBio.01255-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Popov V.L., Korenberg E.I., Nefedova V.V. Ultrastructural evidence of the ehrlichial developmental cycle in naturally infected Ixodes persulcatus ticks in the course of coinfection with Rickettsia, Borrelia, and a flavivirus. Vector Borne Zoonotic Dis. 2007;7:699–716. doi: 10.1089/vbz.2007.0148. [DOI] [PubMed] [Google Scholar]
- 30.Booth T.F., Davies C.R., Jones L.D., Staunton D., Nuttall P.A. Anatomical basis of Thogoto virus infection in BHK cell culture and in the ixodid tick vector, Rhipicephalus appendiculatus. J Gen Virol. 1989;70(Pt 5):1093–1104. doi: 10.1099/0022-1317-70-5-1093. [DOI] [PubMed] [Google Scholar]
- 31.Dickson D.L., Turell M.J. Replication and tissue tropisms of Crimean-Congo hemorrhagic fever virus in experimentally infected adult Hyalomma truncatum (Acari: Ixodidae) J Med Entomol. 1992;29:767–773. doi: 10.1093/jmedent/29.5.767. [DOI] [PubMed] [Google Scholar]
- 32.Meng F., Ding M., Tan Z. Virome analysis of tick-borne viruses in Heilongjiang Province, China. Ticks Tick Borne Dis. 2019;10:412–420. doi: 10.1016/j.ttbdis.2018.12.002. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary material