Abstract
In Austria, all laboratories are legally obligated to forward human and food/environmental L. monocytogenes isolates to the National Reference Laboratory/Center (NRL) for Listeria. Two invasive human isolates of L. monocytogenes serotype 1/2a of the same pulsed-field gel electrophoresis (PFGE) pattern, previously unknown in Austria, were cultured for the first time in January 2016. Five further human isolates, obtained from patients with invasive listeriosis between April 2016 and September 2017, showed this PFGE pattern. In Austria the NRL started to use whole-genome sequencing (WGS) based typing in 2016, using a core genome MLST (cgMLST) scheme developed by Ruppitsch et al. 2015, which contains 1701 target genes. Sequence data are submitted to a publicly available nomenclature server (Ridom GmbH, Münster, Germany) for allocation of the core genome complex type (CT). The seven invasive human isolates differed from each other with zero to two alleles and were allocated to CT1234 (declared as outbreak strain). Among the Austrian strain collection of about 6,000 cgMLST-characterized non-human isolates (i.e., food/environmental isolates) 90 isolates shared CT1234. Out of these, 83 isolates were traced back to one meat processing-company. They differed from the outbreak strain by up to seven alleles; one isolate originated from the company's industrial slicer. The remaining seven CT1234-isolates were obtained from food products of four other companies (five fish-products, one ready-to-eat dumpling and one deer-meat) and differed from the outbreak strain by six to eleven alleles. The outbreak described shows the considerable potential of WGS to identify the source of a listeriosis outbreak. Compared to PFGE analysis, WGS-based typing has higher discriminatory power, yields better data accuracy, and allows higher laboratory through-put at lower cost. Utilization of WGS-based typing results of human and food/ environmental L. monocytogenes isolates by appropriate public health analysts and epidemiologists is indispensable to support a successful outbreak investigation.
Keywords: whole-genome sequencing, pulsed-field gel electrophoresis, outbreak investigation, public health laboratory capacity, public health surveillance
Introduction
Listeriosis is a relatively uncommon disease, which typically causes a severe disease in a high portion of cases and deaths in susceptible population subgroups (1, 2). Listeriosis is a foodborne illness of major public health concern because of the severity of its complications (infections of the central nervous system, septicemia, gastroenteritis and abortion), a hospitalization rate of 98.6% and a case-fatality ratio of 13,8%, as reported by the EU summary report on zoonoses, zoonotic agents and food-borne outbreaks from 2017 (3). The surveillance of listeriosis in the European Union/European Economic Area (EU/EEA) focuses on the severe invasive forms of the disease for which the risk groups are mainly elderly and immunocompromised persons, pregnant women and infants. In 2017, 2,480 confirmed cases of invasive listeriosis were reported by 28 EU/EEA countries, resulting in an overall notification rate of 0.48 per 100,000 population (3). The increasing trend in the number of listeriosis cases in the EU/EEA, probably also due to the increased population size of the elderlies (4, 5), is worrying and calls for utmost attention to be placed on the prevention and control of the disease and outbreaks. The European Center for Disease Prevention and Control (ECDC), the European Food Safety Authority (EFSA) and the European Union Reference Laboratory (EU-RL) for L. monocytogenes have set up a joint database collecting, on a voluntary basis, combined AscI/ApaI PFGE profiles for PFGE typing data for human, food, animal and environmental isolates from public heath institutes and food safety and veterinary authorities to enable detection of listeriosis outbreaks affecting several countries (6). However, technical development is evolving fast and whole genome sequencing (WGS)-based typing methods replaced pulsed-field gel electrophoresis (PFGE) as the gold-standard showing higher accuracy and a superior discriminatory power (7). The outbreak described here, illustrates impressively the considerable potential of WGS based typing to elucidate the source of a listeriosis outbreak.
Background to Outbreak Investigation
In Austria, laboratories have a legal obligation to forward human and food/environmental L. monocytogenes isolates derived from official controls as well as from ownchecks to the NRL. In January 2016, two human isolates of L. monocytogenes serotype 1/2a of the identical pulsed-field gel electrophoresis (PFGE) pattern, previously unknown in Austria, were cultured for the first time. Two environmental isolates of unknown origin, were obtained in January and February 2016, and another 24 food/environmental isolates were obtained between September 2016 and December 2017. In addition five further human isolates from patients with invasive listeriosis, isolated between April 2016 and September 2017 were obtained. All isolates showed this new PFGE pattern. The food and environmental isolates originated from six different laboratories. From January to August 2018, further 69 food/environmental isolates possibly related to the outbreak were sent to the National Reference Laboratory (NRL) for Listeria. In summary, a total of 95 non-human isolates together and seven human outbreak isolates were typed by using WGS cgMLST analysis. No reliable information was available on patients' relevant food consumption.
On 25 January 2018, Austria launched an Urgent Inquiry (UI-460) in The Epidemic Intelligence Information System (EPIS), a web-based communication platform that allows nominated public health experts to exchange technical information to assess whether current and emerging public health threats have a potential impact in the European Union. Aim of the outbreak investigation was to identify the source(s) and to recommend the appropriate public health measures for preventing further cases. Thirteen countries (Denmark, Finland, France, Germany, Ireland, Italy, Luxembourg, The Netherlands, Norway, Slovenia, Spain, Sweden and the United Kingdom) answered via the platform and eight countries reported cases with at least six allelic differences to the Austrian outbreak cluster. Raw data of the sequences were provided from the countries, which allowed a direct comparison with the Austrian database. Eight non-human strains isolated in France and the Netherlands were reported in the European Union Reference Laboratory for Listeria monocytogenes technical report [EURL Lm 2018 (8)] to form a cgMLST cluster with five to seven pairwise allele differences against the outbreak strain.
Materials and Methods
Origin of Isolates, Cultivation, and Genomic DNA Isolation
In Austria, Listeria isolates obtained from food and environmental samples, as well as human isolates, must be sent to the NRL for Listeria by legislation. The non-human isolates are anonymized, provided with unique identifier and information on the type of food matrix only (e.g., meat-product, diary-product, vegetable-product, food-environment) by the sending primary food laboratories. Isolates are cultivated on RAPID'L.Mono™ agar plates (Biorad, Munich, Germany) for species confirmation and subsequently subcultured overnight on Columbia Broth (BD Difco™, Heidelberg, Germany) for extraction of high molecular weight genomic DNA using the HMW MagAttract kit (Qiagen, Hilden, Germany) according to the instructions of the manufacturer for Gram positive bacteria.
Whole Genome Sequencing and Data Analysis
Whole genome sequencing was performed as described previously (9). Briefly, for sequencing, an Illumina MiSeq platform (Illumina Inc., San Diego, CA, USA) was used. Library preparation was carried out using Nextera XT according to the instructions of the manufacturer (Illumina Inc., San Diego, CA, USA). For assembly into draft genomes, raw reads were de novo assembled using SPAdes version 3.11.1 (10). Contigs were filtered for a minimum coverage of 5-fold and minimum length of 200 bp, which resulted in 26–187 contigs at a coverage of 46–148-fold. Classical multilocus sequence typing (MLST) data according to Ragon et al. (11) and genoserotyping data according to Hyden et al. (12) were de novo extracted from WGS sequence data. Assessment of the core genome multilocus sequence typing (cgMLST) results was done using Ridom SeqSphere+ software version 5.1.0 as described by Ruppitsch et al. (13). All isolates had 98.1–99.8% good targets and a minimum spanning tree (MST) was generated in Ridom SeqSphere+ version 5.1.0 for visualization of strain relatedness. For comparison and data harmonization SeqSphere+ results were compared to the Pasteur cgMLST scheme (7) and GenomeGraphR (14). The sequences have been deposited in DDBJ/EMBL/GenBank under the project number PRJNA434392. Raw sequence data for each strain were deposited under SRA accession numbers (Table 1).
Table 1.
Accession numbers of sequences available at NCBI Sequence Read Archive (SRA).
| ID | Accession no. | cgMLST | MLST CC | Genoserotype |
|---|---|---|---|---|
| 5F_CoA | SRR6740436 | 1234 | 155 | IIa |
| 6F_CoA | SRR6740437 | 1234 | 155 | IIa |
| 3E_CoA | SRR6740438 | 1234 | 155 | IIa |
| 4E_CoA | SRR6740439 | 1234 | 155 | IIa |
| 1E_uk | SRR6740440 | 1234 | 155 | IIa |
| 2E_uk | SRR6740441 | 1234 | 155 | IIa |
| 7H | SRR6740442 | 1234 | 155 | IIa |
| 4F_CoA | SRR6740443 | 1234 | 155 | IIa |
| 1F_CoA | SRR6740444 | 1234 | 155 | IIa |
| 2F_CoA | SRR6740445 | 1234 | 155 | IIa |
| 15F_CoA | SRR6740446 | 1234 | 155 | IIa |
| 16F_CoA | SRR6740447 | 1234 | 155 | IIa |
| 9F_CoA | SRR6740448 | 1234 | 155 | IIa |
| 10F_CoA | SRR6740449 | 1234 | 155 | IIa |
| 7F_CoA | SRR6740450 | 1234 | 155 | IIa |
| 8F_CoA | SRR6740451 | 1234 | 155 | IIa |
| 13F_CoA | SRR6740452 | 1234 | 155 | IIa |
| 14F_CoA | SRR6740453 | 1234 | 155 | IIa |
| 11F_CoA | SRR6740454 | 1234 | 155 | IIa |
| 12F_CoA | SRR6740455 | 1234 | 155 | IIa |
| 3F_CoA | SRR6740456 | 1234 | 155 | IIa |
| 28F_CoA | SRR6740457 | 1234 | 155 | IIa |
| 19F_CoA | SRR6740458 | 1234 | 155 | IIa |
| 18F_CoA | SRR6740459 | 1234 | 155 | IIa |
| 17F_CoA | SRR6740460 | 1234 | 155 | IIa |
| 4H | SRR6740461 | 1234 | 155 | IIa |
| 3H | SRR6740462 | 1234 | 155 | IIa |
| 2H | SRR6740463 | 1234 | 155 | IIa |
| 1H | SRR6740464 | 1234 | 155 | IIa |
| 6H | SRR6740465 | 1234 | 155 | IIa |
| 5H | SRR6740466 | 1234 | 155 | IIa |
| 90F_CoA | SRR8184623 | 6743 | 37 | IIa |
| 56F_CoA | SRR8185109 | 1234 | 155 | IIa |
| 55F_CoA | SRR8185110 | 1234 | 155 | IIa |
| 58F_CoA | SRR8185111 | 1234 | 155 | IIa |
| 57F_CoA | SRR8185112 | 1234 | 155 | IIa |
| 52F_CoA | SRR8185113 | 1234 | 155 | IIa |
| 51F_CoA | SRR8185114 | 1234 | 155 | IIa |
| 40F_CoA | SRR8185115 | 1234 | 155 | IIa |
| 39F_CoA | SRR8185116 | 1234 | 155 | IIa |
| 38F_CoA | SRR8185117 | 1234 | 155 | IIa |
| 37F_CoA | SRR8185118 | 1234 | 155 | IIa |
| 36F_CoA | SRR8185119 | 1234 | 155 | IIa |
| 35F_CoA | SRR8185120 | 1234 | 155 | IIa |
| 34F_CoA | SRR8185121 | 1234 | 155 | IIa |
| 33F_CoA | SRR8185122 | 1234 | 155 | IIa |
| 32F_CoA | SRR8185123 | 1234 | 155 | IIa |
| 31F_CoA | SRR8185124 | 1234 | 155 | IIa |
| 72F_CoA | SRR8185125 | 1234 | 155 | IIa |
| 71F_CoA | SRR8185126 | 1234 | 155 | IIa |
| 74F_CoA | SRR8185127 | 1234 | 155 | IIa |
| 78F_CoA | SRR8185128 | 1234 | 155 | IIa |
| 76F_CoA | SRR8185129 | 1234 | 155 | IIa |
| 27F_nonCoA | SRR8185130 | 1234 | 155 | IIa |
| 77F_CoA | SRR8185131 | 1234 | 155 | IIa |
| 73F_CoA | SRR8185132 | 1234 | 155 | IIa |
| 54F_CoA | SRR8185133 | 1234 | 155 | IIa |
| 53F_CoA | SRR8185134 | 1234 | 155 | IIa |
| 89F_CoA | SRR8185135 | 1234 | 155 | IIa |
| 69F_CoA | SRR8185136 | 1234 | 155 | IIa |
| 70F_CoA | SRR8185137 | 1234 | 155 | IIa |
| 65F_CoA | SRR8185138 | 1234 | 155 | IIa |
| 66F_CoA | SRR8185139 | 1234 | 155 | IIa |
| 67F_CoA | SRR8185140 | 1234 | 155 | IIa |
| 68F_CoA | SRR8185141 | 1234 | 155 | IIa |
| 61F_CoA | SRR8185142 | 1234 | 155 | IIa |
| 62F_CoA | SRR8185143 | 1234 | 155 | IIa |
| 63F_CoA | SRR8185144 | 1234 | 155 | IIa |
| 64F_CoA | SRR8185145 | 1234 | 155 | IIa |
| 47F_CoA | SRR8185146 | 1234 | 155 | IIa |
| 48F_CoA | SRR8185147 | 1234 | 155 | IIa |
| 45F_CoA | SRR8185148 | 1234 | 155 | IIa |
| 46F_CoA | SRR8185149 | 1234 | 155 | IIa |
| 43F_CoA | SRR8185150 | 1234 | 155 | IIa |
| 44F_CoA | SRR8185151 | 1234 | 155 | IIa |
| 41F_CoA | SRR8185152 | 1234 | 155 | IIa |
| 42F_CoA | SRR8185153 | 1234 | 155 | IIa |
| 60F_CoA | SRR8185154 | 1234 | 155 | IIa |
| 49F_CoA | SRR8185155 | 1234 | 155 | IIa |
| 50F_CoA | SRR8185156 | 1234 | 155 | IIa |
| 59F_CoA | SRR8185157 | 1234 | 155 | IIa |
| 83F_CoA | SRR8185158 | 5753 | 517 | IIb |
| 29F_CoA | SRR8185159 | 6252 | 155 | IIa |
| 25F_nonCoA | SRR8185160 | 1234 | 155 | IIa |
| 24F_nonCoA | SRR8185161 | 1234 | 155 | IIa |
| 75F_CoA | SRR8185162 | 1234 | 155 | IIa |
| 26F_nonCoA | SRR8185163 | 1234 | 155 | IIa |
| 21F_nonCoA | SRR8185164 | 1234 | 155 | IIa |
| 5E_CoA | SRR8185165 | 1234 | 155 | IIa |
| 23F_nonCoA | SRR8185166 | 1234 | 155 | IIa |
| 22F_nonCoA | SRR8185167 | 1234 | 155 | IIa |
| 81F_CoA | SRR8185168 | 6399 | 451 | IIa |
| 82F_CoA | SRR8185169 | 6424 | 1 | IVb |
| 87F_CoA | SRR8185170 | 1234 | 155 | IIa |
| 84F_CoA | SRR8185171 | 1234 | 155 | IIa |
| 79F_CoA | SRR8185172 | 1234 | 155 | IIa |
| 88F_CoA | SRR8185173 | 1234 | 155 | IIa |
| 80F_CoA | SRR8185174 | 1234 | 155 | IIa |
| 85F_CoA | SRR8185175 | 1234 | 155 | IIa |
| 30F_CoA | SRR8185176 | 1234 | 155 | IIa |
| 86F_CoA | SRR8185177 | 1234 | 155 | IIa |
| 20F-CoA | SRR8186973 | 1234 | 155 | IIa |
BioProject ID: PRJNA434392; Title: LIST_2018_CT1234. E_uk, environmental isolate of unknown origin (at the time of submission); E_CoA, environmental isolate, company A-associated; F_CoA, food-isolate, company A-associated; H, human isolate.
SNP analysis was done with GenomeGraphR Beta 2.7 [Sanaa et al. (14)] using the default settings and a cluster threshold definition of 12 SNPs. All strains were compared with the isolates present in the database.
Results
Austria reported a suspected outbreak due to L. monocytogenes serotype 1/2a of the same PFGE pattern, including seven patients of invasive listeriosis, having occurred in eastern Austria between 2015 and 2017. The cgMLST typing of the seven human invasive isolates revealed a genetically tight cluster, complex type 1234 (CT1234), which corresponds to CT1170 of Institut Pasteur cgMLST scheme [Moura et al. (7)], with zero to two allelic differences from each other. SNP analysis revealed that our clinical isolates differed from each other by 1–4 SNPs. In addition, the closest clinical strain clustering with our isolates differed by 11–12 SNPs and therefore confirmed the CT1234.
On 26 January 2018, the Austrian Ministry of Health mandated the Austrian Agency for Health and Food Safety (AGES) to investigate this suspected outbreak. A confirmed outbreak case was defined as a patient with invasive listeriosis, positive for L. monocytogenes cgMLST CT1234 isolate, which differed by ≤ 2 alleles from a representative outbreak isolate by using cgMLST, and with a disease onset on or after 1 January 2015.
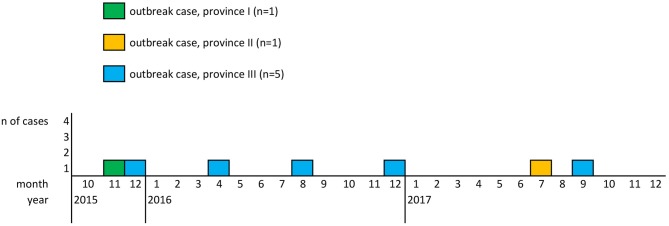
The aforementioned seven patients fulfilled the definition of a confirmed outbreak case. Patients were 29–97 years old (mean: 68; median: 73), five females and two males, with disease onset between November 2015 and September 2017 and residence in three of the nine Austrian provinces. Figure 1 depicts the outbreak cases by month of diagnosis and province of residence.
Figure 1.
Outbreak cases of L. monocytogenes CT1234 by month of disease onset and province of residence, 2015–2017, Austria.
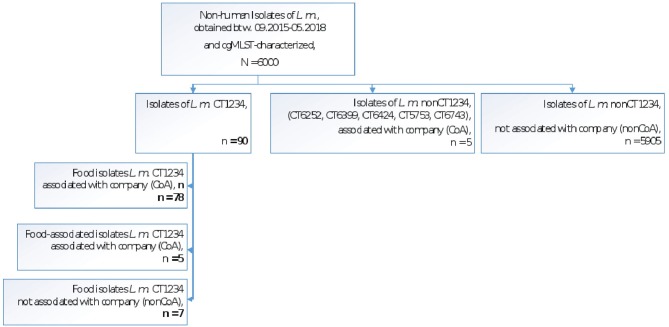
Among the Austrian genome database of about 6,000 non-human isolates (i.e., food/environmental isolates, collected between 2015 and 2018), 90 isolates shared genoserotype IIa, MLST CC155, and cgMLST CT1234. Out of these, 83 isolates were traced back to a meat-processing company (companyA; CoA) in eastern Austria. These food/environmental isolates differed from the outbreak strain by zero to seven alleles and one isolate, originated from the company's industrial slicer. The remaining seven CT1234 isolates were obtained from food products of four other companies (five fish-products, one ready-to-eat dumpling and deer-meat product each) and differed from the main outbreak strain by six to eleven alleles.
Figure 2 illustrates the non-human isolates of the AGES Listeria strain collection by CT1234 allocation and linkage to the meat processing company A.
Figure 2.
Non-human L. monocytogenes isolates of the AGES strain collection (L.m. CT1234, L.m. nonCT1234) with and without company A linkage (CoA, nonCoA).
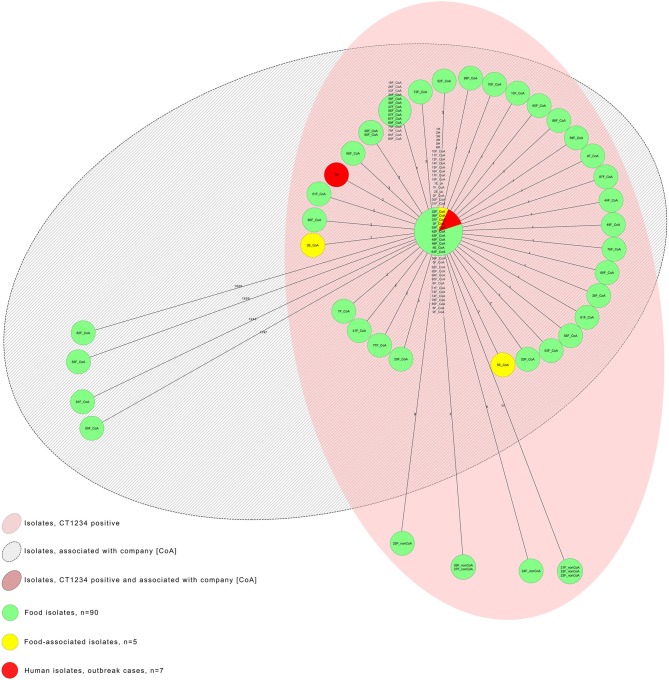
Figure 3 illustrates the MST of the seven human outbreak isolates, of food and environmental isolates of L. monocytogenes, comprising company A associated and non-associated CT1234-isolates (CoA, nonCoA), and company A associated, nonCT1234-isolates.
Figure 3.
Adapted minimum spanning tree of the seven human outbreak isolates, the food and environmental isolates of L. monocytogenes, including the company A associated and non-associated CT1234-isolates (CoA CT1234-isolates and nonCoA CT1234-isolates; n = 90), and the five company A associated nonCT1234-isolates; N = 102.
In November 2017, a 47-year old male developed signs and symptoms compatible with a non-invasive listeriosis (i.e., febrile gastroenteritis) 7 h after consumption of a pizza with sliced ham topping in a restaurant in the Austrian province Tyrol. No patient isolate of L.monocytogenes was available. The official sample taken from the sliced pizza ham at the restaurant tested positive for L. monocytogenes CT1234 with one allelic difference from the outbreak strain (Figure 3: 28F_CoA). Trace-back analyses identified the origin of the ham pizza topping from meat-processing company A.
Public Health Measures
Company A implemented control measures including intensified environmental disinfection, installation of a new slicer and continuous investigation of environmental swabs and newly processed food products for Listeria. All food batches had to be negative for L. monocytogenes before being released to the market. During these activities, further four strains of L. monocytogenes were found in the tested food products of the company. There were of complex types CT6252 (genoserotype IIa, MLST CC155), CT6399 (genoserotype IIa, MLST CC451), CT6424 (genoserotype IVb, MLST CC1), CT6743 (genoserotype IIa, MLST CC37), different from the outbreak CT, CT1234 (Figure 3: food isolates 81F, 82F, 83F, and 90F). After August 2018, the public health authorities found no further L. monocytogenes positive food products.
Discussion
Investigation of listeriosis outbreaks is difficult due to the multitude of possible food vehicles including a broad range of ready-to-eat foods. Pulsed-field gel electrophoresis (PFGE) was the gold standard for strain typing (6) but has become obsolete with the advent of WGS. WGS is highly discriminatory and superior for allocating listeriosis cases to an outbreak (7, 13). Due to this superiority of WGS it is time to stop PFGE (Pulsenet Network resolution, ECDC). A dictionary between PFGE and MLST will allow to screen the PFGE database for previous strains using WGS specific ST or CC (15). However, with the limitations of current WGS technology we cannot create PFGE patterns from WGS data and therefore cannot create a PFGE-WGS dictionary. Despite the availability of technical literature on methods for outbreak investigations, there are no pre-specified formulae to dictate the path that an outbreak investigation is supposed to take (16). Investigations of listeriosis outbreaks provide a unique opportunity to gain new scientific knowledge on the occurrence of L. monocytogenes in the food-processing setting.
In contrast to Europe, the United States have a zero tolerance policy for L. monocytogenes in ready-to-eat foods (17). Commission Regulation (EC) No 2073/2005 of 15 November 2005 on microbiological criteria for foodstuffs requires a limitation of < 100 CFU/g in ready-to-eat food products able to support the growth of L. monocytogenes (other than those intended for infants and for special medical purposes) during the shelf life, when products are placed on the market; these particular food products must be tested for the absence of L. monocytogenes in 25 g before leaving the immediate control of the producing food business operator. Otherwise a challenge test which ensures that the limit of 100 cfu/g at the end of shelf-life will not be exceeded, has to be shown (18). Although the infective dose of L. monocytogenes is unknown and population subgroups differ in vulnerability to Listeria monocytogenes in food, the present European legislation should be sufficient in controlling foodborne listeriosis.
As a consequence of an earlier outbreak of listeriosis in Austria (19, 20), the Federal Ministry of Health had classified 600 food-producing facilities as being at high risk of Listeria contamination and ordered the provinces to conduct inspections on various control measures in a Key Activity Action Campaign entitled “Schwerpunktaktion SPA-A-600” in 2014. The province to which company A was assigned had neglected to complete this requirement. From 2015 to 2017, only one official sample of sliced bacon was obtained on 22 November 2017 by the local food authority at the meat processing company A. The final report, outlining the presence of L. monocytogenes in numbers below 100 CFU/g, was not issued until 1 February 2018. Surprisingly, no challenge test was performed for this food-product, especially considering that a similar outbreak caused by the consumption of bacon that was contaminated with L. monocytogenes caused four fatalities in Bavaria. At the time in 2016, this outbreak led to a public recall and public warning in Austria (21).
For two decades, PFGE was the reference method for L. monocytogenes surveillance and outbreak investigation (2, 22). It is still used for screening but is increasingly replaced by WGS based typing methods (7, 14, 21, 23–26). WGS based typing outperforms PFGE with respect to the discriminatory power, information content, throughput, reproducibility, costs and inter-laboratory data exchange. However, it is important to keep in mind that the differences between the cgMLST schemes of Moura et al. and Ruppitsch et al. can have an impact on the cluster detection (24). For communication on detected clusters it is important to know which core genome scheme, assembler, and assembler version and sequencing technology was used and which average sequencing coverage was achieved.
Based on the current cgMLST analysis of the human outbreak isolates, a difference from each other by only zero to two alleles, and of the majority of the outbreak associated food-isolates, a difference by zero to four alleles should be considered to increase the specificity of linking isolates to L. monocytogenes outbreaks. WGS data not only allow to infer phylogenetic relationships but also to filter for additional information like serotypes (12), virulence- and resistance-genes (7).
Although it is known that SNP analysis provides maximal discriminatory power, results are difficult to standardize and interpret (27). Moreover, the analysis based on single nucleotide polymorphisms (SNPs) showed here identical results to the ones obtained by cgMLST. Expansion of the classical MLST principle to a genome wide gene-by-gene comparison allowed the establishment of databases based on well-defined core genome or whole genome MLST schemes (7, 13, 24, 28). The setup of open accessible databases (Listeria monocytogenes cgMLST at https://www.cgmlst.org/ncs/schema/690488/, BIGSdb-Lm at http://bigsdb.pasteur.fr/listeria) allows the comparison and sharing of data between public health laboratories worldwide and facilitates international source tracking and multinational outbreak investigation (29, 30). These new WGS databases, although only 3 years old, already harbor nearly twice the number of strain complex types (CT) than the >20 year old PulseNet PFGE database demonstrating again the higher discriminative power of WGS based typing. Compared to PFGE, these major improvements in L. monocytogenes typing allow a faster and more discriminative detection of clusters and reduce unnecessary epidemiological investigations. L. monocytogenes is one of the pathogens for which a rapid transition from traditional typing methods to WGS-based typing methods is presently occurring in the public health laboratories of the EU/EEA as well as the PulseNet International network, and it is the first food- and waterborne pathogen for which a comprehensive WGS-assisted real-time surveillance is planned to be established at the EU/EEA level (31). Due to the superiority of WGS for real-time surveillance in a One Health approach, the PulseNet International network and EU/EEA health and food safety authorities move to cgMLST and wgMLST analysis (24, 30).
Conclusions
Compared to PFGE analysis, WGS based typing has a higher discriminatory power, yields better data accuracy, and allows higher laboratory through-put at lower cost, as proven in the current outbreak investigation (26). The meaningful use of WGS based typing data for a successful investigation of a listeriosis outbreak and the appropriate public health measures, requires intense collaboration between the public health and food safety authorities, food microbiologists, typing experts and epidemiologists.
Author Contributions
AP, DS, FA, and WR contributed conception and design of the study, data analysis and interpretation and writing of the manuscript. AM, AL, AS, AC, SM, and SL performed data analysis. SH and BS contributed to the interpretation of data of the work and critically revised the content of the study. All authors contributed to manuscript revision, read and approved the submitted version.
Conflict of Interest Statement
We declare that none of the authors have any commercial and financial relationship to the company Ridom GmbH (Münster, Germany), developer of the Ridom SeqSphere+ software, mentioned in the manuscript. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
Funding. Part of the sequencing-work was funded by a grant awarded under the One Health European Joint Programme (OHEJP) JRP7 LISTADAPT.
References
- 1.Allerberger F, Wagner M. Listeriosis: a resurgent foodborne infection. Clin Microbiol Infect. (2010) 16:16–23. 10.1111/j.1469-0691.2009.03109.x [DOI] [PubMed] [Google Scholar]
- 2.Allerberger F, Bagó Z, Huhulescu S, Pietzka A. Listeriosis: the dark side of refrigeration and ensiling. In: Sing A, editor. Zoonoses - Infections Affecting Humans and Animals. Focus on Public Health Aspects. Heidelberg, DE: Springer Verlag; (2015), p. 249–86. [Google Scholar]
- 3.EFSA and ECDC (European Food Safety Authority and European Centre for Disease Prevention and Control) The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2017. EFSA J. (2018) 16:5500 10.2903/j.efsa.2018.5500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.European Centre for Disease Prevention and Control . Listeria. In: ECDC Annual epidemiological report for 2016. Stockholm: European Centre for Disease Prevention and Control; (2016). [Google Scholar]
- 5.EFSA Panel on Biological Hazards. Ricci A, Allende A, Bolton D, Chemaly M, Davies R, et al. Listeria monocytogenes contamination of ready-to-eat foods and the risk for human health in the EU. EFSA J. (2018) 16:5134 10.2903/j.efsa.2018.5134 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Félix B, Danan C, Van Walle I, Lailler R, Texier T, Lombard B, et al. Building a molecular Listeria monocytogenes database to centralize and share PFGE typing data from food, environmental and animal strains throughout Europe. J Microbiol Methods. (2014) 104:1–8. 10.1016/j.mimet.2014.06.001 [DOI] [PubMed] [Google Scholar]
- 7.Moura A, Criscuolo A, Pouseele H, Maury MM, Leclercq A, Tarr C, et al. Whole genome-based population biology and epidemiological surveillance of Listeria monocytogenes. Nat Microbiol. (2016) 2:16185. 10.1038/nmicrobiol.2016.185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.European Union Reference Laboratory for Listeria monocytogenes technical reports to EFSA and DG SANTE related to UI-460 ST155 Technical report ST155 UI-460 multicountry outbreak of Listeria monocytogenes Version 1, Foodborne pathogens department, Laboratory for Food Safety, Anses, Salmonella Et Listeria. (2018). [Google Scholar]
- 9.Lepuschitz S, Mach R, Springer B, Allerberger F, Ruppitsch W. Draft genome sequence of a community-acquired methicillin-resistant Staphylococcus aureus USA300 Isolate from a River Sample. Genome Announce. (2017) 5:e01166–17. 10.1128/genomeA.01166-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nurk S, Bankevich A, Antipov D, Gurevich AA, Korobeynikov A, Lapidus A, et al. Assembling genomes and mini-metagenomes from chimeric MDA products. J Comput Biol. (2013) 20:714–37. 10.1089/cmb.2013.0084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ragon M, Wirth T, Hollandt F, Lavenir R, Lecuit M, Le Monnier A, & Brisse S. A new perspective on Listeria monocytogenes evolution. PLoS Pathog. 4:e1000146. 10.1371/journal.ppat.1000146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hyden P, Pietzka A, Lennkh A, Murer A, Springer B, Blaschitz M, et al. Whole genome sequence-based serogrouping of Listeria monocytogenes isolates. J Biotechnol. (2016) 235:181–6. 10.1016/j.jbiotec.2016.06.005 [DOI] [PubMed] [Google Scholar]
- 13.Ruppitsch W, Pietzka A, Prior K, Bletz S, Fernandez HL, Allerberger F, et al. Defining and evaluating a core genome multilocus sequence typing scheme for whole-genome sequence-based typing of Listeria monocytogenes. J Clin Microbiol. (2015) 53:2869–76. 10.1128/JCM.01193-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sanaa M, Pouillot R, Vega FG, Strain E, Van Doren JM. GenomeGraphR: a user-friendly open-source web application for foodborne pathogen whole genome sequencing data integration, analysis, and visualization. PLoS ONE. (2019) 14:e0213039. 10.1371/journal.pone.0213039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Maury MM, Tsai YH, Charlier C, Touchon M, Chenal-Francisque V, Leclercq A, et al. (2016). Uncovering Listeria monocytogenes hypervirulence by harnessing its biodiversity. Nat Genet. 48:308–313. 10.1038/ng.3501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jamasji-Pavri A. Transaction cost economics and principal-agent theory: insights into investigations of outbreaks of infectious diseases. In: Roberts JA, editor. The Economics of Infectious Disease. Oxford: Oxford University Press, (2006) p. 261–79. [Google Scholar]
- 17.Montville TJ, Matthews KR. Food Microbiology: An Introduction. 2nd ed. Washington DC: ASM Press, (2008). [Google Scholar]
- 18.European Commission. Commission Regulation (EC) No 2073/2005 of 15 November 2005 on microbiological criteria for foodstuffs. Official Journal. (2005) 338:1–26. [Google Scholar]
- 19.Fretz R, Pichler J, Sagel U, Much P, Ruppitsch W, Pietzka AT, et al. Update: multinational listeriosis outbreak due to 'Quargel', a sour milk curd cheese, caused by two different L. monocytogenes serotype 1/2a strains, 2009-2010. Euro Surveill. (2010) 15:19543. 10.2807/ese.15.16.19543-en [DOI] [PubMed] [Google Scholar]
- 20.Pichler J, Appl G, Pietzka A, Allerberger F. Lessons to be Learned from an Outbreak of Foodborne Listeriosis, Austria 2009–2010. Food Prot Trends. (2011) 31:268–73. [Google Scholar]
- 21.Ruppitsch W, Prager R, Halbedel S, Hyden P, Pietzka A, Huhulescu S. Ongoing outbreak of invasive listeriosis, Germany, 2012 to 2015. Euro Surveill. (2015) 20:30094. 10.2807/1560-7917.ES.2015.20.50.30094 [DOI] [PubMed] [Google Scholar]
- 22.Swaminathan B, Barrett TJ, Hunter SB, Tauxe RV, & CDC. PulseNet Task Force PulseNet: the molecular subtyping network for foodborne bacterial disease surveillance. United States. Emerg Infect Dis. 7:382–9. 10.3201/eid0703.010303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen Y, Gonzalez-Escalona N, Hammack TS, Allard MW, Strain EA, Brown EW. Core genome multilocus sequence typing for identification of globally distributed clonal groups and differentiation of outbreak strains of Listeria monocytogenes. Appl Environ Microb. (2016) 82:6258–72. 10.1128/AEM.01532-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Van Walle I, Björkman JT, Cormican M, Dallman T, Mossong J, Moura A, et al. Retrospective validation of whole genome sequencing-enhanced surveillance of listeriosis in Europe, 2010 to 2015. Euro Surveill. (2018) 23:1700798. 10.2807/1560-7917.ES.2018.23.33.1700798 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schmid D, Allerberger F, Huhulescu S, Pietzka A, Amar C, Kleta S, et al. Whole genome sequencing as a tool to investigate a cluster of seven cases of listeriosis in Austria and Germany, 2011-2013. Clin Microbiol Infect. (2014) 20:431–6. 10.1111/1469-0691.12638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Reimer A, Weedmark K, Petkau A, Peterson C, Walker M, Knox N, et al. Shared genome analyses of notable listeriosis outbreaks, highlighting the critical importance of epidemiological evidence, input datasets and interpretation criteria. Microb Genom. (2019) 5: 237. 10.1099/mgen.0.000237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pightling AW, Pettengill JB, Luo Y, Baugher JD, Rand H, Strain E. Interpreting whole-genome sequence analyses of foodborne bacteria for regulatory applications and outbreak investigations. Front Microbiol. (2018) 10:1482 10.3389/fmicb.2018.01482 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Moura A, Tourdjman M, Leclercq A, Hamelin E, Laurent E, Fredriksen N, et al. Real-Time Whole-Genome Sequencing for Surveillance of Listeria monocytogenes, France. Emerg Infect Dis. 23:1462–70. 10.3201/eid2309.170336 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Schjørring S, Gillesberg Lassen S, Jensen T, Moura A, Kjeldgaard JS, Müller L, et al. Cross-border outbreak of listeriosis caused by cold-smoked salmon, revealed by integrated surveillance and whole genome sequencing (WGS), Denmark and France, 2015 to 2017. Euro Surveill. (2017) 22:17–00762. 10.2807/1560-7917.ES.2017.22.50.17-00762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nadon C, Van Walle I, Gerner-Smidt P, Campos J, Chinen I, Concepcion-Acevedo J, et al. PulseNet International: vision for the implementation of whole genome sequencing (WGS) for global food-borne disease surveillance. Euro Surveill. (2017) 22:30544. 10.2807/1560-7917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Revez J, Espinosa L, Albiger B, Leitmeyer KC, Struelens MJ. ECDC microbiology focal points and experts group. Survey on the use of whole-genome sequencing for infectious diseases surveillance: rapid expansion of European national capacities, 2015-2016. Front Public Health. (2017) 5:347 10.3389/fpubh.2017.00347 [DOI] [PMC free article] [PubMed] [Google Scholar]