Abstract
Background & objectives:
Klebsiella pneumoniae (KP), a common cause of invasive infections, is often extensively drug resistant in India. At present, studies on resistance mechanism and clonal relationship of KP from India are limited. The present study was undertaken to determine the resistance mechanism and clonal relationship of colistin-resistant isolates obtained from various specimens. Carbapenemases were also determined since the isolates were carbapenem resistant.
Methods:
Sixty five isolates from blood, exudates and respiratory specimens collected between 2016 and 2017 were studied. Colistin minimum inhibitory concentration (MIC) was performed by broth-micro dilution method. Multiplex PCR was carried out to determine carbapenemases. Targeted sequencing was performed to determine mutations in mgrB, phoP, phoQ and multilocus sequence typing was performed to determine the prevalent clones.
Results:
Colistin MIC ranged from 4 to 256 μg/ml. SHV, TEM and CTX-M were co-produced in 60 per cent and OXA48-like in 71 per cent. Thirteen isolates had mutations in mgrB. Mutations included a premature stop codon at 21st amino acid, the presence of insertion sequences such as IS903, ISKpn14 and ISKpn26; and elongation of mgrB. Novel mutations were also observed among phoP and phoQ genes. Colistin resistance due to mcr genes was absent. Fifteen clonal types were seen with ST231, ST14 and ST2096 being predominant.
Interpretation & conclusions:
This study revealed the changing trend of carbapenem resistance mechanism predominantly to OXA48-like from NDM. Known mgrB mutations and novel mutations in phoP and phoQ were detected. There was no plasmid-mediated colistin resistance. ST14 and ST231 were international clones associated with carbapenem resistance. Colistin-resistant KP was of diverse clones with predominantly ST231, ST14 and ST2096.
Keywords: Colistin resistance, India, Klebsiella pneumoniae, mgrB, multilocus sequence typing
Klebsiella pneumoniae (KP) causes a wide spectrum of infections including pneumonia, bacteraemia and skin and soft-tissue infections. The incidence of extensively drug-resistant (XDR, non-susceptible to at least one agent in all classes, but susceptible to ≤2 classes of antimicrobials) isolates have been on the rise1. Colistin resistance is mediated by chromosomal mutations and plasmid-borne genes2. Chromosomal mutations are predominantly reported in the genes encoding two-component systems (mgrB, phoP, phoQ, pmrA, pmrB) involved in lipopolysaccharide modification2. Plasmid-mediated mcr1-5 and its variants encoding phosphoethanolamine transferase contribute to colistin resistance.
At present, studies on molecular characterization of XDR KP (carbapenem and colistin resistant) and clonal relatedness are limited in India. This study was performed to determine the molecular resistance mechanisms and its association with clonal relationship among colistin-resistant KP. Chromosomal colistin resistance contributed by mutations in mgrB, phoP and phoQ and plasmid-mediated mcr-1 and mcr-3 were investigated. Multilocus sequence typing (MLST) to look for clonal types and its relationship with colistin-resistant mechanism were determined in isolates collected from various clinical specimens. Since the study isolates were carbapenem-resistant, carbapenemases were also characterized.
Material & Methods
Totally 65 colistin-resistant KP isolated from clinical specimens including blood and body fluids (n=43), exudates (n=11) and respiratory specimens (n=11) collected between 2016 and 2017 at the department of Clinical Microbiology, Christian Medical College, Vellore, India, were selected retrospectively. The study protocol was approved by the Institutional Review Board. The isolates were identified up to species level as per standard procedures3. First colistin-resistant KP isolate from a patient was included in the study.
Phenotypic characterization: Antimicrobial susceptibility testing was performed as per Clinical and Laboratory Standards Institute (CLSI) guidelines and interpreted based on CLSI 2016 and 2017 breakpoints4,5. The panel of antimicrobials tested included ceftazidime (30 μg), cefotaxime (30 μg), cefepime (30 μg), piperacillin/tazobactam (100/10 μg), imipenem (10 μg), meropenem (10 μg), ciprofloxacin (5 μg), gentamicin (10 μg), netilmicin (30 μg), amikacin (30 μg), minocycline (30 μg) and tigecycline (15 μg). All the isolates were carbapenem resistant which was defined as resistant to meropenem with zone diameter of ≤19 mm4,5. Escherichia coli ATCC 25922, Staphylococcus aureus ATCC 25923 and Pseudomonas aeruginosa ATCC 27853 were used as control strains for susceptibility testing as recommended by CLSI4,5. Susceptibility to tigecycline was interpreted with FDA breakpoints (http://www.accessdata.fda.gov/drugsatfda_docs/label/2009/021821s016lbl.pdf). Colistin minimum inhibitory concentration (MIC) was determined by reference broth-micro dilution method6,7. mcr-1 positive E. coli (Courtesy: Dr Olga Perovic, National Institute for Communicable Diseases, Johannesburg, South Africa) was used as positive control and E. coli ATCC 25922 and P. aeruginosa ATCC 27853 were used as negative controls. MIC results were interpreted as per European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines 2016 and 2017, respectively6,7.
Molecular characterization
Characterization of β-lactamases: Bacterial DNA was extracted from 18 h cultures by Qiasymphony (Qiagen, Hilden, Germany) as per manufacturer's instructions. Multiplex PCR was performed for the detection of genes encoding β-lactam resistance which included EBSL (blaSHV, blaTEM, blaPER, blaVEB, blaCTX-M-1, blaCTX-M-2, blaCTX-M-8, blaCTX-M-9 and blaCTX-M-25)8,9, AmpC (blaACT, blaACC, blaCIT, blaDHA and blaFOX)10 and carbapenemases (blaNDM, blaVIM, blaIMP, blaOXA48-like, blaKPC, blaGES and blaSPM)11,12,13. Known positive controls (Courtesy: International Health Management Association Inc., USA) were used.
Characterization of colistin resistance: Chromosomal mutations: PCR for amplification of mgrB, phoP and phoQ genes was performed as described by Jayol et al14. From the amplified product, mutations were determined by targeted sequencing. For analysis of mutations, KP ATCC 35657 was used as reference. To determine insertion sequences present in mgrB, the sequences were run through ISfinder (https://www-is.biotoul.fr/) and the position of insertion was assigned by aligning with reference mgrB. Sequencing was performed using 3500 Genetic Analyzer (Applied Biosystems, USA) as per manufacturer's instructions.
Sorting Intolerant From Tolerant (SIFT) scores for the non-synonymous mutations implicating in the amino acid substitutions were predicted for mgrB, phoP and phoQ genes. A value of <0.05 for a substitution is predicted to be deleterious and affects the function of the protein (https://ionreporter.thermofisher.com/ionreporter/help/GUID-2097F236-C8A2-4E67-862D-0FB5875979AC.html). SIFT score calculates the deleterious effect based on the probability of the substituted amino acid being tolerated at a position.
Plasmid-mediated colistin resistance: To determine plasmid-borne colistin resistance, PCR was performed for mcr-1 and mcr-3 genes, as controls were available for these two mcr variants15,16.
Multilocus sequence typing (MLST): MLST was performed as described by Diancourt et al17. Seven housekeeping genes used to determine clonal type among KP included gapA (coding for glyceraldehyde-3-phosphate), infB (coding for translation initiation factor-2), mdh (coding for malate dehydrogenase), pgi (coding for phosphoglucose isomerase), phoE (coding for phosphorine E), rpoB (coding for β-subunit of RNA polymerase B) and tonB (coding for periplasmic energy transducer). The sequence type was assigned by determining the allele number for each of the housekeeping genes using the database maintained by Pasteur Institute, Paris, France (http://bigsdb.pasteur.fr/klebsiella/).
Genetic analysis: The relatedness of the predicted sequence types was investigated by eBURST V3 software employing the BURST algorithm18. Splits Tree4 programme version 4.14.6 was used to determine the phylogenetic relatedness among the various clonal types based on the seven house-keeping genes that define the MLST19. Using the concatenated sequences of seven housekeeping genes of MLST, phylogenetic tree was constructed using MEGA v.7 (https://www.megasoftware.net/) and meta-data were added using iTOL (https://itol.embl.de/login.cgi).
Results
All the 65 isolates were XDR being resistant to aminoglycosides, cephalosporins and fluoroquinolones and susceptible only to tigecycline; 10 blood isolates were susceptible to minocycline and two isolates each from exudates were susceptible to chloramphenicol and trimethoprim-sulphamethoxazole, respectively. The colistin MIC for the isolates ranged from 4 to 256 μg/ml. The mean colistin MIC value was found to be 19±31.9 μg/ml.
Among the ESBL genes, 60 per cent (n=40) of the isolates co-produced blaSHV, blaTEM and blaCTX-M-15. blaOXA48-like was the predominant carbapenemase gene produced in 71 per cent (n=47) followed by co-production of blaNDM and blaOXA48-like in 11 per cent (n=7), blaNDM in seven per cent (n=5) and blaKPC three per cent (n=2) and four isolates did not produce any of the carbapenemases genes tested. Fig. 1 shows the distribution of β-lactamases among the various clinical specimens.
Fig. 1.

Distribution of β-lactamases in colistin-resistant Klebsiella pneumoniae among various clinical specimens.
Colistin MIC, mutations in mgrB, phoP and phoQ and associated clonal types among the study isolates are shown in the Table. The mutations in mgrB observed among clinical specimens were among diverse clonal types. Fig. 2A–E shows the association of mgrB with insertion sequences. A total of 41 isolates did not have mutations in mgrB and in 11 isolates mgrB gene was absent. Mutations in phoP and phoQ were present among six and nine isolates, respectively. Screening for mcr-1 and mcr-3 genes by PCR was negative for all the isolates.
Table.
Mutations in mgrB, phoP and phoQ genes among the colistin-resistant Klebsiella pneumoniae
| Isolate ID | Source | Colistin MIC (µg/ml) | MLST | Carbapenemase | mgrB | phoP | phoQ | Accession number |
|---|---|---|---|---|---|---|---|---|
| Kp1 | Blood | 16 | ST86 | Nil | Nil | Alanine 114 arginine (0.22) | Leucine 209 cysteine (0.00*) | bPAW96459.1 cNRQZ01000030.1 |
| Kp2 | Blood | 32 | ST11 | Nil | Elongated mgrB of 55 amino acids | Nil | Nil | aPBD34554.1 |
| Kp3 | Blood | 16 | ST14 | OXA48-like | Nil | Nil | Tryptophan 161 leucine (0.30) | cMH211110 |
| Kp4 | Blood | 32 | ST14 | OXA48, NDM | No amplification | Nil | Tryptophan 161 leucine (0.30) | cMH211111 |
| Kp5 | Blood | 32 | ST14 | OXA48-like | Truncated by IS1R of IS1 family | Nil | Nil | aMH590725 |
| Kp6 | Blood | 8 | ST231 | OXA48-like | Premature stop codon at 21st amino acid and change in the sequence of protein after 10th amino acid | Nil | Nil | aMH807824 |
| Kp7 | Blood | 8 | ST15 | KPC | Truncated by IS903B group of IS5 family | Nil | Nil | aMH590724 |
| Kp8 | Blood | 16 | ST231 | OXA48-like | Truncated by ISKpn14 of IS1 family | Nil | Nil | aMH590726 |
| Kp9 | CSF | 16 | ST14 | OXA48-like | Truncated by ISKpn26 of IS5 family | Nil | Changes from 218 to 239 and 436 to 448 amino acids | aMH590727 cMH807825 |
| Kp10 | Exudate | 8 | ST23 | NDM | Nil | Nil | Glycine 117 aspartic acid (1.00) | cMH211112 |
| Kp11 | Exudate | 4 | ST2096 | OXA48-like | Nil | Nil | Valine 370 glutamic acid (0.03*) | cMH211113 |
| Kp12 | Exudate | 32 | ST14 | OXA48, NDM | Cysteine to glycine at 28th amino acid (0.00*) | Threonine 151 Alanine (0.00*) | Nil | aMH337365 bMH211116 |
| Kp13 | Exudate | 16 | ST147 | OXA48, NDM | Nil | Glutamic acid 22 lysine (0.49) | Nil | bMH211117 |
| Kp14 | Exudate | 32 | ST231 | OXA48-like | Premature stop codon at 21st amino acid and change in the sequence of protein after 10th amino acid | Nil | Nil | aMH807823 |
| Kp15 | Respiratory | 32 | ST14 | NDM | Cysteine to glycine at 28th amino acid (0.00*) | Nil | Leucine 172 glutamine (0.01*); Tryptophan 182 serine (0.04*) | aMH337366 cMH211114 |
| Kp16 | Respiratory | 8 | ST231 | OXA48-like | Nil | Nil | Valine444^, phenylalanine 445 glycine (0.00*) | cMH211115 |
| Kp17 | Respiratory | 8 | ST147 | OXA48-like | Methionine to arginine first amino acid (0.00*) | Nil | Nil | aMH590728 |
| Kp18 | Respiratory | 16 | ST231 | OXA48-like | Premature stop codon at 21st amino acid and change in the sequence of protein after 10th amino acid | Nil | Nil | aMH337364 |
| Kp19 | Respiratory | 8 | ST2957 | KPC-9 | No mutation; associated with IS102 of IS5 family | Nil | Proline 424 leucine (0.37) V446W (0.00*), change from 448th amino acid | aPAT25546.1 cCJ307_16160 |
amgrB accession number; bphoP accession number; cphoQ accession number. Numbers in parenthesis indicate the SIFT score; values with *<0.05 show that the mutation affects the protein function. #Position of insertion in mgrB gene. ^Indicates Valine deletion at 444th position in phoQ. MLST, multilocus sequence typing; SIFT, sorting intolerant from tolerant; MIC, minimum inhibitory concentration; IS, insertion sequence
Fig. 2.
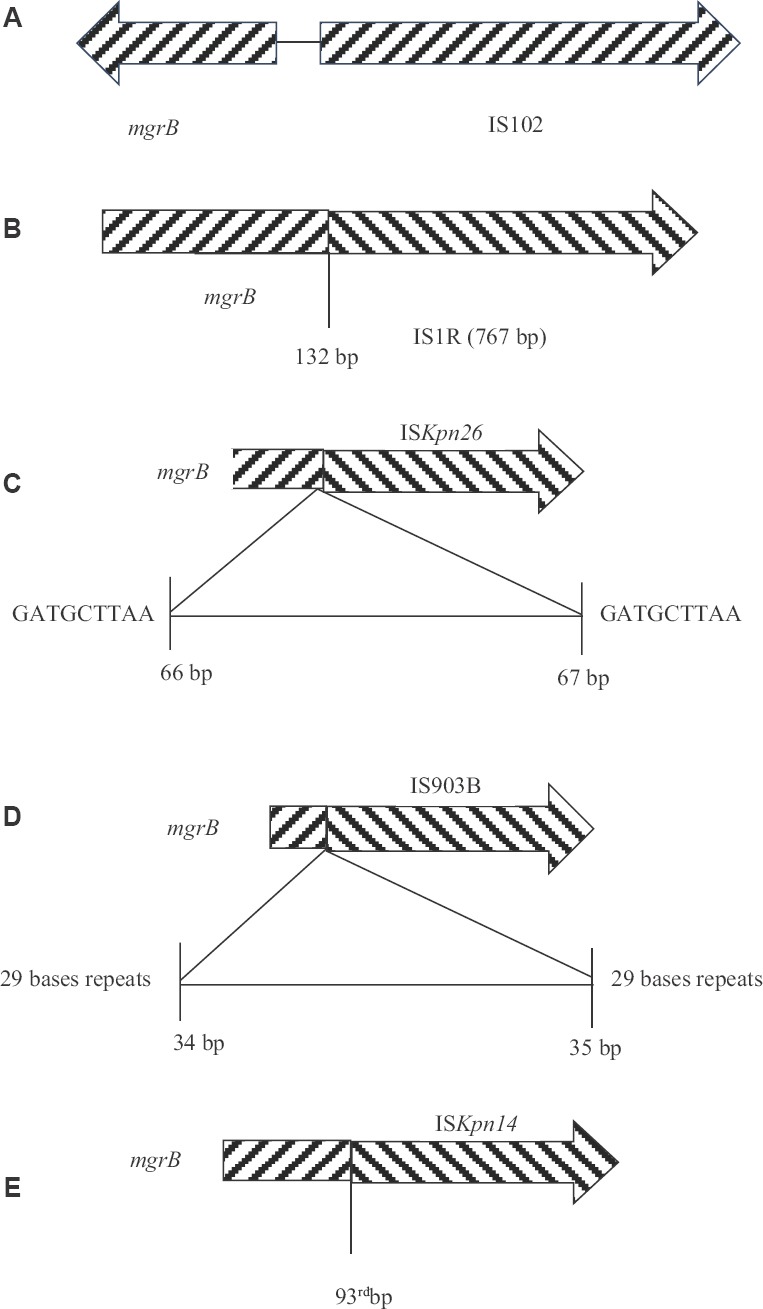
(A) Insertion sequence (IS) associated with mgrB in Kp2; (B) IS1R inserts at 132nd position in mgrB in Kp7; (C) Insertion of ISKpn26 between 66th and 67th bases of mgrB in Kp14 with repeats of 9 bases; (D) Insertion of IS903B between 34th and 35th bases of mgrB in Kp19 with repeats of 29 bases; (E) Insertion of ISKpn14 at 93rd position of mgrB in Kp20.
SIFT score was calculated to predict the effect of amino acid substitutions identified in this study isolates. In mgrB, both the substitutions Met1Ala and Cys28Gly were found deleterious. In phoP, among the three substitutions observed, Ala114Arg, Glu22Lys showed a score of 0.22 and 0.49, while Thr151Ala scored zero and found significant and deleterious. Similarly, in phoQ, mutations such as Leu205Gln, Leu209Cys, Trp215Ser, Val403Glu, Val446Trp and Phe478Gly were predicted to affect the function of the protein. Non-significant mutations of Trp194Leu (phoQ) in Kp7, Kp8 and Glu22Lys (phoP) in Kp13 were also noted.
Totally, 15 different sequence types were obtained among the 65 isolates and these included ST231 (n=23), ST14 (n=13), ST2096 (n=9), ST 147 (n=5), ST78 (n=3), ST15 (n=2), ST16 (n=2) and one isolate each of ST11, ST17, ST23, ST38, ST43, ST86, ST395 and ST2957. Fig. 3 shows the eBURST analysis for the clonal types among the study isolates. ST17 and ST16 belonged to the same clonal complex; while ST14, ST15, ST78 and ST2096 form another clonal complex, CC14. Other sequence types such as ST11, ST23, ST43, ST147 and ST231 were found as singletons.
Fig. 3.
eBURST of colistin-resistant Klebsiella pneumoniae. The clonal types identified in the present study are highlighted. ST17 and ST16 belonged to the same clonal complex; ST14, ST15, ST78 and ST2096 formed a clonal complex. Other sequence types such as ST11, ST23, ST38, ST43, ST86, ST147, ST231 and ST395 were found as singletons.
Among the study isolates, KP harbouring carbapenemases were of diverse sequence types. Overall, ST14 was the most predominant. Among the STs, OXA48-like producers were distributed among diverse clonal types such as ST14, ST16, ST17, ST38, ST43, ST231, ST395 and ST2096. Co-producers of blaNDM with blaOXA48 (n=6) belonged to ST14, ST78 and ST147. Isolates producing blaNDM (n=6) alone belonged to ST14, ST15, ST23 and ST78. Two of the study isolates were blaKPC producers, each belonging to ST15 and ST2957.
Fig. 4 shows the Splits Tree diagram for the sequence types. All the clonal types were closely related except ST2957 which was an outlier. The outlier was identified as K. quasipneumoniae belonging to phylogenetic group II of KP. Fig. 5 shows the maximum likelihood tree constructed based on the concatenated sequences of seven housekeeping genes used for MLST. As shown, isolates of ST231 were OXA48-like carbapenemase producers and was the predominant clone. The isolates of CC14 were more diverse in carbapenemases producing New Delhi metallo-β-lactamase (NDM) and/ or OXA48-like and one isolate with KPC. Fig. 5 also shows the clones present among different specimen sources. The isolates with chromosomal mutations contributing to colistin resistance (Table) are also indicated and the distribution of carbapenemases among these isolates is evident from Fig. 5.
Fig. 4.

Splits Tree for the clonal types identified among colistin-resistant Klebsiella pneumoniae. Fifteen clonal types identified in the study were closely related forming a cluster while ST2957 was distinct. The tree was constructed based on seven housekeeping genes.
Fig. 5.
Maximum likelihood tree for colistin-resistant Klebsiella pneumoniae using concatenated sequences of seven multilocus sequence typing genes.
Discussion
In this study molecular resistance mechanisms were assessed and clonal relatedness of colistin-resistant KP was investigated. SHV, TEM and CTX-M-15 were the prevalent β-lactamases similar to earlier study20. Among the study isolates, OXA48-like was the commonest carbapenemase produced. In this study, colistin resistance was contributed by mutations in mgrB, phoP and phoQ genes in the isolates. Our earlier study identified truncated mgrB of 27 amino acids due to a premature stop codon and also deletion of A at 10th nucleotide2. However, in the present study, these two mutations were absent, but various other mutations were seen. All the mgrB negatives could be due to partial or complete deletion of mgrB as reported previously21. The disruption of mgrB due to insertion elements such as IS1R, IS903, ISKpn14 and ISKpn26 similar to the present study have been reported earlier21,22,23. The mgrB mutations were not associated with specific carbapenemases or specific clones. There was no association between colistin MIC, mgrB mutations and specific clones. Substitutions in the phoP and phoQ genes implicated in colistin resistance have been reported23. However, certain mutations observed in the present study were novel. The significance of these novel mutations were predicted using SIFT scores. These scores revealed that the novel mutations were likely to affect protein function. However, detailed functional properties of mutations need to be determined by cloning studies to substantiate its significance.
Clonal relatedness with antimicrobial resistance profile in colistin-resistant KP was determined. New Delhi metallo-β-lactamase (NDM) producing KP are known to be associated with ST11, ST14, ST147 and ST231 in India and globally2,24,25 while the two KPC KP belong to ST15 and ST2957. International KPC clones such as ST258 and ST512 were absent among the study collection.
The colistin-resistant isolates in the present study were diverse and 15 clones were observed among the isolates. ST14, ST231 and ST2096 were predominant. The truncation of mgrB due to IS5-like elements21,22 and ISKpn1423,26 has been reported among ST258, ST512 and ST101. ISKpn26 has been reported among OXA48-like producer belonging to ST30726. In the present study, these were seen among isolates belonging to ST14, ST15 and ST231. Substitutions previously reported in phoP23,27 and phoQ23,27 were among isolates of ST147 and ST258.
Isolate belonging to ST23 produced NDM; also carried rmpA and rmpA2 genes indicating hypervirulent KP. Earlier, carbapenem-resistant hypervirulent KP belonging to ST14 and ST23128 has been reported. These two clones are classically associated with carbapenem-resistant isolates in India and other countries2,20. A novel ST2957 was observed in the present study. Earlier in India, Shankar et al29 have reported pan-susceptible hypervirulent K. quasipneumoniae of novel clone ST2320.
The association between the MLST sequence types and the molecular resistance mechanisms was not observed. This could be due to the limited relatedness information provided by MLST. Whole genome SNP-based phylogeny may provide further insights into the strain-specific lineage within a particular sequence type/clonal complex30. A previous study revealed that 97 per cent of KPC KP isolates sequenced, regardless of country of isolation, belong to a well-supported clade, corresponding to the complex CC258 based on the whole genome phylogeny30. MLST can be used to understand the epidemiology at a large scale while whole genome SNP phylogeny would be discriminatory in local outbreak situations.
The mechanism of colistin resistance in isolates which lacked mutations in mgrB, phoP and phoQ were not determined. This was a limitation of the study since in 70 per cent of study isolates the mechanism of colistin resistance was unidentified.
In conclusion, this study revealed the changing trend of carbapenem resistance mechanism predominantly to OXA48-like from NDM. There is alarming increase in colistin resistance with novel mutations in chromosomal genes such as mgrB, phoP and phoQ. There was no plasmid-mediated colistin resistance. Diverse clones of colistin-resistant isolates were observed with CG14 being predominant. No known international clones such as ST258 and ST512 were observed. It is important to monitor colistin resistance by using appropriate testing method and also limit colistin usage.
Footnotes
Financial support & sponsorship: Fluid Research Grant of the Christian Medical College, Vellore (IRB min. no. 9616 dated 01.09.2015) and the Indian Council of Medical Research, New Delhi (Ref. No: AMR/TF/55/13ECDII dated 23/10/2013) are acknowledged for financial support.
Conflicts of Interest: None.
References
- 1.Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18:268–81. doi: 10.1111/j.1469-0691.2011.03570.x. [DOI] [PubMed] [Google Scholar]
- 2.Pragasam AK, Shankar C, Veeraraghavan B, Biswas I, Nabarro LE, Inbanathan FY, et al. Molecular mechanisms of colistin resistance in Klebsiella pneumoniae causing bacteremia from India - a first report. Front Microbiol. 2016;7:2135. doi: 10.3389/fmicb.2016.02135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Koshi M. Vellore, India: Christian Medical College and Hospital; 2001. Myer's and Koshi's manual of diagnostic procedures in medical microbiology and immunology/serology. [Google Scholar]
- 4.CLSI Document M100S. 26th ed. Wayne, PA: CLSI; 2016. Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. [Google Scholar]
- 5.CLSI Document M100. 27th ed. Wayne, PA: CLSI; 2017. Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. [Google Scholar]
- 6.The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 6.0. 2016. [accessed on October 15, 2017]. Available from: http://www.eucast.org .
- 7.The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 7.0. 2017. [accessed on October 15, 2017]. Available from: http://www.eucast.org .
- 8.Dallenne C, Da Costa A, Decré D, Favier C, Arlet G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in enterobacteriaceae. J Antimicrob Chemother. 2010;65:490–5. doi: 10.1093/jac/dkp498. [DOI] [PubMed] [Google Scholar]
- 9.Woodford N, Ward ME, Kaufmann ME, Turton J, Fagan EJ, James D, et al. Community and hospital spread of Escherichia coli producing CTX-M extended-spectrum beta-lactamases in the UK. J Antimicrob Chemother. 2004;54:735–43. doi: 10.1093/jac/dkh424. [DOI] [PubMed] [Google Scholar]
- 10.Pérez-Pérez FJ, Hanson ND. Detection of plasmid-mediated AmpC beta-lactamase genes in clinical isolates by using multiplex PCR. J Clin Microbiol. 2002;40:2153–62. doi: 10.1128/JCM.40.6.2153-2162.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Poirel L, Walsh TR, Cuvillier V, Nordmann P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn Microbiol Infect Dis. 2011;70:119–23. doi: 10.1016/j.diagmicrobio.2010.12.002. [DOI] [PubMed] [Google Scholar]
- 12.Ellington MJ, Kistler J, Livermore DM, Woodford N. Multiplex PCR for rapid detection of genes encoding acquired metallo-beta-lactamases. J Antimicrob Chemother. 2007;59:321–2. doi: 10.1093/jac/dkl481. [DOI] [PubMed] [Google Scholar]
- 13.Yigit H, Queenan AM, Anderson GJ, Domenech-Sanchez A, Biddle JW, Steward CD, et al. Novel carbapenem-hydrolyzing beta-lactamase, KPC-1, from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob Agents Chemother. 2001;45:1151–61. doi: 10.1128/AAC.45.4.1151-1161.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jayol A, Nordmann P, Brink A, Poirel L. Heteroresistance to colistin in Klebsiella pneumoniae associated with alterations in the PhoPQ regulatory system. Antimicrob Agents Chemother. 2015;59:2780–4. doi: 10.1128/AAC.05055-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Liu YY, Wang Y, Walsh TR, Yi LX, Zhang R, Spencer J, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect Dis. 2016;16:161–8. doi: 10.1016/S1473-3099(15)00424-7. [DOI] [PubMed] [Google Scholar]
- 16.Yin W, Li H, Shen Y, Liu Z, Wang S, Shen Z, et al. Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. MBio. 2017;8:e00543–17. doi: 10.1128/mBio.00543-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Diancourt L, Passet V, Verhoef J, Grimont PA, Brisse S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J Clin Microbiol. 2005;43:4178–82. doi: 10.1128/JCM.43.8.4178-4182.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Feil EJ, Li BC, Aanensen DM, Hanage WP, Spratt BG. EBURST: Inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J Bacteriol. 2004;186:1518–30. doi: 10.1128/JB.186.5.1518-1530.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Huson DH, Bryant D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol. 2006;23:254–67. doi: 10.1093/molbev/msj030. [DOI] [PubMed] [Google Scholar]
- 20.Veeraraghavan B, Shankar C, Karunasree S, Kumari S, Ravi R, Ralph R. Carbapenem resistant Klebsiella pneumoniae isolated from bloodstream infection: Indian experience. Pathog Glob Health. 2017;111:240–6. doi: 10.1080/20477724.2017.1340128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cannatelli A, Giani T, D’Andrea MM, Di Pilato V, Arena F, Conte V, et al. MgrB inactivation is a common mechanism of colistin resistance in KPC-producing Klebsiella pneumoniae of clinical origin. Antimicrob Agents Chemother. 2014;58:5696–703. doi: 10.1128/AAC.03110-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Poirel L, Jayol A, Bontron S, Villegas MV, Ozdamar M, Türkoglu S, et al. The mgrB gene as a key target for acquired resistance to colistin in Klebsiella pneumoniae. J Antimicrob Chemother. 2015;70:75–80. doi: 10.1093/jac/dku323. [DOI] [PubMed] [Google Scholar]
- 23.Pitt ME, Elliott AG, Cao MD, Ganesamoorthy D, Karaiskos I, Giamarellou H, et al. Multifactorial chromosomal variants regulate polymyxin resistance in extensively drug-resistant Klebsiella pneumoniae. Microb Genom. 2018;4:1–35. doi: 10.1099/mgen.0.000158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Subramanian GK, Soundari PG, Ramanathan V, Krishnan P. Endemic Indian clones of Klebsiella pneumoniae-harbouring New Delhi metallo-beta-lactamase-1 on a hybrid plasmid replicon type: A case of changing New Delhi metallo-beta-lactamase plasmid landscapes in India? Indian J Med Microbiol. 2016;34:286–92. doi: 10.4103/0255-0857.188314. [DOI] [PubMed] [Google Scholar]
- 25.Giske CG, Fröding I, Hasan CM, Turlej-Rogacka A, Toleman M, Livermore D, et al. Diverse sequence types of Klebsiella pneumoniae contribute to the dissemination of blaNDM-1 in India, Sweden, and the United Kingdom. Antimicrob Agents Chemother. 2012;56:2735–8. doi: 10.1128/AAC.06142-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Novović K, Trudić A, Brkić S, Vasiljević Z, Kojić M, Medić D, et al. Molecular epidemiology of colistin-resistant, carbapenemase-producing Klebsiella pneumoniae in Serbia from 2013 to 2016. Antimicrob Agents Chemother. 2017;61 doi: 10.1128/AAC.02550-16. pii: e02550-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jaidane N, Bonnin RA, Mansour W, Girlich D, Creton E, Cotellon G, et al. Genomic insights into colistin-resistant Klebsiella pneumoniae from a Tunisian teaching hospital. Antimicrob Agents Chemother. 2018;62 doi: 10.1128/AAC.01601-17. pii: e01601-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shankar C, Nabarro LE, Devanga Ragupathi NK, Muthuirulandi Sethuvel DP, Daniel JL, Doss CGP, et al. Draft genome sequences of three hypervirulent carbapenem-resistant Klebsiella pneumoniae isolates from bacteremia. Genome Announc. 2016;4 doi: 10.1128/genomeA.01081-16. pii: e01081-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shankar C, Nabarro LEB, Muthuirulandi Sethuvel DP, Raj A, Devanga Ragupathi NK, Doss GP, et al. Draft genome of a hypervirulent Klebsiella quasipneumoniae subsp. similipneumoniae with novel sequence type ST2320 isolated from a chronic liver disease patient. J Glob Antimicrob Resist. 2017;9:30–1. doi: 10.1016/j.jgar.2017.01.004. [DOI] [PubMed] [Google Scholar]
- 30.Gaiarsa S, Comandatore F, Gaibani P, Corbella M, Dalla Valle C, Epis S, et al. Genomic epidemiology of Klebsiella pneumoniae in Italy and novel insights into the origin and global evolution of its resistance to carbapenem antibiotics. Antimicrob Agents Chemother. 2015;59:389–96. doi: 10.1128/AAC.04224-14. [DOI] [PMC free article] [PubMed] [Google Scholar]




